Identification of a Prognostic Microenvironment-Related Gene Signature in Glioblastoma Patients Treated with Carmustine Wafers
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Surgical Procedure
2.2. Isolation and Culture of Glioma Associated Stem Cells (GASC)
2.3. GASC RNA Extraction, Library Preparation and Sequencing
2.4. Functional Enrichment Analysis
2.5. RT2 Profiler™ PCR Array
2.6. Differential Gene Expression Analysis in the TCGA-GBM Dataset
2.7. Survival Analysis
3. Results
3.1. Transcriptomic Characterization of GASC through RNA-Seq
3.2. A Customized RT-PCR Array Identified Four Genes Specifically Modulated in GASC-LS
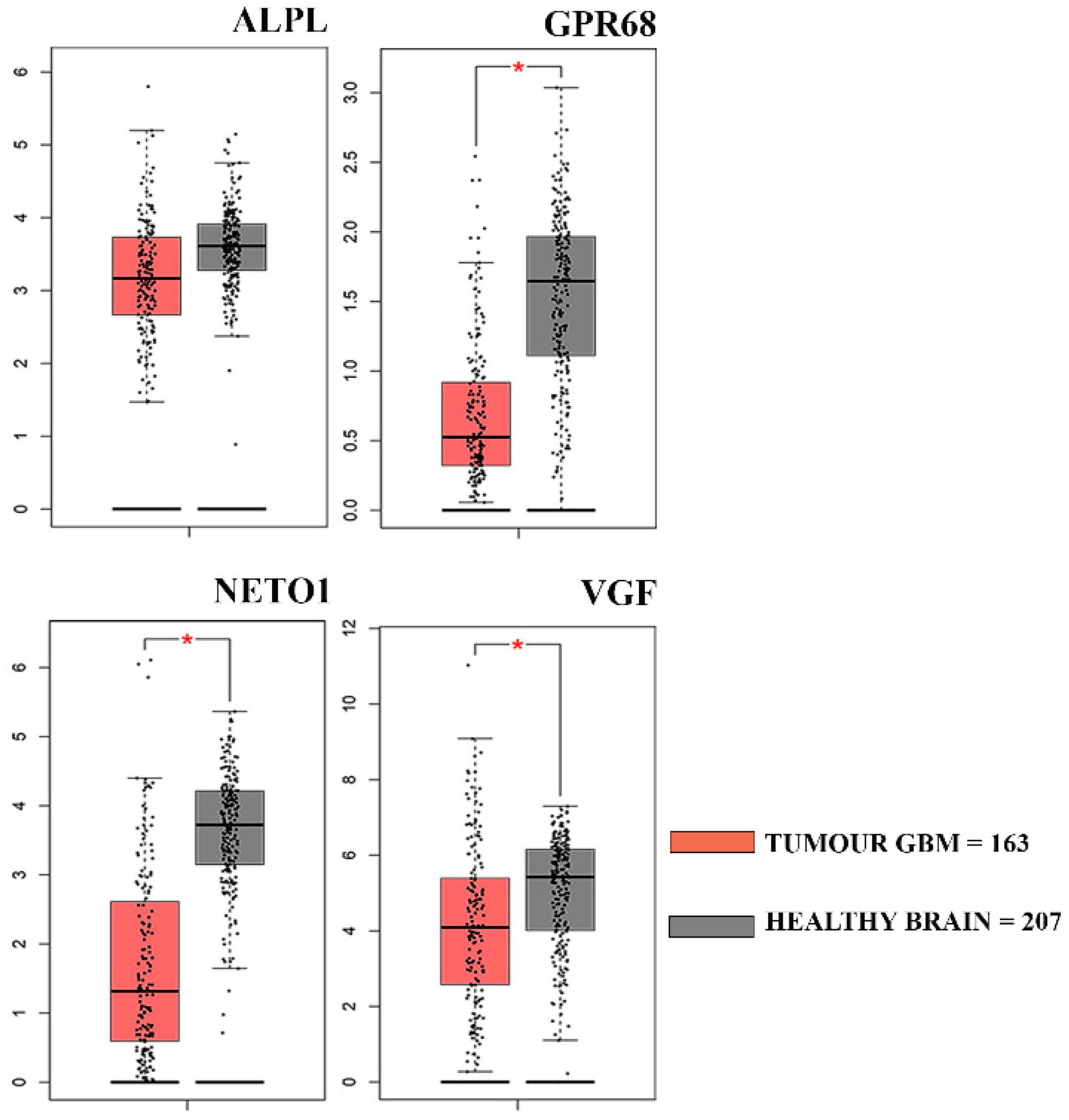
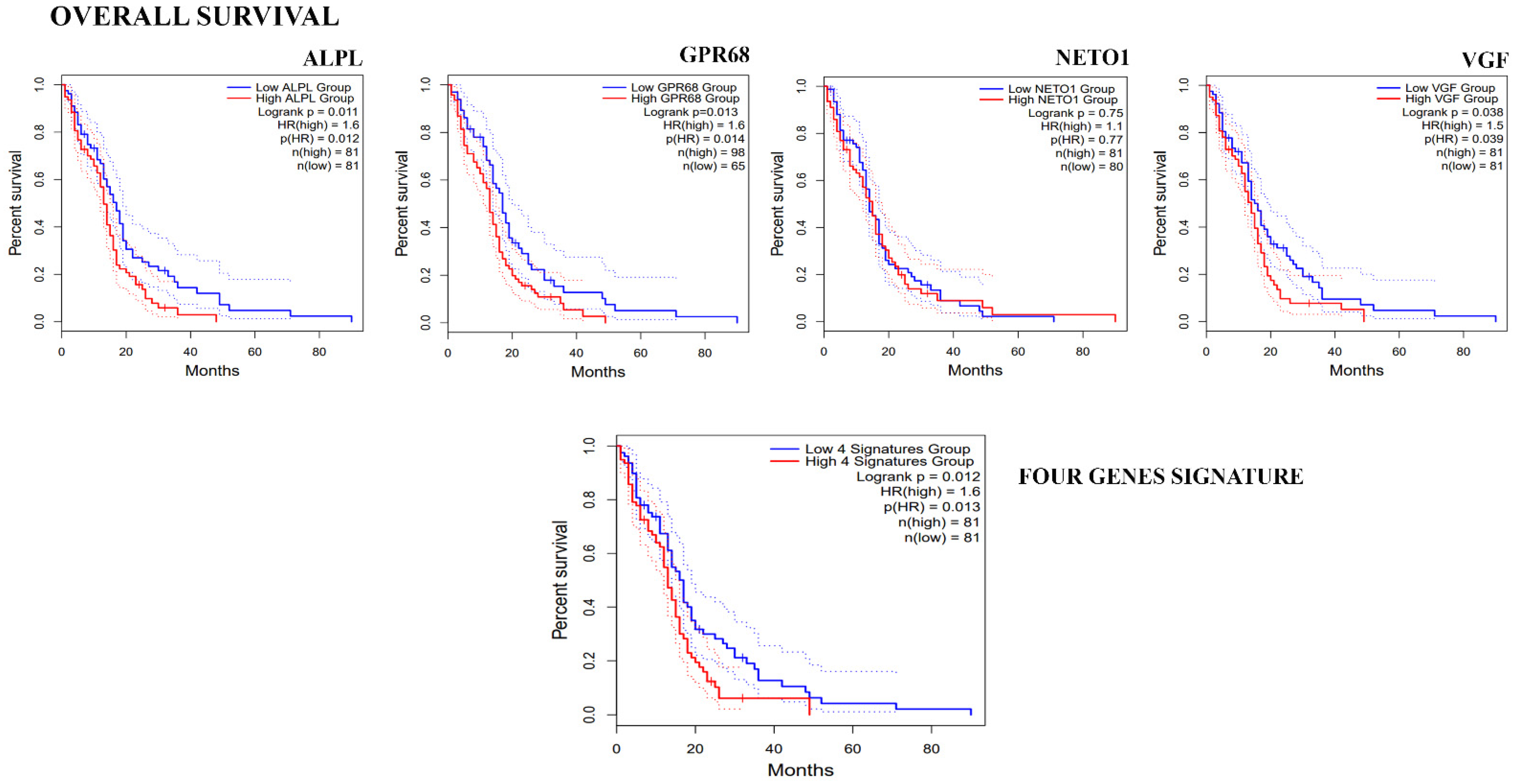
3.3. Clinical Correlation of Genes Down-Regulated in GASC-LS
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Stupp, R.; Mason, W.P.; van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.B.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus Concomitant and Adjuvant Temozolomide for Glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef] [PubMed]
- Ostrom, Q.T.; Gittleman, H.; Liao, P.; Rouse, C.; Chen, Y.; Dowling, J.; Wolinsky, Y.; Kruchko, C.; Barnholtz-Sloan, J. CBTRUS Statistical Report: Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2007–2011. Neuro Oncol. 2014, 16 (Suppl. S4), iv1–iv63. [Google Scholar] [CrossRef] [PubMed]
- Lara-Velazquez, M.; Al-Kharboosh, R.; Jeanneret, S.; Vazquez-Ramos, C.; Mahato, D.; Tavanaiepour, D.; Rahmathulla, G.; Quinones-Hinojosa, A. Advances in Brain Tumor Surgery for Glioblastoma in Adults. Brain Sci. 2017, 7, 166. [Google Scholar] [CrossRef] [PubMed]
- Quail, D.F.; Joyce, J.A. Microenvironmental Regulation of Tumor Progression and Metastasis. Nat. Med. 2013, 19, 1423–1437. [Google Scholar] [CrossRef] [PubMed]
- Jhaveri, N.; Chen, T.C.; Hofman, F.M. Tumor Vasculature and Glioma Stem Cells: Contributions to Glioma Progression. Cancer Lett. 2016, 380, 545–551. [Google Scholar] [CrossRef] [PubMed]
- Lathia, J.D.; Heddleston, J.M.; Venere, M.; Rich, J.N. Deadly Teamwork: Neural Cancer Stem Cells and the Tumor Microenvironment. Cell Stem Cell 2011, 8, 482–485. [Google Scholar] [CrossRef]
- Current Treatments for Brain Tumors. Available online: https://www.bing.com/search?q=Current+Treatments+for+Brain+Tumors.+Available+online%3A+https%3A%2F%2Fbraintumor.org (accessed on 19 March 2022).
- Drugs Approved for Brain Tumors—National Cancer Institute. Available online: https://www.cancer.gov/about-cancer/treatment/drugs/brain (accessed on 19 March 2022).
- Bota, D.A.; Desjardins, A.; Quinn, J.A.; Affronti, M.L.; Friedman, H.S. Interstitial Chemotherapy with Biodegradable BCNU (Gliadel) Wafers in the Treatment of Malignant Gliomas. Ther. Clin. Risk Manag. 2007, 3, 707–715. [Google Scholar]
- Shibahara, I.; Hanihara, M.; Watanabe, T.; Dan, M.; Sato, S.; Kuroda, H.; Inamura, A.; Inukai, M.; Hara, A.; Yasui, Y.; et al. Tumor Microenvironment after Biodegradable BCNU Wafer Implantation: Special Consideration of Immune System. J. Neurooncol. 2018, 137, 417–427. [Google Scholar] [CrossRef]
- Han, S.; Zhang, C.; Li, Q.; Dong, J.; Liu, Y.; Huang, Y.; Jiang, T.; Wu, A. Tumour-Infiltrating CD4+ and CD8+ Lymphocytes as Predictors of Clinical Outcome in Glioma. Br. J. Cancer 2014, 110, 2560–2568. [Google Scholar] [CrossRef]
- Brem, H.; Tamargo, R.J.; Olivi, A.; Pinn, M.; Weingart, J.D.; Wharam, M.; Epstein, J.I. Biodegradable Polymers for Controlled Delivery of Chemotherapy with and without Radiation Therapy in the Monkey Brain. J. Neurosurg. 1994, 80, 283–290. [Google Scholar] [CrossRef]
- Xing, W.; Shao, C.; Qi, Z.; Yang, C.; Wang, Z. The Role of Gliadel Wafers in the Treatment of Newly Diagnosed GBM: A Meta-Analysis. Drug Des. Dev. Ther. 2015, 9, 3341–3348. [Google Scholar] [CrossRef]
- Ashby, L.S.; Smith, K.A.; Stea, B. Gliadel Wafer Implantation Combined with Standard Radiotherapy and Concurrent Followed by Adjuvant Temozolomide for Treatment of Newly Diagnosed High-Grade Glioma: A Systematic Literature Review. World J. Surg. Oncol. 2016, 14, 225. [Google Scholar] [CrossRef] [PubMed]
- Bregy, A.; Shah, A.H.; Diaz, M.V.; Pierce, H.E.; Ames, P.L.; Diaz, D.; Komotar, R.J. The Role of Gliadel Wafers in the Treatment of High-Grade Gliomas. Expert Rev. Anticancer Ther. 2013, 13, 1453–1461. [Google Scholar] [CrossRef]
- Westphal, M.; Hilt, D.C.; Bortey, E.; Delavault, P.; Olivares, R.; Warnke, P.C.; Whittle, I.R.; Jääskeläinen, J.; Ram, Z. A Phase 3 Trial of Local Chemotherapy with Biodegradable Carmustine (BCNU) Wafers (Gliadel Wafers) in Patients with Primary Malignant Glioma. Neuro Oncol. 2003, 5, 79–88. [Google Scholar] [CrossRef]
- Attenello, F.J.; Mukherjee, D.; Datoo, G.; McGirt, M.J.; Bohan, E.; Weingart, J.D.; Olivi, A.; Quinones-Hinojosa, A.; Brem, H. Use of Gliadel (BCNU) Wafer in the Surgical Treatment of Malignant Glioma: A 10-Year Institutional Experience. Ann. Surg. Oncol. 2008, 15, 2887–2893. [Google Scholar] [CrossRef]
- McGirt, M.J.; Than, K.D.; Weingart, J.D.; Chaichana, K.L.; Attenello, F.J.; Olivi, A.; Laterra, J.; Kleinberg, L.R.; Grossman, S.A.; Brem, H.; et al. Gliadel (BCNU) Wafer plus Concomitant Temozolomide Therapy after Primary Resection of Glioblastoma Multiforme. J. Neurosurg. 2009, 110, 583–588. [Google Scholar] [CrossRef] [PubMed]
- Ius, T.; Cesselli, D.; Isola, M.; Toniato, G.; Pauletto, G.; Sciacca, G.; Fabbro, S.; Pegolo, E.; Rizzato, S.; Beltrami, A.P.; et al. Combining Clinical and Molecular Data to Predict the Benefits of Carmustine Wafers in Newly Diagnosed High-Grade Gliomas. Curr. Treat. Options Neurol. 2018, 20, 3. [Google Scholar] [CrossRef]
- National Research Council (US) Committee on a Framework for Developing a New Taxonomy of Disease. Toward Precision Medicine: Building a Knowledge Network for Biomedical Research and a New Taxonomy of Disease; The National Academies Collection: Reports Funded by National Institutes of Health; National Academies Press (US): Washington, DC, USA, 2011; ISBN 978-0-309-22222-8. [Google Scholar]
- Eckel-Passow, J.E.; Lachance, D.H.; Molinaro, A.M.; Walsh, K.M.; Decker, P.A.; Sicotte, H.; Pekmezci, M.; Rice, T.; Kosel, M.L.; Smirnov, I.V.; et al. Glioma Groups Based on 1p/19q, IDH, and TERT Promoter Mutations in Tumors. N. Engl. J. Med. 2015, 372, 2499–2508. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Research Network; Brat, D.J.; Verhaak, R.G.W.; Aldape, K.D.; Yung, W.K.A.; Salama, S.R.; Cooper, L.A.D.; Rheinbay, E.; Miller, C.R.; Vitucci, M.; et al. Comprehensive, Integrative Genomic Analysis of Diffuse Lower-Grade Gliomas. N. Engl. J. Med. 2015, 372, 2481–2498. [Google Scholar] [CrossRef]
- Ceccarelli, M.; Barthel, F.P.; Malta, T.M.; Sabedot, T.S.; Salama, S.R.; Murray, B.A.; Morozova, O.; Newton, Y.; Radenbaugh, A.; Pagnotta, S.M.; et al. Molecular Profiling Reveals Biologically Discrete Subsets and Pathways of Progression in Diffuse Glioma. Cell 2016, 164, 550–563. [Google Scholar] [CrossRef]
- Sankar, P.L.; Parker, L.S. The Precision Medicine Initiative’s All of Us Research Program: An Agenda for Research on Its Ethical, Legal, and Social Issues. Genet Med 2017, 19, 743–750. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Beltrami, A.P.; Cesselli, D.; Bergamin, N.; Marcon, P.; Rigo, S.; Puppato, E.; D’Aurizio, F.; Verardo, R.; Piazza, S.; Pignatelli, A.; et al. Multipotent Cells Can Be Generated in Vitro from Several Adult Human Organs (Heart, Liver, and Bone Marrow). Blood 2007, 110, 3438–3446. [Google Scholar] [CrossRef]
- Bourkoula, E.; Mangoni, D.; Ius, T.; Pucer, A.; Isola, M.; Musiello, D.; Marzinotto, S.; Toffoletto, B.; Sorrentino, M.; Palma, A.; et al. Glioma-Associated Stem Cells: A Novel Class of Tumor-Supporting Cells Able to Predict Prognosis of Human Low-Grade Gliomas. Stem Cells 2014, 32, 1239–1253. [Google Scholar] [CrossRef] [PubMed]
- Manini, I.; Ruaro, M.E.; Sgarra, R.; Bartolini, A.; Caponnetto, F.; Ius, T.; Skrap, M.; Di Loreto, C.; Beltrami, A.P.; Manfioletti, G.; et al. Semaphorin-7A on Exosomes: A Promigratory Signal in the Glioma Microenvironment. Cancers 2019, 11, 758. [Google Scholar] [CrossRef] [PubMed]
- Ius, T.; Ciani, Y.; Ruaro, M.E.; Isola, M.; Sorrentino, M.; Bulfoni, M.; Candotti, V.; Correcig, C.; Bourkoula, E.; Manini, I.; et al. An NF-ΚB Signature Predicts Low-Grade Glioma Prognosis: A Precision Medicine Approach Based on Patient-Derived Stem Cells. Neuro Oncol. 2018, 20, 776–787. [Google Scholar] [CrossRef]
- Manini, I.; Caponnetto, F.; Dalla, E.; Ius, T.; Della Pepa, G.M.; Pegolo, E.; Bartolini, A.; La Rocca, G.; Menna, G.; Di Loreto, C.; et al. Heterogeneity Matters: Different Regions of Glioblastoma Are Characterized by Distinctive Tumor-Supporting Pathways. Cancers 2020, 12, 2960. [Google Scholar] [CrossRef]
- Caponnetto, F.; Dalla, E.; Mangoni, D.; Piazza, S.; Radovic, S.; Ius, T.; Skrap, M.; Di Loreto, C.; Beltrami, A.P.; Manini, I.; et al. The MiRNA Content of Exosomes Released from the Glioma Microenvironment Can Affect Malignant Progression. Biomedicines 2020, 8, 564. [Google Scholar] [CrossRef]
- Babraham Bioinformatics—FastQC a Quality Control Tool for High Throughput Sequence Data. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 19 March 2022).
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize Analysis Results for Multiple Tools and Samples in a Single Report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie Enables Improved Reconstruction of a Transcriptome from RNA-Seq Reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Durinck, S.; Spellman, P.T.; Birney, E.; Huber, W. Mapping Identifiers for the Integration of Genomic Datasets with the R/Bioconductor Package BiomaRt. Nat. Protoc. 2009, 4, 1184–1191. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-D.; Dai, R.-Y.; Chen, Z.; Zhang, Y.-H.; He, X.-Z.; Zhou, J. Efficacy and Safety of Carmustine Wafers in the Treatment of Glioblastoma Multiforme: A Systematic Review. Turk. Neurosurg. 2014, 24, 639–645. [Google Scholar] [CrossRef]
- Gutenberg, A.; Lumenta, C.B.; Braunsdorf, W.E.K.; Sabel, M.; Mehdorn, H.M.; Westphal, M.; Giese, A. The Combination of Carmustine Wafers and Temozolomide for the Treatment of Malignant Gliomas. A Comprehensive Review of the Rationale and Clinical Experience. J. Neurooncol. 2013, 113, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Catacuzzeno, L.; Franciolini, F. Role of KCa3.1 Channels in Modulating Ca2+ Oscillations during Glioblastoma Cell Migration and Invasion. Int. J. Mol. Sci. 2018, 19, 2970. [Google Scholar] [CrossRef]
- Cuddapah, V.A.; Turner, K.L.; Seifert, S.; Sontheimer, H. Bradykinin-Induced Chemotaxis of Human Gliomas Requires the Activation of KCa3.1 and ClC-3. J. Neurosci. 2013, 33, 1427–1440. [Google Scholar] [CrossRef]
- Venkatesh, H.S.; Morishita, W.; Geraghty, A.C.; Silverbush, D.; Gillespie, S.M.; Arzt, M.; Tam, L.T.; Espenel, C.; Ponnuswami, A.; Ni, L.; et al. Electrical and Synaptic Integration of Glioma into Neural Circuits. Nature 2019, 573, 539–545. [Google Scholar] [CrossRef]
- Monje, M.; Borniger, J.C.; D’Silva, N.J.; Deneen, B.; Dirks, P.B.; Fattahi, F.; Frenette, P.S.; Garzia, L.; Gutmann, D.H.; Hanahan, D.; et al. Roadmap for the Emerging Field of Cancer Neuroscience. Cell 2020, 181, 219–222. [Google Scholar] [CrossRef]
- Panyi, G.; Beeton, C.; Felipe, A. Ion Channels and Anti-Cancer Immunity. Philos. Trans. R. Soc. Lond B Biol. Sci. 2014, 369, 20130106. [Google Scholar] [CrossRef] [PubMed]
- Kale, V.P.; Amin, S.G.; Pandey, M.K. Targeting Ion Channels for Cancer Therapy by Repurposing the Approved Drugs. Biochim Biophys. Acta 2015, 1848, 2747–2755. [Google Scholar] [CrossRef] [PubMed]
- Bagal, S.K.; Brown, A.D.; Cox, P.J.; Omoto, K.; Owen, R.M.; Pryde, D.C.; Sidders, B.; Skerratt, S.E.; Stevens, E.B.; Storer, R.I.; et al. Ion Channels as Therapeutic Targets: A Drug Discovery Perspective. J. Med. Chem. 2013, 56, 593–624. [Google Scholar] [CrossRef]
- Arvanitis, C.D.; Ferraro, G.B.; Jain, R.K. The Blood-Brain Barrier and Blood-Tumour Barrier in Brain Tumours and Metastases. Nat. Rev. Cancer 2020, 20, 26–41. [Google Scholar] [CrossRef] [PubMed]
- Alphandéry, E. Nano-Therapies for Glioblastoma Treatment. Cancers 2020, 12, 242. [Google Scholar] [CrossRef]
- Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Ichisaka, T.; Tomoda, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Adult Human Fibroblasts by Defined Factors. Cell 2007, 131, 861–872. [Google Scholar] [CrossRef] [PubMed]
- Rico, L.G.; Juncà, J.; Ward, M.D.; Bradford, J.; Petriz, J. Is Alkaline Phosphatase the Smoking Gun for Highly Refractory Primitive Leukemic Cells? Oncotarget 2016, 7, 72057–72066. [Google Scholar] [CrossRef][Green Version]
- Vassilatis, D.K.; Hohmann, J.G.; Zeng, H.; Li, F.; Ranchalis, J.E.; Mortrud, M.T.; Brown, A.; Rodriguez, S.S.; Weller, J.R.; Wright, A.C.; et al. The G Protein-Coupled Receptor Repertoires of Human and Mouse. Proc. Natl. Acad. Sci. USA 2003, 100, 4903–4908. [Google Scholar] [CrossRef]
- Lynch, J.R.; Wang, J.Y. G Protein-Coupled Receptor Signaling in Stem Cells and Cancer. Int. J. Mol. Sci. 2016, 17, 707. [Google Scholar] [CrossRef]
- O’Hayre, M.; Degese, M.S.; Gutkind, J.S. Novel Insights into G Protein and G Protein-Coupled Receptor Signaling in Cancer. Curr. Opin. Cell Biol. 2014, 27, 126–135. [Google Scholar] [CrossRef]
- Vaupel, P.; Kallinowski, F.; Okunieff, P. Blood Flow, Oxygen and Nutrient Supply, and Metabolic Microenvironment of Human Tumors: A Review. Cancer Res. 1989, 49, 6449–6465. [Google Scholar] [PubMed]
- Gatenby, R.A.; Gillies, R.J. Why Do Cancers Have High Aerobic Glycolysis? Nat. Rev. Cancer 2004, 4, 891–899. [Google Scholar] [CrossRef] [PubMed]
- Cairns, R.; Papandreou, I.; Denko, N. Overcoming Physiologic Barriers to Cancer Treatment by Molecularly Targeting the Tumor Microenvironment. Mol. Cancer Res. 2006, 4, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Damaghi, M.; Wojtkowiak, J.W.; Gillies, R.J. PH Sensing and Regulation in Cancer. Front. Physiol. 2013, 4, 370. [Google Scholar] [CrossRef]
- Justus, C.R.; Dong, L.; Yang, L.V. Acidic Tumor Microenvironment and PH-Sensing G Protein-Coupled Receptors. Front. Physiol. 2013, 4, 354. [Google Scholar] [CrossRef]
- Insel, P.A.; Sriram, K.; Wiley, S.Z.; Wilderman, A.; Katakia, T.; McCann, T.; Yokouchi, H.; Zhang, L.; Corriden, R.; Liu, D.; et al. GPCRomics: GPCR Expression in Cancer Cells and Tumors Identifies New, Potential Biomarkers and Therapeutic Targets. Front. Pharmacol. 2018, 9, 431. [Google Scholar] [CrossRef]
- Wiley, S.Z.; Sriram, K.; Liang, W.; Chang, S.E.; French, R.; McCann, T.; Sicklick, J.; Nishihara, H.; Lowy, A.M.; Insel, P.A. GPR68, a Proton-Sensing GPCR, Mediates Interaction of Cancer-Associated Fibroblasts and Cancer Cells. FASEB J. 2018, 32, 1170–1183. [Google Scholar] [CrossRef]
- Zhang, W.; St-Gelais, F.; Grabner, C.P.; Trinidad, J.C.; Sumioka, A.; Morimoto-Tomita, M.; Kim, K.S.; Straub, C.; Burlingame, A.L.; Howe, J.R.; et al. A Transmembrane Accessory Subunit That Modulates Kainate-Type Glutamate Receptors. Neuron 2009, 61, 385–396. [Google Scholar] [CrossRef]
- Copits, B.A.; Swanson, G.T. Dancing Partners at the Synapse: Auxiliary Subunits That Shape Kainate Receptor Function. Nat. Rev. Neurosci. 2012, 13, 675–686. [Google Scholar] [CrossRef]
- Copits, B.A.; Robbins, J.S.; Frausto, S.; Swanson, G.T. Synaptic Targeting and Functional Modulation of GluK1 Kainate Receptors by the Auxiliary Neuropilin and Tolloid-like (NETO) Proteins. J. Neurosci. 2011, 31, 7334–7340. [Google Scholar] [CrossRef]
- Straub, C.; Hunt, D.L.; Yamasaki, M.; Kim, K.S.; Watanabe, M.; Castillo, P.E.; Tomita, S. Distinct Functions of Kainate Receptors in the Brain Are Determined by the Auxiliary Subunit Neto1. Nat. Neurosci. 2011, 14, 866–873. [Google Scholar] [CrossRef] [PubMed]
- Bowie, D. Ionotropic Glutamate Receptors & CNS Disorders. CNS Neurol. Disord. Drug Targets 2008, 7, 129–143. [Google Scholar] [CrossRef]
- Vincent, P.; Mulle, C. Kainate Receptors in Epilepsy and Excitotoxicity. Neuroscience 2009, 158, 309–323. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wang, W.; Chen, J.; Mao, H.; Liu, Y.; Gu, S.; Liu, Q.; Xi, Q.; Shi, W. High Neuropilin and Tolloid-like 1 Expression Associated with Metastasis and Poor Survival in Epithelial Ovarian Cancer via Regulation of Actin Cytoskeleton. J. Cell. Mol. Med. 2020, 24, 9114–9124. [Google Scholar] [CrossRef] [PubMed]
- Levi, A.; Eldridge, J.D.; Paterson, B.M. Molecular Cloning of a Gene Sequence Regulated by Nerve Growth Factor. Science 1985, 229, 393–395. [Google Scholar] [CrossRef]
- Levi, A.; Ferri, G.-L.; Watson, E.; Possenti, R.; Salton, S.R.J. Processing, Distribution, and Function of VGF, a Neuronal and Endocrine Peptide Precursor. Cell. Mol. Neurobiol. 2004, 24, 517–533. [Google Scholar] [CrossRef]
- Watson, E.; Fargali, S.; Okamoto, H.; Sadahiro, M.; Gordon, R.E.; Chakraborty, T.; Sleeman, M.W.; Salton, S.R. Analysis of Knockout Mice Suggests a Role for VGF in the Control of Fat Storage and Energy Expenditure. BMC Physiol. 2009, 9, 19. [Google Scholar] [CrossRef] [PubMed]
- Hunsberger, J.G.; Newton, S.S.; Bennett, A.H.; Duman, C.H.; Russell, D.S.; Salton, S.R.; Duman, R.S. Antidepressant Actions of the Exercise-Regulated Gene VGF. Nat. Med. 2007, 13, 1476–1482. [Google Scholar] [CrossRef]
- Hwang, W.; Chiu, Y.-F.; Kuo, M.-H.; Lee, K.-L.; Lee, A.-C.; Yu, C.-C.; Chang, J.-L.; Huang, W.-C.; Hsiao, S.-H.; Lin, S.-E.; et al. Expression of Neuroendocrine Factor VGF in Lung Cancer Cells Confers Resistance to EGFR Kinase Inhibitors and Triggers Epithelial-to-Mesenchymal Transition. Cancer Res. 2017, 77, 3013–3026. [Google Scholar] [CrossRef]
- Mani, S.A.; Guo, W.; Liao, M.-J.; Eaton, E.N.; Ayyanan, A.; Zhou, A.Y.; Brooks, M.; Reinhard, F.; Zhang, C.C.; Shipitsin, M.; et al. The Epithelial-Mesenchymal Transition Generates Cells with Properties of Stem Cells. Cell 2008, 133, 704–715. [Google Scholar] [CrossRef]
- Wang, X.; Prager, B.C.; Wu, Q.; Kim, L.J.Y.; Gimple, R.C.; Shi, Y.; Yang, K.; Morton, A.R.; Zhou, W.; Zhu, Z.; et al. Reciprocal Signaling between Glioblastoma Stem Cells and Differentiated Tumor Cells Promotes Malignant Progression. Cell Stem Cell 2018, 22, 514–528.e5. [Google Scholar] [CrossRef] [PubMed]

| GASC-LS UP | |||
|---|---|---|---|
| Gene Set Name | #Genes in Gene Set | #Genes in the Overlap | p-Value |
| REGULATION OF ION TRANSPORT | 1314 | 27 | 4.36 × 10−10 |
| NEUROGENESIS | 1613 | 30 | 4.52 × 10−10 |
| NOTCH_SIGNALING_PATHWAY | 186 | 11 | 2.59 × 10−9 |
| REGULATION_OF_TRANSPORT | 1730 | 29 | 9.28 × 10−9 |
| REGULATION_OF_CELL_DIFFERENTIATION | 1618 | 28 | 8.74 × 10−9 |
| NEURON_DIFFERENTIATION | 1357 | 25 | 1.77 × 10−8 |
| BMI1_DN_MEL18_DN.V1_DN | 147 | 9 | 5.55 × 10−8 |
| SECRETION | 1464 | 25 | 7.61 × 10−8 |
| MORF_DCC | 112 | 8 | 9.39 × 10−8 |
| BEHAVIOR | 541 | 15 | 9.61 × 10−8 |
| POSITIVE_REGULATION_OF_ION_TRANSPORT | 659 | 16 | 2.14 × 10−7 |
| MORF_EPHA7 | 139 | 8 | 4.96 × 10−7 |
| POSITIVE_REGULATION_OF_TRANSPORT | 882 | 18 | 4.66 × 10−7 |
| POSITIVE_REGULATION_OF_MULTICELLULAR_ ORGANISMAL_PROCESS | 1397 | 23 | 4.96 × 10−7 |
| POSITIVE_REGULATION_OF_DEVELOPMENTAL_ PROCESS | 1284 | 22 | 4.62 × 10−7 |
| CENTRAL_NERVOUS_SYSTEM_DEVELOPMENT | 980 | 19 | 4.77 × 10−7 |
| MODULE_137 | 545 | 14 | 6.47 × 10−7 |
| MODULE_100 | 543 | 14 | 6.19 × 10−7 |
| MODULE_66 | 551 | 14 | 7.36 × 10−7 |
| HALLMARK_ANGIOGENESIS | 36 | 5 | 9.28 × 10−7 |
| GASC-LS DOWN | |||
|---|---|---|---|
| Gene Set Name | #Genes in Gene Set | #Genes in the Overlap | p-Value |
| CELL_CELL_SIGNALING | 1672 | 21 | 5.52 × 10−8 |
| MODULE_88 | 834 | 14 | 3.84 × 10−7 |
| MODULE_55 | 831 | 14 | 3.67 × 10−7 |
| ANTI_TREM1_VS_ANTI_TREM1_AND_LPS_MONOCYTE_DN | 195 | 8 | 2.00 × 10−7 |
| UNTREATED_VS_IL2_TREATED_STAT5_AB_KNOCKIN_TCELL_ 2H_UP | 200 | 8 | 2.43 × 10−7 |
| MODULE_64 | 517 | 11 | 7.49 × 10−7 |
| DEFENSE_RESPONSE | 1790 | 20 | 7.52 × 10−7 |
| MODULE_24 | 453 | 10 | 1.76 × 10−6 |
| REGULATION_OF_TRANSPORT | 1730 | 19 | 1.89 × 10−6 |
| LOW_LPS_VS_VEHICLE_TREATED_MONOCYTE_UP | 196 | 7 | 2.99 × 10−6 |
| ANTI_TREM1_AND_LPS_VS_VEHICLE_TREATED_MONOCYTES_UP | 195 | 7 | 2.89 × 10−6 |
| NEUTROPHIL_VS_DC_UP | 199 | 7 | 3.30 × 10−6 |
| REACTOME_DNA_DAMAGE_TELOMERE_STRESS_INDUCED_ SENESCENCE | 80 | 5 | 5.49 × 10−6 |
| MODULE_89 | 14 | 3 | 1.06 × 10−5 |
| REGULATION_OF_IMMUNE_SYSTEM_PROCESS | 1593 | 17 | 1.01 × 10−5 |
| REACTOME_SIGNALING_BY_INTERLEUKINS | 463 | 9 | 1.62 × 10−5 |
| MODULE_203 | 16 | 3 | 1.62 × 10−5 |
| SYNAPTIC_SIGNALING | 712 | 11 | 1.56 × 10−5 |
| REGULATION_OF_ION_TRANSPORT | 1314 | 15 | 1.61 × 10−5 |
| MODULE_90 | 17 | 3 | 1.96 × 10−5 |
| Gene Symbol | Gene Name | Fold Change | p-Value |
|---|---|---|---|
| VGF | VGF nerve growth factor inducible | −5.97 | 2.30 × 10−2 |
| ALPL | Alkaline Phosphatase, liver/bone/Kidney | −5.7 | 1.50 × 10−2 |
| GPR68 | G protein coupled receptor 68 | −2.61 | 1.80 × 10−2 |
| NETO1 | Neuropilin and tolloid-like 1 | −3.52 | 1.30 × 10−2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manini, I.; Dalla, E.; Vendramin, V.; Cesselli, D.; Di Loreto, C.; Skrap, M.; Ius, T. Identification of a Prognostic Microenvironment-Related Gene Signature in Glioblastoma Patients Treated with Carmustine Wafers. Cancers 2022, 14, 3413. https://doi.org/10.3390/cancers14143413
Manini I, Dalla E, Vendramin V, Cesselli D, Di Loreto C, Skrap M, Ius T. Identification of a Prognostic Microenvironment-Related Gene Signature in Glioblastoma Patients Treated with Carmustine Wafers. Cancers. 2022; 14(14):3413. https://doi.org/10.3390/cancers14143413
Chicago/Turabian StyleManini, Ivana, Emiliano Dalla, Vera Vendramin, Daniela Cesselli, Carla Di Loreto, Miran Skrap, and Tamara Ius. 2022. "Identification of a Prognostic Microenvironment-Related Gene Signature in Glioblastoma Patients Treated with Carmustine Wafers" Cancers 14, no. 14: 3413. https://doi.org/10.3390/cancers14143413
APA StyleManini, I., Dalla, E., Vendramin, V., Cesselli, D., Di Loreto, C., Skrap, M., & Ius, T. (2022). Identification of a Prognostic Microenvironment-Related Gene Signature in Glioblastoma Patients Treated with Carmustine Wafers. Cancers, 14(14), 3413. https://doi.org/10.3390/cancers14143413







