High ETV6 Levels Support Aggressive B Lymphoma Cell Survival and Predict Poor Outcome in Diffuse Large B-Cell Lymphoma Patients
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Primary Tumor Specimens
2.2. Forward Phase Protein Arrays
2.3. Histological Evaluation and Immunohistochemical Analysis
2.4. Bioinformatical Software and Analyses
2.5. Gene Expression Data
2.6. Western Blotting
2.7. Lentiviral Constructs and Viral Production
2.8. Cell Viability Assays and Flow Cytometry
2.9. Statistical Analyses
3. Results
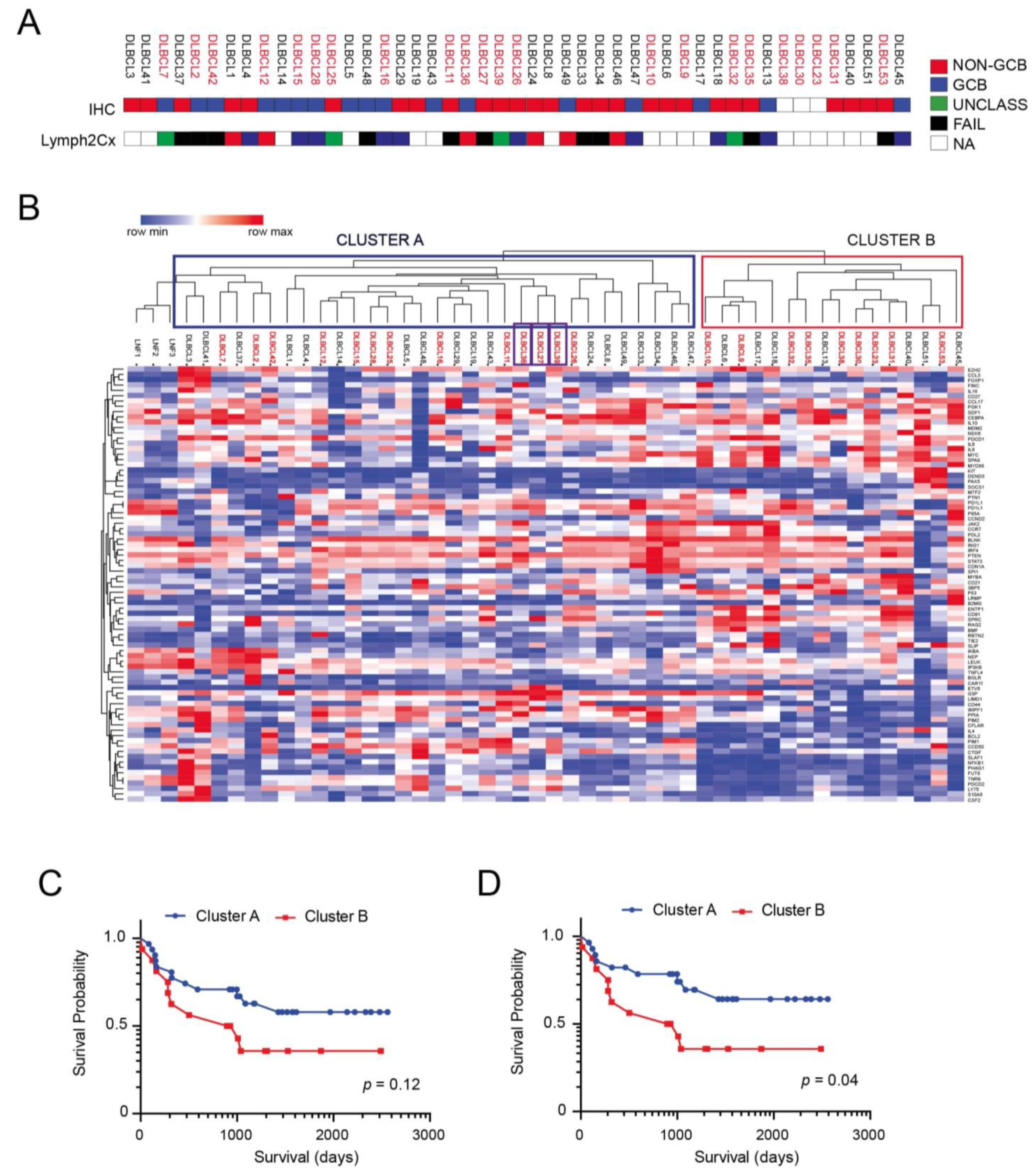
3.1. Antibody Arrays Identify Two Distinct Protein Signatures in Profiled DLBCLs
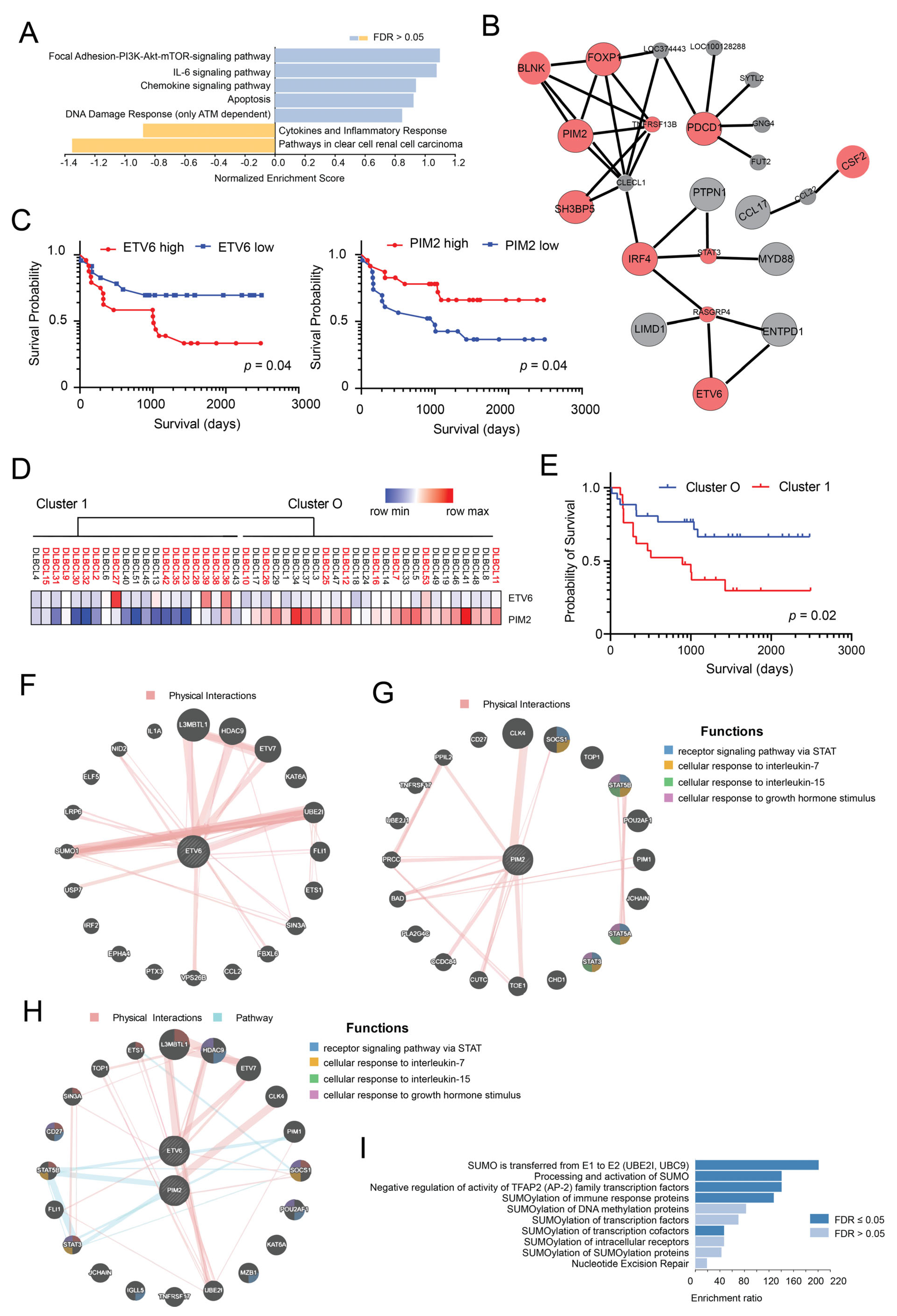
3.2. ETV6 and PIM2 Represent Candidate Proteins Predicting Survival in Aggressive DLBCLs Cases
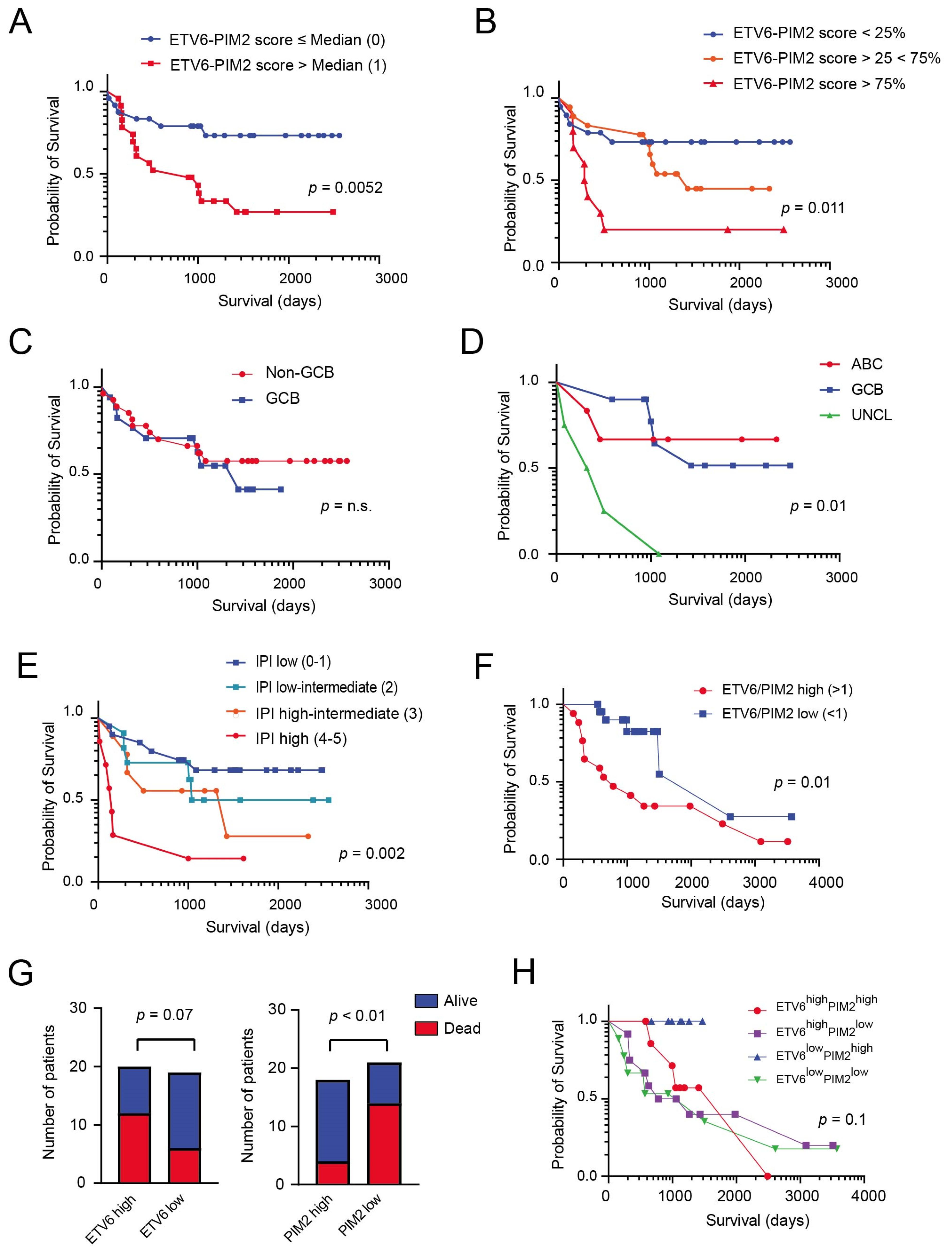
3.3. Generation of a Model Based on the Expression of PIM2 and ETV6 for Predicting Survival in DLBCL
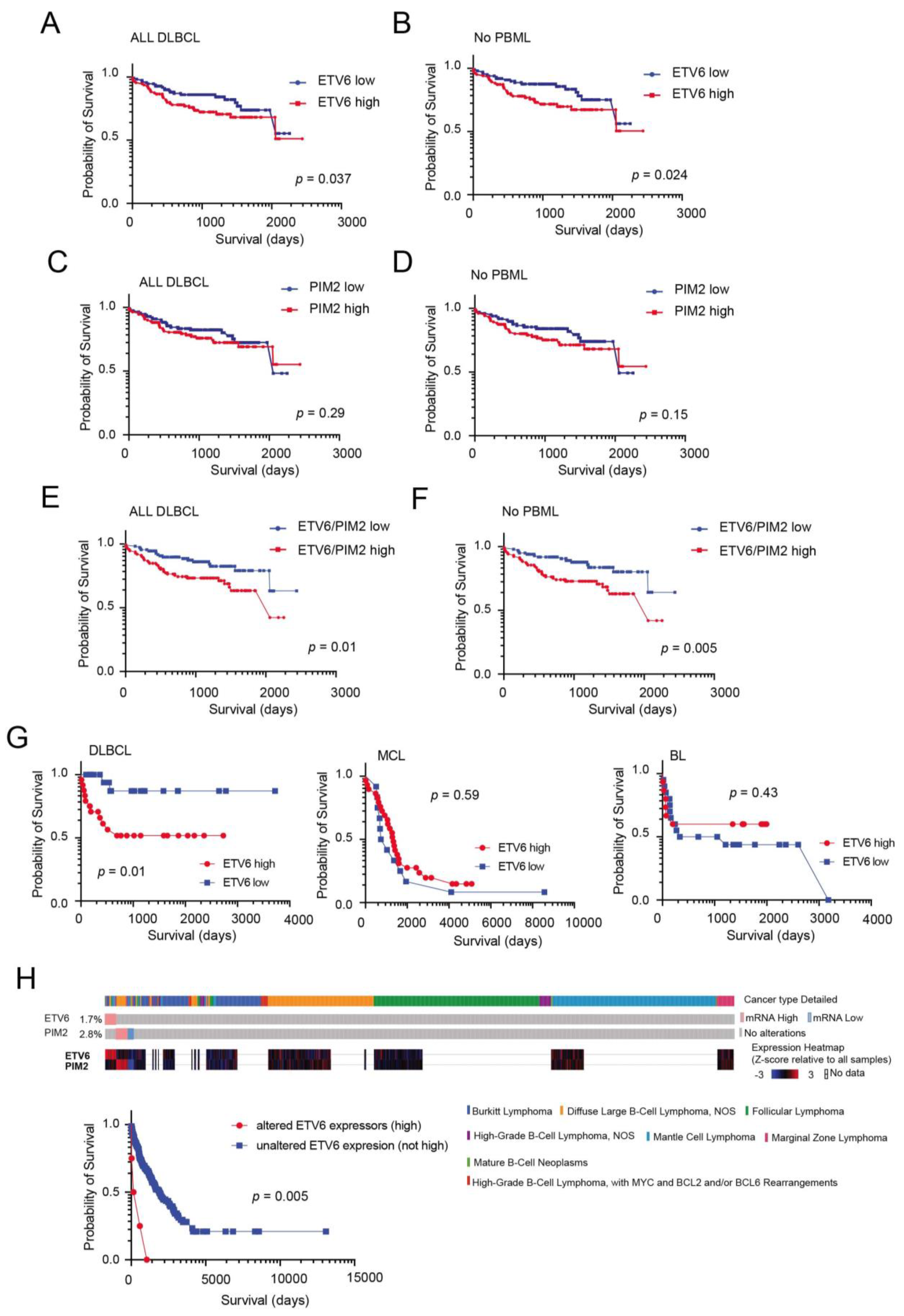
3.4. External Validation and Prognostic Significance of ETV6 Overexpression in Aggressive B-Cell Lymphomas
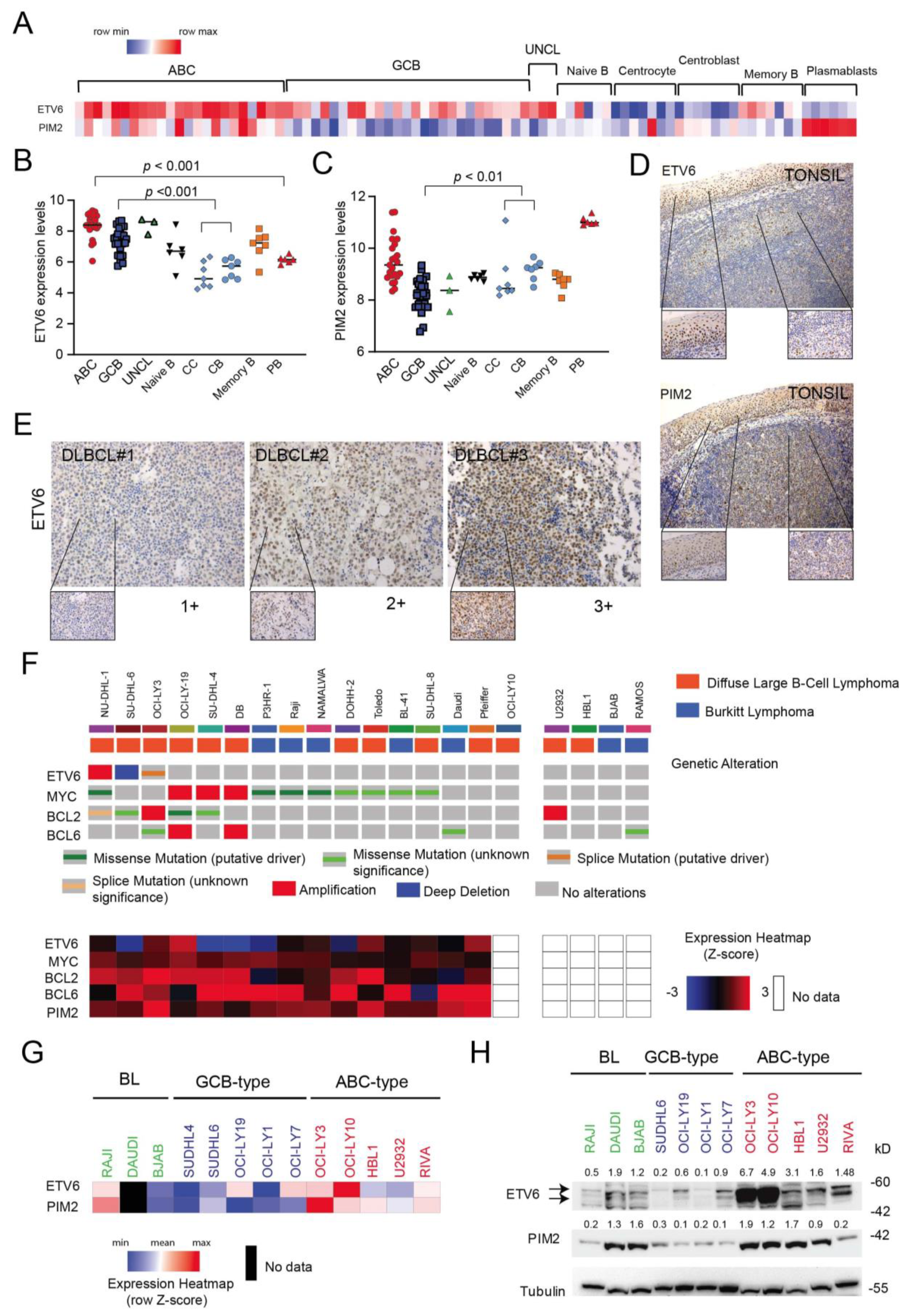
3.5. ETV6 Expression in Normal and Malignant B-Cells
3.6. Inactivation of ETV6 Is Highly Cytotoxic to Aggressive B-Cell Lymphoma Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Coiffier, B. Diffuse large cell lymphoma. Curr. Opin. Oncol. 2001, 13, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Roschewski, M.; Staudt, L.M.; Wilson, W.H. Diffuse large B-cell lymphoma-treatment approaches in the molecular era. Nat. Rev. Clin. Oncol. 2014, 11, 12–23. [Google Scholar] [CrossRef] [PubMed]
- A clinical evaluation of the International Lymphoma Study Group classification of non-Hodgkin’s lymphoma. The Non-Hodgkin’s Lymphoma Classification Project. Blood 1997, 89, 3909–3918. [CrossRef]
- Onaindia, A.; Santiago-Quispe, N.; Iglesias-Martinez, E.; Romero-Abrio, C. Molecular Update and Evolving Classification of Large B-Cell Lymphoma. Cancers 2021, 13, 3352. [Google Scholar] [CrossRef]
- Alizadeh, A.A.; Eisen, M.B.; Davis, R.E.; Ma, C.; Lossos, I.S.; Rosenwald, A.; Boldrick, J.C.; Sabet, H.; Tran, T.; Yu, X.; et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 2000, 403, 503–511. [Google Scholar] [CrossRef]
- Rosenwald, A.; Wright, G.; Chan, W.C.; Connors, J.M.; Campo, E.; Fisher, R.I.; Gascoyne, R.D.; Muller-Hermelink, H.K.; Smeland, E.B.; Giltnane, J.M.; et al. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N. Engl. J. Med. 2002, 346, 1937–1947. [Google Scholar] [CrossRef]
- Wright, G.; Tan, B.; Rosenwald, A.; Hurt, E.H.; Wiestner, A.; Staudt, L.M. A gene expression-based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc. Natl. Acad. Sci. USA 2003, 100, 9991–9996. [Google Scholar] [CrossRef]
- Hans, C.P.; Weisenburger, D.D.; Greiner, T.C.; Gascoyne, R.D.; Delabie, J.; Ott, G.; Muller-Hermelink, H.K.; Campo, E.; Braziel, R.M.; Jaffe, E.S.; et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 2004, 103, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Choi, W.W.; Weisenburger, D.D.; Greiner, T.C.; Piris, M.A.; Banham, A.H.; Delabie, J.; Braziel, R.M.; Geng, H.; Iqbal, J.; Lenz, G.; et al. A new immunostain algorithm classifies diffuse large B-cell lymphoma into molecular subtypes with high accuracy. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2009, 15, 5494–5502. [Google Scholar] [CrossRef] [PubMed]
- Xue, X.; Zeng, N.; Gao, Z.; Du, M.Q. Diffuse large B-cell lymphoma: Sub-classification by massive parallel quantitative RT-PCR. Lab. Investig. A J. Tech. Methods Pathol. 2015, 95, 113–120. [Google Scholar] [CrossRef]
- Alizadeh, A.A.; Gentles, A.J.; Alencar, A.J.; Liu, C.L.; Kohrt, H.E.; Houot, R.; Goldstein, M.J.; Zhao, S.; Natkunam, Y.; Advani, R.H.; et al. Prediction of survival in diffuse large B-cell lymphoma based on the expression of 2 genes reflecting tumor and microenvironment. Blood 2011, 118, 1350–1358. [Google Scholar] [CrossRef] [PubMed]
- Malumbres, R.; Chen, J.; Tibshirani, R.; Johnson, N.A.; Sehn, L.H.; Natkunam, Y.; Briones, J.; Advani, R.; Connors, J.M.; Byrne, G.E.; et al. Paraffin-based 6-gene model predicts outcome in diffuse large B-cell lymphoma patients treated with R-CHOP. Blood 2008, 111, 5509–5514. [Google Scholar] [CrossRef]
- Lossos, I.S.; Czerwinski, D.K.; Alizadeh, A.A.; Wechser, M.A.; Tibshirani, R.; Botstein, D.; Levy, R. Prediction of survival in diffuse large-B-cell lymphoma based on the expression of six genes. N. Engl. J. Med. 2004, 350, 1828–1837. [Google Scholar] [CrossRef]
- Scott, D.W.; Wright, G.W.; Williams, P.M.; Lih, C.J.; Walsh, W.; Jaffe, E.S.; Rosenwald, A.; Campo, E.; Chan, W.C.; Connors, J.M.; et al. Determining cell-of-origin subtypes of diffuse large B-cell lymphoma using gene expression in formalin-fixed paraffin-embedded tissue. Blood 2014, 123, 1214–1217. [Google Scholar] [CrossRef] [PubMed]
- Coutinho, R.; Clear, A.J.; Owen, A.; Wilson, A.; Matthews, J.; Lee, A.; Alvarez, R.; Gomes da Silva, M.; Cabecadas, J.; Calaminici, M.; et al. Poor concordance among nine immunohistochemistry classifiers of cell-of-origin for diffuse large B-cell lymphoma: Implications for therapeutic strategies. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2013, 19, 6686–6695. [Google Scholar] [CrossRef]
- Schroder, C.; Alhamdani, M.S.; Fellenberg, K.; Bauer, A.; Jacob, A.; Hoheisel, J.D. Robust protein profiling with complex antibody microarrays in a dual-colour mode. Methods Mol. Biol. 2011, 785, 203–221. [Google Scholar] [CrossRef] [PubMed]
- Brennan, D.J.; O’Connor, D.P.; Rexhepaj, E.; Ponten, F.; Gallagher, W.M. Antibody-based proteomics: Fast-tracking molecular diagnostics in oncology. Nat. Rev. Cancer 2010, 10, 605–617. [Google Scholar] [CrossRef]
- Swerdlow, S.H.; Campo, E.; Pileri, S.A.; Harris, N.L.; Stein, H.; Siebert, R.; Advani, R.; Ghielmini, M.; Salles, G.A.; Zelenetz, A.D.; et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 2016, 127, 2375–2390. [Google Scholar] [CrossRef]
- Alhamdani, M.S.; Schroder, C.; Hoheisel, J.D. Analysis conditions for proteomic profiling of mammalian tissue and cell extracts with antibody microarrays. Proteomics 2010, 10, 3203–3207. [Google Scholar] [CrossRef]
- Lenz, G.; Wright, G.; Dave, S.S.; Xiao, W.; Powell, J.; Zhao, H.; Xu, W.; Tan, B.; Goldschmidt, N.; Iqbal, J.; et al. Stromal gene signatures in large-B-cell lymphomas. N. Engl. J. Med. 2008, 359, 2313–2323. [Google Scholar] [CrossRef]
- Pizzi, M.; Friziero, A.; Vianello, F.; Binotto, G.; Da Dalt, G.; Zoletto, S.; Carraro, E.; Fassan, M.; Sbaraglia, M.; Sperti, C.; et al. Histology of the spleen in immune thrombocytopenia: Clinical-pathological characterization and prognostic implications. Eur. J. Haematol. 2021, 106, 281–289. [Google Scholar] [CrossRef]
- Cappellesso, R.; Fassan, M.; Hanspeter, E.; Bornschein, J.; d’Amore, E.S.; Cuorvo, L.V.; Mazzoleni, G.; Barbareschi, M.; Pizzi, M.; Guzzardo, V.; et al. HER2 status in gastroesophageal cancer: A tissue microarray study of 1040 cases. Hum. Pathol. 2015, 46, 665–672. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Wang, J.; Jaehnig, E.J.; Shi, Z.; Zhang, B. WebGestalt 2019: Gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019, 47, W199–W205. [Google Scholar] [CrossRef] [PubMed]
- Warde-Farley, D.; Donaldson, S.L.; Comes, O.; Zuberi, K.; Badrawi, R.; Chao, P.; Franz, M.; Grouios, C.; Kazi, F.; Lopes, C.T.; et al. The GeneMANIA prediction server: Biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010, 38, W214–W220. [Google Scholar] [CrossRef]
- Dubois, S.; Tesson, B.; Mareschal, S.; Viailly, P.J.; Bohers, E.; Ruminy, P.; Etancelin, P.; Peyrouze, P.; Copie-Bergman, C.; Fabiani, B.; et al. Refining diffuse large B-cell lymphoma subgroups using integrated analysis of molecular profiles. EBioMedicine 2019, 48, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.C.J.; Tadros, S.; Bouska, A.; Heavican, T.; Yang, H.; Deng, Q.; Moore, D.; Akhter, A.; Hartert, K.; Jain, N.; et al. Subtype-specific and co-occurring genetic alterations in B-cell non-Hodgkin lymphoma. Haematologica 2021. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef]
- Dybkaer, K.; Bogsted, M.; Falgreen, S.; Bodker, J.S.; Kjeldsen, M.K.; Schmitz, A.; Bilgrau, A.E.; Xu-Monette, Z.Y.; Li, L.; Bergkvist, K.S.; et al. Diffuse large B-cell lymphoma classification system that associates normal B-cell subset phenotypes with prognosis. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2015, 33, 1379–1388. [Google Scholar] [CrossRef]
- Ghandi, M.; Huang, F.W.; Jane-Valbuena, J.; Kryukov, G.V.; Lo, C.C.; McDonald, E.R., 3rd; Barretina, J.; Gelfand, E.T.; Bielski, C.M.; Li, H.; et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 2019, 569, 503–508. [Google Scholar] [CrossRef]
- Cheson, B.D.; Fisher, R.I.; Barrington, S.F.; Cavalli, F.; Schwartz, L.H.; Zucca, E.; Lister, T.A. Recommendations for initial evaluation, staging, and response assessment of Hodgkin and non-Hodgkin lymphoma: The Lugano classification. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2014, 32, 3059–3068. [Google Scholar] [CrossRef]
- Liu, Z.; Meng, J.; Li, X.; Zhu, F.; Liu, T.; Wu, G.; Zhang, L. Identification of Hub Genes and Key Pathways Associated with Two Subtypes of Diffuse Large B-Cell Lymphoma Based on Gene Expression Profiling via Integrated Bioinformatics. BioMed Res. Int. 2018, 2018, 3574534. [Google Scholar] [CrossRef]
- Miao, Y.; Medeiros, L.J.; Li, Y.; Li, J.; Young, K.H. Genetic alterations and their clinical implications in DLBCL. Nat. Rev. Clin. Oncol. 2019, 16, 634–652. [Google Scholar] [CrossRef] [PubMed]
- Lacy, S.E.; Barrans, S.L.; Beer, P.A.; Painter, D.; Smith, A.G.; Roman, E.; Cooke, S.L.; Ruiz, C.; Glover, P.; Van Hoppe, S.J.L.; et al. Targeted sequencing in DLBCL, molecular subtypes, and outcomes: A Haematological Malignancy Research Network report. Blood 2020, 135, 1759–1771. [Google Scholar] [CrossRef] [PubMed]
- Chapuy, B.; Stewart, C.; Dunford, A.J.; Kim, J.; Kamburov, A.; Redd, R.A.; Lawrence, M.S.; Roemer, M.G.M.; Li, A.J.; Ziepert, M.; et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat. Med. 2018, 24, 679–690. [Google Scholar] [CrossRef]
- Schmitz, R.; Wright, G.W.; Huang, D.W.; Johnson, C.A.; Phelan, J.D.; Wang, J.Q.; Roulland, S.; Kasbekar, M.; Young, R.M.; Shaffer, A.L.; et al. Genetics and Pathogenesis of Diffuse Large B-Cell Lymphoma. N. Engl. J. Med. 2018, 378, 1396–1407. [Google Scholar] [CrossRef] [PubMed]
- Bolen, C.R.; Klanova, M.; Trneny, M.; Sehn, L.H.; He, J.; Tong, J.; Paulson, J.N.; Kim, E.; Vitolo, U.; Di Rocco, A.; et al. Prognostic impact of somatic mutations in diffuse large B-cell lymphoma and relationship to cell-of-origin: Data from the phase III GOYA study. Haematologica 2020, 105, 2298–2307. [Google Scholar] [CrossRef] [PubMed]
- Hock, H.; Shimamura, A. ETV6 in hematopoiesis and leukemia predisposition. Semin. Hematol. 2017, 54, 98–104. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, J.; Sui, X.; Li, Y.; Lu, K.; Fang, X.; Jiang, Y.; Wang, X. Prognostic and Clinicopathological Value of Survivin in Diffuse Large B-cell Lymphoma: A Meta-Analysis. Medicine 2015, 94, e1432. [Google Scholar] [CrossRef]
- Nakahara, T.; Kita, A.; Yamanaka, K.; Mori, M.; Amino, N.; Takeuchi, M.; Tominaga, F.; Hatakeyama, S.; Kinoyama, I.; Matsuhisa, A.; et al. YM155, a novel small-molecule survivin suppressant, induces regression of established human hormone-refractory prostate tumor xenografts. Cancer Res. 2007, 67, 8014–8021. [Google Scholar] [CrossRef] [PubMed]
- Kita, A.; Nakahara, T.; Yamanaka, K.; Nakano, K.; Nakata, M.; Mori, M.; Kaneko, N.; Koutoku, H.; Izumisawa, N.; Sasamata, M. Antitumor effects of YM155, a novel survivin suppressant, against human aggressive non-Hodgkin lymphoma. Leuk. Res. 2011, 35, 787–792. [Google Scholar] [CrossRef]
- Nakahara, T.; Kita, A.; Yamanaka, K.; Mori, M.; Amino, N.; Takeuchi, M.; Tominaga, F.; Kinoyama, I.; Matsuhisa, A.; Kudou, M.; et al. Broad spectrum and potent antitumor activities of YM155, a novel small-molecule survivin suppressant, in a wide variety of human cancer cell lines and xenograft models. Cancer Sci. 2011, 102, 614–621. [Google Scholar] [CrossRef] [PubMed]
- Tolcher, A.W.; Mita, A.; Lewis, L.D.; Garrett, C.R.; Till, E.; Daud, A.I.; Patnaik, A.; Papadopoulos, K.; Takimoto, C.; Bartels, P.; et al. Phase I and pharmacokinetic study of YM155, a small-molecule inhibitor of survivin. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2008, 26, 5198–5203. [Google Scholar] [CrossRef]
- Vitolo, U.; Novo, M. Bcl-2 inhibition in DLBCL: “The times they are a-changing”? Blood 2021, 137, 577–579. [Google Scholar] [CrossRef] [PubMed]
- Younes, A. Prognostic Significance of Diffuse Large B-Cell Lymphoma Cell of Origin: Seeing the Forest and the Trees. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2015, 33, 2835–2836. [Google Scholar] [CrossRef]
- Lenz, G.; Wright, G.W.; Emre, N.C.; Kohlhammer, H.; Dave, S.S.; Davis, R.E.; Carty, S.; Lam, L.T.; Shaffer, A.L.; Xiao, W.; et al. Molecular subtypes of diffuse large B-cell lymphoma arise by distinct genetic pathways. Proc. Natl. Acad. Sci. USA 2008, 105, 13520–13525. [Google Scholar] [CrossRef]
- Rosenwald, A.; Wright, G.; Leroy, K.; Yu, X.; Gaulard, P.; Gascoyne, R.D.; Chan, W.C.; Zhao, T.; Haioun, C.; Greiner, T.C.; et al. Molecular diagnosis of primary mediastinal B cell lymphoma identifies a clinically favorable subgroup of diffuse large B cell lymphoma related to Hodgkin lymphoma. J. Exp. Med. 2003, 198, 851–862. [Google Scholar] [CrossRef]
- Gomez-Abad, C.; Pisonero, H.; Blanco-Aparicio, C.; Roncador, G.; Gonzalez-Menchen, A.; Martinez-Climent, J.A.; Mata, E.; Rodriguez, M.E.; Munoz-Gonzalez, G.; Sanchez-Beato, M.; et al. PIM2 inhibition as a rational therapeutic approach in B-cell lymphoma. Blood 2011, 118, 5517–5527. [Google Scholar] [CrossRef]
- Brault, L.; Menter, T.; Obermann, E.C.; Knapp, S.; Thommen, S.; Schwaller, J.; Tzankov, A. PIM kinases are progression markers and emerging therapeutic targets in diffuse large B-cell lymphoma. Br. J. Cancer 2012, 107, 491–500. [Google Scholar] [CrossRef] [PubMed]
- Hock, H.; Meade, E.; Medeiros, S.; Schindler, J.W.; Valk, P.J.; Fujiwara, Y.; Orkin, S.H. Tel/Etv6 is an essential and selective regulator of adult hematopoietic stem cell survival. Genes Dev. 2004, 18, 2336–2341. [Google Scholar] [CrossRef]
- Bohlander, S.K. ETV6: A versatile player in leukemogenesis. Semin. Cancer Biol. 2005, 15, 162–174. [Google Scholar] [CrossRef]
- Zhang, M.Y.; Churpek, J.E.; Keel, S.B.; Walsh, T.; Lee, M.K.; Loeb, K.R.; Gulsuner, S.; Pritchard, C.C.; Sanchez-Bonilla, M.; Delrow, J.J.; et al. Germline ETV6 mutations in familial thrombocytopenia and hematologic malignancy. Nat. Genet. 2015, 47, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Sevilla, D.W.; Nandula, S.V.; Colovai, A.I.; Alexander, S.; Murty, V.V.; Alobeid, B.; Bhagat, G. Diffuse large B-cell lymphoma with TEL/ETV6 translocation. Hum. Pathol. 2009, 40, 588–593. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bogusz, A.M. EBV-Negative Monomorphic B-Cell Posttransplant Lymphoproliferative Disorder with Marked Morphologic Pleomorphism and Pathogenic Mutations in ASXL1, BCOR, CDKN2A, NF1, and TP53. Case Rep. Hematol. 2017, 2017, 5083463. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.; Zhang, J.; Davis, N.S.; Moffitt, A.B.; Love, C.L.; Waldrop, A.; Leppa, S.; Pasanen, A.; Meriranta, L.; Karjalainen-Lindsberg, M.L.; et al. Genetic and Functional Drivers of Diffuse Large B Cell Lymphoma. Cell 2017, 171, 481–494.e415. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Dong, S.; Yao, H.; Wen, L.; Qiu, H.; Qin, L.; Ma, L.; Chen, S. ETV6 mutation in a cohort of 970 patients with hematologic malignancies. Haematologica 2014, 99, e176–e178. [Google Scholar] [CrossRef] [PubMed]
- Cheson, B.D.; Bartlett, N.L.; Vose, J.M.; Lopez-Hernandez, A.; Seiz, A.L.; Keating, A.T.; Shamsili, S.; Papadopoulos, K.P. A phase II study of the survivin suppressant YM155 in patients with refractory diffuse large B-cell lymphoma. Cancer 2012, 118, 3128–3134. [Google Scholar] [CrossRef]
- Kaneko, N.; Mitsuoka, K.; Amino, N.; Yamanaka, K.; Kita, A.; Mori, M.; Miyoshi, S.; Kuromitsu, S. Combination of YM155, a survivin suppressant, with bendamustine and rituximab: A new combination therapy to treat relapsed/refractory diffuse large B-cell lymphoma. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2014, 20, 1814–1822. [Google Scholar] [CrossRef] [PubMed]

| Patients and Tumor Characteristics | |||
|---|---|---|---|
| Features | CR (33) | PR or PD (Less than CR; 14) | p Value |
| Median age, range | 67 (20–83) | 73 (47–82) | n.d. |
| Male sex | 17 (51%) | 7 (50%) | 0.93 |
| AAS III-IV | 18 (55%) | 10 (71%) | 0.28 |
| Elevated LDH | 11 (33%) | 8 (57%) | 0.13 |
| ECOG PS ≥ 2 | 3 (9%) | 6 (43%) | 0.007 |
| Extra nodal site ≥ 2 | 2 (6%) | 3 (21%) | 0.12 |
| Bulky > 6 cm | 12 (36%) | 5 (36%) | 0.96 |
| IPI risk group | 0.019 | ||
| Low (0–1) | 17 (52%) | 3 (9%) | |
| Intermediate (2–3) | 14 (42%) | 6 (43%) | |
| Poor (4–5) | 2 (6%) | 5 (36%) | |
| Histology | n.d. | ||
| DLBCL NOS | 31 (94%) | 13 (93%) | |
| DLBCL CNS | 2 (6%) | 1 (7%) | |
| Cell of Origin (Hans) | 0.62 | ||
| GCB | 12 (36%) | 5 (35%) | |
| Non GCB | 21 (63%) | 6 (43%) | |
| Not evaluated | 0 | 3 (21%) | |
| CGA | 0.0015 | ||
| Fit | 15 (45%) | 1 (7%) | |
| Unfit | 4 (12%) | 4 (28%) | |
| Frail | 1 (3%) | 5 (36%) | |
| Not assessed | 13 (39%) | 4 (29%) | |
| Primary treatment | n.d. | ||
| R-CHOP/R-COMP/COMP/R-AC/R-CHOP + HD MTX | 27 | 8 | |
| R-Bendamustine | 0 | 1 | |
| R-CVP | 1 | 3 | |
| MTX/ARA-C ± R | 2 | 1 | |
| R-VACOP-B | 2 | 1 | |
| R-Hyper CVAD | 1 | 0 | |
| Alive at last follow-up | 24 (72%) | 1 (7%) | <0.0001 |
| UniProtKB | Protein Name | Regression Coefficient | p-Value |
|---|---|---|---|
| Q9HC98 | NEK6 | 9.9051 | 0.005 |
| O15181 | CD21 | −3.3531 | 0.0089 |
| P27986 | P85A | −8.8588 | 0.0008 |
| Q02548 | PAX5 | −0.914 | 0.0475 |
| Q15116 | PDCD1 | 13.5509 | 0.0004 |
| Q16342 | PDCD2 | 7.034 | 0.0013 |
| P00558 | PGK1 | 7.2112 | 0.0008 |
| Q9NWQ8 | PAG1 | −16.0977 | 0.0004 |
| Q9P1W9 | PIM2 | −1.1727 | 0.0062 |
| P60484 | PTEN | 3.8517 | 0.0006 |
| P62937 | PPIA | −1.2619 | 0.0647 |
| P18031 | PTPN1 | −4.3549 | 0.0018 |
| Q9BQ51 | PD1L2 | −1.9748 | 0.0027 |
| P55895 | RAG2 | −3.035 | 0.0008 |
| P06703 | S100A6 | −11.2893 | 0.0029 |
| Q9H334 | FOXP1 | 5.85 | 0.0034 |
| P04406 | G3P/GAPDH | −3.6582 | 0.0002 |
| P05112 | IL4 | −4.8767 | 0.0022 |
| P05231 | IL6 | 4.8604 | 0.0011 |
| P27987 | IP3KB/ITPKB | 0.9472 | 0.0514 |
| Q15306 | IRF4 | −3.2923 | 0.0097 |
| O60674 | JAK2 | 1.35 | 0.0318 |
| Q9UGP4 | LIMD1 | 3.15 | <0.0001 |
| O60449 | LY75/DEC-205 | −10.7107 | 0.0005 |
| P10243 | MYBA/MYBL1 | 2.0831 | 0.0959 |
| Q99836 | MYD88 | −3.5678 | 0.0047 |
| O60239 | 3BP5/SH3BP5 | 6.0439 | 0.0003 |
| P61769 | B2MG | −0.8181 | 0.0992 |
| P10415 | BCL2 | 3.0367 | 0.0106 |
| P08236 | BGLR/GUSB | −2.6631 | 0.0287 |
| Q8WV28 | BLNK | 2.4548 | 0.0011 |
| Q92583 | CCL17/TARC | 6.7439 | 0.0014 |
| P10147 | CCL3 | 4.1361 | 0.0105 |
| P32248 | CCR7 | −3.5845 | <0.0001 |
| P26842 | CD27 | −6.0218 | 0.0064 |
| P60033 | CD81 | 7.0502 | 0.0006 |
| P49715 | CEBPA | 4.0571 | 0.0527 |
| P04141 | CSF2/GMCSF | −10.0918 | 0.0032 |
| P29279 | CTGF/IGFBP-8 | −7.2609 | 0.0138 |
| P49961 | ENTP1/CD39 | −4.3636 | 0.0176 |
| P41212 | ETV6/TEL1 | 1.6527 | 0.0017 |
| UniProtKB | Protein Name | HR (95% CI) | p-Value | Regression Coefficient |
|---|---|---|---|---|
| P41212 | ETV6/TEL1 | 1.09–3.43 | 0.0286 | 0.6489 |
| Q9P1W9 | PIM2 | 0.57–0.98 | 0.0346 | −0.2927 |
| Variable | Wald Statistic (Z-Score) | p-Value | Regression Coefficient (b) | Exp(b) | 95% CI of Exp(b) |
|---|---|---|---|---|---|
| IPI score versus | 6.28 | 0.012 | 0.411 | 1.5083 | 1.09–2.08 |
| ETV6-PIM2 score | 4.19 | 0.04 | 1.01 | 2.76 | 1.04–7.28 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marino, D.; Pizzi, M.; Kotova, I.; Schmidt, R.; Schröder, C.; Guzzardo, V.; Talli, I.; Peroni, E.; Finotto, S.; Scapinello, G.; et al. High ETV6 Levels Support Aggressive B Lymphoma Cell Survival and Predict Poor Outcome in Diffuse Large B-Cell Lymphoma Patients. Cancers 2022, 14, 338. https://doi.org/10.3390/cancers14020338
Marino D, Pizzi M, Kotova I, Schmidt R, Schröder C, Guzzardo V, Talli I, Peroni E, Finotto S, Scapinello G, et al. High ETV6 Levels Support Aggressive B Lymphoma Cell Survival and Predict Poor Outcome in Diffuse Large B-Cell Lymphoma Patients. Cancers. 2022; 14(2):338. https://doi.org/10.3390/cancers14020338
Chicago/Turabian StyleMarino, Dario, Marco Pizzi, Iuliia Kotova, Ronny Schmidt, Christoph Schröder, Vincenza Guzzardo, Ilaria Talli, Edoardo Peroni, Silvia Finotto, Greta Scapinello, and et al. 2022. "High ETV6 Levels Support Aggressive B Lymphoma Cell Survival and Predict Poor Outcome in Diffuse Large B-Cell Lymphoma Patients" Cancers 14, no. 2: 338. https://doi.org/10.3390/cancers14020338
APA StyleMarino, D., Pizzi, M., Kotova, I., Schmidt, R., Schröder, C., Guzzardo, V., Talli, I., Peroni, E., Finotto, S., Scapinello, G., Dei Tos, A. P., Piazza, F., Trentin, L., Zagonel, V., & Piovan, E. (2022). High ETV6 Levels Support Aggressive B Lymphoma Cell Survival and Predict Poor Outcome in Diffuse Large B-Cell Lymphoma Patients. Cancers, 14(2), 338. https://doi.org/10.3390/cancers14020338








