Simple Summary
This paper presents an extended version of the Cox regression model to examine heterogeneous effects of risk factors on disease subtypes defined by a continuous biomarker. This approach can be easily applied to cancer studies and is accessible to researchers via user-friendly R scripts.
Abstract
Molecular pathologic diagnosis is important in clinical (oncology) practice. Integration of molecular pathology into epidemiological methods (i.e., molecular pathological epidemiology) allows for investigating the distinct etiology of disease subtypes based on biomarker analyses, thereby contributing to precision medicine and prevention. However, existing approaches for investigating etiological heterogeneity deal with categorical subtypes. We aimed to fully leverage continuous measures available in most biomarker readouts (gene/protein expression levels, signaling pathway activation, immune cell counts, microbiome/microbial abundance in tumor microenvironment, etc.). We present a cause-specific Cox proportional hazards regression model for evaluating how the exposure–disease subtype association changes across continuous subtyping biomarker levels. Utilizing two longitudinal observational prospective cohort studies, we investigated how the association of alcohol intake (a risk factor) with colorectal cancer incidence differed across the continuous values of tumor epigenetic DNA methylation at long interspersed nucleotide element-1 (LINE-1). The heterogeneous alcohol effect was modeled using different functions of the LINE-1 marker to demonstrate the method’s flexibility. This real-world proof-of-principle computational application demonstrates how the new method enables visualizing the trend of the exposure effect over continuous marker levels. The utilization of continuous biomarker data without categorization for investigating etiological heterogeneity can advance our understanding of biological and pathogenic mechanisms.
1. Introduction
In clinical medicine, patients who share common symptoms and disease characteristics are grouped into a certain disease entity. However, molecular pathological diagnosis is a part of routine clinical practice, especially in oncology. Pathogenic mechanisms commonly vary between patients with the same disease entity. Therefore, when appropriate, patients with the disease are subclassified into groups (disease subtypes) based on their molecular pathological diagnosis to improve clinical management and treatment outcomes. Different disease subtypes are regarded as developing through distinct pathological mechanisms, on which risk factors may exert differential influence [1,2,3,4]. Therefore, the disease-subtyping framework and associated etiological heterogeneity have been widely applied in analyses of both neoplastic and non-neoplastic diseases [5,6,7]. For example, subtype heterogeneity has been identified when investigating the specific effects of a polygenic risk score and breastfeeding for breast cancer subtypes: basal-like and ERBB2 (HGNC ID: 3430; so-called HER2)-overexpressing breast cancer [8].
Despite continuous measurement readouts of many biomarkers used for disease subtyping, such continuous biomarker measures are commonly reduced to a small number of categorical levels (sometimes only two or three) to define disease subtypes, which can simplify the statistical analysis and generate readily interpretable data. Therefore, most existing statistical methods for studying etiological heterogeneity have focused on categorical disease subtype settings [9]. However, this categorization leads to reduction of information in the biomarker data, and is prone to bias due to arbitrary selection of cutoff values. For example, a weakness of categorical subtyping is evident when the exposure effect is limited to patients corresponding to extreme ends of the biomarker measures. In such situations, the patients associated with the exposure effect will likely be submerged among other patients not associated with the exposure effect. As a result, analysis using limited disease subtype categories may fail to discover existing exposure–disease associations. To maximize the value of disease subtyping biomarker information, this article presents an analytical framework for assessing the heterogeneity of exposure–disease subtype associations using continuous biomarker measures instead of categorical subtyping [10].
For illustration, we applied the proposed method to assess how the association of alcohol intake with colorectal cancer incidence changes across DNA methylation level at long interspersed nucleotide element-1 (LINE-1), measured in tumors. We used data from two prospective cohort studies, the Nurses’ Health Study (NHS) and Health Professionals Follow-up Study (HPFS).
2. Materials and Methods
To evaluate the association of an exposure with an incident disease in a cohort study, researchers typically use the Cox model [11], in which the hazard function is modeled as
where is the baseline hazard at time , is the possibly time-varying exposure for the i-th individual, the coefficient of , represents the exposure–outcome association, is a vector of potential confounders, which may also be time-varying, for the i-th individual, and is a vector of regression coefficients for . Without further specification, we assumed that the exposure is a scalar throughout this paper for notational simplicity.
Now, it is of interest to evaluate how the association of an exposure with the disease risk changes over the level of a disease marker. Extending Equation (1), we model the cause-specific hazards [12] of the disease subtypes by incorporating a function of the marker’s value as the coefficient of the exposure. Our model is
where is the continuous disease marker (cause), and are the baseline hazard and hazard functions for disease with marker level , and is a given real-valued function of with unknown parameters . The association between the exposure and the disease with marker level can be then represented by the hazard ratio . If the exposure is a -dimensional column vector, its coefficient will also be vector-valued with the form , where is the function of the disease marker corresponding to the coefficient of the k-th element of the exposure, and is a scalar or vector parameter of interest, .
The regression coefficients in the standard Cox model (1) are typically estimated by maximizing the partial likelihood [13]. Under the cause-specific proportional hazards model (2), we can construct the corresponding partial likelihood [14] as follows:
where is the set of all cases and is the time to event, which in a cohort study is typically age at disease diagnosis. Statistical software for the standard Cox model does not work here, as the marker level in is defined only among cases. In this partial likelihood, the subjects in a risk set are assigned the marker value of the case in that risk set so that the numerator and denominator in correspond to the hazard defined at the same marker level. The parameters and in Model (2) can be estimated through maximizing this partial likelihood. Similar to the standard Cox model setting, the variances of the parameter estimates can be estimated based on the inverse of the Hessian matrix.
We suggest using the restricted cubic spline approach [15] to characterize . The restricted cubic spline approach has advantages of parsimony while allowing for great flexibility in characterizing nonlinear curves. A restricted cubic spline function with knots includes one intercept, one linear, and nonlinear terms of Z; that is,
where is the j-th basis function of the restricted cubic spline, evaluated at Z. See Supplementary Material Section S1 for details. If , only includes the intercept and the linear term. The unknown parameter contains the intercept and all the coefficients of the linear and nonlinear terms. The number of knots can be determined using the Akaike information criterion (AIC) or the Bayesian information criterion (BIC) [16], and typically, the knots can be evenly spaced over the distribution of .
We used the likelihood ratio test to test for zero elements of . All elements of being zero implies no exposure–outcome association. Non-zero intercept and zero coefficients of all the linear and nonlinear terms imply an exposure–disease association that is independent of the disease marker. A non-zero coefficient of the linear term along with zero coefficients of all the nonlinear terms implies that the exposure–outcome association increases or decreases linearly over the marker level.
3. Results
3.1. Simulation Study
We conducted a simulation study to assess the finite sample performance of the method when . See Supplementary Material Section S2 for details. This simulation study shows that the point estimate of performs satisfactorily (Table S1 in the Supplementary Material Section S2). When the number of cases was 900, the percent bias of was 4 to 8% in five out of six configurations and 11% in the last configuration. It was 0.3 to 4% in five out of six configurations and 9.7% in the last configuration when the number of cases was increased to 4500. The empirical standard error of decreased by about 60% when the number of cases were increased from 900 to 4500.
3.2. Results of Illustrative Example
We used colorectal cancer (adenocarcinoma) and its subtyping biomarker, LINE-1 methylation (with continuous unitless values) [17], as a disease biomarker example to illustrate the method. We utilized data from ongoing large prospective cohort studies, namely the Nurses’ Health Study (NHS) [18,19] and Health Professionals Follow-up Study (HPFS) [20,21]. The main exposure was cumulative average alcohol intake (0, ≤15, >15 g/day). Detailed descriptions of the study population, assessment of main exposure and covariates, ascertainment of colorectal cancer cases, and quantification of LINE-1 levels are described in Supplementary Material Section S3. The age-standardized characteristics of participants in the two cohorts are summarized in Table S2 (Supplementary Material).
Shown in Figure 1 and Figure S1 (Supplementary Material) are the curves of the hazard ratios (HRs) representing the association between alcohol intake and incidence of colorectal cancer subtype as a function of continuous LINE-1 methylation level. These curves were constructed by plotting over the LINE-1 marker values () within the plausible range (25 to 85). The number of knots considered were . The knots were evenly spaced over the LINE-1 distribution. Figure 1 and Figure S1 were drawn based on the results using the combined cohort, HPFS alone, and NHS alone. We considered two models: the main model, with stratification factors only, and the full model, which adjusted for additional covariates as described in the Methods section. Since the inclusion of additional covariates in the full model had little impact on the set of estimated coefficients , we simply utilized the estimation results from the main model hereafter.
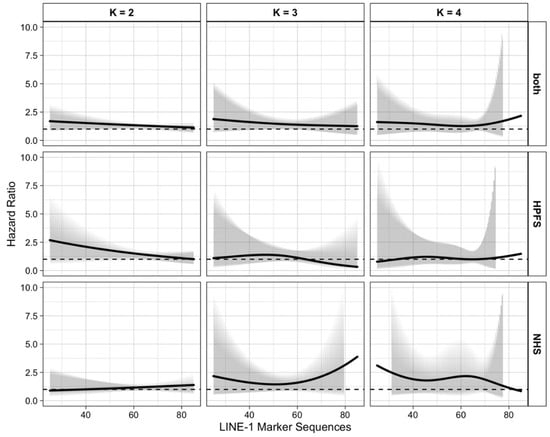
Figure 1.
Heterogeneous Effect of Cumulative Categorical Alcohol Intake (>15 g/day vs. 0 g/day) on continuous subtypes of colorectal cancer; the 3 × 3 plot panel illustrates the combination of three choices of the knot number in and three cohort settings. Abbreviations: HPFS, Health Professionals Follow-up Study; LINE-1, long interspersed nucleotide element-1; NHS, Nurses’ Health Study.
Table 1 and Table S3 (Supplementary Material) present p-values from testing the following null hypotheses for the same choices of knot numbers and cohort settings as in Figure 1 and Figure S1: (i) the intercept and all the coefficients in are zero (the overall test); (ii) all the coefficients in except the intercept are zero (test for heterogeneity); (iii) all the coefficients of the nonlinear terms in are zero (test for nonlinearity). For the NHS cohort and the combined cohort, the linear model () had the smallest BIC and AIC, and for the HPFS cohort, the linear model had the smallest BIC and the model with had the smallest AIC. For the comparison between >15 g/day intake and 0 g/day based on the models with , as shown in Table 1, there were significant associations between alcohol and cancer risk in the HPFS cohort (overall test p < 0.001) and the combined cohort (overall test p < 0.001), but there was insufficient statistical evidence to establish such an association in the NHS cohort. There was insufficient statistical evidence to establish a difference in the comparison of ≤15 g/day intake versus 0 g/day in the NHS, HPFS, or the combined cohort (Table S3). Furthermore, in the comparison of >15 g/day versus 0 g/day in the combined cohort, the heterogeneity tests were statistically significant (p < 0.001) under and the alcohol effect changed with the LINE-1 level linearly (nonlinear test p = 0.54 for ).

Table 1.
Model testing for the association of categorical alcohol intake (>15 g/day vs. 0 g/day) with colorectal cancer incidence, based on the main model for three functional forms and three cohorts.
Table 2 and Table S4 (Supplementary Material) display the estimated HRs, with 95% pointwise confidence intervals, representing the alcohol–cancer association for some plausible LINE-1 values (30 to 80 in steps of 10) for the choices of knot numbers and data settings considered in Figure 1. As shown for the HPFS and combined cohorts in Figure 1 and Table 2, the alcohol–cancer association (for >15 g/day vs. 0 g/day) tended to decrease with increasing LINE-1 methylation level, as seen from the two functions with as selected by AIC and BIC.

Table 2.
Hazard ratio for categorical alcohol intake (>15 g/day vs. 0 g/day) modeled using three functional forms for the LINE-1 marker value in three cohort settings, based on the main model.
4. Discussion
In this paper, we have presented a Cox proportional hazards regression model method to fully utilize a continuous biomarker measure for disease subtyping. This statistical method can examine subtype heterogeneity of diseases in the exposure–disease association with more comprehensive and versatile utilization of continuous marker measurements. The ability of this method to potentially reveal more complicated patterns in subtype heterogeneity can help us gain deeper insights into etiologies in molecular epidemiological research and provide further evidence in the development of personalized precision medicine.
Statistical methods for investigating disease subtype heterogeneity for categorical and ordinal subtypes have been studied previously under several common study designs [9]. However, a concern may be raised about defining discrete subtypes based on categorization of biomarker values when there is little or no evidence supporting biomarker cut-point values that are often arbitrarily determined. In addition, the categorization of a continuous measure of a biomarker can lead to loss of information from the biological and statistical perspectives. The proposed method is less prone to these problems and has the potential to reveal more detailed and granular subtype heterogeneity than established approaches using categorical and ordinal subtypes.
Many biological phenomena and related biomarkers (including expressions of genes and proteins) are continuous in nature [6]. LINE-1 methylation level (i.e., the percentage of the amount of C nucleotides divided by the sum of the amounts of C and T nucleotides at CpG sites), which we used in the illustrative example, is a surrogate marker for genome-wide DNA methylation and widely distributed in colorectal cancer tissue from 20 to 90% [22,23]. Currently, it remains unclear how to set the best cut-points for defining subtypes based on quantitative LINE-1 methylation levels. Accordingly, the proposed method can be applied to this biomarker without using arbitrarily cut-points. Another example for continuous tissue biomarkers is immune cell infiltrates in tumor tissue. Ample evidence supports the biological importance of the immune system in cancer [24,25,26,27]. Tumors exhibit considerably heterogeneous phenotypes according to types and quantities of immune cell infiltrates in tumor tissue [28,29], and higher immune cell infiltrates in cancer have often been associated with better cancer survival [26,30,31,32]. Related to immune cells, microbial species are often quantitatively measured in biospecimens including tumor and normal tissue in population studies [33,34]. Readouts of quantitative microbial assays are continuous in nature without prior knowledge on any biological cut-points (or threshold effect). Categorizations of such variables are often used [35,36,37,38]. However, simple categorizations may lose biological information. It is evident that standardized definitions of tumor subtypes based on immune cell infiltrates or tissue microbiota have not been developed. There is a clear need to analyze tumor biomarker data in a way that exploits the underlying continuous nature of the biomarker.
The real-world application of this method in the two large prospective cohort studies has demonstrated its capability to depict the trend of the exposure effect across continuous molecular marker levels in contrast to use of solely categorical subtypes [10]. Further, this method allows for the flexible modeling of the heterogeneous effect of exposure on the disease of interest across biomarker levels, using models ranging from linear functions, to functions of any hypothesized form, to a case-by-case understanding of the disease.
A user-friendly R program that implements this method is publicly available (https://www.hsph.harvard.edu/molin-wang/software/, accessed on 31 March 2022). This R function fits a Cox regression model for either incidence analysis or post-diagnosis survival analysis, where the model can include one or more exposure variables, a set of confounders (optional), and one or more stratification variables (optional). Left truncation and time-varying covariates, which are common in cohort data analyses, can be handled by putting the data in counting process form [39] before applying our R function. In the counting process data structure, a new data record is created for each questionnaire cycle at which a participant was at risk, with covariates set to their values at the time the questionnaire was returned. Furthermore, in addition to AIC and BIC, the cross validation approach [40] could also be used to choose the number of knots in the restricted cubic spline approach. The proposed method can be easily applied to studies of various diseases and risk factors and is accessible to researchers with limited experience with time-to-event data analysis.
In this article, we follow the nomenclature guideline for gene products using the Human Genome Organization (HUGO) Gene Nomenclature Committee (HGNC) standards, recommended by the expert panel [41].
5. Conclusions
To summarize, we have presented a Cox proportional hazards regression model for analyzing heterogeneous exposure–disease associations across disease subtypes defined by continuous biomarker measures. This method is helpful in decreasing bias caused by arbitrary subtype categorization and in increasing statistical power, as well as flexibility of assumptions about the pattern of pathologic heterogeneity. The utilization of continuous marker data without categorization for investigating subtype heterogeneity will advance our understanding of etiological heterogeneity and possibly contribute to precision medicine.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/cancers14071811/s1, Table S1: Simulation Results for K = 3; 10% censoring rate; 1000 simulation replicates, Table S2: Age-Standardized Characteristics for Study Participants in the NHS (1980–2012) and the HPFS (1986–2012), Table S3: Model Testing for the Association of Alcohol Intake (≤15 g/day Versus 0 g/day) with Colorectal Cancer Incidence, Based on Main Model in Three Functions and Three Settings, Table S4: Hazard Ratio for Alcohol Intake (≤15 g/day Versus 0 g/day) Modeled as Three Functions of LINE-1 Marker Value in Three Cohort Settings, Based on the Main Model. Figure S1: Heterogeneous Effect of Cumulative Categorical Alcohol Intake (≤15 g/day Versus <0 g/day) on Continuous Subtypes of Colorectal Cancer; the 3 × 3 plot panel illustrates the combination of three choices of the knot number in g(ϕ,Z) and three cohort settings.
Author Contributions
Conceptualization, M.W.; methodology, M.W. and R.L.; formal analysis, R.L. and L.X.; investigation, R.L and T.U.; resources, S.O.; data curation, R.L.; writing—original draft preparation, R.L. and T.U.; writing—review and editing, all authors.; supervision, D.Z., S.O. and M.W.; project administration, M.W.; funding acquisition, T.U., S.O. and M.W. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by U.S. National Institutes of Health (NIH) grants [R35 CA197735 to S.O., M.W.; R01 CA151993 to S.O., R01 CA248857 to S.O., U01 CA167552, UM1 CA167552, P01 CA055075, UM1 CA186107, P01 CA087969]; by Cancer Research UK Grand Challenge Award [UK C10674/A27140 to S.O.]. T.U. was supported by a grant from Overseas Research Fellowship from Japan Society for the Promotion of Science [201960541] and Prevent Caner Foundation.
Institutional Review Board Statement
The study was conducted according to the guidelines of the Declaration of Helsinki and approved by the Institutional Review Board of Brigham and Women’s Hospital and Harvard T.H. Chan School of Public Health (2001P001945) on 20 October 2020.
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
The datasets generated and/or analyzed during the current study are not publicly available. Further information including the procedures to obtain and access data from the Nurses’ Health Studies and the Health Professionals Follow-up Study is described at https://www.nurseshealthstudy.org/researchers/, accessed on 1 February 2022 and https://sites.sph.harvard.edu/hpfs/for-collaborators/, accessed on 1 February 2022.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| AIC | Akaike information criterion |
| BIC | Bayesian information criterion |
| HPFS | Health Professionals Follow-up Study |
| HR | hazard ratio |
| LINE-1 | long interspersed nucleotide element-1 |
| NHS | Nurses’ Health Study |
References
- Begg, C.B. A strategy for distinguishing optimal cancer subtypes. Int. J. Cancer 2011, 129, 931–937. [Google Scholar] [CrossRef] [PubMed]
- Begg, C.B.; Zabor, E.C. Detecting and exploiting etiologic heterogeneity in epidemiologic studies. Am. J. Epidemiol. 2012, 176, 512–518. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Begg, C.B.; Zabor, E.C.; Bernstein, J.L.; Bernstein, L.; Press, M.F.; Seshan, V.E. A conceptual and methodological framework for investigating etiologic heterogeneity. Stat. Med. 2013, 32, 5039–5052. [Google Scholar] [CrossRef] [PubMed]
- Richiardi, L.; Barone-Adesi, F.; Pearce, N. Cancer subtypes in aetiological research. Eur. J. Epidemiol. 2017, 32, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Ogino, S.; Chan, A.T.; Fuchs, C.S.; Giovannucci, E. Molecular pathological epidemiology of colorectal neoplasia: An emerging transdisciplinary and interdisciplinary field. Gut 2011, 60, 397–411. [Google Scholar] [CrossRef]
- Ogino, S.; Nishihara, R.; VanderWeele, T.J.; Wang, M.; Nishi, A.; Lochhead, P.; Qian, Z.R.; Zhang, X.; Wu, K.; Nan, H. The role of molecular pathological epidemiology in the study of neoplastic and non-neoplastic diseases in the era of precision medicine. Epidemiology 2016, 27, 602. [Google Scholar] [CrossRef]
- Ogino, S.; Nowak, J.A.; Hamada, T.; Milner, D.A., Jr.; Nishihara, R. Insights into pathogenic interactions among environment, host, and tumor at the crossroads of molecular pathology and epidemiology. Annu. Rev. Pathol. Mech. Dis. 2019, 14, 83–103. [Google Scholar] [CrossRef]
- Holm, J.; Eriksson, L.; Ploner, A.; Eriksson, M.; Rantalainen, M.; Li, J.; Hall, P.; Czene, K. Assessment of breast cancer risk factors reveals subtype heterogeneity. Cancer Res. 2017, 77, 3708–3717. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Spiegelman, D.; Kuchiba, A.; Lochhead, P.; Kim, S.; Chan, A.T.; Poole, E.M.; Tamimi, R.; Tworoger, S.S.; Giovannucci, E. Statistical methods for studying disease subtype heterogeneity. Stat. Med. 2016, 35, 782–800. [Google Scholar] [CrossRef]
- Schernhammer, E.S.; Giovannucci, E.; Kawasaki, T.; Rosner, B.; Fuchs, C.S.; Ogino, S. Dietary folate, alcohol and B vitamins in relation to LINE-1 hypomethylation in colon cancer. Gut 2010, 59, 794–799. [Google Scholar] [CrossRef] [Green Version]
- Cox, D.R. Regression models and life-tables. J. R. Stat. Soc. Ser. B 1972, 34, 187–202. [Google Scholar] [CrossRef]
- Prentice, R.L.; Kalbfleisch, J.D.; Peterson, A.V., Jr.; Flournoy, N.; Farewell, V.T.; Breslow, N.E. The analysis of failure times in the presence of competing risks. Biometrics 1978, 34, 541–554. [Google Scholar] [CrossRef] [PubMed]
- Cox, D.R. Partial likelihood. Biometrika 1975, 62, 269–276. [Google Scholar] [CrossRef]
- Chatterjee, N.; Sinha, S.; Diver, W.R.; Feigelson, H.S. Analysis of cohort studies with multivariate and partially observed disease classification data. Biometrika 2010, 97, 683–698. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Durrleman, S.; Simon, R. Flexible regression models with cubic splines. Stat. Med. 1989, 8, 551–561. [Google Scholar] [CrossRef] [PubMed]
- Burnham, K.P.; Anderson, D.R. Multimodel inference: Understanding AIC and BIC in model selection. Sociol. Methods Res. 2004, 33, 261–304. [Google Scholar] [CrossRef]
- Irahara, N.; Nosho, K.; Baba, Y.; Shima, K.; Lindeman, N.I.; Hazra, A.; Schernhammer, E.S.; Hunter, D.J.; Fuchs, C.S.; Ogino, S. Precision of pyrosequencing assay to measure LINE-1 methylation in colon cancer, normal colonic mucosa, and peripheral blood cells. J. Mol. Diagn. 2010, 12, 177–183. [Google Scholar] [CrossRef]
- Bao, Y.; Bertoia, M.L.; Lenart, E.B.; Stampfer, M.J.; Willett, W.C.; Speizer, F.E.; Chavarro, J.E. Origin, Methods, and Evolution of the Three Nurses’ Health Studies. Am. J. Public Health 2016, 106, 1573–1581. [Google Scholar] [CrossRef]
- Ugai, T.; Vayrynen, J.P.; Haruki, K.; Akimoto, N.; Lau, M.C.; Zhong, R.; Kishikawa, J.; Vayrynen, S.A.; Zhao, M.; Fujiyoshi, K.; et al. Smoking and Incidence of Colorectal Cancer Subclassified by Tumor-Associated Macrophage Infiltrates. J. Natl. Cancer Inst. 2022, 114, 68–77. [Google Scholar] [CrossRef]
- Nishihara, R.; Wu, K.; Lochhead, P.; Morikawa, T.; Liao, X.; Qian, Z.R.; Inamura, K.; Kim, S.A.; Kuchiba, A.; Yamauchi, M.; et al. Long-term colorectal-cancer incidence and mortality after lower endoscopy. N. Engl. J. Med. 2013, 369, 1095–1105. [Google Scholar] [CrossRef] [Green Version]
- Ugai, T.; Haruki, K.; Vayrynen, J.P.; Borowsky, J.; Fujiyoshi, K.; Lau, M.C.; Akimoto, N.; Zhong, R.; Kishikawa, J.; Arima, K.; et al. Coffee Intake of Colorectal Cancer Patients and Prognosis According to Histopathologic Lymphocytic Reaction and T-Cell Infiltrates. Mayo Clin. Proc. 2022, 97, 124–133. [Google Scholar] [CrossRef] [PubMed]
- Baba, Y.; Huttenhower, C.; Nosho, K.; Tanaka, N.; Shima, K.; Hazra, A.; Schernhammer, E.S.; Hunter, D.J.; Giovannucci, E.L.; Fuchs, C.S.; et al. Epigenomic diversity of colorectal cancer indicated by LINE-1 methylation in a database of 869 tumors. Mol. Cancer 2010, 9, 125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Estecio, M.R.; Gharibyan, V.; Shen, L.; Ibrahim, A.E.; Doshi, K.; He, R.; Jelinek, J.; Yang, A.S.; Yan, P.S.; Huang, T.H.; et al. LINE-1 hypomethylation in cancer is highly variable and inversely correlated with microsatellite instability. PLoS ONE 2007, 2, e399. [Google Scholar] [CrossRef] [PubMed]
- Havel, J.J.; Chowell, D.; Chan, T.A. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat. Rev. Cancer 2019, 19, 133–150. [Google Scholar] [CrossRef] [PubMed]
- Paucek, R.D.; Baltimore, D.; Li, G. The Cellular Immunotherapy Revolution: Arming the Immune System for Precision Therapy. Trends Immunol. 2019, 40, 292–309. [Google Scholar] [CrossRef]
- Grizzi, F.; Basso, G.; Borroni, E.M.; Cavalleri, T.; Bianchi, P.; Stifter, S.; Chiriva-Internati, M.; Malesci, A.; Laghi, L. Evolving notions on immune response in colorectal cancer and their implications for biomarker development. Inflamm. Res. 2018, 67, 375–389. [Google Scholar] [CrossRef]
- Kather, J.N.; Halama, N. Harnessing the innate immune system and local immunological microenvironment to treat colorectal cancer. Br. J. Cancer 2019, 120, 871–882. [Google Scholar] [CrossRef] [Green Version]
- Ogino, S.; Giannakis, M. Immunoscore for (colorectal) cancer precision medicine. Lancet 2018, 391, 2084–2086. [Google Scholar] [CrossRef]
- Ogino, S.; Nowak, J.A.; Hamada, T.; Phipps, A.I.; Peters, U.; Milner, D.A., Jr.; Giovannucci, E.L.; Nishihara, R.; Giannakis, M.; Garrett, W.S.; et al. Integrative analysis of exogenous, endogenous, tumour and immune factors for precision medicine. Gut 2018, 67, 1168–1180. [Google Scholar] [CrossRef]
- Le, D.T.; Hubbard-Lucey, V.M.; Morse, M.A.; Heery, C.R.; Dwyer, A.; Marsilje, T.H.; Brodsky, A.N.; Chan, E.; Deming, D.A.; Diaz, L.A., Jr.; et al. A Blueprint to Advance Colorectal Cancer Immunotherapies. Cancer Immunol. Res. 2017, 5, 942–949. [Google Scholar] [CrossRef] [Green Version]
- Kather, J.N.; Halama, N.; Jaeger, D. Genomics and emerging biomarkers for immunotherapy of colorectal cancer. Semin. Cancer Biol. 2018, 52, 189–197. [Google Scholar] [CrossRef] [PubMed]
- Pages, F.; Galon, J.; Dieu-Nosjean, M.C.; Tartour, E.; Sautes-Fridman, C.; Fridman, W.H. Immune infiltration in human tumors: A prognostic factor that should not be ignored. Oncogene 2010, 29, 1093–1102. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hamada, T.; Nowak, J.A.; Milner, D.A., Jr.; Song, M.; Ogino, S. Integration of microbiology, molecular pathology, and epidemiology: A new paradigm to explore the pathogenesis of microbiome-driven neoplasms. J. Pathol. 2019, 247, 615–628. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mima, K.; Kosumi, K.; Baba, Y.; Hamada, T.; Baba, H.; Ogino, S. The microbiome, genetics, and gastrointestinal neoplasms: The evolving field of molecular pathological epidemiology to analyze the tumor-immune-microbiome interaction. Hum. Genet. 2021, 140, 725–746. [Google Scholar] [CrossRef]
- Mima, K.; Nishihara, R.; Qian, Z.R.; Cao, Y.; Sukawa, Y.; Nowak, J.A.; Yang, J.; Dou, R.; Masugi, Y.; Song, M.; et al. Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut 2016, 65, 1973–1980. [Google Scholar] [CrossRef] [Green Version]
- Mima, K.; Cao, Y.; Chan, A.T.; Qian, Z.R.; Nowak, J.A.; Masugi, Y.; Shi, Y.; Song, M.; da Silva, A.; Gu, M.; et al. Fusobacterium nucleatum in Colorectal Carcinoma Tissue According to Tumor Location. Clin. Transl. Gastroenterol. 2016, 7, e200. [Google Scholar] [CrossRef]
- Mehta, R.S.; Nishihara, R.; Cao, Y.; Song, M.; Mima, K.; Qian, Z.R.; Nowak, J.A.; Kosumi, K.; Hamada, T.; Masugi, Y.; et al. Association of Dietary Patterns With Risk of Colorectal Cancer Subtypes Classified by Fusobacterium nucleatum in Tumor Tissue. JAMA Oncol. 2017, 3, 921–927. [Google Scholar] [CrossRef] [Green Version]
- Borowsky, J.; Haruki, K.; Lau, M.C.; Dias Costa, A.; Vayrynen, J.P.; Ugai, T.; Arima, K.; da Silva, A.; Felt, K.D.; Zhao, M.; et al. Association of Fusobacterium nucleatum with Specific T-cell Subsets in the Colorectal Carcinoma Microenvironment. Clin. Cancer Res. 2021, 27, 2816–2826. [Google Scholar] [CrossRef]
- Lin, D.; Fleming, T.R. Proceedings of the First Seattle Symposium in Biostatistics: Survival Analysis: Survival Analysis; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2012; Volume 123. [Google Scholar]
- Verweij, P.J.; Van Houwelingen, H.C. Cross-validation in survival analysis. Stat. Med. 1993, 12, 2305–2314. [Google Scholar] [CrossRef]
- Fujiyoshi, K.; Bruford, E.A.; Mroz, P.; Sims, C.L.; O’Leary, T.J.; Lo, A.W.I.; Chen, N.; Patel, N.R.; Patel, K.P.; Seliger, B.; et al. Opinion: Standardizing gene product nomenclature—A call to action. Proc. Natl. Acad. Sci. USA 2021, 118, e2025207118. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).