Patterns of Whole Exome Sequencing in Resected Cholangiocarcinoma
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Tyson, G.L.; Ilyas, J.A.; Duan, Z.; Green, L.K.; Younes, M.; El-Serag, H.B.; Davila, J.A. Secular trends in the incidence of cholangiocarcinoma in the USA and the impact of misclassification. Dig. Dis. Sci. 2014, 59, 3103–3110. [Google Scholar] [CrossRef]
- Ejaz, A.; Cloyd, J.M.; Pawlik, T.M. Advances in the Diagnosis and Treatment of Patients with Intrahepatic Cholangiocarcinoma. Ann. Surg. Oncol. 2020, 27, 552–560. [Google Scholar] [CrossRef] [PubMed]
- Poultsides, G.A.; Zhu, A.X.; Choti, M.A.; Pawlik, T.M. Intrahepatic cholangiocarcinoma. Surg. Clin. N. Am. 2010, 90, 817–837. [Google Scholar] [CrossRef]
- Diaz-Gonzalez, A.; Vilana, R.; Bianchi, L.; Garcia-Criado, A.; Rimola, J.; Rodriguez de Lope, C.; Ferrer, J.; Ayuso, C.; Da Fonseca, L.G.; Reig, M.; et al. Thermal Ablation for Intrahepatic Cholangiocarcinoma in Cirrhosis: Safety and Efficacy in Non-Surgical Patients. J. Vasc. Interv. Radiol. 2020, 31, 710–719. [Google Scholar] [CrossRef] [PubMed]
- Blechacz, B. Cholangiocarcinoma: Current Knowledge and New Developments. Gut Liver 2017, 11, 13–26. [Google Scholar] [CrossRef] [PubMed]
- Ebata, T.; Hirano, S.; Konishi, M.; Uesaka, K.; Tsuchiya, Y.; Ohtsuka, M.; Kaneoka, Y.; Yamamoto, M.; Ambo, Y.; Shimizu, Y.; et al. Randomized clinical trial of adjuvant gemcitabine chemotherapy versus observation in resected bile duct cancer. Br. J. Surg. 2018, 105, 192–202. [Google Scholar] [CrossRef] [PubMed]
- Edeline, J.; Benabdelghani, M.; Bertaut, A.; Watelet, J.; Hammel, P.; Joly, J.P.; Boudjema, K.; Fartoux, L.; Bouhier-Leporrier, K.; Jouve, J.L.; et al. Gemcitabine and Oxaliplatin Chemotherapy or Surveillance in Resected Biliary Tract Cancer (PRODIGE 12-ACCORD 18-UNICANCER GI): A Randomized Phase III Study. J. Clin. Oncol. 2019, 37, 658–667. [Google Scholar] [CrossRef]
- Primrose, J.N.; Fox, R.P.; Palmer, D.H.; Malik, H.Z.; Prasad, R.; Mirza, D.; Anthony, A.; Corrie, P.; Falk, S.; Finch-Jones, M.; et al. Capecitabine compared with observation in resected biliary tract cancer (BILCAP): A randomised, controlled, multicentre, phase 3 study. Lancet Oncol. 2019, 20, 663–673. [Google Scholar] [CrossRef] [Green Version]
- Shroff, R.T.; Kennedy, E.B.; Bachini, M.; Bekaii-Saab, T.; Crane, C.; Edeline, J.; El-Khoueiry, A.; Feng, M.; Katz, M.H.G.; Primrose, J.; et al. Adjuvant Therapy for Resected Biliary Tract Cancer: ASCO Clinical Practice Guideline. J. Clin. Oncol. 2019, 37, 1015–1027. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valle, J.; Wasan, H.; Palmer, D.H.; Cunningham, D.; Anthoney, A.; Maraveyas, A.; Madhusudan, S.; Iveson, T.; Hughes, S.; Pereira, S.P.; et al. Cisplatin plus gemcitabine versus gemcitabine for biliary tract cancer. N. Engl. J. Med. 2010, 362, 1273–1281. [Google Scholar] [CrossRef] [Green Version]
- Okusaka, T.; Nakachi, K.; Fukutomi, A.; Mizuno, N.; Ohkawa, S.; Funakoshi, A.; Nagino, M.; Kondo, S.; Nagaoka, S.; Funai, J.; et al. Gemcitabine alone or in combination with cisplatin in patients with biliary tract cancer: A comparative multicentre study in Japan. Br. J. Cancer 2010, 103, 469–474. [Google Scholar] [CrossRef] [Green Version]
- Lamarca, A.; Barriuso, J.; McNamara, M.G.; Valle, J.W. Molecular targeted therapies: Ready for "prime time" in biliary tract cancer. J. Hepatol. 2020, 73, 170–185. [Google Scholar] [CrossRef] [Green Version]
- Ueno, M.; Ikeda, M.; Morizane, C.; Kobayashi, S.; Ohno, I.; Kondo, S.; Okano, N.; Kimura, K.; Asada, S.; Namba, Y.; et al. Nivolumab alone or in combination with cisplatin plus gemcitabine in Japanese patients with unresectable or recurrent biliary tract cancer: A non-randomised, multicentre, open-label, phase 1 study. Lancet Gastroenterol. Hepatol. 2019, 4, 611–621. [Google Scholar] [CrossRef]
- Marabelle, A.; Le, D.T.; Ascierto, P.A.; Di Giacomo, A.M.; De Jesus-Acosta, A.; Delord, J.P.; Geva, R.; Gottfried, M.; Penel, N.; Hansen, A.R.; et al. Efficacy of Pembrolizumab in Patients With Noncolorectal High Microsatellite Instability/Mismatch Repair-Deficient Cancer: Results From the Phase II KEYNOTE-158 Study. J. Clin. Oncol. 2020, 38, 1–10. [Google Scholar] [CrossRef]
- Banales, J.M.; Cardinale, V.; Carpino, G.; Marzioni, M.; Andersen, J.B.; Invernizzi, P.; Lind, G.E.; Folseraas, T.; Forbes, S.J.; Fouassier, L.; et al. Expert consensus document: Cholangiocarcinoma: Current knowledge and future perspectives consensus statement from the European Network for the Study of Cholangiocarcinoma (ENS-CCA). Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 261–280. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, H.; Arai, Y.; Totoki, Y.; Shirota, T.; Elzawahry, A.; Kato, M.; Hama, N.; Hosoda, F.; Urushidate, T.; Ohashi, S.; et al. Genomic spectra of biliary tract cancer. Nat. Genet. 2015, 47, 1003–1010. [Google Scholar] [CrossRef] [PubMed]
- Ross, J.S.; Wang, K.; Gay, L.; Al-Rohil, R.; Rand, J.V.; Jones, D.M.; Lee, H.J.; Sheehan, C.E.; Otto, G.A.; Palmer, G.; et al. New routes to targeted therapy of intrahepatic cholangiocarcinomas revealed by next-generation sequencing. Oncologist 2014, 19, 235–242. [Google Scholar] [CrossRef] [Green Version]
- Valle, J.W.; Lamarca, A.; Goyal, L.; Barriuso, J.; Zhu, A.X. New Horizons for Precision Medicine in Biliary Tract Cancers. Cancer Discov. 2017, 7, 943–962. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Verlingue, L.; Malka, D.; Allorant, A.; Massard, C.; Ferté, C.; Lacroix, L.; Rouleau, E.; Auger, N.; Ngo, M.; Nicotra, C.; et al. Precision medicine for patients with advanced biliary tract cancers: An effective strategy within the prospective MOSCATO-01 trial. Eur. J. Cancer (Oxf. Engl. 1990) 2017, 87, 122–130. [Google Scholar] [CrossRef]
- Simile, M.M.; Bagella, P.; Vidili, G.; Spanu, A.; Manetti, R.; Seddaiu, M.A.; Babudieri, S.; Madeddu, G.; Serra, P.A.; Altana, M.; et al. Targeted Therapies in Cholangiocarcinoma: Emerging Evidence from Clinical Trials. Medicina (Kaunas) 2019, 55, 42. [Google Scholar] [CrossRef] [Green Version]
- Thornblade, L.W.; Kessler, J.J.D.M.R. Thermal ablation of intrahepatic cholangiocarcinoma: A narrative review. Dig. Med. Res. 2021, 4. [Google Scholar] [CrossRef]
- Koeppel, F.; Bobard, A.; Lefebvre, C.; Pedrero, M.; Deloger, M.; Boursin, Y.; Richon, C.; Chen-Min-Tao, R.; Robert, G.; Meurice, G.; et al. Added Value of Whole-Exome and Transcriptome Sequencing for Clinical Molecular Screenings of Advanced Cancer Patients With Solid Tumors. Cancer J. (Sudbury Mass.) 2018, 24, 153–162. [Google Scholar] [CrossRef]
- Abou-Alfa, G.K.; Macarulla, T.; Javle, M.M.; Kelley, R.K.; Lubner, S.J.; Adeva, J.; Cleary, J.M.; Catenacci, D.V.; Borad, M.J.; Bridgewater, J.; et al. Ivosidenib in IDH1-mutant, chemotherapy-refractory cholangiocarcinoma (ClarIDHy): A multicentre, randomised, double-blind, placebo-controlled, phase 3 study. Lancet Oncol. 2020, 21, 796–807. [Google Scholar] [CrossRef]
- Banales, J.M.; Marin, J.J.G.; Lamarca, A.; Rodrigues, P.M.; Khan, S.A.; Roberts, L.R.; Cardinale, V.; Carpino, G.; Andersen, J.B.; Braconi, C.; et al. Cholangiocarcinoma 2020: The next horizon in mechanisms and management. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 557–588. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.K.; Gordan, J.D.; Kleinstiver, B.P.; Vu, P.; Najem, M.S.; Yeo, J.C.; Shi, L.; Kato, Y.; Levin, R.S.; Webber, J.T.; et al. Isocitrate Dehydrogenase Mutations Confer Dasatinib Hypersensitivity and SRC Dependence in Intrahepatic Cholangiocarcinoma. Cancer Discov. 2016, 6, 727–739. [Google Scholar] [CrossRef] [Green Version]
- Goyal, L.; Saha, S.K.; Liu, L.Y.; Siravegna, G.; Leshchiner, I.; Ahronian, L.G.; Lennerz, J.K.; Vu, P.; Deshpande, V.; Kambadakone, A.; et al. Polyclonal Secondary FGFR2 Mutations Drive Acquired Resistance to FGFR Inhibition in Patients with FGFR2 Fusion-Positive Cholangiocarcinoma. Cancer Discov. 2017, 7, 252–263. [Google Scholar] [CrossRef] [Green Version]
- Javle, M.; Lowery, M.; Shroff, R.T.; Weiss, K.H.; Springfeld, C.; Borad, M.J.; Ramanathan, R.K.; Goyal, L.; Sadeghi, S.; Macarulla, T.; et al. Phase II Study of BGJ398 in Patients With FGFR-Altered Advanced Cholangiocarcinoma. J. Clin. Oncol. 2018, 36, 276–282. [Google Scholar] [CrossRef]
- Abou-Alfa, G.K.; Sahai, V.; Hollebecque, A.; Vaccaro, G.; Melisi, D.; Al-Rajabi, R.; Paulson, A.S.; Borad, M.J.; Gallinson, D.; Murphy, A.G.; et al. Pemigatinib for previously treated, locally advanced or metastatic cholangiocarcinoma: A multicentre, open-label, phase 2 study. Lancet Oncol. 2020, 21, 671–684. [Google Scholar] [CrossRef]
- Goyal, L.; Shi, L.; Liu, L.Y.; Fece de la Cruz, F.; Lennerz, J.K.; Raghavan, S.; Leschiner, I.; Elagina, L.; Siravegna, G.; Ng, R.W.S.; et al. TAS-120 Overcomes Resistance to ATP-Competitive FGFR Inhibitors in Patients with FGFR2 Fusion-Positive Intrahepatic Cholangiocarcinoma. Cancer Discov. 2019, 9, 1064–1079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Javle, M.; Bekaii-Saab, T.; Jain, A.; Wang, Y.; Kelley, R.K.; Wang, K.; Kang, H.C.; Catenacci, D.; Ali, S.; Krishnan, S.; et al. Biliary cancer: Utility of next-generation sequencing for clinical management. Cancer 2016, 122, 3838–3847. [Google Scholar] [CrossRef] [Green Version]
- Subbiah, V.; Lassen, U.; Élez, E.; Italiano, A.; Curigliano, G.; Javle, M.; de Braud, F.; Prager, G.W.; Greil, R.; Stein, A.; et al. Dabrafenib plus trametinib in patients with BRAFV600E-mutated biliary tract cancer (ROAR): A phase 2, open-label, single-arm, multicentre basket trial. Lancet Oncol. 2020, 21, 1234–1243. [Google Scholar] [CrossRef]
- Bian, J.L.; Wang, M.M.; Tong, E.J.; Sun, J.; Li, M.; Miao, Z.B.; Li, Y.L.; Zhu, B.H.; Xu, J.J. Benefit of everolimus in treatment of an intrahepatic cholangiocarcinoma patient with a PIK3CA mutation. World J. Gastroenterol. 2017, 23, 4311–4316. [Google Scholar] [CrossRef]
- Brandi, G.; Farioli, A.; Astolfi, A.; Biasco, G.; Tavolari, S. Genetic heterogeneity in cholangiocarcinoma: A major challenge for targeted therapies. Oncotarget 2015, 6, 14744–14753. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Forner, A.; Vidili, G.; Rengo, M.; Bujanda, L.; Ponz-Sarvisé, M.; Lamarca, A. Clinical presentation, diagnosis and staging of cholangiocarcinoma. Liver Int. Off. J. Int. Assoc. Study Liver 2019, 39 (Suppl. 1), 98–107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ettrich, T.J.; Schwerdel, D.; Dolnik, A.; Beuter, F.; Blätte, T.J.; Schmidt, S.A.; Stanescu-Siegmund, N.; Steinacker, J.; Marienfeld, R.; Kleger, A.; et al. Genotyping of circulating tumor DNA in cholangiocarcinoma reveals diagnostic and prognostic information. Sci. Rep. 2019, 9, 13261. [Google Scholar] [CrossRef] [Green Version]

| All Patients n = 114 | Patients Receiving Sequencing n = 36 | Patients Not Receiving Sequencing n = 78 | p-Value | |
|---|---|---|---|---|
| Demographics | ||||
| Mean age (years ± SD) (at diagnosis) | 64.5 ± 11.8 | 60.7 (12.9) | 66.3 (10.9) | 0.02 |
| Female, n (%) | 51 (44.7) | 17 (47.2) | 34. (43.6) | 0.72 |
| Race, n (%) | ||||
| Asian | 30 (26.3) | 9 (25.0) | 21 (26.9) | 0.71 |
| Black | 0 (0) | 0 (0) | 0 (0) | |
| Hispanic | 30 (26.3) | 8 (22.2) | 22 (28.2) | |
| Middle Eastern | 0 (0) | 0 (0) | 0 (0) | |
| White | 54 (47.4) | 19 (52.3) | 35 (44.9) | |
| Cholangiocarcinoma risk factors, n (%) | ||||
| Viral hepatitis | 18 (15.8) | 5 (13.9) | 13 (16.7) | 0.71 |
| Primary sclerosing cholangitis | 1 (0.9) | 1 (2.8) | 0 (0) | 0.14 |
| Alcohol abuse | 5 (4.4) | 2 (5.6) | 3 (3.9) | 0.68 |
| Obese (BMI > 30) | 25 (22.1) | 6 (16.7) | 19 (24.7) | 0.34 |
| Diabetes | 27 (23.7) | 5 (13.9) | 22 (28.2) | 0.10 |
| Clinical characteristics, n (%) | ||||
| Obstructive jaundice at diagnosis | 46 (40.4) | 13 (36.1) | 33 (42.3) | 0.53 |
| Tumor site | 0.69 | |||
| Intrahepatic, n (%) | 61 (53.5) | 21 (58.3) | 40 (51.3) | |
| Hilar, n (%) | 17 (14.9) | 4 (11.1) | 13 (16.7) | |
| Extrahepatic distal, n (%) | 36 (31.6) | 11 (30.6) | 25 (32.1) | |
| Tumor grade, n (%) | 0.73 | |||
| Well differentiated | 9 (7.9) | 2 (5.6) | 7 (9.0) | |
| Moderately differentiated | 52 (45.6) | 18 (50.0) | 34 (43.6) | |
| Poorly differentiated | 53 (46.5) | 16 (44.4) | 37 (47.4) | |
| Mean tumor size, cm (SD) | 4.6 (2.5) | 4.5 (2.9) | 4.4 (2.3) | 0.80 |
| Multifocal disease | 16 (14.0) | 8 (22.2) | 8 (10.3) | 0.29 |
| Bilobar disease | 4 (3.5) | 1 (2.8) | 3 (3.9) | 0.77 |
| T-stage | 0.40 | |||
| 1 | 26 (22.9) | 8 (22.8) | 18 (24.3) | |
| 2 | 40 (36.7) | 16 (45.7) | 24 (32.4) | |
| 3 | 40 (36.7) | 11 (31.4) | 29 (39.2) | |
| 4 | 3 (2.8) | 0 (0) | 3 (4.1) | |
| N-stage | 0.34 | |||
| 0 | 57 (58.2) | 22 (68.8) | 35 (53.0) | |
| 1 | 37 (37.8) | 9 (28.1) | 28 (42.4) | |
| 2 | 4 (4.1) | 1 (3.1) | 3 (4.6) | |
| AJCC 8th Edition—Pathological Stage | 0.40 | |||
| IA | 12 (11.1) | 3 (8.8) | 9 (12.2) | |
| IB | 28 (25.9) | 7 (20.6) | 21 (28.4) | |
| IIA | 25 (23.2) | 12 (35.3) | 4 (5.4) | |
| IIB | 28 (25.9) | 7 (20.6) | 21 (28.4) | |
| IIIA | 10 (9.3) | 2 (5.9) | 8 (10.8) | |
| IIIB | 21 (19.4) | 7 (20.6) | 14 (18.9) | |
| IIIC | 3 (2.8) | 0 (0) | 3 (4.1) | |
| IV | 2 (1.9) | 0 (0) | 2 (2.7) |
| All Patients n = 114 | Patients Receiving Sequencing n = 36 | Patients Not Receiving Sequencing n = 78 | p-Value | |
|---|---|---|---|---|
| Systemic chemotherapy | 83 (72.8) | 28 (77.8) | 55 (70.5) | 0.42 |
| Radiation therapy | 43 (38.4) | 15 (41.7) | 28 (36.8) | 0.62 |
| Surgery | 0.64 | |||
| Liver resection without bile duct resection | 61 (53.5) | 21 (58.3) | 40 (51.3) | |
| Resection requiring bile duct resection/reconstruction | 21 (18.4) | 7 (19.4) | 14 (18.0) | |
| Pancreaticoduodenectomy | 32 (28.1) | 8 (22.2) | 24 (30.8) | |
| Margin status | 0.32 | |||
| R0 | 86 (75.4) | 26 (72.2) | 60 (76.9) | |
| R1 | 25 (21.9) | 10 (27.8) | 15 (19.2) | |
| R2 | 3 (2.6) | 0 (0) | 3 (3.9) | |
| Period of treatment | 0.06 | |||
| Early (2010–2013) | 25 (21.9) | 3 (8.3) | 22 (28.2) | |
| Mid (2014–2016) | 41 (36.0) | 16 (44.4) | 25 (32.1) | |
| Late (2017–2020) | 48 (42.1) | 17 (47.2) | 31 (39.7) | |
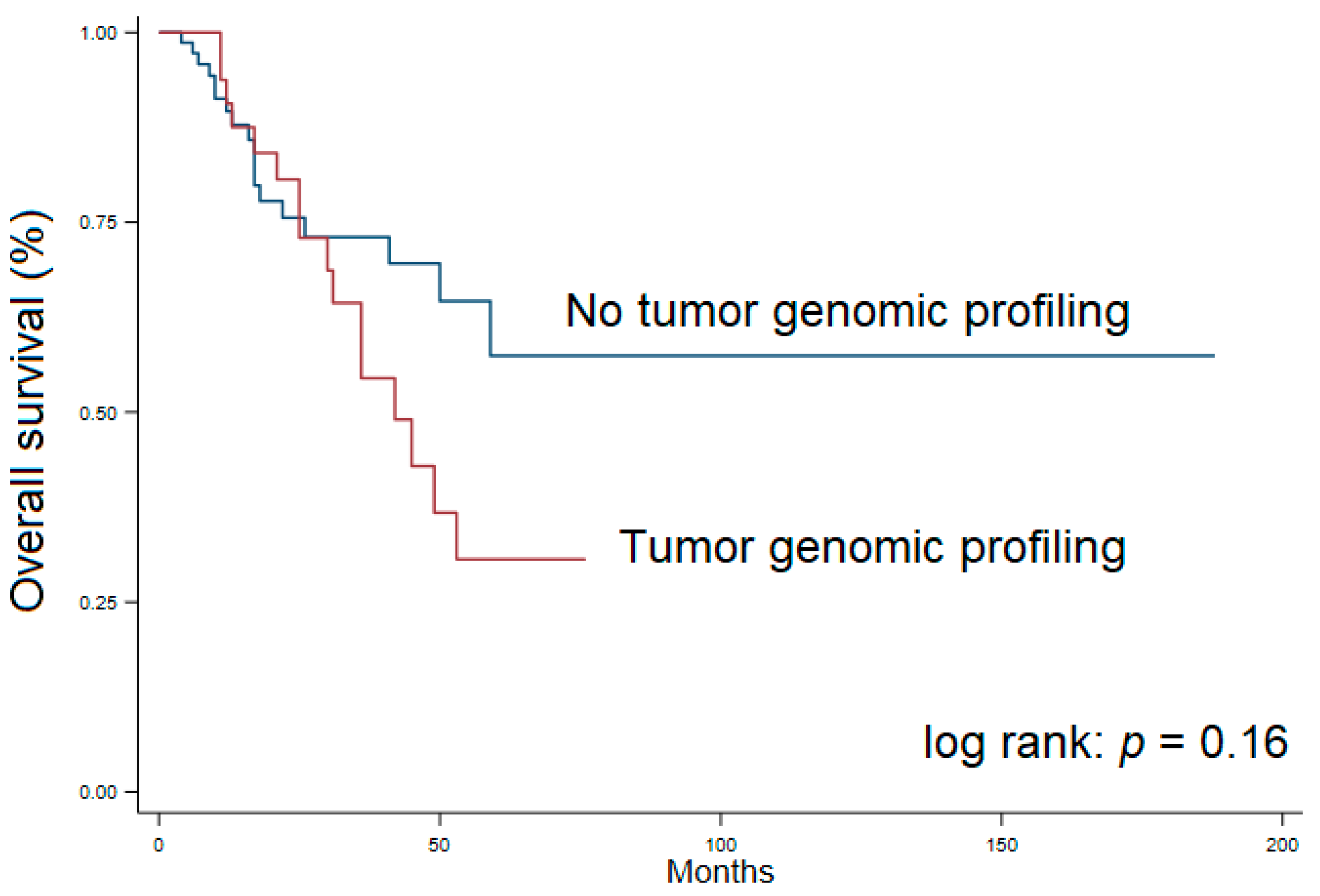
| Median overall survival—months (25%, 75%) | 59 (25, NR) | 42 (25, NR) | NR (26, NR) | 0.16 |
| Genes Assessed (WE: Whole Exome) | Actionable Mutations Identified | Tumor Mutational Burden (per MBP) | Identified Mutations/Alterations | Drugs Associated with Identified Mutations | Trials Identified | Therapy Received | Survival (Months) * AWD, † NED at Last Follow-up | |
|---|---|---|---|---|---|---|---|---|
| 1. | 324 | 2 | - | KRAS G12S LRP1B D478N | KRAS Cobimetinib Trametinib | 8 | Gemcitabine/Cisplatinum 5-FU FOLFOX Pembrolizumab | 53 |
| 2. | 48 | 0 | - | - | 0 | 0 | Gemcitabine/Cisplatinum | 68 |
| 3. | 324 | 2 | - | KRAS G12V TP53 R248Q | KRAS Trametinib | 6 | Gemcitabine/Cisplatinum Capecitabine | 21 |
| 4. | WE | 1 | 1 | NRAS Q61R | 0 | 11 | Gemcitabine/Cisplatinum FOLFOX | 76 * |
| 5. | 324 | 2 | - | BRAF V600E CDKN2A/B | BRAF Dabrafenib Regorafenib Trametinib Vemurafenib | 10 | Gemcitabine/Cisplatinum Capecitabine Dabrafenib/Trametinib | 31 |
| 6. | 50 | 2 | - | TP53 R213 NRAS G12D | MEK Trametinib mTOR Everolimus Temsirolimus | 5 | Gemcitabine/Cisplatinum | 13 |
| 7. | 324 | 6 | - | ARID1A E1763 & Q372 NF2 447 PTPN11 G503V CDKN2A EP300 | NF2 Everolimus Lapatinib Temsirolimus Trametinib PTPN11 Trametinib | 10 | Sorafenib Pembrolizumab | 45 |
| 8. | WE | 4 | 1 | KRAS G12V TP53 S125G CDK6 amplification CDKN2A L16 | CDK6 Abemaciclib Palbociclib Ribociclib | 28 | - | 68 * |
| 9. | 324 | 2 | - | BRAF V600E CDKN2A loss | BRAF Cobimetinib Dabrafenib Regorafenib Trametinib Vemurafenib | 4 | Gemcitabine/Cisplatinum | 59 * |
| 10. | 324 | 5 | - | ARID1A Y1719 IDH2 R172W TP53 Y220C BAP1 123-1 LRP1B R295 | None | 1 | - | 8 * |
| 11. | 324 | 6 | - | PIK3CA M1004I TP53 C141W ATRX A419V GATA6 amplification MCL1 amplification U2AF1 S34F | PIK3CA Everolimus Temsirolimus | 4 | Gemcitabine/Cisplatinum Pembrolizumab | 53 * |
| 12. | 324 | 7 | 5 | ERBB2 amplification KRAS amplification MET amplification CDKN2A p16INK4a & p14ARF TET2 R1572W | ERBB2 Afatinib Lapatinib Neratinib Pertuzumab Traztuzumab Ado-traztuzumab Traztuzumab-dkst KRAS Cobimetinib Trametinib MET Cabozantinib Crizotinib | 19 | Gemcitabine/Cisplatinum FOFOX Capecitabine | 35 * |
| 13. | 324 | 1 | - | IDH1 | None | 0 | Gemcitabine/Cisplatinum CAPOX | 36 |
| 14. | - | 2 | - | KRAS TP53 | None | 0 | Gemcitabine/Cisplatinum FOLFOX Atezolizumab Cobimetinib | 42 |
| 15. | 49 | 1 | - | PIK3CA E542K | None | 0 | - | 10 * |
| 16. | 48 | 1 | - | KRAS G12V | None | 0 | Gemcitabine/Cisplatinum FOLFIRI Capecitabine | 25 |
| 17. | WE | 0 | <1 | - | None | 0 | Gemcitabine/Cisplatinum FOLFOX 5-FU | 41 * |
| 18. | 324 & 48 | 5 | 3 | EGFR A289V BAP1 V616 TP53 V173M MET amplification | MET Crizotinib Cabozantinib EGFR Pantitumumab Osimertinib Lapatinib Gefitinib Erlotinib Cetuximab Afatinib | 10 | Gemcitabine/Cisplatinum FOLFOX FOLFIRI | 11 |
| 19. | 159 | 1 | 5 | IDH1 R132L | RRM1 positive Gemcitabine | 4 | Gemcitabine/Cisplatinum Capecitabine | 43 * |
| 20. | 159 | 1 | 5 | BAP1 E278 | None | 0 | Gemcitabine/Paclitaxel FOLFOX FOLFIRI Sorafenib | 30 |
| 21. | 48 | 3 | - | KRAS G12D TP53 H193R SMAD4 R361C | None | 0 | - | 11 |
| 22. | WE | 1 | <1 | TP53 V216G | None | 1 | Gemcitabine/Cisplatinum | 33 * |
| 23. | 324 | 3 | - | KRAS G12V TP53 A276D KDM6A P334 | KRAS Cobimetinib Trametinib | 0 | - | 18 * |
| 24. | 324 | 9 | 4 | PIK3CA E545K AKT1 W80R FGFR2 C382R CDKN2A/B loss SMAD4 R361C & R361H TP53 R213 | AKT1 & PIK3CA Everolimus Temsirolimus FGFR2 Pazopanib Ponatinib | 19 | Gemcitabine/Cisplatinum Capecitabine | 12 |
| 25. | 324 | 2 | - | BRAF V600E TBX3 942-1G | BRAF Cobimetinib Dabrafenib Trametinib Regorafenib Vemurafenib | 9 | Gemcitabine Capecitabine Pembrolizumab Dabrafenib/Trametinib Lenvatinib | 25 |
| 26. | WE | 3 | 2 | CCND1 amplification TERT C124C4 TSC1 Y48 | CCND1 Abemaciclib Palbociclib Ribociclib TSC1 Everolimus Temsirolimus | 17 | Pembrolizumab | 14 * |
| 27. | 49 | 2 | - | KRAS G13D GNAS R201H | None | 36 | Gemcitabine/Cisplatinum Capecitabine CAPOX | 25 * |
| 28. | WE | 2 | 1 | KRAS G12D PIK3CA H1047R | PIK3CA Copanlisib Everolimus Temsirolimus | 10 | Gemcitabine/Cisplatinum Capecitabine | 22 * |
| 29. | WE | 1 | <1 | BAP1 F15_T16 | 0 | 4 | Gemcitabine/Cisplatinum | 17 |
| 30. | 168 | 0 | 0 | MDM4 amplification NOTCH2 amplification FAM48C amplification PDGFRA amplification KIT amplification HIST2H3D amplification HIST2H3C amplification MCL1 amplification IL10 amplification FGFR3 amplification WHSC1 gain FGFR2 loss NFKB1A loss FGFR3 TACC3 | 0 | 0 | Gemcitabine Floxuridine | 49 |
| 31. | WE | 3 | 1 | CDKN2A H83N FGF3 amplification FGF4 amplification | FGF3 Sorafenib FGF4 Pazopanib Sorafenib | 10 | Gemcitabine/Cisplatinum Capecitabine | 15 † |
| 32. | WE | 1 | <1 | GNAS R844C | None | 6 | - | 7 † |
| 33. | WE | 1 | 1 | IDH1 R132C | IDH1 Ivosidinib | 0 | Capecitabine | 3 * |
| 34. | 48 | 4 | - | TP53 P152 ARID2 F1537 GNAS amplification ZNF217 amplificaiton | None | 6 | Gemcitabine/Cisplatinum 5-FU Pembrolizumab | 36 |
| 35. | WE | 0 | <1 | - | None | 0 | Gemcitabine/Cisplatinum | 22 † |
| 36. | WE | 4 | <1 | KRAS G12D IDH1 R132L KDM5C G1452 MDM2 amplification | IDH1 Ivosidinib KDM5C Sunitinib | 15 | Gemcitabine/Cisplatinum Ivosidinib | 26 * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thornblade, L.W.; Wong, P.; Li, D.; Warner, S.G.; Chang, S.; Raoof, M.; Kessler, J.; Amini, A.; Lin, J.; Chung, V.; et al. Patterns of Whole Exome Sequencing in Resected Cholangiocarcinoma. Cancers 2021, 13, 4062. https://doi.org/10.3390/cancers13164062
Thornblade LW, Wong P, Li D, Warner SG, Chang S, Raoof M, Kessler J, Amini A, Lin J, Chung V, et al. Patterns of Whole Exome Sequencing in Resected Cholangiocarcinoma. Cancers. 2021; 13(16):4062. https://doi.org/10.3390/cancers13164062
Chicago/Turabian StyleThornblade, Lucas W., Paul Wong, Daneng Li, Susanne G. Warner, Sue Chang, Mustafa Raoof, Jonathan Kessler, Arya Amini, James Lin, Vincent Chung, and et al. 2021. "Patterns of Whole Exome Sequencing in Resected Cholangiocarcinoma" Cancers 13, no. 16: 4062. https://doi.org/10.3390/cancers13164062
APA StyleThornblade, L. W., Wong, P., Li, D., Warner, S. G., Chang, S., Raoof, M., Kessler, J., Amini, A., Lin, J., Chung, V., Singh, G., Fong, Y., & Melstrom, L. G. (2021). Patterns of Whole Exome Sequencing in Resected Cholangiocarcinoma. Cancers, 13(16), 4062. https://doi.org/10.3390/cancers13164062







