miRNAs and lncRNAs as Novel Therapeutic Targets to Improve Cancer Immunotherapy
Abstract
Simple Summary
Abstract
1. Cancer Immunotherapy
2. Noncoding RNA Biogenesis and Function: An Overview
3. In Silico Approaches to Investigate Immune-Related ncRNA
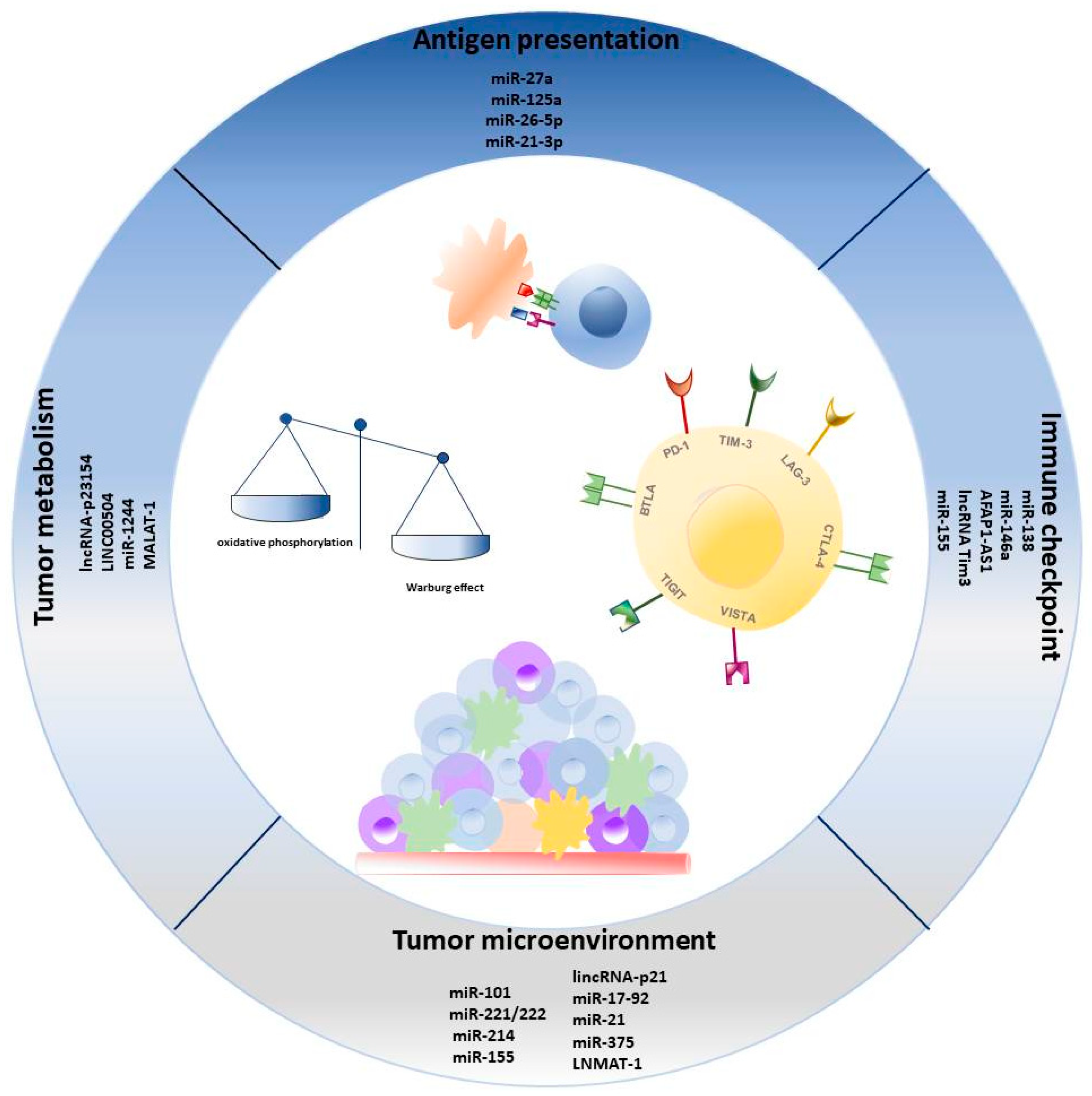
4. ncRNA in Immune Escape
4.1. ncRNAs Regulate Tumor Antigen Presentation
4.2. Role of ncRNAs in Tumor Metabolism
4.3. ncRNAs as Crucial Players in TME
4.4. ncRNA and Immune Checkpoint
5. ncRNAs in Immunotherapy Resistance
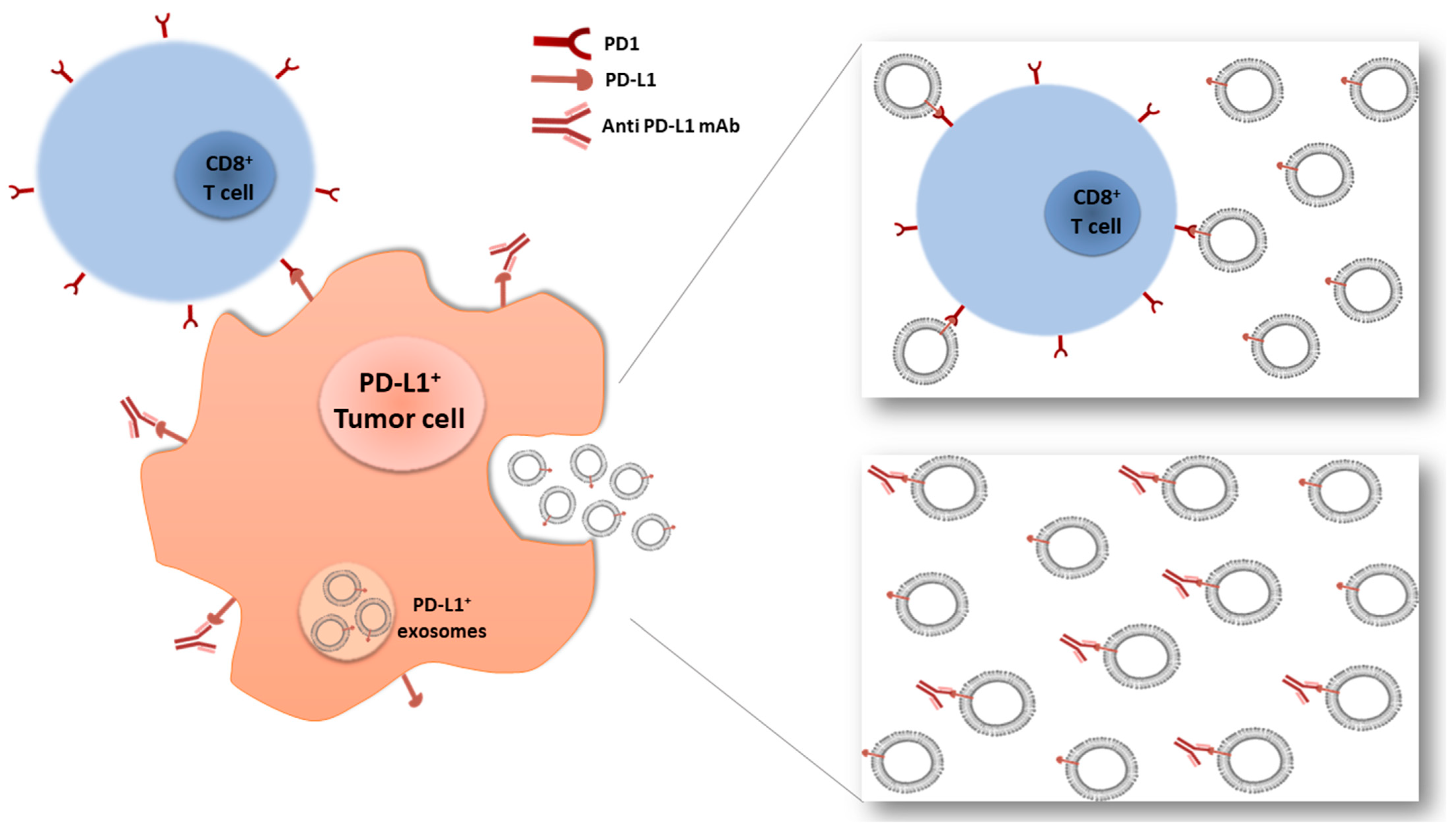
6. Role of Exosomal ncRNA as Anticancer and Drug Resistance Cargo
| Regulatory Function | ncRNA Name | Target Name | Target Modulation | Tumor Type | Ref. | |
|---|---|---|---|---|---|---|
| miRNAs | lncRNAs | |||||
| Tumor antigen presentation | miR-27a | TAP2 | Downregulation of MHC-I expression | Esophageal adenocarcinoma | [47] | |
| miR-26-5p and miR-21-3p | TAP1 | Downregulation of TAP1 and reduced expression of HLA class I cell surface antigens | Melanoma | [49] | ||
| Tumor metabolism | lncRNA p23154 | miR-378a-3p | Repression of Glut1 expression through miR-378a-3p binding to the UTR of the gene | Oral squamous cell carcinoma | [51] | |
| lncRNA LINC00504 | miR-1244 | Stimulation of aerobic glycolysis through PKM2, HK2, and PDK1 | Ovarian cancer | [52] | ||
| miR133a-3p | GABARAPL1 | Blockade of glutaminolysis by the reduction of the expression level of core enzymes including GLS and GDH | Gastric cancer | [54] | ||
| MALAT-1 | Promote VEGF expression not only through a direct pathway, but also through miRNAs, which deserves other studies | Immunosuppressive Properties of Mesenchymal Stem Cells (MSC) by Inducing VEGF and IDO | Mesenchymal stem cells | [55] | ||
| Tumor Microenvironment | miR-149 | IL-6 | Inhibition of the activation of tumor-promoting fibroblasts by the reduction of IL-6 expression | Gastric Cancer | [63] | |
| miR-101 | CXCL12 | Inhibition of the interaction between fibroblasts and cancer cells by the downregulation of CXCL12 | Lung cancer | [64] | ||
| miR-222 | LBR | Downregulation of LBR by inducing normal fibroblasts to show the cancer-associated fibroblast (CAF) characteristics | Breast cancer | [65] | ||
| miR-221 | A20 | Stimulatory action in breast cancer cells and in main component of the TME such asCAFs, through the involvement of A20/c-Rel/CTGF signaling | Breast cancer | [66,67,68,69,70,71] | ||
| lincRNA-p21 | P53 | Direct targeting on p53, abolishment of MDM2 degradation to p53 by facilitating phenotype maintenance of Tumor-associated macrophages (TAMs) | Breast Cancer | [86] | ||
| LNMAT1 | CCL2 | CCL2 upregulation, macrophage recruitment, and metastasis spreading | Bladder cancer | [87] | ||
| miR-375 | PXN and TNS3 | Destabilization of PXN and TNS3, TAM infiltration | Breast cancer | [88] | ||
| miR-21 | OPG | OPG downmodulation and RANKL upregulation by playing a role in bone resorption/apposition balance | Multiple Myeloma | [81] | ||
| Immune Checkpoint | miR-138 | CTLA-4 and PD-1 | Inhibition of human checkpoint expression in Tregs. Downmodulation of CTLA-4, PD-1, and FoxP3 in CD4+ T cells | Glioma | [91] | |
| miR-155 | IL7R | Repression of IL7R expression in response to activation signals by regulating T cell survival, homeostasis, and proliferation | Melanoma | [79] | ||
| miR-34 | PDL1 | Downregulation of PDL-1 | Non small cell lung cancer (NSCLC) | [95] | ||
| miR-146a | IFNY-STAT1 | Upregulation of PDL-1 | Melanoma | [96] | ||
| lncRNA AFAP1-AS1 | PDC1 | Upregulation of PD-1 | Nasopharyngeal carcinoma | [97] | ||
| lncRNA Tim3 | TIM3 | Binding of Tim3 and nuclear translocation of Bat3 | Hepatocellular carcinoma | [99] | ||
| miR-155 | BTLA | Downregulation of BTLA surface expression | Tumor microenvironment | [101] | ||
| Immunotherapy resistance | NEAT1 | miR-155/Tim-3 | Downregulation of miR-155 and Tim-3 upregulation | Hepatocellular carcinoma | [105] | |
| MALAT1 | Upregulation, through miR-195, of PD-L1 Inhibition of MALAT1 interaction with miR-101, modulation of cisplatin, and temozolomide resistance | Diffuse large B-cell lymphoma Lung cancer and glioblastoma | [72,110,111] | |||
| ExosomalncRNAs As drug resistance cargo | lncRNA Olfr29-ps1 | miR-214-3p | Sponging of miR-214-3p and downregulation of miR-214-3p, which target MyD88 to modulate differentiation and function of MDSCs | Tumor microenvironment | [113] | |
| Lnc-chop | CHOP and the C/EBPβ isoform liver-enriched inhibitory protein | Activation of C/EBPβ, upregulation of arginase-1, NO synthase 2, NADPH oxidase 2, and cyclooxygenase-2 | Tumor microenvironment | [114] | ||
| lnc- POU3F3 | TGF-β | Upregulation of TGF-β, distribution of Tregs in peripheral blood, enhance cell proliferation of gastric cancer | Gastric cancer | [116] | ||
| miR-23a-3p | PTEN, AKT | PDL-1 upregulation in macrophages | Hepatocarcinoma | [123] | ||
| hsa-miR-24-3p, hsa-miR-891a, hsa-miR-106a-5p, hsa-miR-20a-5p, and hsa-miR-1908 | MARK1 | Downregulation of the MARK1 signaling pathway | Nasopharyngeal carcinoma | [124] | ||
| ZFAS1 | D1, Bcl2, N-cadherin, Slug, Snail, Twist, Bax and E-cadherin | Upregulation of D1, Bcl2, N-cadherin, Slug, Snail, Twist, and ZEB1 and downregulation of Bax and E-cadherin | Gastric cancer | [125] | ||
| MALAT-1 | cyclinD1, cyclinD2 and CDK | Upregulation of cyclinD1, cyclinD2, and CDK; tumor growth promotion; migration; and apoptosis prevention in lung cancer cell lines | NSCLC | [126] | ||
| lncRNA UCA1 | E-cadherin, vimentin, MMP9 proteins | Decreasing of E-cadherin, increasing of vimentin and MMP9 | Bladder cancer | [127] | ||
| lncRNA SNHG16 | acts as ce-RNA via sponging miR-16-5p | De-repression of miR-16-5p targets, SMAD5 among these | Breast cancer | [128] | ||
| lncRNA RPPH1 | TUBB3 | Interaction with TUBB3 to prevent its ubiquitination, macrophage M2 polarization, metastasis spreading, and proliferation of colon cancer cells | Colorectal cancer metastasis | [129] | ||
| lncRNA LINK-A | PtdIns (3,4,5) P3, inhibitory GCPRs, E3 ubiquitin ligase TRIM71 | Enhancement of K48–polyubiquitination-mediated degradation of the antigen peptide-loading complex (PLC), and Rb and p53 | Triple negative breast cancer | [106,107] | ||
| miR-21-5p and miR-155-5p | BRG1 | Downregulation of BRG1 leading to colorectal cancer cells migration and invasion. | Colorectal cancer metastasis | [130] | ||
| Algorithm or Signature ID | Freely Available | Description | Reference |
|---|---|---|---|
| ImmunemiR | http://www.biominingbu.org/immunemir/, accessed on 17 September 2020 | repository for immune-related disease and miRNA associations | Prabahar A. et al. [35] |
| ncRI | http://www.jianglab.cn/ncRI/, accessed on 27 March 2021 | comprehensive repository of ncRNAs and their rolesin inflammatory disease | Wang S. et al. [36] |
| IRlncRs | https://rdcu.be/ceXm7, accessed on 27 March 2021 | immune-related risk score (IRRS) in RCC | Jiang Y. et al. [37] |
| 11 immune-related lncRNAs signature | https://rdcu.be/ceXFC, accessed on 17 September 2020 | immune-related lncRNAs for glioma risk score formula | Xia P. et al. [38] |
| ImmLnc | https://rdcu.be/ceXEP, accessed on 17 September 2020 | integrated algorithm for identifying lncRNA regulators of immune-related pathways | Li Y. et al. [39] |
| 16 lncRNAs signatures | https://rdcu.be/ceXGg, accessed on 17 September 2020 | lncRNAs model for automatic microsatellite instability (MSI) classification using a machine learning technology | Chen T. et al. [40] |
7. ncRNA-Based Therapeutic Approaches
8. ncRNAs and Immune Checkpoint Inhibitors: Current Clinical Evidence and Future Perspectives
9. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- O’Donnell, J.S.; Teng, M.W.L.; Smyth, M.J. Cancer immunoediting and resistance to T cell-based immunotherapy. Nat. Rev. Clin. Oncol. 2019, 16, 151–167. [Google Scholar] [CrossRef]
- Chen, D.S.; Mellman, I. Oncology meets immunology: The cancer-immunity cycle. Immunity 2013, 39, 1–10. [Google Scholar] [CrossRef]
- Dobosz, P.; Dzieciatkowski, T. The Intriguing History of Cancer Immunotherapy. Front. Immunol. 2019, 10, 2965. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Vacchelli, E.; Bravo-San Pedro, J.M.; Buque, A.; Senovilla, L.; Baracco, E.E.; Bloy, N.; Castoldi, F.; Abastado, J.P.; Agostinis, P.; et al. Classification of current anticancer immunotherapies. Oncotarget 2014, 5, 12472–12508. [Google Scholar] [CrossRef] [PubMed]
- Kapranov, P.; Willingham, A.T.; Gingeras, T.R. Genome-wide transcription and the implications for genomic organization. Nat. Rev. Genet. 2007, 8, 413–423. [Google Scholar] [CrossRef]
- Abulwerdi, F.A.; Xu, W.; Ageeli, A.A.; Yonkunas, M.J.; Arun, G.; Nam, H.; Schneekloth, J.S., Jr.; Dayie, T.K.; Spector, D.; Baird, N.; et al. Selective Small-Molecule Targeting of a Triple Helix Encoded by the Long Noncoding RNA, MALAT1. ACS Chem. Biol. 2019, 14, 223–235. [Google Scholar] [CrossRef] [PubMed]
- Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.C.; Maeda, N.; Oyama, R.; Ravasi, T.; Lenhard, B.; Wells, C.; et al. The transcriptional landscape of the mammalian genome. Science 2005, 309, 1559–1563. [Google Scholar] [CrossRef]
- Seal, R.L.; Chen, L.L.; Griffiths-Jones, S.; Lowe, T.M.; Mathews, M.B.; O’Reilly, D.; Pierce, A.J.; Stadler, P.F.; Ulitsky, I.; Wolin, S.L.; et al. A guide to naming human non-coding RNA genes. EMBO J. 2020, 39, e103777. [Google Scholar] [CrossRef]
- Huttenhofer, A.; Schattner, P.; Polacek, N. Non-coding RNAs: Hope or hype? Trends Genet. TIG 2005, 21, 289–297. [Google Scholar] [CrossRef]
- Saini, H.K.; Griffiths-Jones, S.; Enright, A.J. Genomic analysis of human microRNA transcripts. Proc. Natl. Acad. Sci. USA 2007, 104, 17719–17724. [Google Scholar] [CrossRef]
- Carthew, R.W.; Sontheimer, E.J. Origins and Mechanisms of miRNAs and siRNAs. Cell 2009, 136, 642–655. [Google Scholar] [CrossRef] [PubMed]
- Ruby, J.G.; Jan, C.H.; Bartel, D.P. Intronic microRNA precursors that bypass Drosha processing. Nature 2007, 448, 83–86. [Google Scholar] [CrossRef]
- Bohnsack, M.T.; Czaplinski, K.; Gorlich, D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 2004, 10, 185–191. [Google Scholar] [CrossRef]
- Hutvagner, G.; McLachlan, J.; Pasquinelli, A.E.; Balint, E.; Tuschl, T.; Zamore, P.D. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 2001, 293, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Kawamata, T.; Seitz, H.; Tomari, Y. Structural determinants of miRNAs for RISC loading and slicer-independent unwinding. Nat. Struct. Mol. Biol. 2009, 16, 953–960. [Google Scholar] [CrossRef] [PubMed]
- Hibio, N.; Hino, K.; Shimizu, E.; Nagata, Y.; Ui-Tei, K. Stability of miRNA 5’terminal and seed regions is correlated with experimentally observed miRNA-mediated silencing efficacy. Sci. Rep. 2012, 2, 996. [Google Scholar] [CrossRef] [PubMed]
- Eulalio, A.; Huntzinger, E.; Izaurralde, E. Getting to the root of miRNA-mediated gene silencing. Cell 2008, 132, 9–14. [Google Scholar] [CrossRef]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from repression to activation: microRNAs can up-regulate translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Bang, C.; Thum, T. Circulating microRNAs as biomarkers and potential paracrine mediators of cardiovascular disease. Circ. Cardiovasc. Genet. 2010, 3, 484–488. [Google Scholar] [CrossRef] [PubMed]
- Viereck, J.; Bang, C.; Foinquinos, A.; Thum, T. Regulatory RNAs and paracrine networks in the heart. Cardiovasc. Res. 2014, 102, 290–301. [Google Scholar] [CrossRef]
- Nie, L.; Wu, H.J.; Hsu, J.M.; Chang, S.S.; Labaff, A.M.; Li, C.W.; Wang, Y.; Hsu, J.L.; Hung, M.C. Long non-coding RNAs: Versatile master regulators of gene expression and crucial players in cancer. Am. J. Transl. Res. 2012, 4, 127–150. [Google Scholar] [PubMed]
- Louro, R.; Smirnova, A.S.; Verjovski-Almeida, S. Long intronic noncoding RNA transcription: Expression noise or expression choice? Genomics 2009, 93, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef] [PubMed]
- Bond, C.S.; Fox, A.H. Paraspeckles: Nuclear bodies built on long noncoding RNA. J. Cell Biol. 2009, 186, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D.; Huarte, M.; Zuk, O.; Carey, B.W.; Cassady, J.P.; et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009, 458, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.A.; Valenstein, M.L.; Yario, T.A.; Tycowski, K.T.; Steitz, J.A. Formation of triple-helical structures by the 3’-end sequences of MALAT1 and MENbeta noncoding RNAs. Proc. Natl. Acad. Sci. USA 2012, 109, 19202–19207. [Google Scholar] [CrossRef]
- Rackham, O.; Shearwood, A.M.; Mercer, T.R.; Davies, S.M.; Mattick, J.S.; Filipovska, A. Long noncoding RNAs are generated from the mitochondrial genome and regulated by nuclear-encoded proteins. RNA 2011, 17, 2085–2093. [Google Scholar] [CrossRef]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, M.T.; Amodio, N.; Tassone, P.; Tagliaferri, P. Functional Analysis of microRNA in Multiple Myeloma. Methods Mol. Biol. 2016, 1375, 181–194. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.W.; Huang, K.; Yang, C.; Kang, C.S. Non-coding RNAs as regulators in epigenetics (Review). Oncol. Rep. 2017, 37, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.T.; Bartolomei, M.S. X-inactivation, imprinting, and long noncoding RNAs in health and disease. Cell 2013, 152, 1308–1323. [Google Scholar] [CrossRef]
- Sleutels, F.; Zwart, R.; Barlow, D.P. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 2002, 415, 810–813. [Google Scholar] [CrossRef]
- Fatica, A.; Bozzoni, I. Long non-coding RNAs: New players in cell differentiation and development. Nat. Rev. Genet. 2014, 15, 7–21. [Google Scholar] [CrossRef] [PubMed]
- Grote, P.; Herrmann, B.G. The long non-coding RNA Fendrr links epigenetic control mechanisms to gene regulatory networks in mammalian embryogenesis. RNA Biol. 2013, 10, 1579–1585. [Google Scholar] [CrossRef] [PubMed]
- Grote, P.; Herrmann, B.G. Long noncoding RNAs in organogenesis: Making the difference. Trends Genet. TIG 2015, 31, 329–335. [Google Scholar] [CrossRef]
- Li, Z.; Rana, T.M. Decoding the noncoding: Prospective of lncRNA-mediated innate immune regulation. RNA Biol. 2014, 11, 979–985. [Google Scholar] [CrossRef]
- Prabahar, A.; Natarajan, J. ImmunemiR—A Database of Prioritized Immune miRNA Disease Associations and its Interactome. MicroRNA 2017, 6, 71–78. [Google Scholar] [CrossRef]
- Wang, S.; Zhou, S.; Liu, H.; Meng, Q.; Ma, X.; Liu, H.; Wang, L.; Jiang, W. ncRI: A manually curated database for experimentally validated non-coding RNAs in inflammation. BMC Genom. 2020, 21, 380. [Google Scholar] [CrossRef]
- Jiang, Y.; Gou, X.; Wei, Z.; Tan, J.; Yu, H.; Zhou, X.; Li, X. Bioinformatics profiling integrating a three immune-related long non-coding RNA signature as a prognostic model for clear cell renal cell carcinoma. Cancer Cell Int. 2020, 20, 166. [Google Scholar] [CrossRef]
- Xia, P.; Li, Q.; Wu, G.; Huang, Y. An Immune-Related lncRNA Signature to Predict Survival In Glioma Patients. Cell. Mol. Neurobiol. 2020. [Google Scholar] [CrossRef]
- Li, Y.; Jiang, T.; Zhou, W.; Li, J.; Li, X.; Wang, Q.; Jin, X.; Yin, J.; Chen, L.; Zhang, Y.; et al. Pan-cancer characterization of immune-related lncRNAs identifies potential oncogenic biomarkers. Nat. Commun. 2020, 11, 1000. [Google Scholar] [CrossRef]
- Chen, T.; Zhang, C.; Liu, Y.; Zhao, Y.; Lin, D.; Hu, Y.; Yu, J.; Li, G. A gastric cancer LncRNAs model for MSI and survival prediction based on support vector machine. BMC Genom. 2019, 20, 846. [Google Scholar] [CrossRef]
- Liu, L.; Wang, Q.; Qiu, Z.; Kang, Y.; Liu, J.; Ning, S.; Yin, Y.; Pang, D.; Xu, S. Noncoding RNAs: The shot callers in tumor immune escape. Signal Transduct. Target. Ther. 2020, 5, 102. [Google Scholar] [CrossRef] [PubMed]
- Beatty, G.L.; Gladney, W.L. Immune escape mechanisms as a guide for cancer immunotherapy. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2015, 21, 687–692. [Google Scholar] [CrossRef]
- Lee, M.Y.; Jeon, J.W.; Sievers, C.; Allen, C.T. Antigen processing and presentation in cancer immunotherapy. J. Immunother. Cancer 2020, 8. [Google Scholar] [CrossRef] [PubMed]
- Colangelo, T.; Polcaro, G.; Ziccardi, P.; Pucci, B.; Muccillo, L.; Galgani, M.; Fucci, A.; Milone, M.R.; Budillon, A.; Santopaolo, M.; et al. Proteomic screening identifies calreticulin as a miR-27a direct target repressing MHC class I cell surface exposure in colorectal cancer. Cell Death Dis. 2016, 7, e2120. [Google Scholar] [CrossRef] [PubMed]
- Mari, L.; Hoefnagel, S.J.M.; Zito, D.; van de Meent, M.; van Endert, P.; Calpe, S.; Sancho Serra, M.D.C.; Heemskerk, M.H.M.; van Laarhoven, H.W.M.; Hulshof, M.; et al. microRNA 125a Regulates MHC-I Expression on Esophageal Adenocarcinoma Cells, Associated With Suppression of Antitumor Immune Response and Poor Outcomes of Patients. Gastroenterology 2018, 155, 784–798. [Google Scholar] [CrossRef] [PubMed]
- Lazaridou, M.F.; Massa, C.; Handke, D.; Mueller, A.; Friedrich, M.; Subbarayan, K.; Tretbar, S.; Dummer, R.; Koelblinger, P.; Seliger, B. Identification of microRNAs Targeting the Transporter Associated with Antigen Processing TAP1 in Melanoma. J. Clin. Med. 2020, 9, 2690. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Huang, Z.; Sheng, W.; Xu, M.D. Emerging roles of long non-coding RNAs in tumor metabolism. J. Hematol. Oncol. 2018, 11, 106. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, X.; Wang, Z.; Hu, Q.; Wu, J.; Li, Y.; Ren, X.; Wu, T.; Tao, X.; Chen, X.; et al. LncRNA-p23154 promotes the invasion-metastasis potential of oral squamous cell carcinoma by regulating Glut1-mediated glycolysis. Cancer Lett. 2018, 434, 172–183. [Google Scholar] [CrossRef]
- Liu, Y.; He, X.; Chen, Y.; Cao, D. Long non-coding RNA LINC00504 regulates the Warburg effect in ovarian cancer through inhibition of miR-1244. Mol. Cell. Biochem. 2020, 464, 39–50. [Google Scholar] [CrossRef]
- Ortiz-Pedraza, Y.; Munoz-Bello, J.O.; Olmedo-Nieva, L.; Contreras-Paredes, A.; Martinez-Ramirez, I.; Langley, E.; Lizano, M. Non-Coding RNAs as Key Regulators of Glutaminolysis in Cancer. Int. J. Mol. Sci. 2020, 21, 2872. [Google Scholar] [CrossRef]
- Zhang, X.; Li, Z.; Xuan, Z.; Xu, P.; Wang, W.; Chen, Z.; Wang, S.; Sun, G.; Xu, J.; Xu, Z. Novel role of miR-133a-3p in repressing gastric cancer growth and metastasis via blocking autophagy-mediated glutaminolysis. J. Exp. Clin. Cancer Res. CR 2018, 37, 320. [Google Scholar] [CrossRef]
- Li, X.; Song, Y.; Liu, F.; Liu, D.; Miao, H.; Ren, J.; Xu, J.; Ding, L.; Hu, Y.; Wang, Z.; et al. Long Non-Coding RNA MALAT1 Promotes Proliferation, Angiogenesis, and Immunosuppressive Properties of Mesenchymal Stem Cells by Inducing VEGF and IDO. J. Cell. Biochem. 2017, 118, 2780–2791. [Google Scholar] [CrossRef]
- Galon, J.; Bruni, D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat. Rev. Drug Discov. 2019, 18, 197–218. [Google Scholar] [CrossRef]
- Zhang, L.; Xu, X.; Su, X. Noncoding RNAs in cancer immunity: Functions, regulatory mechanisms, and clinical application. Mol. Cancer 2020, 19, 48. [Google Scholar] [CrossRef]
- Xu, S.J.; Hu, H.T.; Li, H.L.; Chang, S. The Role of miRNAs in Immune Cell Development, Immune Cell Activation, and Tumor Immunity: With a Focus on Macrophages and Natural Killer Cells. Cells 2019, 8, 1140. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Yang, J.; Yu, J.; Liu, X.; Yu, C.; Hu, J.; Shi, H.; Ma, X. Long Non-coding RNAs: Emerging Roles in the Immunosuppressive Tumor Microenvironment. Front. Oncol. 2020, 10, 48. [Google Scholar] [CrossRef] [PubMed]
- Raimondi, L.; Amodio, N.; Di Martino, M.T.; Altomare, E.; Leotta, M.; Caracciolo, D.; Gulla, A.; Neri, A.; Taverna, S.; D’Aquila, P.; et al. Targeting of multiple myeloma-related angiogenesis by miR-199a-5p mimics: In vitro and in vivo anti-tumor activity. Oncotarget 2014, 5, 3039–3054. [Google Scholar] [CrossRef] [PubMed]
- Botta, C.; Cuce, M.; Pitari, M.R.; Caracciolo, D.; Gulla, A.; Morelli, E.; Riillo, C.; Biamonte, L.; Gallo Cantafio, M.E.; Prabhala, R.; et al. MiR-29b antagonizes the pro-inflammatory tumor-promoting activity of multiple myeloma-educated dendritic cells. Leukemia 2018, 32, 1003–1015. [Google Scholar] [CrossRef]
- Drak Alsibai, K.; Meseure, D. Tumor microenvironment and noncoding RNAs as co-drivers of epithelial-mesenchymal transition and cancer metastasis. Dev. Dyn. Off. Publ. Am. Assoc. Anat. 2018, 247, 405–431. [Google Scholar] [CrossRef]
- Li, P.; Shan, J.X.; Chen, X.H.; Zhang, D.; Su, L.P.; Huang, X.Y.; Yu, B.Q.; Zhi, Q.M.; Li, C.L.; Wang, Y.Q.; et al. Epigenetic silencing of microRNA-149 in cancer-associated fibroblasts mediates prostaglandin E2/interleukin-6 signaling in the tumor microenvironment. Cell Res. 2015, 25, 588–603. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, J.; Liu, Y.; Wu, W.; Li, X.; Wu, Y.; Chen, H.; Zhang, K.; Gu, L. miR-101 represses lung cancer by inhibiting interaction of fibroblasts and cancer cells by down-regulating CXCL12. Biomed. Pharmacother. Biomed. Pharmacother. 2015, 74, 215–221. [Google Scholar] [CrossRef]
- Chatterjee, A.; Jana, S.; Chatterjee, S.; Wastall, L.M.; Mandal, G.; Nargis, N.; Roy, H.; Hughes, T.A.; Bhattacharyya, A. MicroRNA-222 reprogrammed cancer-associated fibroblasts enhance growth and metastasis of breast cancer. Br. J. Cancer 2019, 121, 679–689. [Google Scholar] [CrossRef]
- Di Martino, M.T.; Campani, V.; Misso, G.; Gallo Cantafio, M.E.; Gulla, A.; Foresta, U.; Guzzi, P.H.; Castellano, M.; Grimaldi, A.; Gigantino, V.; et al. In vivo activity of miR-34a mimics delivered by stable nucleic acid lipid particles (SNALPs) against multiple myeloma. PLoS ONE 2014, 9, e90005. [Google Scholar] [CrossRef]
- Gulla, A.; Di Martino, M.T.; Gallo Cantafio, M.E.; Morelli, E.; Amodio, N.; Botta, C.; Pitari, M.R.; Lio, S.G.; Britti, D.; Stamato, M.A.; et al. A 13 mer LNA-i-miR-221 Inhibitor Restores Drug Sensitivity in Melphalan-Refractory Multiple Myeloma Cells. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2016, 22, 1222–1233. [Google Scholar] [CrossRef] [PubMed]
- Gallo Cantafio, M.E.; Nielsen, B.S.; Mignogna, C.; Arbitrio, M.; Botta, C.; Frandsen, N.M.; Rolfo, C.; Tagliaferri, P.; Tassone, P.; Di Martino, M.T. Pharmacokinetics and Pharmacodynamics of a 13-mer LNA-inhibitor-miR-221 in Mice and Non-human Primates. Mol. Ther. Nucleic Acids 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, M.T.; Arbitrio, M.; Fonsi, M.; Erratico, C.A.; Scionti, F.; Caracciolo, D.; Tagliaferri, P.; Tassone, P. Allometric Scaling Approaches for Predicting Human Pharmacokinetic of a Locked Nucleic Acid Oligonucleotide Targeting Cancer-Associated miR-221. Cancers 2019, 12, 27. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, M.T.; Arbitrio, M.; Caracciolo, D.; Scionti, F.; Tagliaferri, P.; Tassone, P. Dose-Finding Study and Pharmacokinetics Profile of the Novel 13-Mer Antisense miR-221 Inhibitor in Sprague-Dawley Rats. Mol. Ther. Nucleic Acids 2020, 20, 73–85. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, M.T.; Gulla, A.; Cantafio, M.E.; Lionetti, M.; Leone, E.; Amodio, N.; Guzzi, P.H.; Foresta, U.; Conforti, F.; Cannataro, M.; et al. In vitro and in vivo anti-tumor activity of miR-221/222 inhibitors in multiple myeloma. Oncotarget 2013, 4, 242–255. [Google Scholar] [CrossRef]
- Santolla, M.F.; Lappano, R.; Cirillo, F.; Rigiracciolo, D.C.; Sebastiani, A.; Abonante, S.; Tassone, P.; Tagliaferri, P.; Di Martino, M.T.; Maggiolini, M.; et al. miR-221 stimulates breast cancer cells and cancer-associated fibroblasts (CAFs) through selective interference with the A20/c-Rel/CTGF signaling. J. Exp. Clin. Cancer Res. CR 2018, 37, 94. [Google Scholar] [CrossRef]
- Wang, R.; Sun, Y.; Yu, W.; Yan, Y.; Qiao, M.; Jiang, R.; Guan, W.; Wang, L. Downregulation of miRNA-214 in cancer-associated fibroblasts contributes to migration and invasion of gastric cancer cells through targeting FGF9 and inducing EMT. J. Exp. Clin. Cancer Res. CR 2019, 38, 20. [Google Scholar] [CrossRef]
- Colvin, E.K.; Howell, V.M.; Mok, S.C.; Samimi, G.; Vafaee, F. Expression of long noncoding RNAs in cancer-associated fibroblasts linked to patient survival in ovarian cancer. Cancer Sci. 2020, 111, 1805–1817. [Google Scholar] [CrossRef]
- Ding, L.; Ren, J.; Zhang, D.; Li, Y.; Huang, X.; Hu, Q.; Wang, H.; Song, Y.; Ni, Y.; Hou, Y. A novel stromal lncRNA signature reprograms fibroblasts to promote the growth of oral squamous cell carcinoma via LncRNA-CAF/interleukin-33. Carcinogenesis 2018, 39, 397–406. [Google Scholar] [CrossRef]
- Zarogoulidis, P.; Petanidis, S.; Domvri, K.; Kioseoglou, E.; Anestakis, D.; Freitag, L.; Zarogoulidis, K.; Hohenforst-Schmidt, W.; Eberhardt, W. Autophagy inhibition upregulates CD4(+) tumor infiltrating lymphocyte expression via miR-155 regulation and TRAIL activation. Mol. Oncol. 2016, 10, 1516–1531. [Google Scholar] [CrossRef] [PubMed]
- Shang, B.; Liu, Y.; Jiang, S.J.; Liu, Y. Prognostic value of tumor-infiltrating FoxP3+ regulatory T cells in cancers: A systematic review and meta-analysis. Sci. Rep. 2015, 5, 15179. [Google Scholar] [CrossRef] [PubMed]
- Knochelmann, H.M.; Dwyer, C.J.; Bailey, S.R.; Amaya, S.M.; Elston, D.M.; Mazza-McCrann, J.M.; Paulos, C.M. When worlds collide: Th17 and Treg cells in cancer and autoimmunity. Cell. Mol. Immunol. 2018, 15, 458–469. [Google Scholar] [CrossRef] [PubMed]
- Kunze-Schumacher, H.; Krueger, A. The Role of MicroRNAs in Development and Function of Regulatory T Cells—Lessons for a Better Understanding of MicroRNA Biology. Front. Immunol. 2020, 11, 2185. [Google Scholar] [CrossRef] [PubMed]
- Brajic, A.; Franckaert, D.; Burton, O.; Bornschein, S.; Calvanese, A.L.; Demeyer, S.; Cools, J.; Dooley, J.; Schlenner, S.; Liston, A. The Long Non-coding RNA Flatr Anticipates Foxp3 Expression in Regulatory T Cells. Front. Immunol. 2018, 9, 1989. [Google Scholar] [CrossRef] [PubMed]
- Pitari, M.R.; Rossi, M.; Amodio, N.; Botta, C.; Morelli, E.; Federico, C.; Gulla, A.; Caracciolo, D.; Di Martino, M.T.; Arbitrio, M.; et al. Inhibition of miR-21 restores RANKL/OPG ratio in multiple myeloma-derived bone marrow stromal cells and impairs the resorbing activity of mature osteoclasts. Oncotarget 2015, 6, 27343–27358. [Google Scholar] [CrossRef]
- Botta, C.; Di Martino, M.T.; Ciliberto, D.; Cuce, M.; Correale, P.; Rossi, M.; Tagliaferri, P.; Tassone, P. A gene expression inflammatory signature specifically predicts multiple myeloma evolution and patients survival. Blood Cancer J. 2016, 6, e511. [Google Scholar] [CrossRef]
- Rossi, M.; Altomare, E.; Botta, C.; Gallo Cantafio, M.E.; Sarvide, S.; Caracciolo, D.; Riillo, C.; Gaspari, M.; Taverna, D.; Conforti, F.; et al. miR-21 antagonism abrogates Th17 tumor promoting functions in multiple myeloma. Leukemia 2020. [Google Scholar] [CrossRef] [PubMed]
- Laviron, M.; Boissonnas, A. Ontogeny of Tumor-Associated Macrophages. Front. Immunol. 2019, 10, 1799. [Google Scholar] [CrossRef]
- Mohapatra, S.; Pioppini, C.; Ozpolat, B.; Calin, G.A. Non-coding RNAs regulation of macrophage polarization in cancer. Mol. Cancer 2021, 20, 24. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Tian, Y.; Guo, F.; Yu, B.; Li, J.; Xu, H.; Su, Z. LincRNA-p21 knockdown reversed tumor-associated macrophages function by promoting MDM2 to antagonize* p53 activation and alleviate breast cancer development. Cancer Immunol. Immunother. CII 2020, 69, 835–846. [Google Scholar] [CrossRef]
- Chen, C.; He, W.; Huang, J.; Wang, B.; Li, H.; Cai, Q.; Su, F.; Bi, J.; Liu, H.; Zhang, B.; et al. LNMAT1 promotes lymphatic metastasis of bladder cancer via CCL2 dependent macrophage recruitment. Nat. Commun. 2018, 9, 3826. [Google Scholar] [CrossRef] [PubMed]
- Frank, A.C.; Ebersberger, S.; Fink, A.F.; Lampe, S.; Weigert, A.; Schmid, T.; Ebersberger, I.; Syed, S.N.; Brune, B. Apoptotic tumor cell-derived microRNA-375 uses CD36 to alter the tumor-associated macrophage phenotype. Nat. Commun. 2019, 10, 1135. [Google Scholar] [CrossRef]
- Sckisel, G.D.; Bouchlaka, M.N.; Monjazeb, A.M.; Crittenden, M.; Curti, B.D.; Wilkins, D.E.; Alderson, K.A.; Sungur, C.M.; Ames, E.; Mirsoian, A.; et al. Out-of-Sequence Signal 3 Paralyzes Primary CD4(+) T-Cell-Dependent Immunity. Immunity 2015, 43, 240–250. [Google Scholar] [CrossRef]
- Qin, S.; Xu, L.; Yi, M.; Yu, S.; Wu, K.; Luo, S. Novel immune checkpoint targets: Moving beyond PD-1 and CTLA-4. Mol. Cancer 2019, 18, 155. [Google Scholar] [CrossRef]
- Wei, J.; Nduom, E.K.; Kong, L.Y.; Hashimoto, Y.; Xu, S.; Gabrusiewicz, K.; Ling, X.; Huang, N.; Qiao, W.; Zhou, S.; et al. MiR-138 exerts anti-glioma efficacy by targeting immune checkpoints. Neuro-Oncology 2016, 18, 639–648. [Google Scholar] [CrossRef]
- Huffaker, T.B.; Lee, S.H.; Tang, W.W.; Wallace, J.A.; Alexander, M.; Runtsch, M.C.; Larsen, D.K.; Thompson, J.; Ramstead, A.G.; Voth, W.P.; et al. Antitumor immunity is defective in T cell-specific microRNA-155-deficient mice and is rescued by immune checkpoint blockade. J. Biol. Chem. 2017, 292, 18530–18541. [Google Scholar] [CrossRef]
- Shin, D.S.; Zaretsky, J.M.; Escuin-Ordinas, H.; Garcia-Diaz, A.; Hu-Lieskovan, S.; Kalbasi, A.; Grasso, C.S.; Hugo, W.; Sandoval, S.; Torrejon, D.Y.; et al. Primary Resistance to PD-1 Blockade Mediated by JAK1/2 Mutations. Cancer Discov. 2017, 7, 188–201. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Hu, Y.; Hu, M.; Li, B. Development of PD-1/PD-L1 Pathway in Tumor Immune Microenvironment and Treatment for Non-Small Cell Lung Cancer. Sci. Rep. 2015, 5, 13110. [Google Scholar] [CrossRef] [PubMed]
- Cortez, M.A.; Ivan, C.; Valdecanas, D.; Wang, X.; Peltier, H.J.; Ye, Y.; Araujo, L.; Carbone, D.P.; Shilo, K.; Giri, D.K.; et al. PDL1 Regulation by p53 via miR-34. J. Natl. Cancer Inst. 2016, 108. [Google Scholar] [CrossRef]
- Mastroianni, J.; Stickel, N.; Andrlova, H.; Hanke, K.; Melchinger, W.; Duquesne, S.; Schmidt, D.; Falk, M.; Andrieux, G.; Pfeifer, D.; et al. miR-146a Controls Immune Response in the Melanoma Microenvironment. Cancer Res. 2019, 79, 183–195. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; He, Y.; Shi, L.; Yang, L.; Wang, J.; Lian, Y.; Fan, C.; Zhang, P.; Guo, C.; Zhang, S.; et al. Co-expression of AFAP1-AS1 and PD-1 predicts poor prognosis in nasopharyngeal carcinoma. Oncotarget 2017, 8, 39001–39011. [Google Scholar] [CrossRef]
- Wolf, Y.; Anderson, A.C.; Kuchroo, V.K. TIM3 comes of age as an inhibitory receptor. Nat. Rev. Immunol. 2020, 20, 173–185. [Google Scholar] [CrossRef]
- Ji, J.; Yin, Y.; Ju, H.; Xu, X.; Liu, W.; Fu, Q.; Hu, J.; Zhang, X.; Sun, B. Long non-coding RNA Lnc-Tim3 exacerbates CD8 T cell exhaustion via binding to Tim-3 and inducing nuclear translocation of Bat3 in HCC. Cell Death Dis. 2018, 9, 478. [Google Scholar] [CrossRef]
- Watanabe, N.; Gavrieli, M.; Sedy, J.R.; Yang, J.; Fallarino, F.; Loftin, S.K.; Hurchla, M.A.; Zimmerman, N.; Sim, J.; Zang, X.; et al. BTLA is a lymphocyte inhibitory receptor with similarities to CTLA-4 and PD-1. Nat. Immunol. 2003, 4, 670–679. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Nie, W.; Jin, Y.; Zhuo, A.; Zang, Y.; Xiu, Q. B and T Lymphocyte Attenuator is a Target of miR-155 during Naive CD4+ T Cell Activation. Iran. J. Immunol. IJI 2016, 13, 89–99. [Google Scholar]
- Sade-Feldman, M.; Jiao, Y.J.; Chen, J.H.; Rooney, M.S.; Barzily-Rokni, M.; Eliane, J.P.; Bjorgaard, S.L.; Hammond, M.R.; Vitzthum, H.; Blackmon, S.M.; et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 2017, 8, 1136. [Google Scholar] [CrossRef]
- Kim, J.M.; Chen, D.S. Immune escape to PD-L1/PD-1 blockade: Seven steps to success (or failure). Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2016, 27, 1492–1504. [Google Scholar] [CrossRef]
- Heward, J.A.; Lindsay, M.A. Long non-coding RNAs in the regulation of the immune response. Trends Immunol. 2014, 35, 408–419. [Google Scholar] [CrossRef] [PubMed]
- Yan, K.; Fu, Y.; Zhu, N.; Wang, Z.; Hong, J.L.; Li, Y.; Li, W.J.; Zhang, H.B.; Song, J.H. Repression of lncRNA NEAT1 enhances the antitumor activity of CD8(+)T cells against hepatocellular carcinoma via regulating miR-155/Tim-3. Int. J. Biochem. Cell Biol. 2019, 110, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.; Ye, Y.; Chan, L.C.; Li, Y.; Liang, K.; Lin, A.; Egranov, S.D.; Zhang, Y.; Xia, W.; Gong, J.; et al. Oncogenic lncRNA downregulates cancer cell antigen presentation and intrinsic tumor suppression. Nat. Immunol. 2019, 20, 835–851. [Google Scholar] [CrossRef]
- Blees, A.; Januliene, D.; Hofmann, T.; Koller, N.; Schmidt, C.; Trowitzsch, S.; Moeller, A.; Tampe, R. Structure of the human MHC-I peptide-loading complex. Nature 2017, 551, 525–528. [Google Scholar] [CrossRef] [PubMed]
- Pistoia, V.; Morandi, F.; Wang, X.; Ferrone, S. Soluble HLA-G: Are they clinically relevant? Semin. Cancer Biol. 2007, 17, 469–479. [Google Scholar] [CrossRef]
- Charpentier, M.; Croyal, M.; Carbonnelle, D.; Fortun, A.; Florenceau, L.; Rabu, C.; Krempf, M.; Labarriere, N.; Lang, F. IRES-dependent translation of the long non coding RNA meloe in melanoma cells produces the most immunogenic MELOE antigens. Oncotarget 2016, 7, 59704–59713. [Google Scholar] [CrossRef]
- Wang, H.; Wang, L.; Zhang, G.; Lu, C.; Chu, H.; Yang, R.; Zhao, G. MALAT1/miR-101-3p/MCL1 axis mediates cisplatin resistance in lung cancer. Oncotarget 2018, 9, 7501–7512. [Google Scholar] [CrossRef] [PubMed]
- Cai, T.; Liu, Y.; Xiao, J. Long noncoding RNA MALAT1 knockdown reverses chemoresistance to temozolomide via promoting microRNA-101 in glioblastoma. Cancer Med. 2018, 7, 1404–1415. [Google Scholar] [CrossRef]
- Gao, Q.; Qiu, S.J.; Fan, J.; Zhou, J.; Wang, X.Y.; Xiao, Y.S.; Xu, Y.; Li, Y.W.; Tang, Z.Y. Intratumoral balance of regulatory and cytotoxic T cells is associated with prognosis of hepatocellular carcinoma after resection. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2007, 25, 2586–2593. [Google Scholar] [CrossRef]
- Shang, W.; Gao, Y.; Tang, Z.; Zhang, Y.; Yang, R. The Pseudogene Olfr29-ps1 Promotes the Suppressive Function and Differentiation of Monocytic MDSCs. Cancer Immunol. Res. 2019, 7, 813–827. [Google Scholar] [CrossRef]
- Gao, Y.; Wang, T.; Li, Y.; Zhang, Y.; Yang, R. Lnc-chop Promotes Immunosuppressive Function of Myeloid-Derived Suppressor Cells in Tumor and Inflammatory Environments. J. Immunol. 2018, 200, 2603–2614. [Google Scholar] [CrossRef]
- Xia, M.; Liu, J.; Liu, S.; Chen, K.; Lin, H.; Jiang, M.; Xu, X.; Xue, Y.; Liu, W.; Gu, Y.; et al. Ash1l and lnc-Smad3 coordinate Smad3 locus accessibility to modulate iTreg polarization and T cell autoimmunity. Nat. Commun. 2017, 8, 15818. [Google Scholar] [CrossRef] [PubMed]
- Xiong, G.; Yang, L.; Chen, Y.; Fan, Z. Linc-POU3F3 promotes cell proliferation in gastric cancer via increasing T-reg distribution. Am. J. Transl. Res. 2015, 7, 2262–2269. [Google Scholar]
- Pegtel, D.M.; Gould, S.J. Exosomes. Annu. Rev. Biochem. 2019, 88, 487–514. [Google Scholar] [CrossRef] [PubMed]
- Fu, Q.; Zhang, Q.; Lou, Y.; Yang, J.; Nie, G.; Chen, Q.; Chen, Y.; Zhang, J.; Wang, J.; Wei, T.; et al. Primary tumor-derived exosomes facilitate metastasis by regulating adhesion of circulating tumor cells via SMAD3 in liver cancer. Oncogene 2018, 37, 6105–6118. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.W.; Wieckowski, E.; Taylor, D.D.; Reichert, T.E.; Watkins, S.; Whiteside, T.L. Fas ligand-positive membranous vesicles isolated from sera of patients with oral cancer induce apoptosis of activated T lymphocytes. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2005, 11, 1010–1020. [Google Scholar]
- Ning, Y.; Shen, K.; Wu, Q.; Sun, X.; Bai, Y.; Xie, Y.; Pan, J.; Qi, C. Tumor exosomes block dendritic cells maturation to decrease the T cell immune response. Immunol. Lett. 2018, 199, 36–43. [Google Scholar] [CrossRef]
- Morrissey, S.M.; Yan, J. Exosomal PD-L1: Roles in Tumor Progression and Immunotherapy. Trends Cancer 2020, 6, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Aung, T.; Chapuy, B.; Vogel, D.; Wenzel, D.; Oppermann, M.; Lahmann, M.; Weinhage, T.; Menck, K.; Hupfeld, T.; Koch, R.; et al. Exosomal evasion of humoral immunotherapy in aggressive B-cell lymphoma modulated by ATP-binding cassette transporter A3. Proc. Natl. Acad. Sci. USA 2011, 108, 15336–15341. [Google Scholar] [CrossRef]
- Liu, J.; Fan, L.; Yu, H.; Zhang, J.; He, Y.; Feng, D.; Wang, F.; Li, X.; Liu, Q.; Li, Y.; et al. Endoplasmic Reticulum Stress Causes Liver Cancer Cells to Release Exosomal miR-23a-3p and Up-regulate Programmed Death Ligand 1 Expression in Macrophages. Hepatology 2019, 70, 241–258. [Google Scholar] [CrossRef] [PubMed]
- Ye, S.B.; Li, Z.L.; Luo, D.H.; Huang, B.J.; Chen, Y.S.; Zhang, X.S.; Cui, J.; Zeng, Y.X.; Li, J. Tumor-derived exosomes promote tumor progression and T-cell dysfunction through the regulation of enriched exosomal microRNAs in human nasopharyngeal carcinoma. Oncotarget 2014, 5, 5439–5452. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Liang, W.; Fu, M.; Huang, Z.H.; Li, X.; Zhang, W.; Zhang, P.; Qian, H.; Jiang, P.C.; Xu, W.R.; et al. Exosomes-mediated transfer of long noncoding RNA ZFAS1 promotes gastric cancer progression. J. Cancer Res. Clin. Oncol. 2017, 143, 991–1004. [Google Scholar] [CrossRef]
- Zhang, R.; Xia, Y.; Wang, Z.; Zheng, J.; Chen, Y.; Li, X.; Wang, Y.; Ming, H. Serum long non coding RNA MALAT-1 protected by exosomes is up-regulated and promotes cell proliferation and migration in non-small cell lung cancer. Biochem. Biophys. Res. Commun. 2017, 490, 406–414. [Google Scholar] [CrossRef]
- Xue, M.; Chen, W.; Xiang, A.; Wang, R.; Chen, H.; Pan, J.; Pang, H.; An, H.; Wang, X.; Hou, H.; et al. Hypoxic exosomes facilitate bladder tumor growth and development through transferring long non-coding RNA-UCA1. Mol. Cancer 2017, 16, 143. [Google Scholar] [CrossRef] [PubMed]
- Ni, C.; Fang, Q.Q.; Chen, W.Z.; Jiang, J.X.; Jiang, Z.; Ye, J.; Zhang, T.; Yang, L.; Meng, F.B.; Xia, W.J.; et al. Breast cancer-derived exosomes transmit lncRNA SNHG16 to induce CD73+gammadelta1 Treg cells. Signal Transduct. Target. Ther. 2020, 5, 41. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.X.; Liu, H.S.; Wang, F.W.; Xiong, L.; Zhou, C.; Hu, T.; He, X.W.; Wu, X.J.; Xie, D.; Wu, X.R.; et al. LncRNA RPPH1 promotes colorectal cancer metastasis by interacting with TUBB3 and by promoting exosomes-mediated macrophage M2 polarization. Cell Death Dis. 2019, 10, 829. [Google Scholar] [CrossRef] [PubMed]
- Lan, J.; Sun, L.; Xu, F.; Liu, L.; Hu, F.; Song, D.; Hou, Z.; Wu, W.; Luo, X.; Wang, J.; et al. M2 Macrophage-Derived Exosomes Promote Cell Migration and Invasion in Colon Cancer. Cancer Res. 2019, 79, 146–158. [Google Scholar] [CrossRef]
- Rupaimoole, R.; Slack, F.J. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017, 16, 203–222. [Google Scholar] [CrossRef]
- Garzon, R.; Marcucci, G.; Croce, C.M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 2010, 9, 775–789. [Google Scholar] [CrossRef]
- Misso, G.; Zarone, M.R.; Grimaldi, A.; Di Martino, M.T.; Lombardi, A.; Kawasaki, H.; Stiuso, P.; Tassone, P.; Tagliaferri, P.; Caraglia, M. Non Coding RNAs: A New Avenue for the Self-Tailoring of Blood Cancer Treatment. Curr. Drug Targets 2017, 18, 35–55. [Google Scholar] [CrossRef] [PubMed]
- Amodio, N.; Di Martino, M.T.; Neri, A.; Tagliaferri, P.; Tassone, P. Non-coding RNA: A novel opportunity for the personalized treatment of multiple myeloma. Expert Opin. Biol. Ther. 2013, 13, S125–S137. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.P.; Berkhout, B. miRNA cassettes in viral vectors: Problems and solutions. Biochim. Biophys. Acta 2011, 1809, 732–745. [Google Scholar] [CrossRef]
- Wang, H.; Jiang, Y.; Peng, H.; Chen, Y.; Zhu, P.; Huang, Y. Recent progress in microRNA delivery for cancer therapy by non-viral synthetic vectors. Adv. Drug Deliv. Rev. 2015, 81, 142–160. [Google Scholar] [CrossRef] [PubMed]
- Hutvagner, G.; Simard, M.J.; Mello, C.C.; Zamore, P.D. Sequence-specific inhibition of small RNA function. PLoS Biol. 2004, 2, E98. [Google Scholar] [CrossRef] [PubMed]
- Vester, B.; Wengel, J. LNA (locked nucleic acid): High-affinity targeting of complementary RNA and DNA. Biochemistry 2004, 43, 13233–13241. [Google Scholar] [CrossRef] [PubMed]
- Leone, E.; Morelli, E.; Di Martino, M.T.; Amodio, N.; Foresta, U.; Gulla, A.; Rossi, M.; Neri, A.; Giordano, A.; Munshi, N.C.; et al. Targeting miR-21 inhibits in vitro and in vivo multiple myeloma cell growth. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2013, 19, 2096–2106. [Google Scholar] [CrossRef]
- Ebert, M.S.; Neilson, J.R.; Sharp, P.A. MicroRNA sponges: Competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 2007, 4, 721–726. [Google Scholar] [CrossRef]
- Choi, W.Y.; Giraldez, A.J.; Schier, A.F. Target protectors reveal dampening and balancing of Nodal agonist and antagonist by miR-430. Science 2007, 318, 271–274. [Google Scholar] [CrossRef]
- Wahlestedt, C.; Salmi, P.; Good, L.; Kela, J.; Johnsson, T.; Hokfelt, T.; Broberger, C.; Porreca, F.; Lai, J.; Ren, K.; et al. Potent and nontoxic antisense oligonucleotides containing locked nucleic acids. Proc. Natl. Acad. Sci. USA 2000, 97, 5633–5638. [Google Scholar] [CrossRef]
- Morelli, E.; Biamonte, L.; Federico, C.; Amodio, N.; Di Martino, M.T.; Gallo Cantafio, M.E.; Manzoni, M.; Scionti, F.; Samur, M.K.; Gulla, A.; et al. Therapeutic vulnerability of multiple myeloma to MIR17PTi, a first-in-class inhibitor of pri-miR-17-92. Blood 2018, 132, 1050–1063. [Google Scholar] [CrossRef]
- Koch, L. Functional genomics: Screening for lncRNA function. Nat. Rev. Genet. 2017, 18, 70. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.J.; Horlbeck, M.A.; Cho, S.W.; Birk, H.S.; Malatesta, M.; He, D.; Attenello, F.J.; Villalta, J.E.; Cho, M.Y.; Chen, Y.; et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science 2017, 355. [Google Scholar] [CrossRef] [PubMed]
- Adriaens, C.; Standaert, L.; Barra, J.; Latil, M.; Verfaillie, A.; Kalev, P.; Boeckx, B.; Wijnhoven, P.W.; Radaelli, E.; Vermi, W.; et al. p53 induces formation of NEAT1 lncRNA-containing paraspeckles that modulate replication stress response and chemosensitivity. Nat. Med. 2016, 22, 861–868. [Google Scholar] [CrossRef] [PubMed]
- Mendell, J.T. Targeting a Long Noncoding RNA in Breast Cancer. N. Engl. J. Med. 2016, 374, 2287–2289. [Google Scholar] [CrossRef]
- Quirico, L.; Orso, F.; Esposito, C.L.; Bertone, S.; Coppo, R.; Conti, L.; Catuogno, S.; Cavallo, F.; de Franciscis, V.; Taverna, D. Axl-148b chimeric aptamers inhibit breast cancer and melanoma progression. Int. J. Biol. Sci. 2020, 16, 1238–1251. [Google Scholar] [CrossRef]
- Kumar Kulabhusan, P.; Hussain, B.; Yuce, M. Current Perspectives on Aptamers as Diagnostic Tools and Therapeutic Agents. Pharmaceutics 2020, 12, 646. [Google Scholar] [CrossRef]
- Catuogno, S.; Di Martino, M.T.; Nuzzo, S.; Esposito, C.L.; Tassone, P.; de Franciscis, V. An Anti-BCMA RNA Aptamer for miRNA Intracellular Delivery. Mol. Ther. Nucleic Acids 2019, 18, 981–990. [Google Scholar] [CrossRef]
- Gu, S.; Jin, L.; Zhang, Y.; Huang, Y.; Zhang, F.; Valdmanis, P.N.; Kay, M.A. The loop position of shRNAs and pre-miRNAs is critical for the accuracy of dicer processing in vivo. Cell 2012, 151, 900–911. [Google Scholar] [CrossRef]
- Watanabe, C.; Cuellar, T.L.; Haley, B. Quantitative evaluation of first, second, and third generation hairpin systems reveals the limit of mammalian vector-based RNAi. RNA Biol. 2016, 13, 25–33. [Google Scholar] [CrossRef]
- Connelly, C.M.; Moon, M.H.; Schneekloth, J.S., Jr. The Emerging Role of RNA as a Therapeutic Target for Small Molecules. Cell Chem. Biol. 2016, 23, 1077–1090. [Google Scholar] [CrossRef] [PubMed]
- Donlic, A.; Hargrove, A.E. Targeting RNA in mammalian systems with small molecules. Wiley Interdiscip. Rev. RNA 2018, 9, e1477. [Google Scholar] [CrossRef] [PubMed]
- Warner, K.D.; Hajdin, C.E.; Weeks, K.M. Principles for targeting RNA with drug-like small molecules. Nat. Rev. Drug Discov. 2018, 17, 547–558. [Google Scholar] [CrossRef] [PubMed]
- Costales, M.G.; Aikawa, H.; Li, Y.; Childs-Disney, J.L.; Abegg, D.; Hoch, D.G.; Pradeep Velagapudi, S.; Nakai, Y.; Khan, T.; Wang, K.W.; et al. Small-molecule targeted recruitment of a nuclease to cleave an oncogenic RNA in a mouse model of metastatic cancer. Proc. Natl. Acad. Sci. USA 2020, 117, 2406–2411. [Google Scholar] [CrossRef]
- Costales, M.G.; Childs-Disney, J.L.; Haniff, H.S.; Disney, M.D. How We Think about Targeting RNA with Small Molecules. J. Med. Chem. 2020, 63, 8880–8900. [Google Scholar] [CrossRef]
- Cruz, J.A.; Westhof, E. The dynamic landscapes of RNA architecture. Cell 2009, 136, 604–609. [Google Scholar] [CrossRef] [PubMed]
- Butcher, S.E.; Pyle, A.M. The molecular interactions that stabilize RNA tertiary structure: RNA motifs, patterns, and networks. Acc. Chem. Res. 2011, 44, 1302–1311. [Google Scholar] [CrossRef]
- Jones, C.P.; Ferre-D’Amare, A.R. RNA quaternary structure and global symmetry. Trends Biochem. Sci. 2015, 40, 211–220. [Google Scholar] [CrossRef]
- Schlick, T. Adventures with RNA graphs. Methods 2018, 143, 16–33. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, C.; Sharma, A.R.; Sharma, G.; Doss, C.G.P.; Lee, S.S. Therapeutic miRNA and siRNA: Moving from Bench to Clinic as Next Generation Medicine. Mol. Ther. Nucleic Acids 2017, 8, 132–143. [Google Scholar] [CrossRef] [PubMed]
- Grillone, K.; Riillo, C.; Scionti, F.; Rocca, R.; Tradigo, G.; Guzzi, P.H.; Alcaro, S.; Di Martino, M.T.; Tagliaferri, P.; Tassone, P. Non-coding RNAs in cancer: Platforms and strategies for investigating the genomic “dark matter”. J. Exp. Clin. Cancer Res. CR 2020, 39, 117. [Google Scholar] [CrossRef]
- Caracciolo, D.; Montesano, M.; Altomare, E.; Scionti, F.; Di Martino, M.T.; Tagliaferri, P.; Tassone, P. The potential role of miRNAs in multiple myeloma therapy. Expert Rev. Hematol. 2018, 11, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Janssen, H.L.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y.; et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Beg, M.S.; Brenner, A.J.; Sachdev, J.; Borad, M.; Kang, Y.K.; Stoudemire, J.; Smith, S.; Bader, A.G.; Kim, S.; Hong, D.S. Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Investig. New Drugs 2017, 35, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Job, C.; De Mercoyrol, L.; Job, D. A slow kinetic transient in RNA synthesis catalysed by wheat-germ RNA polymerase II. Biochem. J. 1988, 253, 281–285. [Google Scholar] [CrossRef]
- Scognamiglio, I.; Di Martino, M.T.; Campani, V.; Virgilio, A.; Galeone, A.; Gulla, A.; Gallo Cantafio, M.E.; Misso, G.; Tagliaferri, P.; Tassone, P.; et al. Transferrin-conjugated SNALPs encapsulating 2’-O-methylated miR-34a for the treatment of multiple myeloma. BioMed Res. Int. 2014, 2014, 217365. [Google Scholar] [CrossRef]
- Misso, G.; Di Martino, M.T.; De Rosa, G.; Farooqi, A.A.; Lombardi, A.; Campani, V.; Zarone, M.R.; Gulla, A.; Tagliaferri, P.; Tassone, P.; et al. Mir-34: A new weapon against cancer? Mol. Ther. Nucleic acids 2014, 3, e194. [Google Scholar] [CrossRef]
- Zarone, M.R.; Misso, G.; Grimaldi, A.; Zappavigna, S.; Russo, M.; Amler, E.; Di Martino, M.T.; Amodio, N.; Tagliaferri, P.; Tassone, P.; et al. Evidence of novel miR-34a-based therapeutic approaches for multiple myeloma treatment. Sci. Rep. 2017, 7, 17949. [Google Scholar] [CrossRef] [PubMed]
- Cosco, D.; Cilurzo, F.; Maiuolo, J.; Federico, C.; Di Martino, M.T.; Cristiano, M.C.; Tassone, P.; Fresta, M.; Paolino, D. Delivery of miR-34a by chitosan/PLGA nanoplexes for the anticancer treatment of multiple myeloma. Sci. Rep. 2015, 5, 17579. [Google Scholar] [CrossRef] [PubMed]
- van Zandwijk, N.; Pavlakis, N.; Kao, S.C.; Linton, A.; Boyer, M.J.; Clarke, S.; Huynh, Y.; Chrzanowska, A.; Fulham, M.J.; Bailey, D.L.; et al. Safety and activity of microRNA-loaded minicells in patients with recurrent malignant pleural mesothelioma: A first-in-man, phase 1, open-label, dose-escalation study. Lancet Oncol. 2017, 18, 1386–1396. [Google Scholar] [CrossRef]
- MacDiarmid, J.A.; Mugridge, N.B.; Weiss, J.C.; Phillips, L.; Burn, A.L.; Paulin, R.P.; Haasdyk, J.E.; Dickson, K.A.; Brahmbhatt, V.N.; Pattison, S.T.; et al. Bacterially derived 400 nm particles for encapsulation and cancer cell targeting of chemotherapeutics. Cancer Cell 2007, 11, 431–445. [Google Scholar] [CrossRef]
- Viteri, S.; Rosell, R. An innovative mesothelioma treatment based on miR-16 mimic loaded EGFR targeted minicells (TargomiRs). Transl. Lung Cancer Res. 2018, 7, S1–S4. [Google Scholar] [CrossRef] [PubMed]
- Seto, A.G.; Beatty, X.; Lynch, J.M.; Hermreck, M.; Tetzlaff, M.; Duvic, M.; Jackson, A.L. Cobomarsen, an oligonucleotide inhibitor of miR-155, co-ordinately regulates multiple survival pathways to reduce cellular proliferation and survival in cutaneous T-cell lymphoma. Br. J. Haematol. 2018, 183, 428–444. [Google Scholar] [CrossRef] [PubMed]
- Franzoni, S.; Morbioli, L.; Turtoro, A.; Solazzo, L.; Greco, A.; Arbitrio, M.; Tagliaferri, P.; Tassone, P.; Di Martino, M.T.; Breda, M. Development and validation of bioanalytical methods for LNA-i-miR-221 quantification in human plasma and urine by LC-MS/MS. J. Pharm. Biomed. Anal. 2020, 188, 113451. [Google Scholar] [CrossRef]
- Franzoni, S.; Vezzelli, A.; Turtoro, A.; Solazzo, L.; Greco, A.; Tassone, P.; Di Martino, M.T.; Breda, M. Development and validation of a bioanalytical method for quantification of LNA-i-miR-221, a 13-mer oligonucleotide, in rat plasma using LC-MS/MS. J. Pharm. Biomed. Anal. 2018, 150, 300–307. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, M.T.; Gulla, A.; Gallo Cantafio, M.E.; Altomare, E.; Amodio, N.; Leone, E.; Morelli, E.; Lio, S.G.; Caracciolo, D.; Rossi, M.; et al. In vitro and in vivo activity of a novel locked nucleic acid (LNA)-inhibitor-miR-221 against multiple myeloma cells. PLoS ONE 2014, 9, e89659. [Google Scholar] [CrossRef]
- Di Martino, M.T.; Rossi, M.; Caracciolo, D.; Gulla, A.; Tagliaferri, P.; Tassone, P. Mir-221/222 are promising targets for innovative anticancer therapy. Expert Opin. Ther. Targets 2016, 20, 1099–1108. [Google Scholar] [CrossRef]
- Sharma, P.; Allison, J.P. Dissecting the mechanisms of immune checkpoint therapy. Nat. Rev. Immunol. 2020, 20, 75–76. [Google Scholar] [CrossRef] [PubMed]
- Fares, C.M.; Van Allen, E.M.; Drake, C.G.; Allison, J.P.; Hu-Lieskovan, S. Mechanisms of Resistance to Immune Checkpoint Blockade: Why Does Checkpoint Inhibitor Immunotherapy Not Work for All Patients? Am. Soc. Clin. Oncol. Educ. Book 2019, 39, 147–164. [Google Scholar] [CrossRef]
- Sudo, K.; Kato, K.; Matsuzaki, J.; Takizawa, S.; Aoki, Y.; Shoji, H.; Iwasa, S.; Honma, Y.; Takashima, A.; Sakamoto, H.; et al. Identification of serum microRNAs predicting the response of esophageal squamous-cell carcinoma to nivolumab. Jpn. J. Clin. Oncol. 2020, 50, 114–121. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Zhang, W.; Li, A.; Chen, Y.; Ou, Q.; He, Z.; Zhang, Y.; Liu, R.; Yao, H.; Song, E. Association of Long Noncoding RNA Biomarkers with Clinical Immune Subtype and Prediction of Immunotherapy Response in Patients with Cancer. JAMA Netw. Open 2020, 3, e202149. [Google Scholar] [CrossRef]
- Xu, J.; Shi, A.; Long, Z.; Xu, L.; Liao, G.; Deng, C.; Yan, M.; Xie, A.; Luo, T.; Huang, J.; et al. Capturing functional long non-coding RNAs through integrating large-scale causal relations from gene perturbation experiments. EBioMedicine 2018, 35, 369–380. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Tao, Z.; Hai, B.; Liang, H.; Shi, Y.; Wang, T.; Song, W.; Chen, Y.; OuYang, J.; Chen, J.; et al. miR-424(322) reverses chemoresistance via T-cell immune response activation by blocking the PD-L1 immune checkpoint. Nat. Commun. 2016, 7, 11406. [Google Scholar] [CrossRef]
- Li, Q.; Johnston, N.; Zheng, X.; Wang, H.; Zhang, X.; Gao, D.; Min, W. miR-28 modulates exhaustive differentiation of T cells through silencing programmed cell death-1 and regulating cytokine secretion. Oncotarget 2016, 7, 53735–53750. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Chen, Z.; Chen, Y.; Wang, X.; Tang, N. MIR155HG is a prognostic biomarker and associated with immune infiltration and immune checkpoint molecules expression in multiple cancers. Cancer Med. 2019, 8, 7161–7173. [Google Scholar] [CrossRef] [PubMed]
- Gallo Cantafio, M.E.; Grillone, K.; Caracciolo, D.; Scionti, F.; Arbitrio, M.; Barbieri, V.; Pensabene, L.; Guzzi, P.H.; Di Martino, M.T. From Single Level Analysis to Multi-Omics Integrative Approaches: A Powerful Strategy towards the Precision Oncology. High-Throughput 2018, 7, 33. [Google Scholar] [CrossRef]
- Guzzi, P.H.; Di Martino, M.T.; Tagliaferri, P.; Tassone, P.; Cannataro, M. Analysis of miRNA, mRNA, and TF interactions through network-based methods. EURASIP J. Bioinform. Syst. Biol. 2015, 2015, 4. [Google Scholar] [CrossRef]
- Di Martino, M.T.; Guzzi, P.H.; Caracciolo, D.; Agnelli, L.; Neri, A.; Walker, B.A.; Morgan, G.J.; Cannataro, M.; Tassone, P.; Tagliaferri, P. Integrated analysis of microRNAs, transcription factors and target genes expression discloses a specific molecular architecture of hyperdiploid multiple myeloma. Oncotarget 2015, 6, 19132–19147. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Martino, M.T.; Riillo, C.; Scionti, F.; Grillone, K.; Polerà, N.; Caracciolo, D.; Arbitrio, M.; Tagliaferri, P.; Tassone, P. miRNAs and lncRNAs as Novel Therapeutic Targets to Improve Cancer Immunotherapy. Cancers 2021, 13, 1587. https://doi.org/10.3390/cancers13071587
Di Martino MT, Riillo C, Scionti F, Grillone K, Polerà N, Caracciolo D, Arbitrio M, Tagliaferri P, Tassone P. miRNAs and lncRNAs as Novel Therapeutic Targets to Improve Cancer Immunotherapy. Cancers. 2021; 13(7):1587. https://doi.org/10.3390/cancers13071587
Chicago/Turabian StyleDi Martino, Maria Teresa, Caterina Riillo, Francesca Scionti, Katia Grillone, Nicoletta Polerà, Daniele Caracciolo, Mariamena Arbitrio, Pierosandro Tagliaferri, and Pierfrancesco Tassone. 2021. "miRNAs and lncRNAs as Novel Therapeutic Targets to Improve Cancer Immunotherapy" Cancers 13, no. 7: 1587. https://doi.org/10.3390/cancers13071587
APA StyleDi Martino, M. T., Riillo, C., Scionti, F., Grillone, K., Polerà, N., Caracciolo, D., Arbitrio, M., Tagliaferri, P., & Tassone, P. (2021). miRNAs and lncRNAs as Novel Therapeutic Targets to Improve Cancer Immunotherapy. Cancers, 13(7), 1587. https://doi.org/10.3390/cancers13071587










