Identification of Prognostic Genes in Gliomas Based on Increased Microenvironment Stiffness
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Identification of Stiffness-Dependent Genes
2.2. TCGA Glioma Datasets
2.3. Functional Enrichment Analysis
2.4. Survival Analysis
2.5. CGGA Validation Datasets
2.6. GSE16011 Validation Datasets
2.7. Multivariate Cox Regression
3. Results
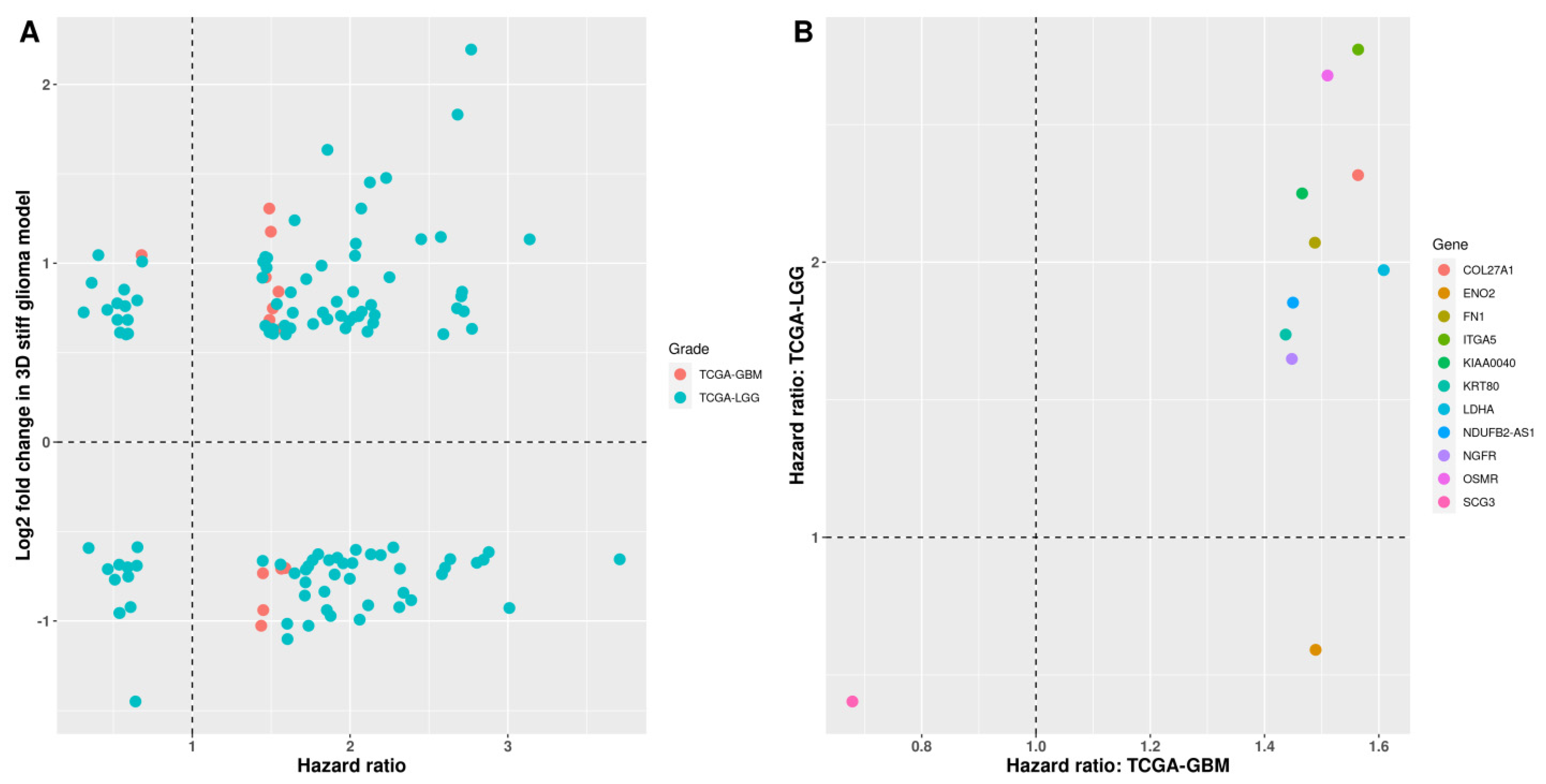
3.1. Identification of Stiffness-Dependent Genes from 3D-Printed Glioma Models
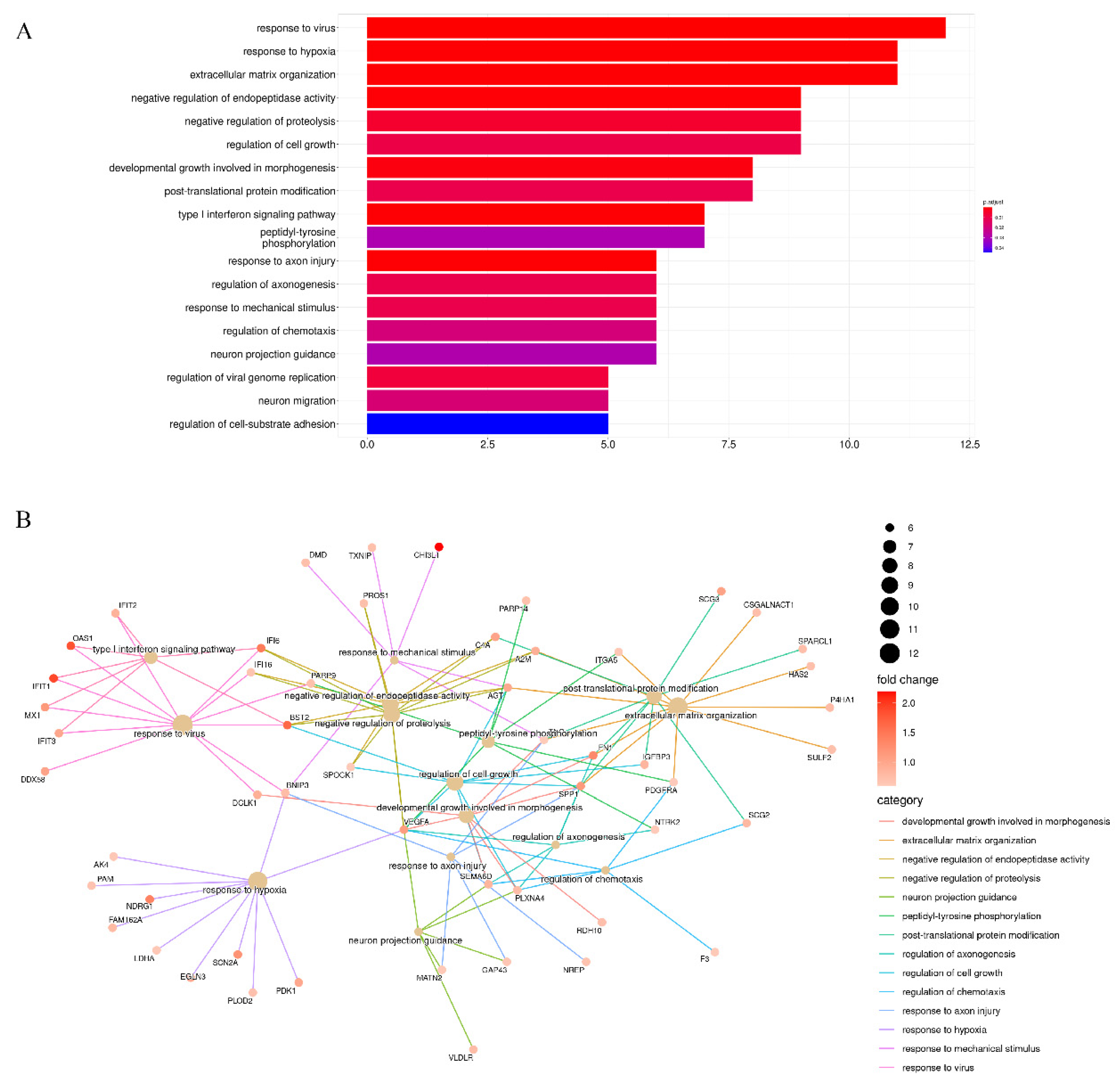
3.2. Enrichment Analysis of Stiffness-Dependent Genes
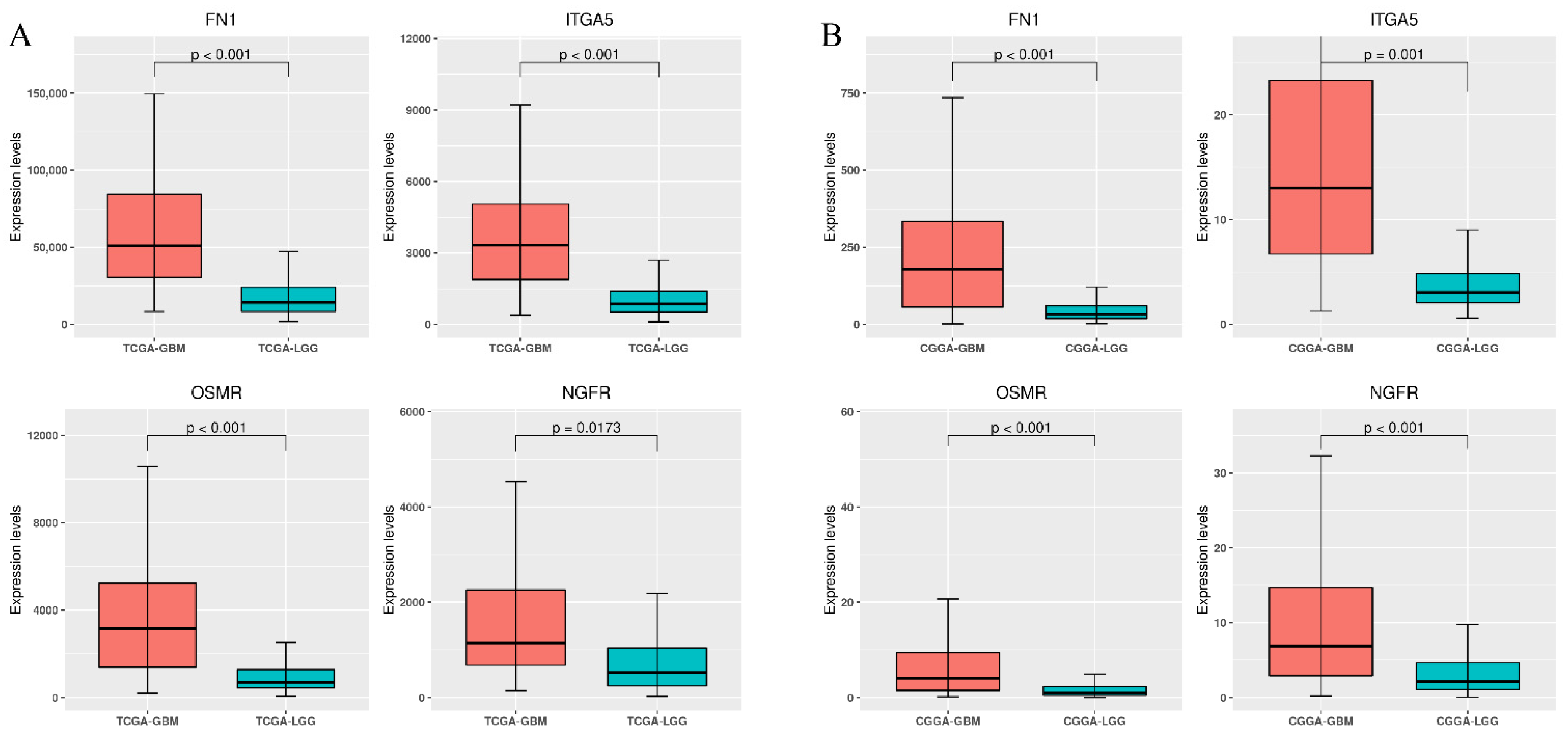
3.3. Differential Expression of Stiffness-Dependent Genes in TCGA Glioma Datasets
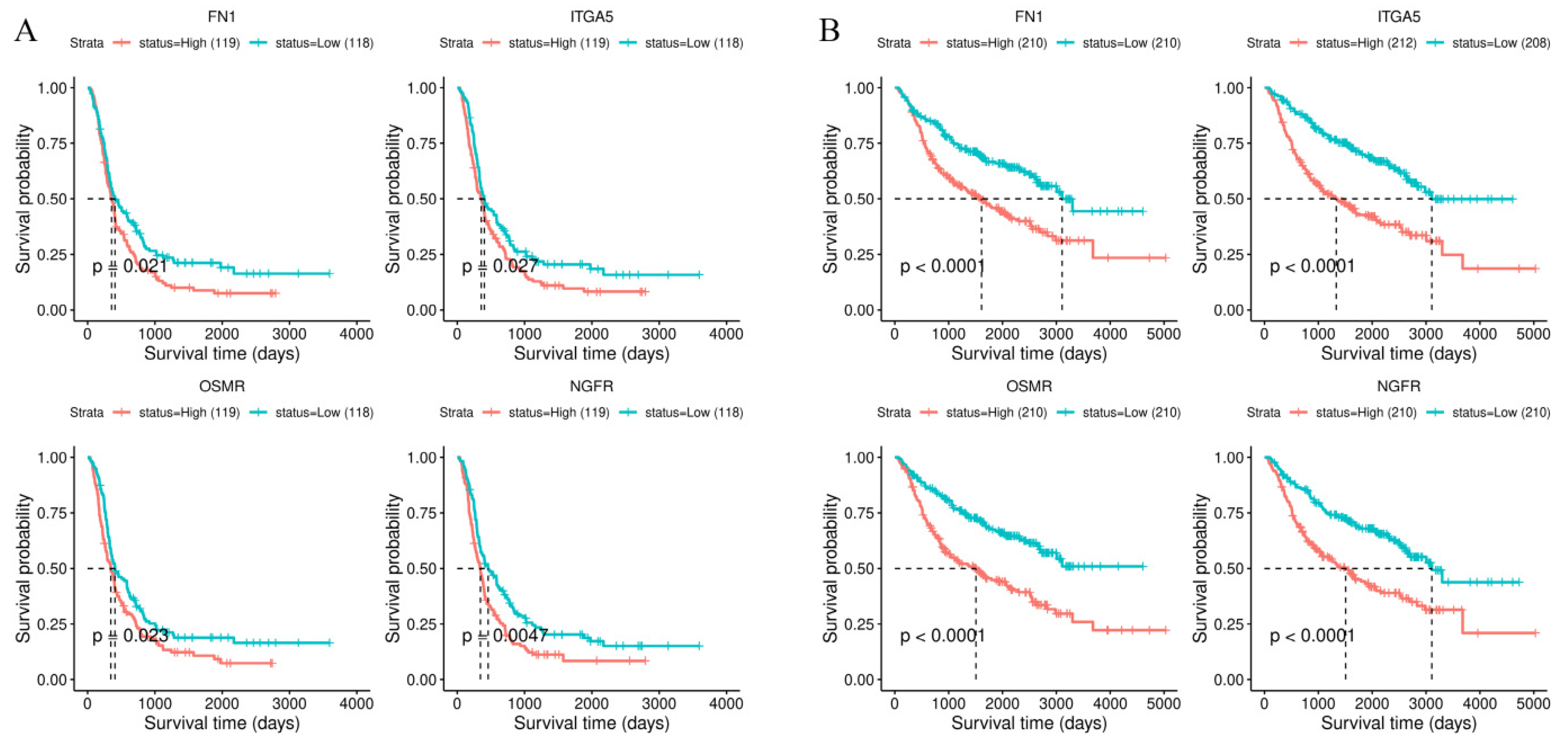
3.4. Survival Analysis of Stiffness-Dependent Genes
3.5. Validation Using the CGGA Database
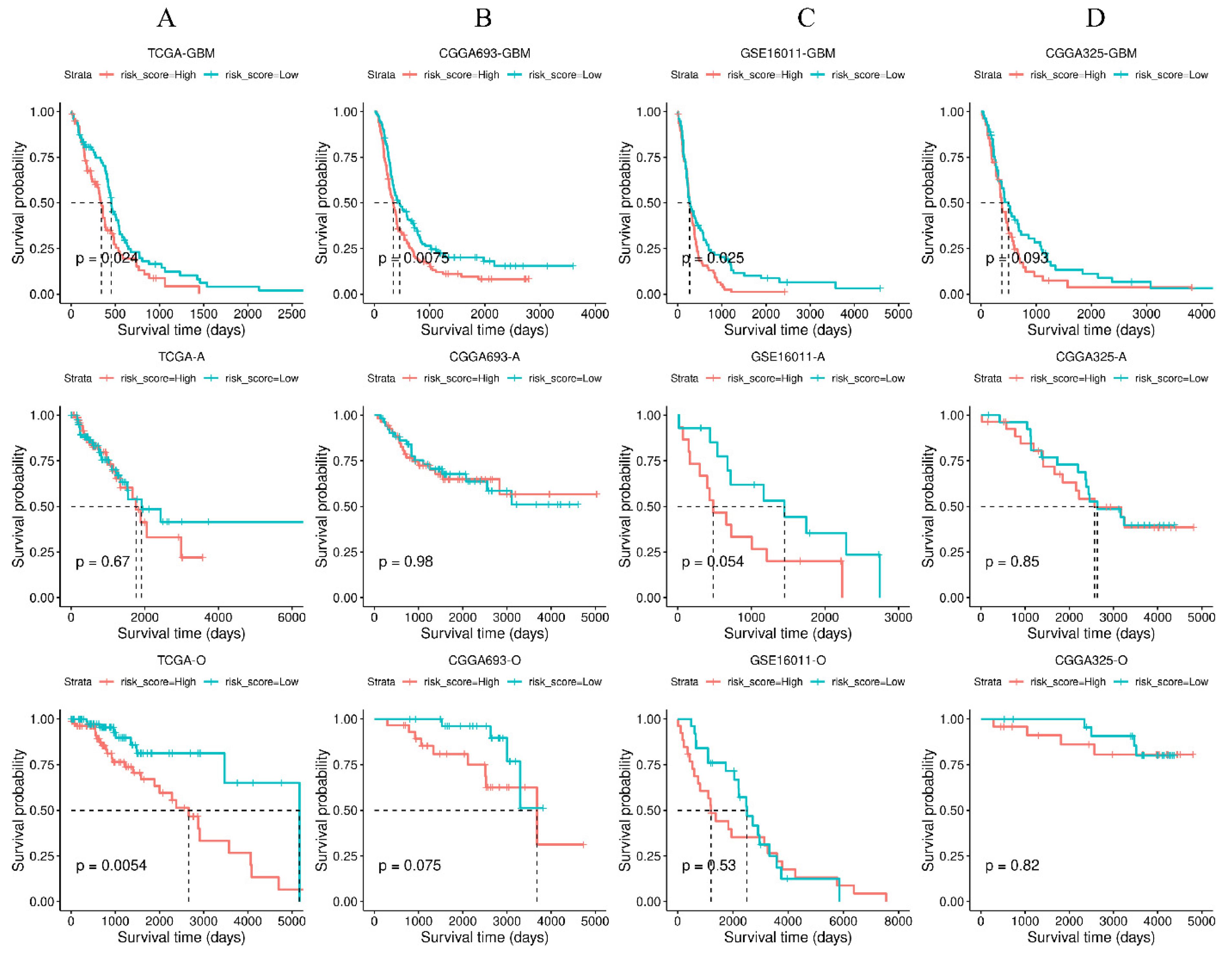
3.6. Validation of the Four-Gene Signature in Independent Cohorts
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Iacob, G.; Dinca, E.B. Current data and strategy in glioblastoma multiforme. J. Med. Life 2009, 2, 386–393. [Google Scholar] [PubMed]
- Affronti, M.L.; Heery, C.R.; Herndon, J.E., 2nd; Rich, J.N.; Reardon, D.A.; Desjardins, A.; Vredenburgh, J.J.; Friedman, A.H.; Bigner, D.D.; Friedman, H.S. Overall survival of newly diagnosed glioblastoma patients receiving carmustine wafers followed by radiation and concurrent temozolomide plus rotational multiagent chemotherapy. Cancer 2009, 115, 3501–3511. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.W.; Hoadley, K.A.; Purdom, E.; Wang, V.; Qi, Y.; Wilkerson, M.D.; Miller, C.R.; Ding, L.; Golub, T.; Mesirov, J.P.; et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010, 17, 98–110. [Google Scholar] [CrossRef] [Green Version]
- Brennan, C.W.; Verhaak, R.G.; McKenna, A.; Campos, B.; Noushmehr, H.; Salama, S.R.; Zheng, S.; Chakravarty, D.; Sanborn, J.Z.; Berman, S.H.; et al. The somatic genomic landscape of glioblastoma. Cell 2013, 155, 462–477. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Li, H.; Luo, K.; Sharma, A.; Sun, X. Prognostic gene expression signature revealed the involvement of mutational pathways in cancer genome. J. Cancer 2020, 11, 4510–4520. [Google Scholar] [CrossRef]
- Xu, P.; Yang, J.; Liu, J.; Yang, X.; Liao, J.; Yuan, F.; Xu, Y.; Liu, B.; Chen, Q. Identification of glioblastoma gene prognosis modules based on weighted gene co-expression network analysis. BMC Med. Genom. 2018, 11, 96. [Google Scholar] [CrossRef] [Green Version]
- Hsu, J.B.; Chang, T.H.; Lee, G.A.; Lee, T.Y.; Chen, C.Y. Identification of potential biomarkers related to glioma survival by gene expression profile analysis. BMC Med. Genom. 2019, 11, 34. [Google Scholar] [CrossRef]
- Lin, S.; Xu, H.; Zhang, A.; Ni, Y.; Xu, Y.; Meng, T.; Wang, M.; Lou, M. Prognosis Analysis and Validation of m(6)A Signature and Tumor Immune Microenvironment in Glioma. Front. Oncol. 2020, 10, 541401. [Google Scholar] [CrossRef]
- Guo, Q.; Xiao, X.; Zhang, J. MYD88 Is a Potential Prognostic Gene and Immune Signature of Tumor Microenvironment for Gliomas. Front. Oncol. 2021, 11, 654388. [Google Scholar] [CrossRef]
- Wang, G.; Zhou, H.; Tian, L.; Yan, T.; Han, X.; Chen, P.; Li, H.; Wang, W.; Xiao, Z.; Hou, L.; et al. A Prognostic DNA Damage Repair Genes Signature and Its Impact on Immune Cell Infiltration in Glioma. Front. Oncol. 2021, 11, 682932. [Google Scholar] [CrossRef]
- Li, Y.; Deng, G.; Zhang, H.; Qi, Y.; Gao, L.; Tan, Y.; Hu, P.; Wang, Y.; Liu, B.; Chen, Q. Weighted gene correlation network analysis identifies microenvironment-related genes signature as prognostic candidate for Grade II/III glioma. Aging 2020, 12, 22122–22138. [Google Scholar] [CrossRef] [PubMed]
- Bangasser, B.L.; Shamsan, G.A.; Chan, C.E.; Opoku, K.N.; Tüzel, E.; Schlichtmann, B.W.; Kasim, J.A.; Fuller, B.J.; McCullough, B.R.; Rosenfeld, S.S.; et al. Shifting the optimal stiffness for cell migration. Nat. Commun. 2017, 8, 15313. [Google Scholar] [CrossRef] [Green Version]
- Wei, S.C.; Fattet, L.; Tsai, J.H.; Guo, Y.; Pai, V.H.; Majeski, H.E.; Chen, A.C.; Sah, R.L.; Taylor, S.S.; Engler, A.J.; et al. Matrix stiffness drives epithelial-mesenchymal transition and tumour metastasis through a TWIST1-G3BP2 mechanotransduction pathway. Nat. Cell Biol. 2015, 17, 678–688. [Google Scholar] [CrossRef]
- Stowers, R.S.; Shcherbina, A.; Israeli, J.; Gruber, J.J.; Chang, J.; Nam, S.; Rabiee, A.; Teruel, M.N.; Snyder, M.P.; Kundaje, A.; et al. Matrix stiffness induces a tumorigenic phenotype in mammary epithelium through changes in chromatin accessibility. Nat. Biomed. Eng. 2019, 3, 1009–1019. [Google Scholar] [CrossRef] [PubMed]
- Micalet, A.; Moeendarbary, E.; Cheema, U. 3D In Vitro Models for Investigating the Role of Stiffness in Cancer Invasion. ACS Biomater. Sci. Eng. 2021. [Google Scholar] [CrossRef] [PubMed]
- Miroshnikova, Y.A.; Mouw, J.K.; Barnes, J.M.; Pickup, M.W.; Lakins, J.N.; Kim, Y.; Lobo, K.; Persson, A.I.; Reis, G.F.; McKnight, T.R.; et al. Tissue mechanics promote IDH1-dependent HIF1α–tenascin C feedback to regulate glioblastoma aggression. Nat. Cell Biol. 2016, 18, 1336–1345. [Google Scholar] [CrossRef]
- Umesh, V.; Rape, A.D.; Ulrich, T.A.; Kumar, S. Microenvironmental stiffness enhances glioma cell proliferation by stimulating epidermal growth factor receptor signaling. PLoS ONE 2014, 9, e101771. [Google Scholar] [CrossRef] [Green Version]
- Tao, B.; Song, Y.; Wu, Y.; Yang, X.; Peng, T.; Peng, L.; Xia, K.; Xia, X.; Chen, L.; Zhong, C. Matrix stiffness promotes glioma cell stemness by activating BCL9L/Wnt/β-catenin signaling. Aging 2021, 13, 5284–5296. [Google Scholar] [CrossRef] [PubMed]
- Erickson, A.E.; Lan Levengood, S.K.; Sun, J.; Chang, F.-C.; Zhang, M. Fabrication and Characterization of Chitosan-Hyaluronic Acid Scaffolds with Varying Stiffness for Glioblastoma Cell Culture. Adv. Healthc. Mater. 2018, 7, e1800295. [Google Scholar] [CrossRef]
- Zhu, D.; Trinh, P.; Li, J.; Grant, G.A.; Yang, F. Gradient hydrogels for screening stiffness effects on patient-derived glioblastoma xenograft cellfates in 3D. J. Biomed. Mater. Research. Part A 2021, 109, 1027–1035. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Zhang, K.N.; Wang, Q.; Li, G.; Zeng, F.; Zhang, Y.; Wu, F.; Chai, R.; Wang, Z.; Zhang, C.; et al. Chinese Glioma Genome Atlas (CGGA): A Comprehensive Resource with Functional Genomic Data from Chinese Glioma Patients. Genom. Proteom. Bioinform. 2021, 19, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.; Tiwari, S.K.; Agrawal, K.; Tan, M.; Dang, J.; Tam, T.; Tian, J.; Wan, X.; Schimelman, J.; You, S.; et al. Rapid 3D Bioprinting of Glioblastoma Model Mimicking Native Biophysical Heterogeneity. Small 2021, 17, e2006050. [Google Scholar] [CrossRef] [PubMed]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets-update. Nucleic. Acids Res. 2013, 41, D991–D995. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [Green Version]
- Colaprico, A.; Silva, T.C.; Olsen, C.; Garofano, L.; Cava, C.; Garolini, D.; Sabedot, T.S.; Malta, T.M.; Pagnotta, S.M.; Castiglioni, I.; et al. TCGAbiolinks: An R/Bioconductor package for integrative analysis of TCGA data. Nucleic. Acids Res. 2016, 44, e71. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Gravendeel, L.; Kouwenhoven, M.; Gevaert, O.; de Rooi, J.; Stubbs, A.; Duijm, E.; Daemen, A.; Bleeker, F.; Bralten, L.; Kloosterhof, N.; et al. Intrinsic gene expression profiles of gliomas are a better predictor of survival than histology. Cancer Res. 2010, 70, 808. [Google Scholar] [CrossRef]
- Ye, Y.; Dai, Q.J.; Qi, H.B. A novel defined pyroptosis-related gene signature for predicting the prognosis of ovarian cancer. Cell Death Discov. 2021, 7, 71. [Google Scholar] [CrossRef]
- Zuo, S.G.; Dai, G.P.; Ren, X.Q. Identification of a 6-gene signature predicting prognosis for colorectal cancer. Cancer Cell Int. 2019, 19, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Y.F.; Wu, J.H.; Huang, W.W.; Weng, S.W.; Wang, B.C.; Chen, Y.M.; Wang, H. Development and validation of a hypoxia-immune-based microenvironment gene signature for risk stratification in gastric cancer. J. Transl. Med. 2020, 18, 201. [Google Scholar] [CrossRef] [PubMed]
- An, N.; Yu, Z.; Yang, X. Expression Differentiation Is Not Helpful in Identifying Prognostic Genes Based on TCGA Datasets. Mol. Ther. Nucleic Acids 2018, 11, 292–299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Louis, D.N.; Perry, A.; Wesseling, P.; Brat, D.J.; Cree, I.A.; Figarella-Branger, D.; Hawkins, C.; Ng, H.K.; Pfister, S.M.; Reifenberger, G.; et al. The 2021 WHO Classification of Tumors of the Central Nervous System: A summary. Neuro Oncol. 2021, 23, 1231–1251. [Google Scholar] [CrossRef] [PubMed]
- Spada, S.; Tocci, A.; Di Modugno, F.; Nisticò, P. Fibronectin as a multiregulatory molecule crucial in tumor matrisome: From structural and functional features to clinical practice in oncology. J. Exp. Clin. Cancer Res. CR 2021, 40, 102. [Google Scholar] [CrossRef] [PubMed]
- Serres, E.; Debarbieux, F.; Stanchi, F.; Maggiorella, L.; Grall, D.; Turchi, L.; Burel-Vandenbos, F.; Figarella-Branger, D.; Virolle, T.; Rougon, G.; et al. Fibronectin expression in glioblastomas promotes cell cohesion, collective invasion of basement membrane in vitro and orthotopic tumor growth in mice. Oncogene 2014, 33, 3451–3462. [Google Scholar] [CrossRef]
- Efthymiou, G.; Saint, A.; Ruff, M.; Rekad, Z.; Ciais, D.; Van Obberghen-Schilling, E. Shaping Up the Tumor Microenvironment With Cellular Fibronectin. Front. Oncol. 2020, 10, 641. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Carraher, C.; Schwarzbauer, J.E. Assembly of fibronectin extracellular matrix. Annu. Rev. Cell Dev. Biol. 2010, 26, 397–419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wierzbicka-Patynowski, I.; Schwarzbauer, J.E. The ins and outs of fibronectin matrix assembly. J. Cell Sci. 2003, 116, 3269–3276. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, C.-W.; Yang, C.-H.; Lin, Y.-H.; Hou, Y.-C.; Cheng, T.-J.; Chang, S.-T.; Huang, Y.-H.; Chung, S.-T.; Chio, C.-C.; Shan, Y.-S.; et al. The Fibronectin Expression Determines the Distinct Progressions of Malignant Gliomas via Transforming Growth Factor-Beta Pathway. Int. J. Mol. Sci. 2021, 22, 3782. [Google Scholar] [CrossRef]
- Yu, Q.; Xiao, W.K.; Sun, S.P.; Sohrabi, A.; Liang, J.; Seidlits, S.K. Extracellular Matrix Proteins Confer Cell Adhesion-Mediated Drug Resistance Through Integrin alpha(v) in Glioblastoma Cells. Front. Cell Dev. Biol. 2021, 9, 616580. [Google Scholar] [CrossRef]
- Yuan, L.; Siegel, M.; Choi, K.; Khosla, C.; Miller, C.R.; Jackson, E.N.; Piwnica-Worms, D.; Rich, K.M. Transglutaminase 2 inhibitor, KCC009, disrupts fibronectin assembly in the extracellular matrix and sensitizes orthotopic glioblastomas to chemotherapy. Oncogene 2007, 26, 2563–2573. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, C.H.; Wang, Y.; Sims, M.; Cai, C.; Pfeffer, L.M. MicroRNA-1 suppresses glioblastoma in preclinical models by targeting fibronectin. Cancer Lett. 2019, 465, 59–67. [Google Scholar] [CrossRef]
- Saw, P.E.; Xu, X.D.; Kang, B.R.; Lee, J.; Lee, Y.S.; Kim, C.; Kim, H.; Kang, S.H.; Na, Y.J.; Moon, H.J.; et al. Extra-domain B of fibronectin as an alternative target for drug delivery and a cancer diagnostic and prognostic biomarker for malignant glioma. Theranostics 2021, 11, 941–957. [Google Scholar] [CrossRef]
- Echavidre, W.; Picco, V.; Faraggi, M.; Montemagno, C. Integrin-alpha v beta 3 as a Therapeutic Target in Glioblastoma: Back to the Future? Pharmaceutics 2022, 14, 1053. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wan, Q.; Yan, W. Integrin α5/ITGA5 Promotes The Proliferation, Migration, Invasion And Progression Of Oral Squamous Carcinoma By Epithelial-Mesenchymal Transition. Cancer Manag. Res. 2019, 11, 9609–9620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuninty, P.R.; Bansal, R.; De Geus, S.W.L.; Mardhian, D.F.; Schnittert, J.; van Baarlen, J.; Storm, G.; Bijlsma, M.F.; van Laarhoven, H.W.; Metselaar, J.M.; et al. ITGA5 inhibition in pancreatic stellate cells attenuates desmoplasia and potentiates efficacy of chemotherapy in pancreatic cancer. Sci. Adv. 2019, 5, eaax2770. [Google Scholar] [CrossRef] [Green Version]
- Zhu, H.; Wang, G.; Zhu, H.; Xu, A. ITGA5 is a prognostic biomarker and correlated with immune infiltration in gastrointestinal tumors. BMC Cancer 2021, 21, 269. [Google Scholar] [CrossRef]
- Mallawaaratchy, D.M.; Buckland, M.E.; McDonald, K.L.; Li, C.C.Y.; Ly, L.; Sykes, E.K.; Christopherson, R.I.; Kaufman, K.L. Membrane proteome analysis of glioblastoma cell invasion. J. Neuropathol. Exp. Neurol. 2015, 74, 425–441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, Y.; Wu, M.; Liu, Y.; Hu, L.; Wu, H.; Xie, L.; Liu, Z.; Wu, A.; Chen, L.; Xu, C. ITGA5 Predicts Dual-Drug Resistance to Temozolomide and Bevacizumab in Glioma. Front. Oncol. 2021, 11, 769592. [Google Scholar] [CrossRef]
- Zhao, L.; Chen, H.J.; Guo, Z.D.; Fu, K.L.; Yao, L.L.; Fu, L.; Guo, W.X.; Wen, X.J.; Jacobson, O.; Zhang, X.Z.; et al. Targeted Radionuclide Therapy in Patient-Derived Xenografts Using Lu-177-EB-RGD. Mol. Cancer Ther. 2020, 19, 2034–2043. [Google Scholar] [CrossRef]
- Guo, Q.; Guan, G.-F.; Cao, J.-Y.; Zou, C.-Y.; Zhu, C.; Cheng, W.; Xu, X.-Y.; Lin, Z.-G.; Cheng, P.; Wu, A.-H. Overexpression of oncostatin M receptor regulates local immune response in glioblastoma. J. Cell. Physiol. 2019, 234, 15496–15509. [Google Scholar] [CrossRef] [PubMed]
- Jahani-Asl, A.; Yin, H.; Soleimani, V.D.; Haque, T.; Luchman, H.A.; Chang, N.C.; Sincennes, M.-C.; Puram, S.V.; Scott, A.M.; Lorimer, I.A.J.; et al. Control of glioblastoma tumorigenesis by feed-forward cytokine signaling. Nat. Neurosci. 2016, 19, 798–806. [Google Scholar] [CrossRef] [Green Version]
- Greenall, S.A.; Johns, T.G. EGFRvIII: The promiscuous mutation. Cell Death Discov. 2016, 2, 16049. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharanek, A.; Burban, A.; Laaper, M.; Heckel, E.; Joyal, J.-S.; Soleimani, V.D.; Jahani-Asl, A. OSMR controls glioma stem cell respiration and confers resistance of glioblastoma to ionizing radiation. Nat. Commun. 2020, 11, 4116. [Google Scholar] [CrossRef] [PubMed]
- Johnston, A.L.M.; Lun, X.; Rahn, J.J.; Liacini, A.; Wang, L.; Hamilton, M.G.; Parney, I.F.; Hempstead, B.L.; Robbins, S.M.; Forsyth, P.A.; et al. The p75 neurotrophin receptor is a central regulator of glioma invasion. PLoS Biol. 2007, 5, e212. [Google Scholar] [CrossRef]
- Zhou, X.; Hao, Q.; Liao, P.; Luo, S.; Zhang, M.; Hu, G.; Liu, H.; Zhang, Y.; Cao, B.; Baddoo, M.; et al. Nerve growth factor receptor negates the tumor suppressor p53 as a feedback regulator. eLife 2016, 5, e15099. [Google Scholar] [CrossRef] [Green Version]
- Parsi, S.; Soltani, B.M.; Hosseini, E.; Tousi, S.E.; Mowla, S.J. Experimental verification of a predicted intronic microRNA in human NGFR gene with a potential pro-apoptotic function. PLoS ONE 2012, 7, e35561. [Google Scholar] [CrossRef]
- DeSisto, J.A.; Flannery, P.; Lemma, R.; Pathak, A.; Mestnik, S.; Philips, N.; Bales, N.J.; Kashyap, T.; Moroze, E.; Venkataraman, S.; et al. Exportin 1 Inhibition Induces Nerve Growth Factor Receptor Expression to Inhibit the NF-κB Pathway in Preclinical Models of Pediatric High-Grade Glioma. Mol. Cancer Ther. 2020, 19, 540–551. [Google Scholar] [CrossRef] [Green Version]
- Boshuizen, J.; Vredevoogd, D.W.; Krijgsman, O.; Ligtenberg, M.A.; Blankenstein, S.; de Bruijn, B.; Frederick, D.T.; Kenski, J.C.N.; Parren, M.; Bruggemann, M.; et al. Reversal of pre-existing NGFR-driven tumor and immune therapy resistance. Nat. Commun. 2020, 11, 3946. [Google Scholar] [CrossRef]
- Prasad, B.; Tian, Y.; Li, X. Large-Scale Analysis Reveals Gene Signature for Survival Prediction in Primary Glioblastoma. Mol. Neurobiol. 2020, 57, 5235–5246. [Google Scholar] [CrossRef]
- Xiao, K.; Liu, Q.; Peng, G.; Su, J.; Qin, C.Y.; Wang, X.Y. Identification and validation of a three-gene signature as a candidate prognostic biomarker for lower grade glioma. PeerJ 2020, 8, e8312. [Google Scholar] [CrossRef]
- Liu, J.; Li, G. Identification and validation of a risk signature based on extracellular matrix-related genes in gliomas. Medicine 2021, 100, e25603. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.; Liu, S.; Qi, F.; Li, X.; Yu, S.; Feng, J.; Zheng, Y. Screening TCGA database for prognostic genes in lower grade glioma microenvironment. Ann. Transl. Med. 2020, 8, 209. [Google Scholar] [CrossRef]
- Gong, X.; Liu, L.; Xiong, J.; Li, X.; Xu, J.; Xiao, Y.; Li, J.; Luo, X.; Mao, D.; Liu, L. Construction of a Prognostic Gene Signature Associated with Immune Infiltration in Glioma: A Comprehensive Analysis Based on the CGGA. J. Oncol. 2021, 2021, 6620159. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, C.-R.; Chang, R.-S.; Chen, C.-S. Identification of Prognostic Genes in Gliomas Based on Increased Microenvironment Stiffness. Cancers 2022, 14, 3659. https://doi.org/10.3390/cancers14153659
Chen C-R, Chang R-S, Chen C-S. Identification of Prognostic Genes in Gliomas Based on Increased Microenvironment Stiffness. Cancers. 2022; 14(15):3659. https://doi.org/10.3390/cancers14153659
Chicago/Turabian StyleChen, Chaang-Ray, Rong-Shing Chang, and Chi-Shuo Chen. 2022. "Identification of Prognostic Genes in Gliomas Based on Increased Microenvironment Stiffness" Cancers 14, no. 15: 3659. https://doi.org/10.3390/cancers14153659
APA StyleChen, C.-R., Chang, R.-S., & Chen, C.-S. (2022). Identification of Prognostic Genes in Gliomas Based on Increased Microenvironment Stiffness. Cancers, 14(15), 3659. https://doi.org/10.3390/cancers14153659






