Simple Summary
Tau protein is a microtubule-associated protein, widely known for its role in neurodegenerative diseases. Recent studies show that Tau can also be involved in the progression of many cancers as well as in cancer treatment resistance. This work overviews the role of Tau in tumorigenic processes with a focus on glioblastoma which exhibits a high expression level of this protein. Unraveling the role of Tau in glioblastoma and cancers, in general, will boost our understanding of the Tau protein and more significantly may open new avenues of strategies for curing cancer patients, in particular those with glioblastoma.
Abstract
Despite being extensively studied for several decades, the microtubule-associated protein Tau has not finished revealing its secrets. For long, Tau has been known for its ability to promote microtubule assembly. A less known feature of Tau is its capability to bind to cancer-related protein kinases, suggesting a possible role of Tau in modulating microtubule-independent cellular pathways that are associated with oncogenesis. With the intention of finding new therapeutic targets for cancer, it appears essential to examine the interaction of Tau with these kinases and their consequences. This review aims at collecting the literature data supporting the relationship between Tau and cancer with a particular focus on glioblastoma tumors in which the pathological significance of Tau remains largely unexplored. We will first treat this subject from a mechanistic point of view showing the pivotal role of Tau in oncogenic processes. Then, we will discuss the involvement of Tau in dysregulating critical pathways in glioblastoma. Finally, we will outline promising strategies to target Tau protein for the therapy of glioblastoma.
1. Introduction
The microtubule cytoskeleton forms a functional network in eukaryotic cells. It participates in many cellular activities such as cell division, motility, morphogenesis, and intracellular trafficking of both macromolecules and organelles [1]. These processes depend on the dynamic instability of microtubules which is tightly regulated by associated proteins such as Tau. Tau is a protein that stabilizes and promotes the assembly of microtubules [2,3,4]. Interest in Tau stems from its critical role in neurodegenerative diseases, the so-called tauopathies. Indeed, the experimental models of tauopathies strongly suggest that Tau-mediated neurodegeneration results from a combination of toxic gain of function and the loss of normal Tau function [5,6,7,8,9]. Tau has also been reported to be implicated in different types of cancer. Consequently, many researchers have started to shift their attention toward understanding the role of this protein in oncogenesis [10,11,12,13]. Indeed, many studies have reported abnormal expression of Tau in breast, stomach, prostate, and brain cancer cells [14,15]. Furthermore, the expression level of Tau has been linked to resistance to anti-microtubule agents in cancer [12,16,17].
Recent advances in our understanding of Tau protein and its different localizations inside and outside neurons show that Tau has a wide interactome, including cancer-related kinase proteins. These interactions are likely to play an important role in a diverse range of cellular pathways such as cell signaling, cell motility, cellular metabolism, or genomic modifications as well as cancers, including glioblastoma, the deadliest brain tumor. Glioblastoma multiform (GBM) accounts for 52–75% of diagnosed gliomas with a median survival of patients of only 15 months [18,19,20]. Standard therapy, which consists of resection of the tumor, radiotherapy, followed by adjuvant chemotherapy with the alkylating agent temozolomide (TMZ) is unable to extend a patient’s life for more than two and a half months [21,22,23]. In addition, the efficacy of TMZ-based chemotherapy is limited in clinical application, mainly due to drug resistance [24,25]. In this context, the pathological significance of Tau protein remains poorly examined, considering that GBM tumors express variable levels of Tau [15,26]. Given the complicated nature of GBM progression, it would not be surprising that there exists a complex array of mechanisms involving Tau and diverse cellular signaling pathways where pharmacological intervention would allow opportunities to treat the disease.
Here, we review the understudied microtubule-independent functions of Tau, and its interplay with cellular signaling. Furthermore, we present Tau as a vulnerable target for cancer systems and a credible oncotarget for GBM. Moreover, we examine some of the Tau protein interactants that are initially described in neurodegenerative disorders, and which should be explored in glioma and/or GBM. Finally, we review interesting approaches to target Tau and examine efforts to apply them for GBM treatment.
2. Tau Structure and Regulation
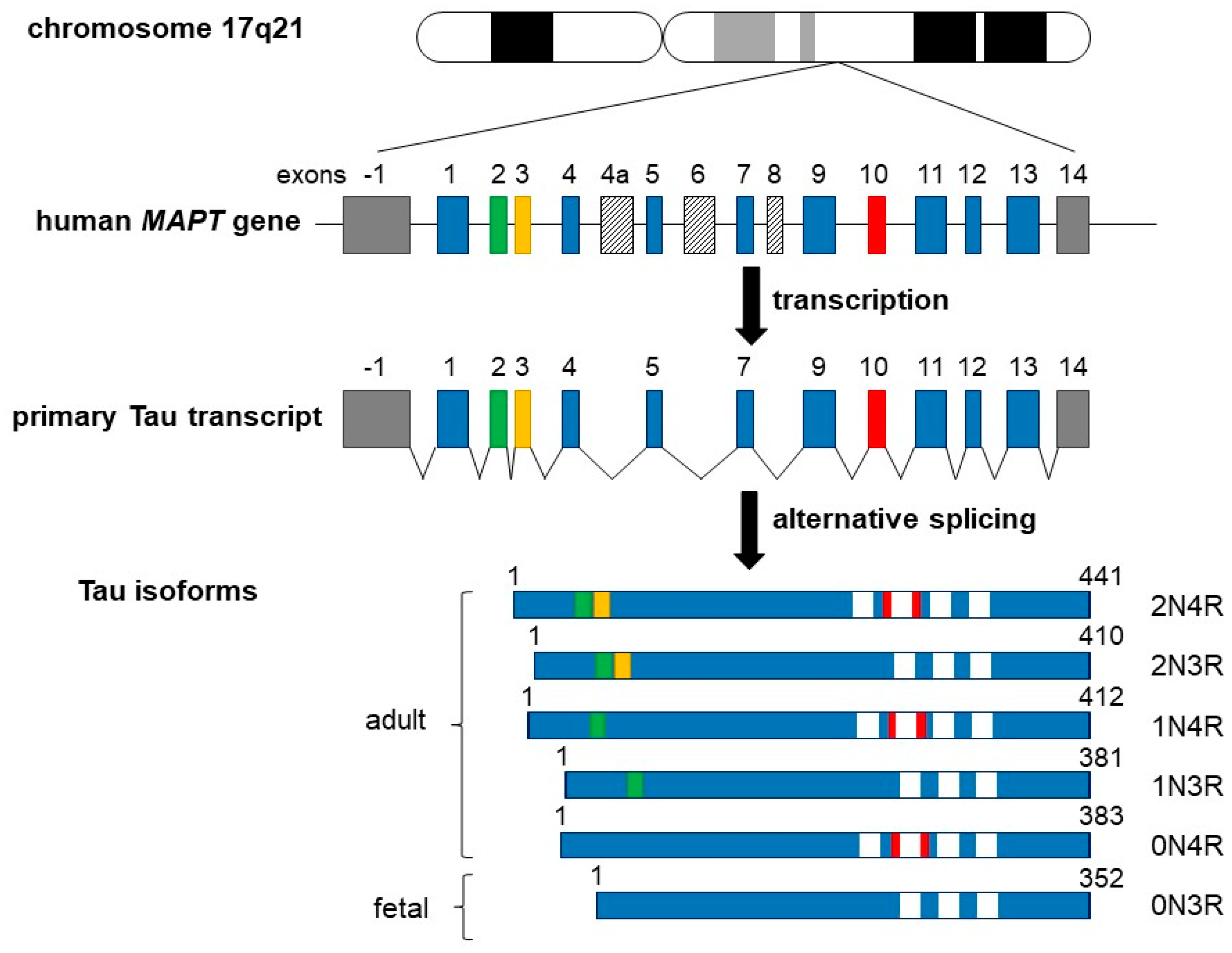
In the adult human brain, there are six predominant Tau isoforms, with a primary structure consisting of 352 to 441 amino acids [27,28,29,30] (see also Figure 1 for details). Tau protein is encoded by the MAPT gene which comprises 16 exons encoding a single mRNA undergoing alternative splicing of exons 2–3 and 10. Tau isoforms can be distinguished by the presence of zero, one, or two N-terminal inserts of 29 residues each, involving the splicing of exons 2 and/or 3. Exon 10 can be spliced in (four repeats) or out (three repeats), encoding different microtubule-binding domain (MTBD) sequences [27,30,31,32,33,34,35]. The spliced mRNA gives therefore the six isoforms 0N3R, 0N4R, 1N3R, 1N4R, 2N3R, and 2N4R in which “N” indicates the number of N-terminal inserts and “R” represents the number of MTBD sequences. Note that the shortest Tau isoform without exons 2, 3, 4a, and 10 (also called “fetal Tau”) is predominantly expressed in the fetal brain [36,37].
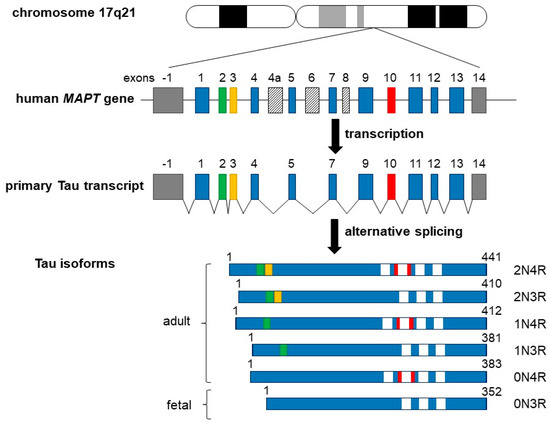
Figure 1.
Schematic representation of the human MAPT (Tau) gene, the primary Tau transcript, and the six Tau protein isoforms. The MAPT gene is located over 100 kb of the long arm of chromosome 17 at position 17q21. It contains 16 exons (upper panel), while the tau primary transcript contains 13 exons (middle panel). Exon −1 (a part of the gene promotor) and 14 (gray boxes) are transcribed but not translated. Exons 4a, 6, and 8 (slashed boxes) are not transcribed in it. Six different Tau isoforms are shown in lower panel, which have been translated by alternatively spliced exons 2, 3, and 10, and constitutive exons 1, 4, 5, 7, 9, 11, 12, and 13, which form six different mRNAs. These isoforms vary due to the presence or absence of one or two 29 amino acids inserts, which are encoded by exon 2 (green box) and exon 3 (yellow box) in the N-terminal part in combination with either three (R1, R3, and R4) or four (R1–R4) repeat regions at carboxyl end (white boxes). The additional microtubule-binding domain is encoded by exon 10 (red box). Adult Tau includes all six Tau isoforms, including the largest isoform of 441 amino acids containing all inserts and other isoforms as indicated. The shortest 352 amino acids isoform is the only one found only in fetal brain.
Tau protein is a highly soluble and heat-stable protein and is usually considered an intrinsically disordered protein in solution [38,39,40,41,42,43,44,45]. Based on the longest 2N4R isoform, the first 44 amino acids contain a glycine-rich sequence that encompasses the two highly acidic regions (N1 and N2) [30]. Moreover, threonine (or alanine) residues are present in a higher proportion at the N-terminal region [46]. Regarding the possible function of this N-terminal region, it has been reported that Tau protein can be associated with different cell membrane proteins such as apolipoprotein A1 (preferentially with the 2N isoform) or synaptophysin (preferentially with the 0N isoform) [47], two key proteins involved in the neurite outgrowth and synaptic dysfunction [48,49].
Moreover, the N-terminal region also contains two proline-rich domains (PRD) followed by the four MTBD repeats (R1–R4) belonging to the C-terminal domain of Tau protein. Apart from their well-described involvement in binding to microtubules, the PRD regions also play a pivotal role in the interaction of Tau with diverse components of other cytoskeletons and of the plasma membrane, as we mention later [50,51,52,53,54]. Moreover, the ability of Tau to bind microtubules is mediated by its MTBD in the C-terminal region. A single MTBD is sufficient for its binding to the microtubule. Each of the MTBDs repeats has a highly homologous 18 amino acid repeat region containing a KXGS motif that can be phosphorylated [28,29,55]. Additionally, it has been shown that 3R and 4R Tau isoforms can adopt many different structures to differentially regulate their interaction with microtubules [56]. Furthermore, it has been reported that 4R Tau is more prone to phosphorylation in vitro and it is more likely to aggregate into paired helical filaments (PHFs) than the 3R Tau, a finding that could be relevant to the understanding of tauopathies such as AD [57]. In cancer, doubts remain concerning the impact of post-translational modifications such as the phosphorylation of specific residues of Tau on its function. At least, an accumulation of hyperphosphorylated Tau in the periphery of GBM tumors in patients has been reported, suggesting a possible role of Tau phosphorylation in the invasiveness of cancer cells [58].
Overall, tubulin, undoubtedly, remains the main ligand for the Tau protein. Under physiological conditions, up to 90% of Tau is bound to microtubules and therefore unavailable for other interactions [59]. In cancer, Tau plays a pivotal role in modifying the response to microtubules targeting chemotherapies by directly modulating microtubules and their participation in the neoplastic process [10,11,13,60]. Moreover, the binding of Tau to microtubules is very dynamic [61], which could explain why Tau can be present in intracellular compartments normally devoid of microtubules such as the nucleus. In fact, it is this presence in the nucleus that prompted and hinted to researchers that Tau has additional functions independent of its interaction with microtubules in cells. Many reports now show that Tau can co-locate and interact with multiple non-cytoskeletal proteins, many of which are involved in important cancer signaling pathways. In the following, we will explore some of these additional functions of Tau and explain how relevant they are likely to be to oncological processes.
3. Relationship between Tau Expression and Dysfunctions in Cancer
Tau is found with an abnormally high expression in various types of cancer such as breast cancer, ovarian cancer, gastric cancer, prostate cancer, pediatric neuroblastoma, glioma, and more [14,15,16,62,63]. Regarding these cancers, the aberrant expression of Tau is not necessarily related to the progression of this pathology. For example, high expression of Tau in several types of carcinomas, including breast cancer, prostate cancer, and gliomas [60,64,65], has been associated with improved patient prognosis. However, it seems to be related to worse survival in other cancer types such as ovarian cancer [63,66]. The question is still open regarding whether Tau expression represents a predictive marker for (neo-) adjuvant chemotherapy response or not. In vitro, Tau protein competes with microtubule-stabilizing agents such as taxane for controlling microtubule dynamic and Tau expression loss may render microtubules more vulnerable to the effects of these agents [61]. However, if we take breast cancer as an example, it appears clearly that Tau protein expression is insufficient for identifying a subset of patients with carcinomas that may benefit more from taxane-based chemotherapy [67]. For more details on the prognostic value of Tau and resistance to microtubule-targeting drugs in cancer, interested readers can refer to the review written by Papin and Paganetti [68].
Regarding its functions, cumulative studies are suggesting that the role of Tau goes far beyond its microtubule-stabilizing function and that Tau is involved in many key signaling pathways, such as cell differentiation and proliferation, morphogenesis, motility, etc. At first sight, the microtubule-stabilizing function of Tau appears relevant to cancer because the microtubule dynamic ensures several critical functions such as cell motility, cytoplasmic transport, and cell division [69]. Other functions of Tau independent of interaction with microtubules are expected to be relevant to cancer. The following paragraphs will explain each demonstrated function and how it may be associated with cancer, such as genomic instability, cell cycle, mitosis and proliferation, cell migration, and angiogenesis.
3.1. Tau and Genomic Instability
One of the hallmarks of cancer is genomic instability manifesting through sequence instability and a loss of chromosomal integrity [70]. Alterations in many gene sequences confer new functional abilities for survival, proliferation, and invasiveness. Chromosomal instability can provide changes in sequence structure or the allelic number of tumor genes thus amplifying an oncogene or suppressing tumor-suppressing genes. Numerous studies brought proof that Tau can translocate to the nuclei of neurons [71,72,73,74], as well as in non-neuronal cells [75]. Using an immunofluorescence approach to label diverse tumor cells, Binder’s team reported for the first time in the 1990s the presence of Tau in the nuclei, especially near the nucleolar organizing regions of acrocentric chromosomes [76,77]. Since, it has been proposed that Tau is a DNA protector, notably by preventing stress-induced DNA breaks [78,79,80]. Indeed, the presence of Tau in the nucleus contributes to the maintenance of the DNA double helix structure and DNA winding [72,73,81]. Moreover, it has been reported that Tau functions in the nucleus are regulated by hyperphosphorylation–dephosphorylation processes, allowing Tau to bind to and protect DNA under oxidative stress conditions [45]. So, it is tempting to think that Tau could affect the progression of some cancers through a specific DNA binding site.
Other evidence that Tau is involved in chromosome integrity is the fact that specific mutations in the protein enable diverse chromosomal aberrations in cells. Rossi et al. have recently conducted a retrospective cohort study in which they analyzed cancer incidence in 15 families affected by frontotemporal lobar degeneration (FTLD) bearing MAPT mutations (detected by sequencing of exons 1, 9–13) [82]. Their study reveals that the FTLD family members have a significantly higher risk of developing cancer compared to control families. Moreover, bioinformatics analysis of pathways associated with the Tau protein interactome showed that the third of proteins interacting with Tau is involved in DNA damage recognition and repair, cell cycle checkpoints and phase transition, chromatin and telomere maintenance processes, and response to radiation stressors [82]. Taken together, this epidemiologic investigation supports another indirect role of Tau in DNA protection/repair systems.
3.2. Tau in Cell Cycle and Mitosis
The incidence of Tau in the cell cycle has been primarily explored in AD, notably to understand chromosome mis-segregation in brain cells and in peripheral tissues [83,84,85,86]. Physiologically, neurons charged with neurofibrillary tangles (NFT) hyperphosphorylated Tau undergo an abnormal cell cycle entry before apoptosis [87,88]. In the case of cancer, it is surprising that the impact of Tau phosphorylation on mitosis has been poorly examined considering the significant proliferative activity of tumor cells. In this regard, a recent study focused on the effect of Tau on mitosis: Flores-Rodriguez et al. have shown that the quantity of Tau phosphorylated on Thr231 in the PRD region (pT231-Tau) correlates to phases of cell division in the SH-S5Y neuroblastoma cell model [89]. Indeed, the authors at first observed an increased amount of pT231-Tau concomitantly with the placement of mitotic spindles and condensation of chromosomes during prophase and metaphase up to fill the entire cellular cytoplasm. Then, the quantity of pT231-Tau decreased markedly to diffuse granular staining during anaphase and telophase. These results suggest that the role of pT231-Tau is to maintain the integrity of the DNA and chromosomes during mitosis. In addition, the presence of Tau protein, maybe unphosphorylated here, could be involved in the separation of sister chromatids. To clarify, Bougé and Parmentier have recently reported the effect of Tau expression on chromosome mis-segregation in vivo [90]. Using models of wing disc epithelium of Drosophilia and tumor HeLa cells, the authors show that an excess of Tau induces mitotic arrest, with the presence of monopolar spindles. Interestingly, they also demonstrated that the mitotic defect occurs through the inhibition of the kinesin Klp61F, the Drosophilia homolog of Kif11 which ensures chromatid transport onto spindles. In cancer, it is hence questionable whether Tau could play a role in carcinogenesis or tumor progression, notably by defecting programmed cell death, necrosis, or senescence [91].
Moreover, many microtubule destabilizing and stabilizing compounds, called microtubule targeting agents (MTA) such as vinca alkaloid or taxoid, have been widely used as antimitotic agents for their induction of cell cycle arrest and apoptosis [92,93]. However, the relationship between the expression of Tau observed in many cancers and the relapse of patients who have followed a therapeutic protocol involving MTA is still conflicted [12,94]. One hypothesis may imply MTA-mediated activation of different cell death pathways such as autophagy, a mechanism in which the quantity of Tau could promote. Indeed, the potential association between autophagy inhibition and Tau has been recently investigated in prostate cancer cells where the presence of aberrant monoastral mitotic spindles has been evidenced to correlate both with the accumulation of Tau oligomers and with a recovered cytotoxic effect of docetaxel [95]. Interestingly, the authors suggest that cancer could maintain a high Tau protein turnover to avoid the formation of toxic oligomers, mainly during mitosis, through defective autophagy. Therefore, interfering with Tau turnover could represent an effective strategy to counteract the resistance to antimitotic agents such as MTA.
3.3. Tau in Cell Migration
Another characteristic of cancer is exacerbated cell migration. This process involves strong cell shape modifications and thus heavily relies on the redistribution and cooperation of cytoskeletal filamentous proteins, notably actin filaments and microtubules. While the requirement of crosstalk between microtubules and actin is not questioned, mechanisms underlying the microtubule-actin organization by cross-linkers remain largely unexplored, and little is known about the functional contribution of Tau protein. The impact of Tau overexpression on cell motility was investigated by Morris and coll. using an IDH wild-type human GBM-derived LN229 cell line [96]. The authors showed that high Tau expression selectively reduced cell migration without affecting cellular proliferation or apoptosis. As a reminder, isocitrate dehydrogenases (IDH) are enzymes that catalyze the oxidative decarboxylation of carbohydrates in the Krebs cycle to produce ATP in the mitochondria [97].
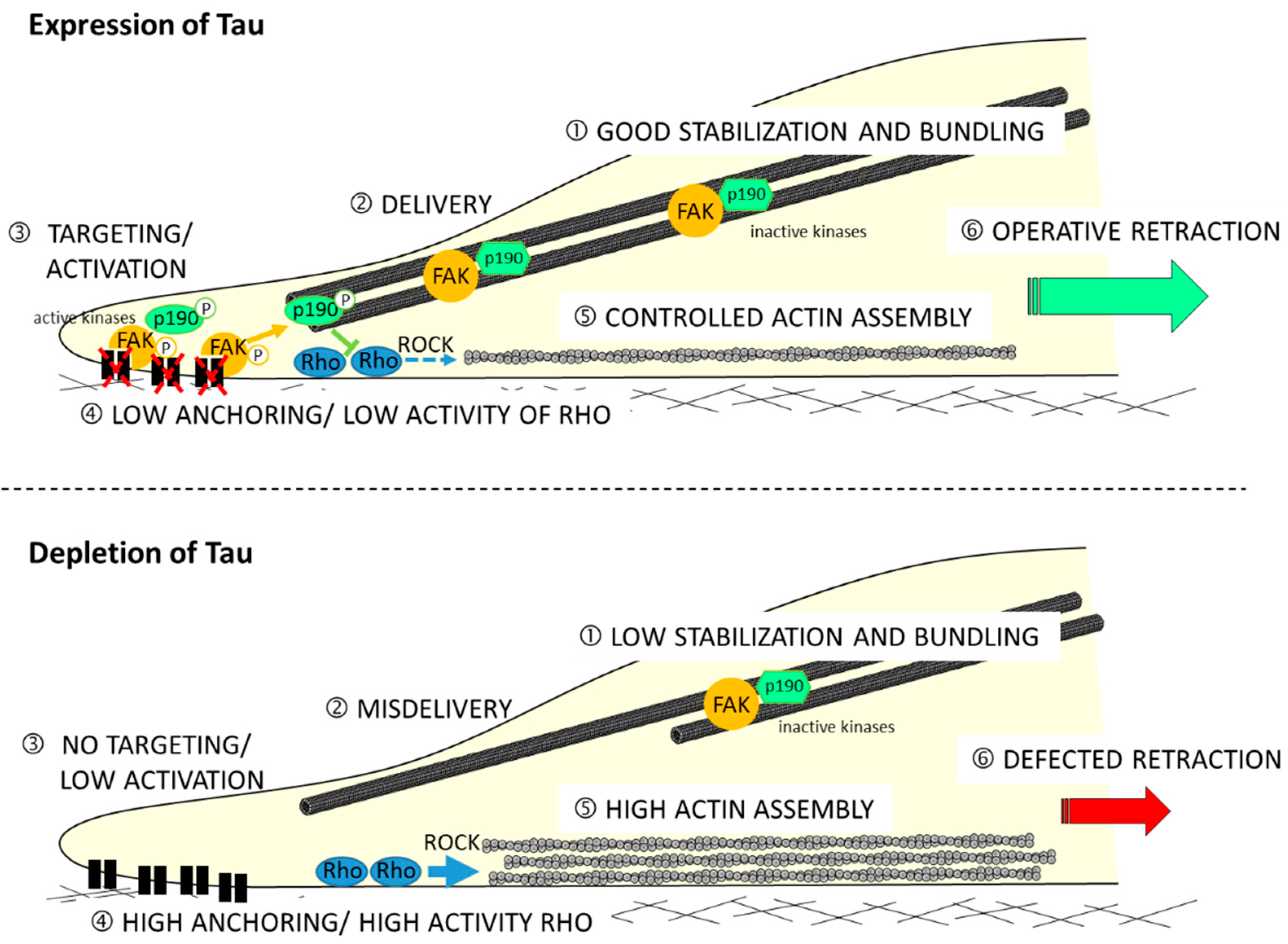
Moreover, the mechanism showing how Tau protein is involved in the migration process is not described. Recently, by using the shRNA approach to deplete Tau in GBM-derived U87MG cells (shTau cells), the authors determined its impact on cytoskeletal coordination during migration [98]. Results have revealed a 36% reduced motility in the shTau cells compared to the control cells with no impact on directionality. By analyzing the spatial organization of microtubule and actin networks, they observed extensive actin cables but limited microtubule bundling at the back of polarized shTau cells. This remodeling of cytoskeletons resulted in weak coordination between protrusion at the leading edge and retraction of the trailing edge of shTau cells. Authors also demonstrated that the remodeling of the actin cytoskeleton depends on the relocation of Rho-associated protein kinase 1 (ROCK1), as well as regulating kinases p190-RhoGAP and FAK. They therefore propose the following model (see Figure 2).
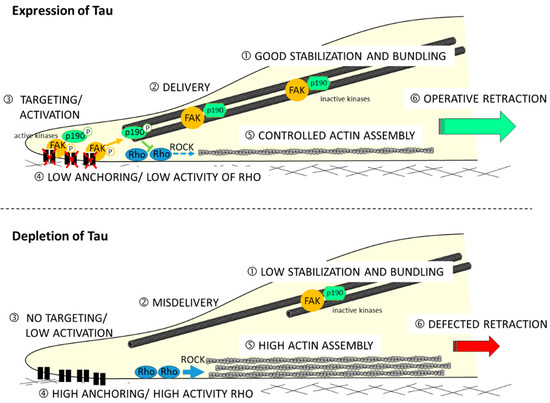
Figure 2.
Tau regulates the microtubule-dependent migration of glioblastoma U87MG cells via the Rho-ROCK signaling pathway. Directional cell migration requires spatiotemporal control of focal adhesion formation and dissociation: (upper model) in Tau-expressing cell, good stabilization and bundling of microtubules (1) are positioned at the tail of polarized cells; during migration, the active phosphorylated forms of FAK and p190-RhoGAP proteins are delivered (2) to focal adhesion at the rear of cells (3), thus reducing anchoring and limiting activity of the Rho-ROCK axis (4); this could moderate actin polymerization (5); and good tail retraction results on, which contributes to a coordinated back-to-front movement of cell; (lower schema) in Tau-depleted cells, microtubules are less stabilized and weakly organized in bundles in the back of cells (1); the delivery of FAK and p190-RhoGAP proteins to the back of cells is defected (3); high anchoring on focal adhesion and removal of the restrictive activity of the Rho-ROCK pathways (4) leads to excessive actin polymerization (5); and bad tail retraction is observed.
In addition, it is worth mentioning that Tau can also directly bind to actin as proven by many studies [99,100,101,102,103]. In this regard, Tau exhibits a similar binding mode to microtubules and actin filaments. Indeed, using reconstituted high-viscosity networks of microtubules and actin filaments from purified tubulin, actin, and Tau, Farias et al. reported that Tau can interact with each polymer individually by MTBD regions [104]. Moreover, the ability of Tau to bundle and crosslink actin is regulated by phosphorylation at its serine and tyrosine residues in the PRD region [105,106,107]. As mentioned above, studies by He et al. demonstrated, with Tau mutants having the PRD region but lacking the MTBD region, that Tau can induce the assembly of monomeric actin to filamentous actin polymers and promote F-actin bundling [53]. Thus, the way in which Tau governs the crosstalk between microtubules and actin cytoskeletons remains an unsolved puzzle, with the latest studies showing that both MTBD and PRD regions of Tau are important for their interaction with actin.
3.4. Tau and Angiogenesis
Angiogenesis is defined as the formation of new blood vessels from pre-existing ones. This process involves the migration, proliferation, and differentiation of endothelial cells, which line the inside wall of blood vessels. The involvement of Tau has been mainly explored in tauopathies including AD for which we can assist in active neoangiogenesis [108,109,110,111,112,113,114]. To better understand the role played by Tau in this process, Bennett and coll. made use of the tetracycline-repressible gene expression cassette to deplete P301L-mutated Tau in aged rTg4510 mice [115]. Although Tau expression neither affected vascular remodeling nor vascular barrier functions, the authors evidenced that many genes involved in endothelial senescence and in recruiting leukocytes to the endothelium were upregulated in the microvessels (including, i.e., Serpine1, IL8, ICAM-2 and TIE1). Altogether, these findings show that pathological Tau deteriorates brain microvasculature and increases the expression of senescence genes in endothelial cells, which may contribute to the impairment of cerebral blood flow in AD.
Excessive abnormal angiogenesis plays a crucial role in cancer, particularly for solid tumors in which there is an ever-greater nutrient and oxygen demand [116]. Neoangiogenesis triggered by tissue hypoxia and the progression of neoplastic lesions occurs in several key steps [117,118,119,120]. The initially avascular tumor cell mass, upon reaching a critical size, cannot ensure its further growth. Hypoxic cells of the tumor mass, via the secretion of proangiogenic factors such as vascular endothelial growth factor (VEGF), Angiopoietin-1 and 2 (Ang1/2), or platelet-derived growth factor (PDGF), initiate angiogenic switch by stimulating peripheral endothelial cells of parenchymal tissue microvasculature. To our knowledge, the only study showing the involvement of Tau in cancer angiogenesis is the collaborative work led by Prof. Sanchez-Gomez on IDH mutant gliomas [65]. Here, the authors first showed that the expression of Tau depends on the genetic status of IDH1/2 and that it hinders the progression of the tumors. Then and more interestingly, they found that Tau protein inhibits angiogenesis and favors vascular normalization in gliomas expressing wild-type EGFR. The mechanism behind this inhibition is shown to be a result of Tau’s opposition to the EGFR–NF-κB–TAZ signaling pathway which favors the secretion of vascular components (αSMA and CD248). By blocking this signaling pathway, Tau could impede the plasticity of the tumor cells and their capacity to generate mesenchymal-pericyte-like cells necessary for tumor progression. However, an aberrant vasculature is observed when Tau or when EGFR are mutated. The relationship between Tau and EGFR proteins will be further examined when we discuss the RTK-Ras/MAPK/ERK signaling pathway.
4. Tau in Glioblastoma
A limited understanding of the underlying biology of GBM contributes to the lack of clinical advances in its therapy. For two decades, many studies were conducted to establish gene expression profiles of GBMs which permitted its grouping into several subclasses reflecting patients’ survival outcomes [121,122,123]. In 2010, Verhaak et al. analyzed The Cancer Genome Atlas (TCGA) Research Network data [124] and identified four molecular subtypes of GBMs based on distinct gene expression signatures reflecting the cell types of origin: classical (from astrocytic cells), mesenchymal (from astroglial cells), proneural (from oligodendrocytes), and neural (from neural cells) [123]. Recently, Berman’s team examined 543 cases of GBM from the TCGA, and the authors were able to shed light on core pathway alterations responsible for the occurrence and the progression of gliomas: EGFR, IDH1, NF1, PDGFRA, PIK3R1, PIK3CA, PTEN, RB1, and p53 [125]. The fact that Tau can be a ligand of many kinases and transcription factors drives us to discuss their impact on GBM progression. Table 1 below will overview Tau’s interactions with partner signaling pathways and their effects on cellular responses.

Table 1.
Overview of Tau interactions with important effectors, kinases, and transcription factors, involved in altered signaling pathways and their effects on cellular responses in glioma/glioblastoma. The involvement of Tau protein in each signaling pathway is indicated in “Tau status” column, as being “up-stream” or “downstream” of the pathway. n.d: not determined; n.a.: not applicable; ?: pathway, effector, or Tau status not specified in referred studies.
4.1. Tau and Altered Pathways in Glioblastoma
4.1.1. The Receptor Tyrosine Kinases Signaling Pathways
A large group of genes in all eukaryotes encodes for membrane-spanning surface receptors, among them we have one large family supplying intrinsic protein tyrosine kinase activity, the receptor tyrosine kinases (RTKs). RTKs regulate proliferation, survival, differentiation, migration, and angiogenesis through the activation of two major downstream pathways Ras/MAPK/ERK and PI3K/AKT. The function of RTKs is to catalyze the transfer of the γ phosphate of ATP to hydroxyl groups of tyrosine residues on target proteins which serve as docking sites for cytoplasmic signaling effectors. Numerous reviews focus on six major tyrosine kinase receptors; the epidermal growth factor receptor (EGFR), the vascular endothelial growth factor receptor (VEGFR), the platelet-derived growth factor receptor (PDGFR), the hepatocyte growth factor receptor (HGFR/c-MET), the fibroblast growth factor receptor (FGFR) and the insulin-like growth factor 1 receptor (IGF-1R) (see also well-described reviews [147,148,149]). Alteration of these receptors is commonly observed and at least one RTK was found altered in 67.3% of GBM; the EGFR in 57.4% of GBM, PDGFR in 13.1% of GBM, HGFR/c-MET in 1.6% of GBM and FGFR in 3.2% of GBM [125]. Additionally, the RTK/Ras/MAPK/ERK and RTK/PI3K/AKT pathways were found to be altered in more than eight out of ten patients [125]. However, the importance of the Ras protein and downstream protein signaling remains unclear to the pathogenicity of GBM. Apart from GBM, Makino and coll. recently conducted a multiplexed sequencing on 242 glioma tumors for mutations of IDH1/2, H3F3A, HIST1H3B, and TERT genes concomitantly to RAS isotypes [150]. Overall, they found very little correlation between RAS mutations and these other genes, as well as no clear association, was identified between RAS mutations and the histological phenotype of glioma. The accurate diagnosis of GBM involving Ras signaling needs further discussion.
Tau and the RTK/Ras/MAPK/ERK Pathway
Mitogen-activated protein kinases (MAPKs) are highly conserved serine/threonine protein kinases that could be directly activated by extracellular stimuli, typically including oxidative stress, growth factors, and pro-inflammatory factors [151,152]. MAPK includes ERK1/2, p38, and JNK. They can induce many cellular responses, such as proliferation, invasiveness, differentiation, and autophagy [153,154]. The role of abnormal activation of the RTK/Ras/MAPK/ERK pathway through genetic alterations in cancer has been extensively investigated [151]. In GBM, the specific gain-of-function mutation of EGFR, and more specifically the constitutive activation of EGRF variant type III (EGFRvIII), is considered as a driver mutation to activate the MAPK/ERK pathway which consequently leads to tumorigenesis [155,156,157].
Gargini et al. recently characterized the effects of the high expression of Tau protein in a subset of gliomas, especially in IDH mutant tumors, where it seemed to impair tumor aggressiveness [65]. Based on in silico gene analysis, it was found that the EGFR/MAPK pathway was co-upregulated with Tau. Using mice orthotopically injected with RG1 cells expressing the IDH1 gene or its inactive variant IDH1R132H, the authors also showed that IDH1R132H tumors were less aggressive and expressed more Tau protein in comparison to IDH wild-type tumors. More interestingly, they also revealed an inverse correlation between the Tau levels and phosphorylated EGFR in tumors. Note that no difference was observed for mice developing tumors expressing EGFRvIII and/or Tau protein. Thus, these results strongly suggest a negative effect of IDH and Tau on the EGFR/MAPK/ERK pathway. On downstream of the signaling pathway, the authors demonstrated that Tau blocked the mesenchymalization of EGFR glioma cells by inhibiting NF-κB phosphorylation and by reducing the amount of TAZ protein. Altogether, their results indicated that Tau expression can induce changes in the glioma phenotype, through the regulation of the EGFR/TAZ/NF-κB pathway. However, once again, the transposition of these conclusions from gliomas to GBM cannot be completed so simply, as these two models present numerous histopathological, genotypic, and protein differences.
Tau and the RTK/PI3K/AKT Pathway
The RTK/PI3K/AKT pathway is activated by transmembrane tyrosine kinase growth factor receptors, transmembrane integrins, and G-protein-coupled receptors. Upon activation of these receptors, the phosphoinositide 3 kinase (PI3K) translocates to the plasma membrane and produces phosphatidylinositol 3,4,5-triphosphate (PIP3) from phosphatidylinositol bisphosphate (PIP2) [158,159,160]. A cascade of phosphorylation follows, first for AKT (at threonine 308 and serine 473), then the downstream effectors of PI3K/AKT signaling, mTOR kinase. Moreover, AKT indirectly inhibits apoptosis by phosphorylating and inactivating the pro-apoptotic protein GSK3 [161]. Interestingly, Tau is one of the well-described targets of mTOR and GSK3 [162,163].
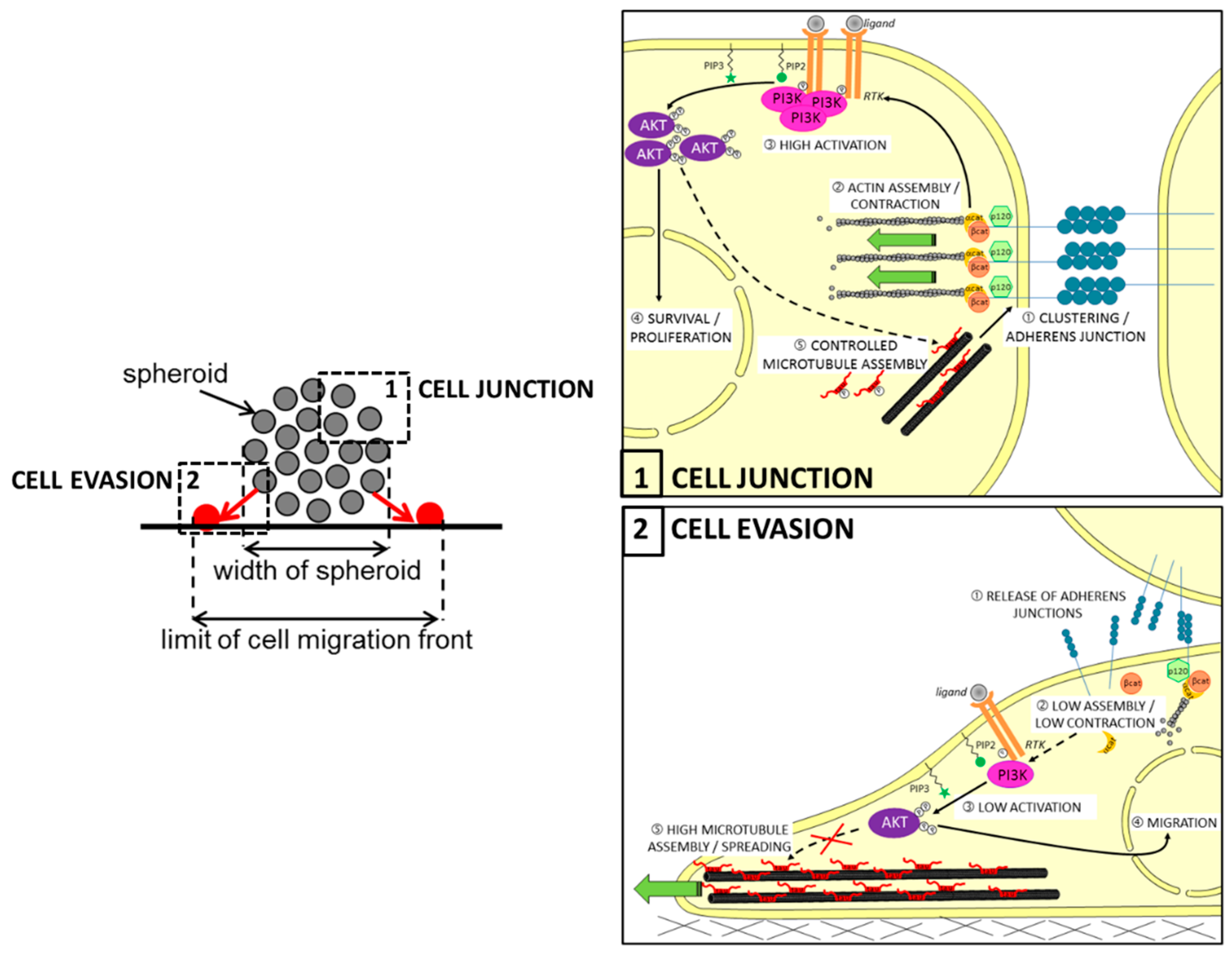
It has been previously reported that Tau can interact with some of the components of the RTK/PI3K/AKT pathway. For example, several in vitro studies showed that Tau can associate with the regulatory subunit p85 of PI3K through the SH3 domain of the subunit [14,128]. Although the consequences were not explored here, it is tempting to think that direct Tau-to-PI3K interaction places Tau in the pathway upstream, as previously described for prostate cancer [127]. In this regard, Pagano et al. recently conducted in silico analysis of reverse phase protein arrays (RPPA) data from a cohort of GBM patients (TCGA-GBM dataset). They found a significantly lower level of activated AKT, notably phosphorylated on serine 473 (pAKTSer473) in human GBM expressing low levels of Tau RNA (MAPT gene, with p-value = 0.0078). The authors confirmed this result both by performing a PathScan® intracellular signaling array and then by immunoblotting on cell lysates. Moreover, treatment of Tau-expressing U87MG cells by the PI3K inhibitor LY294002 significantly reduced the growth of multi-cellular spheroids (MCS), while the AKT inhibitor perifosine mainly disrupted their evasion. As expected, both growths of MCSs and evasions of Tau-depleted cells from MCS were not significantly affected by the two treatments. More interestingly, the depletion of Tau reduced paracellular permeability into MCS originating from the mislocation of the N-cadherin protein at the cell membrane. Altogether, these findings showed for the first time that Tau is an upstream regulator of the PI3K/AKT signaling, likely through the remodeling of microtubules favoring the correct delivery of N-cadherin into the membrane; hence, the AKT activity is necessary for Tau-dependent cell migration during the EMT, while the upstream PI3K activity is rather involved in the control of growth and survival of GBM cell tumors (see also Figure 3).
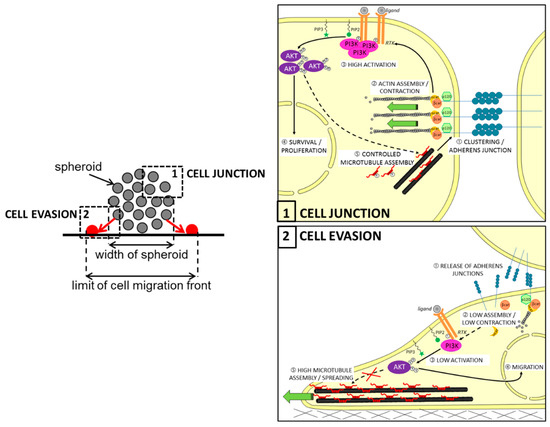
Figure 3.
Proposed model for the role of Tau in regulating multicellular spheroid organization, growth, and migration via the PI3K-AKT signaling pathway in glioblastoma. Upper left panel: principle of cell evasion and cell junction in spheroid: in the zone (1), the N-cadherin junctions maintain cell-cell adhesion necessary for the wholeness of spheroid; in the zone (2), cells closed to the support leave spheroid to migrate individually (in red). Upper right panel: in the context of PTEN-null mutated cells, Tau could contribute by microtubule-dependent function to (1) stabilize the N-cadherin/β-catenin complexes in clusters to reinforce cell–cell adherens and to (2) promote the actin assembly required for cell contraction; Tau promotes indirectly activations cascade of the RTK/PI3K-AKT signaling pathway (3), resulting in survival and proliferation of cells (4); also, active AKT may control the microtubule assembly by increasing Tau phosphorylation (5). Lower left panel: following the adhesion of cells to the support, the release of adherens junctions further promotes cell detachment from spheroid (1); this results in reduced assembly of actin and weak contraction of the cell (2), then low activation of the RTK/PI3K/AKT pathway (3); active AKT is regulating genes involved in cell migration (4), as well as Tau-promoted assembly of microtubules contributes to cell spreading. β-cat: β-catenin; α-cat: α-catenin; PIP2: phosphatidyl-inositol 4,5; PIP3: phosphatidylinositol-3,4,5-trisphosphate.
Tau and PTEN to Regulate the RTK/PI3K/AKT Pathway
The tumor suppressor gene PTEN (phosphatase and tensin homolog deleted on chromosome 10) is the second most frequently mutated gene after P53 in many human sporadic and hereditary cancers [164,165]. PTEN antagonizes the PI3K/AKT pathway by dephosphorylating PIP3 to PIP2. The lipid phosphatase activity of PTEN is critical for its tumor-suppressor function. Indeed, it has been reported that PTEN-null mice die at early embryonic stages, and heterozygous PTEN knockout mice develop numerous tumors [166,167,168].
The role of Tau protein in the PTEN/PI3K/AKT pathway has been poorly explored in cancers [169,170,171], and at most it has been reported that Tau expression is inversely correlated to PTEN mutation/deletion in various cancers including gliomas [65]. In GBM, PTEN mutation or deletion is observed in more than 40% of patients, making it one of the most frequent genomic events of the disease [164,172,173,174,175]. Its dominant-negative effect is seen in the constitutive activation of the PI3K/AKT signaling pathway. In this context, recent results on PTEN-deleted U87MG cells showed that the depletion of Tau was associated with reduced AKT activation, as demonstrated with low pAKTSer473 [129]. Again, Pagano et al. also found a positive association between pAKTSer473 and MAPT RNA expression in the publicly available TCGA human GBM datasets, suggesting that Tau contributes to AKT activation in a context of a loss of function of PTEN. In agreement with these results, a recent publication revealed a positive association between increased phosphorylation of AKT on Ser473 in patient tissue samples and progression from anaplastic astrocytoma to GBM [130]. Despite all these observations, the mechanism of action relating to Tau and PTEN remains unidentified. In opposition, the relationship between Tau and PTEN has been extensively examined in AD pathogenesis [176,177,178,179,180]. Indeed, it has been observed that a significant loss of PTEN correlates with a dramatic increase in the concentration of phosphorylated Tau at Ser214 in the PRD region in NFTs [179]. Mechanistically, PTEN downregulates AKT, which in turn activates the Glycogen synthase kinase 3 beta (GSK3β) phosphorylating Tau at multiple sites in vitro and in vivo [181,182,183,184]. Altogether, it would be interesting to examine if the pAKTSer473-Tau-PTEN triad may define a prognostic marker of poor GBM outcome.
4.1.2. The Src Family Kinases Signaling Pathways
Src designates a family of proto-oncogenes encoding a family of non-receptor tyrosine kinases, called Src family kinases (SFK) (see [185,186,187] for details). c-Src was the first protein kinase to be described as capable of phosphorylating tyrosine residues [188]. SFKs interact with multiple cell surface receptors including integrin and RTKs such as EGFR, PDGFR, and VEGFR [189]. They are activated rapidly upon receptor activation, and they participate in the transduction and regulation of signaling events involving cell adhesion, migration, invasion, proliferation, apoptosis, and angiogenesis [185]. Src has been quite well-explored in the context of cancer, notably for its impact on PI3K/Akt/mTOR, MAPK, and PDGF signaling pathways [190,191]. In addition, it has been reported that six members of SFKs (Fyn, c-Src, Yes, Lyn, Lck, Hck) are expressed in gliomas and actively contribute to the malignancy of tumors [192,193,194,195,196,197]. The following section will talk about the role of two major SFKs, Src and Fyn, in GBM, and will focus especially on their interaction with Tau.
Tau and Src Protein
In GBM, Src protein is neither overexpressed nor mutated but it is highly hyperactivated compared to normal brain tissue leading to the aberrant transduction of its signaling cascades [124,194]. In addition, the role of Src in GBM invasion is demonstrated in transgenic mouse models where GBM invasion was significantly decreased in Src knockout mice compared with control mice, suggesting that Src expression by normal brain tissue is required for invasion and infiltration of GBM cells [198]. Moreover, the hyperactivation of Src in GBM also contributes to processes such as inflammation and metabolism, which help in the establishment of the tumor microenvironment and its development [199]. Overall, these data could indicate that the hyperactivation of Src is mainly a consequence of the aberrant activity of other signaling pathways, i.e., RTKs as suggested by [200,201].
As with all SFKs, Src consists of SH domains that include the SH3 domain. This domain can recognize the proline-rich region of proteins such as the PXXP motif of the PRD region of Tau, as it has already been demonstrated experimentally both in vitro and in cells [54,202]. A potential consequence of the SH3 interaction is the upregulation of tyrosine kinase activity. Indeed, Lee’s team demonstrated in PDGF-stimulated fibroblasts that Tau appeared to prime Src for activation following PDGF stimulation, as reflected by changes in Src-mediated actin rearrangements [99]. In addition, Tau-expressing cells showed sustained actin breakdown, even in the absence of PDGF stimulation, unlike cells that lacked Tau protein. These observations indicate that the interaction of Tau with Src may serve as a mechanism for coupling extracellular signals to the actin cytoskeletal system. Interestingly, the authors have demonstrated that microtubule association by Tau was not required for the observed changes in actin morphology. These findings are interesting because they suggest that targeting Tau can mediate the function of the protein, and thus it can give a prognostic advantage for GBM patients.
Tau and Fyn Protein
In GBM, Fyn was found to be overexpressed in patient samples relative to both normal brain and all other tumors, with varying patterns of expression [192,203]. This kinase has also been shown to be downstream of the commonly mutated RTKs in GBM, such as EGFR, PDGFR, and c-MET [144]. In addition, genetic and pharmacologic inhibitions of Fyn block EGFR-dependent motility and tumor growth both in vitro and in vivo [192]. Recently, Lowenstein’s team evaluated the role of Fyn using genetically engineered glioma mice models (GEMMs) [204]. The team clearly demonstrated that Fyn knockdown reduced tumor progression and significantly increased survival in diverse immune-competent GEMMs of glioma. More interestingly, examination of glioma immune infiltrates displayed a reduction in the amount and in the activity of immune suppressive myeloid-derived cells in the Fyn glioma tumor microenvironment. These findings suggest that targeting Fyn in GBM may help both in reducing tumor cell proliferation and in making the immune microenvironment more engaged in tumor striking.
Interestingly, Tau protein also interacts with Fyn through the SH3 domain. Curiously, little or no study was interested in the relationship between Fyn and Tau in a cancer context. In the 1990s, many studies demonstrated that the pattern of Fyn expression in the brain correlates well with that of Tau and that the two proteins are localized in the growth cones of neurons [145,146,205,206,207]. They are also expressed in oligodendrocytes where they participate in myelinogenesis [208,209]. For these reasons, it would be interesting to further explore the role of Tau and Fyn in GBM tumorigenesis.
4.1.3. The p53 Signaling Pathway
The transcription factor, p53, is critical for many important cellular functions involving genome integrity, such as cell cycle control, DNA damage response, and apoptosis. The p53 protein is encoded by the TP53 gene located on chromosome 17p13.1. In cancer, p53 is considered a broad suppressor factor suppressing over 2500 genes involved in tumorigenesis and tumor invasion. Under normal physiological conditions, and in response to diverse stress signals such as DNA damage, p53 induces DNA repair, cell cycle arrest, and cell apoptosis. Thus, an inactivating mutation in the TP53 gene or its negative regulation results in tumorigenesis. Most GBM patients (about 87% of cases) and cell models (about 94%) exhibit dysregulation of the p53/ARF/Mdm2 pathway [124,210]. Based on multidimensional and comprehensive characterization of more than 500 GBMs, Brennan and coll. described the landscape of somatic genomic alterations in which mutation or deletion of TP53 is identified in about 28% of cases, amplification of the negative p53 regulator Mdm2/4 in 15%, and/or deletion or mutation of its own negative regulator CDKN2A in 58% of patients [125]. Moreover, dysregulated p53 pathway components contribute to cell invasion, migration, proliferation, evasion of apoptosis, and cancer cell stemness of GBM (see the well-detailed review [211]).
The relationship between Tau and p53 has been mainly explored in neurodegenerative diseases [212,213,214,215]. Indeed, it has been observed that the levels of p53 are increased in the AD brain, which is also accompanied by an accumulation of hyperphosphorylated Tau (see review [216]). Meanwhile, no direct relationship between Tau and p53 has so far been clearly demonstrated in GBM. However, Tau is necessary to transport p53 into the nucleus in response to DNA damage. In this context, Giannakakou and coll. demonstrated in numerous tumor cell lines that the trafficking of both wild-type and mutant p53 was granted by a functional microtubule network since cell treatment with microtubule-disrupting agents, vincristine, and taxol, drastically reduced nuclear accumulation of p53 [132]. Another way, through which Tau may be implicated in this pathway is by modulating p53 and/or other p53-pathway components. To support this statement, we can cite a recent study demonstrating the effects of Tau on the p53 function [133]. The authors showed in the neuroblastoma cell model that Tau downregulation impacted p53 stability which affected cell fate by increasing cellular senescence and by decreasing apoptosis. They also showed that Tau modulates the DNA damage response by post-translational modification of p53 and Mdm2 [133].
Altogether, it is tempting to think that Tau could be involved in the suppressive role of p53, both directly and indirectly most likely due to its microtubule-stabilizing function. In addition, the Tau protein can interact with many other protein kinases with significant roles in gliomagenesis. The following chapters list some of these proteins and their roles in GBM.
4.2. Other Altered Kinase Activity Characterized in Tauopathies
4.2.1. The Glycogen Synthase Kinase 3 Signaling Pathway
Glycogen synthase kinase-3 (GSK3) is an unusual serine/threonine kinase involved in diverse cellular functions ranging from glucose metabolism and energy homeostasis to proliferation and apoptosis [217]. GSK3 is considered of some interest because it has some unconventional characteristics for a kinase: being constitutively active, its substrates usually need to be pre-phosphorylated by another kinase, and it is inhibited, rather than activated, in response to stimulation of the two main signaling pathways known to impinge on GSK3, (1) the PI3K/AKT-dependent pathway that is triggered by insulin and growth factors, and (2) the Wnt signaling pathway that is required for embryonic development. Major findings focus on the fact that GSK3, particularly the beta isoform of GSK3 (GSK3β), is a key kinase contributing to abnormal phosphorylation of Tau in NFTs in AD [218,219,220].
Multiple studies have tried to identify the role of this kinase in the glioma and GBM progression [134,221,222,223], and its relationship with Tau can so far only be assumed by circumstantial evidence. In fact, GSK3β could impact Tau via 14-3-3 proteins which constitute a family of highly conserved and ubiquitously expressed small phosphoserine-binding proteins [224]. This family of proteins has seven members which can interact with multiple phospho-proteins. The isoform ζ was reported to be overexpressed and to play a role in GBM tumorigenesis [135,136]. Interestingly, 14-3-3 ζ mediates Tau hyperphosphorylation by multiple kinases [137,138], including active S9-phosphorylated GSK3β kinase. Mechanistically, 14-3-3 ζ promotes the stabilization of the Tau-GSK3β protein complex [139,140]. Another way by which 14-3-3 could interfere with Tau was reported by Joo and coll. who evidenced that overexpression of the σ isoform promotes Tau detachment from microtubules in neurons [225]. Overall, we should further examine how GSK3β/14-3-3 could contribute to GBM tumorigenesis by acting on the microtubule-stabilizing function of Tau.
4.2.2. The Cyclin-Dependent Kinase 5 Signaling Pathway
Cyclin-dependent kinases (CDKs) are a family of serine/threonine protein kinases first discovered for their role in regulating the cell cycle, but are also involved in regulating transcription, mRNA processing, and the differentiation of nerve cells (see book of [226]). CDKs require cyclins for their activity as cell cycle regulators in proliferating cells.
CDK5, however, is an unconventional member, because (i) it does not participate in cell cycle progression, and (ii) it is activated by the binding of its special regulatory subunits p35 or p39 instead of cyclin like the other CDKs. Indeed, CDK5 plays an important role in nervous system development [227,228,229], as well as in angiogenesis, apoptosis, myogenesis, vesicular transport, and senescence in non-neuronal cells [230,231,232,233,234]. In addition, CDK5 dysregulation has been associated with neurodegenerative disorders but its contribution to cancer has been, until recently, overlooked mostly because of the absence of any mutation in tumors. It is now obvious that CDK5 is dysregulated in multiple cancer types, and it needs to receive more attention [235].
In GBM, CDK5 is widely overexpressed and can reach up to 83% higher expression level than normal brain tissue [236]. Additionally, Liu et al. showed that CDK5 promoted GBM cell survival, cell migration, and cell invasion by phosphorylating PIKE-A (PI3K enhancer) and by stimulating its GTPase activity which in turn activates nuclear AKT. Furthermore, CDK5 knock-down in glioma stem cells reduced the self-renewal capacity of GSCs both in vitro and in drosophila xenografts [237] and lead to apoptosis in GBM cell models [238]. Owing to all of this, it is unsurprising that CDK5 is emerging as a potential therapeutic target for GBM, and Tau as a substrate of this important kinase suggests that more attention should be given to these two proteins in this context. Especially since Tau can simultaneously bind to CDK5 and the above-mentioned GSK3β kinase [142]. This binding leads to the forming of a complex in which CDK5 phosphorylates Tau at S235 and primes it for phosphorylation of T231 by GSK3β; similarly, CDK5 primes Tau for the sequential phosphorylation of S400 and S396 by GSK3β by phosphorylating Tau at S404. As in the brains of transgenic mice overexpressing CDK5 activity, CDK5 and GSK3β co-localize with hyperphosphorylated Tau [143]. To note, some forms hyperphosphorylated forms of Tau have already been detected in some colon cancer cells such as HCT116 cells [239], and in some prostate cancer cells [14].
5. Innovative Therapeutic Strategies Targeting Tau Protein for Glioblastoma
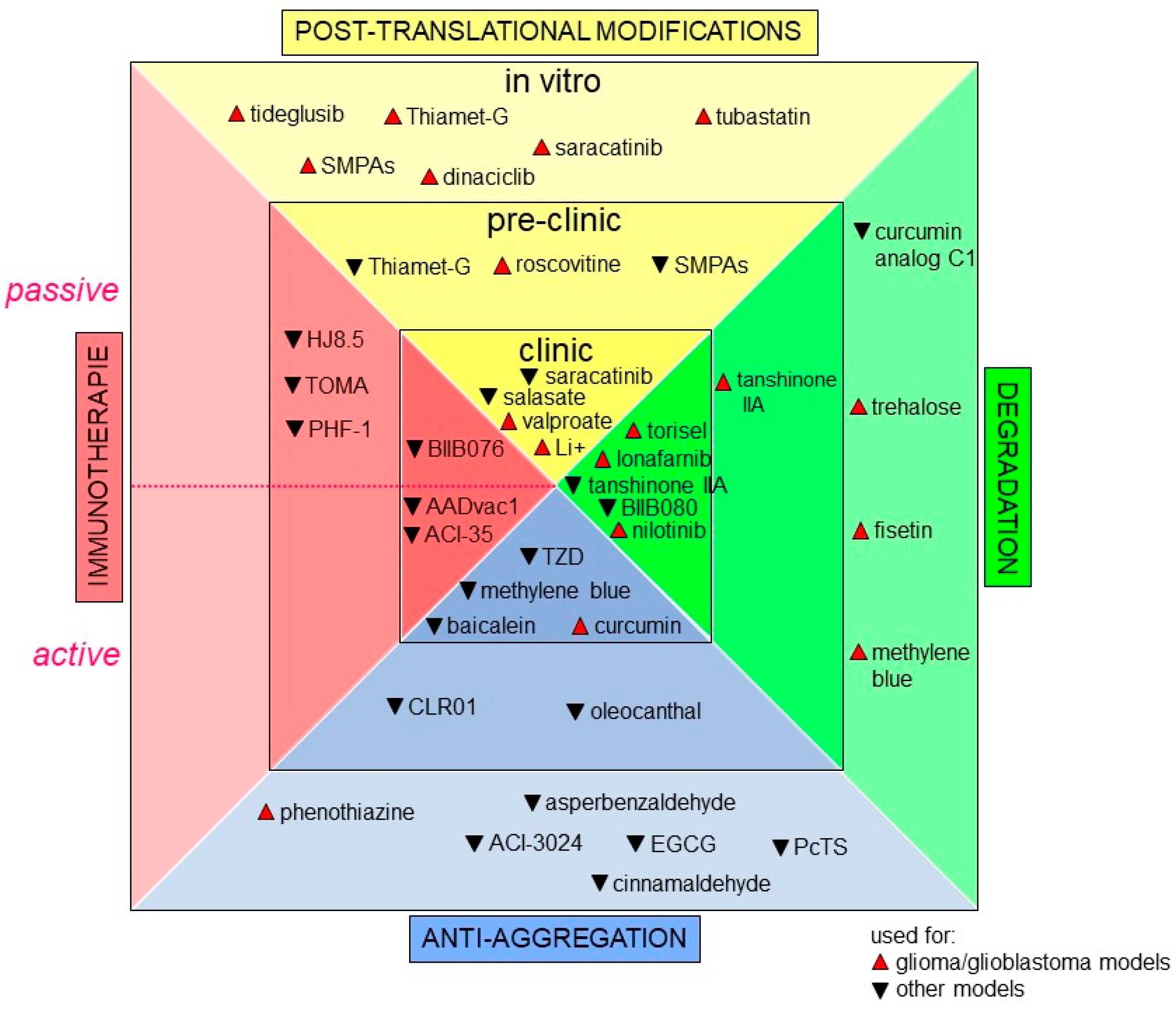
Increasing attention is now being paid to precision medicine. Indeed, scientific and medical communities agree to say that healthcare should be finely tuned to each individual based on molecular profiling instead of the “on drug fits all model” currently used [240,241,242]. In GBM, the current standard therapy, using TMZ, only adds 2.5 months to the median survival of patients [22,243]. Hence, it appears crucial to find druggable cancer-associated proteins. Since Tau protein has a major role in many signaling pathways in GBM, it is tempting to think that targeting this protein would be a good promising therapeutic strategy for GBM and maybe even for other cancer types. In GBM, a major challenge for Tau targeting drug discovery is to maximize the probability that the discovered drug, by either biochemical or phenotypic methods, will lead to clinical efficacy and improved disease management [244,245,246,247,248]. This is even more important since many non-cancer cell types such as neurons also express Tau protein. In this regard, these cells can become targets for cytotoxic drugs, and it is crucial to preserve them during therapy. Moreover, key hallmarks of malignancy are clearly not regulated by a single signaling pathway, as we reviewed some of the above. Unfortunately, drugs targeting the elements of essential and core signaling pathways altered in GBM such as EGFR, IDH1, p53, PI3K and more have failed to transfer into efficient clinical agents [249]. One of the reasons behind these failures could be attributed to the significant heterogeneity of GBM tumors [250,251]. Other pitfalls should be overcome with cell-based models that often do not accurately represent the true in vivo situation. For example, the blood–brain barrier remains relatively impermeant to the majority of GBM marker-targeting drugs (see the well-detailed review [252]). Till now, no drug targeting Tau protein has been designed for cancer treatment per se, but many promising compounds developed as a treatment strategy for tauopathies also deserve our attention [253]. We will now summarize approaches consisting of indirect targeting of Tau protein, such as inducing post-translational modifications; or of direct targeting, such as (i) inhibiting Tau aggregation, (ii) active and passive immunotherapies, and (iii) reducing Tau levels in the cell [254,255]. In this chapter, we will talk about some of Tau targeting approaches as an attempt to provide a starting ground for drug development for GBM and other cancers. Figure 4 below summarizes new promising drugs, as well as drug repurposing, all targeting Tau protein.
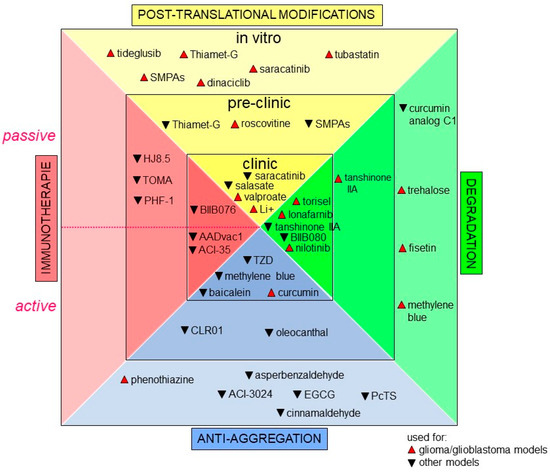

Figure 4.
Tau targeting approaches in glioma/ glioblastoma and other pathologies. Tau targeting agents are grouped according to their targeting approach and to their phase of development, in vitro, preclinical, or clinical. Drugs developed for cancer are in red and denoted with upright triangles. Drugs developed for other conditions are in black with an upside triangle. The dashed line in immunotherapies separates the drugs used for passive immunotherapies (upper half) and active immunotherapies (lower half).
5.1. Playing on Post-Translational Modifications of Tau
In GBM, regulating Tau post-translational modifications could also denote a good strategy to control Tau pathogenicity given that Tau post-translational state of Tau in GBM is studied. The best-described post-modification of Tau is its level of phosphorylation which promotes its aggregation among other pathological defects. In addition to the kinases and the proteins described above, Tau has other interacting proteins that deserve further examination since they were also proven to have an important role in GBM. Among them, we mention the HSP70 chaperone system protein which mediates ubiquitinylation of aberrant Tau species for selective elimination [256,257], the PP2A protein which promotes the dephosphorylation of Tau [258], the HAT/HDAC proteins which can acetylate/deacetylate Tau [259], or even the OGT/OGA proteins which transfer or remove GlcNAc to Tau. Several compounds targeting this set of Tau-regulating proteins are currently being in trial for GBM to examine their potential therapeutic benefit.
5.1.1. Inhibitors of Tau Hyperphosphorylation
Src Kinase Inhibitor Saractinib
Apart from the usual EGFR/PI3K/AKT pathway, the tyrosine-protein Src kinase family is currently being targeted. Among promising candidates, we cite saracatinib (AZD0530), a small molecule Src and Fyn inhibitor. Initially developed for AD treatment, Yun and coll. recently evaluated, in vitro and in vivo, the beneficial effect of the combination of saracatinib and radiation on GBM-derived cells U251 [260]. They observed a saracatinib-induced radiosensitization of cells. This was associated with the persistence of radiation-induced γH2AX foci, with no specific cell cycle and mitotic index changes. Interestingly, the authors showed that treatment with saracatinib also blocked the EGFR/PI3K/AKT pathway activation induced by radiation. As the U251 cell model displays a quite high expression of Tau [106], it is tempting to suppose that a reduction in the phosphorylation of Tau induced by the inhibition of the Src pathway (and of the EGFR/PI3K/AKT one) may represent a novel good therapeutic approach of GBMs, though it remains to verify Tau phosphorylation level in these cells.
GSK3β Kinase Inhibitors
A second interesting Tau kinase is GSK3β which is overexpressed in GBM as mentioned above. Its suppression in vitro has been shown to induce apoptosis of cancer cells. In this sense, many chemical GSK3β inhibitors have been explored, initially to reverse Tau pathogenicity in AD such as lithium (Li+), valproate (VPA), and tideglusib (TDG) (see the in-depth review of [261]).
Concerning Li+, Nowicki and coll. reported by experimental evidence that Li+ potently and reversibly blocked glioma cell migration and invasion in all tested cell lines [223]. Indeed, Li+ caused the retraction of the long protrusions at the front of the migrating cell, which was fully reversible upon withdrawal of treatment. Moreover, the authors demonstrated that the degree of GSK3β inhibition was inversely correlated with the degree of invasion, suggesting that GSK3β inhibition directly impacts cytoskeleton-regulated migration of glioma cells. In addition, it is important to note that significant effects of Li+ on glioma migration were observed at high concentrations of around 5 mM, a dose with severe toxic effects in a clinical context.
In addition, the clinical effectiveness of VPA in adjuvant GBM treatment has yet to be entirely investigated, and especially due to the lack of randomized prospective phase III trials that show indication of progression-free or overall survival benefit [262,263]. To better understand the interest of VPA in GBM therapy, the work of Hu et al. provides some experimental evidence [264]. Using AD transgenic mice as well as human neuroblastoma SH-SY5Y cells, they determined that VPA inhibited both the activities of GSK3β and CDK5 kinases, and consequently reduced Tau hyperphosphorylation.
Concerning TDG, no clinical trial is underway to our knowledge to examine the beneficial effects of this GSK3β inhibitor in cancer therapy. However, few in vitro results deserve our attention [265,266]. By combining TDG with X-ray radiation treatment on 2D and 3D cultured GBM-derived U251 and U118 cells, Fares and Abou-Kheir’s teams found reduced cell proliferation, cell viability, and migration in a dose- and time-dependent manner [267]. These results suggest strongly that TDG may serve as a potential adjuvant radiotherapeutic agent to better target GBM tumors. Given our limited progress in improving GBM survival, we suggest that the repositioning of these drugs, initially explored for AD, deserves attention. Therefore, it would be interesting to hold them to clinical efficacy demonstration for GBM.
CDK Inhibitors
CDK inhibitors (CDKi) are developed with great caution. Some of them are in pre-clinical and clinical trials, particularly those targeting CDK5 [268,269,270,271]. In this context, the best-described CDK5 inhibitor is roscovitine (seliciclib), a small molecule that inhibits CDKs through direct competition at the ATP-binding site. It is a broad-range purine inhibitor of CDK5, as well as CDK2/7. Roscovitine is widely used as a biological tool in cell cycle, cancer, apoptosis, and neurobiology studies and even clinical trials using roscovitine have been conducted on patients with cancer [268]. Using the RG2 rat glioma model, Yakisich and coll. found that roscovitine caused a high decrease in glioma cell proliferation coupled with a strong inhibition of DNA synthesis. The cytotoxic activity of roscovitine could have originated from both an early effect on the DNA replication machinery and a late effect on cell cycle progression. Here, it is important to remember that CDK5 inhibition enhances Tau phosphorylation by activating GSK3β. Therefore, it is tempting to think that CDKi-induced Tau phosphorylation could also contribute to defected cell cycle progression. On the other hand, to our knowledge, none of these studies shows a direct correlation between the anti-tumor activity of roscovitine and the expression and/or phosphorylation of Tau, which deserves to be examined.
5.1.2. Compounds Stimulating Tau Dephosphorylation: Case of PP2A Activation
The serine/threonine protein phosphatase 2A (PP2A) is the most active enzyme in dephosphorylating Tau protein [258]. Therefore, stimulating PP2A activity may represent an alternative strategy to kinase inhibition. In this topic, Sangodkar and coll. reported that improved small-molecule activators of PP2A (SMAPs), synthetic tricyclic sulfonamide, were able to inhibit the growth of KRAS-mutant lung cancer in mice xenografts and mice transgenic models through binding to the PP2A Aα scaffold subunit driving by such activation of the protein [272]. Hence, SMAPs can directly activate PP2A, leading to cell death and lung tumor suppression by inhibiting MAPK signaling pathway. Recently, the same research team demonstrated clearly that the two SMAPs DBK-1154 and DBK-1160 decreased Tau phosphorylation in two cell models [273]. These data reveal that pharmacological PP2A activation may be a novel therapeutic strategy for cancer treatment in general and for GBM therapy in particular.
5.1.3. Compounds Regulating Tau Acetylation/Deacetylation: HAT/HDAC Proteins
Epigenetic regulation of genes plays a key role in numerous disorders. In this process, histone acetylation controls the ease with which transcription factors can access DNA to regulate gene expression. Two classes of enzymes, histone acetyltransferases (HAT) and histone deacetylases (HDAC), control histone acetylation and deacetylation, respectively [274]. In GBM, many HAT mutations such as those of lysine acetyltransferase MOZ and MORF proteins have been reported [275]. To our knowledge, no study in oncology has reported that Tau can be acetylated by HAT proteins. However, an indirect relationship between the two proteins could occur through p53 which has been identified as a MOZ/MORF non-histone protein acetylation substrate [276]. Recently, a clinicians and researchers’ consortium study found that two MOZ and MORF inhibitors, WM-8014 and WM-1119, showed promise for lymphoma therapy [277]. As oligomeric Tau can be a substrate for p53 [214], it should be interesting to deeply examine the beneficial effect of HAT inhibitors as a new therapeutic option in GBM.
An alternative approach for the broad, yet selective, acetylation of Tau protein by HAT is the inhibition of HDAC activity. In the past two decades, in vitro studies have shown significant promise in HDAC inhibitors (HDACis) that could synergize with other compounds in cancer treatment (see the reviews by [278,279]). This could provide a rationale to apply these synergistic ways in GBM as well. Recently, Perez and coll. demonstrated that low concentrations of an HDAC6 inhibitor, vorinostat (SAHA) induced changes in microtubule cytoskeleton properties in U87MG cells [280]. Here, they focused on End-protein 1 (EB1), a MAP stabilizing GTP cap on microtubule plus ends. They found that micromolar concentrations of SAHA caused a strong decrease in EB1 expression. More interestingly, they evidenced that SAHA also induced an increase in tubulin acetylation, a marker of the stabilized microtubule, as well as reduced cell migration. To note, this U87MG cell model exhibits a strong expression level of Tau protein [98,129]. Furthermore, the association of HDAC6 with Tau protein has already been evidenced and it was shown to occur through HDAC6 deacetylase and ZnF UBP domains. The consequence of this association consists of a reduction in HDAC6 deacetylase activity on α-tubulin [259,281]. Moreover, Tau acetylation at lysine residues in KXGS motifs can promote Tau hyperphosphorylation and favors its accumulation [282,283]. As suggested by Honoré’s team [280], low doses of SAHA could represent an interesting therapeutic option for GBM, especially in patients with EB1 and/or Tau overexpressing tumors.
5.1.4. Inhibitors of Tau O-GlcNAcetylation
O-GlcNAcylation refers to the posttranslational modification of O-linkage of N-acetyl-glucosamine (GlcNAc) moieties to serine and threonine residues on proteins. This is regulated by the O-GlcNAc transferase (OGT) and the O-GlcNAcase (OGA), which transfers GlcNAc to and removes GlcNAc from proteins, respectively. The newly emerged OGA selective inhibitor thiamet-G has shown its potential benefit in the therapy of Tau hyperphosphorylation-associated neurodegenerative disorders [284]. In cancers, Ding and coll. reported that thiamet-G significantly sensitized diverse human leukemia cell lines to paclitaxel, with an approximate 10-fold leftward shift of IC50. Interestingly, they observed that paclitaxel combined with thiamet-G resulted in more profound perturbations on microtubule stability than did either one alone. One of the explanations could be that the O-GlcAcetylation of Tau promotes its microtubule-stabilizing function by preventing its hyperphosphorylation [285]. These findings thus suggest that OGA inhibitors such as thiamet-G may serve as a potential adjuvant chemotherapeutic agent for the treatment of GBM.
5.2. Reducing Tau Levels in the Cell
The rationale behind reducing Tau quantity in cancer cells stems from the contribution of this protein to tumorigenesis, as exposed above. Two strategies, in addition to antibodies use, were explored to reduce Tau levels, (i) directly inhibiting Tau protein expression with antisense oligonucleotides (ASO), or (ii) inducing Tau clearance.
5.2.1. Antisense Oligonucleotides Assayed in AD That Could Be Used in GBM
Although there are many ASO designed for AD treatment purposes, only BIIB080 also known as IONIS-MAPTRx or ISIS 814907 has made it into clinical trials for AD as the first and only ASO targeting Tau expression (NCT03186989). This ASO was designed with some modifications to improve its nuclease resistance, increase its cellular uptake, and to induce RNase H-mediated degradation of Tau mRNA as described in a review by the authors [286]. The preclinical study showed that this ASO not only reduces human Tau levels in PS19 mice, but it also decreases Tau mRNA and protein expression in non-human primate Cynomolgus monkeys after intrathecal administration [287]. The primary results of phase I/II clinical trials for AD based on this ASO are to be expected in 2022 (NCT03186989). It could be interesting to explore this approach for cancers, especially in GBM.
5.2.2. Tau Clearance by the Autophagy-Lysosomal System
This strategy consists of stimulating Tau protein degradation mechanisms. In addition to the ubiquitination-proteasome system [288], targeting the autophagy–lysosome system (ALS) could represent an interesting therapeutic avenue [289,290,291,292]. ALS includes three major autophagy pathways, macroautophagy, microautophagy, and chaperone-mediated autophagy (CMA), all leading to lysosome degradation of proteins and cargos [293]. Overall, all modifications of Tau, such as mutation [290,291,292,293,294] and/or post-translational modification such as acetylation [283], impair its correct clearance by this pathway and interferes with global lysosomal functioning. This leads to Tau accumulation inside the cell. Interestingly, in AD and related tauopathies and even in GBM, CMA is dysfunctional [295]. CMA involves Heat shock cognate 70 kDa protein (Hsc70), a member of the Heat shock protein 70 family (Hsp70), that identifies and escorts substrates to the lysosome for degradation [296]. In this way, Jinwal and coll. found that Hsc70 rapidly engages Tau after microtubule destabilization and increases the accumulation of the protein within the cell [297]. Moreover, they showed that cell treatment with methylene blue, well-described to inhibit Hsc70 ATPase activity [290], leads to better Tau degradation.
Hsc70 inhibitors aside, numerous compounds natural or synthetic have been reported to increase Tau clearance in cells such as EA-1 [298], Li+ [299], Trehalose [300], the mTor inhibitor temsirolimus (torisel©) [301], nilotinib [302], lonafarnib [303], curcumin analog C1 [304], the flavonoid Fisetin [305], Flubendazole (fluvermal©) [306,307], Bromhexine [308], tanshinone IIA [309], and other molecules via activation of one or multiple Tau degradation pathways. Some of these cited compounds are already in phase 2 clinical trials (see the review of [310]).
5.3. Therapies Primarily Developed for Tauopathies
5.3.1. Modulation of Tau Aggregation
Likewise, in the regulation of Tau post-translational modifications, the lack of knowledge on the Tau aggregation state in GBM makes it hard to assess the potential efficiency of aggregation inhibitors for treatment. In contrary to neurons, Tau oligomers accumulation in cells was not toxic such as in prostate cancer but they sensitized them to radiotherapy [95]. Alternatively, we can also propose that modulators of Tau aggregation may sequester monomeric Tau making it unavailable for other interactions; therefore, this approach might show some benefit in GBM therapy. Interestingly, Gandini and coll. recently adopted a new strategy that consisted of targeting both Tau aggregation and phosphorylation by GSK3β at the same time and identified some thiazolidinedione derivatives that had this potential in the AD model [311]. Molecules that have dual actions on Tau and major kinases may be the most suitable to be used for GBM.
Regarding the search for new Tau aggregation modulators, the structure–activity studies carried out in the context of neurodegenerative diseases showed that Tau protein is prone to aberrant conformational changes that cause its aggregation and the formation of neurotoxic PHFs. Therefore, these inhibitors can act directly on Tau aggregates dissociation, as well as prevent the formation of Tau aggregates. It is this last propriety that may be a potential therapeutic strategy for GBM tumors. These inhibitors can be grouped into two classes, covalent and non-covalent Tau aggregation inhibitors [312]. The covalent Tau aggregation inhibitors act mainly on monomeric Tau by covalently and directly modifying it, but they can also act on non-monomeric Tau proteins by covalently altering bonds between or within non-monomeric Tau proteins [312]. Examples include many chemical compounds with aldehyde groups such as oleocanthal, cinnamaldehyde, and asperbenzaldehyde, some flavonoids such as baicalein, some molecules that belong to the aminothienopyridazines family, and finally some phenothiazines such as methylene blue. Moreover, non-covalent Tau aggregation inhibitors are structurally more diverse; they can act on different Tau species and have various modes of action which range from transiently interacting with Tau and depressing its aggregation by blocking lysine residues or by sequestering the protein. Examples of this group include: phthalocyanine tetrasulfonate (PcTS; [313]), phenothiazines, triarylmethines, curcumin, and some of the derivatives such as PE859 and some molecular tweezers such as CLR01 among other small or large molecules [314,315].
5.3.2. Immunotherapies against Tau Protein
Tau immunotherapies have advanced from proof-of-concept studies to over a dozen clinical trials for AD and other tauopathies, of which there are ten clinical trials on passive immunotherapy and two on active immunotherapy (see well-detailed reviews in [316,317,318]). Briefly, passive immunotherapy consists of antibody administration which will lead to Tau protein clearance, while active immunotherapy is a vaccine developed against Tau protein to establish immunity in patients. Although we can fairly say that the field is still in its infancy for cancer, this interesting approach exhibits some interesting aspects, as we will discuss below.
Passive Immunotherapy
As mentioned above, the purpose of passive Tau immunotherapy is to neutralize pathogenic Tau forms from the brain to reduce its neurotoxicity. Evidently, this type of immunotherapy has mainly been explored in neuropathologies such as AD. To date, several antibodies have been developed to target monomeric or aggregated Tau, phospho-specific Tau, conformational forms of Tau, or a combination of more than one Tau form. As for their mode of action, some antibodies can only act extracellularly where they sequester extracellular Tau and prevent its aggregation or sequester Tau aggregates and promote their phagocytosis by microglia, preventing the propagation of Tauopathy to other neurons [319,320,321]. Other antibodies can act both extracellularly, in a similar manner to the previous ones, and intracellularly by sequestering cytosolic Tau; they promote its degradation by addressing aggregates to the lysosome system [322,323,324]. In the clinic, most of the trials are ongoing and their detailed outcome has yet to be published. Although worth mentioning, the implementation of this therapeutic strategy to GBM may seem implausible. Additionally, and as mentioned before, the lack of knowledge on Tau forms found in GBM makes it hard to predict which antibody may be more suitable for use.
Active Immunotherapy
The attractive side of this approach is that vaccines would induce autologous antibody production by patients without generating antidrug antibodies. Nevertheless, vaccines are not without issues themselves, off-target immune response presents a big concern for Tau vaccine development [325]. This means that the choice and the criteria of choice of the Tau fragment to be used as a vaccine should be very well studied. Regardless, if Tau vaccines show to be effective, we can imagine a preventive usage of vaccines for people more susceptible to developing tauopathies. Indeed, active Tau immunotherapy has been exclusively examined in AD therapy with two ongoing clinical trials [317]. The first Tau-directed vaccine tested in clinical trials was AADvac1, developed by Axon Neuroscience SE. These antibodies are expected to prevent Tau protein aggregating, facilitate the removal of Tau aggregates, and prevent the spreading of the pathology, slowing or halting the progression of the disease. The Tau fragment used corresponds to 151–391 residues of the 4R isoform and hence, the antibody recognizes an epitope in the microtubule-binding domain repeat region. The second phospho-Tau-specific vaccine is ACI-35 which was initially developed by AC Immune and later licensed by Janssen. Here, the Tau fragment used corresponds to 393–408 residues in the C-terminal end of the protein. This fragment includes several phosphorylated serine residues (S396/S404) known to be pathologically phosphorylated by GSK3 kinases in AD. In the future, when more research is completed in this context, we can reassess the benefit of using Tau vaccines for GBM.
6. Conclusions and Perspective
Through this review, we explained how Tau could be involved in cancer and particularly in GBM. Firstly, we showed how this protein can be implicated in cancer by exploring its role in various cancer hallmarks, and then we discussed some of the pathways highly altered in GBM and how Tau protein can be involved in them. We also mentioned some of the Tau interactors that have demonstrated roles in GBM. Lastly, we summarized some of the innovative therapeutic approaches targeting Tau for the treatment of GBM. Through the evidence presented above, we believe that this area of research is worth exploring and we hope that this review will inspire researchers to investigate the role of Tau protein in this type of cancer.
7. Limitations of This Review
One of the clear limitations of this review is the lack of research completed on Tau in GBMs and most of the arguments made are also based on results of research completed on other types of cancer or through the known functions of Tau that can be relevant in this topic.
Author Contributions
Conceptualization, G.B.; writing—original draft preparation, R.H. and G.B.; writing—review and editing, H.K., A.P., V.P., M.R. and F.D.; supervision, G.B.; All authors have read and agreed to the published version of the manuscript.
Funding
This study received no funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
This study was partly supported by research funding from the Canceropôle Provence Alpes Côte d’Azur, Association pour la Recherche sur les Tumeurs Cérébrales (ARTC-Sud), Institut National du Cancer and Région Sud and R.H. received a student grant from the Algerian ministry of higher education and scientific research.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Mitchison, T.; Kirschner, M. Dynamic Instability of Microtubule Growth. Nature 1984, 312, 237–242. [Google Scholar] [CrossRef] [PubMed]
- Weingarten, M.D.; Lockwood, A.H.; Hwo, S.Y.; Kirschner, M.W. A Protein Factor Essential for Microtubule Assembly. Proc. Natl. Acad. Sci. USA 1975, 72, 1858–1862. [Google Scholar] [CrossRef] [PubMed]
- Drubin, D.G.; Kirschner, M.W. Tau Protein Function in Living Cells. J. Cell Biol. 1986, 103, 2739–2746. [Google Scholar] [CrossRef] [PubMed]
- Cleveland, D.W.; Hwo, S.Y.; Kirschner, M.W. Purification of Tau, a Microtubule-Associated Protein That Induces Assembly of Microtubules from Purified Tubulin. J. Mol. Biol. 1977, 116, 207–225. [Google Scholar] [CrossRef]
- Lee, G.; Rook, S.L. Expression of Tau Protein in Non-Neuronal Cells: Microtubule Binding and Stabilization. J. Cell Sci. 1992, 102, 227–237. [Google Scholar] [CrossRef]
- Drewes, G.; Ebneth, A.; Preuss, U.; Mandelkow, E.-M.; Mandelkow, E. MARK, a Novel Family of Protein Kinases That Phosphorylate Microtubule-Associated Proteins and Trigger Microtubule Disruption. Cell 1997, 89, 297–308. [Google Scholar] [CrossRef]
- Bunker, J.M.; Wilson, L.; Jordan, M.A.; Feinstein, S.C. Modulation of Microtubule Dynamics by Tau in Living Cells: Implications for Development and Neurodegeneration. Mol. Biol. Cell 2004, 15, 2720–2728. [Google Scholar] [CrossRef]
- Feinstein, S.C.; Wilson, L. Inability of Tau to Properly Regulate Neuronal Microtubule Dynamics: A Loss-of-Function Mechanism by Which Tau Might Mediate Neuronal Cell Death. Biochim. Biophys. Acta 2005, 1739, 268–279. [Google Scholar] [CrossRef]
- LeBoeuf, A.C.; Levy, S.F.; Gaylord, M.; Bhattacharya, A.; Singh, A.K.; Jordan, M.A.; Wilson, L.; Feinstein, S.C. FTDP-17 mutations in Tau alter the regulation of microtubule dynamics: An “alternative core” model for normal and pathological Tau action. J. Biol. Chem. 2008, 283, 36406–36415. [Google Scholar] [CrossRef]
- Huang, L.C.; Ye, J.C.; Hsieh, C.H.; Chen, L.M.; Lin, T.Y.; Hung, Y.C.; Chang, W.C. PTEN, Tau-AP-3, Thymidylate Synthase Immunohistochemistry Scoring Expression in Patients with Uterine Leiomyomas, Uterine Smooth Muscle Tumors of Uncertain Malignancy Potential and Uterine Leiomyosarcomas. Eur. J. Gynaecol. Oncol. 2011, 32, 496–499. [Google Scholar]
- Zaatiti, H.; Abdallah, J.; Nasr, Z.; Khazen, G.; Sandler, A.; Abou-Antoun, T.J. Tumorigenic Proteins Upregulated in the MYCN-Amplified IMR-32 Human Neuroblastoma Cells Promote Proliferation and Migration. Int. J. Oncol. 2018, 52, 787–803. [Google Scholar] [CrossRef] [PubMed]
- Wagner, P.; Wang, B.; Clark, E.; Lee, H.; Rouzier, R.; Pusztai, L. Microtubule Associated Protein (MAP)-Tau: A Novel Mediator of Paclitaxel Sensitivity In Vitro and In Vivo. Cell Cycle 2005, 4, 1149–1152. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Zhang, Z.; Sun, L.; Fang, Y.; Xu, X.; Zhou, G. MiR-186 Regulates Chemo-Sensitivity to Paclitaxel via Targeting MAPT in Non-Small Cell Lung Cancer (NSCLC). Mol. Biosyst. 2016, 12, 3417–3424. [Google Scholar] [CrossRef]
- Souter, S.; Lee, G. Microtubule-Associated Protein Tau In Human Prostate Cancer Cells: Isoforms, Phosphorylation, And Interactions. J. Cell. Biochem. 2009, 108, 555–564. [Google Scholar] [CrossRef]
- Miyazono, M.; Iwaki, T.; Kitamoto, T.; Shin, R.-W.; Fukui, M.; Tateishi, J. Widespread Distribution of Tau in the Astrocytic Elements of Glial Tumors. Acta Neuropathol. 1993, 86, 236–241. [Google Scholar] [CrossRef] [PubMed]
- Rouzier, R.; Rajan, R.; Wagner, P.; Hess, K.R.; Gold, D.L.; Stec, J.; Ayers, M.; Ross, J.S.; Zhang, P.; Buchholz, T.A.; et al. Microtubule-Associated Protein Tau: A Marker of Paclitaxel Sensitivity in Breast Cancer. Proc. Natl. Acad. Sci. USA 2005, 102, 8315–8320. [Google Scholar] [CrossRef]
- Pusztai, L.; Jeong, J.-H.; Gong, Y.; Ross, J.S.; Kim, C.; Paik, S.; Rouzier, R.; Andre, F.; Hortobagyi, G.N.; Wolmark, N.; et al. Evaluation of Microtubule-Associated Protein-Tau Expression As a Prognostic and Predictive Marker in the NSABP-B 28 Randomized Clinical Trial. J. Clin. Oncol. 2009, 27, 4287–4292. [Google Scholar] [CrossRef] [PubMed]
- Lara-Velazquez, M.; Al-Kharboosh, R.; Jeanneret, S.; Vazquez-Ramos, C.; Mahato, D.; Tavanaiepour, D.; Rahmathulla, G.; Quinones-Hinojosa, A. Advances in Brain Tumor Surgery for Glioblastoma in Adults. Brain Sci. 2017, 7, 166. [Google Scholar] [CrossRef]
- De Vleeschouwer, S.; Bergers, G. Glioblastoma: To Target the Tumor Cell or the Microenvironment? In Glioblastoma; De Vleeschouwer, S., Ed.; Codon Publications: Brisbane, Australia, 2017; pp. 315–340. ISBN 978-0-9944381-2-6. [Google Scholar]
- Figarella-Branger, D.; Chappe, C.; Padovani, L.; Mercurio, S.; Colin, C.; Forest, F.; Bouvier, C. Glial and glioneuronal tumors in adults and children: Main genetic alterations and towards a histomolecular classification. Bull. Cancer 2013, 100, 715–726. [Google Scholar] [CrossRef]
- Eramo, A.; Ricci-Vitiani, L.; Zeuner, A.; Pallini, R.; Lotti, F.; Sette, G.; Pilozzi, E.; Larocca, L.M.; Peschle, C.; De Maria, R. Chemotherapy Resistance of Glioblastoma Stem Cells. Cell Death Differ. 2006, 13, 1238–1241. [Google Scholar] [CrossRef]
- Stupp, R.; Mason, W.P.; van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.B.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus Concomitant and Adjuvant Temozolomide for Glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef] [PubMed]
- Alifieris, C.; Trafalis, D.T. Glioblastoma Multiforme: Pathogenesis and Treatment. Pharmacol. Ther. 2015, 152, 63–82. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Liang, R.F.; Wang, X.; Mao, Q.; Liu, Y.H. BKM120 Sensitizes C6 Glioma Cells to Temozolomide via Suppression of the PI3K/Akt/NF-κB/MGMT Signaling Pathway. Oncol. Lett. 2017, 14, 6597–6603. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gao, Y.-T.; Chen, X.-B.; Liu, H.-L. Up-Regulation of MiR-370-3p Restores Glioblastoma Multiforme Sensitivity to Temozolomide by Influencing MGMT Expression. Sci. Rep. 2016, 6, 32972. [Google Scholar] [CrossRef] [PubMed]
- Couchie, D.; Fages, C.; Bridoux, A.M.; Rolland, B.; Tardy, M.; Nunez, J. Microtubule-Associated Proteins and in Vitro Astrocyte Differentiation. J. Cell Biol. 1985, 101, 2095–2103. [Google Scholar] [CrossRef] [PubMed]
- Goedert, M.; Spillantini, M.G.; Jakes, R.; Rutherford, D.; Crowther, R.A. Multiple Isoforms of Human Microtubule-Associated Protein Tau: Sequences and Localization in Neurofibrillary Tangles of Alzheimer’s Disease. Neuron 1989, 3, 519–526. [Google Scholar] [CrossRef]
- Goode, B.L.; Feinstein, S.C. Identification of a Novel Microtubule Binding and Assembly Domain in the Developmentally Regulated Inter-Repeat Region of Tau. J. Cell Biol. 1994, 124, 769–782. [Google Scholar] [CrossRef]
- Gustke, N.; Trinczek, B.; Biernat, J.; Mandelkow, E.M.; Mandelkow, E. Domains of Tau Protein and Interactions with Microtubules. Biochemistry 1994, 33, 9511–9522. [Google Scholar] [CrossRef]
- Pedersen, J.T.; Sigurdsson, E.M. Tau Immunotherapy for Alzheimer’s Disease. Trends Mol. Med. 2015, 21, 394–402. [Google Scholar] [CrossRef]
- Goedert, M.; Spillantini, M.G.; Potier, M.C.; Ulrich, J.; Crowther, R.A. Cloning and Sequencing of the CDNA Encoding an Isoform of Microtubule-Associated Protein Tau Containing Four Tandem Repeats: Differential Expression of Tau Protein MRNAs in Human Brain. EMBO J. 1989, 8, 393–399. [Google Scholar] [CrossRef]
- Ittner, L.M.; Götz, J. Amyloid-β and Tau—A Toxic Pas de Deux in Alzheimer’s Disease. Nat. Rev. Neurosci. 2011, 12, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Kolarova, M.; García-Sierra, F.; Bartos, A.; Ricny, J.; Ripova, D. Structure and Pathology of Tau Protein in Alzheimer Disease. Int. J. Alzheimers Dis. 2012, 2012, 731526. [Google Scholar] [CrossRef] [PubMed]
- Martin, L.; Latypova, X.; Terro, F. Post-Translational Modifications of Tau Protein: Implications for Alzheimer’s Disease. Neurochem. Int. 2011, 58, 458–471. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Lu, S.X.; Ouyang, X.; Melchor, J.; Lee, J.; Terracina, G.; Wang, X.; Hyde, L.; Hess, J.F.; Parker, E.M.; et al. Analysis of Tau Post-Translational Modifications in RTg4510 Mice, a Model of Tau Pathology. Mol. Neurodegener. 2015, 10, 14. [Google Scholar] [CrossRef] [PubMed]
- Goedert, M.; Jakes, R. Expression of Separate Isoforms of Human Tau Protein: Correlation with the Tau Pattern in Brain and Effects on Tubulin Polymerization. EMBO J. 1990, 9, 4225–4230. [Google Scholar] [CrossRef] [PubMed]
- Kosik, K.S.; Orecchio, L.D.; Bakalis, S.; Neve, R.L. Developmentally Regulated Expression of Specific Tau Sequences. Neuron 1989, 2, 1389–1397. [Google Scholar] [CrossRef]
- Avila, J.; Jiménez, J.S.; Sayas, C.L.; Bolós, M.; Zabala, J.C.; Rivas, G.; Hernández, F. Tau Structures. Front. Aging Neurosci. 2016, 8, 262. [Google Scholar] [CrossRef]
- Cleveland, D.W.; Hwo, S.Y.; Kirschner, M.W. Physical and Chemical Properties of Purified Tau Factor and the Role of Tau in Microtubule Assembly. J. Mol. Biol. 1977, 116, 227–247. [Google Scholar] [CrossRef]
- Woody, R.W.; Clark, D.C.; Roberts, G.C.K.; Martin, S.R.; Bayley, P.M. Molecular Flexibility in Microtubule Proteins: Proton Nuclear Magnetic Resonance Characterization. Biochemistry 1983, 22, 2186–2192. [Google Scholar] [CrossRef]
- Schweers, O.; Schönbrunn-Hanebeck, E.; Marx, A.; Mandelkow, E. Structural Studies of Tau Protein and Alzheimer Paired Helical Filaments Show No Evidence for Beta-Structure. J. Biol. Chem. 1994, 269, 24290–24297. [Google Scholar] [CrossRef]
- Jeganathan, S.; Chinnathambi, S.; Mandelkow, E.-M.; Mandelkow, E. Conformations of Microtubule-Associated Protein Tau Mapped by Fluorescence Resonance Energy Transfer. In Amyloid Proteins: Methods and Protocols; Sigurdsson, E.M., Calero, M., Gasset, M., Eds.; Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2012; pp. 85–99. ISBN 978-1-61779-551-0. [Google Scholar]
- De Bessa, T.; Breuzard, G.; Allegro, D.; Devred, F.; Peyrot, V.; Barbier, P. Tau Interaction with Tubulin and Microtubules: From Purified Proteins to Cells. In Tau Protein: Methods and Protocols; Smet-Nocca, C., Ed.; Methods in Molecular Biology; Springer: New York, NY, USA, 2017; pp. 61–85. ISBN 978-1-4939-6598-4. [Google Scholar]
- Di Maïo, I.L.; Barbier, P.; Allegro, D.; Brault, C.; Peyrot, V. Quantitative Analysis of Tau-Microtubule Interaction Using FRET. Int. J. Mol. Sci. 2014, 15, 14697–14714. [Google Scholar] [CrossRef] [PubMed]
- Qi, H.; Cantrelle, F.-X.; Benhelli-Mokrani, H.; Smet-Nocca, C.; Buée, L.; Lippens, G.; Bonnefoy, E.; Galas, M.-C.; Landrieu, I. Nuclear Magnetic Resonance Spectroscopy Characterization of Interaction of Tau with DNA and Its Regulation by Phosphorylation. Biochemistry 2015, 54, 1525–1533. [Google Scholar] [CrossRef] [PubMed]
- Avila, J.; León-Espinosa, G.; García, E.; García-Escudero, V.; Hernández, F.; DeFelipe, J. Tau Phosphorylation by GSK3 in Different Conditions. Int. J. Alzheimers Dis. 2012, 2012, e578373. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Song, X.; Nisbet, R.; Götz, J. Co-immunoprecipitation with Tau Isoform-specific Antibodies Reveals Distinct Protein Interactions and Highlights a Putative Role for 2N Tau in Disease. J. Biol. Chem. 2016, 291, 8173–8188. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, A.; Novak, M.; Grundke-Iqbal, I.; Iqbal, K. Regulation of Phosphorylation of Tau by Cyclin-Dependent Kinase 5 and Glycogen Synthase Kinase-3 at Substrate Level. FEBS Lett. 2006, 580, 5925–5933. [Google Scholar] [CrossRef] [PubMed]
- Thiel, G. Synapsin I, Synapsin II, and Synaptophysin: Marker Proteins of Synaptic Vesicles. Brain Pathol. 1993, 3, 87–95. [Google Scholar] [CrossRef]
- Gong, C.-X.; Liu, F.; Grundke-Iqbal, I.; Iqbal, K. Post-Translational Modifications of Tau Protein in Alzheimer’s Disease. J. Neural Transm. 2005, 112, 813–838. [Google Scholar] [CrossRef]
- Maas, T.; Eidenmüller, J.; Brandt, R. Interaction of Tau with the Neural Membrane Cortex Is Regulated by Phosphorylation at Sites That Are Modified in Paired Helical Filaments. J. Biol. Chem. 2000, 275, 15733–15740. [Google Scholar] [CrossRef]
- Yu, J.-Z.; Rasenick, M.M. Tau Associates with Actin in Differentiating PC12 Cells. FASEB J. 2006, 20, 1452–1461. [Google Scholar] [CrossRef]
- He, H.J.; Wang, X.S.; Pan, R.; Wang, D.L.; Liu, M.N.; He, R.Q. The Proline-Rich Domain of Tau Plays a Role in Interactions with Actin. BMC Cell Biol. 2009, 10, 81. [Google Scholar] [CrossRef]
- Lee, G.; Newman, S.T.; Gard, D.L.; Band, H.; Panchamoorthy, G. Tau Interacts with Src-Family Non-Receptor Tyrosine Kinases. J. Cell Sci. 1998, 111, 3167–3177. [Google Scholar] [CrossRef] [PubMed]
- Devred, F.; Douillard, S.; Briand, C.; Peyrot, V. First Tau Repeat Domain Binding to Growing and Taxol-Stabilized Microtubules, and Serine 262 Residue Phosphorylation. FEBS Lett. 2002, 523, 247–251. [Google Scholar] [CrossRef]
- Goode, B.L.; Chau, M.; Denis, P.E.; Feinstein, S.C. Structural and Functional Differences between 3-Repeat and 4-Repeat Tau Isoforms. Implications for Normal Tau Function and the Onset of Neurodegenetative Disease. J. Biol. Chem. 2000, 275, 38182–38189. [Google Scholar] [CrossRef]
- Alonso, A.d.C.; Mederlyova, A.; Novak, M.; Grundke-Iqbal, I.; Iqbal, K. Promotion of Hyperphosphorylation by Frontotemporal Dementia Tau Mutations. J. Biol. Chem. 2004, 279, 34873–34881. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Kim, D.; Ju, S.; Shin, S.; Cho, I.; Park, S.-H.; Grailhe, R.; Lee, C.; Kim, Y.K. Glioblastoma-Secreted Soluble CD44 Activates Tau Pathology in the Brain. Exp. Mol. Med. 2018, 50, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Barbier, P.; Zejneli, O.; Martinho, M.; Lasorsa, A.; Belle, V.; Smet-Nocca, C.; Tsvetkov, P.O.; Devred, F.; Landrieu, I. Role of Tau as a Microtubule-Associated Protein: Structural and Functional Aspects. Front. Aging Neurosci. 2019, 11, 204. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, N.; Shao, G.; Qian, J.; Shen, D.; Fei, Y.; Mao, W.; Wu, D. Relationship between Gastric Cancer Tau Protein Expression and Paclitaxel Sensitivity. Pathol. Oncol. Res. 2013, 19, 429–435. [Google Scholar] [CrossRef] [PubMed]
- Breuzard, G.; Hubert, P.; Nouar, R.; Bessa, T.D.; Devred, F.; Barbier, P.; Sturgis, J.N.; Peyrot, V. Molecular Mechanisms of Tau Binding to Microtubules and Its Role in Microtubule Dynamics in Live Cells. J. Cell Sci. 2013, 126, 2810–2819. [Google Scholar] [CrossRef]
- Mimori, K.; Sadanaga, N.; Yoshikawa, Y.; Ishikawa, K.; Hashimoto, M.; Tanaka, F.; Sasaki, A.; Inoue, H.; Sugimachi, K.; Mori, M. Reduced Tau Expression in Gastric Cancer Can Identify Candidates for Successful Paclitaxel Treatment. Br. J. Cancer 2006, 94, 1894–1897. [Google Scholar] [CrossRef]
- Smoter, M.; Bodnar, L.; Grala, B.; Stec, R.; Zieniuk, K.; Kozlowski, W.; Szczylik, C. Tau Protein as a Potential Predictive Marker in Epithelial Ovarian Cancer Patients Treated with Paclitaxel/Platinum First-Line Chemotherapy. J. Exp. Clin. Cancer Res. 2013, 32, 25. [Google Scholar] [CrossRef]
- Cirak, Y.; Sarsik, B.; Cakar, B.; Sen, S.; Simsir, A.; Uslu, R. Predictive and Prognostic Values of Tau and BubR1 Protein in Prostate Cancer and Their Relationship to the Gleason Score. Med. Oncol. 2013, 30, 526. [Google Scholar] [CrossRef] [PubMed]
- Gargini, R.; Segura-Collar, B.; Herránz, B.; García-Escudero, V.; Romero-Bravo, A.; Núñez, F.J.; García-Pérez, D.; Gutiérrez-Guamán, J.; Ayuso-Sacido, A.; Seoane, J.; et al. The IDH-TAU-EGFR Triad Defines the Neovascular Landscape of Diffuse Gliomas. Sci. Transl. Med. 2020, 12, eaax1501. [Google Scholar] [CrossRef]
- Steffensen, K.D.; Smoter, M.; Waldstrøm, M.; Grala, B.; Bodnar, L.; Stec, R.; Szczylik, C.; Jakobsen, A. Resistance to First Line Platinum Paclitaxel Chemotherapy in Serous Epithelial Ovarian Cancer: The Prediction Value of ERCC1 and Tau Expression. Int. J. Oncol. 2014, 44, 1736–1744. [Google Scholar] [CrossRef]
- Bonneau, C.; Gurard-Levin, Z.A.; Andre, F.; Pusztai, L.; Rouzier, R. Predictive and Prognostic Value of the Tau Protein in Breast Cancer. Anticancer Res. 2015, 35, 5179–5184. [Google Scholar]
- Papin, S.; Paganetti, P. Emerging Evidences for an Implication of the Neurodegeneration-Associated Protein TAU in Cancer. Brain Sci. 2020, 10, 862. [Google Scholar] [CrossRef] [PubMed]
- Cirillo, L.; Gotta, M.; Meraldi, P. The Elephant in the Room: The Role of Microtubules in Cancer. In Cell Division Machinery and Disease; Gotta, M., Meraldi, P., Eds.; Advances in Experimental Medicine and Biology; Springer International Publishing: Cham, Switzerland, 2017; pp. 93–124. ISBN 978-3-319-57127-0. [Google Scholar]
- Langie, S.A.S.; Koppen, G.; Desaulniers, D.; Al-Mulla, F.; Al-Temaimi, R.; Amedei, A.; Azqueta, A.; Bisson, W.H.; Brown, D.; Brunborg, G.; et al. Causes of Genome Instability: The Effect of Low Dose Chemical Exposures in Modern Society. Carcinogenesis 2015, 36, S61–S88. [Google Scholar] [CrossRef] [PubMed]
- Brady, R.M.; Zinkowski, R.P.; Binder, L.I. Presence of Tau in Isolated Nuclei from Human Brain. Neurobiol. Aging 1995, 16, 479–486. [Google Scholar] [CrossRef]
- Greenwood, J.A.; Johnson, G.V.W. Localization and in Situ Phosphorylation State of Nuclear Tau. Exp. Cell Res. 1995, 220, 332–337. [Google Scholar] [CrossRef]
- Frost, B.; Hemberg, M.; Lewis, J.; Feany, M.B. Tau Promotes Neurodegeneration through Global Chromatin Relaxation. Nat. Neurosci. 2014, 17, 357–366. [Google Scholar] [CrossRef]
- Bukar Maina, M.; Al-Hilaly, Y.K.; Serpell, L.C. Nuclear Tau and Its Potential Role in Alzheimer’s Disease. Biomolecules 2016, 6, 9. [Google Scholar] [CrossRef]
- Cross, D.C.; Muñoz, J.P.; Hernández, P.; Maccioni, R.B. Nuclear and Cytoplasmic Tau Proteins from Human Nonneuronal Cells Share Common Structural and Functional Features with Brain Tau. J. Cell. Biochem. 2000, 78, 305–317. [Google Scholar] [CrossRef]
- Loomis, P.A.; Howard, T.H.; Castleberry, R.P.; Binder, L.I. Identification of Nuclear Tau Isoforms in Human Neuroblastoma Cells. Proc. Natl. Acad. Sci. USA 1990, 87, 8422–8426. [Google Scholar] [CrossRef] [PubMed]
- Thurston, V.C.; Zinkowski, R.P.; Binder, L.I. Tau as a Nucleolar Protein in Human Nonneural Cells in Vitro and in Vivo. Chromosoma 1996, 105, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Sultan, A.; Nesslany, F.; Violet, M.; Bégard, S.; Loyens, A.; Talahari, S.; Mansuroglu, Z.; Marzin, D.; Sergeant, N.; Humez, S.; et al. Nuclear Tau, a Key Player in Neuronal DNA Protection. J. Biol. Chem. 2011, 286, 4566–4575. [Google Scholar] [CrossRef] [PubMed]
- Violet, M.; Delattre, L.; Tardivel, M.; Sultan, A.; Chauderlier, A.; Caillierez, R.; Talahari, S.; Nesslany, F.; Lefebvre, B.; Bonnefoy, E.; et al. A Major Role for Tau in Neuronal DNA and RNA Protection in Vivo under Physiological and Hyperthermic Conditions. Front. Cell. Neurosci. 2014, 8, 84. [Google Scholar] [CrossRef] [PubMed]
- Violet, M.; Chauderlier, A.; Delattre, L.; Tardivel, M.; Chouala, M.S.; Sultan, A.; Marciniak, E.; Humez, S.; Binder, L.; Kayed, R.; et al. Prefibrillar Tau Oligomers Alter the Nucleic Acid Protective Function of Tau in Hippocampal Neurons in Vivo. Neurobiol. Dis. 2015, 82, 540–551. [Google Scholar] [CrossRef] [PubMed]
- Hua, Q.; He, R. Tau Could Protect DNA Double Helix Structure. Biochim. Biophys. Acta 2003, 1645, 205–211. [Google Scholar] [CrossRef]
- Rossi, G.; Conconi, D.; Panzeri, E.; Redaelli, S.; Piccoli, E.; Paoletta, L.; Dalprà, L.; Tagliavini, F. Mutations in MAPT Gene Cause Chromosome Instability and Introduce Copy Number Variations Widely in the Genome. J. Alzheimers Dis. 2013, 33, 969–982. [Google Scholar] [CrossRef]
- Migliore, L.; Testa, A.; Scarpato, R.; Pavese, N.; Petrozzi, L.; Bonuccelli, U. Spontaneous and Induced Aneuploidy in Peripheral Blood Lymphocytes of Patients with Alzheimer’s Disease. Hum. Genet. 1997, 101, 299–305. [Google Scholar] [CrossRef]
- Mosch, B.; Morawski, M.; Mittag, A.; Lenz, D.; Tarnok, A.; Arendt, T. Aneuploidy and DNA Replication in the Normal Human Brain and Alzheimer’s Disease. J. Neurosci. 2007, 27, 6859–6867. [Google Scholar] [CrossRef]
- Iourov, I.Y.; Vorsanova, S.G.; Liehr, T.; Yurov, Y.B. Aneuploidy in the Normal, Alzheimer’s Disease and Ataxia-Telangiectasia Brain: Differential Expression and Pathological Meaning. Neurobiol. Dis. 2009, 34, 212–220. [Google Scholar] [CrossRef] [PubMed]
- Yurov, Y.B.; Vorsanova, S.G.; Liehr, T.; Kolotii, A.D.; Iourov, I.Y. X Chromosome Aneuploidy in the Alzheimer’s Disease Brain. Mol. Cytogenet. 2014, 7, 20. [Google Scholar] [CrossRef] [PubMed]
- Pope, W.B.; Lambert, M.P.; Leypold, B.; Seupaul, R.; Sletten, L.; Krafft, G.; Klein, W.L. Microtubule-Associated Protein Tau Is Hyperphosphorylated during Mitosis in the Human Neuroblastoma Cell Line SH-SY5Y. Exp. Neurol. 1994, 126, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Preuss, U.; Döring, F.; Illenberger, S.; Mandelkow, E.M. Cell Cycle-Dependent Phosphorylation and Microtubule Binding of Tau Protein Stably Transfected into Chinese Hamster Ovary Cells. Mol. Biol. Cell. 1995, 6, 1397–1410. [Google Scholar] [CrossRef] [PubMed]
- Flores-Rodríguez, P.; Harrington, C.R.; Wischik, C.M.; Ibarra-Bracamontes, V.; Zarco, N.; Navarrete, A.; Martínez-Maldonado, A.; Guadarrama-Ortíz, P.; Villanueva-Fierro, I.; Ontiveros-Torres, M.A.; et al. Phospho-Tau Protein Expression in the Cell Cycle of SH-SY5Y Neuroblastoma Cells: A Morphological Study. J. Alzheimers Dis. 2019, 71, 631–645. [Google Scholar] [CrossRef]
- Bougé, A.-L.; Parmentier, M.-L. Tau Excess Impairs Mitosis and Kinesin-5 Function, Leading to Aneuploidy and Cell Death. Dis. Model. Mech. 2016, 9, 307–319. [Google Scholar] [CrossRef]
- Giam, M.; Rancati, G. Aneuploidy and Chromosomal Instability in Cancer: A Jackpot to Chaos. Cell Div. 2015, 10, 3. [Google Scholar] [CrossRef]
- Jordan, M.A.; Thrower, D.; Wilson, L. Mechanism of Inhibition of Cell Proliferation by Vinca Alkaloids1. Cancer Res. 1991, 51, 2212–2222. [Google Scholar]
- Jordan, M.A.; Wilson, L. Microtubules as a Target for Anticancer Drugs. Nat. Rev. Cancer 2004, 4, 253–265. [Google Scholar] [CrossRef]
- Spicakova, T.; O’Brien, M.M.; Duran, G.E.; Sweet-Cordero, A.; Sikic, B.I. Expression and Silencing of the Microtubule-Associated Protein Tau in Breast Cancer Cells. Mol. Cancer Ther. 2010, 9, 2970–2981. [Google Scholar] [CrossRef][Green Version]
- Martellucci, S.; Clementi, L.; Sabetta, S.; Muzi, P.; Mattei, V.; Bologna, M.; Angelucci, A. Tau Oligomers Accumulation Sensitizes Prostate Cancer Cells to Docetaxel Treatment. J. Cancer Res. Clin. Oncol. 2021, 147, 1957–1971. [Google Scholar] [CrossRef] [PubMed]
- Nakata, S.; Price, A.; Eberhart, C.; Morris, M. Increased Tau Expression Correlates with IDH Mutation in Infiltrating Gliomas and Impairs Cell Migration. J. Neuropathol. Exp. Neurol. 2020, 79, 493–499. [Google Scholar] [CrossRef]
- Yan, H.; Bigner, D.D.; Velculescu, V.; Parsons, D.W. Mutant Metabolic Enzymes Are at the Origin of Gliomas. Cancer Res. 2009, 69, 9157–9159. [Google Scholar] [CrossRef] [PubMed]
- Breuzard, G.; Pagano, A.; Bastonero, S.; Malesinski, S.; Parat, F.; Barbier, P.; Peyrot, V.; Kovacic, H. Tau Regulates the Microtubule-Dependent Migration of Glioblastoma Cells via the Rho-ROCK Signaling Pathway. J. Cell Sci. 2019, 132, jcs222851. [Google Scholar] [CrossRef] [PubMed]
- Sharma, V.M.; Litersky, J.M.; Bhaskar, K.; Lee, G. Tau Impacts on Growth-Factor-Stimulated Actin Remodeling. J. Cell Sci. 2007, 120, 748–757. [Google Scholar] [CrossRef]
- Griffith, L.M.; Pollard, T.D. Evidence for Actin Filament-Microtubule Interaction Mediated by Microtubule-Associated Proteins. J. Cell Biol. 1978, 78, 958–965. [Google Scholar] [CrossRef]
- Griffith, L.M.; Pollard, T.D. The Interaction of Actin Filaments with Microtubules and Microtubule-Associated Proteins. J. Biol. Chem. 1982, 257, 9143–9151. [Google Scholar] [CrossRef]
- Pollard, T.D.; Selden, S.C.; Maupin, P. Interaction of Actin Filaments with Microtubules. J. Cell Biol. 1984, 99, 33s–37s. [Google Scholar] [CrossRef]
- Selden, S.C.; Pollard, T.D. Interaction of Actin Filaments with Microtubules Is Mediated by Microtubule-Associated Proteins and Regulated by Phosphorylation. Ann. N. Y. Acad. Sci. 1986, 466, 803–812. [Google Scholar] [CrossRef]
- Farias, G.A.; Muñoz, J.P.; Garrido, J.; Maccioni, R.B. Tubulin, Actin, and Tau Protein Interactions and the Study of Their Macromolecular Assemblies. J. Cell. Biochem. 2002, 85, 315–324. [Google Scholar] [CrossRef]
- Kadowaki, T.; Fujita-Yamaguchi, Y.; Nishida, E.; Takaku, F.; Akiyama, T.; Kathuria, S.; Akanuma, Y.; Kasuga, M. Phosphorylation of Tubulin and Microtubule-Associated Proteins by the Purified Insulin Receptor Kinase. J. Biol. Chem. 1985, 260, 4016–4020. [Google Scholar] [CrossRef]
- Hoshi, M.; Nishida, E.; Miyata, Y.; Sakai, H.; Miyoshi, T.; Ogawara, H.; Akiyama, T. Protein Kinase C Phosphorylates Tau and Induces Its Functional Alterations. FEBS Lett. 1987, 217, 237–241. [Google Scholar] [CrossRef]
- Zhu, D.; Kosik, K.S.; Meigs, T.E.; Yanamadala, V.; Denker, B.M. Gα12 Directly Interacts with PP2A: Evidence for gα12-stimulated pp2a phosphatase activity and dephosphorylation of microtubule-associated protein, tau. J. Biol. Chem. 2004, 279, 54983–54986. [Google Scholar] [CrossRef]
- Siedlak, S.L.; Cras, P.; Kawai, M.; Richey, P.; Perry, G. Basic Fibroblast Growth Factor Binding Is a Marker for Extracellular Neurofibrillary Tangles in Alzheimer Disease. J. Histochem. Cytochem. 1991, 39, 899–904. [Google Scholar] [CrossRef] [PubMed]
- Deininger, M.H.; Fimmen, B.A.; Thal, D.R.; Schluesener, H.J.; Meyermann, R. Aberrant Neuronal and Paracellular Deposition of Endostatin in Brains of Patients with Alzheimer’s Disease. J. Neurosci. 2002, 22, 10621–10626. [Google Scholar] [CrossRef]
- Ułamek-Kozioł, M.; Czuczwar, S.J.; Januszewski, S.; Pluta, R. Proteomic and Genomic Changes in Tau Protein, Which Are Associated with Alzheimer’s Disease after Ischemia-Reperfusion Brain Injury. Int. J. Mol. Sci. 2020, 21, 892. [Google Scholar] [CrossRef]
- Jeong, H.; Shin, J.Y.; Lee, K.; Lee, S.-J.; Chong, H.-J.; Jeong, H.; Jeon, Y.-E.; Shin, D.-S.; Jang, S.; Kim, K.H.; et al. Caffeoyl-Prolyl-Histidine Amide Inhibits Fyn and Alleviates Atopic Dermatitis-Like Phenotypes via Suppression of NF-ΚB Activation. Int. J. Mol. Sci. 2020, 21, 7160. [Google Scholar] [CrossRef]
- Merlini, M.; Wanner, D.; Nitsch, R.M. Tau Pathology-Dependent Remodelling of Cerebral Arteries Precedes Alzheimer’s Disease-Related Microvascular Cerebral Amyloid Angiopathy. Acta Neuropathol. 2016, 131, 737–752. [Google Scholar] [CrossRef]
- Castillo-Carranza, D.L.; Nilson, A.N.; Van Skike, C.E.; Jahrling, J.B.; Patel, K.; Garach, P.; Gerson, J.E.; Sengupta, U.; Abisambra, J.; Nelson, P.; et al. Cerebral Microvascular Accumulation of Tau Oligomers in Alzheimer’s Disease and Related Tauopathies. Aging Dis. 2017, 8, 257–266. [Google Scholar] [CrossRef]
- Bennett, R.E.; Robbins, A.B.; Hu, M.; Cao, X.; Betensky, R.A.; Clark, T.; Das, S.; Hyman, B.T. Tau Induces Blood Vessel Abnormalities and Angiogenesis-Related Gene Expression in P301L Transgenic Mice and Human Alzheimer’s Disease. Proc. Natl. Acad. Sci. USA 2018, 115, E1289–E1298. [Google Scholar] [CrossRef]
- Bennett, R.E.; Hu, M.; Fernandes, A.; Perez-Rando, M.; Robbins, A.; Kamath, T.; Dujardin, S.; Hyman, B.T. Tau Reduction in Aged Mice Does Not Impact Microangiopathy. Acta Neuropathol. Commun. 2020, 8, 137. [Google Scholar] [CrossRef] [PubMed]
- Pouysségur, J.; Dayan, F.; Mazure, N.M. Hypoxia Signalling in Cancer and Approaches to Enforce Tumour Regression. Nature 2006, 441, 437–443. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Folkman, J. Patterns and Emerging Mechanisms of the Angiogenic Switch during Tumorigenesis. Cell 1996, 86, 353–364. [Google Scholar] [CrossRef]
- Bergers, G.; Benjamin, L.E. Tumorigenesis and the Angiogenic Switch. Nat. Rev. Cancer 2003, 3, 401–410. [Google Scholar] [CrossRef]
- Ribatti, D.; Nico, B.; Crivellato, E.; Roccaro, A.M.; Vacca, A. The History of the Angiogenic Switch Concept. Leukemia 2007, 21, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Jászai, J.; Schmidt, M.H.H. Trends and Challenges in Tumor Anti-Angiogenic Therapies. Cells 2019, 8, 1102. [Google Scholar] [CrossRef]
- Freije, W.A.; Castro-Vargas, F.E.; Fang, Z.; Horvath, S.; Cloughesy, T.; Liau, L.M.; Mischel, P.S.; Nelson, S.F. Gene Expression Profiling of Gliomas Strongly Predicts Survival. Cancer Res. 2004, 64, 6503–6510. [Google Scholar] [CrossRef]
- Phillips, H.S.; Kharbanda, S.; Chen, R.; Forrest, W.F.; Soriano, R.H.; Wu, T.D.; Misra, A.; Nigro, J.M.; Colman, H.; Soroceanu, L.; et al. Molecular Subclasses of High-Grade Glioma Predict Prognosis, Delineate a Pattern of Disease Progression, and Resemble Stages in Neurogenesis. Cancer Cell 2006, 9, 157–173. [Google Scholar] [CrossRef]
- Verhaak, R.G.W.; Hoadley, K.A.; Purdom, E.; Wang, V.; Qi, Y.; Wilkerson, M.D.; Miller, C.R.; Ding, L.; Golub, T.; Mesirov, J.P.; et al. Integrated Genomic Analysis Identifies Clinically Relevant Subtypes of Glioblastoma Characterized by Abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010, 17, 98–110. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Research Network Comprehensive Genomic Characterization Defines Human Glioblastoma Genes and Core Pathways. Nature 2008, 455, 1061–1068. [CrossRef]
- Brennan, C.W.; Verhaak, R.G.W.; McKenna, A.; Campos, B.; Noushmehr, H.; Salama, S.R.; Zheng, S.; Chakravarty, D.; Sanborn, J.Z.; Berman, S.H.; et al. The Somatic Genomic Landscape of Glioblastoma. Cell 2013, 155, 462–477. [Google Scholar] [CrossRef] [PubMed]
- Murakami, C.; Ikota, H.; Nobusawa, S.; Nakata, S.; Yamazaki, T.; Hashiba, Y.; Hirato, J.; Yokoo, H. Oligodendroglioma Showing Pleomorphic Xanthoastrocytoma-like Perivascular Microlesion: With IDH1, TERT Promoter Mutation and 1p/19q Codeletion Detected in Both Components. Pathol. Int. 2020, 70, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yu, Y.; Liu, W.; Liu, W.; Li, Z.; Wei, Z.; Jiang, R. Microtubule-Associated Protein Tau Is Associated with the Resistance to Docetaxel in Prostate Cancer Cell Lines. Res. Rep. Urol. 2017, 9, 71–77. [Google Scholar] [CrossRef]
- Reynolds, C.H.; Garwood, C.J.; Wray, S.; Price, C.; Kellie, S.; Perera, T.; Zvelebil, M.; Yang, A.; Sheppard, P.W.; Varndell, I.M.; et al. Phosphorylation Regulates Tau Interactions with Src Homology 3 Domains of Phosphatidylinositol 3-Kinase, Phospholipase Cγ1, Grb2, and Src Family Kinases. J. Biol. Chem. 2008, 283, 18177–18186. [Google Scholar] [CrossRef] [PubMed]
- Pagano, A.; Breuzard, G.; Parat, F.; Tchoghandjian, A.; Figarella-Branger, D.; De Bessa, T.C.; Garrouste, F.; Douence, A.; Barbier, P.; Kovacic, H. Tau Regulates Glioblastoma Progression, 3D Cell Organization, Growth and Migration via the PI3K-AKT Axis. Cancers 2021, 13, 5818. [Google Scholar] [CrossRef]
- Wu, Y.-H.; Huang, Y.-F.; Chen, C.-C.; Chou, C.-Y. Akt Inhibitor SC66 Promotes Cell Sensitivity to Cisplatin in Chemoresistant Ovarian Cancer Cells through Inhibition of COL11A1 Expression. Cell Death Dis. 2019, 10, 322. [Google Scholar] [CrossRef]
- Sekino, Y.; Han, X.; Babasaki, T.; Goto, K.; Inoue, S.; Hayashi, T.; Teishima, J.; Shiota, M.; Takeshima, Y.; Yasui, W.; et al. Microtubule-Associated Protein Tau (MAPT) Promotes Bicalutamide Resistance and Is Associated with Survival in Prostate Cancer. Urol. Oncol. Semin. Orig. Investig. 2020, 38, 795.e1–795.e8. [Google Scholar] [CrossRef]
- Giannakakou, P.; Sackett, D.L.; Ward, Y.; Webster, K.R.; Blagosklonny, M.V.; Fojo, T. P53 Is Associated with Cellular Microtubules and Is Transported to the Nucleus by Dynein. Nat. Cell Biol. 2000, 2, 709–717. [Google Scholar] [CrossRef]
- Sola, M.; Magrin, C.; Pedrioli, G.; Pinton, S.; Salvadè, A.; Papin, S.; Paganetti, P. Tau Affects P53 Function and Cell Fate during the DNA Damage Response. Commun. Biol. 2020, 3, 245. [Google Scholar] [CrossRef]
- Miyashita, K.; Kawakami, K.; Nakada, M.; Mai, W.; Shakoori, A.; Fujisawa, H.; Hayashi, Y.; Hamada, J.; Minamoto, T. Potential Therapeutic Effect of Glycogen Synthase Kinase 3beta Inhibition against Human Glioblastoma. Clin. Cancer Res. 2009, 15, 887–897. [Google Scholar] [CrossRef]
- Liang, S.; Shen, G.; Liu, Q.; Xu, Y.; Zhou, L.; Xiao, S.; Xu, Z.; Gong, F.; You, C.; Wei, Y. Isoform-Specific Expression and Characterization of 14-3-3 Proteins in Human Glioma Tissues Discovered by Stable Isotope Labeling with Amino Acids in Cell Culture-Based Proteomic Analysis. Proteom. Clin. Appl. 2009, 3, 743–753. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xu, H.; Wang, Q.; Wang, S.; Xiong, N. 14-3-3ζ Promotes Gliomas Cells Invasion by Regulating Snail through the PI3K/AKT Signaling. Cancer Med. 2019, 8, 783–794. [Google Scholar] [CrossRef] [PubMed]
- Hashiguchi, M.; Sobue, K.; Paudel, H.K. 14-3-3ζ Is an Effector of Tau Protein Phosphorylation. J. Biol. Chem. 2000, 275, 25247–25254. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, H.Y.; Li, T.; MacDonald, R.; Cho, C.M.; Leclerc, N.; Paudel, H.K. Interaction of 14-3-3ζ with Microtubule-Associated Protein Tau within Alzheimer’s Disease Neurofibrillary Tangles. Biochemistry 2013, 52, 6445–6455. [Google Scholar] [CrossRef] [PubMed]
- Agarwal-Mawal, A.; Qureshi, H.Y.; Cafferty, P.W.; Yuan, Z.; Han, D.; Lin, R.; Paudel, H.K. 14-3-3 Connects Glycogen Synthase Kinase-3β to Tau within a Brain Microtubule-Associated Tau Phosphorylation Complex. J. Biol. Chem. 2003, 278, 12722–12728. [Google Scholar] [CrossRef]
- Yuan, Z.; Agarwal-Mawal, A.; Paudel, H.K. 14-3-3 Binds to and Mediates Phosphorylation of Microtubule-Associated Tau Protein by Ser9-Phosphorylated Glycogen Synthase Kinase 3beta in the Brain. J. Biol. Chem. 2004, 279, 26105–26114. [Google Scholar] [CrossRef]
- Martin, D.; Brown-Luedi, M.; Chiquet-Ehrismann, R. Tenascin-C Signaling through Induction of 14-3-3 Tau. J. Cell Biol. 2003, 160, 171–175. [Google Scholar] [CrossRef]
- Li, T.; Hawkes, C.; Qureshi, H.Y.; Kar, S.; Paudel, H.K. Cyclin-Dependent Protein Kinase 5 Primes Microtubule-Associated Protein Tau Site-Specifically for Glycogen Synthase Kinase 3β. Biochemistry 2006, 45, 3134–3145. [Google Scholar] [CrossRef]
- Noble, W.; Olm, V.; Takata, K.; Casey, E.; Mary, O.; Meyerson, J.; Gaynor, K.; LaFrancois, J.; Wang, L.; Kondo, T.; et al. Cdk5 Is a Key Factor in Tau Aggregation and Tangle Formation in Vivo. Neuron 2003, 38, 555–565. [Google Scholar] [CrossRef]
- Qin, A.; Musket, A.; Musich, P.R.; Schweitzer, J.B.; Xie, Q. Receptor Tyrosine Kinases as Druggable Targets in Glioblastoma: Do Signaling Pathways Matter? Neurooncol. Adv. 2021, 3, vdab133. [Google Scholar] [CrossRef]
- Bare, D.J.; Lauder, J.M.; Wilkie, M.B.; Maness, P.F. P59fyn in Rat Brain Is Localized in Developing Axonal Tracts and Subpopulations of Adult Neurons and Glia. Oncogene 1993, 8, 1429–1436. [Google Scholar] [PubMed]
- Kempf, M.; Clement, A.; Faissner, A.; Lee, G.; Brandt, R. Tau Binds to the Distal Axon Early in Development of Polarity in a Microtubule- and Microfilament-Dependent Manner. J. Neurosci. 1996, 16, 5583–5592. [Google Scholar] [CrossRef] [PubMed]
- Tilak, M.; Holborn, J.; New, L.A.; Lalonde, J.; Jones, N. Receptor Tyrosine Kinase Signaling and Targeting in Glioblastoma Multiforme. Int. J. Mol. Sci. 2021, 22, 1831. [Google Scholar] [CrossRef] [PubMed]
- Pearson, J.R.D.; Regad, T. Targeting Cellular Pathways in Glioblastoma Multiforme. Signal Transduct. Target. Ther. 2017, 2, 1–11. [Google Scholar] [CrossRef]
- Schlessinger, J. Cell Signaling by Receptor Tyrosine Kinases. Cell 2000, 103, 211–225. [Google Scholar] [CrossRef]
- Makino, Y.; Arakawa, Y.; Yoshioka, E.; Shofuda, T.; Minamiguchi, S.; Kawauchi, T.; Tanji, M.; Kanematsu, D.; Nonaka, M.; Okita, Y.; et al. Infrequent RAS Mutation Is Not Associated with Specific Histological Phenotype in Gliomas. BMC Cancer 2021, 21, 1025. [Google Scholar] [CrossRef]
- Dhillon, A.S.; Hagan, S.; Rath, O.; Kolch, W. MAP Kinase Signalling Pathways in Cancer. Oncogene 2007, 26, 3279–3290. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Cheng, J.; North, B.J.; Wei, W. Functional Analyses of Major Cancer-Related Signaling Pathways in Alzheimer’s Disease Etiology. Biochim. Biophys. Acta Rev. Cancer 2017, 1868, 341–358. [Google Scholar] [CrossRef]
- Roberts, P.J.; Der, C.J. Targeting the Raf-MEK-ERK Mitogen-Activated Protein Kinase Cascade for the Treatment of Cancer. Oncogene 2007, 26, 3291–3310. [Google Scholar] [CrossRef]
- du Plessis, M.; Davis, T.; Loos, B.; Pretorius, E.; de Villiers, W.J.S.; Engelbrecht, A.M. Molecular Regulation of Autophagy in a Pro-Inflammatory Tumour Microenvironment: New Insight into the Role of Serum Amyloid A. Cytokine Growth Factor Rev. 2021, 59, 71–83. [Google Scholar] [CrossRef]
- Gan, H.K.; Kaye, A.H.; Luwor, R.B. The EGFRvIII Variant in Glioblastoma Multiforme. J. Clin. Neurosci. 2009, 16, 748–754. [Google Scholar] [CrossRef] [PubMed]
- An, Z.; Aksoy, O.; Zheng, T.; Fan, Q.-W.; Weiss, W.A. Epidermal Growth Factor Receptor (EGFR) and EGFRvIII in Glioblastoma (GBM): Signaling Pathways and Targeted Therapies. Oncogene 2018, 37, 1561–1575. [Google Scholar] [CrossRef]
- Chen, S.; Yang, L.; Li, Z.; Zhuo, S.; Yan, B.; Zhang, Z.; Zhang, J.; Feng, H.; Yang, K. EGFR/EGFRvIII Partly Regulates the Tumourigenesis of Glioblastoma through the SOX9-GLUT3 Axis. Am. J. Transl. Res. 2021, 13, 6055–6065. [Google Scholar] [PubMed]
- Shibasaki, F.; Homma, Y.; Takenawa, T. Two Types of Phosphatidylinositol 3-Kinase from Bovine Thymus. Monomer and Heterodimer Form. J. Biol. Chem. 1991, 266, 8108–8114. [Google Scholar] [CrossRef]
- Koyama, S.; Yu, H.; Dalgarno, D.C.; Shin, T.B.; Zydowsky, L.D.; Schreiber, S.L. Structure of the Pl3K SH3 Domain and Analysis of the SH3 Family. Cell 1993, 72, 945–952. [Google Scholar] [CrossRef]
- Franke, T.F.; Kaplan, D.R.; Cantley, L.C. PI3K: Downstream AKTion Blocks Apoptosis. Cell 1997, 88, 435–437. [Google Scholar] [CrossRef]
- Fang, X.; Yu, S.X.; Lu, Y.; Bast, R.C.; Woodgett, J.R.; Mills, G.B. Phosphorylation and Inactivation of Glycogen Synthase Kinase 3 by Protein Kinase A. Proc. Natl. Acad. Sci. USA 2000, 97, 11960–11965. [Google Scholar] [CrossRef]
- Cai, Z.; Chen, G.; He, W.; Xiao, M.; Yan, L.-J. Activation of MTOR: A Culprit of Alzheimer’s Disease? Neuropsychiatr. Dis. Treat. 2015, 11, 1015–1030. [Google Scholar] [CrossRef]
- Morris, G.; Berk, M.; Maes, M.; Puri, B.K. Could Alzheimer’s Disease Originate in the Periphery and If So How So? Mol. Neurobiol. 2019, 56, 406–434. [Google Scholar] [CrossRef]
- Li, J.; Yen, C.; Liaw, D.; Podsypanina, K.; Bose, S.; Wang, S.I.; Puc, J.; Miliaresis, C.; Rodgers, L.; McCombie, R.; et al. PTEN, a Putative Protein Tyrosine Phosphatase Gene Mutated in Human Brain, Breast, and Prostate Cancer. Science 1997, 275, 1943–1947. [Google Scholar] [CrossRef]
- Stokoe, D. PTEN. Curr. Biol. 2001, 11, R502. [Google Scholar] [CrossRef]
- Stambolic, V.; MacPherson, D.; Sas, D.; Lin, Y.; Snow, B.; Jang, Y.; Benchimol, S.; Mak, T.W. Regulation of PTEN Transcription by P53. Mol. Cell 2001, 8, 317–325. [Google Scholar] [CrossRef]
- Suzuki, A.; de la Pompa, J.L.; Stambolic, V.; Elia, A.J.; Sasaki, T.; Barrantes, I.d.B.; Ho, A.; Wakeham, A.; Ltie, A.; Khoo, W.; et al. High Cancer Susceptibility and Embryonic Lethality Associated with Mutation of the PTEN Tumor Suppressor Gene in Mice. Curr. Biol. 1998, 8, 1169–1178. [Google Scholar] [CrossRef]
- Podsypanina, K.; Ellenson, L.H.; Nemes, A.; Gu, J.; Tamura, M.; Yamada, K.M.; Cordon-Cardo, C.; Catoretti, G.; Fisher, P.E.; Parsons, R. Mutation of Pten/Mmac1 in Mice Causes Neoplasia in Multiple Organ Systems. Proc. Natl. Acad. Sci. USA 1999, 96, 1563–1568. [Google Scholar] [CrossRef] [PubMed]
- Bamias, A.; Karina, M.; Papakostas, P.; Kostopoulos, I.; Bobos, M.; Vourli, G.; Samantas, E.; Christodoulou, C.; Pentheroudakis, G.; Pectasides, D.; et al. A Randomized Phase III Study of Adjuvant Platinum/Docetaxel Chemotherapy with or without Radiation Therapy in Patients with Gastric Cancer. Cancer Chemother. Pharmacol. 2010, 65, 1009–1021. [Google Scholar] [CrossRef]
- Schroeder, C.; Grell, J.; Hube-Magg, C.; Kluth, M.; Lang, D.; Simon, R.; Höflmayer, D.; Minner, S.; Burandt, E.; Clauditz, T.S.; et al. Aberrant Expression of the Microtubule-Associated Protein Tau Is an Independent Prognostic Feature in Prostate Cancer. BMC Cancer 2019, 19, 193. [Google Scholar] [CrossRef]
- Koo, H.; Choi, S.W.; Cho, H.J.; Lee, I.-H.; Kong, D.-S.; Seol, H.J.; Lee, J.-I.; Choi, J.-W.; Sa, J.K.; Nam, D.-H. Ethnic Delineation of Primary Glioblastoma Genome. Cancer Med. 2020, 9, 7352–7359. [Google Scholar] [CrossRef]
- Bonneau, D.; Longy, M. Mutations of the Human PTEN Gene. Hum. Mutat. 2000, 16, 109–122. [Google Scholar] [CrossRef]
- Leslie, N.R.; Downes, C.P. PTEN Function: How Normal Cells Control It and Tumour Cells Lose It. Biochem. J. 2004, 382, 1–11. [Google Scholar] [CrossRef]
- Lv, S.; Teugels, E.; Sadones, J.; De Brakeleer, S.; Duerinck, J.; Du Four, S.; Michotte, A.; De Grève, J.; Neyns, B. Correlation of EGFR, IDH1 and PTEN Status with the Outcome of Patients with Recurrent Glioblastoma Treated in a Phase II Clinical Trial with the EGFR-Blocking Monoclonal Antibody Cetuximab. Int. J. Oncol. 2012, 41, 1029–1035. [Google Scholar] [CrossRef]
- Choi, S.W.; Lee, Y.; Shin, K.; Koo, H.; Kim, D.; Sa, J.K.; Cho, H.J.; Shin, H.; Lee, S.J.; Kim, H.; et al. Mutation-Specific Non-Canonical Pathway of PTEN as a Distinct Therapeutic Target for Glioblastoma. Cell Death Dis. 2021, 12, 374. [Google Scholar] [CrossRef]
- Pyo, H.; Lovati, E.; Pasinetti, G.M.; Ksiezak-Reding, H. Phosphorylation of Tau at THR212 and SER214 in Human Neuronal and Glial Cultures: The Role of AKT. Neuroscience 2004, 127, 649–658. [Google Scholar] [CrossRef]
- Griffin, R.J.; Moloney, A.; Kelliher, M.; Johnston, J.A.; Ravid, R.; Dockery, P.; O’Connor, R.; O’Neill, C. Activation of Akt/PKB, Increased Phosphorylation of Akt Substrates and Loss and Altered Distribution of Akt and PTEN Are Features of Alzheimer’s Disease Pathology. J. Neurochem. 2005, 93, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Kerr, F.; Rickle, A.; Nayeem, N.; Brandner, S.; Cowburn, R.F.; Lovestone, S. PTEN, a Negative Regulator of PI3 Kinase Signalling, Alters Tau Phosphorylation in Cells by Mechanisms Independent of GSK-3. FEBS Lett. 2006, 580, 3121–3128. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, F.; Bulloj, A.; Zhang, Y.-W.; Tong, G.; Zhang, Z.; Liao, F.-F.; Xu, H. Tumor-Suppressor PTEN Affects Tau Phosphorylation, Aggregation, and Binding to Microtubules. FASEB J. 2006, 20, 1272–1274. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Chen, B.; Xu, W.-F.; Liu, R.-F.; Yang, J.; Yu, C.-X. Effects of PTEN Inhibition on Regulation of Tau Phosphorylation in an Okadaic Acid-Induced Neurodegeneration Model. Int. J. Dev. Neurosci. 2012, 30, 411–419. [Google Scholar] [CrossRef] [PubMed]
- Singh, T.J.; Zaidi, T.; Grundke-Iqbal, I.; Iqbal, K. Modulation of GSK-3-Catalyzed Phosphorylation of Microtubule-Associated Protein Tau by Non-Proline-Dependent Protein Kinases. FEBS Lett. 1995, 358, 4–8. [Google Scholar] [CrossRef]
- Wagner, U.; Utton, M.; Gallo, J.M.; Miller, C.C. Cellular Phosphorylation of Tau by GSK-3 Beta Influences Tau Binding to Microtubules and Microtubule Organisation. J. Cell Sci. 1996, 109, 1537–1543. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.J.; Zhang, J.Y.; Li, H.L.; Fang, Z.Y.; Wang, Q.; Deng, H.M.; Gong, C.X.; Grundke-Iqbal, I.; Iqbal, K.; Wang, J.Z. Tau Becomes a More Favorable Substrate for GSK-3 When It Is Prephosphorylated by PKA in Rat Brain. J. Biol. Chem. 2004, 279, 50078–50088. [Google Scholar] [CrossRef]
- Avila, J.; Hernández, F. GSK-3 Inhibitors for Alzheimer’s Disease. Expert Rev. Neurother. 2007, 7, 1527–1533. [Google Scholar] [CrossRef]
- Thomas, S.M.; Brugge, J.S. Cellular Functions Regulated by Src Family Kinases. Annu. Rev. Cell Dev. Biol. 1997, 13, 513–609. [Google Scholar] [CrossRef] [PubMed]
- Parsons, S.J.; Parsons, J.T. Src Family Kinases, Key Regulators of Signal Transduction. Oncogene 2004, 23, 7906–7909. [Google Scholar] [CrossRef] [PubMed]
- Portugal, C.C.; Almeida, T.O.; Socodato, R.; Relvas, J.B. Src Family Kinases (SFKs): Critical Regulators of Microglial Homeostatic Functions and Neurodegeneration in Parkinson’s and Alzheimer’s Diseases. FEBS J. 2021. [Google Scholar] [CrossRef] [PubMed]
- Stehelin, D.; Varmus, H.E.; Bishop, J.M.; Vogt, P.K. DNA Related to the Transforming Gene(s) of Avian Sarcoma Viruses Is Present in Normal Avian DNA. Nature 1976, 260, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Arias-Salgado, E.G.; Lizano, S.; Sarkar, S.; Brugge, J.S.; Ginsberg, M.H.; Shattil, S.J. Src Kinase Activation by Direct Interaction with the Integrin Beta Cytoplasmic Domain. Proc. Natl. Acad. Sci. USA 2003, 100, 13298–13302. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Lee, F.Y.; Bhalla, K.N.; Wu, J. Potent Inhibition of Platelet-Derived Growth Factor-Induced Responses in Vascular Smooth Muscle Cells by BMS-354825 (Dasatinib). Mol. Pharmacol. 2006, 69, 1527–1533. [Google Scholar] [CrossRef]
- Schittenhelm, M.M.; Shiraga, S.; Schroeder, A.; Corbin, A.S.; Griffith, D.; Lee, F.Y.; Bokemeyer, C.; Deininger, M.W.N.; Druker, B.J.; Heinrich, M.C. Dasatinib (BMS-354825), a Dual SRC/ABL Kinase Inhibitor, Inhibits the Kinase Activity of Wild-Type, Juxtamembrane, and Activation Loop Mutant KIT Isoforms Associated with Human Malignancies. Cancer Res. 2006, 66, 473–481. [Google Scholar] [CrossRef]
- Lu, K.V.; Zhu, S.; Cvrljevic, A.; Huang, T.T.; Sarkaria, S.; Ahkavan, D.; Dang, J.; Dinca, E.B.; Plaisier, S.B.; Oderberg, I.; et al. Fyn and SRC Are Effectors of Oncogenic Epidermal Growth Factor Receptor Signaling in Glioblastoma Patients. Cancer Res. 2009, 69, 6889–6898. [Google Scholar] [CrossRef]
- Kleber, S.; Sancho-Martinez, I.; Wiestler, B.; Beisel, A.; Gieffers, C.; Hill, O.; Thiemann, M.; Mueller, W.; Sykora, J.; Kuhn, A.; et al. Yes and PI3K Bind CD95 to Signal Invasion of Glioblastoma. Cancer Cell 2008, 13, 235–248. [Google Scholar] [CrossRef]
- Stettner, M.R.; Wang, W.; Nabors, L.B.; Bharara, S.; Flynn, D.C.; Grammer, J.R.; Gillespie, G.Y.; Gladson, C.L. Lyn Kinase Activity Is the Predominant Cellular SRC Kinase Activity in Glioblastoma Tumor Cells. Cancer Res. 2005, 65, 5535–5543. [Google Scholar] [CrossRef]
- Liu, W.M.; Huang, P.; Kar, N.; Burgett, M.; Muller-Greven, G.; Nowacki, A.S.; Distelhorst, C.W.; Lathia, J.D.; Rich, J.N.; Kappes, J.C.; et al. Lyn Facilitates Glioblastoma Cell Survival under Conditions of Nutrient Deprivation by Promoting Autophagy. PLoS ONE 2013, 8, e70804. [Google Scholar] [CrossRef] [PubMed]
- Zepecki, J.P.; Snyder, K.M.; Moreno, M.M.; Fajardo, E.; Fiser, A.; Ness, J.; Sarkar, A.; Toms, S.A.; Tapinos, N. Regulation of Human Glioma Cell Migration, Tumor Growth, and Stemness Gene Expression Using a Lck Targeted Inhibitor. Oncogene 2019, 38, 1734–1750. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ying, C.; Zhang, A.; Xu, H.; Jiang, Y.; Lou, M. HCK Promotes Glioblastoma Progression by TGFβ Signaling. Biosci. Rep. 2020, 40, BSR20200975. [Google Scholar] [CrossRef]
- Lund, C.V.; Nguyen, M.T.N.; Owens, G.C.; Pakchoian, A.J.; Shaterian, A.; Kruse, C.A.; Eliceiri, B.P. Reduced Glioma Infiltration in Src-Deficient Mice. J. Neurooncol. 2006, 78, 19–29. [Google Scholar] [CrossRef] [PubMed]
- Cirotti, C.; Contadini, C.; Barilà, D. SRC Kinase in Glioblastoma: News from an Old Acquaintance. Cancers 2020, 12, 1558. [Google Scholar] [CrossRef]
- Bjorge, J.D.; Jakymiw, A.; Fujita, D.J. Selected Glimpses into the Activation and Function of Src Kinase. Oncogene 2000, 19, 5620–5635. [Google Scholar] [CrossRef] [PubMed]
- Ahluwalia, M.S.; de Groot, J.; Liu, W.; Gladson, C.L. Targeting SRC in Glioblastoma Tumors and Brain Metastases: Rationale and Preclinical Studies. Cancer Lett. 2010, 298, 139–149. [Google Scholar] [CrossRef]
- Bhaskar, K.; Yen, S.-H.; Lee, G. Disease-Related Modifications in Tau Affect the Interaction between Fyn and Tau. J. Biol. Chem. 2005, 280, 35119–35125. [Google Scholar] [CrossRef]
- Kant, S.; Kesarwani, P.; Guastella, A.R.; Kumar, P.; Graham, S.F.; Buelow, K.L.; Nakano, I.; Chinnaiyan, P. Perhexiline Demonstrates FYN-Mediated Antitumor Activity in Glioblastoma. Mol. Cancer Ther. 2020, 19, 1415–1422. [Google Scholar] [CrossRef]
- Comba, A.; Dunn, P.J.; Argento, A.E.; Kadiyala, P.; Ventosa, M.; Patel, P.; Zamler, D.B.; Núñez, F.J.; Zhao, L.; Castro, M.G.; et al. Fyn Tyrosine Kinase, a Downstream Target of Receptor Tyrosine Kinases, Modulates Antiglioma Immune Responses. Neuro-Oncology 2020, 22, 806–818. [Google Scholar] [CrossRef]
- Gordon-Weeks, P.R.; Gary Mansfield, S.; Curran, I. Direct Visualisation of the Soluble Pool of Tubulin in the Neuronal Growth Cone: Immunofluorescence Studies Following Taxol Polymerisation. Dev. Brain Res. 1989, 49, 305–310. [Google Scholar] [CrossRef]
- DiTella, M.; Feiguin, F.; Morfini, G.; Cáceres, A. Microfilament-associated growth cone component depends upon Tau for its intracellular localization. Cell Motil. 1994, 29, 117–130. [Google Scholar] [CrossRef] [PubMed]
- Helmke, S.; Pfenninger, K.H. Growth Cone Enrichment and Cytoskeletal Association of Non-Receptor Tyrosine Kinases. Cell Motil. Cytoskeleton 1995, 30, 194–207. [Google Scholar] [CrossRef] [PubMed]
- LoPresti, P.; Szuchet, S.; Papasozomenos, S.C.; Zinkowski, R.P.; Binder, L.I. Functional Implications for the Microtubule-Associated Protein Tau: Localization in Oligodendrocytes. Proc. Natl. Acad. Sci. USA 1995, 92, 10369–10373. [Google Scholar] [CrossRef]
- Umemori, H.; Satot, S.; Yagi, T.; Aizawal, S.; Yamamoto, T. Initial Events of Myelination Involve Fyn Tyrosine Kinase Signalling. Nature 1994, 367, 572–576. [Google Scholar] [CrossRef]
- Ishii, N.; Maier, D.; Merlo, A.; Tada, M.; Sawamura, Y.; Diserens, A.C.; Van Meir, E.G. Frequent Co-Alterations of TP53, P16/CDKN2A, P14ARF, PTEN Tumor Suppressor Genes in Human Glioma Cell Lines. Brain Pathol. 1999, 9, 469–479. [Google Scholar] [CrossRef]
- Zhang, Y.; Dube, C.; Gibert, M.; Cruickshanks, N.; Wang, B.; Coughlan, M.; Yang, Y.; Setiady, I.; Deveau, C.; Saoud, K.; et al. The P53 Pathway in Glioblastoma. Cancers 2018, 10, 297. [Google Scholar] [CrossRef]
- Baquero, J.; Varriano, S.; Ordonez, M.; Kuczaj, P.; Murphy, M.R.; Aruggoda, G.; Lundine, D.; Morozova, V.; Makki, A.E.; Alonso, A.D.C.; et al. Nuclear Tau, P53 and Pin1 Regulate PARN-Mediated Deadenylation and Gene Expression. Front. Mol. Neurosci. 2019, 12, 242. [Google Scholar] [CrossRef]
- Chou, P.-Y.; Lin, S.-R.; Lee, M.-H.; Schultz, L.; Sze, C.-I.; Chang, N.-S. A P53/TIAF1/WWOX Triad Exerts Cancer Suppression but May Cause Brain Protein Aggregation Due to P53/WWOX Functional Antagonism. Cell Commun. Signal. 2019, 17, 76. [Google Scholar] [CrossRef]
- Farmer, K.M.; Ghag, G.; Puangmalai, N.; Montalbano, M.; Bhatt, N.; Kayed, R. P53 Aggregation, Interactions with Tau, and Impaired DNA Damage Response in Alzheimer’s Disease. Acta Neuropathol. Commun. 2020, 8, 132. [Google Scholar] [CrossRef]
- Montalbano, M.; McAllen, S.; Cascio, F.L.; Sengupta, U.; Garcia, S.; Bhatt, N.; Ellsworth, A.; Heidelman, E.A.; Johnson, O.D.; Doskocil, S.; et al. TDP-43 and Tau Oligomers in Alzheimer’s Disease, Amyotrophic Lateral Sclerosis, and Frontotemporal Dementia. Neurobiol. Dis. 2020, 146, 105130. [Google Scholar] [CrossRef] [PubMed]
- Jazvinšćak Jembrek, M.; Slade, N.; Hof, P.R.; Šimić, G. The Interactions of P53 with Tau and Aß as Potential Therapeutic Targets for Alzheimer’s Disease. Prog. Neurobiol. 2018, 168, 104–127. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, K.M.; Bhave, S.R.; Ferraro, D.J.; Jaboin, J.J.; Hallahan, D.E.; Thotala, D. GSK-3 β: A Bifunctional Role in Cell Death Pathways. Int. J. Cell Biol. 2012, 2012, e930710. [Google Scholar] [CrossRef]
- Hanger, D.P.; Hughes, K.; Woodgett, J.R.; Brion, J.-P.; Anderton, B.H. Glycogen Synthase Kinase-3 Induces Alzheimer’s Disease-like Phosphorylation of Tau: Generation of Paired Helical Filament Epitopes and Neuronal Localisation of the Kinase. Neurosci. Lett. 1992, 147, 58–62. [Google Scholar] [CrossRef]
- Mandelkow, E.M.; Drewes, G.; Biernat, J.; Gustke, N.; Van Lint, J.; Vandenheede, J.R.; Mandelkow, E. Glycogen Synthase Kinase-3 and the Alzheimer-like State of Microtubule-Associated Protein Tau. FEBS Lett. 1992, 314, 315–321. [Google Scholar] [CrossRef]
- Sun, W.; Qureshi, H.Y.; Cafferty, P.W.; Sobue, K.; Agarwal-Mawal, A.; Neufield, K.D.; Paudel, H.K. Glycogen Synthase Kinase-3beta Is Complexed with Tau Protein in Brain Microtubules. J. Biol. Chem. 2002, 277, 11933–11940. [Google Scholar] [CrossRef]
- Korur, S.; Huber, R.M.; Sivasankaran, B.; Petrich, M.; Jr, P.M.; Hemmings, B.A.; Merlo, A.; Lino, M.M. GSK3β Regulates Differentiation and Growth Arrest in Glioblastoma. PLoS ONE 2009, 4, e7443. [Google Scholar] [CrossRef]
- Kotliarova, S.; Pastorino, S.; Kovell, L.C.; Kotliarov, Y.; Song, H.; Zhang, W.; Bailey, R.; Maric, D.; Zenklusen, J.C.; Lee, J.; et al. Glycogen Synthase Kinase 3 Inhibition Induces Glioma Cell Death through C-MYC, NF-ΚB and Glucose Regulation. Cancer Res. 2008, 68, 6643–6651. [Google Scholar] [CrossRef]
- Nowicki, M.O.; Dmitrieva, N.; Stein, A.M.; Cutter, J.L.; Godlewski, J.; Saeki, Y.; Nita, M.; Berens, M.E.; Sander, L.M.; Newton, H.B.; et al. Lithium Inhibits Invasion of Glioma Cells; Possible Involvement of Glycogen Synthase Kinase-3. Neuro-Oncology 2008, 10, 690–699. [Google Scholar] [CrossRef]
- Aitken, A.; Collinge, D.B.; van Heusden, B.P.; Isobe, T.; Roseboom, P.H.; Rosenfeld, G.; Soll, J. 14-3-3 Proteins: A Highly Conserved, Widespread Family of Eukaryotic Proteins. Trends Biochem. Sci. 1992, 17, 498–501. [Google Scholar] [CrossRef]
- Joo, Y.; Schumacher, B.; Landrieu, I.; Bartel, M.; Smet-Nocca, C.; Jang, A.; Choi, H.S.; Jeon, N.L.; Chang, K.-A.; Kim, H.-S.; et al. Involvement of 14-3-3 in Tubulin Instability and Impaired Axon Development Is Mediated by Tau. FASEB J. 2015, 29, 4133–4144. [Google Scholar] [CrossRef] [PubMed]
- Morgan, D. The Cell Cycle: Principles of Control; New Science Press: London, UK, 2006; 297p. [Google Scholar]
- Lagace, D.C.; Benavides, D.R.; Kansy, J.W.; Mapelli, M.; Greengard, P.; Bibb, J.A.; Eisch, A.J. Cdk5 Is Essential for Adult Hippocampal Neurogenesis. Proc. Natl. Acad. Sci. USA 2008, 105, 18567–18571. [Google Scholar] [CrossRef] [PubMed]
- Tan, T.C.; Valova, V.A.; Malladi, C.S.; Graham, M.E.; Berven, L.A.; Jupp, O.J.; Hansra, G.; McClure, S.J.; Sarcevic, B.; Boadle, R.A.; et al. Cdk5 Is Essential for Synaptic Vesicle Endocytosis. Nat. Cell Biol. 2003, 5, 701–710. [Google Scholar] [CrossRef] [PubMed]
- Dhariwala, F.A.; Rajadhyaksha, M.S. An Unusual Member of the Cdk Family: Cdk5. Cell. Mol. Neurobiol. 2008, 28, 351–369. [Google Scholar] [CrossRef]
- Liebl, J.; Weitensteiner, S.B.; Vereb, G.; Takács, L.; Fürst, R.; Vollmar, A.M.; Zahler, S. Cyclin-Dependent Kinase 5 Regulates Endothelial Cell Migration and Angiogenesis. J. Biol. Chem. 2010, 285, 35932–35943. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.-S.; Alexander, K.; Santiago, P.; Hinds, P.W. ERM Proteins and Cdk5 in Cellular Senescence. Cell Cycle Georget. Tex. 2003, 2, 517–520. [Google Scholar] [CrossRef]
- Lazaro, J.B.; Kitzmann, M.; Poul, M.A.; Vandromme, M.; Lamb, N.J.; Fernandez, A. Cyclin Dependent Kinase 5, Cdk5, Is a Positive Regulator of Myogenesis in Mouse C2 Cells. J. Cell Sci. 1997, 110, 1251–1260. [Google Scholar] [CrossRef]
- Chapman, D.E.; Reddy, B.J.N.; Huy, B.; Bovyn, M.J.; Cruz, S.J.S.; Al-Shammari, Z.M.; Han, H.; Wang, W.; Smith, D.S.; Gross, S.P. Regulation of in Vivo Dynein Force Production by CDK5 and 14-3-3ε and KIAA0528. Nat. Commun. 2019, 10, 228. [Google Scholar] [CrossRef]
- Contreras-Vallejos, E.; Utreras, E.; Gonzalez-Billault, C. Going out of the Brain: Non-Nervous System Physiological and Pathological Functions of Cdk5. Cell. Signal. 2012, 24, 44–52. [Google Scholar] [CrossRef]
- Do, P.A.; Lee, C.H. The Role of CDK5 in Tumours and Tumour Microenvironments. Cancers 2020, 13, 101. [Google Scholar] [CrossRef]
- Yushan, R.; Wenjie, C.; Suning, H.; Yiwu, D.; Tengfei, Z.; Madushi, W.M.; Feifei, L.; Changwen, Z.; Xin, W.; Roodrajeetsing, G.; et al. Insights into the Clinical Value of Cyclin-Dependent Kinase 5 in Glioma: A Retrospective Study. World J. Surg. Oncol. 2015, 13, 223. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mukherjee, S.; Tucker-Burden, C.; Kaissi, E.; Newsam, A.; Duggireddy, H.; Chau, M.; Zhang, C.; Diwedi, B.; Rupji, M.; Seby, S.; et al. CDK5 Inhibition Resolves PKA/CAMP-Independent Activation of CREB1 Signaling in Glioma Stem Cells. Cell Rep. 2018, 23, 1651–1664. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, X.; Lv, P.; Yu, H.; Jiang, X. CDK5 Knockdown Inhibits Proliferation and Induces Apoptosis and Cell Cycle Arrest in Human Glioblastoma. J. Cancer 2021, 12, 3958–3966. [Google Scholar] [CrossRef] [PubMed]
- Huda, M.N.; Kim, D.H.; Erdene-Ochir, E.; Kim, Y.S.; Pan, C.-H. Expression, Phosphorylation, Localization, and Microtubule Binding of Tau in Colorectal Cell Lines. Appl. Biol. Chem. 2016, 59, 807–812. [Google Scholar] [CrossRef]
- Pauli, C.; Hopkins, B.D.; Prandi, D.; Shaw, R.; Fedrizzi, T.; Sboner, A.; Sailer, V.; Augello, M.; Puca, L.; Rosati, R.; et al. Personalized In Vitro and In Vivo Cancer Models to Guide Precision Medicine. Cancer Discov. 2017, 7, 462–477. [Google Scholar] [CrossRef]
- Aguirre, A.J.; Nowak, J.A.; Camarda, N.D.; Moffitt, R.A.; Ghazani, A.A.; Hazar-Rethinam, M.; Raghavan, S.; Kim, J.; Brais, L.K.; Ragon, D.; et al. Real-Time Genomic Characterization of Advanced Pancreatic Cancer to Enable Precision Medicine. Cancer Discov. 2018, 8, 1096–1111. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhou, L.; Xie, N.; Nice, E.C.; Zhang, T.; Cui, Y.; Huang, C. Overcoming Cancer Therapeutic Bottleneck by Drug Repurposing. Signal Transduct. Target. Ther. 2020, 5, 113. [Google Scholar] [CrossRef]
- Stupp, R.; Brada, M.; van den Bent, M.J.; Tonn, J.-C.; Pentheroudakis, G. ESMO Guidelines Working Group High-Grade Glioma: ESMO Clinical Practice Guidelines for Diagnosis, Treatment and Follow-Up. Ann. Oncol. 2014, 25, iii93–iii101. [Google Scholar] [CrossRef]
- Moffat, J.G.; Rudolph, J.; Bailey, D. Phenotypic Screening in Cancer Drug Discovery—Past, Present and Future. Nat. Rev. Drug Discov. 2014, 13, 588–602. [Google Scholar] [CrossRef]
- Chen, H.I.; Song, H.; Ming, G.-L. Applications of Human Brain Organoids to Clinical Problems. Dev. Dyn. 2019, 248, 53–64. [Google Scholar] [CrossRef]
- Loong, H.H.-F.; Wong, A.M.; Chan, D.T.-M.; Cheung, M.S.-H.; Chow, C.; Ding, X.; Chan, A.K.-Y.; Johnston, P.A.; Lau, J.Y.-W.; Poon, W.S.; et al. Patient-Derived Tumor Organoid Predicts Drugs Response in Glioblastoma: A Step Forward in Personalized Cancer Therapy? J. Clin. Neurosci. 2020, 78, 400–402. [Google Scholar] [CrossRef] [PubMed]
- Klein, E.; Hau, A.-C.; Oudin, A.; Golebiewska, A.; Niclou, S.P. Glioblastoma Organoids: Pre-Clinical Applications and Challenges in the Context of Immunotherapy. Front. Oncol. 2020, 10, 604121. [Google Scholar] [CrossRef] [PubMed]
- Vincent, F.; Nueda, A.; Lee, J.; Schenone, M.; Prunotto, M.; Mercola, M. Phenotypic Drug Discovery: Recent Successes, Lessons Learned and New Directions. Nat. Rev. Drug Discov. 2022, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Delgado-Martín, B.; Medina, M.Á. Advances in the Knowledge of the Molecular Biology of Glioblastoma and Its Impact in Patient Diagnosis, Stratification, and Treatment. Adv. Sci. 2020, 7, 1902971. [Google Scholar] [CrossRef]
- Bougnaud, S.; Golebiewska, A.; Oudin, A.; Keunen, O.; Harter, P.N.; Mäder, L.; Azuaje, F.; Fritah, S.; Stieber, D.; Kaoma, T.; et al. Molecular Crosstalk between Tumour and Brain Parenchyma Instructs Histopathological Features in Glioblastoma. Oncotarget 2016, 7, 31955–31971. [Google Scholar] [CrossRef]
- Perrin, S.L.; Samuel, M.S.; Koszyca, B.; Brown, M.P.; Ebert, L.M.; Oksdath, M.; Gomez, G.A. Glioblastoma Heterogeneity and the Tumour Microenvironment: Implications for Preclinical Research and Development of New Treatments. Biochem. Soc. Trans. 2019, 47, 625–638. [Google Scholar] [CrossRef]
- Mitusova, K.; Peltek, O.O.; Karpov, T.E.; Muslimov, A.R.; Zyuzin, M.V.; Timin, A.S. Overcoming the Blood-Brain Barrier for the Therapy of Malignant Brain Tumor: Current Status and Prospects of Drug Delivery Approaches. J. Nanobiotechnol. 2022, 20, 412. [Google Scholar] [CrossRef] [PubMed]
- Congdon, E.E.; Sigurdsson, E.M. Tau-Targeting Therapies for Alzheimer Disease. Nat. Rev. Neurol. 2018, 14, 399–415. [Google Scholar] [CrossRef]
- Jadhav, S.; Avila, J.; Schöll, M.; Kovacs, G.G.; Kövari, E.; Skrabana, R.; Evans, L.D.; Kontsekova, E.; Malawska, B.; de Silva, R.; et al. A Walk through Tau Therapeutic Strategies. Acta Neuropathol. Commun. 2019, 7, 22. [Google Scholar] [CrossRef]
- Yu, T.-W.; Lane, H.-Y.; Lin, C.-H. Novel Therapeutic Approaches for Alzheimer’s Disease: An Updated Review. Int. J. Mol. Sci. 2021, 22, 8208. [Google Scholar] [CrossRef]
- Hatakeyama, S.; Matsumoto, M.; Kamura, T.; Murayama, M.; Chui, D.-H.; Planel, E.; Takahashi, R.; Nakayama, K.I.; Takashima, A. U-Box Protein Carboxyl Terminus of Hsc70-Interacting Protein (CHIP) Mediates Poly-Ubiquitylation Preferentially on Four-Repeat Tau and Is Involved in Neurodegeneration of Tauopathy. J. Neurochem. 2004, 91, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Petrucelli, L.; Dickson, D.; Kehoe, K.; Taylor, J.; Snyder, H.; Grover, A.; De Lucia, M.; McGowan, E.; Lewis, J.; Prihar, G.; et al. CHIP and Hsp70 Regulate Tau Ubiquitination, Degradation and Aggregation. Hum. Mol. Genet. 2004, 13, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.; Zhang, H.; Xie, J.; Meng, D.; Wang, X.; Ke, D.; Zeng, J.; Liu, R. Protein Phosphatase 2A as a Drug Target in the Treatment of Cancer and Alzheimer’s Disease. Curr. Med. Sci. 2020, 40, 1–8. [Google Scholar] [CrossRef]
- Ding, H.; Dolan, P.J.; Johnson, G.V.W. Histone Deacetylase 6 Interacts with the Microtubule-Associated Protein Tau. J. Neurochem. 2008, 106, 2119–2130. [Google Scholar] [CrossRef] [PubMed]
- Yun, H.S.; Lee, J.; Kil, W.J.; Kramp, T.R.; Tofilon, P.J.; Camphausen, K. The Radiosensitizing Effect of AZD0530 in Glioblastoma and Glioblastoma Stem-Like Cells. Mol. Cancer Ther. 2021, 20, 1672–1679. [Google Scholar] [CrossRef]
- Khan, I.; Tantray, M.A.; Alam, M.S.; Hamid, H. Natural and Synthetic Bioactive Inhibitors of Glycogen Synthase Kinase. Eur. J. Med. Chem. 2017, 125, 464–477. [Google Scholar] [CrossRef]
- Happold, C.; Gorlia, T.; Chinot, O.; Gilbert, M.R.; Nabors, L.B.; Wick, W.; Pugh, S.L.; Hegi, M.; Cloughesy, T.; Roth, P.; et al. Does Valproic Acid or Levetiracetam Improve Survival in Glioblastoma? A Pooled Analysis of Prospective Clinical Trials in Newly Diagnosed Glioblastoma. J. Clin. Oncol. 2016, 34, 731–739. [Google Scholar] [CrossRef]
- Fay, M.F.; Head, R.; Sminia, P.; Dowson, N.; Cosgrove, L.; Rose, S.E.; Martin, J.H. Valproate in Adjuvant Glioblastoma Treatment. J. Clin. Oncol. 2016, 34, 3105–3107. [Google Scholar] [CrossRef][Green Version]
- Hu, J.-P.; Xie, J.-W.; Wang, C.-Y.; Wang, T.; Wang, X.; Wang, S.-L.; Teng, W.-P.; Wang, Z.-Y. Valproate Reduces Tau Phosphorylation via Cyclin-Dependent Kinase 5 and Glycogen Synthase Kinase 3 Signaling Pathways. Brain Res. Bull. 2011, 85, 194–200. [Google Scholar] [CrossRef]
- Bahmad, H.F.; Chalhoub, R.M.; Harati, H.; Bou-Gharios, J.; Assi, S.; Ballout, F.; Monzer, A.; Msheik, H.; Araji, T.; Elajami, M.K.; et al. Tideglusib Attenuates Growth of Neuroblastoma Cancer Stem/Progenitor Cells in Vitro and in Vivo by Specifically Targeting GSK-3β. Pharmacol. Rep. 2021, 73, 211–226. [Google Scholar] [CrossRef]
- Wei, D.; Zhu, X.; Li, S.; Liu, G.; Wang, Y.; Wang, W.; Zhang, Q.; Jiang, S. Tideglusib Suppresses Stem-Cell-like Features and Progression of Osteosarcoma by Inhibiting GSK-3β/NOTCH1 Signaling. Biochem. Biophys. Res. Commun. 2021, 554, 206–213. [Google Scholar] [CrossRef] [PubMed]
- Bou-Gharios, J.; Assi, S.; Bahmad, H.F.; Kharroubi, H.; Araji, T.; Chalhoub, R.M.; Ballout, F.; Harati, H.; Fares, Y.; Abou-Kheir, W. The Potential Use of Tideglusib as an Adjuvant Radio-Therapeutic Treatment for Glioblastoma Multiforme Cancer Stem-like Cells. Pharmacol. Rep. 2021, 73, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Cicenas, J.; Kalyan, K.; Sorokinas, A.; Stankunas, E.; Levy, J.; Meskinyte, I.; Stankevicius, V.; Kaupinis, A.; Valius, M. Roscovitine in Cancer and Other Diseases. Ann. Transl. Med. 2015, 3, 6. [Google Scholar] [CrossRef]
- Ghia, P.; Scarfò, L.; Perez, S.; Pathiraja, K.; Derosier, M.; Small, K.; McCrary Sisk, C.; Patton, N. Efficacy and Safety of Dinaciclib vs Ofatumumab in Patients with Relapsed/Refractory Chronic Lymphocytic Leukemia. Blood 2017, 129, 1876–1878. [Google Scholar] [CrossRef]
- Murphy, A.G.; Zahurak, M.; Shah, M.; Weekes, C.D.; Hansen, A.; Siu, L.L.; Spreafico, A.; LoConte, N.; Anders, N.M.; Miles, T.; et al. A Phase I Study of Dinaciclib in Combination With MK-2206 in Patients With Advanced Pancreatic Cancer. Clin. Transl. Sci. 2020, 13, 1178–1188. [Google Scholar] [CrossRef]
- Meijer, L.; Hery-Arnaud, G.; Leven, C.; Nowak, E.; Hillion, S.; Renaudineau, Y.; Durieu, I.; Chiron, R.; Prevotat, A.; Fajac, I.; et al. Safety and Pharmacokinetics of Roscovitine (Seliciclib) in Cystic Fibrosis Patients Chronically Infected with Pseudomonas Aeruginosa, a Randomized, Placebo-Controlled Study. J. Cyst. Fibros. 2021, 21, 529–536. [Google Scholar] [CrossRef]
- Sangodkar, J.; Perl, A.; Tohme, R.; Kiselar, J.; Kastrinsky, D.B.; Zaware, N.; Izadmehr, S.; Mazhar, S.; Wiredja, D.D.; O’Connor, C.M.; et al. Activation of Tumor Suppressor Protein PP2A Inhibits KRAS-Driven Tumor Growth. J. Clin. Investig. 2017, 127, 2081–2090. [Google Scholar] [CrossRef]
- Wei, H.; Zhang, H.-L.; Wang, X.-C.; Xie, J.-Z.; An, D.-D.; Wan, L.; Wang, J.-Z.; Zeng, Y.; Shu, X.-J.; Westermarck, J.; et al. Direct Activation of Protein Phosphatase 2A (PP2A) by Tricyclic Sulfonamides Ameliorates Alzheimer’s Disease Pathogenesis in Cell and Animal Models. Neurother. J. Am. Soc. Exp. Neurother. 2020, 17, 1087–1103. [Google Scholar] [CrossRef]
- Yang, X.-J.; Seto, E. HATs and HDACs: From Structure, Function and Regulation to Novel Strategies for Therapy and Prevention. Oncogene 2007, 26, 5310–5318. [Google Scholar] [CrossRef] [PubMed]
- Lv, D.; Jia, F.; Hou, Y.; Sang, Y.; Alvarez, A.A.; Zhang, W.; Gao, W.-Q.; Hu, B.; Cheng, S.-Y.; Ge, J.; et al. Histone Acetyltransferase KAT6A Upregulates PI3K/AKT Signaling through TRIM24 Binding. Cancer Res. 2017, 77, 6190–6201. [Google Scholar] [CrossRef] [PubMed]
- Rokudai, S.; Laptenko, O.; Arnal, S.M.; Taya, Y.; Kitabayashi, I.; Prives, C. MOZ Increases P53 Acetylation and Premature Senescence through Its Complex Formation with PML. Proc. Natl. Acad. Sci. USA 2013, 110, 3895–3900. [Google Scholar] [CrossRef] [PubMed]
- Baell, J.B.; Leaver, D.J.; Hermans, S.J.; Kelly, G.L.; Brennan, M.S.; Downer, N.L.; Nguyen, N.; Wichmann, J.; McRae, H.M.; Yang, Y.; et al. Inhibitors of Histone Acetyltransferases KAT6A/B Induce Senescence and Arrest Tumour Growth. Nature 2018, 560, 253–257. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Zhang, M.; Zhou, Y.; Guo, W.; Yi, M.; Zhang, Z.; Ding, Y.; Wang, Y. The Application of Histone Deacetylases Inhibitors in Glioblastoma. J. Exp. Clin. Cancer Res. 2020, 39, 138. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; Zhang, H.; Jiao, M.; Han, B.; Zhang, Z.; Li, J.; Zhang, Q. Histone Deacetylase 6 Inhibitors with Blood-Brain Barrier Penetration as a Potential Strategy for CNS-Disorders Therapy. Eur. J. Med. Chem. 2022, 229, 114090. [Google Scholar] [CrossRef] [PubMed]
- Perez, T.; Bergès, R.; Maccario, H.; Oddoux, S.; Honoré, S. Low Concentrations of Vorinostat Decrease EB1 Expression in GBM Cells and Affect Microtubule Dynamics, Cell Survival and Migration. Oncotarget 2021, 12, 304–315. [Google Scholar] [CrossRef]
- Balmik, A.A.; Chidambaram, H.; Dangi, A.; Marelli, U.K.; Chinnathambi, S. HDAC6 ZnF UBP as the Modifier of Tau Structure and Function. Biochemistry 2020, 59, 4546–4562. [Google Scholar] [CrossRef]
- Min, S.-W.; Cho, S.-H.; Zhou, Y.; Schroeder, S.; Haroutunian, V.; Seeley, W.W.; Huang, E.J.; Shen, Y.; Masliah, E.; Mukherjee, C.; et al. Acetylation of Tau Inhibits Its Degradation and Contributes to Tauopathy. Neuron 2010, 67, 953–966. [Google Scholar] [CrossRef]
- Caballero, B.; Bourdenx, M.; Luengo, E.; Diaz, A.; Sohn, P.D.; Chen, X.; Wang, C.; Juste, Y.R.; Wegmann, S.; Patel, B.; et al. Acetylated Tau Inhibits Chaperone-Mediated Autophagy and Promotes Tau Pathology Propagation in Mice. Nat. Commun. 2021, 12, 2238. [Google Scholar] [CrossRef]
- Hastings, N.B.; Wang, X.; Song, L.; Butts, B.D.; Grotz, D.; Hargreaves, R.; Fred Hess, J.; Hong, K.-L.K.; Huang, C.R.-R.; Hyde, L.; et al. Inhibition of O-GlcNAcase Leads to Elevation of O-GlcNAc Tau and Reduction of Tauopathy and Cerebrospinal Fluid Tau in RTg4510 Mice. Mol. Neurodegener. 2017, 12, 39. [Google Scholar] [CrossRef]
- Yuzwa, S.A.; Macauley, M.S.; Heinonen, J.E.; Shan, X.; Dennis, R.J.; He, Y.; Whitworth, G.E.; Stubbs, K.A.; McEachern, E.J.; Davies, G.J.; et al. A Potent Mechanism-Inspired O-GlcNAcase Inhibitor That Blocks Phosphorylation of Tau in Vivo. Nat. Chem. Biol. 2008, 4, 483–490. [Google Scholar] [CrossRef]
- DeVos, S.L.; Miller, T.M. Antisense Oligonucleotides: Treating Neurodegeneration at the Level of RNA. Neurother. J. Am. Soc. Exp. Neurother. 2013, 10, 486–497. [Google Scholar] [CrossRef] [PubMed]
- DeVos, S.L.; Miller, R.L.; Schoch, K.M.; Holmes, B.B.; Kebodeaux, C.S.; Wegener, A.J.; Chen, G.; Shen, T.; Tran, H.; Nichols, B.; et al. Tau Reduction Prevents Neuronal Loss and Reverses Pathological Tau Deposition and Seeding in Mice with Tauopathy. Sci. Transl. Med. 2017, 9, eaag0481. [Google Scholar] [CrossRef] [PubMed]
- David, D.C.; Layfield, R.; Serpell, L.; Narain, Y.; Goedert, M.; Spillantini, M.G. Proteasomal Degradation of Tau Protein. J. Neurochem. 2002, 83, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Dolan, P.J.; Johnson, G.V.W. A Caspase Cleaved Form of Tau Is Preferentially Degraded through the Autophagy Pathway. J. Biol. Chem. 2010, 285, 21978–21987. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Martinez-Vicente, M.; Krüger, U.; Kaushik, S.; Wong, E.; Mandelkow, E.-M.; Cuervo, A.M.; Mandelkow, E. Synergy and Antagonism of Macroautophagy and Chaperone-Mediated Autophagy in a Cell Model of Pathological Tau Aggregation. Autophagy 2010, 6, 182–183. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Mandelkow, E. Degradation of Tau Protein by Autophagy and Proteasomal Pathways. Biochem. Soc. Trans. 2012, 40, 644–652. [Google Scholar] [CrossRef]
- Lee, M.J.; Lee, J.H.; Rubinsztein, D.C. Tau Degradation: The Ubiquitin–Proteasome System versus the Autophagy-Lysosome System. Prog. Neurobiol. 2013, 105, 49–59. [Google Scholar] [CrossRef]
- Parzych, K.R.; Klionsky, D.J. An Overview of Autophagy: Morphology, Mechanism, and Regulation. Antioxid. Redox Signal. 2014, 20, 460–473. [Google Scholar] [CrossRef]
- Caballero, B.; Wang, Y.; Diaz, A.; Tasset, I.; Juste, Y.R.; Stiller, B.; Mandelkow, E.-M.; Mandelkow, E.; Cuervo, A.M. Interplay of Pathogenic Forms of Human Tau with Different Autophagic Pathways. Aging Cell. 2018, 17, e12692. [Google Scholar] [CrossRef]
- Auzmendi-Iriarte, J.; Matheu, A. Impact of Chaperone-Mediated Autophagy in Brain Aging: Neurodegenerative Diseases and Glioblastoma. Front. Aging Neurosci. 2021, 12, 509. [Google Scholar] [CrossRef]
- Massey, A.; Kiffin, R.; Cuervo, A.M. Pathophysiology of Chaperone-Mediated Autophagy. Int. J. Biochem. Cell Biol. 2004, 36, 2420–2434. [Google Scholar] [CrossRef] [PubMed]
- Jinwal, U.K.; O’Leary, J.C.; Borysov, S.I.; Jones, J.R.; Li, Q.; Koren, J.; Abisambra, J.F.; Vestal, G.D.; Lawson, L.Y.; Johnson, A.G.; et al. Hsc70 Rapidly Engages Tau after Microtubule Destabilization. J. Biol. Chem. 2010, 285, 16798–16805. [Google Scholar] [CrossRef]
- Bi, X.; Haque, T.S.; Zhou, J.; Skillman, A.G.; Lin, B.; Lee, C.E.; Kuntz, I.D.; Ellman, J.A.; Lynch, G. Novel Cathepsin D Inhibitors Block the Formation of Hyperphosphorylated Tau Fragments in Hippocampus. J. Neurochem. 2000, 74, 1469–1477. [Google Scholar] [CrossRef] [PubMed]
- Shimada, K.; Motoi, Y.; Ishiguro, K.; Kambe, T.; Matsumoto, S.; Itaya, M.; Kunichika, M.; Mori, H.; Shinohara, A.; Chiba, M.; et al. Long-Term Oral Lithium Treatment Attenuates Motor Disturbance in Tauopathy Model Mice: Implications of Autophagy Promotion. Neurobiol. Dis. 2012, 46, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer, V.; Goedert, M. Stimulation of Autophagy Is Neuroprotective in a Mouse Model of Human Tauopathy. Autophagy 2012, 8, 1686–1687. [Google Scholar] [CrossRef]
- Jiang, T.; Yu, J.-T.; Zhu, X.-C.; Zhang, Q.-Q.; Cao, L.; Wang, H.-F.; Tan, M.-S.; Gao, Q.; Qin, H.; Zhang, Y.-D.; et al. Temsirolimus Attenuates Tauopathy in Vitro and in Vivo by Targeting Tau Hyperphosphorylation and Autophagic Clearance. Neuropharmacology 2014, 85, 121–130. [Google Scholar] [CrossRef]
- Hebron, M.L.; Javidnia, M.; Moussa, C.E.-H. Tau Clearance Improves Astrocytic Function and Brain Glutamate-Glutamine Cycle. J. Neurol. Sci. 2018, 391, 90–99. [Google Scholar] [CrossRef]
- Hernandez, I.; Luna, G.; Rauch, J.N.; Reis, S.A.; Giroux, M.; Karch, C.M.; Boctor, D.; Sibih, Y.E.; Storm, N.J.; Diaz, A.; et al. A Farnesyltransferase Inhibitor Activates Lysosomes and Reduces Tau Pathology in Mice with Tauopathy. Sci. Transl. Med. 2019, 11, eaat3005. [Google Scholar] [CrossRef]
- Song, J.-X.; Malampati, S.; Zeng, Y.; Durairajan, S.S.K.; Yang, C.-B.; Tong, B.C.-K.; Iyaswamy, A.; Shang, W.-B.; Sreenivasmurthy, S.G.; Zhu, Z.; et al. A Small Molecule Transcription Factor EB Activator Ameliorates Beta-Amyloid Precursor Protein and Tau Pathology in Alzheimer’s Disease Models. Aging Cell 2020, 19, e13069. [Google Scholar] [CrossRef]
- Currais, A.; Prior, M.; Dargusch, R.; Armando, A.; Ehren, J.; Schubert, D.; Quehenberger, O.; Maher, P. Modulation of P25 and Inflammatory Pathways by Fisetin Maintains Cognitive Function in Alzheimer’s Disease Transgenic Mice. Aging Cell 2014, 13, 379–390. [Google Scholar] [CrossRef]
- Spagnuolo, P.A.; Hu, J.; Hurren, R.; Wang, X.; Gronda, M.; Sukhai, M.A.; Di Meo, A.; Boss, J.; Ashali, I.; Beheshti Zavareh, R.; et al. The Antihelmintic Flubendazole Inhibits Microtubule Function through a Mechanism Distinct from Vinca Alkaloids and Displays Preclinical Activity in Leukemia and Myeloma. Blood 2010, 115, 4824–4833. [Google Scholar] [CrossRef] [PubMed]
- Hou, Z.-J.; Luo, X.; Zhang, W.; Peng, F.; Cui, B.; Wu, S.-J.; Zheng, F.-M.; Xu, J.; Xu, L.-Z.; Long, Z.-J.; et al. Flubendazole, FDA-Approved Anthelmintic, Targets Breast Cancer Stem-like Cells. Oncotarget 2015, 6, 6326–6340. [Google Scholar] [CrossRef]
- Chauhan, S.; Ahmed, Z.; Bradfute, S.B.; Arko-Mensah, J.; Mandell, M.A.; Won Choi, S.; Kimura, T.; Blanchet, F.; Waller, A.; Mudd, M.H.; et al. Pharmaceutical Screen Identifies Novel Target Processes for Activation of Autophagy with a Broad Translational Potential. Nat. Commun. 2015, 6, 8620. [Google Scholar] [CrossRef] [PubMed]
- Cai, N.; Chen, J.; Bi, D.; Gu, L.; Yao, L.; Li, X.; Li, H.; Xu, H.; Hu, Z.; Liu, Q.; et al. Specific Degradation of Endogenous Tau Protein and Inhibition of Tau Fibrillation by Tanshinone IIA through the Ubiquitin–Proteasome Pathway. J. Agric. Food Chem. 2020, 68, 2054–2062. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Bhaskar, K. Degradation and Transmission of Tau by Autophagic-Endolysosomal Networks and Potential Therapeutic Targets for Tauopathy. Front. Mol. Neurosci. 2020, 13, 199. [Google Scholar] [CrossRef]
- Gandini, A.; Bartolini, M.; Tedesco, D.; Martinez-Gonzalez, L.; Roca, C.; Campillo, N.E.; Zaldivar-Diez, J.; Perez, C.; Zuccheri, G.; Miti, A.; et al. Tau-Centric Multitarget Approach for Alzheimer’s Disease: Development of First-in-Class Dual Glycogen Synthase Kinase 3β and Tau-Aggregation Inhibitors. J. Med. Chem. 2018, 61, 7640–7656. [Google Scholar] [CrossRef]
- Cisek, K.; Cooper, G.L.; Huseby, C.J.; Kuret, J. Structure and Mechanism of Action of Tau Aggregation Inhibitors. Curr. Alzheimer Res. 2014, 11, 918–927. [Google Scholar] [CrossRef]
- Akoury, E.; Pickhardt, M.; Gajda, M.; Biernat, J.; Mandelkow, E.; Zweckstetter, M. Mechanistic Basis of Phenothiazine-Driven Inhibition of Tau Aggregation. Angew. Chem. Int. Ed. 2013, 52, 3511–3515. [Google Scholar] [CrossRef]
- Okuda, M.; Hijikuro, I.; Fujita, Y.; Wu, X.; Nakayama, S.; Sakata, Y.; Noguchi, Y.; Ogo, M.; Akasofu, S.; Ito, Y.; et al. PE859, a Novel Tau Aggregation Inhibitor, Reduces Aggregated Tau and Prevents Onset and Progression of Neural Dysfunction In Vivo. PLoS ONE 2015, 10, e0117511. [Google Scholar] [CrossRef]
- Okuda, M.; Hijikuro, I.; Fujita, Y.; Teruya, T.; Kawakami, H.; Takahashi, T.; Sugimoto, H. Design and Synthesis of Curcumin Derivatives as Tau and Amyloid β Dual Aggregation Inhibitors. Bioorg. Med. Chem. Lett. 2016, 26, 5024–5028. [Google Scholar] [CrossRef]
- Sandusky-Beltran, L.A.; Sigurdsson, E.M. Tau Immunotherapies: Lessons Learned, Current Status and Future Considerations. Neuropharmacology 2020, 175, 108104. [Google Scholar] [CrossRef]
- Ji, C.; Sigurdsson, E.M. Current Status of Clinical Trials on Tau Immunotherapies. Drugs 2021, 81, 1135–1152. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, S.K.; Joly-Amado, A.; Gordon, M.N.; Morgan, D. Tau-Directed Immunotherapy: A Promising Strategy for Treating Alzheimer’s Disease and Other Tauopathies. J. Neuroimmune Pharmacol. 2016, 11, 9–25. [Google Scholar] [CrossRef] [PubMed]
- Funk, K.E.; Mirbaha, H.; Jiang, H.; Holtzman, D.M.; Diamond, M.I. Distinct Therapeutic Mechanisms of Tau Antibodies: Promoting Microglial Clearance Versus Blocking Neuronal Uptake. J. Biol. Chem. 2015, 290, 21652–21662. [Google Scholar] [CrossRef]
- Luo, W.; Liu, W.; Hu, X.; Hanna, M.; Caravaca, A.; Paul, S.M. Microglial Internalization and Degradation of Pathological Tau Is Enhanced by an Anti-Tau Monoclonal Antibody. Sci. Rep. 2015, 5, 11161. [Google Scholar] [CrossRef] [PubMed]
- Congdon, E.E.; Lin, Y.; Rajamohamedsait, H.B.; Shamir, D.B.; Krishnaswamy, S.; Rajamohamedsait, W.J.; Rasool, S.; Gonzalez, V.; Levenga, J.; Gu, J.; et al. Affinity of Tau Antibodies for Solubilized Pathological Tau Species but Not Their Immunogen or Insoluble Tau Aggregates Predicts in Vivo and Ex Vivo Efficacy. Mol. Neurodegener. 2016, 11, 62. [Google Scholar] [CrossRef]
- Krishnamurthy, P.; Deng, Y.; Sigurdsson, E. Mechanistic Studies of Antibody-Mediated Clearance of Tau Aggregates Using an Ex Vivo Brain Slice Model. Front. Psychiatry 2011, 2, 59. [Google Scholar] [CrossRef]
- Gu, J.; Congdon, E.E.; Sigurdsson, E.M. Two Novel Tau Antibodies Targeting the 396/404 Region Are Primarily Taken up by Neurons and Reduce Tau Protein Pathology. J. Biol. Chem. 2013, 288, 33081–33095. [Google Scholar] [CrossRef]
- McEwan, W.A.; Falcon, B.; Vaysburd, M.; Clift, D.; Oblak, A.L.; Ghetti, B.; Goedert, M.; James, L.C. Cytosolic Fc Receptor TRIM21 Inhibits Seeded Tau Aggregation. Proc. Natl. Acad. Sci. USA 2017, 114, 574–579. [Google Scholar] [CrossRef]
- Shahpasand, K.; Sepehri Shamloo, A.; Nabavi, S.M.; Ping Lu, K.; Zhen Zhou, X. Tau Immunotherapy: Hopes and Hindrances. Hum. Vaccines Immunother. 2017, 14, 277–284. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).