Maraviroc Prevents HCC Development by Suppressing Macrophages and the Liver Progenitor Cell Response in a Murine Chronic Liver Disease Model
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals and Animal Model
2.2. Serum Levels of Alanine Amino Transferase
2.3. Immunohistochemical Staining
2.4. Quantitation of Immunohistochemical Staining
2.5. LPC Cell Lines and Cell Culture
2.6. Isolation and Culture of Murine Bone Marrow Macrophages (BMMOs)
2.7. RNA Extraction and Gene Expression Quantification
2.8. Western Blotting and Protein Quantitation
2.9. Proliferation Assay
2.10. RNA-Seq
2.11. Single-Nucleus RNA Sequencing (snRNA-Seq)
2.12. Measurement of Impedance-Based Wound Healing of Confluent BMMO Cultures
2.13. Transwell Migration Assay
2.14. β-Arrestin Recruitment
2.15. Statistical Analyses
3. Results
3.1. Maraviroc Inhibited Human and Mouse CCR5
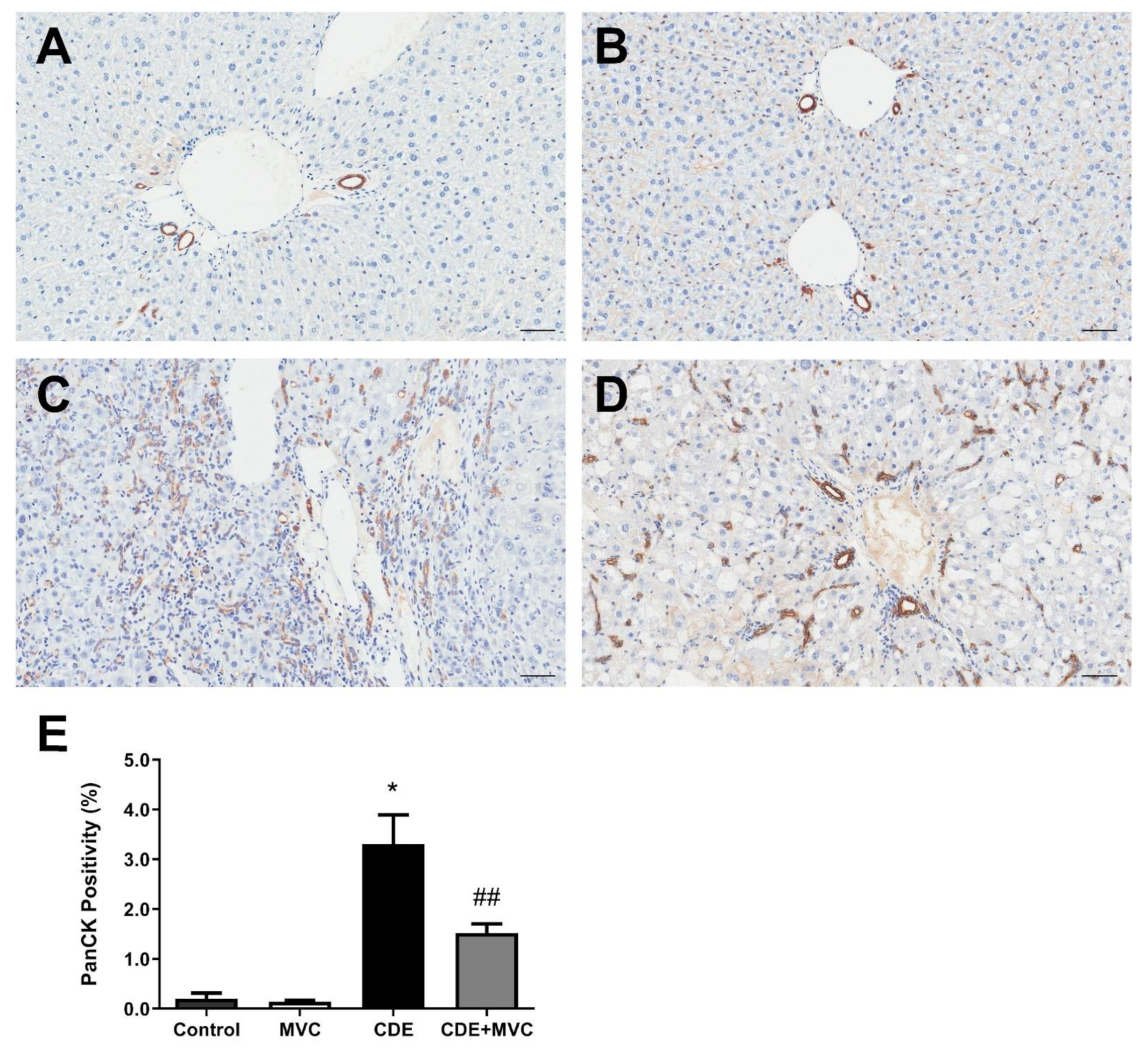
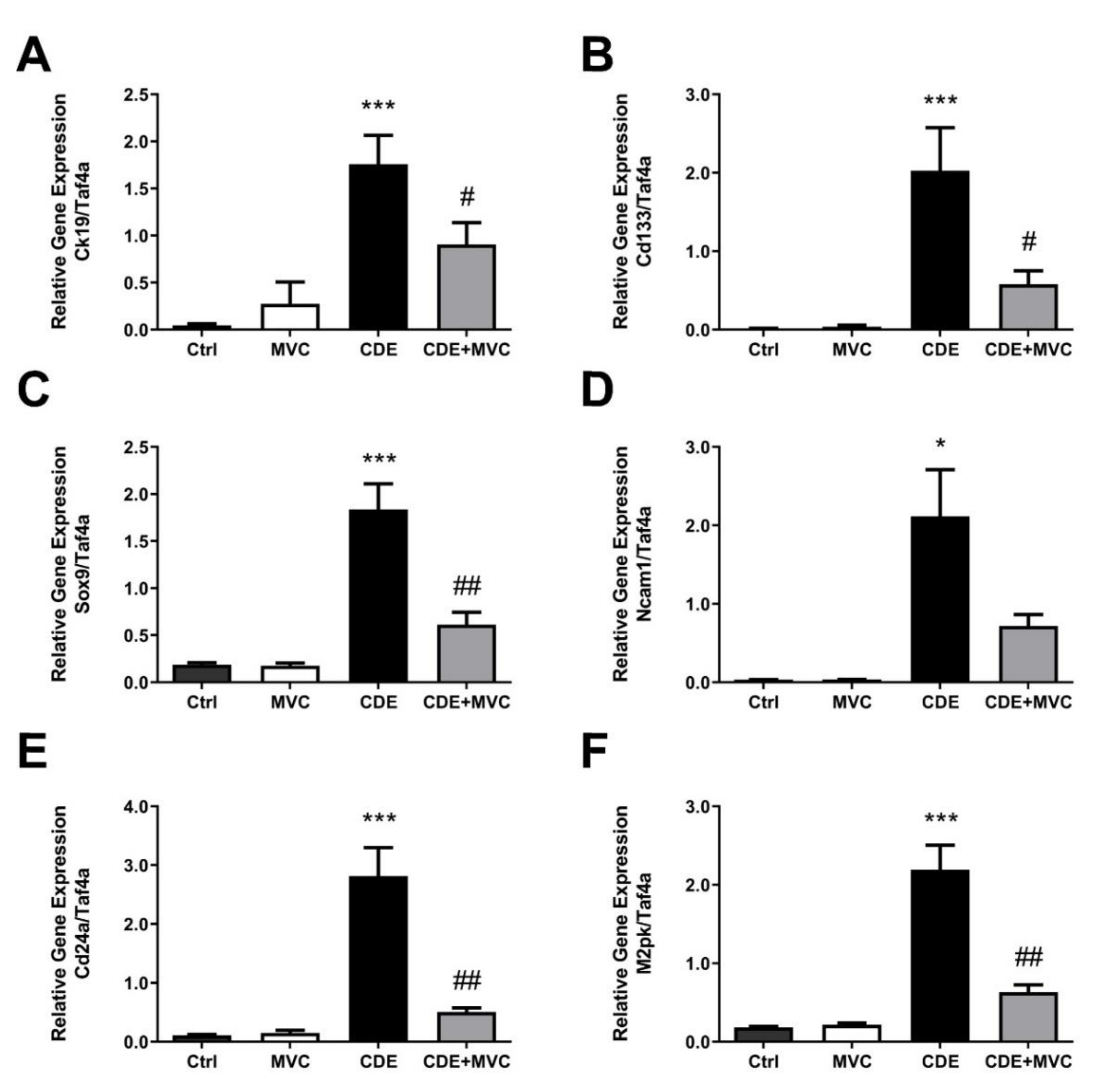
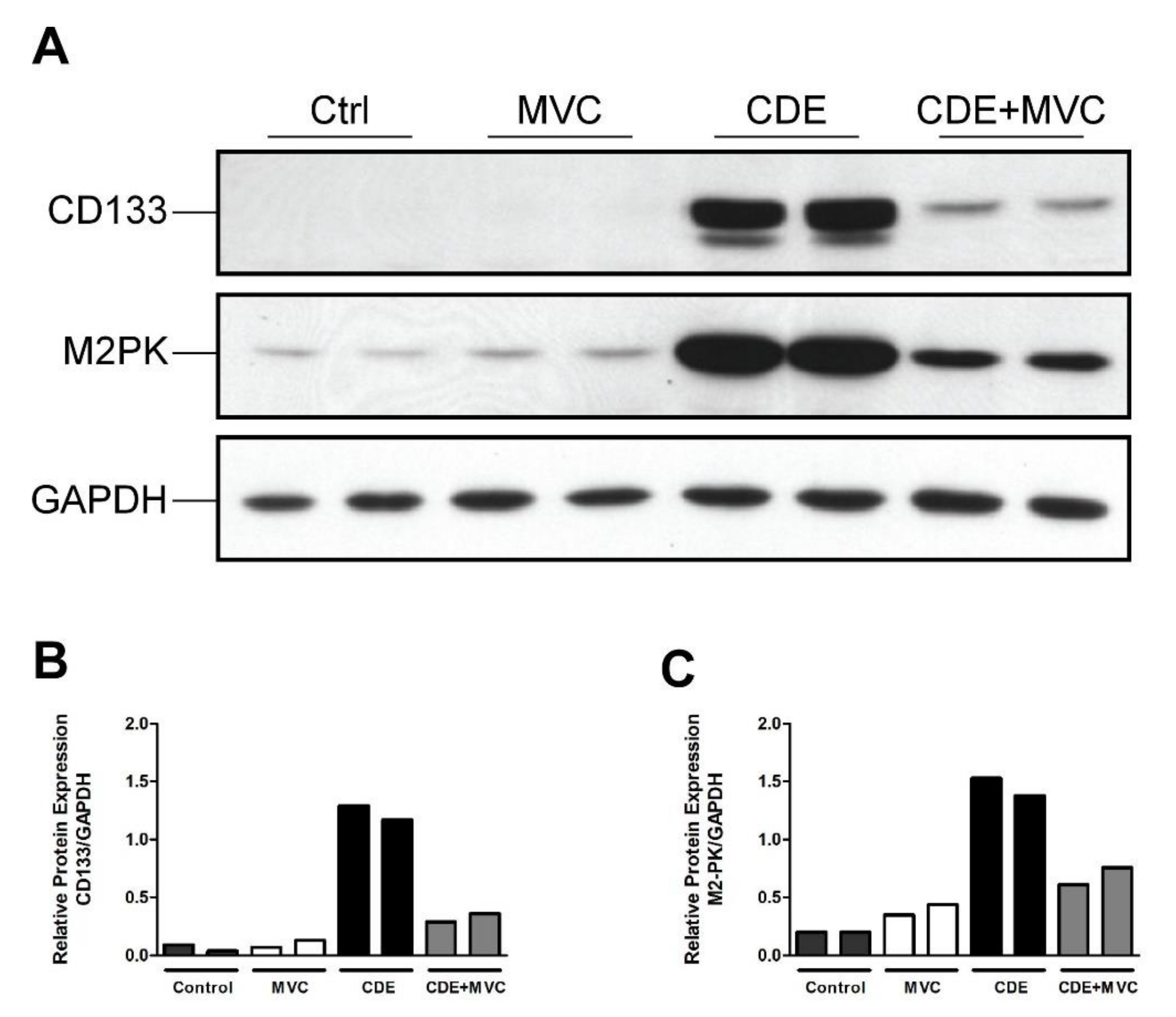
3.2. MVC Treatment Significantly Reduced CDE-Diet-Induced Liver Progenitor Cell Proliferation
3.3. MVC Reduced CDE-Induced Phosphorylation of AKT and STAT3
3.4. CCL5 and MVC Did Not Alter Proliferation of LPC Lines
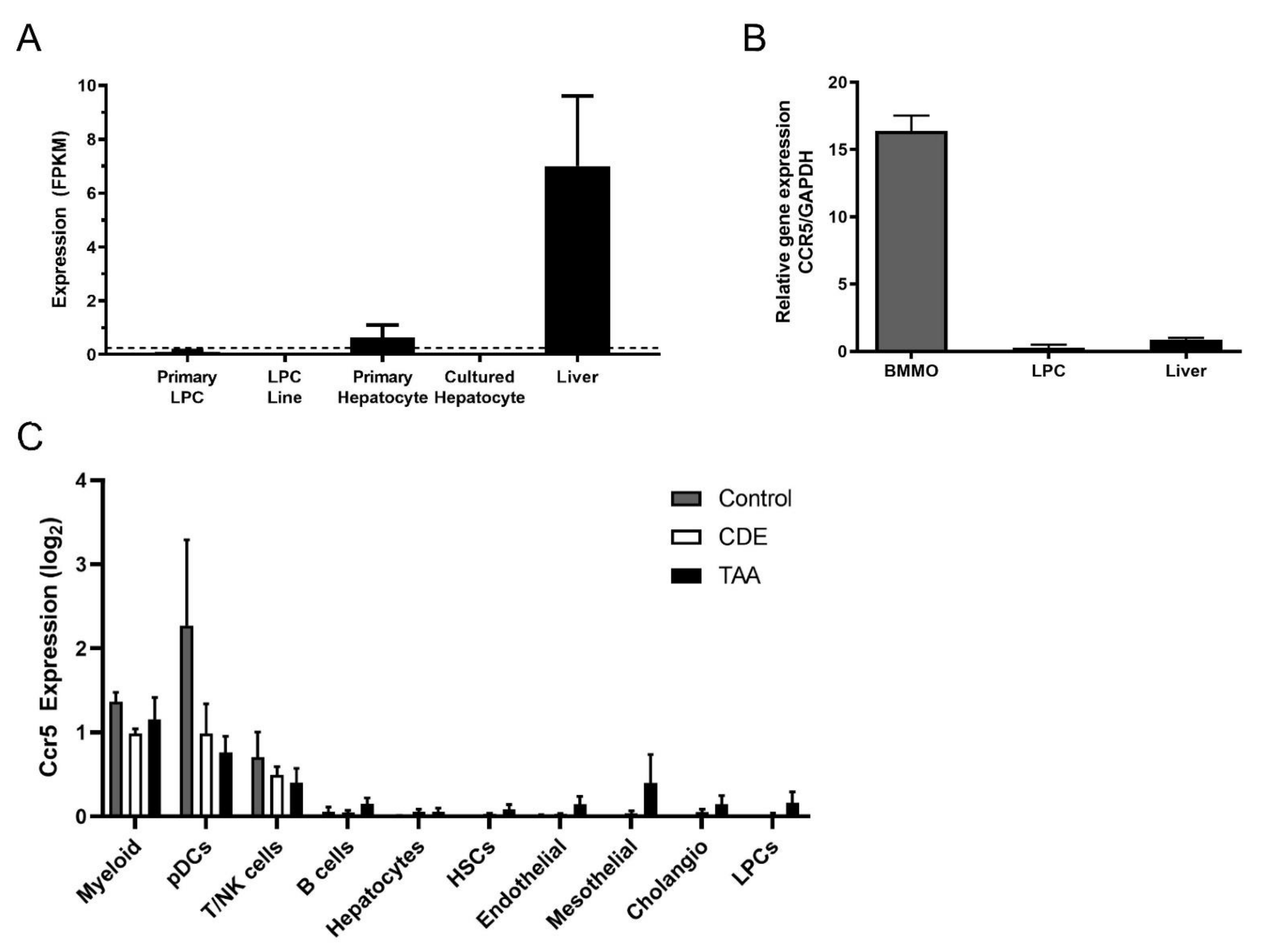
3.5. Ccr5 Was Expressed by CDE Liver and Macrophages but Not by LPCs
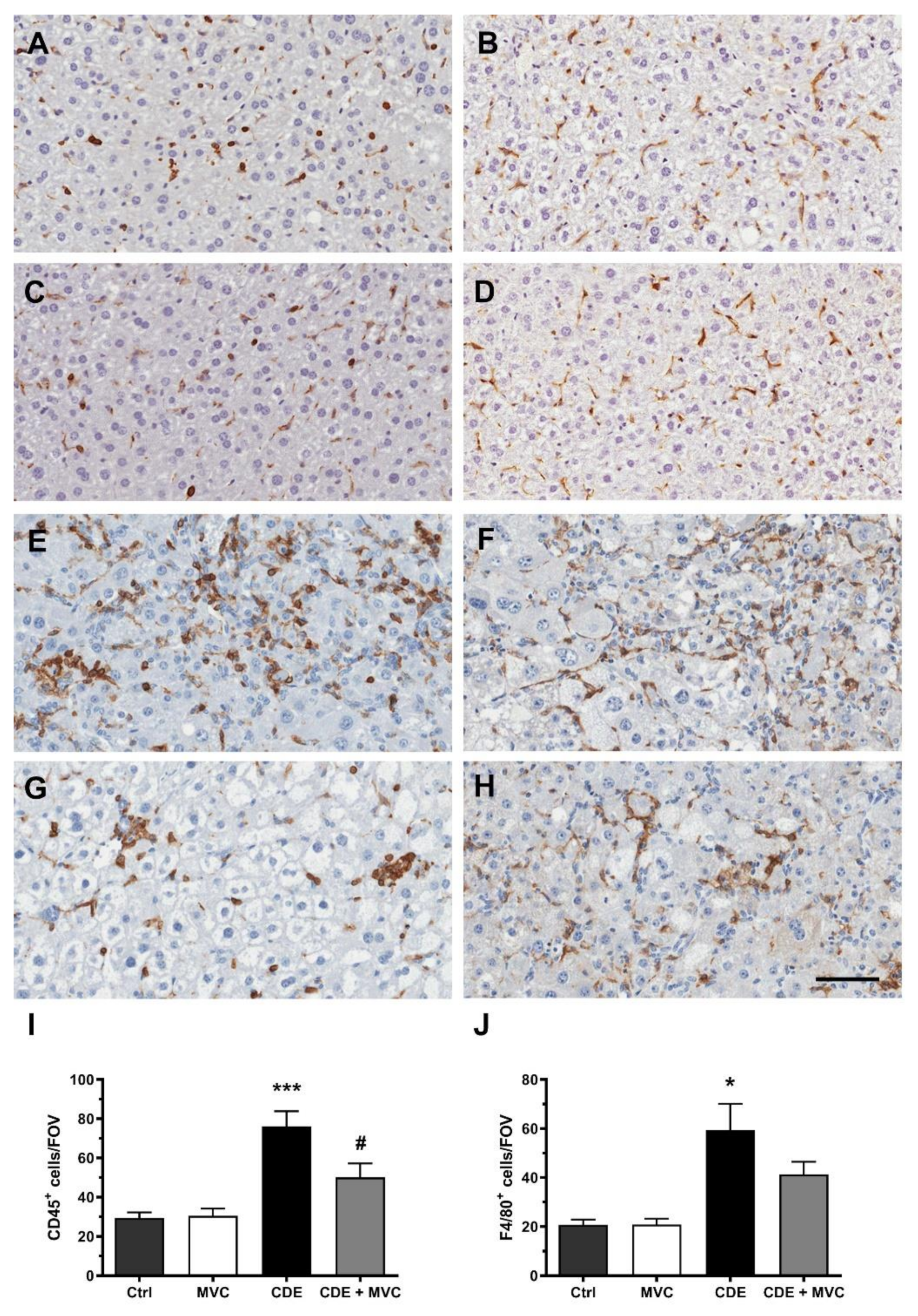
3.6. MVC Treatment Significantly Reduced the Inflammatory Response to the CDE Diet
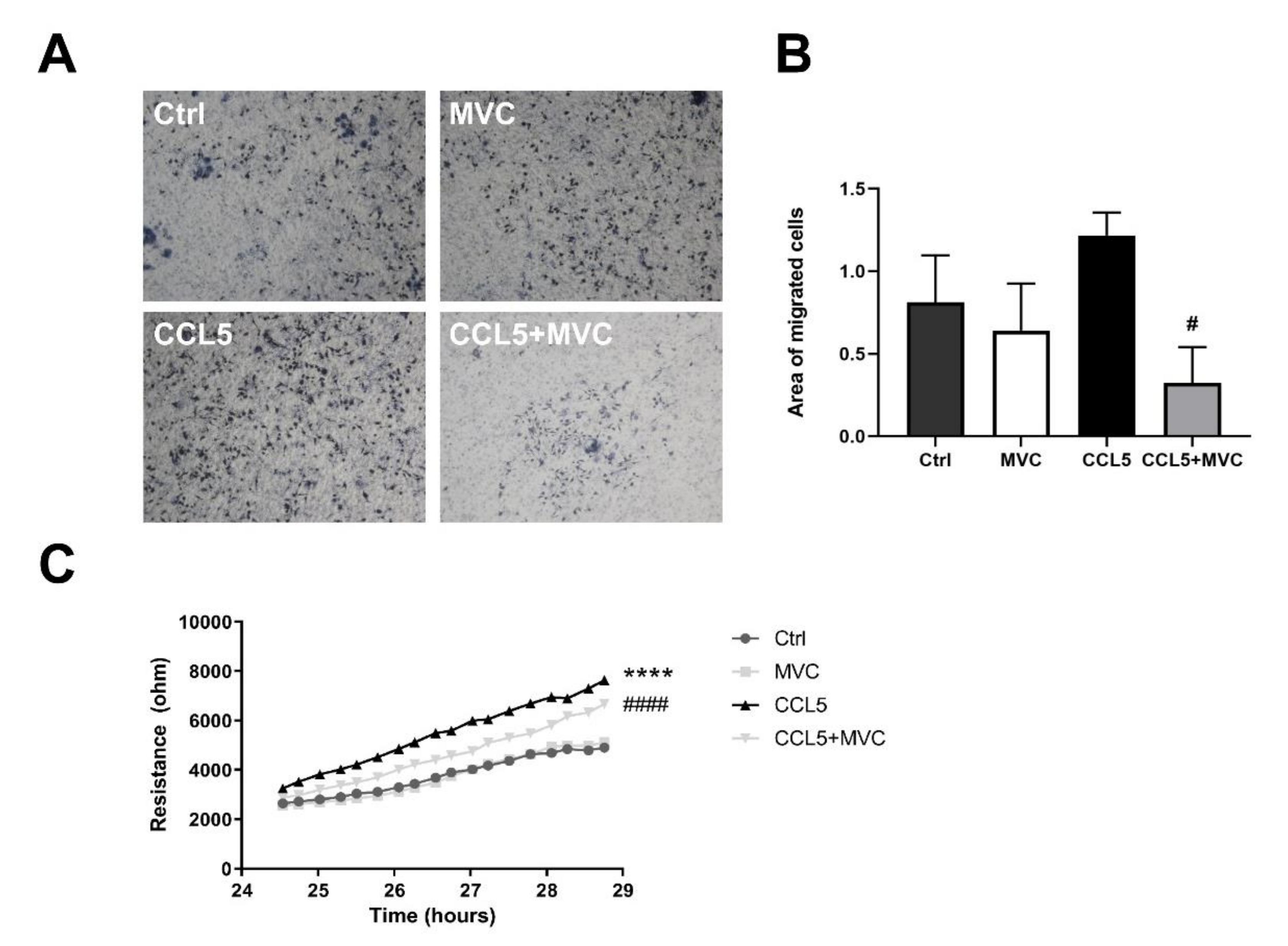
3.7. MVC-Treated Macrophages Exhibited Reduced Migration
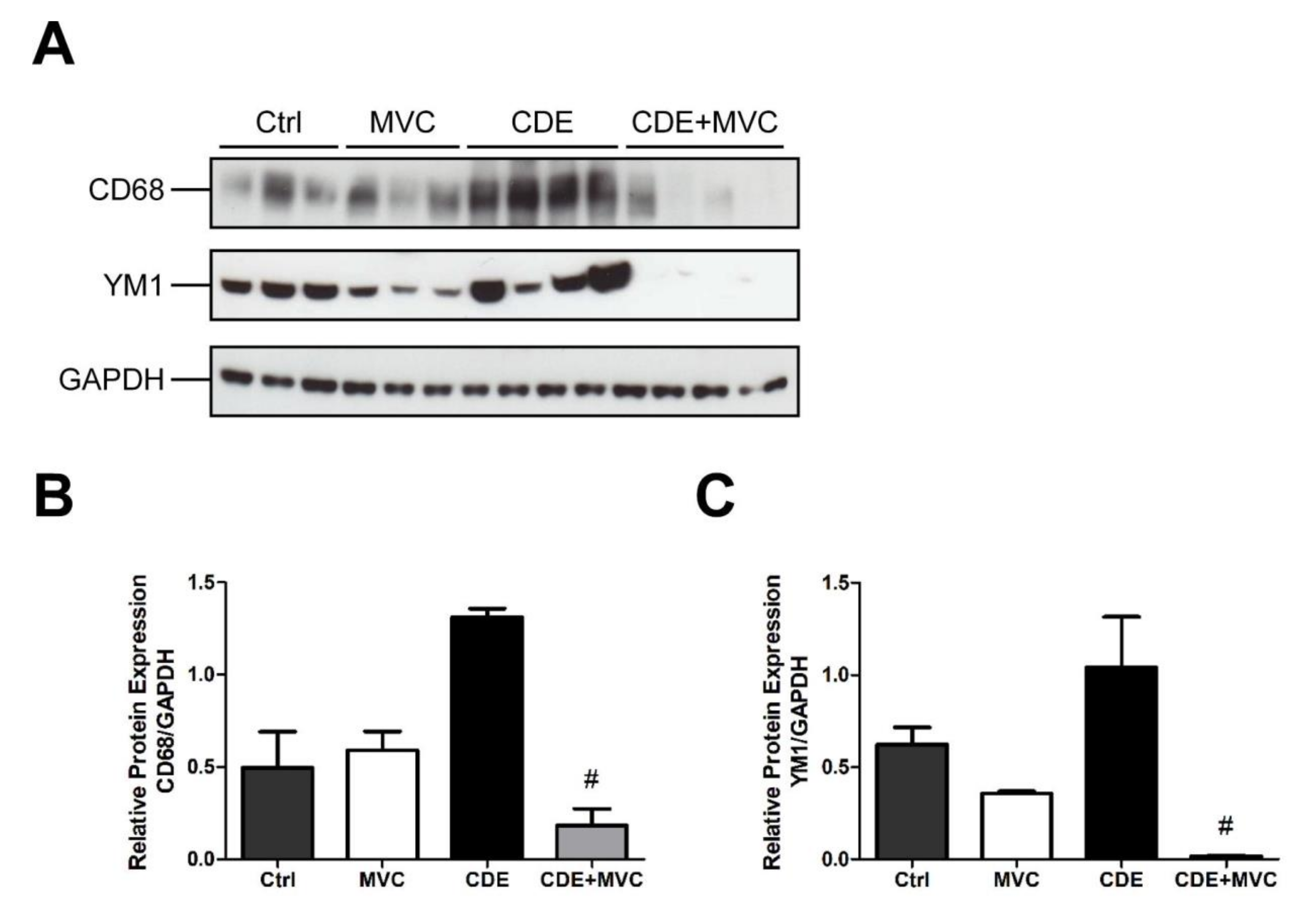
3.8. MVC Modified Macrophage Polarization Induced by CDE
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- El-Serag, H.B. Hepatocellular carcinoma. N. Engl. J. Med. 2011, 365, 1118–1127. [Google Scholar] [CrossRef]
- Sun, B.; Karin, M. Obesity, inflammation, and liver cancer. J. Hepatol. 2012, 56, 704–713. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Chen, L.; Zern, M.A.; Theise, N.D.; Diehl, A.M.; Liu, P.; Duan, Y. The diversity and plasticity of adult hepatic progenitor cells and their niche. Liver Int. 2017, 37, 1260–1271. [Google Scholar] [CrossRef]
- Hsia, C.C.; Evarts, R.P.; Nakatsukasa, H.; Marsden, E.R.; Thorgeirsson, S.S. Occurrence of oval-type cells in hepatitis B virus—Associated human hepatocarcinogenesis. Hepatology 1992, 16, 1327–1333. [Google Scholar] [CrossRef] [PubMed]
- Lowes, K.N.; Brennan, B.A.; Yeoh, G.; Olynyk, J.K. Oval cell numbers in human chronic liver diseases are directly related to disease severity. Am. J. Pathol. 1999, 154, 537–541. [Google Scholar] [CrossRef]
- Sancho-Bru, P.; Altamirano, J.; Rodrigo-Torres, D.; Coll, M.; Millán, C.; Lozano, J.J.; Miquel, R.; Arroyo, V.; Caballería, J.; Ginès, P.; et al. Liver progenitor cell markers correlate with liver damage and predict short-term mortality in patients with alcoholic hepatitis. Hepatology 2012, 55, 1931–1941. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.-R.; Xu, Y.; Yu, B.; Zhou, J.; Qiu, S.-J.; Shi, G.-M.; Zhang, B.-H.; Wu, W.-Z.; Shi, Y.-H.; Wu, B.; et al. High expression levels of putative hepatic stem/progenitor cell biomarkers related to tumour angiogenesis and poor prognosis of hepatocellular carcinoma. Gut 2010, 59, 953–962. [Google Scholar] [CrossRef]
- Braun, L.; Goyette, M.; Yaswen, P.; Thompson, N.L.; Fausto, N. Growth in culture and tumorigenicity after transfection with the ras oncogene of liver epithelial cells from carcinogen-treated rats. Cancer Res. 1987, 47, 4116–4124. [Google Scholar]
- Shachaf, C.M.; Kopelman, A.M.; Arvanitis, C.; Karlsson, A.; Beer, S.; Mandl, S.; Bachmann, M.; Borowsky, A.D.; Ruebner, B.; Cardiff, R.D.; et al. MYC inactivation uncovers pluripotent differentiation and tumour dormancy in hepatocellular cancer. Nature 2004, 431, 1112–1117. [Google Scholar] [CrossRef]
- Zender, L.; Spector, M.S.; Xue, W.; Flemming, P.; Cordon-Cardo, C.; Silke, J.; Fan, S.-T.; Luk, J.; Wigler, M.; Hannon, G.J.; et al. Identification and validation of oncogenes in liver cancer using an integrative oncogenomic approach. Cell 2006, 125, 1253–1267. [Google Scholar] [CrossRef]
- Dumble, M.L.; Croager, E.J.; Yeoh, G.; A Quail, E. Generation and characterization of p53 null transformed hepatic progenitor cells: Oval cells give rise to hepatocellular carcinoma. Carcinogenesis 2002, 23, 435–445. [Google Scholar] [CrossRef] [PubMed]
- Coussens, L.M. and Z. Werb, Inflammation and cancer. Nature 2002, 420, 860–867. [Google Scholar] [CrossRef]
- Lu, C.-F.; Huang, C.-S.; Tjiu, J.-W.; Chiang, C.-P. Infiltrating macrophage count: A significant predictor for the progression and prognosis of oral squamous cell carcinomas in Taiwan. Head Neck 2010, 32, 18–25. [Google Scholar] [CrossRef]
- Helm, O.; Held-Feindt, J.; Grage-Griebenow, E.; Reiling, N.; Ungefroren, H.; Vogel, I.; Krüger, U.; Becker, T.; Ebsen, M.; Röcken, C.; et al. Tumor-associated macrophages exhibit pro- and anti-inflammatory properties by which they impact on pancreatic tumorigenesis. Int. J. Cancer 2014, 135, 843–861. [Google Scholar] [CrossRef] [PubMed]
- Marra, F.; Tacke, F. Roles for chemokines in liver disease. Gastroenterology 2014, 147, 577–594. [Google Scholar] [CrossRef]
- Aldinucci, D.; Colombatti, A. The inflammatory chemokine CCL5 and cancer progression. Mediat. Inflamm. 2014, 2014, 292376. [Google Scholar]
- Van Hul, N.K.M.; Abarca-Quinones, J.; Sempoux, C.; Horsmans, Y.; Leclercq, I.A. Relation between liver progenitor cell expansion and extracellular matrix deposition in a CDE-induced murine model of chronic liver injury. Hepatology 2009, 49, 1625–1635. [Google Scholar] [CrossRef]
- Berres, M.-L.; Koenen, R.; Rueland, A.; Zaldivar, M.M.; Heinrichs, D.; Sahin, H.; Schmitz, P.; Streetz, K.L.; Berg, T.; Gassler, N.; et al. Antagonism of the chemokine Ccl5 ameliorates experimental liver fibrosis in mice. J. Clin. Investig. 2010, 120, 4129–4140. [Google Scholar] [CrossRef]
- Seki, E.; De Minicis, S.; Gwak, G.-Y.; Kluwe, J.; Inokuchi, S.; Bursill, C.; Llovet, J.M.; Brenner, D.; Schwabe, R.F. CCR1 and CCR5 promote hepatic fibrosis in mice. J. Clin. Investig. 2009, 119, 1858–1870. [Google Scholar] [CrossRef]
- Schwabe, R.F.; Bataller, R.; Brenner, D.A. Human hepatic stellate cells express CCR5 and RANTES to induce proliferation and migration. Am. J. Physiol.-Gastrointest. Liver Physiol. 2003, 285, G949–G958. [Google Scholar] [CrossRef]
- Ruddell, R.G.; Knight, B.; Tirnitz-Parker, J.E.E.; Akhurst, B.; Summerville, L.; Subramaniam, V.N.; Olynyk, J.K.; Ramm, G.A. Lymphotoxin-beta receptor signaling regulates hepatic stellate cell function and wound healing in a murine model of chronic liver injury. Hepatology 2009, 49, 227–239. [Google Scholar] [CrossRef]
- Fätkenheuer, G.; Pozniak, A.L.; A Johnson, M.; Plettenberg, A.; Staszewski, S.; Hoepelman, A.I.M.; Saag, M.S.; Goebel, F.D.; Rockstroh, J.K.; Dezube, B.J.; et al. Efficacy of short-term monotherapy with maraviroc, a new CCR5 antagonist, in patients infected with HIV-1. Nat. Med. 2005, 11, 1170–1172. [Google Scholar] [CrossRef]
- Gonzalez, E.O.; Boix, V.; Deltoro, M.G.; Aldeguer, J.L.; Portilla, J.; Montero, M.; Belda, E.B.; Abril, V.; Gutiérrez, F.; Minguez, C.; et al. The effects of Maraviroc on liver fibrosis in HIV/HCV co-infected patients. J. Int. AIDS Soc. 2014, 17 (Suppl. 3), 19643. [Google Scholar]
- Knight, B.; Yeoh, G.; Husk, K.L.; Ly, T.; Abraham, L.J.; Yu, C.; Rhim, J.A.; Fausto, N. Impaired preneoplastic changes and liver tumor formation in tumor necrosis factor receptor type 1 knockout mice. J. Exp. Med. 2000, 192, 1809–1818. [Google Scholar] [CrossRef]
- Akhurst, B.; Matthews, V.; Husk, K.; Smyth, M.J.; Abraham, L.J.; Yeoh, G.C. Differential lymphotoxin-beta and interferon gamma signaling during mouse liver regeneration induced by chronic and acute injury. Hepatology 2005, 41, 327–335. [Google Scholar] [CrossRef] [PubMed]
- Tirnitz-Parker, J.E.E.; Viebahn, C.S.; Jakubowski, A.; Klopcic, B.R.S.; Olynyk, J.K.; Yeoh, G.C.T.; Knight, B. Tumor necrosis factor–like weak inducer of apoptosis is a mitogen for liver progenitor cells. Hepatology 2010, 52, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Elsegood, C.L.; Chan, C.W.; Degli-Esposti, M.A.; Wikstrom, M.E.; Domenichini, A.; Lazarus, K.; van Rooijen, N.; Ganss, R.; Olynyk, J.K.; Yeoh, G.C. Kupffer cell-monocyte communication is essential for initiating murine liver progenitor cell-mediated liver regeneration. Hepatology 2015, 62, 1272–1284. [Google Scholar] [CrossRef]
- Ochoa-Callejero, L.; Pérez-Martínez, L.; Rubio-Mediavilla, S.; Oteo, J.A.; Martínez, A.; Blanco, J.-R. Maraviroc, a CCR5 antagonist, prevents development of hepatocellular carcinoma in a mouse model. PLoS ONE 2013, 8, e53992. [Google Scholar] [CrossRef] [PubMed]
- Neff, C.P.; Ndolo, T.; Tandon, A.; Habu, Y.; Akkina, R. Oral pre-exposure prophylaxis by anti-retrovirals raltegravir and maraviroc protects against HIV-1 vaginal transmission in a humanized mouse model. PLoS ONE 2010, 5, e15257. [Google Scholar] [CrossRef]
- Passman, A.M.; Strauss, R.P.; McSpadden, S.B.; Finch-Edmondson, M.L.; Woo, K.H.; Diepeveen, L.A.; London, R.; Callus, B.A.; Yeoh, G.C. A modified choline-deficient, ethionine-supplemented diet reduces morbidity and retains a liver progenitor cell response in mice. Dis. Model. Mech. 2015, 8, 1635–1641. [Google Scholar] [CrossRef]
- Tirnitz-Parker, J.E.E.; Tonkin, J.N.; Knight, B.; Olynyk, J.K.; Yeoh, G.C.T. Isolation, culture and immortalisation of hepatic oval cells from adult mice fed a choline-deficient, ethionine-supplemented diet. Int. J. Biochem. Cell Biol. 2007, 39, 2226–2239. [Google Scholar] [CrossRef] [PubMed]
- Akhurst, B.; Croager, E.J.; Farley-Roche, C.A.; Ong, J.K.; Dumble, M.L.; Knight, B.; Yeoh, G.C. A modified choline-deficient, ethionine-supplemented diet protocol effectively induces oval cells in mouse liver. Hepatology 2001, 34, 519–522. [Google Scholar] [CrossRef] [PubMed]
- Denisenko, E.; Guo, B.; Jones, M.; Hou, R.; de Kock, L.; Lassmann, T.; Poppe, D.; Clément, O.; Simmons, R.; Lister, R.; et al. Systematic assessment of tissue dissociation and storage biases in single-cell and single-nucleus RNA-seq workflows. Genome Biol. 2020, 21, 130. [Google Scholar] [CrossRef]
- Keese, C.R.; Wegener, J.; Walker, S.R.; Giaever, I. Electrical wound-healing assay for cells in vitro. Proc. Natl. Acad. Sci. USA 2004, 101, 1554–1559. [Google Scholar] [CrossRef] [PubMed]
- Dixon, A.S.; Schwinn, M.K.; Hall, M.P.; Zimmerman, K.; Otto, P.; Lubben, T.H.; Butler, B.L.; Binkowski, B.F.; Machleidt, T.; Kirkland, T.A.; et al. NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem. Biol. 2016, 11, 400–408. [Google Scholar] [CrossRef]
- Szpakowska, M.; Meyrath, M.; Reynders, N.; Counson, M.; Hanson, J.; Steyaert, J.; Chevigné, A. Mutational analysis of the extracellular disulphide bridges of the atypical chemokine receptor ACKR3/CXCR7 uncovers multiple binding and activation modes for its chemokine and endogenous non-chemokine agonists. Biochem. Pharmacol. 2018, 153, 299–309. [Google Scholar] [CrossRef]
- Szpakowska, M.; Nevins, A.M.; Meyrath, M.; Rhainds, D.; D’Huys, T.; Guité-Vinet, F.; Dupuis, N.; Gauthier, P.-A.; Counson, M.; Kleist, A.; et al. Different contributions of chemokine N-terminal features attest to a different ligand binding mode and a bias towards activation of ACKR3/CXCR7 compared with CXCR4 and CXCR3. Br. J. Pharmacol. 2018, 175, 1419–1438. [Google Scholar] [CrossRef] [PubMed]
- Perry, C.M. Maraviroc. Drugs 2010, 70, 1189–1213. [Google Scholar] [CrossRef]
- Kitade, M.; Factor, V.M.; Andersen, J.B.; Tomokuni, A.; Kaji, K.; Akita, H.; Holczbauer, A.; Seo, D.; Marquardt, J.; Conner, E.A.; et al. Specific fate decisions in adult hepatic progenitor cells driven by MET and EGFR signaling. Genes Dev. 2013, 27, 1706–1717. [Google Scholar] [CrossRef]
- Okano, J.-I.; Shiota, G.; Matsumoto, K.; Yasui, S.; Kurimasa, A.; Hisatome, I.; Steinberg, P.; Murawaki, Y. Hepatocyte growth factor exerts a proliferative effect on oval cells through the PI3K/AKT signaling pathway. Biochem. Biophys. Res. Commun. 2003, 309, 298–304. [Google Scholar] [CrossRef]
- Hao, N.-B.; Lü, M.-H.; Fan, Y.-H.; Cao, Y.-L.; Zhang, Z.-R.; Yang, S.-M. Macrophages in tumor microenvironments and the progression of tumors. Clin. Dev. Immunol. 2012, 2012, 948098. [Google Scholar] [CrossRef]
- Kumar, A.T.; Knops, A.; Swendseid, B.; Martinez-Outschoom, U.; Harshyne, L.; Philp, N.; Rodeck, U.; Luginbuhl, A.; Cognetti, D.; Johnson, J.; et al. Prognostic Significance of Tumor-Associated Macrophage Content in Head and Neck Squamous Cell Carcinoma: A Meta-Analysis. Front. Oncol. 2019, 9, 656. [Google Scholar] [CrossRef]
- Mantovani, A.; Sica, A.; Locati, M. Macrophage polarization comes of age. Immunity 2005, 23, 344–346. [Google Scholar] [CrossRef]
- Loomba, R.; Sanyal, A.J. The global NAFLD epidemic. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 686–690. [Google Scholar] [CrossRef]
- Dawson, D.A.; Goldstein, R.B.; Saha, T.D.; Grant, B.F. Changes in alcohol consumption: United States, 2001–2002 to 2012–2013. Drug Alcohol Depend. 2015, 148, 56–61. [Google Scholar] [CrossRef]
- Soria, G.; Ofri-Shahak, M.; Haas, I.; Yaal-Hahoshen, N.; Leider-Trejo, L.; Leibovich-Rivkin, T.; Weitzenfeld, P.; Meshel, T.; Shabtai, E.; Gutman, M.; et al. Inflammatory mediators in breast cancer: Coordinated expression of TNFalpha & IL-1beta with CCL2 & CCL5 and effects on epithelial-to-mesenchymal transition. BMC Cancer 2011, 11, 130. [Google Scholar]
- Mohs, A.; Kuttkat, N.; Reißing, J.; Zimmermann, H.W.; Sonntag, R.; Proudfoot, A.; Youssef, S.A.; de Bruin, A.; Cubero, F.J.; Trautwein, C. Functional role of CCL5/RANTES for HCC progression during chronic liver disease. J. Hepatol. 2017, 66, 743–753. [Google Scholar] [CrossRef]
- Velasco-Velázquez, M.; Jiao, X.; De La Fuente-Granada, M.; Pestell, T.G.; Ertel, A.; Lisanti, M.; Pestell, R.G. CCR5 antagonist blocks metastasis of basal breast cancer cells. Cancer Res. 2012, 72, 3839–3850. [Google Scholar] [CrossRef] [PubMed]
- Knight, B.; Matthews, V.B.; Akhurst, B.; Croager, E.J.; Klinken, E.; Abraham, L.J.; Olynyk, J.K.; Yeoh, G. Liver inflammation and cytokine production, but not acute phase protein synthesis, accompany the adult liver progenitor (oval) cell response to chronic liver injury. Immunol. Cell Biol. 2005, 83, 364–374. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Liu, T.; Cong, M.; Wu, X.; Bai, Y.; Yin, C.; An, W.; Wang, B.; Jia, J.; You, H. Expression of extracellular matrix genes in cultured hepatic oval cells: An origin of hepatic stellate cells through transforming growth factor beta? Liver Int. 2009, 29, 575–584. [Google Scholar] [CrossRef] [PubMed]
- Chuang, J.-Y.; Yang, W.-H.; Chen, H.-T.; Tan, T.-W.; Lin, Y.-T.; Hsu, C.-J.; Fong, Y.-C.; Tang, C.-H. CCL5/CCR5 axis promotes the motility of human oral cancer cells. J. Cell Physiol. 2009, 220, 418–426. [Google Scholar] [CrossRef]
- González-Arriagada, W.A.; Lozano-Burgos, C.; Zúñiga-Moreta, R.; González-Díaz, P.; Coletta, R.D. Clinicopathological significance of chemokine receptor (CCR 1, CCR 3, CCR 4, CCR 5, CCR 7 and CXCR 4) expression in head and neck squamous cell carcinomas. J. Oral Pathol. Med. 2018, 47, 755–763. [Google Scholar] [CrossRef] [PubMed]
- Arberas, H.; Guardo, A.C.; Bargallo, M.E.; Maleno, M.J.; Calvo, M.; Blanco, J.L.; Garcia, F.; Gatell, J.M.; Plana, M. In vitro effects of the CCR5 inhibitor maraviroc on human T cell function. J. Antimicrob. Chemother. 2012, 68, 577–586. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Ren, H.-Y.; Shi, Y.-J.; Liu, W. In vitro immunological effects of blocking CCR5 on T cells. Inflammation 2015, 38, 902–910. [Google Scholar] [CrossRef]
- Pozo-Balado, M.M.; Martínez-Bonet, M.; Rosado-Sánchez, I.; Ruiz-Mateos, E.; Méndez-Lagares, G.; Rodríguez-Méndez, M.M.; Vidal, F.; Muñoz-Fernández, M.A.; Pacheco, Y.M.; Leal, M. Maraviroc Reduces the Regulatory T-Cell Frequency in Antiretroviral-Naive HIV-Infected Subjects. J. Infect. Dis. 2014, 210, 890–898. [Google Scholar] [CrossRef][Green Version]
- Mencarelli, A.; Cipriani, S.; Francisci, D.; Santucci, L.; Baldelli, F.; Distrutti, E.; Fiorucci, S. Highly specific blockade of CCR5 inhibits leukocyte trafficking and reduces mucosal inflammation in murine colitis. Sci. Rep. 2016, 6, 30802. [Google Scholar] [CrossRef]
- Tanabe, Y.; Sasaki, S.; Mukaida, N.; Baba, T. Blockade of the chemokine receptor, CCR5, reduces the growth of orthotopically injected colon cancer cells via limiting cancerassociated fibroblast accumulation. Oncotarget 2016, 7, 48335. [Google Scholar] [CrossRef]
- Cipriani, S.; Francisci, D.; Mencarelli, A.; Renga, B.; Schiaroli, E.; D’Amore, C.; Baldelli, F.; Fiorucci, S. Efficacy of CCR5 antagonist maraviroc in reducing the early, ritonavir induced, atherogenesis and the advanced plaque progression in mice. Circulation 2013, 127, 2114–2124. [Google Scholar] [CrossRef]
- Rossi, R.; Lichtner, M.; De Rosa, A.; Sauzullo, I.; Mengoni, F.; Massetti, A.P.; Mastroianni, C.M.; Vullo, V. In vitro effect of anti-human immunodeficiency virus CCR5 antagonist maraviroc on chemotactic activity of monocytes, macrophages and dendritic cells. Clin. ExImmunol. 2011, 166, 184–190. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Matute, P.; Pichel, J.G.; Iñiguez, M.; Recio-Fernández, E.; Pérez-Martínez, L.; Torrens, R.; Blanco, J.R.; Oteo, J.A. Maraviroc ameliorates the increased adipose tissue macrophage recruitment induced by a high-fat diet in a mouse model of obesity. Antivir. Ther. 2017, 22, 163–168. [Google Scholar] [CrossRef]
- Chien, H.-C.; Chan, P.-C.; Tu, C.-C.; Day, Y.-J.; Hung, L.-M.; Juan, C.-C.; Tian, Y.-F.; Hsieh, P.-S. Importance of PLC-Dependent PI3K/AKT and AMPK Signaling in RANTES/CCR5 Mediated Macrophage Chemotaxis. Chin. J. Physiol. 2018, 61, 266–279. [Google Scholar] [CrossRef] [PubMed]
- Davies, R.A.; Knight, B.; Tian, Y.W.; Yeoh, G.C.; Olynyk, J.K. Hepatic oval cell response to the choline-deficient, ethionine supplemented model of murine liver injury is attenuated by the administration of a cyclo-oxygenase 2 inhibitor. Carcinogenesis 2006, 27, 1607–1616. [Google Scholar] [CrossRef]
- Xu, Y.; Zhao, W.; Xu, J.; Li, J.; Hong, Z.; Yin, Z.; Wang, X. Activated hepatic stellate cells promote liver cancer by induction of myeloid-derived suppressor cells through cyclooxygenase-2. Oncotarget 2016, 7, 8866. [Google Scholar] [CrossRef] [PubMed]
- Mantovani, A.; Marchesi, F.; Malesci, A.; Laghi, L.; Allavena, P. Tumour-associated macrophages as treatment targets in oncology. Nat. Rev. Clin. Oncol. 2017, 14, 399. [Google Scholar] [CrossRef] [PubMed]
- Azenshtein, E.; Luboshits, G.; Shina, S.; Neumark, E.; Shahbazian, D.; Weil, M.; Wigler, N.; Keydar, I.; Ben-Baruch, A. The CC chemokine RANTES in breast carcinoma progression: Regulation of expression and potential mechanisms of promalignant activity. Cancer Res. 2002, 62, 1093–1102. [Google Scholar] [PubMed]
- Frankenberger, C.; Rabe, D.; Bainer, R.; Sankarasharma, D.; Chada, K.; Krausz, T.; Gilad, Y.; Becker, L.; Rosner, M.R. Metastasis suppressors regulate the tumor microenvironment by blocking recruitment of prometastatic tumor-associated macrophages. Cancer Res. 2015, 75, 4063–4073. [Google Scholar] [CrossRef]
- Robinson, S.C.; Scott, K.A.; Wilson, J.L.; Thompson, R.G.; Proudfoot, A.E.I.; Balkwill, F.R. A chemokine receptor antagonist inhibits experimental breast tumor growth. Cancer Res. 2003, 63, 8360–8365. [Google Scholar]
- Halama, N.; Zoernig, I.; Berthel, A.; Kahlert, C.; Klupp, F.; Suarez-Carmona, M.; Suetterlin, T.; Brand, K.; Krauss, J.; Lasitschka, F.; et al. Tumoral immune cell exploitation in colorectal cancer metastases can be targeted effectively by anti-CCR5 therapy in cancer patients. Cancer Cell 2016, 29, 587–601. [Google Scholar] [CrossRef]
- Wang, H.-C.; Chen, C.-W.; Yang, C.-L.; Tsai, I.-M.; Hou, Y.-C.; Chen, C.-J.; Shan, Y.-S. Tumor-associated macrophages promote epigenetic silencing of gelsolin through DNA methyltransferase 1 in gastric cancer cells. Cancer Immunol. Res. 2017, 5, 885–897. [Google Scholar] [CrossRef]
- Han, X.; Qu, B. Expression of chemotactic factor CCL5 in gastric cancer tissue and its correlation with macrophage marker CD86. Biomed. Res. 2017, 28. [Google Scholar]
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Hadj Khodabakhshi, A.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1–10. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Passman, A.M.; Strauss, R.P.; McSpadden, S.B.; Finch-Edmondson, M.; Andrewartha, N.; Woo, K.H.; Diepeveen, L.A.; Zhao, W.; Fernández-Irigoyen, J.; Santamaría, E.; et al. Maraviroc Prevents HCC Development by Suppressing Macrophages and the Liver Progenitor Cell Response in a Murine Chronic Liver Disease Model. Cancers 2021, 13, 4935. https://doi.org/10.3390/cancers13194935
Passman AM, Strauss RP, McSpadden SB, Finch-Edmondson M, Andrewartha N, Woo KH, Diepeveen LA, Zhao W, Fernández-Irigoyen J, Santamaría E, et al. Maraviroc Prevents HCC Development by Suppressing Macrophages and the Liver Progenitor Cell Response in a Murine Chronic Liver Disease Model. Cancers. 2021; 13(19):4935. https://doi.org/10.3390/cancers13194935
Chicago/Turabian StylePassman, Adam M., Robyn P. Strauss, Sarah B. McSpadden, Megan Finch-Edmondson, Neil Andrewartha, Ken H. Woo, Luke A. Diepeveen, Weihao Zhao, Joaquín Fernández-Irigoyen, Enrique Santamaría, and et al. 2021. "Maraviroc Prevents HCC Development by Suppressing Macrophages and the Liver Progenitor Cell Response in a Murine Chronic Liver Disease Model" Cancers 13, no. 19: 4935. https://doi.org/10.3390/cancers13194935
APA StylePassman, A. M., Strauss, R. P., McSpadden, S. B., Finch-Edmondson, M., Andrewartha, N., Woo, K. H., Diepeveen, L. A., Zhao, W., Fernández-Irigoyen, J., Santamaría, E., Medina-Ruiz, L., Szpakowska, M., Chevigné, A., Park, H., Carlessi, R., Tirnitz-Parker, J. E. E., Blanco, J. R., London, R., Callus, B. A., ... Ochoa-Callejero, L. (2021). Maraviroc Prevents HCC Development by Suppressing Macrophages and the Liver Progenitor Cell Response in a Murine Chronic Liver Disease Model. Cancers, 13(19), 4935. https://doi.org/10.3390/cancers13194935









