CHEK2 Pathogenic Variants in Greek Breast Cancer Patients: Evidence for Strong Associations with Estrogen Receptor Positivity, Overuse of Risk-Reducing Procedures and Population Founder Effects
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients Study Group
2.2. Genetic Analysis of CHEK2 Gene
2.3. Haplotype Analysis
2.4. Age Estimation of CHEK2, c.499G>A, c.549G>C and c.592+3A>T Variants
2.5. Statistical Analysis
3. Results
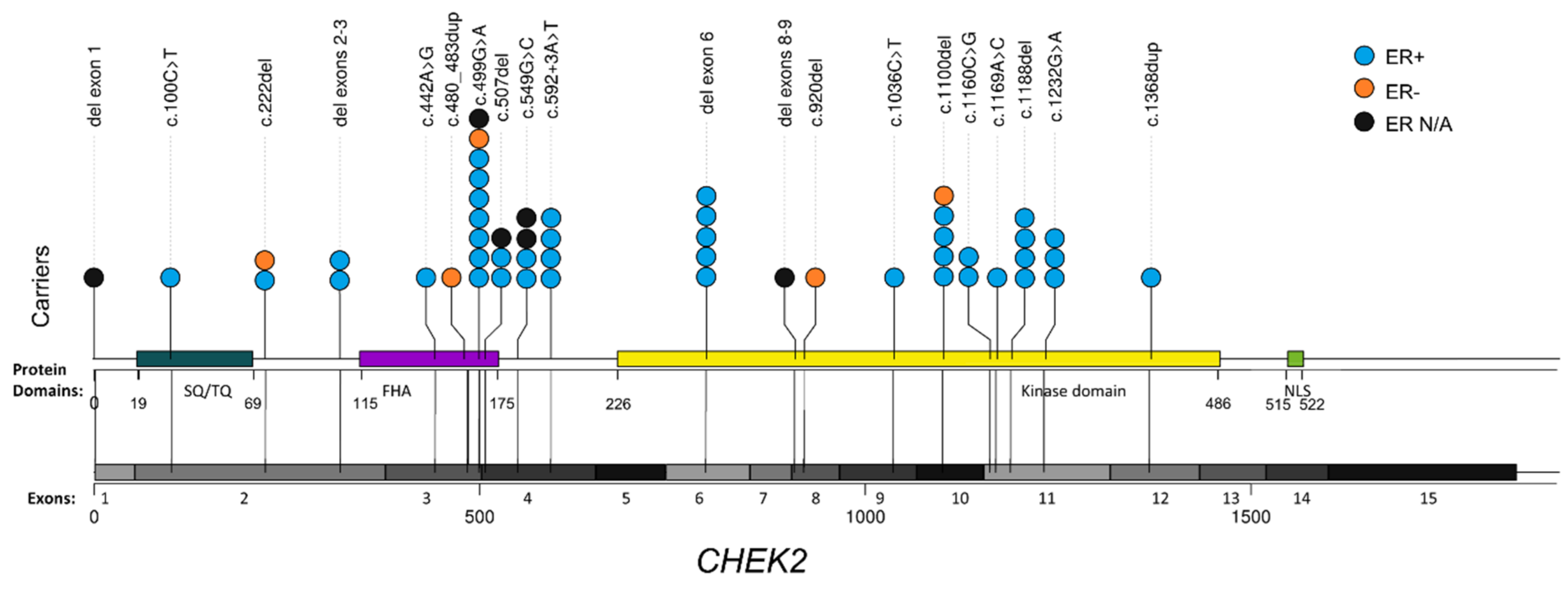
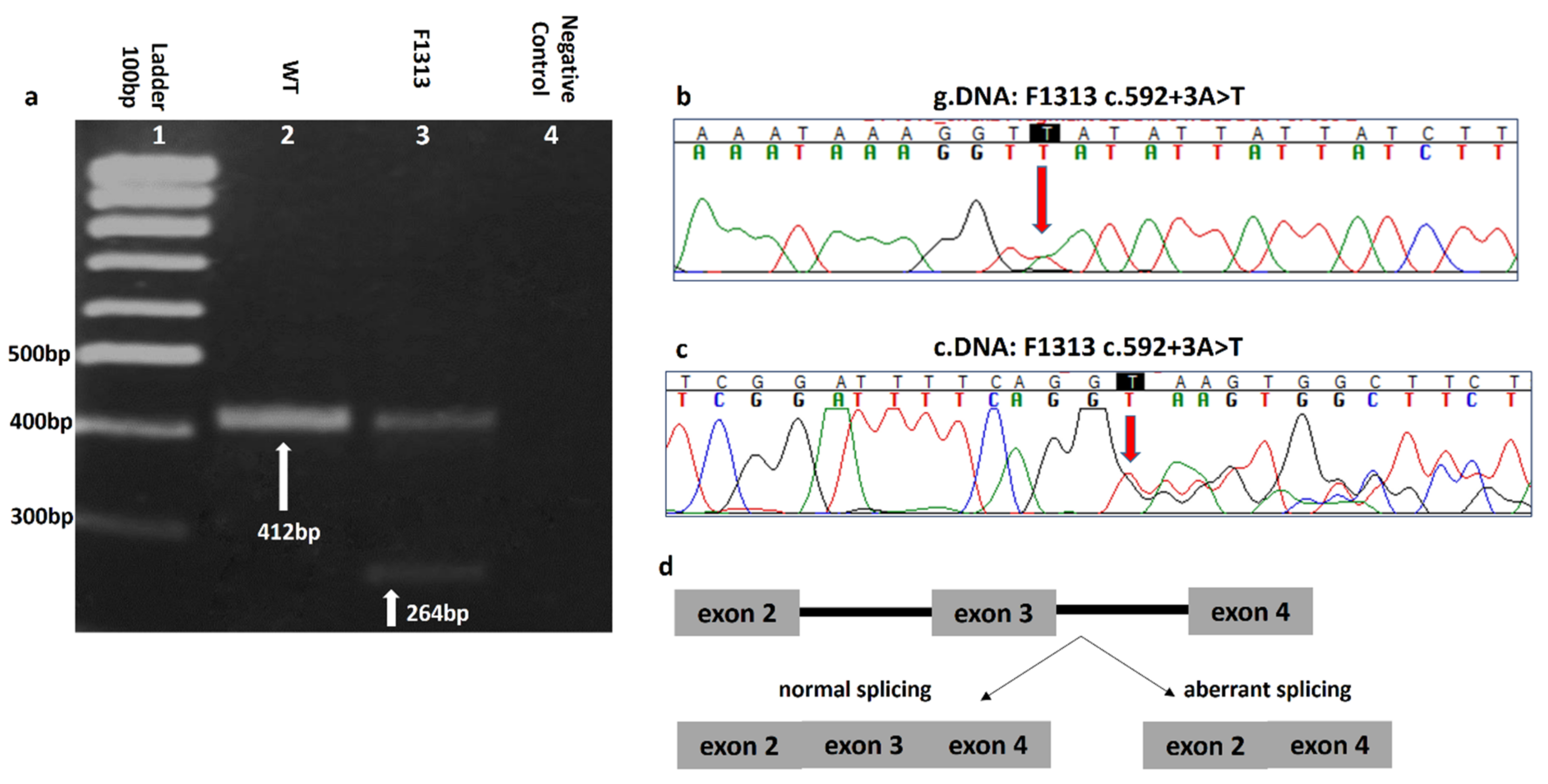
3.1. CHEK2 Pathogenic/Likely Pathogenic Variant Spectrum
3.2. Clinical and Histopathological Features of CHEK2 Variant Carriers
3.3. Surgical Management
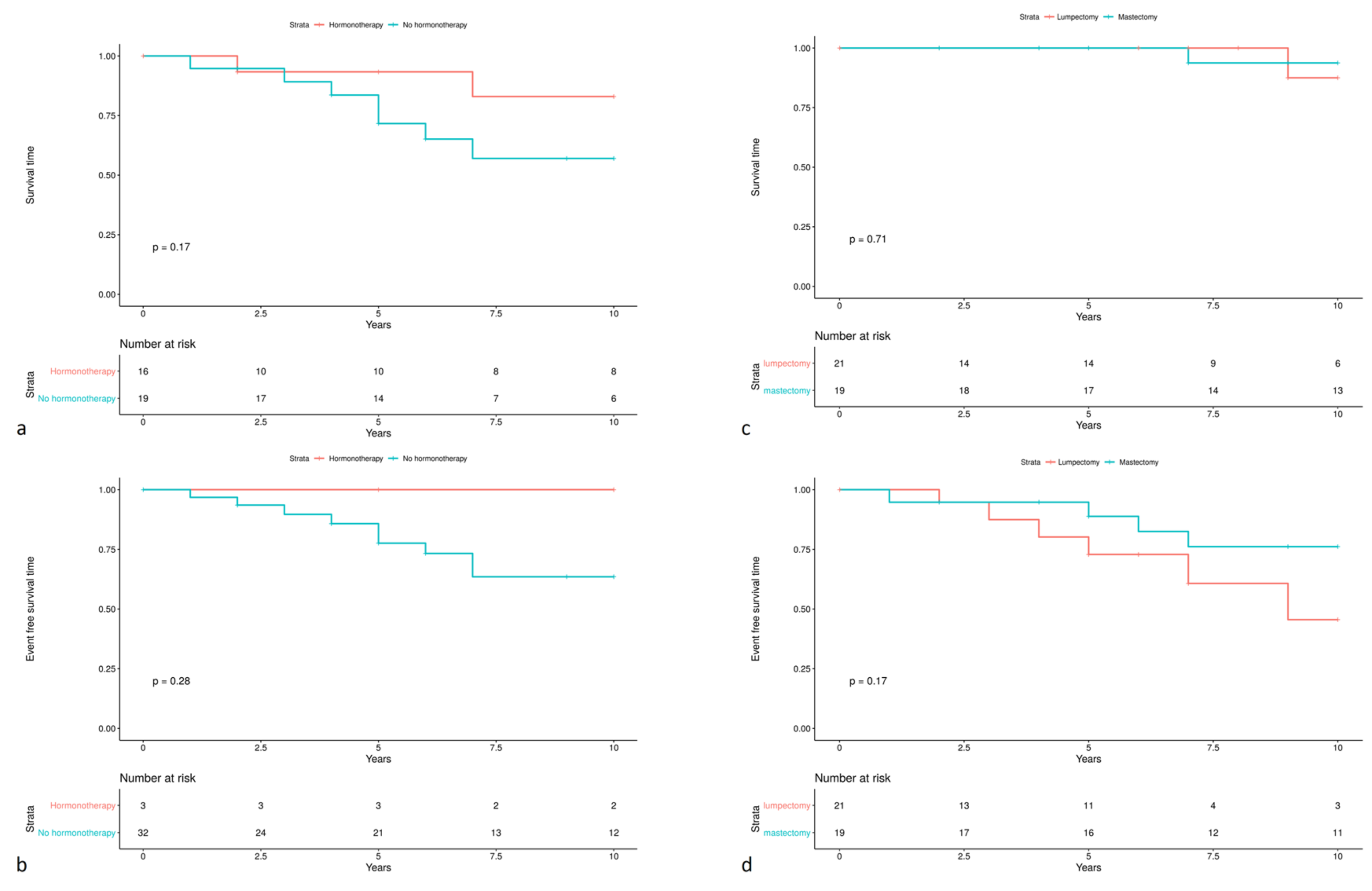
3.4. Patient Outcomes
3.5. Associations with Family History
3.6. Cascade Testing
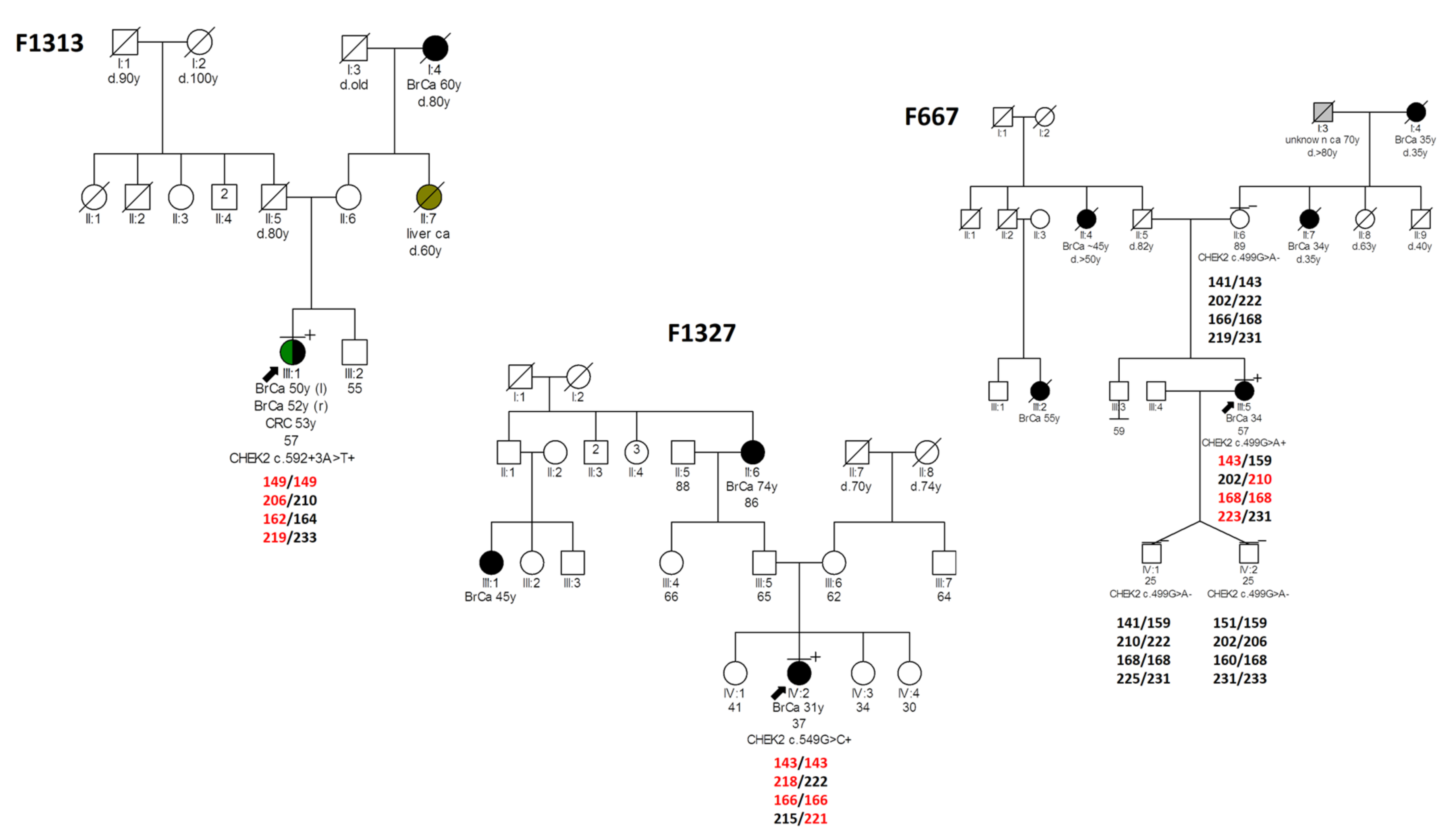
3.7. Haplotype Analysis of Variants with Possible Founder Effect and Age Estimation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Couch, F.J.; Shimelis, H.; Hu, C.; Hart, S.N.; Polley, E.C.; Na, J.; Hallberg, E.; Moore, R.; Thomas, A.; Lilyquist, J.; et al. Associations Between Cancer Predisposition Testing Panel Genes and Breast Cancer. JAMA Oncol. 2017, 3, 1190–1196. [Google Scholar] [CrossRef]
- Easton, D.F.; Pharoah, P.D.; Antoniou, A.C.; Tischkowitz, M.; Tavtigian, S.V.; Nathanson, K.L.; Devilee, P.; Meindl, A.; Couch, F.J.; Southey, M.; et al. Gene-panel sequencing and the prediction of breast-cancer risk. N. Engl. J. Med. 2015, 372, 2243–2257. [Google Scholar] [CrossRef]
- Kurian, A.W.; Li, Y.; Hamilton, A.S.; Ward, K.C.; Hawley, S.T.; Morrow, M.; McLeod, M.C.; Jagsi, R.; Katz, S.J. Gaps in Incorporating Germline Genetic Testing Into Treatment Decision-Making for Early-Stage Breast Cancer. J. Clin. Oncol. 2017, 35, 2232–2239. [Google Scholar] [CrossRef]
- Breast Cancer Association Consortium; Dorling, L.; Carvalho, S.; Allen, J.; Gonzalez-Neira, A.; Luccarini, C.; Wahlstrom, C.; Pooley, K.A.; Parsons, M.T.; Fortuno, C.; et al. Breast Cancer Risk Genes—Association Analysis in More than 113,000 Women. N. Engl. J. Med. 2021, 384, 428–439. [Google Scholar] [CrossRef]
- Hu, C.; Hart, S.N.; Gnanaolivu, R.; Huang, H.; Lee, K.Y.; Na, J.; Gao, C.; Lilyquist, J.; Yadav, S.; Boddicker, N.J.; et al. A Population-Based Study of Genes Previously Implicated in Breast Cancer. N. Engl. J. Med. 2021, 384, 440–451. [Google Scholar] [CrossRef]
- National Comprehensive Cancer Network. Genetic/Familial High-Risk Assessment: Breast, Ovarian, and Pancreatic V.1.2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/genetics_screening.pdf (accessed on 8 September 2020).
- Cybulski, C.; Gorski, B.; Huzarski, T.; Masojc, B.; Mierzejewski, M.; Debniak, T.; Teodorczyk, U.; Byrski, T.; Gronwald, J.; Matyjasik, J.; et al. CHEK2 is a multiorgan cancer susceptibility gene. Am. J. Hum. Genet. 2004, 75, 1131–1135. [Google Scholar] [CrossRef]
- Ma, X.; Zhang, B.; Zheng, W. Genetic variants associated with colorectal cancer risk: Comprehensive research synopsis, meta-analysis, and epidemiological evidence. Gut 2014, 63, 326–336. [Google Scholar] [CrossRef]
- Siolek, M.; Cybulski, C.; Gasior-Perczak, D.; Kowalik, A.; Kozak-Klonowska, B.; Kowalska, A.; Chlopek, M.; Kluzniak, W.; Wokolorczyk, D.; Palyga, I.; et al. CHEK2 mutations and the risk of papillary thyroid cancer. Int. J. Cancer 2015, 137, 548–552. [Google Scholar] [CrossRef]
- Tung, N.; Domchek, S.M.; Stadler, Z.; Nathanson, K.L.; Couch, F.; Garber, J.E.; Offit, K.; Robson, M.E. Counselling framework for moderate-penetrance cancer-susceptibility mutations. Nat. Rev. Clin. Oncol. 2016, 13, 581–588. [Google Scholar] [CrossRef]
- Apostolou, P.; Fostira, F.; Mollaki, V.; Delimitsou, A.; Vlassi, M.; Pentheroudakis, G.; Faliakou, E.; Kollia, P.; Fountzilas, G.; Yannoukakos, D.; et al. Characterization and prevalence of two novel CHEK2 large deletions in Greek breast cancer patients. J. Hum. Genet. 2018, 63, 877–886. [Google Scholar] [CrossRef]
- Walsh, T.; Casadei, S.; Coats, K.H.; Swisher, E.; Stray, S.M.; Higgins, J.; Roach, K.C.; Mandell, J.; Lee, M.K.; Ciernikova, S.; et al. Spectrum of mutations in BRCA1, BRCA2, CHEK2, and TP53 in families at high risk of breast cancer. JAMA 2006, 295, 1379–1388. [Google Scholar] [CrossRef]
- Cybulski, C.; Gorski, B.; Huzarski, T.; Byrski, T.; Gronwald, J.; Debniak, T.; Wokolorczyk, D.; Jakubowska, A.; Kowalska, E.; Oszurek, O.; et al. CHEK2-positive breast cancers in young Polish women. Clin. Cancer Res. 2006, 12, 4832–4835. [Google Scholar] [CrossRef]
- Apostolou, P.; Fostira, F.; Papamentzelopoulou, M.; Michelli, M.; Panopoulos, C.; Fountzilas, G.; Konstantopoulou, I.; Voutsinas, G.E.; Yannoukakos, D. CHEK2 c.1100delC allele is rarely identified in Greek breast cancer cases. Cancer Genet. 2015, 208, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Delimitsou, A.; Fostira, F.; Kalfakakou, D.; Apostolou, P.; Konstantopoulou, I.; Kroupis, C.; Papavassiliou, A.G.; Kleibl, Z.; Stratikos, E.; Voutsinas, G.E.; et al. Functional characterization of CHEK2 variants in a Saccharomyces cerevisiae system. Hum. Mutat. 2019, 40, 631–648. [Google Scholar] [CrossRef]
- Fostira, F.; Kostantopoulou, I.; Apostolou, P.; Papamentzelopoulou, M.S.; Papadimitriou, C.; Faliakou, E.; Christodoulou, C.; Boukovinas, I.; Razis, E.; Tryfonopoulos, D.; et al. One in three highly selected Greek patients with breast cancer carries a loss-of-function variant in a cancer susceptibility gene. J. Med. Genet. 2020, 57, 53–61. [Google Scholar] [CrossRef]
- Rannala, B.; Bertorelle, G. Using linked markers to infer the age of a mutation. Hum. Mutat. 2001, 18, 87–100. [Google Scholar] [CrossRef] [PubMed]
- Kraus, C.; Hoyer, J.; Vasileiou, G.; Wunderle, M.; Lux, M.P.; Fasching, P.A.; Krumbiegel, M.; Uebe, S.; Reuter, M.; Beckmann, M.W.; et al. Gene panel sequencing in familial breast/ovarian cancer patients identifies multiple novel mutations also in genes others than BRCA1/2. Int. J. Cancer 2017, 140, 95–102. [Google Scholar] [CrossRef]
- Reese, M.G.; Eeckman, F.H.; Kulp, D.; Haussler, D. Improved splice site detection in Genie. J. Comput. Biol. 1997, 4, 311–323. [Google Scholar] [CrossRef] [PubMed]
- Cybulski, C.; Wokolorczyk, D.; Jakubowska, A.; Huzarski, T.; Byrski, T.; Gronwald, J.; Masojc, B.; Deebniak, T.; Gorski, B.; Blecharz, P.; et al. Risk of breast cancer in women with a CHEK2 mutation with and without a family history of breast cancer. J. Clin. Oncol. 2011, 29, 3747–3752. [Google Scholar] [CrossRef]
- Kilpivaara, O.; Bartkova, J.; Eerola, H.; Syrjakoski, K.; Vahteristo, P.; Lukas, J.; Blomqvist, C.; Holli, K.; Heikkila, P.; Sauter, G.; et al. Correlation of CHEK2 protein expression and c.1100delC mutation status with tumor characteristics among unselected breast cancer patients. Int. J. Cancer 2005, 113, 575–580. [Google Scholar] [CrossRef]
- Schmidt, M.K.; Hogervorst, F.; van Hien, R.; Cornelissen, S.; Broeks, A.; Adank, M.A.; Meijers, H.; Waisfisz, Q.; Hollestelle, A.; Schutte, M.; et al. Age- and Tumor Subtype-Specific Breast Cancer Risk Estimates for CHEK2*1100delC Carriers. J. Clin. Oncol. 2016, 34, 2750–2760. [Google Scholar] [CrossRef] [PubMed]
- Narod, S.A. Testing for CHEK2 in the cancer genetics clinic: Ready for prime time? Clin. Genet. 2010, 78, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Cybulski, C.; Huzarski, T.; Byrski, T.; Gronwald, J.; Debniak, T.; Jakubowska, A.; Gorski, B.; Wokolorczyk, D.; Masojc, B.; Narod, S.A.; et al. Estrogen receptor status in CHEK2-positive breast cancers: Implications for chemoprevention. Clin. Genet. 2009, 75, 72–78. [Google Scholar] [CrossRef] [PubMed]
- De Bock, G.H.; Mourits, M.J.; Schutte, M.; Krol-Warmerdam, E.M.; Seynaeve, C.; Blom, J.; Brekelmans, C.T.; Meijers-Heijboer, H.; van Asperen, C.J.; Cornelisse, C.J.; et al. Association between the CHEK2*1100delC germ line mutation and estrogen receptor status. Int. J. Gynecol. Cancer 2006, 16 (Suppl. 2), 552–555. [Google Scholar] [CrossRef] [PubMed]
- Rainville, I.; Hatcher, S.; Rosenthal, E.; Larson, K.; Bernhisel, R.; Meek, S.; Gorringe, H.; Mundt, E.; Manley, S. High risk of breast cancer in women with biallelic pathogenic variants in CHEK2. Breast Cancer Res. Treat. 2020, 180, 503–509. [Google Scholar] [CrossRef]
- Greville-Heygate, S.L.; Maishman, T.; Tapper, W.J.; Cutress, R.I.; Copson, E.; Dunning, A.M.; Haywood, L.; Jones, L.J.; Eccles, D.M. Pathogenic Variants in CHEK2 Are Associated With an Adverse Prognosis in Symptomatic Early-Onset Breast Cancer. JCO Precis. Oncol. 2020, 4. [Google Scholar] [CrossRef]
- De Bock, G.H.; Schutte, M.; Krol-Warmerdam, E.M.; Seynaeve, C.; Blom, J.; Brekelmans, C.T.; Meijers-Heijboer, H.; van Asperen, C.J.; Cornelisse, C.J.; Devilee, P.; et al. Tumour characteristics and prognosis of breast cancer patients carrying the germline CHEK2*1100delC variant. J. Med. Genet. 2004, 41, 731–735. [Google Scholar] [CrossRef]
- Meijers-Heijboer, H.; van den Ouweland, A.; Klijn, J.; Wasielewski, M.; de Snoo, A.; Oldenburg, R.; Hollestelle, A.; Houben, M.; Crepin, E.; van Veghel-Plandsoen, M.; et al. Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat. Genet. 2002, 31, 55–59. [Google Scholar] [CrossRef]
- Vahteristo, P.; Bartkova, J.; Eerola, H.; Syrjakoski, K.; Ojala, S.; Kilpivaara, O.; Tamminen, A.; Kononen, J.; Aittomaki, K.; Heikkila, P.; et al. A CHEK2 genetic variant contributing to a substantial fraction of familial breast cancer. Am. J. Hum. Genet. 2002, 71, 432–438. [Google Scholar] [CrossRef]
- Kurian, A.W.; Ward, K.C.; Abrahamse, P.; Hamilton, A.S.; Deapen, D.; Morrow, M.; Jagsi, R.; Katz, S.J. Association of Germline Genetic Testing Results With Locoregional and Systemic Therapy in Patients With Breast Cancer. JAMA Oncol. 2020, 6, e196400. [Google Scholar] [CrossRef]
- Cragun, D.; Weidner, A.; Tezak, A.; Clouse, K.; Pal, T. Cancer risk management among female BRCA1/2, PALB2, CHEK2, and ATM carriers. Breast Cancer Res. Treat. 2020, 182, 421–428. [Google Scholar] [CrossRef] [PubMed]
- Fillon, M. Low-cost online cascade test may persuade relatives to investigate their own cancer risk. CA Cancer J. Clin. 2019, 69, 86–87. [Google Scholar] [CrossRef] [PubMed]
- Courtney, E.; Chok, A.K.; Ting Ang, Z.L.; Shaw, T.; Li, S.T.; Yuen, J.; Ngeow, J. Impact of free cancer predisposition cascade genetic testing on uptake in Singapore. NPJ Genom. Med. 2019, 4, 22. [Google Scholar] [CrossRef]
- Caswell-Jin, J.L.; Zimmer, A.D.; Stedden, W.; Kingham, K.E.; Zhou, A.Y.; Kurian, A.W. Cascade Genetic Testing of Relatives for Hereditary Cancer Risk: Results of an Online Initiative. J. Natl. Cancer Inst. 2019, 111, 95–98. [Google Scholar] [CrossRef] [PubMed]
| Patient ID | F/M | Variant (DNA level, NM _007194.4) | BrCa 1st dx (y) | Histology | Grade | ER | PR | HER2 | Hormonotherapy (Years) | Surgery | Other ca dx (y) | Prophylactic Mastectomy | RRSO | Distant Met (y) | FH |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F876 | F | g.(29130716_29137756)_(29137822_?)del * [del exon 1] | 47 ** | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | Lumpectomy | relapse 56 | N.A. | N.A | N.A | Yes |
| F993 | F | c.100C>T | 47 | Ductal | 2 | + | + | - | Yes (14 -in process) | Mastectomy | No | No | No | No | Yes |
| F1912 | F | c.222del | 61 | Ductal | 2 | - | - | - | No | Lumpectomy | No | No | No | No | N.A. |
| F2217 | F | c.222del | 42 | DCIS | 3 | + | + | - | Yes (5) | Lumpectomy | No | No | No | No | Yes |
| F1081 | F | c.320-4172_593-3895del [del exons 2-3] | 36 | Ductal | 3 | + | - | + | Yes (10) | Mastectomy | No | CPM | RRSO | No | No |
| F3519 | F | c.320-4172_593-3895del [del exons 2-3] | 41 | Mixed (ductal/lobular) | 2 | + | + | N.A. | Yes (2 -in process) | Lumpectomy | No | No | No | No | No |
| F5055 | F | c.442A>G | 39 | Ductal | 3 | + | + | - | Yes (3 -in process) | Mastectomy | No | No | No | No | No |
| F508 | F | c.480_483dup | 39 | Mixed (ductal/lobular) | 2 | - | - | N.A. | No | Μastectomy | BrCa 58 | CPM | RRSO | No | Yes |
| F667 | F | c.499G>A | 34 | Ductal | 1 | + | + | N.A. | No | Lumpectomy | No | No | No | No | Yes |
| F1326 | F | c.499G>A | 38 | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | Lumpectomy | N.A. | N.A. | N.A | N.A | Yes |
| F2050 | F | c.499G>A | 46 | Ductal | 1 | + | + | - | Yes (10) | Mastectomy | No | No | RRSO | No | Yes |
| F2276 | M | c.499G>A | 41 | DCIS | N.A. | + | + | - | N.A. | Lumpectomy | relapse 45 | N.A. | N.A. | N.A. | |
| F2469 | F | c.499G>A | 35 | Ductal | 3 | + | + | - | No | Lumpectomy | No | PBM | No | No | No |
| F2605 | F | c.499G>A | 39 | Ductal | 3 | - | - | - | No | Mastectomy | No | CPM | No | No | No |
| F2774 | F | c.499G>A | 38 | Ductal | 2 | + | - | - | Yes (5) | Mastectomy | BrCa 44 | No | RRSO | No | No |
| F3445 | F | c.499G>A | 50 | DCIS | 3 | + | + | + | Yes (5) | Mastectomy | No | CPM | RRSO | No | Yes |
| F3754 | F | c.499G>A | 40 | Ductal | 2 | + | - | - | Yes (1 -in process) | Lumpectomy | No | No | No | No | Yes |
| F322 | F | c.507del | 51 | Ν.A. | Ν.A. | N.A. | N.A. | N.A. | Ν.A. | Lumpectomy | BrCa 67 | Ν.A. | Ν.A. | Ν.A. | Yes |
| F2305 | F | c.507del | 41 | DCIS | 3 | + | + | - | Yes (4) | Lumpectomy | relapse 45 | PBM | No | No | Yes |
| F3127 | F | c.507del | 44 | Ductal | 3 | + | - | - | Yes (5) | Lumpectomy | BrCa 49 | No | No | No | No |
| F459 | F | c.549G>C | 46 | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | Lumpectomy | N.A. | N.A. | N.A. | N.A. | Yes |
| F498 | F | c.549G>C | 41 | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | Bil-Mastectomy | No | PBM | RRSO | Yes (69) | No |
| F1018 | F | c.549G>C | 36 | Ductal | N.A. | + | + | - | Yes (8) | Lumpectomy | relapse 39 | No | No | No | Yes |
| F1327 | F | c.549G>C | 31 | DCIS | 2 | + | + | - | Yes (10) | Mastectomy | No | No | No | No | Yes |
| F1313 | F | c.592+3A>T | 50 | Lobular | 1 | + | + | - | Yes (10) | Lumpectomy | BrCa 52 CRC 53 | No | No | No | Yes |
| F2482 | F | c.592+3A>T | 33 | Ductal | 3 | + | + | - | Yes (10) | Mastectomy | No | No | No | No | Yes |
| F3247 | F | c.592+3A>T | 36 | DCIS | 2 | + | + | - | Yes (5) | Lumpectomy | No | No | No | No | No |
| F3710 | F | c.592+3A>T | 50 | Ductal | 3 | + | + | N.A. | Yes (6 months -in process) | Lumpectomy | No | No | No | No | Yes |
| F1070 | F | c.793-1557_847-485del [del exon 6] | 57 | Lobular | 3 | + | + | + | Yes (5,5) | Mastectomy | No | No | RRSO | No | No |
| F1136 | F | c.793-1557_847-485del [del exon 6] | 42 | Ductal | 2 | + | + | + | N.A. | Lumpectomy | No | N.A. | RRSO | Yes (48) | Yes |
| F1933 | F | c.793-1557_847-485del [del exon 6] | 55 | Ductal | 2 | + | - | + | No | Mastectomy | No | No | No | No | Yes |
| F1937 | F | c.793-1557_847-485del [del exon 6] | 38 | Ductal | 2 | + | + | + | Yes (3) | Mastectomy | BrCa 50 | No | No | No | Yes |
| F1938 | F | c.793-1557_847-485del [del exon 6] | 48 | Lobular | 2 | + | + | - | Yes (5) | Mastectomy | No | No | No | No | Yes |
| F2034 | F | c.909-2028_1096-698del [del exons 8-9] | 42 | N.A. | N.A. | N.A. | N.A. | N.A. | Yes (N.A.) | Mastectomy | No | No | RRSO | No | Yes |
| F5540s | F | c.920del | 44 | Ductal | 3 | - | - | + | No | No | No | No | No | Yes (44) | No |
| F1741 | F | c.1036C>T | 40 | Lobular | 3 | + | + | - | Yes (5) | Mastectomy | No | No | No | Yes (45) | Yes |
| F1556 | F | c.1100del | 41 | Mixed (ductal/lobular) | 2 | + | + | - | Yes (8) | Mastectomy | BrCa 42 | No | No | No | Yes |
| F1629 | F | c.1100del | 45 | Mixed (ductal/lobular) | 3 | + | - | - | Yes (29 -in process) | Mastectomy | relapse 50 & 63 | No | RRSO | No | No |
| F1630 | F | c.1100del | 49 | Ductal | 2 | + | + | + | N.A. | Lumpectomy | N.A. | N.A. | N.A. | N.A. | No |
| F1631 | F | c.1100del | 47 | Ductal | 3 | - | - | + | No | Lumpectomy | N.A. | N.A. | N.A. | N.A. | No |
| F3412 | F | c.1100del | 37 | Ductal | 3 | + | + | - | Yes (5) | Mastectomy | BrCa 50 | No | No | No | No |
| F619 | F | c.1160C>G | 43 | Ductal | N.A. | + | - | - | N.A. | Lumpectomy | N.A. | N.A. | N.A. | Yes (43) | Yes |
| F1960 | F | c.1160C>G | 53 | Ductal | N.A. | + | + | N.A. | Yes (6 months) | Mastectomy | No | No | No | No | Yes |
| F3477 | F | c.1169A>C | 33 | Ductal | 2 | + | - | - | Yes (10) | Μastectomy | Νο | CPM | No | No | Yes |
| F2839 | F | c.1188del | 30 | Ductal | 3 | + | + | - | N.A. | Lumpectomy | N.A. | N.A. | N.A. | N.A. | N.A. |
| F2891 | F | c.1188del | 47 | Ductal | 2 | + | N.A. | N.A. | Yes (7) | Lumpectomy | BrCa 63 | No | RRSO | No | No |
| F2983 | F | c.1188del | 47 | Ductal | 2 | + | + | + | Yes (5) | Mastectomy | No | CPM | RRSO | Yes (51, 58) | Yes |
| F3696 | F | c.1188del | 35 | Ductal | 2 | + | + | - | Yes (10) | Mastectomy | No | No | No | Yes (49) | No |
| F2261 | F | c.1232G>A | 53 | Ductal | 3 | + | + | - | N.A. | Lumpectomy | No | CPM | RRSO | No | Yes |
| F2790 | F | c.1232G>A | 38 | Ductal | 2 | + | N.A. | N.A. | Yes (14) | Lumpectomy | BrCa 45 | No | RRSO | Yes (52) | No |
| F3158 | F | c.1232G>A | 42 | Ductal | 3 | + | + | - | Yes (10) | Lumpectomy | No | No | No | No | Yes |
| F802 | F | c.1368dup | 48 | Ductal | 2 | + | + | - | Yes (10) | Lumpectomy | No | No | No | No | Yes |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Apostolou, P.; Dellatola, V.; Papadimitriou, C.; Kalfakakou, D.; Fountzilas, E.; Faliakou, E.; Fountzilas, G.; Romanidou, O.; Konstantopoulou, I.; Fostira, F. CHEK2 Pathogenic Variants in Greek Breast Cancer Patients: Evidence for Strong Associations with Estrogen Receptor Positivity, Overuse of Risk-Reducing Procedures and Population Founder Effects. Cancers 2021, 13, 2106. https://doi.org/10.3390/cancers13092106
Apostolou P, Dellatola V, Papadimitriou C, Kalfakakou D, Fountzilas E, Faliakou E, Fountzilas G, Romanidou O, Konstantopoulou I, Fostira F. CHEK2 Pathogenic Variants in Greek Breast Cancer Patients: Evidence for Strong Associations with Estrogen Receptor Positivity, Overuse of Risk-Reducing Procedures and Population Founder Effects. Cancers. 2021; 13(9):2106. https://doi.org/10.3390/cancers13092106
Chicago/Turabian StyleApostolou, Paraskevi, Vasiliki Dellatola, Christos Papadimitriou, Despoina Kalfakakou, Elena Fountzilas, Eleni Faliakou, Georgios Fountzilas, Ourania Romanidou, Irene Konstantopoulou, and Florentia Fostira. 2021. "CHEK2 Pathogenic Variants in Greek Breast Cancer Patients: Evidence for Strong Associations with Estrogen Receptor Positivity, Overuse of Risk-Reducing Procedures and Population Founder Effects" Cancers 13, no. 9: 2106. https://doi.org/10.3390/cancers13092106
APA StyleApostolou, P., Dellatola, V., Papadimitriou, C., Kalfakakou, D., Fountzilas, E., Faliakou, E., Fountzilas, G., Romanidou, O., Konstantopoulou, I., & Fostira, F. (2021). CHEK2 Pathogenic Variants in Greek Breast Cancer Patients: Evidence for Strong Associations with Estrogen Receptor Positivity, Overuse of Risk-Reducing Procedures and Population Founder Effects. Cancers, 13(9), 2106. https://doi.org/10.3390/cancers13092106







