Machine Learning and Deep Learning Applications in Multiple Myeloma Diagnosis, Prognosis, and Treatment Selection
Abstract
Simple Summary
Abstract
1. Introduction
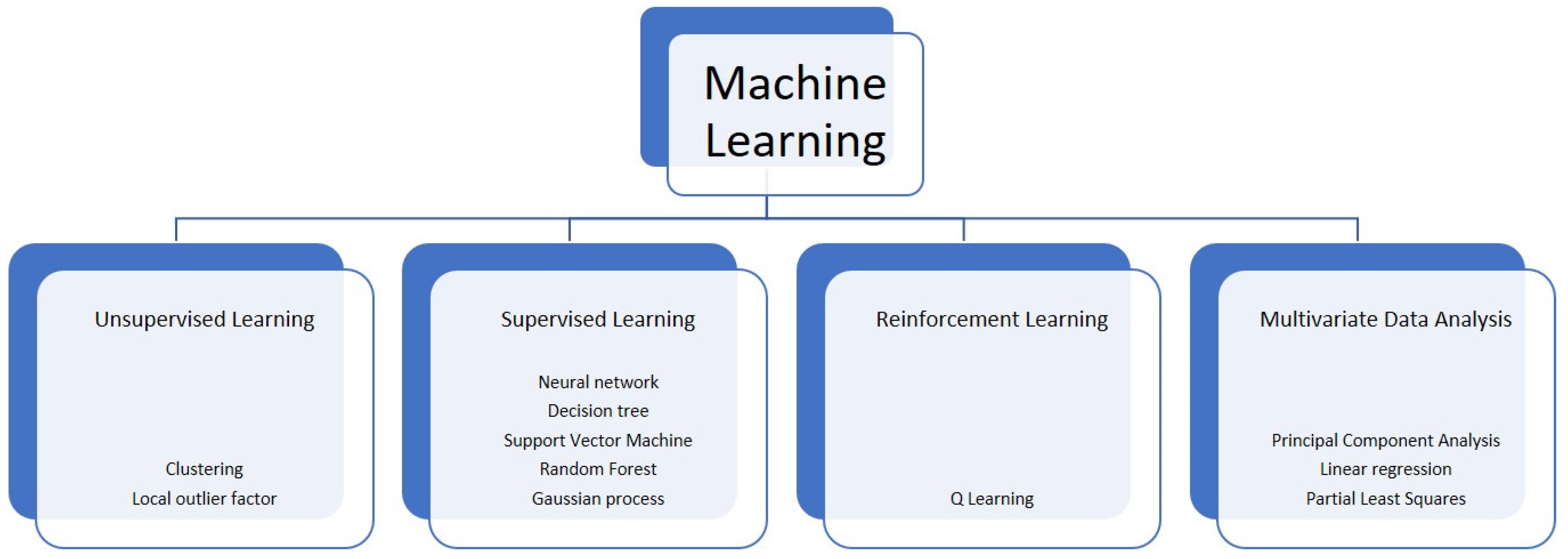
General Considerations on Machine Learning and Deep Learning
2. Machine Learning and Multiple Myeloma Diagnosis
2.1. Machine Learning and Bone Lesions Identification in Multiple Myeloma Patients
2.2. Machine Learning and Multiple Myeloma Prognosis
2.3. Machine Learning and Prediction of Clinical Drug Response
2.4. Machine Learning and Apoptosis in Multiple Myeloma
3. Conclusions, Challenges, and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Yu, K.-H.; Beam, A.L.; Kohane, I.S. Artificial intelligence in healthcare. Nat. Biomed. Eng. 2018, 2, 719–731. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Jiang, Y.; Zhi, H.; Dong, Y.; Li, H.; Ma, S.; Wang, Y.; Dong, Q.; Shen, H.; Wang, Y. Artificial intelligence in healthcare: Past, present and future. Stroke Vasc. Neurol. 2017, 2, 230–243. [Google Scholar] [CrossRef] [PubMed]
- Reddy, S.; Fox, J.; Purohit, M.P. Artificial intelligence-enabled healthcare delivery. J. R. Soc. Med. 2018, 112, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Decary, M. Artificial intelligence in healthcare: An essential guide for health leaders. Healthc. Manag. Forum 2019, 33, 10–18. [Google Scholar] [CrossRef]
- Lynch, C.J.; Liston, C. New machine-learning technologies for computer-aided diagnosis. Nat. Med. 2018, 24, 1304–1305. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.; Malhi, A.K. Using deep learning to enhance head and neck cancer diagnosis and classification. In Proceedings of the 2018 IEEE international Conference on System, Computation, Automation and Networking (Icscan), Pondicherry, India, 6–7 July 2018; pp. 1–6. [Google Scholar] [CrossRef]
- Chen, Y.; Li, Y.; Narayan, R.; Subramanian, A.; Xie, X. Gene expression inference with deep learning. Bioinformatics 2016, 32, 1832–1839. [Google Scholar] [CrossRef]
- Koh, P.W.; Pierson, E.; Kundaje, A. Denoising genome-wide histone ChIP-seq with convolutional neural networks. Bioinformatics 2017, 33, i225–i233. [Google Scholar] [CrossRef]
- Baptista, D.; Ferreira, P.; Rocha, M. Deep learning for drug response prediction in cancer. Brief. Bioinform. 2020, 22, 360–379. [Google Scholar] [CrossRef]
- Russell, S.J.; Norvig, P. Artificial Intelligence: A Modern Approach; Pearson: Boston, MA, USA, 2020; ISBN 978-0-13-461099-3. [Google Scholar]
- English, M.; Kumar, C.; Ditterline, B.L.; Drazin, D.; Dietz, N. Machine Learning in Neuro-Oncology, Epilepsy, Alzheimer’s Disease, and Schizophrenia. Acta Neurochir. Suppl. 2022, 134, 349–361. [Google Scholar] [CrossRef]
- Eckardt, J.-N.; Wendt, K.; Bornhäuser, M.; Middeke, J.M. Reinforcement Learning for Precision Oncology. Cancers 2021, 13, 4624. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Tran, K.A.; Kondrashova, O.; Bradley, A.; Williams, E.D.; Pearson, J.V.; Waddell, N. Deep learning in cancer diagnosis, prognosis and treatment selection. Genome Med. 2021, 13, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Montesinos-López, O.A.; Montesinos-López, A.; Pérez-Rodríguez, P.; Barrón-López, J.A.; Martini, J.W.R.; Fajardo-Flores, S.B.; Gaytan-Lugo, L.S.; Santana-Mancilla, P.C.; Crossa, J. A review of deep learning applications for genomic selection. BMC Genom. 2021, 22, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Bin Tufail, A.; Ma, Y.-K.; Kaabar, M.K.A.; Martínez, F.; Junejo, A.R.; Ullah, I.; Khan, R. Deep Learning in Cancer Diagnosis and Prognosis Prediction: A Minireview on Challenges, Recent Trends, and Future Directions. Comput. Math. Methods Med. 2021, 2021, 1–28. [Google Scholar] [CrossRef]
- Libbrecht, M.; Noble, W.S. Machine learning applications in genetics and genomics. Nat. Rev. Genet. 2015, 16, 321–332. [Google Scholar] [CrossRef]
- Jones, W.; Alasoo, K.; Fishman, D.; Parts, L. Computational biology: Deep learning. Emerg. Top. Life Sci. 2017, 1, 257–274. [Google Scholar] [CrossRef]
- Wainberg, M.; Merico, D.; Delong, A.; Frey, B.J. Deep learning in biomedicine. Nat. Biotechnol. 2018, 36, 829–838. [Google Scholar] [CrossRef]
- Zou, J.; Huss, M.; Abid, A.; Mohammadi, P.; Torkamani, A.; Telenti, A. A primer on deep learning in genomics. Nat. Genet. 2018, 51, 12–18. [Google Scholar] [CrossRef]
- Walsh, S.; de Jong, E.E.; van Timmeren, J.E.; Ibrahim, A.; Compter, I.; Peerlings, J.; Sanduleanu, S.; Refaee, T.; Keek, S.; Larue, R.T.; et al. Decision Support Systems in Oncology. JCO Clin. Cancer Inform. 2019, 3, 1–9. [Google Scholar] [CrossRef]
- Hemminki, K.; Försti, A.; Houlston, R.; Sud, A. Epidemiology, genetics and treatment of multiple myeloma and precursor diseases. Int. J. Cancer 2021, 149, 1980–1996. [Google Scholar] [CrossRef]
- Blimark, C.H.; Vangsted, A.J.; Klausen, T.W.; Gregersen, H.; Szabo, A.G.; Hermansen, E.; Wålinder, G.; Knut-Bojanowska, D.; Zang, C.; Turesson, I.; et al. Outcome data from >10 000 multiple myeloma patients in the Danish and Swedish national registries. Eur. J. Haematol. 2021, 108, 99–108. [Google Scholar] [CrossRef]
- Yan, W.; Shi, H.; He, T.; Chen, J.; Wang, C.; Liao, A.; Yang, W.; Wang, H. Employment of Artificial Intelligence Based on Routine Laboratory Results for the Early Diagnosis of Multiple Myeloma. Front. Oncol. 2021, 11, 933. [Google Scholar] [CrossRef] [PubMed]
- Chandradevan, R.; Aljudi, A.A.; Drumheller, B.R.; Kunananthaseelan, N.; Amgad, M.; Gutman, D.A.; Cooper, L.A.D.; Jaye, D.L. Machine-based detection and classification for bone marrow aspirate differential counts: Initial development focusing on nonneoplastic cells. Lab. Investig. 2020, 100, 98–109. [Google Scholar] [CrossRef] [PubMed]
- Aghaeepour, N.; Finak, G.; The FlowCAP Consortium; The DREAM Consortium; Hoos, H.; Mosmann, T.R.; Brinkman, R.; Gottardo, R.; Scheuermann, R.H. Critical assessment of automated flow cytometry data analysis techniques. Nat. Methods 2013, 10, 228–238. [Google Scholar] [CrossRef] [PubMed]
- Clichet, V.; Harrivel, V.; Delette, C.; Guiheneuf, E.; Gautier, M.; Morel, P.; Assouan, D.; Merlusca, L.; Beaumont, M.; Lebon, D.; et al. Accurate classification of plasma cell dyscrasias is achieved by combining artificial intelligence and flow cytometry. Br. J. Haematol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, X.; Yu, X.; Chen, D.; Liu, A. Diagnosis of human malignancies using laser-induced breakdown spectroscopy in combination with chemometric methods. Spectrochim. Acta Part B At. Spectrosc. 2018, 139, 63–69. [Google Scholar] [CrossRef]
- Chen, X.; Zhang, Y.; Li, X.; Yang, Z.; Liu, A.; Yu, X. Diagnosis and staging of multiple myeloma using serum-based laser-induced breakdown spectroscopy combined with machine learning methods. Biomed. Opt. Express 2021, 12, 3584–3596. [Google Scholar] [CrossRef]
- Xu, L.; Tetteh, G.; Lipkova, J.; Zhao, Y.; Li, H.; Christ, P.; Piraud, M.; Bück, A.; Shi, K.; Menze, B.H. Automated Whole-Body Bone Lesion Detection for Multiple Myeloma on 68Ga-Pentixafor PET/CT Imaging Using Deep Learning Methods. Contrast Media Mol. Imaging 2018, 2018, 1–11. [Google Scholar] [CrossRef]
- Mesguich, C.; Hindie, E.; de Senneville, B.D.; Tlili, G.; Pinaquy, J.-B.; Marit, G.; Saut, O. Improved 18-FDG PET/CT diagnosis of multiple myeloma diffuse disease by radiomics analysis. Nucl. Med. Commun. 2021, 42, 1135–1143. [Google Scholar] [CrossRef]
- Xiong, X.; Wang, J.; Hu, S.; Dai, Y.; Zhang, Y.; Hu, C. Differentiating Between Multiple Myeloma and Metastasis Subtypes of Lumbar Vertebra Lesions Using Machine Learning–Based Radiomics. Front. Oncol. 2021, 11, 128. [Google Scholar] [CrossRef]
- Bhattacharyya, S.; Epstein, J.; Suva, L.J. Biomarkers that Discriminate Multiple Myeloma Patients with or without Skeletal Involvement Detected Using SELDI-TOF Mass Spectrometry and Statistical and Machine Learning Tools. Dis. Markers 2006, 22, 245–255. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Howell, D.A.; Hart, R.I.; Smith, A.G.; MacLeod, U.; Patmore, R.; Cook, G.; Roman, E. Myeloma: Patient accounts of their pathways to diagnosis. PLoS ONE 2018, 13, e0194788. [Google Scholar] [CrossRef] [PubMed]
- Pawlyn, C.; Cairns, D.; Kaiser, M.; Striha, A.; Jones, J.; Shah, V.; Jenner, M.; Drayson, M.; Owen, R.; Gregory, W.; et al. The relative importance of factors predicting outcome for myeloma patients at different ages: Results from 3894 patients in the Myeloma XI trial. Leukemia 2020, 34, 604–612. [Google Scholar] [CrossRef] [PubMed]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning—Data Mining, Inference, and Prediction; Springer: New York, NY, USA, 2001. [Google Scholar]
- Craig, F.E.; Foon, K.A. Flow cytometric immunophenotyping for hematologic neoplasms. Blood 2008, 111, 3941–3967. [Google Scholar] [CrossRef] [PubMed]
- Van der Maaten, L.; Hinton, G. Visualizing high-dimensional data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Beam, A.L.; Kohane, I.S. Big data and machine learning in healthcare. JAMA 2018, 319, 1317–1318. [Google Scholar] [CrossRef]
- Lhermitte, L.; on behalf of Mthe EuroFlow Consortium; Mejstrikova, E.; van der Sluijs-Gelling, A.J.; Grigore, G.E.; Sedek, L.; Bras, A.E.; Gaipa, G.; da Costa, E.S.; Novakova, M.; et al. Automated database-guided expert-supervised orientation for immunophenotypic diagnosis and classification of acute leukemia. Leukemia 2017, 32, 874–881. [Google Scholar] [CrossRef]
- Ni, W.; Hu, B.; Zheng, C.; Tong, Y.; Wang, L.; Li, Q.-Q.; Tong, X.; Han, Y. Automated analysis of acute myeloid leukemia minimal residual disease using a support vector machine. Oncotarget 2016, 7, 71915–71921. [Google Scholar] [CrossRef]
- Hahn, D.W.; Omenetto, N. Laser-Induced Breakdown Spectroscopy (LIBS), Part I: Review of basic diagnostics and plas-ma-particle interactions: Still-challenging issues within the analytical plasma community. Appl. Spectrosc. 2010, 64, 335A–336A. [Google Scholar] [CrossRef]
- Gaudiuso, R.; Ewusi-Annan, E.; Melikechi, N.; Sun, X.; Liu, B.; Campesato, L.F.; Merghoub, T. Using LIBS to diagnose mel-anoma in biomedical fluids deposited on solid substrates: Limits of direct spectral analysis and capability of machine learning. Spectrochim. Acta Part B 2018, 146, 106–114. [Google Scholar] [CrossRef]
- Teng, G.; Wang, Q.; Zhang, H.; Xiaogli, W.; Yang, H.; Qi, X.; Cui, X.; Idrees, B.S.; Wei, K.; Khan, M.N. Discrimination of infil-trative glioma boundary based on laser-induced breakdown spectroscopy. Spectrochim. Acta Part B 2020, 165, 105787. [Google Scholar] [CrossRef]
- Gangemi, S.; Allegra, A.; Alonci, A.; Cristani, M.; Russo, S.; Speciale, A.; Penna, G.; Spatari, G.; Cannavò, A.; Bellomo, G.; et al. Increase of novel biomarkers for oxidative stress in patients with plasma cell disorders and in multiple myeloma patients with bone lesions. Inflamm. Res. 2012, 61, 1063–1067. [Google Scholar] [CrossRef] [PubMed]
- Dutoit, J.C.; Verstraete, K.L. MRI in multiple myeloma: A pictorial review of diagnostic and post-treatment findings. Insights Imaging 2016, 7, 553–569. [Google Scholar] [CrossRef] [PubMed]
- Cavo, M.; Terpos, E.; Nanni, C.; Moreau, P.; Lentzsch, S.; Zweegman, S.; Hillengass, J.; Engelhardt, M.; Usmani, S.Z.; Vesole, D.H.; et al. Role of 18F-FDG PET/CT in the diagnosis and management of multiple myeloma and other plasma cell disorders: A consensus statement by the International Myeloma Working Group. Lancet Oncol. 2017, 18, e206–e217. [Google Scholar] [CrossRef]
- Moreau, P.; Attal, M.; Caillot, D.; Macro, M.; Karlin, L.; Garderet, L.; Facon, T.; Benboubker, L.; Escoffre-Barbe, M.; Stoppa, A.-M.; et al. Prospective Evaluation of Magnetic Resonance Imaging and [18F]Fluorodeoxyglucose Positron Emission Tomography-Computed Tomography at Diagnosis and Before Maintenance Therapy in Symptomatic Patients with Multiple Myeloma Included in the IFM/DFCI 2009 Trial: Results of the IMAJEM Study. J. Clin. Oncol. 2017, 35, 2911–2918. [Google Scholar] [CrossRef]
- Vag, T.; Gerngross, C.; Herhaus, P.; Eiber, M.; Philipp-Abbrederis, K.; Graner, F.-P.; Ettl, J.; Keller, U.; Wester, H.-J.; Schwaiger, M. First Experience with Chemokine Receptor CXCR4–Targeted PET Imaging of Patients with Solid Cancers. J. Nucl. Med. 2016, 57, 741–746. [Google Scholar] [CrossRef]
- Philipp-Abbrederis, K.; Herrmann, K.; Knop, S.; Schottelius, M.; Eiber, M.; Lueckerath, K.; Pietschmann, E.; Habringer, S.; Gerngroß, C.; Franke, K.; et al. In vivo molecular imaging of chemokine receptor CXCR 4 expression in patients with advanced multiple myeloma. EMBO Mol. Med. 2015, 7, 477–487. [Google Scholar] [CrossRef]
- Tang, N.; Tornatore, P.; Weinberger, S.R. Current developments in SELDI affinity technology. Mass Spectrom. Rev. 2003, 23, 34–44. [Google Scholar] [CrossRef]
- Petricoin, E.F.; Ardekani, A.M.; Hitt, B.A.; Levine, P.J.; Fusaro, V.A.; Steinberg, S.M.; Mills, G.B.; Simone, C.; Fishman, D.A.; Kohn, E.C.; et al. Use of proteomic patterns in serum to identify ovarian cancer. Lancet 2002, 359, 572–577. [Google Scholar] [CrossRef]
- Kourou, K.; Exarchos, K.P.; Papaloukas, C.; Sakaloglou, P.; Exarchos, T.; Fotiadis, D.I. Applied machine learning in cancer research: A systematic review for patient diagnosis, classification and prognosis. Comput. Struct. Biotechnol. J. 2021, 19, 5546–5555. [Google Scholar] [CrossRef]
- Farswan, A.; Gupta, A.; Gupta, R.; Hazra, S.; Khan, S.; Kumar, L.; Sharma, A. AI-supported modified risk staging for multiple myeloma cancer useful in real-world scenario. Transl. Oncol. 2021, 14, 101157. [Google Scholar] [CrossRef] [PubMed]
- Flores-Montero, J.; Sanoja-Flores, L.; Paiva, B.; Puig, N.; García-Sánchez, O.; Böttcher, S.; Van Der Velden, V.H.J.; Pérez-Morán, J.-J.; Vidriales, M.-B.; García-Sanz, R.; et al. Next Generation Flow for highly sensitive and standardized detection of minimal residual disease in multiple myeloma. Leukemia 2017, 31, 2094–2103. [Google Scholar] [CrossRef] [PubMed]
- Orgueira, A.M.; Pérez, M.S.G.; Arias, J.Á.D.; Rodríguez, B.A.; Vence, N.A.; López, B.; Blanco, A.A.; Pérez, L.B.; Raíndo, A.P.; López, M.C.; et al. Survival prediction and treatment optimization of multiple myeloma patients using machine-learning models based on clinical and gene expression data. Leukemia 2021, 35, 2924–2935. [Google Scholar] [CrossRef] [PubMed]
- Ban, C.; Yang, F.; Wei, M.; Liu, Q.; Wang, J.; Chen, L.; Lu, L.; Xie, D.; Liu, L.; Huang, J. Integrative Analysis of Gene Expression Through One-Class Logistic Regression Machine Learning Identifies Stemness Features in Multiple Myeloma. Front. Genet. 2021, 12. [Google Scholar] [CrossRef]
- Mitra, A.; Harding, T.; Mukherjee, U.K.; Jang, J.S.; Li, Y.; Hongzheng, R.; Jen, J.; Sonneveld, P.; Kumar, S.; Kuehl, W.M.; et al. A gene expression signature distinguishes innate response and resistance to proteasome inhibitors in multiple myeloma. Blood Cancer J. 2017, 7, e581. [Google Scholar] [CrossRef]
- Ubels, J.; Sonneveld, P.; Van Vliet, M.H.; De Ridder, J. Gene Networks Constructed Through Simulated Treatment Learning can Predict Proteasome Inhibitor Benefit in Multiple Myeloma. Clin. Cancer Res. 2020, 26, 5952–5961. [Google Scholar] [CrossRef]
- Borisov, N.; Sergeeva, A.; Suntsova, M.; Raevskiy, M.; Gaifullin, N.; Mendeleeva, L.; Gudkov, A.; Nareiko, M.; Garazha, A.; Tkachev, V.; et al. Machine Learning Applicability for Classification of PAD/VCD Chemotherapy Response Using 53 Multiple Myeloma RNA Sequencing Profiles. Front. Oncol. 2021, 11, 1124. [Google Scholar] [CrossRef]
- Povoa, L.V.; Ribeiro, C.H.C.; da Silva, I.T. Machine learning predicts treatment sensitivity in multiple myeloma based on molecular and clinical information coupled with drug response. PLoS ONE 2021, 16, e0254596. [Google Scholar] [CrossRef]
- Durie, B.G.; Salmon, S.E. A clinical staging system for multiple myeloma correlation of measured myeloma cell mass with presenting clinical features, response to treatment, and survival. Cancer 1975, 36, 842–854. [Google Scholar] [CrossRef]
- Greipp, P.R.; Miguel, J.S.; Durie, B.G.; Crowley, J.J.; Barlogie, B.; Bladé, J.; Boccadoro, M.; Child, J.A.; Avet-Loiseau, H.; Kyle, R.A.; et al. International Staging System for Multiple Myeloma. J. Clin. Oncol. 2005, 23, 3412–3420. [Google Scholar] [CrossRef]
- Palumbo, A.; Avet-Loiseau, H.; Oliva, S.; Lokhorst, H.M.; Goldschmidt, H.; Rosinol, L.; Richardson, P.; Caltagirone, S.; Lahuerta, J.J.; Facon, T.; et al. Revised International Staging System for Multiple Myeloma: A Report From International Myeloma Working Group. J. Clin. Oncol. 2015, 33, 2863–2869. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, K. Latest treatment strategies aiming for a cure in transplant-eligible multiple myeloma patients: How I cure younger MM patients with lower cost. Int. J. Hematol. 2020, 111, 512–518. [Google Scholar] [CrossRef] [PubMed]
- Cremers, E.M.; Westers, T.M.; Alhan, C.; Cali, C.; Visser-Wisselaar, H.A.; Chitu, D.A.; van der Velden, V.H.; Marvelde, J.G.T.; Klein, S.K.; Muus, P.; et al. Implementation of erythroid lineage analysis by flow cytometry in diagnostic models for myelodysplastic syndromes. Haematologica 2016, 102, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Ettari, R.; Zappalà, M.; Grasso, S.; Musolino, C.; Innao, V.; Allegra, A. Immunoproteasome-selective and non-selective inhibitors: A promising approach for the treatment of multiple myeloma. Pharmacol. Ther. 2018, 182, 176–192. [Google Scholar] [CrossRef]
- Allegra, A.; Sant’Antonio, E.; Penna, G.; Alonci, A.; D’Angelo, A.; Russo, S.; Cannavò, A.; Gerace, D.; Musolino, C. Novel therapeutic strategies in multiple myeloma: Role of the heat shock protein inhibitors. Eur. J. Haematol. 2010, 86, 93–110. [Google Scholar] [CrossRef]
- Allegra, A.; Penna, G.; Alonci, A.; Russo, S.; Greve, B.; Innao, V.; Minardi, V.; Musolino, C. Monoclonal antibodies: Potential new therapeutic treatment against multiple myeloma. Eur. J. Haematol. 2013, 90, 441–468. [Google Scholar] [CrossRef]
- Kumar, S.; Rajkumar, S.V. Many facets of bortezomib resistance/susceptibility. Blood 2008, 112, 2177–2178. [Google Scholar] [CrossRef]
- Allegra, A.; Ettari, R.; Innao, V.; Bitto, A. Potential Role of microRNAs in inducing Drug Resistance in Patients with Multiple Myeloma. Cells 2021, 10, 448. [Google Scholar] [CrossRef]
- Paiva, B.; Corchete, L.A.; Vidriales, M.-B.; Puig, N.; Maiso, P.; Rodriguez, I.; Alignani, D.; Burgos, L.; Sanchez, M.-L.; Barcena, P.; et al. Phenotypic and genomic analysis of multiple myeloma minimal residual disease tumor cells: A new model to understand chemoresistance. Blood 2016, 127, 1896–1906. [Google Scholar] [CrossRef]
- Moreaux, J.; Klein, B.; Bataille, R.; Descamps, G.; Maiga, S.; Hose, D.; Goldschmidt, H.; Jauch, A.; Rème, T.; Jourdan, M.; et al. A high-risk signature for patients with multiple myeloma established from the molecular classification of human mye-loma cell lines. Haematologica 2011, 96, 574–582. [Google Scholar] [CrossRef]
- Kuiper, R.; Broyl, A.; De Knegt, Y.; Van Vliet, M.H.; Van Beers, E.H.; Van Der Holt, B.; El Jarari, L.; Mulligan, G.; Gregory, W.; Morgan, G.; et al. A gene expression signature for high-risk multiple myeloma. Leukemia 2012, 26, 2406–2413. [Google Scholar] [CrossRef]
- Zhan, F.; Barlogie, B.; Mulligan, G.; Shaughnessy, J.D., Jr.; Bryant, B. High-risk myeloma: A gene expression based risk-stratification model for newly diagnosed multiple myeloma treated with high-dose therapy is predictive of outcome in relapsed disease treated with single-agent bortezomib or high-dose dexamethasone. Blood 2008, 111, 968–969. [Google Scholar] [CrossRef]
- Allegra, A.; Alonci, A.; Gerace, D.; Russo, S.; Innao, V.; Calabrò, L.; Musolino, C. New orally active proteasome inhibitors in multiple myeloma. Leuk. Res. 2014, 38, 1–9. [Google Scholar] [CrossRef]
- Bernau, C.; Riester, M.; Boulesteix, A.-L.; Parmigiani, G.; Huttenhower, C.; Waldron, L.; Trippa, L. Cross-study validation for the assessment of prediction algorithms. Bioinformatics 2014, 30, i105–i112. [Google Scholar] [CrossRef]
- Ubels, J.; Sonneveld, P.; van Beers, E.H.; Broijl, A.; van Vliet, M.H.; de Ridder, J. Predicting treatment benefit in multiple myeloma through simulation of alternative treatment effects. Nat. Commun. 2018, 9, 2943. [Google Scholar] [CrossRef]
- Gomez-Bougie, P.; Wuillème-Toumi, S.; Ménoret, E.; Trichet, V.; Robillard, N.; Philippe, M.; Bataille, R.; Amiot, M. Noxa Up-regulation and Mcl-1 Cleavage Are Associated to Apoptosis Induction by Bortezomib in Multiple Myeloma. Cancer Res. 2007, 67, 5418–5424. [Google Scholar] [CrossRef]
- Podar, K.; le Gouill, S.; Zhang, J.; Opferman, J.T.; Zorn, E.; Tai, Y.-T.; Hideshima, T.; Amiot, M.; Chauhan, D.; Harousseau, J.-L.; et al. A pivotal role for Mcl-1 in Bortezomib-induced apoptosis. Oncogene 2008, 27, 721–731. [Google Scholar] [CrossRef][Green Version]
- Gong, J.-N.; Khong, T.; Segal, D.; Yao, Y.; Riffkin, C.D.; Garnier, J.-M.; Khaw, S.L.; Lessene, G.; Spencer, A.; Herold, M.J.; et al. Hierarchy for targeting prosurvival BCL2 family proteins in multiple myeloma: Pivotal role of MCL1. Blood 2016, 128, 1834–1844. [Google Scholar] [CrossRef]
- Bodet, L.; Bougie, P.G.; Touzeau, C.; Dousset, C.; Descamps, G.; Maïga, S.; Avet-Loiseau, H.; Bataille, R.; Moreau, P.; le Gouill, S.; et al. ABT-737 is highly effective against molecular subgroups of multiple myeloma. Blood 2011, 118, 3901–3910. [Google Scholar] [CrossRef]
- Kline, M.P.; Rajkumar, S.V.; Timm, M.M.; Kimlinger, T.K.; Haug, J.L.; Lust, J.A.; Greipp, P.R.; Kumar, S. ABT-737, an inhibitor of Bcl-2 family proteins, is a potent inducer of apoptosis in multiple myeloma cells. Leukemia 2007, 21, 1549–1560. [Google Scholar] [CrossRef]
- Kumar, S.; Kaufman, J.L.; Gasparetto, C.; Mikhael, J.; Vij, R.; Pegourie, B.; Benboubker, L.; Facon, T.; Amiot, M.; Moreau, P.; et al. Efficacy of venetoclax as targeted therapy for relapsed/refractory t(11;14) multiple myeloma. Blood 2017, 130, 2401–2409. [Google Scholar] [CrossRef] [PubMed]
- Teh, C.E.; Gong, J.-N.; Segal, D.; Tan, T.; Vandenberg, C.J.; Fedele, P.L.; Low, M.S.Y.; Grigoriadis, G.; Harrison, S.J.; Strasser, A.; et al. Deep profiling of apoptotic pathways with mass cytometry identifies a synergistic drug combination for killing myeloma cells. Cell Death Differ. 2020, 27, 2217–2233. [Google Scholar] [CrossRef] [PubMed]
- Topol, E.J. High-performance medicine: The convergence of human and artificial intelligence. Nat. Med. 2019, 25, 44–56. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Baxter, S.L.; Xu, J.; Xu, J.; Zhou, X.; Zhang, K. The practical implementation of artificial intelligence technologies in medicine. Nat. Med. 2019, 25, 30–36. [Google Scholar] [CrossRef]
- Ståhl, P.L.; Salmén, F.; Vickovic, S.; Lundmark, A.; Navarro, J.F.; Magnusson, J.; Giacomello, S.; Asp, M.; Westholm, J.O.; Huss, M.; et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 2016, 353, 78–82. [Google Scholar] [CrossRef]
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Math, M.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 2012, 366, 883–892. [Google Scholar] [CrossRef]
- He, B.; Bergenstråhle, L.; Stenbeck, L.; Abid, A.; Andersson, A.; Borg, Å.; Maaskola, J.; Lundeberg, J.; Zou, J. Integrating spatial gene expression and breast tumour morphology via deep learning. Nat. Biomed. Eng. 2020, 4, 827–834. [Google Scholar] [CrossRef]
- Confalonieri, R.; Coba, L.; Wagner, B.; Besold, T.R. A historical perspective of explainable Artificial Intelligence. WIREs Data Min. Knowl. Discov. 2020, 11, e1391. [Google Scholar] [CrossRef]
- Rubin, D.L. Artificial Intelligence in Imaging: The Radiologist’s Role. J. Am. Coll. Radiol. 2019, 16, 1309–1317. [Google Scholar] [CrossRef]
- Savadjiev, P.; Chong, J.; Dohan, A.; Vakalopoulou, M.; Reinhold, C.; Paragios, N.; Gallix, B. Demystification of AI-driven medical image interpretation: Past, present and future. Eur. Radiol. 2018, 29, 1616–1624. [Google Scholar] [CrossRef]
- Price, W.N.; Gerke, S.; Cohen, I.G. Potential Liability for Physicians Using Artificial Intelligence. JAMA J. Am. Med. Assoc. 2019, 322, 1765. [Google Scholar] [CrossRef]
- Price, W.N.; Gerke, S.; Cohen, I.G. How Much Can Potential Jurors Tell Us About Liability for Medical Artificial Intelligence? J. Nucl. Med. 2020, 62, 15–16. [Google Scholar] [CrossRef]
- Coppola, F.; Faggioni, L.; Gabelloni, M.; De Vietro, F.; Mendola, V.; Cattabriga, A.; Cocozza, M.A.; Vara, G.; Piccinino, A.; Monaco, S.L.; et al. Human, All too Human? An All-around Appraisal of the “Artificial Intelligence Revolution” in Medical Imaging. Front. Psychol. 2021, 12, 4296. [Google Scholar] [CrossRef]
- Triberti, S.; Durosini, I.; Pravettoni, G. A “Third Wheel” Effect in Health Decision Making Involving Artificial Entities: A Psychological Perspective. Front. Public Health 2020, 8, 117. [Google Scholar] [CrossRef]
- Coppola, F.; Faggioni, L.; Regge, D.; Giovagnoni, A.; Golfieri, R.; Bibbolino, C.; Miele, V.; Neri, E.; Grassi, R. Artificial intelligence: Radiologists’ expectations and opinions gleaned from a nationwide online survey. Radiol. Med. 2020, 126, 63–71. [Google Scholar] [CrossRef]

| Diagnosis | |||
|---|---|---|---|
| Parameters | AI Tools | Ref. | Key Findings |
| Blood and biochemical exams | Gradient boosting decisional tree | [24] | A ML approach on standard laboratory findings enhances the percentage of early detection |
| Differential cell counts of bone marrow aspirate | VGG16 convolutional network | [25] | Bone marrow aspirate differential counts employing ML techniques |
| Cytofluorimetric analysis of bone marrow aspirate | FlowCAP | [26] | Computerized methods for cytofluorimetric analysis |
| Gradient boosting machine technique | [27] | Classification of plasma cell dyscrasias by combining AI and flow cytometry | |
| Laser-induced breakdown spectroscopy analysis | Quadratic discriminant analysis, k-Nearest Neighbour | [28] | Diagnosis of malignancies using serum-based laser induced breakdown spectroscopy and chemometric methods |
| K-Nearest Neighbour, Support Vector Machine, Artificial Neural Networks | [29] | Diagnosis of malignancies using serum-based laser induced breakdown spectroscopy in combination with ML methods can serve as fast technique for MM diagnosis and staging | |
| Bone Lesions Identification | |||
| Techniques | AI tools | Ref. | Key findings |
| PET and CT | Convolutional neural network (v-Net, w-Net) | [30] | 68Ga-Pentixaflor PET/CT and DL techniques to detect MM whole-body bone lesions |
| PET and CT | Random Forest | [31] | Radiomics analysis of 18-FDG PET/CT image with ML overcame the limitations of visual analysis |
| MRI | Naïve Bayes, Support Vector Machine, k-Nearest Neighbour, Random Forest, Artificial Neural Networks | [32] | ML radiomics is able to differentiate between MM and metastasis subtypes of lumbar vertebra lesions |
| SELDI-TOF-MS (mass peaks with mass-to-charge ratios) | Random Forest, Partial least squares discriminant analysis | [33] | SELDI-TOF-MS and ML tools discriminate MM patients with and without skeletal involvement |
| Prognosis | ||||
|---|---|---|---|---|
| Parameters | AI Tools | Ref. | Key Findings | |
| Laboratory parameters | k-adaptive partitioning | [54] | AI-supported modified risk staging for multiple myeloma | |
| Beta2microglobulin | Infinicyt software | [55] | Next-Generation Flow and ML for highly sensitive detection of minimal residual disease | |
| Gene expression profile, ISS stage, first line therapy | Random Forest | [56] | Survival prediction and treatment optimization using ML models based on clinical and gene expression data | |
| mRNA expression-based steamness index | One-class logistic regression | [57] | Analysis of gene expression via one-class logistic regression ML identifies stemness features in MM | |
| Prediction of Response to Treatment | ||||
| Drugs | Parameters | AI tools | Ref. | |
| Bortezomib, carfilzomib, ixazomib, oprozomib | Gene expression profile | Random Forest | [58] | A gene expression signature distinguishes resistance to proteasome inhibitors |
| Proteasome inhibitors | Gene complex | Simulated Treatment learning signature | [59] | Gene networks constructed using simulated treatment learning can predict proteasome inhibitor benefit |
| PAD, VCD | Gene evaluation | Random Forest, Support Vector Machine, Ridge Regression, Binomial Naïve Bayes, Multi-layer perception | [60] | ML applicability for classification of chemotherapy response using 53 MM RNA-sequencing profiles |
| Five first-line treatments (Bor-Cyc-Dex, Bor-Dex, Bor-Len-Dex, Len-Dex, Non-treatment) | Clinical markers, gene evaluation | Multi Learning Training approach | [61] | ML predicts treatment sensitivity in MM based on molecular and clinical information coupled with drug response |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Allegra, A.; Tonacci, A.; Sciaccotta, R.; Genovese, S.; Musolino, C.; Pioggia, G.; Gangemi, S. Machine Learning and Deep Learning Applications in Multiple Myeloma Diagnosis, Prognosis, and Treatment Selection. Cancers 2022, 14, 606. https://doi.org/10.3390/cancers14030606
Allegra A, Tonacci A, Sciaccotta R, Genovese S, Musolino C, Pioggia G, Gangemi S. Machine Learning and Deep Learning Applications in Multiple Myeloma Diagnosis, Prognosis, and Treatment Selection. Cancers. 2022; 14(3):606. https://doi.org/10.3390/cancers14030606
Chicago/Turabian StyleAllegra, Alessandro, Alessandro Tonacci, Raffaele Sciaccotta, Sara Genovese, Caterina Musolino, Giovanni Pioggia, and Sebastiano Gangemi. 2022. "Machine Learning and Deep Learning Applications in Multiple Myeloma Diagnosis, Prognosis, and Treatment Selection" Cancers 14, no. 3: 606. https://doi.org/10.3390/cancers14030606
APA StyleAllegra, A., Tonacci, A., Sciaccotta, R., Genovese, S., Musolino, C., Pioggia, G., & Gangemi, S. (2022). Machine Learning and Deep Learning Applications in Multiple Myeloma Diagnosis, Prognosis, and Treatment Selection. Cancers, 14(3), 606. https://doi.org/10.3390/cancers14030606








