Activated Regulatory T-Cells, Dysfunctional and Senescent T-Cells Hinder the Immunity in Pancreatic Cancer
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
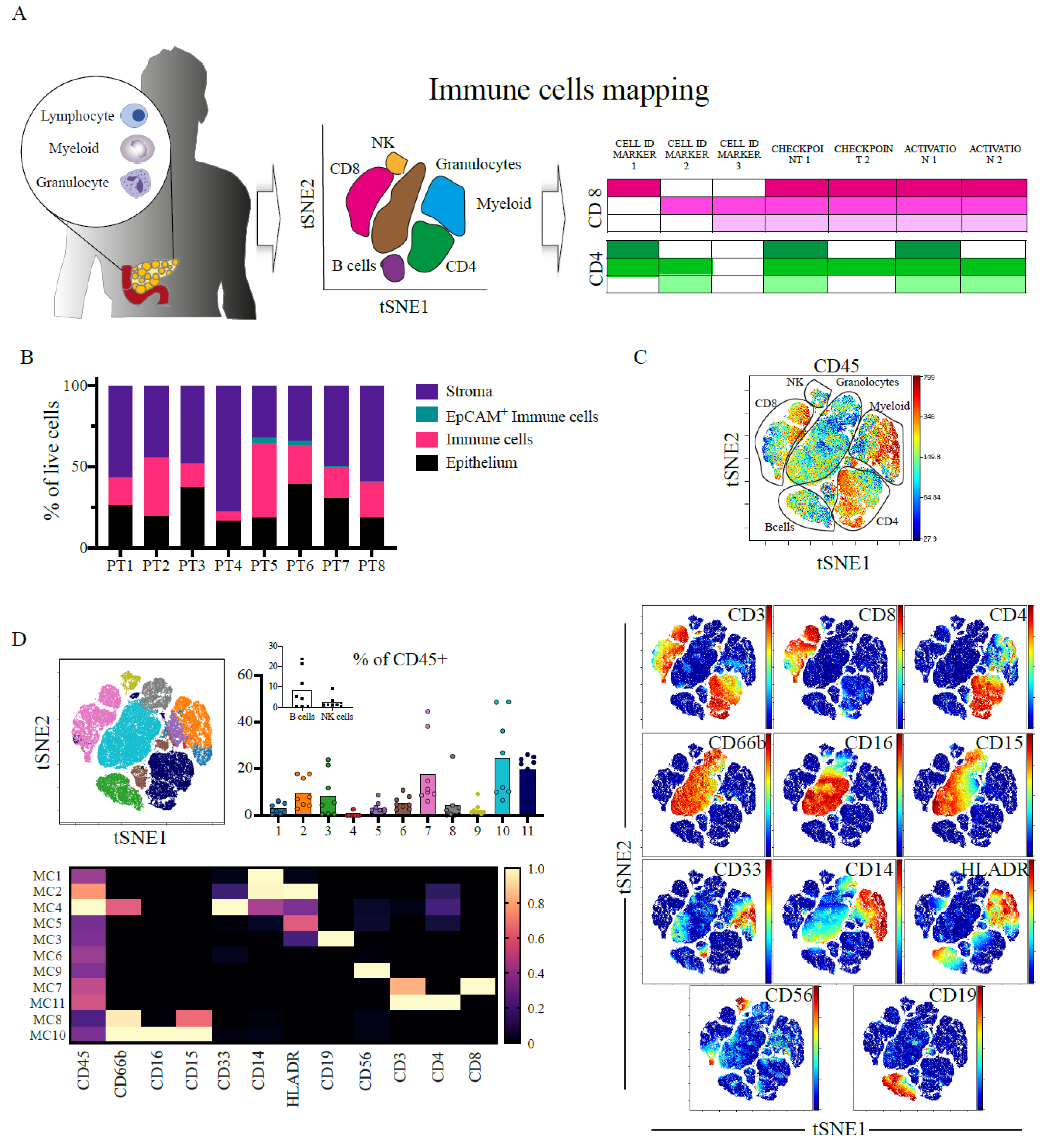
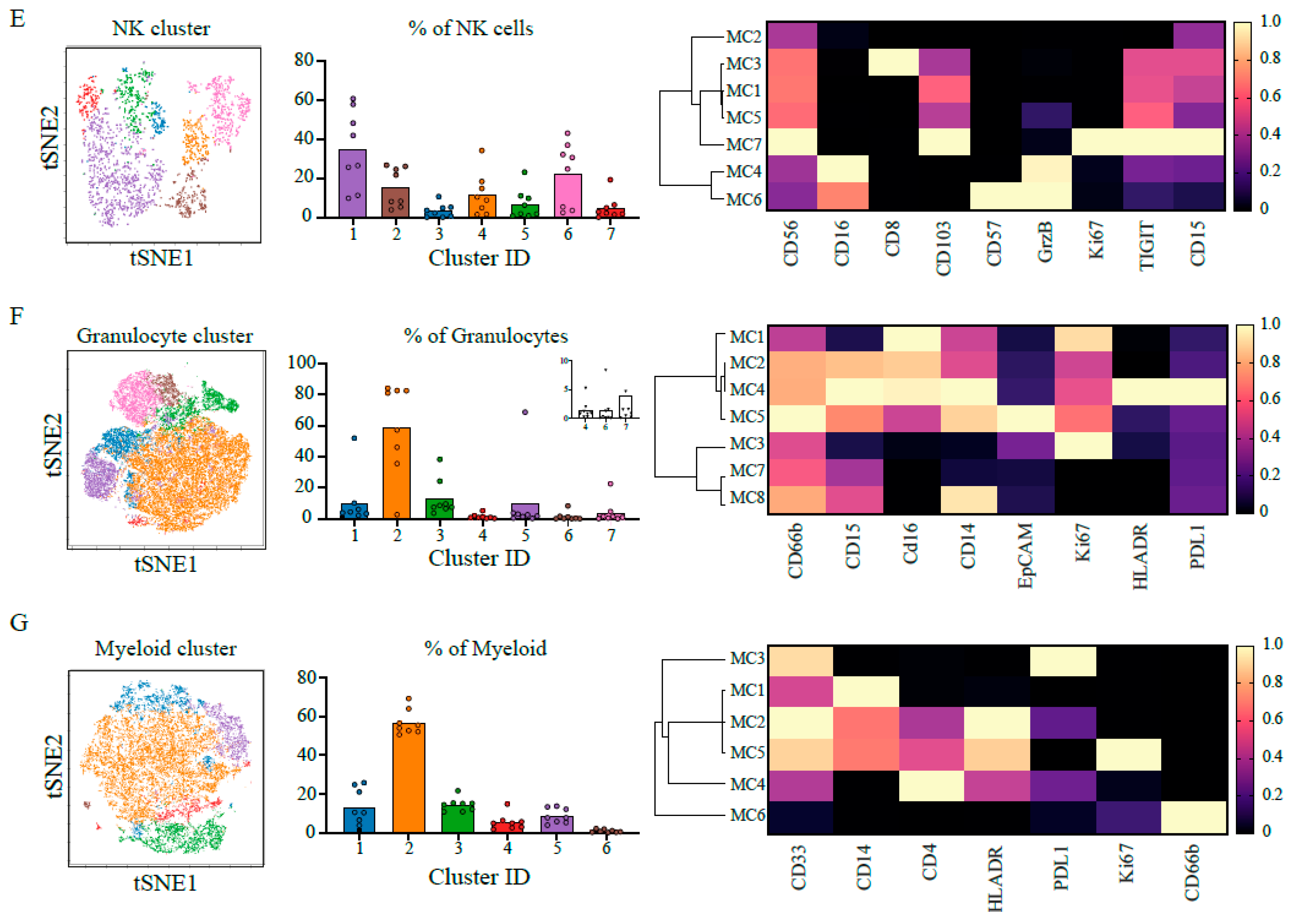
2.1. Heterogenous and Suppressive Innate Immune Cell Composition within Pancreatic Cancer
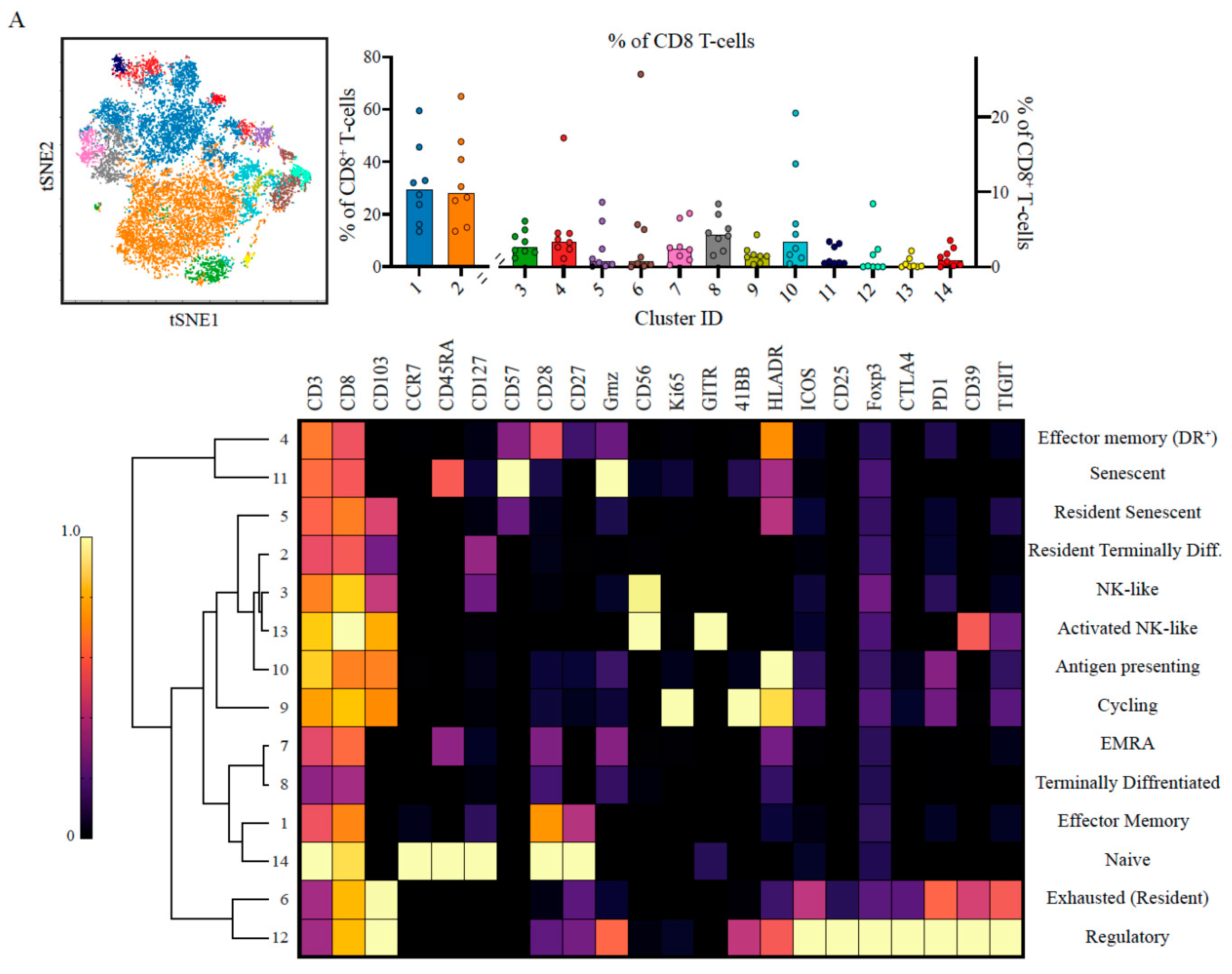
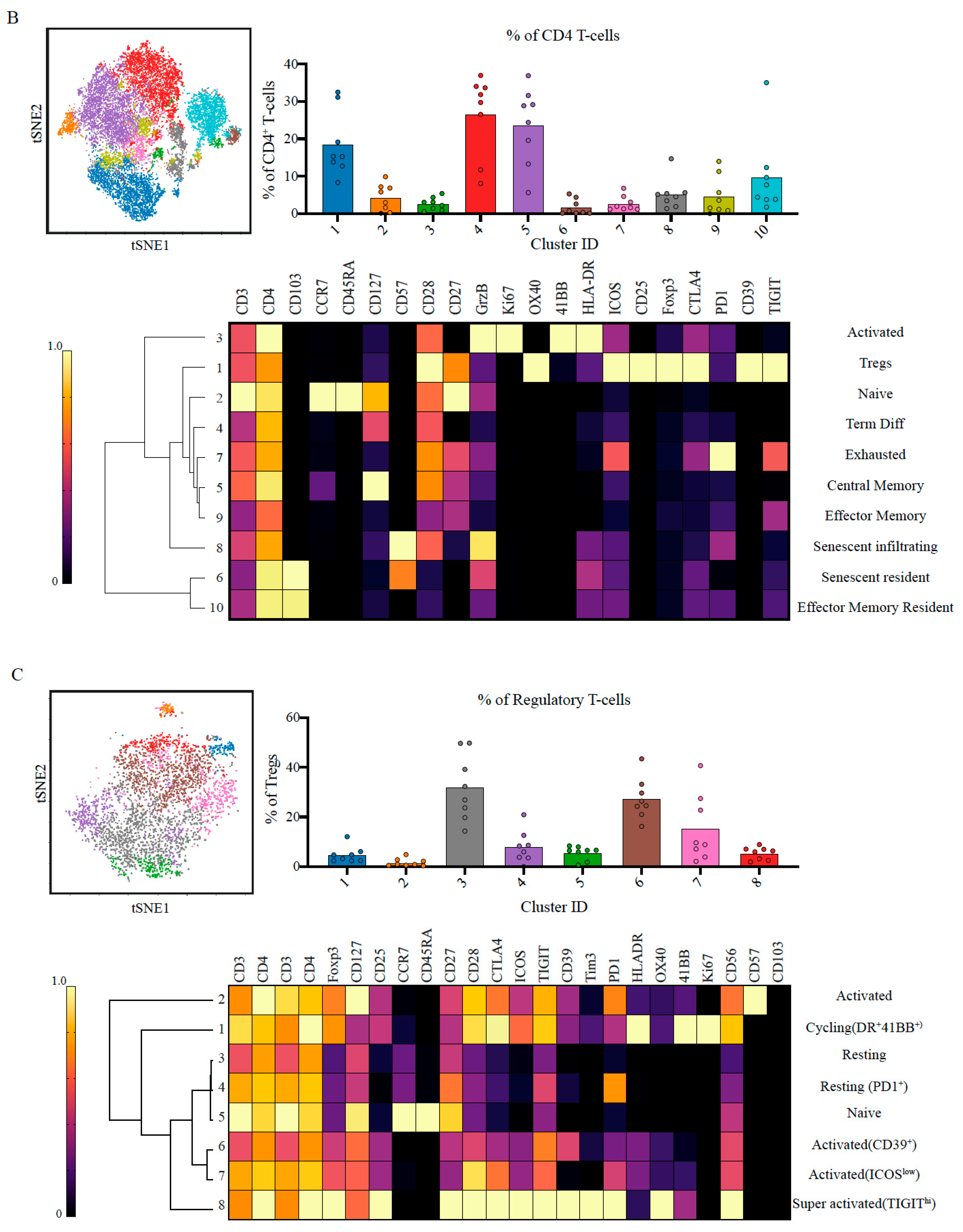
2.2. Suppressive and Non-Tumour Responsive T-Cells Predominate PDAC Microenvironment
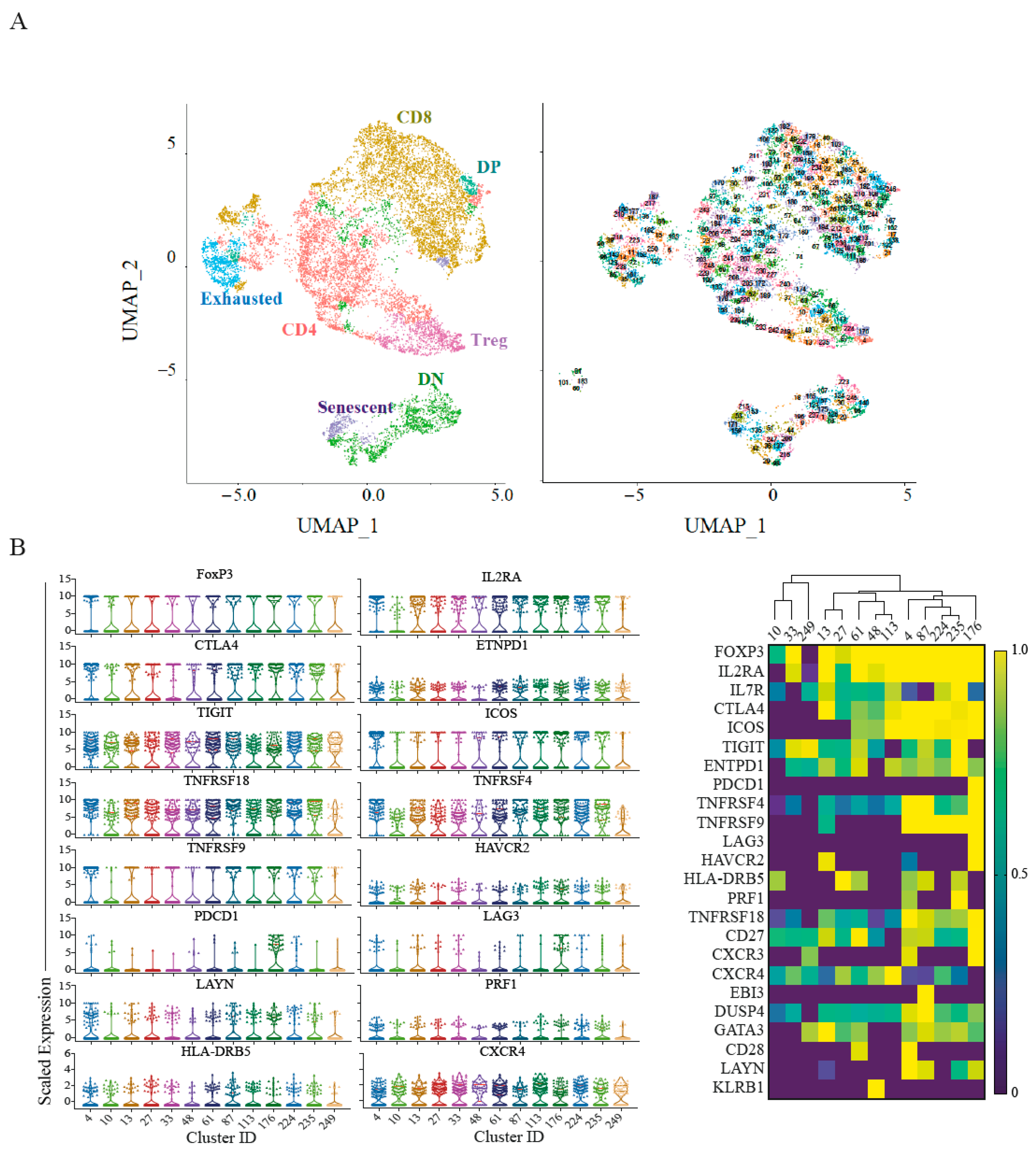
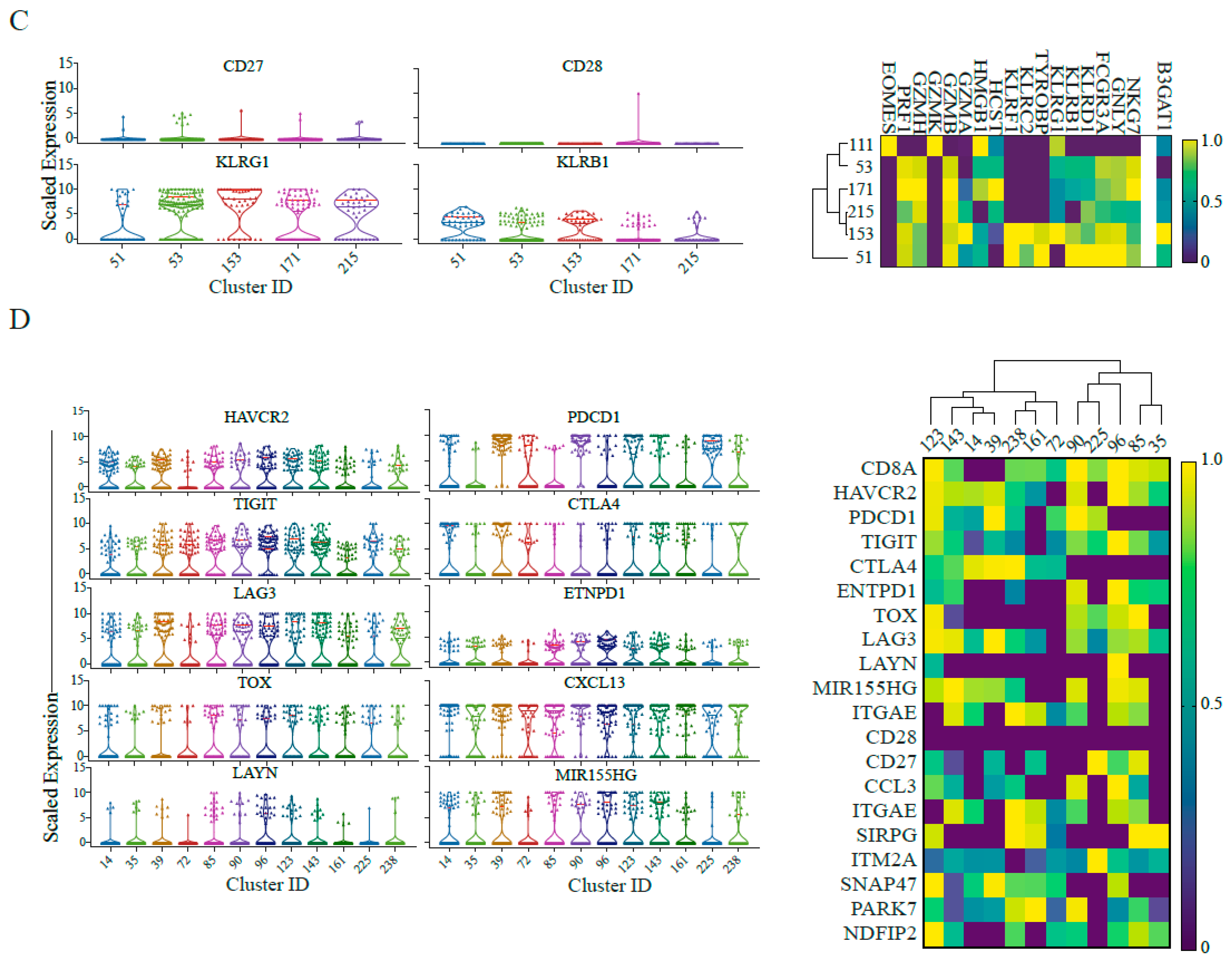
2.3. Single-Cell RNA Sequencing Validates Senescence and Regulatory Signatures in Tumour Infiltrating T-Cells
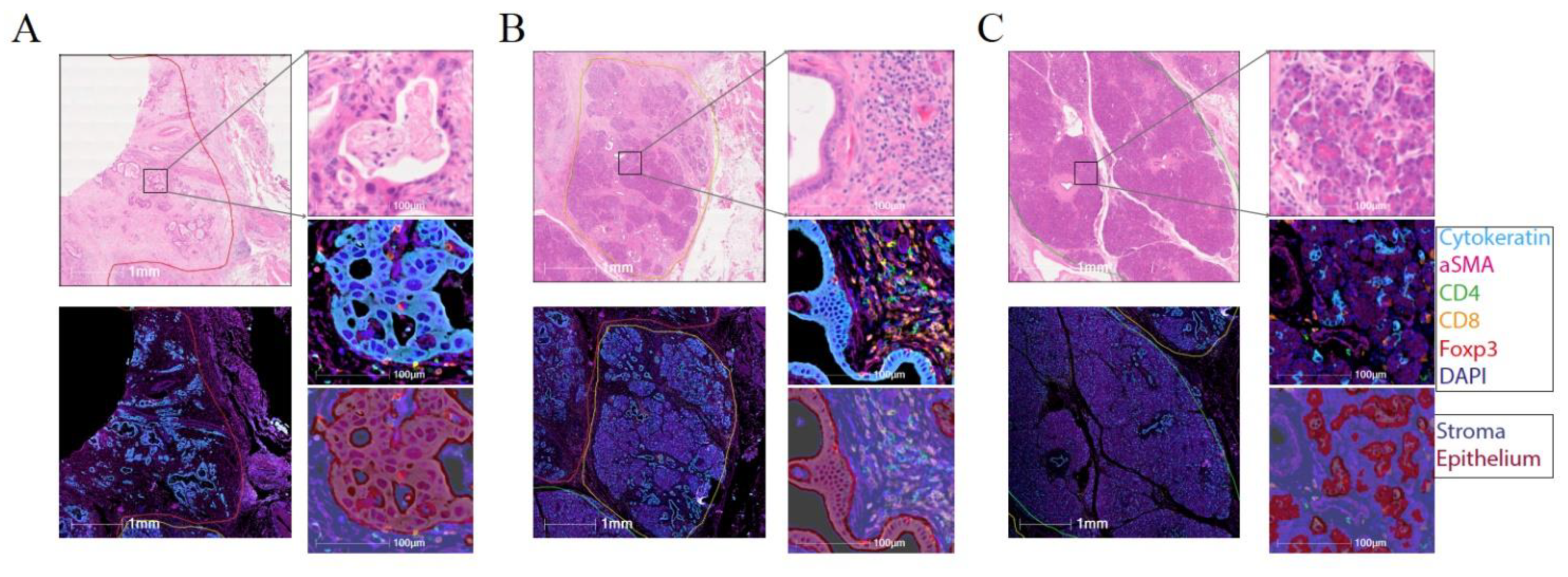
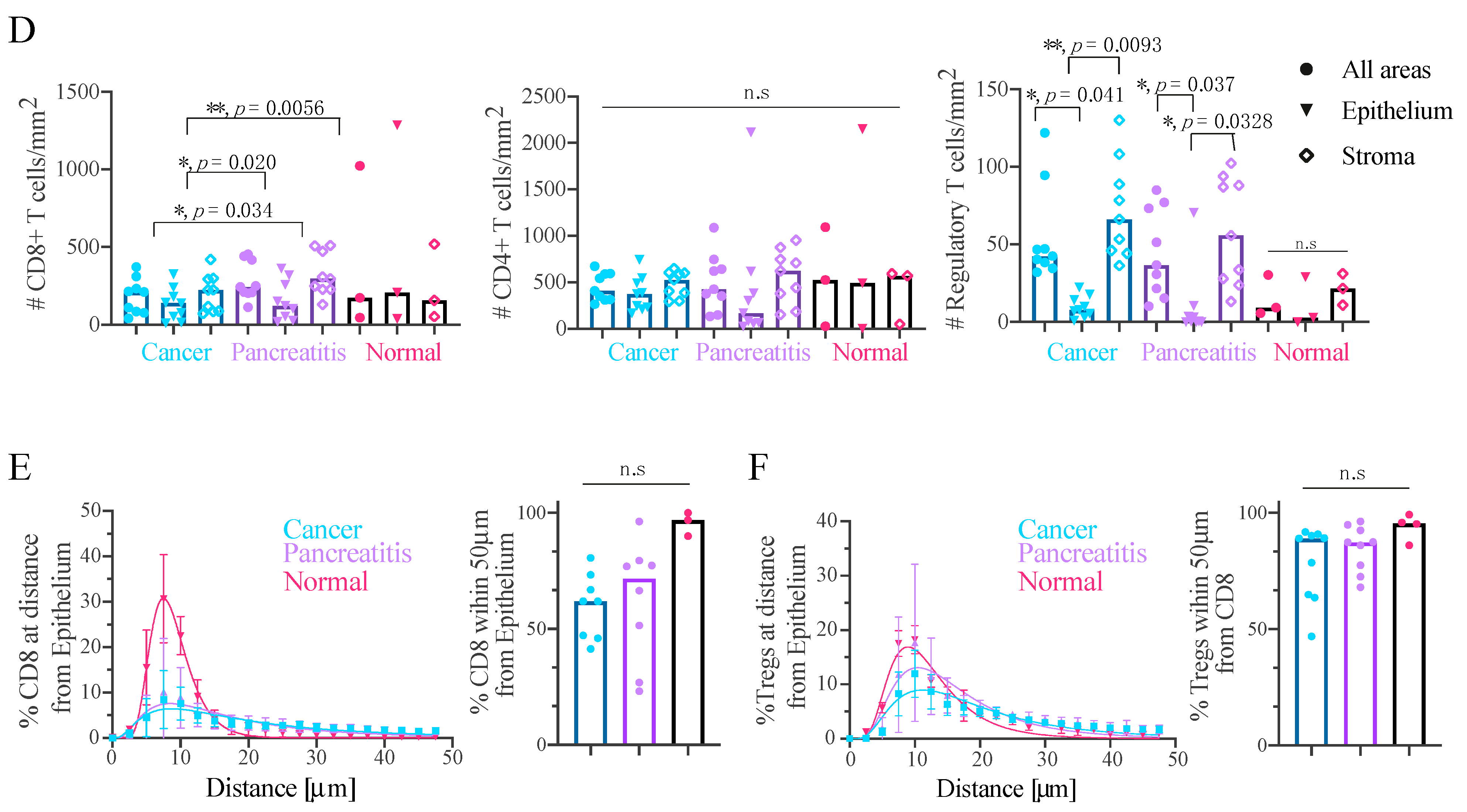
2.4. Effector T-Cells Are Uniformly Distributed within Pancreatic Tumour and Tregs Are Restricted to the Stroma
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Patient Recruitment
5.2. Sample Collection
5.3. PBMC Isolation
5.4. Tissue Digestion
5.5. CyTOF Sample Preparation
5.6. CyTOF Data Acquisition
5.7. CyTOF Data Analysis
5.8. Collection of Histological Sections
5.9. Multiplex Immunohistochemistry
5.10. Multiplex Immunohistochemistry-Image Analysis
5.11. Single-Cell RNA Sequencing Analysis: Pre-Processing, Integration and Batch Correction
5.12. Single-Cell RNA Sequencing Analysis: Single Cell Clustering and Annotation
5.13. Single-Cell RNA Sequencing Analysis: Data Analysis and Figures
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kamisawa, T.; Wood, L.D.; Itoi, T.; Takaori, K. Pancreatic cancer. Lancet 2016, 388, 73–85. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Conroy, T.; Hammel, P.; Hebbar, M.; Ben Abdelghani, M.; Wei, A.C.; Raoul, J.-L.; Choné, L.; Francois, E.; Artru, P.; Biagi, J.J. FOLFIRINOX or gemcitabine as adjuvant therapy for pancreatic cancer. N. Engl. J. Med. 2018, 379, 2395–2406. [Google Scholar] [CrossRef]
- De Santiago, I.; Yau, C.; Heij, L.; Middleton, M.R.; Markowetz, F.; Grabsch, H.I.; Dustin, M.L.; Sivakumar, S. Immunophenotypes of pancreatic ductal adenocarcinoma: Meta-analysis of transcriptional subtypes. Int. J. Cancer 2019, 145, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Ino, Y.; Yamazaki-Itoh, R.; Shimada, K.; Iwasaki, M.; Kosuge, T.; Kanai, Y.; Hiraoka, N. Immune cell infiltration as an indicator of the immune microenvironment of pancreatic cancer. Br. J. Cancer 2013, 108, 914–923. [Google Scholar] [CrossRef] [PubMed]
- Clark, C.E.; Hingorani, S.R.; Mick, R.; Combs, C.; Tuveson, D.A.; Vonderheide, R.H. Dynamics of the immune reaction to pancreatic cancer from inception to invasion. Cancer Res. 2007, 67, 9518–9527. [Google Scholar] [CrossRef]
- Knudsen, E.S.; Vail, P.; Balaji, U.; Ngo, H.; Botros, I.W.; Makarov, V.; Riaz, N.; Balachandran, V.; Leach, S.; Thompson, D.M. Stratification of pancreatic ductal adenocarcinoma: Combinatorial genetic, stromal, and immunologic markers. Clin. Cancer Res. 2017, 23, 4429–4440. [Google Scholar] [CrossRef]
- Bayne, L.J.; Beatty, G.L.; Jhala, N.; Clark, C.E.; Rhim, A.D.; Stanger, B.Z.; Vonderheide, R.H. Tumor-derived granulocyte-macrophage colony-stimulating factor regulates myeloid inflammation and T cell immunity in pancreatic cancer. Cancer Cell 2012, 21, 822–835. [Google Scholar] [CrossRef]
- Roth, S.; Zamzow, K.; Gaida, M.M.; Heikenwälder, M.; Tjaden, C.; Hinz, U.; Bose, P.; Michalski, C.W.; Hackert, T. Evolution of the immune landscape during progression of pancreatic intraductal papillary mucinous neoplasms to invasive cancer. EBioMedicine 2020, 54. [Google Scholar] [CrossRef]
- Martinez-Useros, J.; Martin-Galan, M.; Garcia-Foncillas, J. The Match between Molecular Subtypes, Histology and Microenvironment of Pancreatic Cancer and Its Relevance for Chemoresistance. Cancers 2021, 13, 322. [Google Scholar] [CrossRef] [PubMed]
- Brahmer, J.; Reckamp, K.L.; Baas, P.; Crinò, L.; Eberhardt, W.E.; Poddubskaya, E.; Antonia, S.; Pluzanski, A.; Vokes, E.E.; Holgado, E. Nivolumab versus docetaxel in advanced squamous-cell non–small-cell lung cancer. N. Engl. J. Med. 2015, 373, 123–135. [Google Scholar] [CrossRef]
- Robert, C.; Schachter, J.; Long, G.V.; Arance, A.; Grob, J.J.; Mortier, L.; Daud, A.; Carlino, M.S.; McNeil, C.; Lotem, M. Pembrolizumab versus ipilimumab in advanced melanoma. N. Engl. J. Med. 2015, 372, 2521–2532. [Google Scholar] [CrossRef]
- Aglietta, M.; Barone, C.; Sawyer, M.; Moore, M.; Miller, W., Jr.; Bagalà, C.; Colombi, F.; Cagnazzo, C.; Gioeni, L.; Wang, E. A phase I dose escalation trial of tremelimumab (CP-675,206) in combination with gemcitabine in chemotherapy-naive patients with metastatic pancreatic cancer. Ann. Oncol. 2014, 25, 1750–1755. [Google Scholar] [CrossRef]
- Beatty, G.L.; Torigian, D.A.; Chiorean, E.G.; Saboury, B.; Brothers, A.; Alavi, A.; Troxel, A.B.; Sun, W.; Teitelbaum, U.R.; Vonderheide, R.H. A phase I study of an agonist CD40 monoclonal antibody (CP-870,893) in combination with gemcitabine in patients with advanced pancreatic ductal adenocarcinoma. Clin. Cancer Res. 2013, 19, 6286–6295. [Google Scholar] [CrossRef] [PubMed]
- O’Reilly, E.M.; Oh, D.-Y.; Dhani, N.; Renouf, D.J.; Lee, M.A.; Sun, W.; Fisher, G.; Hezel, A.; Chang, S.-C.; Vlahovic, G. Durvalumab with or without tremelimumab for patients with metastatic pancreatic ductal adenocarcinoma: A phase 2 randomized clinical trial. JAMA Oncol. 2019, 5, 1431–1438. [Google Scholar] [CrossRef]
- Young, K.; Hughes, D.J.; Cunningham, D.; Starling, N. Immunotherapy and pancreatic cancer: Unique challenges and potential opportunities. Ther. Adv. Med. Oncol. 2018, 10, 1758835918816281. [Google Scholar] [CrossRef]
- Mahalingam, D.; Goel, S.; Aparo, S.; Patel Arora, S.; Noronha, N.; Tran, H.; Chakrabarty, R.; Selvaggi, G.; Gutierrez, A.; Coffey, M. A phase II study of pelareorep (REOLYSIN®) in combination with gemcitabine for patients with advanced pancreatic adenocarcinoma. Cancers 2018, 10, 160. [Google Scholar] [CrossRef] [PubMed]
- Carstens, J.L.; De Sampaio, P.C.; Yang, D.; Barua, S.; Wang, H.; Rao, A.; Allison, J.P.; LeBleu, V.S.; Kalluri, R. Spatial computation of intratumoral T cells correlates with survival of patients with pancreatic cancer. Nat. Commun. 2017, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Wartenberg, M.; Zlobec, I.; Perren, A.; Koelzer, V.H.; Gloor, B.; Lugli, A.; Eva, K. Accumulation of FOXP3+ T-cells in the tumor microenvironment is associated with an epithelial-mesenchymal-transition-type tumor budding phenotype and is an independent prognostic factor in surgically resected pancreatic ductal adenocarcinoma. Oncotarget 2015, 6, 4190. [Google Scholar] [CrossRef] [PubMed]
- Krishnamoorthy, M.; Lenehan, J.G.; Burton, J.P.; Maleki Vareki, S. Immunomodulation in Pancreatic Cancer. Cancers 2020, 12, 3340. [Google Scholar] [CrossRef] [PubMed]
- Jang, J.-E.; Hajdu, C.H.; Liot, C.; Miller, G.; Dustin, M.L.; Bar-Sagi, D. Crosstalk between regulatory T cells and tumor-associated dendritic cells negates anti-tumor immunity in pancreatic cancer. Cell Rep. 2017, 20, 558–571. [Google Scholar] [CrossRef] [PubMed]
- Leao, I.C.; Ganesan, P.; Armstrong, T.D.; Jaffee, E.M. Effective Depletion of Regulatory T Cells Allows the Recruitment of Mesothelin-Specific CD8+ T Cells to the Antitumor Immune Response Against a Mesothelin-Expressing Mouse Pancreatic Adenocarcinoma. Clin. Transl. Sci. 2008, 1, 228–239. [Google Scholar] [CrossRef]
- Zhang, Y.; Lazarus, J.; Steele, N.G.; Yan, W.; Lee, H.-J.; Nwosu, Z.C.; Halbrook, C.J.; Menjivar, R.E.; Kemp, S.B.; Sirihorachai, V.; et al. Regulatory T cell depletion alters the tumor microenvironment and accelerates pancreatic carcinogenesis. Cancer Discov. 2020. [Google Scholar] [CrossRef] [PubMed]
- Popovic, A.; Jaffee, E.M.; Zaidi, N. Emerging strategies for combination checkpoint modulators in cancer immunotherapy. J. Clin. Investig. 2018, 128, 3209–3218. [Google Scholar] [CrossRef]
- Mahoney, K.M.; Rennert, P.D.; Freeman, G.J. Combination cancer immunotherapy and new immunomodulatory targets. Nat. Rev. Drug Discov. 2015, 14, 561–584. [Google Scholar] [CrossRef]
- Pandol, S.; Edderkaoui, M.; Gukovsky, I.; Lugea, A.; Gukovskaya, A. Desmoplasia of Pancreatic Ductal Adenocarcinoma. Clin. Gastroenterol. Hepatol. 2009, 7, S44–S47. [Google Scholar] [CrossRef]
- Khaled, Y.S.; Ammori, B.J.; Elkord, E. Increased Levels of Granulocytic Myeloid-Derived Suppressor Cells in Peripheral Blood and Tumour Tissue of Pancreatic Cancer Patients. J. Immunol. Res. 2014, 2014, 879897. [Google Scholar] [CrossRef] [PubMed]
- Pillay, J.; Tak, T.; Kamp, V.M.; Koenderman, L. Immune suppression by neutrophils and granulocytic myeloid-derived suppressor cells: Similarities and differences. Cell Mol. Life Sci. 2013, 70, 3813–3827. [Google Scholar] [CrossRef]
- Singhal, S.; Bhojnagarwala, P.S.; O’Brien, S.; Moon, E.K.; Garfall, A.L.; Rao, A.S.; Quatromoni, J.G.; Stephen, T.L.; Litzky, L.; Deshpande, C.; et al. Origin and Role of a Subset of Tumor-Associated Neutrophils with Antigen-Presenting Cell Features in Early-Stage Human Lung Cancer. Cancer Cell 2016, 30, 120–135. [Google Scholar] [CrossRef]
- Panni, R.Z.; Sanford, D.E.; Belt, B.A.; Mitchem, J.B.; Worley, L.A.; Goetz, B.D.; Mukherjee, P.; Wang-Gillam, A.; Link, D.C.; DeNardo, D.G.; et al. Tumor-induced STAT3 activation in monocytic myeloid-derived suppressor cells enhances stemness and mesenchymal properties in human pancreatic cancer. Cancer Immunol. Immunother. 2014, 63, 513–528. [Google Scholar] [CrossRef]
- Hsu, B.E.; Tabariès, S.; Johnson, R.M.; Andrzejewski, S.; Senecal, J.; Lehuédé, C.; Annis, M.G.; Ma, E.H.; Völs, S.; Ramsay, L. Immature low-density neutrophils exhibit metabolic flexibility that facilitates breast cancer liver metastasis. Cell Rep. 2019, 27, 3902–3915.e3906. [Google Scholar] [CrossRef] [PubMed]
- Burel, J.G.; Pomaznoy, M.; Lindestam Arlehamn, C.S.; Weiskopf, D.; da Silva Antunes, R.; Jung, Y.; Babor, M.; Schulten, V.; Seumois, G.; Greenbaum, J.A.; et al. Circulating T cell-monocyte complexes are markers of immune perturbations. Elife 2019, 8, e46045. [Google Scholar] [CrossRef]
- Pangrazzi, L.; Weinberger, B. T cells, aging and senescence. Exp. Gerontol. 2020, 134, 110887. [Google Scholar] [CrossRef]
- Pereira, B.I.; de Maeyer, R.P.H.; Covre, L.P.; Nehar-Belaid, D.; Lanna, A.; Ward, S.; Marches, R.; Chambers, E.S.; Gomes, D.C.O.; Riddell, N.E.; et al. Sestrins induce natural killer function in senescent-like CD8+ T cells. Nat. Immunol. 2020, 21, 684–694. [Google Scholar] [CrossRef]
- Chou, J.P.; Effros, R.B. T cell replicative senescence in human aging. Curr. Pharm. Des. 2013, 19, 1680–1698. [Google Scholar] [CrossRef]
- Liu, X.; Hoft, D.F.; Peng, G. Senescent T cells within suppressive tumor microenvironments: Emerging target for tumor immunotherapy. J. Clin. Investig. 2020, 130, 1073–1083. [Google Scholar] [CrossRef] [PubMed]
- Ghorani, E.; Reading, J.L.; Henry, J.Y.; Massy, M.R.d.; Rosenthal, R.; Turati, V.; Joshi, K.; Furness, A.J.S.; Ben Aissa, A.; Saini, S.K.; et al. The T cell differentiation landscape is shaped by tumour mutations in lung cancer. Nat. Cancer 2020, 1, 546–561. [Google Scholar] [CrossRef] [PubMed]
- Fourcade, J.; Sun, Z.; Chauvin, J.-M.; Ka, M.; Davar, D.; Pagliano, O.; Wang, H.; Saada, S.; Menna, C.; Amin, R.; et al. CD226 opposes TIGIT to disrupt Tregs in melanoma. JCI Insight 2018, 3. [Google Scholar] [CrossRef]
- Fuhrman, C.A.; Yeh, W.-I.; Seay, H.R.; Saikumar Lakshmi, P.; Chopra, G.; Zhang, L.; Perry, D.J.; McClymont, S.A.; Yadav, M.; Lopez, M.-C.; et al. Divergent Phenotypes of Human Regulatory T Cells Expressing the Receptors TIGIT and CD226. J. Immunol. 2015. [Google Scholar] [CrossRef]
- Joller, N.; Lozano, E.; Burkett, P.R.; Patel, B.; Xiao, S.; Zhu, C.; Xia, J.; Tan, T.G.; Sefik, E.; Yajnik, V.; et al. Treg cells expressing the coinhibitory molecule TIGIT selectively inhibit proinflammatory Th1 and Th17 cell responses. Immunity 2014, 40, 569–581. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Mo, L.; Cai, X.; Wei, L.; Xie, Z.; Li, H.; Li, J.; Hu, Z. ICOS signal facilitates Foxp3 transcription to favor suppressive function of regulatory T cells. Int. J. Med. Sci. 2018, 15, 666–673. [Google Scholar] [CrossRef] [PubMed]
- Vocanson, M.; Rozieres, A.; Hennino, A.; Poyet, G.; Gaillard, V.; Renaudineau, S.; Achachi, A.; Benetiere, J.; Kaiserlian, D.; Dubois, B.; et al. Inducible costimulator (ICOS) is a marker for highly suppressive antigen-specific T cells sharing features of TH17/TH1 and regulatory T cells. J. Allergy Clin. Immunol. 2010, 126, 280–289.e7. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Sun, B.F.; Chen, C.Y.; Zhou, J.Y.; Chen, Y.S.; Chen, H.; Liu, L.; Huang, D.; Jiang, J.; Cui, G.S.; et al. Single-cell RNA-seq highlights intra-tumoral heterogeneity and malignant progression in pancreatic ductal adenocarcinoma. Cell Res. 2019, 29, 725–738. [Google Scholar] [CrossRef]
- Khan, O.; Giles, J.R.; McDonald, S.; Manne, S.; Ngiow, S.F.; Patel, K.P.; Werner, M.T.; Huang, A.C.; Alexander, K.A.; Wu, J.E.; et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 2019, 571, 211–218. [Google Scholar] [CrossRef]
- Zheng, C.; Zheng, L.; Yoo, J.-K.; Guo, H.; Zhang, Y.; Guo, X.; Kang, B.; Hu, R.; Huang, J.Y.; Zhang, Q.; et al. Landscape of Infiltrating T Cells in Liver Cancer Revealed by Single-Cell Sequencing. Cell 2017, 169, 1342–1356.e1316. [Google Scholar] [CrossRef]
- Hiraoka, N.; Onozato, K.; Kosuge, T.; Hirohashi, S. Prevalence of FOXP3+ Regulatory T Cells Increases During the Progression of Pancreatic Ductal Adenocarcinoma and Its Premalignant Lesions. Clin. Cancer Res. 2006, 12, 5423–5434. [Google Scholar] [CrossRef] [PubMed]
- Ho, W.J.; Jaffee, E.M.; Zheng, L. The tumour microenvironment in pancreatic cancer—Clinical challenges and opportunities. Nat. Rev. Clin. Oncol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Steele, N.G.; Carpenter, E.S.; Kemp, S.B.; Sirihorachai, V.R.; The, S.; Delrosario, L.; Lazarus, J.; Amir, E.-A.D.; Gunchick, V.; Espinoza, C. Multimodal mapping of the tumor and peripheral blood immune landscape in human pancreatic cancer. Nat. Cancer 2020, 1–16. [Google Scholar] [CrossRef]
- Lucca, L.E.; Axisa, P.-P.; Singer, E.R.; Nolan, N.M.; Dominguez-Villar, M.; Hafler, D.A. TIGIT signaling restores suppressor function of Th1 Tregs. JCI Insight 2019, 4, e124427. [Google Scholar] [CrossRef]
- Jin, H.-s.; Ko, M.; Choi, D.-s.; Kim, J.H.; Lee, D.-h.; Kang, S.-H.; Kim, I.; Lee, H.J.; Choi, E.K.; Kim, K.-p.; et al. CD226hiCD8+ T Cells Are a Prerequisite for Anti-TIGIT Immunotherapy. Cancer Immunol. Res. 2020. [Google Scholar] [CrossRef]
- Luke, J.J.; Powderly, J.D.; Merchan, J.R.; Barve, M.; Hotson, A.; Mobasher, M.; Kwei, L.; Luciano, G.; Buggy, J.; Piccione, E. Immunobiology, preliminary safety, and efficacy of CPI-006, an anti-CD73 antibody with immune modulating activity, in a phase 1 trial in advanced cancers. J. Clin. Oncol. 2019, 37, 2505. [Google Scholar] [CrossRef]
- Sainson, R.C.; Thotakura, A.K.; Kosmac, M.; Borhis, G.; Parveen, N.; Kimber, R.; Carvalho, J.; Henderson, S.J.; Pryke, K.L.; Okell, T. An Antibody Targeting ICOS Increases Intratumoral Cytotoxic to Regulatory T-cell Ratio and Induces Tumor Regression. Cancer Immunol. Res. 2020, 8, 1568–1582. [Google Scholar] [CrossRef]
- Wu, P.F.; Lu, Z.P.; Cai, B.B.; Tian, L.; Zou, C.; Jiang, K.R.; Miao, Y. Role of CXCL12/CXCR4 signaling axis in pancreatic cancer. Chin. Med. J. 2013, 126, 3371–3374. [Google Scholar]
- Moo-Young, T.A.; Larson, J.W.; Belt, B.A.; Tan, M.C.; Hawkins, W.G.; Eberlein, T.J.; Goedegebuure, P.S.; Linehan, D.C. Tumor-derived TGF-beta mediates conversion of CD4+Foxp3+ regulatory T cells in a murine model of pancreas cancer. J. Immunother. 2009, 32, 12–21. [Google Scholar] [CrossRef]
- Sakamoto, H.; Attiyeh, M.A.; Gerold, J.M.; Makohon-Moore, A.P.; Hayashi, A.; Hong, J.; Kappagantula, R.; Zhang, L.; Melchor, J.P.; Reiter, J.G.; et al. The Evolutionary Origins of Recurrent Pancreatic Cancer. Cancer Discov. 2020, 10, 792–805. [Google Scholar] [CrossRef]
- Kasakovski, D.; Xu, L.; Li, Y. T cell senescence and CAR-T cell exhaustion in hematological malignancies. J. Hematol. Oncol. 2018, 11, 91. [Google Scholar] [CrossRef] [PubMed]
- Nakagami, H. Cellular senescence and senescence-associated T cells as a potential therapeutic target. Geriatr. Gerontol. Int. 2020, 20, 97–100. [Google Scholar] [CrossRef] [PubMed]
- Lanna, A.; Henson, S.M.; Escors, D.; Akbar, A.N. The kinase p38 activated by the metabolic regulator AMPK and scaffold TAB1 drives the senescence of human T cells. Nat. Immunol. 2014, 15, 965–972. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Shao, Q.; Peng, G. Exhaustion and senescence: Two crucial dysfunctional states of T cells in the tumor microenvironment. Cell. Mol. Immunol. 2020, 17, 27–35. [Google Scholar] [CrossRef]
- Liu, X.; Mo, W.; Ye, J.; Li, L.; Zhang, Y.; Hsueh, E.C.; Hoft, D.F.; Peng, G. Regulatory T cells trigger effector T cell DNA damage and senescence caused by metabolic competition. Nat. Commun. 2018, 9, 249. [Google Scholar] [CrossRef]
- Huber, M.; Brehm, C.U.; Gress, T.M.; Buchholz, M.; Alashkar Alhamwe, B.; von Strandmann, E.P.; Slater, E.P.; Bartsch, J.W.; Bauer, C.; Lauth, M. The Immune Microenvironment in Pancreatic Cancer. Int. J. Mol. Sci. 2020, 21, 7307. [Google Scholar] [CrossRef] [PubMed]
- Malik, N.K.; May, K.S.; Chandrasekhar, R.; Wee, W.; Flaherty, L.; Iyer, R.; Gibbs, J.; Kuvshinoff, B.; Wilding, G.; Warren, G. Treatment of locally advanced unresectable pancreatic cancer: A 10-year experience. J. Gastrointest. Oncol. 2012, 3, 326. [Google Scholar] [PubMed]
- Butler, A.; Hoffman, P.; Smibert, P.; Papalexi, E.; Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 2018, 36, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Korsunsky, I.; Millard, N.; Fan, J.; Slowikowski, K.; Zhang, F.; Wei, K.; Baglaenko, Y.; Brenner, M.; Loh, P.-R.; Raychaudhuri, S. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 2019, 16, 1289–1296. [Google Scholar] [CrossRef]
- Yang, C.; Siebert, J.R.; Burns, R.; Gerbec, Z.J.; Bonacci, B.; Rymaszewski, A.; Rau, M.; Riese, M.J.; Rao, S.; Carlson, K.-S.; et al. Heterogeneity of human bone marrow and blood natural killer cells defined by single-cell transcriptome. Nat. Commun. 2019, 10, 3931. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sivakumar, S.; Abu-Shah, E.; Ahern, D.J.; Arbe-Barnes, E.H.; Jainarayanan, A.K.; Mangal, N.; Reddy, S.; Rendek, A.; Easton, A.; Kurz, E.; et al. Activated Regulatory T-Cells, Dysfunctional and Senescent T-Cells Hinder the Immunity in Pancreatic Cancer. Cancers 2021, 13, 1776. https://doi.org/10.3390/cancers13081776
Sivakumar S, Abu-Shah E, Ahern DJ, Arbe-Barnes EH, Jainarayanan AK, Mangal N, Reddy S, Rendek A, Easton A, Kurz E, et al. Activated Regulatory T-Cells, Dysfunctional and Senescent T-Cells Hinder the Immunity in Pancreatic Cancer. Cancers. 2021; 13(8):1776. https://doi.org/10.3390/cancers13081776
Chicago/Turabian StyleSivakumar, Shivan, Enas Abu-Shah, David J. Ahern, Edward H. Arbe-Barnes, Ashwin K. Jainarayanan, Nagina Mangal, Srikanth Reddy, Aniko Rendek, Alistair Easton, Elke Kurz, and et al. 2021. "Activated Regulatory T-Cells, Dysfunctional and Senescent T-Cells Hinder the Immunity in Pancreatic Cancer" Cancers 13, no. 8: 1776. https://doi.org/10.3390/cancers13081776
APA StyleSivakumar, S., Abu-Shah, E., Ahern, D. J., Arbe-Barnes, E. H., Jainarayanan, A. K., Mangal, N., Reddy, S., Rendek, A., Easton, A., Kurz, E., Silva, M., Soonawalla, Z., Heij, L. R., Bashford-Rogers, R., Middleton, M. R., & Dustin, M. L. (2021). Activated Regulatory T-Cells, Dysfunctional and Senescent T-Cells Hinder the Immunity in Pancreatic Cancer. Cancers, 13(8), 1776. https://doi.org/10.3390/cancers13081776






