Simple Summary
Breast cancer is the most common cancer in women worldwide. Although many studies have aimed to understand the genetic basis of breast cancer, leading to increasingly accurate diagnoses, only a few molecular biomarkers are used in clinical practice to predict response to therapy. Current studies aim to develop more personalized therapies to decrease the adverse effects of chemotherapy. Personalized medicine not only requires clinical, but also molecular characterization of tumors, which allows the use of more effective drugs for each patient. The aim of this study was to identify potential molecular biomarkers that can predict the response to therapy after neoadjuvant chemotherapy in patients with breast cancer. In this review, we summarize genomic, transcriptomic, and proteomic biomarkers that can help predict the response to therapy.
Abstract
Neoadjuvant chemotherapy (NAC) is often used to treat locally advanced disease for tumor downstaging, thus improving the chances of breast-conserving surgery. From the NAC response, it is possible to obtain prognostic information as patients may reach a pathological complete response (pCR). Those who do might have significant advantages in terms of survival rates. Breast cancer (BC) is a heterogeneous disease that requires personalized treatment strategies. The development of targeted therapies depends on identifying biomarkers that can be used to assess treatment efficacy as well as the discovery of new and more accurate therapeutic agents. With the development of new “OMICS” technologies, i.e., genomics, transcriptomics, and proteomics, among others, the discovery of new biomarkers is increasingly being used in the context of clinical practice, bringing us closer to personalized management of BC treatment. The aim of this review is to compile the main biomarkers that predict pCR in BC after NAC.
1. Introduction
Breast cancer (BC) is the most commonly diagnosed malignancy and is responsible for the highest number of deaths among women worldwide [1]. Furthermore, BC is heterogeneous and presents different morphological and biological characteristics, thus leading to different clinical behaviors and responses [2]. Therefore, BCs are classified according to their characteristics, histological type, and expression of tumor markers, which develop from genetic and molecular changes in breast tissue cells [3,4].
Neoadjuvant chemotherapy (NAC) is an important treatment strategy for BC patients, with the aim of reducing staging and monitoring response to treatment for prognostic purposes, thereby increasing pathological complete response rate (pCR) [5]. pCR is an important long-term clinical outcome for patients with BC, as patients who achieve pCR with neoadjuvant therapy tend to have better disease-free survival (DFS) and overall survival (OS) compared with patients with residual invasive disease [6,7].
pCR is defined as the complete disappearance of all invasive breast carcinoma cells and axillary lymph nodes (ypT0/ypN0), and is determined pathologically in the resected tissue after NAC [8]. Predicting which patients will achieve pCR or have residual disease (RD) may help suggest and plan a specific treatment according to patient’s characteristics, thus enabling personalized therapy. Clinical staging, axillary lymph node status, and human epidermal growth factor receptor-2 (HER2) positivity are associated with cancer recurrence rates after NAC [9]. RD is defined by the presence of breast cancer cells in the tumor bed and/or positive lymph nodes after surgical removal. Patients with documented RD are usually associated with a worse prognosis than those who achieve pCR, although RD can have a heterogeneous prognosis in each patient [10,11]. Studies have attempted to identify molecular biomarkers that could monitor patients with early pCR and avoid overtreatment in this population. However, these markers require larger studies with long-term follow-up, and for this reason, they currently lack clinical validation [12,13].
The identification of efficient molecular markers that can predict sensitivity to chemotherapy, demonstrate higher rates of pCR, and identify patients that can benefit from NAC in clinical practice has been a challenge in many recent studies. However, molecular markers can be effective in avoiding unnecessary treatments and associated toxicities for BC patients that do not respond to NAC [14]. Since histologically similar tumors may demonstrate different prognoses and responses to therapy, some molecular subtypes of BC can have high rates of pCR to NAC, while others may not have the same benefits from being exposed to the same treatment. Therefore, there is a need for predictive biomarkers to select patients who will not benefit from NAC in order to offer new therapeutic approaches to these patients [15]. NAC offers an opportunity to identify biomarkers that are predictive of the response to such treatment in patients with BC.
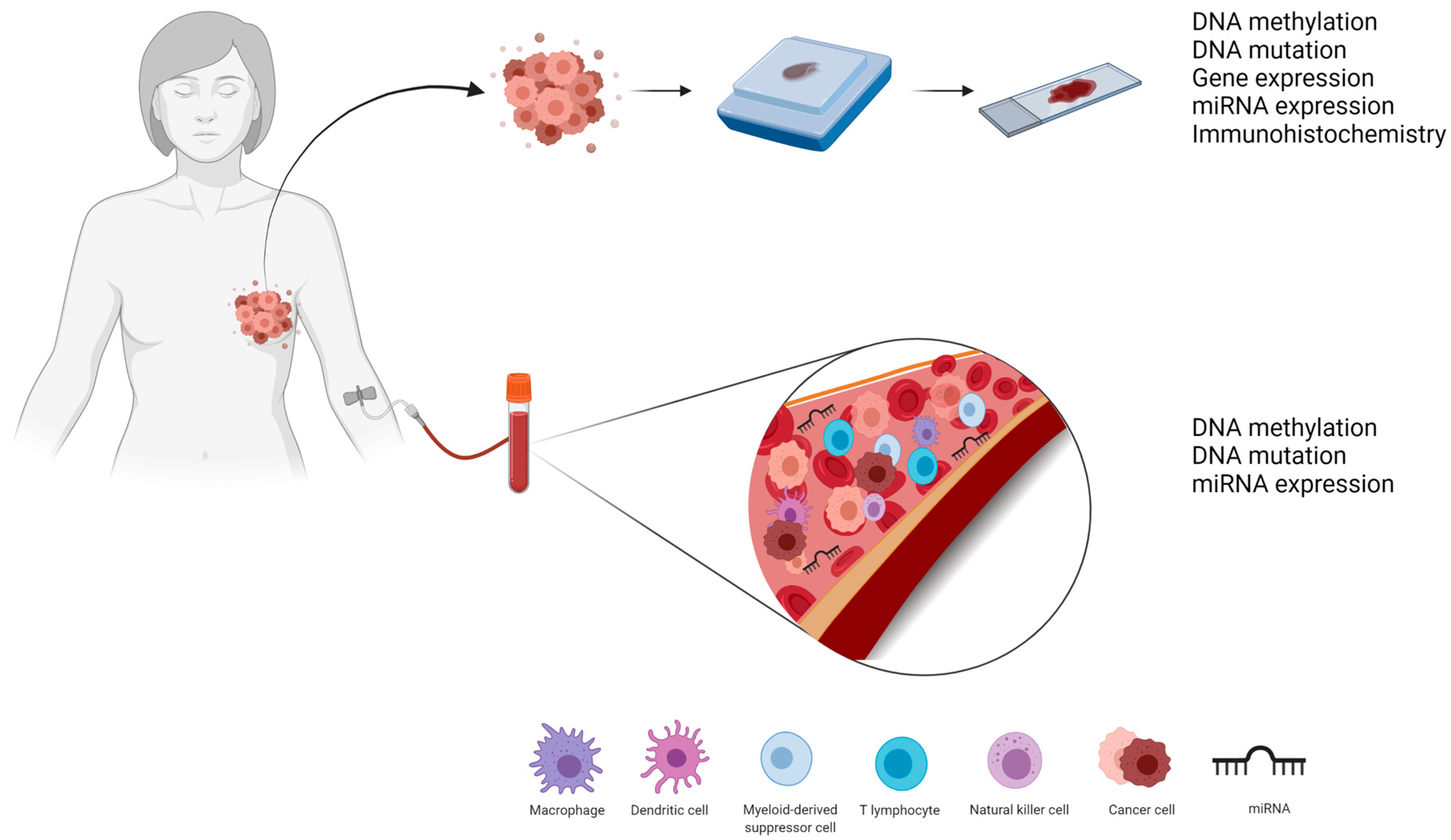
Biomarkers that use “omics” technologies, i.e., genomics, transcriptomics, and proteomics, in BC research have gained recognition in the scientific community. These omics analyses involve the identification of biomolecules responsible for each step of cell function control from DNA replication (genomics markers) to transcriptional events and post-transcriptional regulation (transcriptomic markers) to protein translation (proteomic markers). These markers can be identified not only in tumor tissues but also by liquid biopsy (Figure 1) [16,17], which could assist in the development of new drugs and in the identification and monitoring of patients who will respond and benefit from this treatment [18,19].

Figure 1.
Tumor and liquid biopsies can help identify pCR biomarkers as they can provide information at the genomic (DNA methylation and DNA mutation), transcriptomic (mRNA and miRNA expression), and proteomic (immunohistochemistry) levels.
Despite the importance of pCR markers for therapy selection, we identified only a few studies that explored this potential and demonstrated that many molecules are differentially expressed at the genomic, transcriptomic, and proteomic levels, and can be used as effective biomarkers of NAC response.
2. Genomic: DNA as Biomarkers of NAC Response in BC Patients
The potential use of genomic markers for diagnosis and predicting prognosis, and response to treatment has been increasingly studied. DNA mutations, DNA methylation, and circulating tumor DNA (ctDNA) are among the main classes of genomic biomarkers. These molecules can be identified in tumor tissues or biofluids, such as blood, serum, or plasma samples.
2.1. DNA Mutation
Mutations in genes such as oncogenes or tumor suppressor genes have been widely studied because of their potential as predictors of prognosis. Furthermore, it is possible to predict their impact on tumor development and progression. Table 1 summarizes the mutations that predict pCR in patients with BC. Dysfunctions in DNA repair pathways can occur because of genetic mutations that compromise genomic integrity. Genomic instability is an important hallmark of carcinogenesis, and cellular machinery plays an important role in maintaining this stability [20]. For example, homologous recombination is necessary for repairing DNA double-strand breaks. Some genes have already been described in the literature, such as BRCA1/2, which encodes proteins necessary for homologous recombination repairing [13].

Table 1.
DNA mutations as biomarkers of pCR or non-pCR after NAC in BC patients.
Several studies have evaluated many gene mutations to identify sensitive and specific biomarkers that can predict patient response to different treatments and assess their impact on development and tumor progression. In BC settings, germline mutations of BRCA1/2 genes are frequent in patients with the triple negative breast cancer (TNBC) molecular subtype. A study evaluated the pCR rate in TNBC patients, with or without a mutation in one of these genes, who received NAC with epirubicin, cyclophosphamide, docetaxel, and bevacizumab. The findings showed that therapy with bevacizumab promoted pCR in patients with BRCA1/2 mutations [21].
Another widely studied mutation is the PIK3CA gene mutation. This gene encodes the p110α catalytic subunit of the phosphatidylinositol 3-kinase signaling pathway, one of the intracellular pathways often related to BC [22]. Using whole exome sequencing (WES), Shi et al. identified a correlation between the PIK3CA gene mutation and resistance to trastuzumab treatment associated with non-pCR. Furthermore, these findings indicate that treatment with lapatinib provided better outcomes in patients with HER2-positive molecular phenotype BC who had PIK3CA driver mutation [23].
A second study also evaluated pCR based on the PIK3CA mutation in HER2-positive or TNBC patients who received NAC with paclitaxel and doxorubicin. Patients with TNBC also received bevacizumab and carboplatin. However, in this study, the researchers evaluated only exons 9 and 20, and not the complete gene, through DNA sequencing. By performing a multivariate analysis, it was possible to associate the PIK3CA H1047R hotspot mutation with non-pCR in TNBC patients [22].
A study evaluated pCR in patients with early BC who received NAC with capecitabine, docetaxel (for HER2-negative patients), and trastuzumab (HER2-positive patients). This analysis was performed by identifying mutations in the TP53 gene, a tumor suppressor that is involved in the regulation of cell proliferation, survival, and genomic integrity in BC. The authors showed that the most frequent mutation was missense and that patients with the mutation had higher pCR rates, indicating the effectiveness of NAC [24].
In the context of identifying potential predictive genomic biomarkers, the expression of the TOP2A gene has been studied using the fluorescent in situ hybridization (FISH) technique. TOP2A encodes topoisomerase IIα, a key enzyme in DNA replication, and one of the molecular targets of anthracyclines, and is mutated in a significant percentage of HER2-overexpressing BC patients. Thus, several studies have evaluated the relationship between this gene and its ability to predict resistance to anthracyclines in BC. Desmedt et al. evaluated patients with estrogen receptor-negative (ER-negative) and HER2-positive molecular phenotypes who were treated with anthracycline (epirubicin) and taxane. The findings showed that TOP2A amplification was correlated with pCR in patients who received anthracycline alone [25].
Tibau et al. also evaluated the relationship between TOP2A and anthracycline resistance. Because this gene is located on chromosome 17, close to the centromere, the authors also evaluated the duplication of the centromere on this chromosome (CEP17), which may be a biomarker for genomic instability and DNA repair dysfunction. In this study, HER2-positive BC patients who underwent NAC with either fluorouracil, or taxanes combined with anthracyclines and cyclophosphamide, and were treated before trastuzumab approval, were evaluated. Multivariate analysis showed that the presence of CEP17 duplication, as well as TOP2A amplification, showed a high percentage of pCR [26].
Besides point mutations, it is also possible to use biomarkers based on DNA expression profiles, such as the homologous recombination deficiency (HRD) score, which includes telomeric allelic imbalance (TAI, defined as the number of regions with allelic imbalance that extend to one of the subtelomeres), large-scale state transitions (LST, defined as the number of chromosomal breaks between adjacent regions), and loss of heterozygosity (LOH, defined as allele-specific copy number for each sub-chromosomal region). Kaklamani et al. investigated HRD associated with mutation status for BRCA1/2 genes and BRCA1 promoter methylation through DNA sequencing. An important finding was that patients who had a BRCA1/2 germline mutation or methylation had an HRD score above the threshold. In this study, which aimed to evaluate these profiles based on DNA and protein expression as potential predictors of therapeutic response, it was possible to predict pCR to treatment with carboplatin and eribulin in patients with early stage TNBC [27].
2.2. DNA Methylation
DNA methylation, a type of epigenetic alteration that is involved in carcinogenesis, consists of the addition of a methyl group in the promoter region of a gene related to the tumor development process. These changes occur at high rates and contribute to the loss of epigenetic regulation, which can be crucial in early stages of carcinogenesis. The most common alteration is DNA hypermethylation of CpG dinucleotide islands, which increases the probability of sporadic mutation by deamination of 5-methylcytosine to thymine, resulting in point mutations and abnormal protein translation [28,29]. Some studies have shown that epigenetic mechanisms such as DNA methylation occur more often in patients who had pCR compared to women who had RD. Accordingly, Table 2 summarizes the studies that demonstrated whether DNA methylation could predict pCR in BC patients.

Table 2.
DNA methylation as a biomarker of pCR or non-pCR after NAC in BC patients.
Almeida et al. investigated genome-wide DNA methylation patterns in BC patients and correlated the variations with gene expression data from The Cancer Genome Atlas (TCGA) and Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) databases. Their study suggests that both hypermethylation and hypomethylation of CpG may be crucial events in BC and identifies three new diagnostic and prognostic biomarker candidates for DNA methylation [29].
Studies have demonstrated the consequences of DNA methylation in the cancer research landscape. Fujii et al. identified methylation of the promoter CpG island of the HSD17B4 gene through genome-wide methylation analysis in tumor samples from patients with HER2-positive BC. Using DNA sequencing and performing a multivariate analysis, it was possible to predict pCR for treatment with trastuzumab, paclitaxel, or anthracycline through methylation of that gene that encodes the 17β-hydroxysteroid dehydrogenase type 4 enzyme [30].
Another study evaluated the response to NAC using DNA methylation profiles. Connolly et al. used a methylation panel, which investigated 10 genes (HIST1H3C, AKR1B1, GPX7, HOXB4, TMEFF2, RASGRF2, COL6A2, ARHGEF7, TM6SF1, and RASSF1A), to evaluate tumor tissue and serum samples from patients with HER2-negative BC. These genes were selected from previous studies [32,33] that identified their methylation in breast tumors at all stages and in the serum of patients with metastatic BC. By performing exploratory analyses with univariate and multivariate logistic regression models, it was possible to associate the high cumulative methylation index with non-pCR in patients who underwent NAC with carboplatin and nab-paclitaxel or vorinostat [31].
2.3. Circulating Tumor DNA
The detection of ctDNA through liquid biopsy has already been used in clinical practice to monitor cancer. However, studies have been performed to improve the technique and identify precise biomarkers. These molecules have been studied because they can be obtained using a minimally invasive approach as ctDNA is evaluated through plasma samples.
Malignant cells release cell-free DNA molecules into the bloodstream, thus allowing tumor progression. Studies have evaluated the potential of these molecules as prognostic factors and their response to treatment biomarkers. The first published study of ctDNA and BC was published more than a decade ago [34], and since then, several studies have examined techniques used to identify highly sensitive and specific biomarkers [35].
A recent study evaluated the efficacy of NAC with paclitaxel and/or anthracycline in BC patients using ctDNA expression through WES. This study identified that high expression of ctDNA was associated with non-pCR, thus suggesting it to be a considerable biomarker of early response prediction in the neoadjuvant setting in different molecular subtypes [36]. Moreover, this dynamic monitoring during treatment can facilitate the evaluation of new agents, providing greater sensitivity to the effectiveness of the treatment.
3. Transcriptomic: mRNA and miRNAs as Biomarkers of NAC Response in BC Patients
Studies have demonstrated that transcriptomic biomarkers can predict a patient’s response to NAC. The main classes of these biomarkers are the expression of genes and the miRNAs, which may originate from tumor or liquid biopsy.
3.1. Gene Expression Panels
With the discovery of microarray technology, it is possible to analyze the expression of several genes simultaneously. Thus, Perou et al. were able to use gene expression assays to identify five molecular subtypes in BC: (1) baseline as, Erb−B2+, (2) normal breast, (3) luminal A, (4) luminal B, and (5) luminal C [37]. Later, this classification underwent several modifications, and it was widely accepted as a method to identify the prognostic significance of BC, in which the ER+ HER2− and luminal A tumors demonstrated a better prognosis, while the baseline and non-baseline triple-negative tumors had worse prognosis [38].
Over the past few years and with the advancement of scientific research, it has been possible to identify several biomarkers for BC, and many gene expression signatures have become commercially available as prognostic tools for this neoplasm. Oncotype DX gene panels (RS; Genomic Health, Redwood City, CA, USA) [39], Mammaprint [40] (Agendia, Amsterdam, the Netherlands), EndoPredict (EP; Myriad Genetics, Cologne, Germany) [41], Prediction Analysis of Microarray 50 (PAM50) Risk of Recurrence, Prossigna Kit (Prosigna; NanoString Technologies, Seattle, WA, USA) [42], and breast cancer index (BCI; Biotheranostics, San Diego, CA, USA) [43] are some examples of panels that explore and derive conclusions about tumor recurrence and relapse [44,45]. The main multigene expression signatures (MES) used as biomarkers for pCR in BC that are currently available on the market are shown in Table 3.

Table 3.
Commercial panels for prognostic evaluation of BC patients using mRNA gene expression.
Indeed, the biomarkers that predict patients’ response to NAC offer an opportunity for personalized service, better response rates to therapy, reduced adverse effects, and cost savings for the public health system by avoiding overtreatment in patients who will have non-pCR [45]. Because there are now different molecular signatures, some studies have pointed out that certain commercial gene expression panels may be useful in stratifying patients who will have pCR. Currently, the most commonly used panel is the Oncotype DX, which consists of a panel that assesses the expression of 21 genes in tumor tissue. The test result, considered as Recurrence Score, is able to provide information on the probability of tumor recurrence, as well as the chance of the patient presenting pCR in the face of NAC administration [46].
3.2. Differentially Expressed miRNA
From genome-wide miRNA expression analysis, it was possible to identify several miRNAs that were differentially expressed in BC tissue [47]. Since then, many studies have reported the importance of this molecule in different tumor phenotypes [48]. One recent approach was the ability of miRNA expression profiles to classify breast tumors according to histopathological variables, which are currently used to indicate responsiveness to neoadjuvant therapy [49,50,51,52,53,54,55]. As a result, these molecules are highlighted as potential predictive biomarkers that can allow the individualization of BC treatment and a better selection of patients who could respond to NAC.
Evidence has shown that miRNAs can be differentially expressed in the bloodstream of patients with pCR to NAC compared with patients with RD. Circulating miRNAs (ct-miRNAs) originate from the tumor tissue and migrate into the bloodstream, which makes it possible to identify the specific biological characteristics of the tumor [56,57]. With the advancement of technology in recent years, the detection of ct-miRNAs from body fluids has been made possible, and the evaluation of ct-miRNA expression has shown that it has great potential as a biomarker for early detection, drug resistance, tumor recurrence, and clinical outcome prediction of patients on cancer therapy [58], especially for monitoring of BC patient treatment [59].
Seven articles were identified in this context. These studies evaluated differential miRNA expression and investigated the association between miRNAs and pCR or non-pCR in BC patients who underwent NAC (Table 4). The results obtained from the high-throughput miRNA profile assessment identified four significant signatures between HER2-positive patient groups that received lapatinib at T0 and T1 and the group that received lapatinib and trastuzumab at T1, demonstrating promising evidence for future analyses using ct-miRNAs to assess the response to anti-HER2 agents. However, the authors stated that confirmatory studies in independent case series are needed to validate and evaluate the generalization of these ct-miRNA signatures. The data presented in this study may have direct implications for future clinical trials, as miRNA analyzed in plasma can be a promising strategy for predicting response to trastuzumab as monotherapy and can be used to guide de-escalation therapy [52]. Cosimo et al. identified increased levels of ct-miRNAs, from which ct-miR-148a-3p and ct-miR-374a-5p were significantly associated with pCR after NAC in patients with HER2-positive BC [49]. Using univariate and multivariate models, it was verified that miR-155 and miR-301 indicated a better pCR. This study evaluated the expression of miRNAs from the isolation of total plasma exosomes from patients with TNBC and HER2-positive patients before NAC. It was possible to identify a network of deregulated exosomal miRNAs with specific expression patterns in exosomes of HER2-positive and TNBC patients that are also associated with clinicopathological parameters and pCR within each molecular subtype of BC [51].

Table 4.
miRNAs as biomarkers of pCR or non-pCR after NAC in BC patients.
García-García et al. also demonstrated that miR-145-5p low expression was associated with high pCR rates in patients with TNBC who received cisplatin/doxorubicin-based neoadjuvant treatment. In contrast, patients with higher levels of miR-145-5p expression did not respond to chemotherapy regimens and had worse outcomes [53]. In addition, this study suggested that miR-145-5p could be a predictor of pCR. Our hypothesis is that patients with a worse prognosis may respond better because of the proliferative index. García-García et al. performed functional in vitro assays and verified that miR-145 mimics were able to decrease cell line proliferation of TNBC (MDA-MB-231), and a high expression level of miR-145-5p was identified in patients with non-pCR after NAC regimen [53]. On the other hand, another study demonstrated that the expression of ct-miR-21 could accurately distinguish clinical responders from non-responders, but it was not possible to distinguish those with pCR from those with RD [50].
Bearing in mind that ct-miRNAs, acting as potential predictive and prognostic biomarkers, may be able to identify patients who will have pCR allows us to individualize the treatment of BC and better select patients for NAC. Although rapid and continuous advances are being made in regard to the use of differentially expressed miRNAs as biomarkers of pCR prediction, this area of research still has many obstacles to overcome before its implementation in the management of BC patients’ clinical practice. To date, few studies have evaluated pCR after NAC treatment. Therefore, to validate these miRNA as effective biomarkers for the identification of patients who will achieve pCR, large clinical trials are needed to support these preliminary findings. Current obstacles to overcome include identifying methods for evaluating miRNA expression profiles that are specific, sensitive, and highly accurate at low cost. Additionally, research on the discovery of new biomarkers and more accessible technologies is essential, and the identification of a biomarker that could predict or potentially monitor the tumor’s response to NAC could revolutionize the way chemotherapeutic drugs are administered, bringing us closer to personalized management of BC.
4. Proteomic: Proteins as Biomarkers of NAC Response in BC Patients
Protein biomarkers are widely used in clinical practice to assess the prognosis of patients. Different studies have reported differential expression of proteins as biomarkers of pCR in molecular subtypes of BC (Table 5). Currently, many protein biomarkers have been identified in BC tissues and/or from the tumor-infiltrating immune system [60]. The advent of protein analysis in BC made it possible to obtain prognostic markers [61] and identify molecular subtypes [62] using immunohistochemistry (IHC). These biomarkers are currently available and can guide the clinical management of targeted therapy. IHC is a quick and inexpensive assay that provides important diagnostic and prognostic information [63,64].

Table 5.
Proteins as biomarkers of pCR or non-pCR after NAC in BC patients.
Proliferation markers can predict systemic responses to NAC in some molecular subtypes of BC. Ki-67 is a non-histone nuclear protein expressed during all cell cycle phases, except the G0 phase. Therefore, Ki-67 is used as a marker for tumor proliferation [71]. This marker was identified by IHC analysis, where the levels of Ki-67 expression were associated with the percentage of tumor cells stained positively among the total number of malignant cells evaluated [72].
The use of Ki-67 has been reported in previous BC studies, which demonstrated that this protein expression can predict the response to NAC [73,74]. Yoshioka et al. demonstrated that a high Ki-67 expression in tumors before treatment was associated with higher rates of pCR, and a high Ki-67 expression in post-treatment tumors was strongly correlated with low DFS and OS, regardless of subtype [65].
Another protein that is related to pCR is carbonic anhydrase IX (CAIX), which is a transmembrane protein and one of the only two isoenzymes of carbonic anhydrase associated with tumors that may be involved in cell proliferation and transformation [66]. Alves et al. first described CAIX expression as a predictor of pCR and its association with DFS and OS in patients with locally advanced BC treated with NAC using doxorubicin, cyclophosphamide, and paclitaxel [66].
Studies have shown that in patients with TNBC, the immune system can influence the chemotherapy response. One example is programmed cell death-ligand 1 (PD-L1), a transmembrane protein expressed in a variety of cells, including epithelial cells, vascular endothelial cells, macrophages, myeloid dendritic cells, and B cells [75]. Cerbelli et al. investigated the role of PD-L1 expression in predicting the pathological response to NAC in TNBC. Before NAC, biopsies showed that PD-L1 in ≥25% of tumor cells predicted pCR in TNBC. A possible explanation for these findings is that PD-L1 expression may be associated with a subpopulation of TNBC with more aggressive behavior, with a probability of responding to chemotherapy [67].
Similarly, FK506 binding protein 12 (FKBP12) is a cytoplasmic protein expressed with multiple functions in the transduction of cell signaling [76] and has been reported as a predictive biomarker for the effectiveness of anthracycline-based chemotherapy in BC. Xing et al. demonstrated that the loss of FKBP12 was specifically correlated with poor prognosis and increased resistance to anthracycline-based chemotherapy. Patients with low FKBP12 expression had a significantly lower rate of pCR [68].
A study with a female Japanese population with locally advanced BC showed that lower levels of MGMT protein expression were associated with higher pCR rates when compared with women with normal expression levels of MGMT protein [69]. MGMT is a DNA repair protein that removes alkylating agents from DNA [77].
Furthermore, annexins are a large multifunctional family of phospholipid-binding proteins regulated by Ca2+ [78]. Annexin A1 (ANXA1) is linked to phospholipids involved in inflammation, immune response, and reactivity of mast cells and is associated with the aggressive phenotype of TNBC [79]. Annexin A2 (ANXA2) is a calcium-binding cytoskeleton protein located on the extracellular surface of endothelial cells and in various types of tumor cells [80]. It has been shown that the expression of ANXA2 in breast tumors can be a biomarker for predicting BC outcome in high-risk groups [81]. Chuthapisith reported that the proteins ANXA1 and ANXA2 are predictors of pCR, as it was demonstrated that the presence of ANXA2 in conjunction with ANXA1 could be a potential marker of non-pCR in BC [70].
5. Final Considerations
Our review found several studies that evaluated potential molecular markers as predictors of pCR. The main markers are gene mutations, DNA methylation, and the expression of miRNAs and proteins. Reaching ypT0/ypN0 is strongly associated with a great impact on improving overall and progression-free survival in BC patients, as it is independent of nodal status, and apparently of greater benefit in patients with TNBC [8,82]. Predicting which patients will benefit from NAC is one of the main reasons for researching non-invasive response markers. Despite the existence of various studies to identify biomarkers associated with pathological response, there is still no ideal molecular marker that can be used in clinical practice to distinguish resistant and sensitive patients and, thus, help define possible changes in treatment for patients without pCR [83]. The aim of this review was to compile the main genomic, transcriptomic, and proteomic signatures that were tested for pCR.
The search for biomarkers has been the target of many studies, as they can be used for diagnosis, prognosis, and drug selection in BC [84,85]. There are biomarkers that have already been validated by clinical trials and, therefore, may be available to assist in clinical practice, and promising biomarkers that still need to be better explored and validated [83].
The evolution of methods, such as artificial intelligence-powered imaging analysis, use of high-performance molecular profiling, and computational tools allow the implementation of personalized medicine and aid in prognosis and risk stratification. These methods can also be used for scaling or avoiding therapies, and predicting response to treatment [86,87].
The identification of clinically useful biomarkers is challenging due to several limitations, including tumor heterogeneity, since a single biomarker may not have sufficient sensitivity and specificity to predict response to therapy and tumor behavior [88]. The lack of standardized protocols and precise cutoff values, the need for a complete assessment of sensitivity, specificity and reproducibility are also obstacles for the validation of these biomarkers in clinical practice [89]. The development of affordable biomarkers also poses a challenge due to the high cost needed for use in clinical practice [90]. Therefore, defining which biomarkers that might be clinically applicable to discern between responders and non-responders to NAC is a challenge.
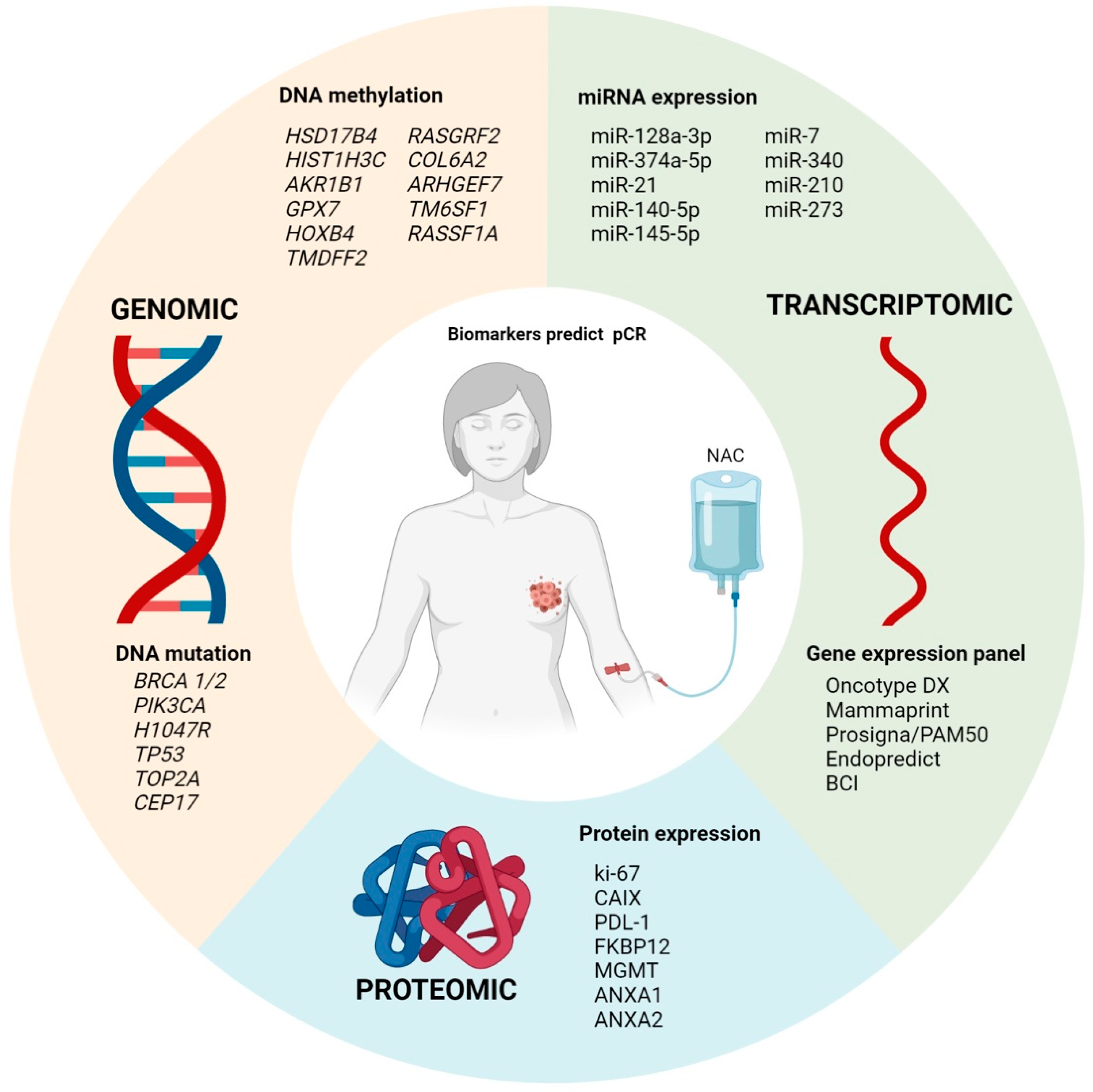
In this review, we highlighted several studies that used the omics approach to identify new biomarkers as potential predictors for target therapy (Figure 2). The use of these biomarkers, although scarcely used in clinical practice, has been shown to be sufficiently accurate to distinguish patients who will achieve pCR. However, further studies with larger cohorts and clinically controlled and randomized groups need to be conducted to validate these findings.

Figure 2.
Summary of the main pCR markers at genomic, transcriptomic, and proteomic levels.
Current research has not yet identified any predictive molecular biomarkers for pCR in BC patients, which are sufficiently robust and can be used in the clinical management of patients with pCR or RD. Hence, it is essential to identify genomic, transcriptomic, and proteomic markers that are specific, sensitive, and accurate. In this review, we demonstrated that different biomarkers may be important in predicting a patient’s response to distinct treatments. Thus, it can minimize the adverse effects and toxicity commonly caused by these drugs and anticipate cases in which patients will not benefit from certain drugs.
Author Contributions
A.J.A.d.F.: Developed and led the overall study, conducted the data reviews and analysis, and wrote the original draft preparation. R.L.C.: Conducted the data reviews and analysis and critically read the manuscript. M.B.V.: Participated in preparing the manuscript and critically reading the manuscript. C.M.T.H.F.: Participated in preparing the manuscript and critically reading the manuscript. V.D.d.S.: Participated in preparing the manuscript and critically reading the manuscript. C.d.P.S.: Participated in designing and developing the study and critically reading the manuscript. M.M.C.M.: Conceived the study, provided advice during study development, and prepared the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Departamento de Ciência e Tecnologia–DECIT, Ministério da Saúde (grant #879848/2018).
Acknowledgments
The authors would like to thank the events sector, especially Caio Fernando de Oliveira. In addition, we would like to thank the sources of funding, Departamento de Ciência e Tecnologia–DECIT, Ministério da Saúde (grant #879848/2018), and the Research Incentive Program of Barretos Cancer Hospital (PAIP).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harbeck, N.; Penault-Llorca, F.; Cortes, J.; Gnant, M.; Houssami, N.; Poortmans, P.; Ruddy, K.; Tsang, J.; Cardoso, F. Breast Cancer. Nat. Rev. Dis. Primers 2019, 5, 66. [Google Scholar] [CrossRef] [PubMed]
- Li, C.I.; Uribe, D.J.; Daling, J.R. Clinical Characteristics of Different Histologic Types of Breast Cancer. Br. J. Cancer 2005, 93, 1046–1052. [Google Scholar] [CrossRef]
- Prat, A.; Perou, C.M. Deconstructing the Molecular Portraits of Breast Cancer. Mol. Oncol. 2011, 5, 5–23. [Google Scholar] [CrossRef]
- Early Breast Cancer Trialists’ Collaborative Group (EBCTCG). Long-Term Outcomes for Neoadjuvant versus Adjuvant Chemotherapy in Early Breast Cancer: Meta-Analysis of Individual Patient Data from Ten Randomised Trials. Lancet Oncol. 2018, 19, 27–39. [Google Scholar] [CrossRef] [Green Version]
- Kong, X.; Moran, M.S.; Zhang, N.; Haffty, B.; Yang, Q. Meta-Analysis Confirms Achieving Pathological Complete Response after Neoadjuvant Chemotherapy Predicts Favourable Prognosis for Breast Cancer Patients. Eur. J. Cancer 2011, 47, 2084–2090. [Google Scholar] [CrossRef]
- I-SPY2 Trial Consortium. Association of Event-Free and Distant Recurrence—Free Survival with Individual-Level Pathologic Complete Response in Neoadjuvant Treatment of Stages 2 and 3 Breast Cancer: Three-Year Follow-up Analysis for the I-SPY2 Adaptively Randomized Clinical Trial. JAMA Oncol. 2020, 6, 1355–1362. [Google Scholar] [CrossRef]
- Von Minckwitz, G.; Untch, M.; Blohmer, J.-U.; Costa, S.D.; Eidtmann, H.; Fasching, P.A.; Gerber, B.; Eiermann, W.; Hilfrich, J.; Huober, J.; et al. Definition and Impact of Pathologic Complete Response on Prognosis After Neoadjuvant Chemotherapy in Various Intrinsic Breast Cancer Subtypes. J. Clin. Oncol. 2012, 30, 1796–1804. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asaoka, M.; Narui, K.; Suganuma, N.; Chishima, T.; Yamada, A.; Sugae, S.; Kawai, S.; Uenaka, N.; Teraoka, S.; Miyahara, K.; et al. Clinical and Pathological Predictors of Recurrence in Breast Cancer Patients Achieving Pathological Complete Response to Neoadjuvant Chemotherapy. Eur. J. Surg. Oncol. 2019, 45, 2289–2294. [Google Scholar] [CrossRef] [PubMed]
- Bossuyt, V.; Provenzano, E.; Symmans, W.F.; Boughey, J.C.; Coles, C.; Curigliano, G.; Dixon, J.M.; Esserman, L.J.; Fastner, G.; Kuehn, T.; et al. Recommendations for Standardized Pathological Characterization of Residual Disease for Neoadjuvant Clinical Trials of Breast Cancer by the BIG-NABCG Collaboration. Ann. Oncol. 2015, 26, 1280–1291. [Google Scholar] [CrossRef]
- Symmans, W.F.; Peintinger, F.; Hatzis, C.; Rajan, R.; Kuerer, H.; Valero, V.; Assad, L.; Poniecka, A.; Hennessy, B.; Green, M.; et al. Measurement of Residual Breast Cancer Burden to Predict Survival after Neoadjuvant Chemotherapy. J. Clin. Oncol. 2007, 25, 4414–4422. [Google Scholar] [CrossRef]
- McDonald, B.R.; Contente-Cuomo, T.; Sammut, S.-J.; Odenheimer-Bergman, A.; Ernst, B.; Perdigones, N.; Chin, S.-F.; Farooq, M.; Mejia, R.; Cronin, P.A.; et al. Personalized Circulating Tumor DNA Analysis to Detect Residual Disease after Neoadjuvant Therapy in Breast Cancer. Sci. Transl. Med. 2019, 11, eaax7392. [Google Scholar] [CrossRef]
- Guan, G.; Wang, Y.; Sun, Q.; Wang, L.; Xie, F.; Yan, J.; Huang, H.; Liu, H. Utility of Urinary CtDNA to Monitoring Minimal Residual Disease in Early Breast Cancer Patients. Cancer Biomark. 2020, 28, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Katayama, A.; Miligy, I.M.; Shiino, S.; Toss, M.S.; Eldib, K.; Kurozumi, S.; Quinn, C.M.; Badr, N.; Murray, C.; Provenzano, E.; et al. Predictors of Pathological Complete Response to Neoadjuvant Treatment and Changes to Post-Neoadjuvant HER2 Status in HER2-Positive Invasive Breast Cancer. Mod. Pathol. 2021, 34, 1271–1281. [Google Scholar] [CrossRef]
- Rouzier, R.; Perou, C.M.; Symmans, W.F.; Ibrahim, N.; Cristofanilli, M.; Anderson, K.; Hess, K.R.; Stec, J.; Ayers, M.; Wagner, P.; et al. Breast Cancer Molecular Subtypes Respond Differently to Preoperative Chemotherapy. Clin. Cancer Res. 2005, 11, 5678–5685. [Google Scholar] [CrossRef] [Green Version]
- Edenberg, E.R.; Downey, M.; Toczyski, D. Polymerase Stalling during Replication, Transcription and Translation. Curr. Biol. 2014, 24, R445–R452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Banys-Paluchowski, M.; Krawczyk, N.; Fehm, T. Liquid Biopsy in Breast Cancer. Geburtshilfe Frauenheilkd 2020, 80, 1093–1104. [Google Scholar] [CrossRef] [PubMed]
- McCartney, A.; Vignoli, A.; Biganzoli, L.; Love, R.; Tenori, L.; Luchinat, C.; Leo, A.D. Metabolomics in Breast Cancer: A Decade in Review. Cancer Treat. Rev. 2018, 67, 88–96. [Google Scholar] [CrossRef]
- Wood, S.L.; Westbrook, J.A.; Brown, J.E. Omic-Profiling in Breast Cancer Metastasis to Bone: Implications for Mechanisms, Biomarkers and Treatment. Cancer Treat. Rev. 2014, 40, 139–152. [Google Scholar] [CrossRef]
- Ma, J.; Setton, J.; Lee, N.Y.; Riaz, N.; Powell, S.N. The Therapeutic Significance of Mutational Signatures from DNA Repair Deficiency in Cancer. Nat. Commun. 2018, 9, 3292. [Google Scholar] [CrossRef]
- Fasching, P.A.; Loibl, S.; Hu, C.; Hart, S.N.; Shimelis, H.; Moore, R.; Schem, C.; Tesch, H.; Untch, M.; Hilfrich, J.; et al. BRCA1/2 Mutations and Bevacizumab in the Neoadjuvant Treatment of Breast Cancer: Response and Prognosis Results in Patients With Triple-Negative Breast Cancer From the GeparQuinto Study. J. Clin. Oncol. 2018, 36, 2281–2287. [Google Scholar] [CrossRef]
- Guo, S.; Loibl, S.; von Minckwitz, G.; Darb-Esfahani, S.; Lederer, B.; Denkert, C. PIK3CA H1047R Mutation Associated with a Lower Pathological Complete Response Rate in Triple-Negative Breast Cancer Patients Treated with Anthracycline-Taxane–Based Neoadjuvant Chemotherapy. Cancer Res. Treat. 2020, 52, 689–696. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, W.; Jiang, T.; Nuciforo, P.; Hatzis, C.; Holmes, E.; Harbeck, N.; Sotiriou, C.; Peña, L.; Loi, S.; Rosa, D.D.; et al. Pathway Level Alterations Rather than Mutations in Single Genes Predict Response to HER2-Targeted Therapies in the Neo-ALTTO Trial. Ann. Oncol. 2017, 28, 128–135. [Google Scholar] [CrossRef] [PubMed]
- Glück, S.; Ross, J.S.; Royce, M.; McKenna, E.F.; Perou, C.M.; Avisar, E.; Wu, L. TP53 Genomics Predict Higher Clinical and Pathologic Tumor Response in Operable Early-Stage Breast Cancer Treated with Docetaxel-Capecitabine ± Trastuzumab. Breast Cancer Res. Treat. 2012, 132, 781–791. [Google Scholar] [CrossRef] [PubMed]
- Desmedt, C.; Di Leo, A.; de Azambuja, E.; Larsimont, D.; Haibe-Kains, B.; Selleslags, J.; Delaloge, S.; Duhem, C.; Kains, J.-P.; Carly, B.; et al. Multifactorial Approach to Predicting Resistance to Anthracyclines. J. Clin. Oncol. 2011. [Google Scholar] [CrossRef]
- Tibau, A.; López-Vilaró, L.; Pérez-Olabarria, M.; Vázquez, T.; Pons, C.; Gich, I.; Alonso, C.; Ojeda, B.; Ramón y Cajal, T.; Lerma, E.; et al. Chromosome 17 Centromere Duplication and Responsiveness to Anthracycline-Based Neoadjuvant Chemotherapy in Breast Cancer. Neoplasia 2014, 16, 861–867. [Google Scholar] [CrossRef] [Green Version]
- Kaklamani, V.G.; Jeruss, J.S.; Hughes, E.; Siziopikou, K.; Timms, K.M.; Gutin, A.; Abkevich, V.; Sangale, Z.; Solimeno, C.; Brown, K.L.; et al. Phase II Neoadjuvant Clinical Trial of Carboplatin and Eribulin in Women with Triple Negative Early-Stage Breast Cancer (NCT01372579). Breast Cancer Res. Treat. 2015, 151, 629–638. [Google Scholar] [CrossRef]
- Rodríguez-Paredes, M.; Esteller, M. Cancer Epigenetics Reaches Mainstream Oncology. Nat. Med. 2011, 17, 330–339. [Google Scholar] [CrossRef] [PubMed]
- De Almeida, B.P.; Apolónio, J.D.; Binnie, A.; Castelo-Branco, P. Roadmap of DNA Methylation in Breast Cancer Identifies Novel Prognostic Biomarkers. BMC Cancer 2019, 19, 219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujii, S.; Yamashita, S.; Yamaguchi, T.; Takahashi, M.; Hozumi, Y.; Ushijima, T.; Mukai, H. Pathological Complete Response of HER2-Positive Breast Cancer to Trastuzumab and Chemotherapy Can Be Predicted by HSD17B4 Methylation. Oncotarget 2017, 8, 19039–19048. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Connolly, R.M.; Fackler, M.J.; Zhang, Z.; Zhou, X.C.; Goetz, M.P.; Boughey, J.C.; Walsh, B.; Carpenter, J.T.; Storniolo, A.M.; Watkins, S.P.; et al. Tumor and Serum DNA Methylation in Women Receiving Preoperative Chemotherapy with or without Vorinostat in TBCRC008. Breast Cancer Res. Treat. 2018, 167, 107–116. [Google Scholar] [CrossRef]
- Fackler, M.J.; Umbricht, C.; Williams, D.; Argani, P.; Cruz, L.-A.; Merino, V.F.; Teo, W.W.; Zhang, Z.; Huang, P.; Visvananthan, K.; et al. Genome-Wide Methylation Analysis Identifies Genes Specific to Breast Cancer Hormone Receptor Status and Risk of Recurrence. Cancer Res. 2011, 71, 6195–6207. [Google Scholar] [CrossRef] [Green Version]
- Fackler, M.J.; Bujanda, Z.L.; Umbricht, C.; Teo, W.W.; Cho, S.; Zhang, Z.; Visvanathan, K.; Jeter, S.; Argani, P.; Wang, C.; et al. Novel Methylated Biomarkers and a Robust Assay to Detect Circulating Tumor DNA in Metastatic Breast Cancer. Cancer Res. 2014, 74, 2160–2170. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, Z.H.; Li, L.H.; Hua, D. Quantitative Analysis of Plasma Circulating DNA at Diagnosis and during Follow-up of Breast Cancer Patients. Cancer Lett. 2006, 243, 64–70. [Google Scholar] [CrossRef]
- Rohanizadegan, M. Analysis of Circulating Tumor DNA in Breast Cancer as a Diagnostic and Prognostic Biomarker. Cancer Genet. 2018, 228–229, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Magbanua, M.J.M.; Swigart, L.B.; Wu, H.-T.; Hirst, G.L.; Yau, C.; Wolf, D.M.; Tin, A.; Salari, R.; Shchegrova, S.; Pawar, H.; et al. Circulating Tumor DNA in Neoadjuvant-Treated Breast Cancer Reflects Response and Survival. Ann. Oncol. 2021, 32, 229–239. [Google Scholar] [CrossRef] [PubMed]
- Perou, C.M.; Sørlie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular Portraits of Human Breast Tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.C.; Hilsenbeck, S.G.; Fuqua, S.A. The Promise of Microarrays in the Management and Treatment of Breast Cancer. Breast Cancer Res. 2005, 7, 100–104. [Google Scholar] [CrossRef] [Green Version]
- Cobleigh, M.A.; Tabesh, B.; Bitterman, P.; Baker, J.; Cronin, M.; Liu, M.-L.; Borchik, R.; Mosquera, J.-M.; Walker, M.G.; Shak, S. Tumor Gene Expression and Prognosis in Breast Cancer Patients with 10 or More Positive Lymph Nodes. Clin. Cancer Res. 2005, 11, 8623–8631. [Google Scholar] [CrossRef] [Green Version]
- Cardoso, F.; van’t Veer, L.J.; Bogaerts, J.; Slaets, L.; Viale, G.; Delaloge, S.; Pierga, J.-Y.; Brain, E.; Causeret, S.; DeLorenzi, M.; et al. 70-Gene Signature as an Aid to Treatment Decisions in Early-Stage Breast Cancer. N. Engl. J. Med. 2016, 375, 717–729. [Google Scholar] [CrossRef] [Green Version]
- Filipits, M.; Rudas, M.; Jakesz, R.; Dubsky, P.; Fitzal, F.; Singer, C.F.; Dietze, O.; Greil, R.; Jelen, A.; Sevelda, P.; et al. A New Molecular Predictor of Distant Recurrence in ER-Positive, HER2-Negative Breast Cancer Adds Independent Information to Conventional Clinical Risk Factors. Clin. Cancer Res. 2011, 17, 6012–6020. [Google Scholar] [CrossRef] [Green Version]
- Nielsen, T.O.; Parker, J.S.; Leung, S.; Voduc, D.; Ebbert, M.; Vickery, T.; Davies, S.R.; Snider, J.; Stijleman, I.J.; Reed, J.; et al. A Comparison of PAM50 Intrinsic Subtyping with Immunohistochemistry and Clinical Prognostic Factors in Tamoxifen-Treated Estrogen Receptor–Positive Breast Cancer. Clin. Cancer Res. 2010, 16, 5222–5232. [Google Scholar] [CrossRef] [Green Version]
- Jerevall, P.-L.; Ma, X.-J.; Li, H.; Salunga, R.; Kesty, N.C.; Erlander, M.G.; Sgroi, D.C.; Holmlund, B.; Skoog, L.; Fornander, T.; et al. Prognostic Utility of HOXB13:IL17BR and Molecular Grade Index in Early-Stage Breast Cancer Patients from the Stockholm Trial. Br. J. Cancer 2011, 104, 1762–1769. [Google Scholar] [CrossRef]
- Yamashita, T.; Honda, M.; Kaneko, S. Application of Serial Analysis of Gene Expression in Cancer Research. Curr. Pharm. Biotechnol. 2008, 9, 375–382. [Google Scholar] [CrossRef]
- Mazo, C.; Barron, S.; Mooney, C.; Gallagher, W.M. Multi-Gene Prognostic Signatures and Prediction of Pathological Complete Response to Neoadjuvant Chemotherapy in ER-Positive, HER2-Negative Breast Cancer Patients. Cancers 2020, 12, 1133. [Google Scholar] [CrossRef] [PubMed]
- Pease, A.M.; Riba, L.A.; Gruner, R.A.; Tung, N.M.; James, T.A. Oncotype DX® Recurrence Score as a Predictor of Response to Neoadjuvant Chemotherapy. Ann. Surg. Oncol. 2019, 26, 366–371. [Google Scholar] [CrossRef]
- Iorio, M.V.; Ferracin, M.; Liu, C.-G.; Veronese, A.; Spizzo, R.; Sabbioni, S.; Magri, E.; Pedriali, M.; Fabbri, M.; Campiglio, M.; et al. MicroRNA Gene Expression Deregulation in Human Breast Cancer. Cancer Res. 2005, 65, 7065–7070. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Casey, M.; Sweeney, K.J.; Brown, J.A.L.; Kerin, M.J. Exploring Circulating Micro-RNA in the Neoadjuvant Treatment of Breast Cancer. Int. J. Cancer 2016, 139, 12–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Cosimo, S.; Appierto, V.; Pizzamiglio, S.; Silvestri, M.; Baselga, J.; Piccart, M.; Huober, J.; Izquierdo, M.; de la Pena, L.; Hilbers, F.S.; et al. Early Modulation of Circulating MicroRNAs Levels in HER2-Positive Breast Cancer Patients Treated with Trastuzumab-Based Neoadjuvant Therapy. Int. J. Mol. Sci. 2020, 21, 1386. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; Su, F.; Lv, X.; Zhang, W.; Shang, X.; Zhang, Y.; Zhang, J. Serum MicroRNA-21 Predicted Treatment Outcome and Survival in HER2-Positive Breast Cancer Patients Receiving Neoadjuvant Chemotherapy Combined with Trastuzumab. Cancer Chemother. Pharmacol. 2019, 84, 1039–1049. [Google Scholar] [CrossRef] [PubMed]
- Stevic, I.; Müller, V.; Weber, K.; Fasching, P.A.; Karn, T.; Marmé, F.; Schem, C.; Stickeler, E.; Denkert, C.; van Mackelenbergh, M.; et al. Specific MicroRNA Signatures in Exosomes of Triple-Negative and HER2-Positive Breast Cancer Patients Undergoing Neoadjuvant Therapy within the GeparSixto Trial. BMC Med. 2018, 16, 179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cosimo, S.D.; Appierto, V.; Pizzamiglio, S.; Tiberio, P.; Iorio, M.V.; Hilbers, F.; de Azambuja, E.; de la Peña, L.; Izquierdo, M.; Huober, J.; et al. Plasma MiRNA Levels for Predicting Therapeutic Response to Neoadjuvant Treatment in HER2-Positive Breast Cancer: Results from the NeoALTTO Trial. Clin. Cancer Res. 2019, 25, 3887–3895. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- García-García, F.; Salinas-Vera, Y.M.; García-Vázquez, R.; Marchat, L.A.; Rodríguez-Cuevas, S.; López-González, J.S.; Carlos-Reyes, Á.; Ramos-Payán, R.; Aguilar-Medina, M.; Pérez-Plasencia, C.; et al. MiR-145-5p Is Associated with Pathological Complete Response to Neoadjuvant Chemotherapy and Impairs Cell Proliferation by Targeting TGFβR2 in Breast Cancer. Oncol. Rep. 2019, 41, 3527–3534. [Google Scholar] [CrossRef]
- Raychaudhuri, M.; Bronger, H.; Buchner, T.; Kiechle, M.; Weichert, W.; Avril, S. MicroRNAs MiR-7 and MiR-340 Predict Response to Neoadjuvant Chemotherapy in Breast Cancer. Breast Cancer Res. Treat. 2017, 162, 511–521. [Google Scholar] [CrossRef] [Green Version]
- Müller, V.; Gade, S.; Steinbach, B.; Loibl, S.; von Minckwitz, G.; Untch, M.; Schwedler, K.; Lübbe, K.; Schem, C.; Fasching, P.A.; et al. Changes in Serum Levels of MiR-21, MiR-210, and MiR-373 in HER2-Positive Breast Cancer Patients Undergoing Neoadjuvant Therapy: A Translational Research Project within the Geparquinto Trial. Breast Cancer Res. Treat. 2014, 147, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Gasparello, J.; Allegretti, M.; Tremante, E.; Fabbri, E.; Amoreo, C.A.; Romania, P.; Melucci, E.; Messana, K.; Borgatti, M.; Giacomini, P.; et al. Liquid Biopsy in Mice Bearing Colorectal Carcinoma Xenografts: Gateways Regulating the Levels of Circulating Tumor DNA (CtDNA) and MiRNA (CtmiRNA). J. Exp. Clin. Cancer Res. 2018, 37, 124. [Google Scholar] [CrossRef] [Green Version]
- Souza, K.C.B.; Evangelista, A.F.; Leal, L.F.; Souza, C.P.; Vieira, R.A.; Causin, R.L.; Neuber, A.C.; Pessoa, D.P.; Passos, G.A.S.; Reis, R.M.V.; et al. Identification of Cell-Free Circulating MicroRNAs for the Detection of Early Breast Cancer and Molecular Subtyping. J. Oncol. 2019, 2019, 8393769. [Google Scholar] [CrossRef] [Green Version]
- Alix-Panabières, C.; Pantel, K. Clinical Applications of Circulating Tumor Cells and Circulating Tumor DNA as Liquid Biopsy. Cancer Discov. 2016, 6, 479–491. [Google Scholar] [CrossRef] [Green Version]
- Openshaw, M.R.; Page, K.; Fernandez-Garcia, D.; Guttery, D.; Shaw, J.A. The Role of CtDNA Detection and the Potential of the Liquid Biopsy for Breast Cancer Monitoring. Expert Rev. Mol. Diagn. 2016, 16, 751–755. [Google Scholar] [CrossRef]
- Denkert, C.; Liedtke, C.; Tutt, A.; von Minckwitz, G. Molecular Alterations in Triple-Negative Breast Cancer—the Road to New Treatment Strategies. Lancet 2017, 389, 2430–2442. [Google Scholar] [CrossRef] [Green Version]
- Geiger, T.; Madden, S.F.; Gallagher, W.M.; Cox, J.; Mann, M. Proteomic Portrait of Human Breast Cancer Progression Identifies Novel Prognostic Markers. Cancer Res. 2012, 72, 2428–2439. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yanovich, G.; Agmon, H.; Harel, M.; Sonnenblick, A.; Peretz, T.; Geiger, T. Clinical Proteomics of Breast Cancer Reveals a Novel Layer of Breast Cancer Classification. Cancer Res. 2018, 78, 6001–6010. [Google Scholar] [CrossRef] [Green Version]
- Mueller, C.; Haymond, A.; Davis, J.B.; Williams, A.; Espina, V. Protein Biomarkers for Subtyping Breast Cancer and Implications for Future Research. Expert Rev. Proteom. 2018, 15, 131–152. [Google Scholar] [CrossRef]
- Allred, D.C.; Harvey, J.M.; Berardo, M.; Clark, G.M. Prognostic and Predictive Factors in Breast Cancer by Immunohistochemical Analysis. Mod. Pathol. 1998, 11, 155–168. [Google Scholar] [PubMed]
- Yoshioka, T.; Hosoda, M.; Yamamoto, M.; Taguchi, K.; Hatanaka, K.C.; Takakuwa, E.; Hatanaka, Y.; Matsuno, Y.; Yamashita, H. Prognostic Significance of Pathologic Complete Response and Ki67 Expression after Neoadjuvant Chemotherapy in Breast Cancer. Breast Cancer 2015, 22, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Alves, W.E.F.M.; Bonatelli, M.; Dufloth, R.; Kerr, L.M.; Carrara, G.F.A.; da Costa, R.F.A.; Scapulatempo-Neto, C.; Tiezzi, D.; da Costa Vieira, R.A.; Pinheiro, C. CAIX Is a Predictor of Pathological Complete Response and Is Associated with Higher Survival in Locally Advanced Breast Cancer Submitted to Neoadjuvant Chemotherapy. BMC Cancer 2019, 19, 1173. [Google Scholar] [CrossRef] [Green Version]
- Cerbelli, B.; Pernazza, A.; Botticelli, A.; Fortunato, L.; Monti, M.; Sciattella, P.; Campagna, D.; Mazzuca, F.; Mauri, M.; Naso, G.; et al. PD-L1 Expression in TNBC: A Predictive Biomarker of Response to Neoadjuvant Chemotherapy? BioMed Res. Int. 2017, 2017, e1750925. [Google Scholar] [CrossRef]
- Xing, M.; Wang, J.; Yang, Q.; Wang, Y.; Li, J.; Xiong, J.; Zhou, S. FKBP12 Is a Predictive Biomarker for Efficacy of Anthracycline-Based Chemotherapy in Breast Cancer. Cancer Chemother. Pharmacol. 2019, 84, 861–872. [Google Scholar] [CrossRef] [Green Version]
- Nakai, K.; Mitomi, H.; Alkam, Y.; Arakawa, A.; Yao, T.; Tokuda, E.; Saito, M.; Kasumi, F. Predictive Value of MGMT, HMLH1, HMSH2 and BRCA1 Protein Expression for Pathological Complete Response to Neoadjuvant Chemotherapy in Basal-like Breast Cancer Patients. Cancer Chemother. Pharmacol. 2012, 69, 923–930. [Google Scholar] [CrossRef]
- Chuthapisith, S.; Bean, B.E.; Cowley, G.; Eremin, J.M.; Samphao, S.; Layfield, R.; Kerr, I.D.; Wiseman, J.; El-Sheemy, M.; Sreenivasan, T.; et al. Annexins in Human Breast Cancer: Possible Predictors of Pathological Response to Neoadjuvant Chemotherapy. Eur. J. Cancer 2009, 45, 1274–1281. [Google Scholar] [CrossRef]
- Yerushalmi, R.; Woods, R.; Ravdin, P.M.; Hayes, M.M.; Gelmon, K.A. Ki67 in Breast Cancer: Prognostic and Predictive Potential. Lancet Oncol. 2010, 11, 174–183. [Google Scholar] [CrossRef]
- Dowsett, M.; Nielsen, T.O.; A’Hern, R.; Bartlett, J.; Coombes, R.C.; Cuzick, J.; Ellis, M.; Henry, N.L.; Hugh, J.C.; Lively, T.; et al. Assessment of Ki67 in Breast Cancer: Recommendations from the International Ki67 in Breast Cancer Working Group. J. Natl. Cancer Inst. 2011, 103, 1656–1664. [Google Scholar] [CrossRef] [Green Version]
- Faneyte, I.F.; Schrama, J.G.; Peterse, J.L.; Remijnse, P.L.; Rodenhuis, S.; van de Vijver, M.J. Breast Cancer Response to Neoadjuvant Chemotherapy: Predictive Markers and Relation with Outcome. Br. J. Cancer 2003, 88, 406–412. [Google Scholar] [CrossRef] [Green Version]
- Lee, H.; Ko, H.; Seol, H.; Noh, D.-Y.; Han, W.; Kim, T.-Y.; Im, S.-A.; Park, I.A. Expression of Immunohistochemical Markers before and after Neoadjuvant Chemotherapy in Breast Carcinoma, and Their Use as Predictors of Response. J. Breast Cancer 2013, 16, 395–403. [Google Scholar] [CrossRef] [Green Version]
- Botti, G.; Collina, F.; Scognamiglio, G.; Rao, F.; Peluso, V.; De Cecio, R.; Piezzo, M.; Landi, G.; De Laurentiis, M.; Cantile, M.; et al. Programmed Death Ligand 1 (PD-L1) Tumor Expression Is Associated with a Better Prognosis and Diabetic Disease in Triple Negative Breast Cancer Patients. Int. J. Mol. Sci. 2017, 18, 459. [Google Scholar] [CrossRef]
- Harding, M.W.; Galat, A.; Uehling, D.E.; Schreiber, S.L. A Receptor for the Immuno-Suppressant FK506 Is a Cis–Trans Peptidyl-Prolyl Isomerase. Nature 1989, 341, 758–760. [Google Scholar] [CrossRef]
- Schaff, L.R.; Yan, D.; Thyparambil, S.; Tian, Y.; Cecchi, F.; Rosenblum, M.; Reiner, A.S.; Panageas, K.S.; Hembrough, T.; Lin, A.L. Characterization of MGMT and EGFR Protein Expression in Glioblastoma and Association with Survival. J. Neurooncol. 2020, 146, 163–170. [Google Scholar] [CrossRef]
- Moss, S.E.; Morgan, R.O. The Annexins. Genome Biol. 2004, 5, 219. [Google Scholar] [CrossRef] [Green Version]
- Okano, M.; Oshi, M.; Butash, A.L.; Katsuta, E.; Tachibana, K.; Saito, K.; Okayama, H.; Peng, X.; Yan, L.; Kono, K.; et al. Triple-Negative Breast Cancer with High Levels of Annexin A1 Expression Is Associated with Mast Cell Infiltration, Inflammation, and Angiogenesis. Int. J. Mol. Sci. 2019, 20, 4197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lokman, N.A.; Ween, M.P.; Oehler, M.K.; Ricciardelli, C. The Role of Annexin A2 in Tumorigenesis and Cancer Progression. Cancer Microenviron. 2011, 4, 199–208. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beyene, D.A.; Kanarek, N.F.; Naab, T.J.; Ricks-Santi, L.L.; Hudson, T.S. Annexin 2 Protein Expression Is Associated with Breast Cancer Subtypes in African American Women. Heliyon 2020, 6, e03241. [Google Scholar] [CrossRef]
- Cortazar, P.; Zhang, L.; Untch, M.; Mehta, K.; Costantino, J.P.; Wolmark, N.; Bonnefoi, H.; Cameron, D.; Gianni, L.; Valagussa, P.; et al. Pathological Complete Response and Long-Term Clinical Benefit in Breast Cancer: The CTNeoBC Pooled Analysis. Lancet 2014, 384, 164–172. [Google Scholar] [CrossRef] [Green Version]
- Chica-Parrado, M.R.; Godoy-Ortiz, A.; Jiménez, B.; Ribelles, N.; Barragan, I.; Alba, E. Resistance to Neoadjuvant Treatment in Breast Cancer: Clinicopathological and Molecular Predictors. Cancers 2020, 12, 2012. [Google Scholar] [CrossRef] [PubMed]
- Bianchini, G.; Kiermaier, A.; Bianchi, G.V.; Im, Y.-H.; Pienkowski, T.; Liu, M.-C.; Tseng, L.-M.; Dowsett, M.; Zabaglo, L.; Kirk, S.; et al. Biomarker Analysis of the NeoSphere Study: Pertuzumab, Trastuzumab, and Docetaxel versus Trastuzumab plus Docetaxel, Pertuzumab plus Trastuzumab, or Pertuzumab plus Docetaxel for the Neoadjuvant Treatment of HER2-Positive Breast Cancer. Breast Cancer Res. 2017, 19, 16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harris, L.N.; Ismaila, N.; McShane, L.M.; Andre, F.; Collyar, D.E.; Gonzalez-Angulo, A.M.; Hammond, E.H.; Kuderer, N.M.; Liu, M.C.; Mennel, R.G.; et al. Use of Biomarkers to Guide Decisions on Adjuvant Systemic Therapy for Women With Early-Stage Invasive Breast Cancer: American Society of Clinical Oncology Clinical Practice Guideline. J. Clin. Oncol. 2016, 34, 1134–1150. [Google Scholar] [CrossRef] [Green Version]
- Shamai, G.; Binenbaum, Y.; Slossberg, R.; Duek, I.; Gil, Z.; Kimmel, R. Artificial Intelligence Algorithms to Assess Hormonal Status From Tissue Microarrays in Patients With Breast Cancer. JAMA Netw. Open 2019, 2, e197700. [Google Scholar] [CrossRef] [PubMed]
- Liefaard, M.C.; Lips, E.H.; Wesseling, J.; Hylton, N.M.; Lou, B.; Mansi, T.; Pusztai, L. The Way of the Future: Personalizing Treatment Plans Through Technology. Am. Soc. Clin. Oncol. Educ. Book 2021, 41, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Haque, W.; Verma, V.; Hatch, S.; Suzanne Klimberg, V.; Brian Butler, E.; Teh, B.S. Response Rates and Pathologic Complete Response by Breast Cancer Molecular Subtype Following Neoadjuvant Chemotherapy. Breast Cancer Res. Treat. 2018, 170, 559–567. [Google Scholar] [CrossRef]
- Wu, H.-J.; Chu, P.-Y. Recent Discoveries of Macromolecule- and Cell-Based Biomarkers and Therapeutic Implications in Breast Cancer. Int. J. Mol. Sci. 2021, 22, 636. [Google Scholar] [CrossRef]
- Goossens, N.; Nakagawa, S.; Sun, X.; Hoshida, Y. Cancer Biomarker Discovery and Validation. Transl. Cancer Res. 2015, 4, 256–269. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).