1. Introduction
Over the past decade, advancements in immune checkpoint blockade treatment have expanded the toolkit available to clinicians in the battle against cancer [
1,
2,
3,
4,
5]. Used alone or in combination with other therapeutic agents, immune checkpoint inhibitors target components of the immune system to unleash the anti-tumor immune function of patients [
6]. Despite tremendous success and future promise, there are factors that pose challenges to reaching full potential in immune checkpoint therapies. With many regulatory components and their interactions involved in the anti-tumor immune response, it is difficult to thoroughly evaluate all the possible candidates. For candidates that are subjected to preclinical examination, there is the risk that they may be prematurely rejected due to the lack of sensitivity in the animal model system, or they may not translate well in clinical trials due to differences in the immune system between humans and model animals [
7]. In these contexts, mathematical and computational models can potentially serve as a complementary platform to facilitate and expedite the process of identifying effective therapeutic agents.
Quantitative systems pharmacology (QSP) models have become increasingly adopted as a powerful technique in drug discovery by academic researchers, the pharmaceutical industry and government regulators [
8,
9,
10,
11,
12,
13]. Typically composed of systems of ordinary differential equations, this type of model can integrate a large network of interacting components in a mechanistic manner and predict the behavior of the entire body as a complex system under different internal and external perturbations from disease and therapies. This feature is crucial in modeling immune checkpoint blockade treatment of cancer, considering the wide range of organ systems involved in mounting an adaptive immune response against tumor neoantigens and system-wise changes induced by the therapy when they attempt to curb the ability of cancer cell to evade immune surveillance [
14,
15]. In order to understand different factors contributing to a successful immune checkpoint therapy while addressing the complexity of the system, QSP models have been developed for drug development in generic solid tumor [
16,
17,
18,
19] as well as different specific human cancer types, including non-small-cell lung cancer (NSCLC) [
20,
21], breast cancer [
22,
23,
24], and melanoma [
25].
Despite the holistic approach the QSP models take to capture tumor growth in patients, one hallmark of cancer that is not reflected in this type of models is intratumoral heterogeneity (ITH) [
26,
27,
28,
29]. Tumor heterogeneity is observed in various forms, including spatial heterogeneity of cancer cells themselves and heterogeneity of immune cells that could play key roles in immune checkpoint blockade treatment [
30,
31,
32]. The recent development of digital pathology techniques has made spatial information on the tumor microenvironment such as multiplexed immunohistochemistry and immunofluorescence more abundantly available [
33,
34,
35]. Using these tools, researchers are able to quantitatively evaluate the characteristics of tumor microenvironment [
36,
37,
38] and correlate these characteristics with the effectiveness of treatment strategies involving immune checkpoint inhibitors [
39,
40,
41]. In this context, spatio-temporal agent-based models (ABM) are a useful tool to quantitatively investigate the generation of these spatial heterogeneities and their implications for treatment efficacy. In ABM, individual agents sense their local environment, interact with neighboring agents based on locally defined rules and produce global emergent behavior. In the field of immuno-oncology, studies have been performed with cancer cells and immune cells, represented as interacting cellular agents, to investigate the formation of spatial patterns of the cells and their response to checkpoint inhibition [
42,
43,
44]. These models provide insight into the effectiveness of certain immune checkpoint inhibitors with tumor heterogeneity accounted for; nevertheless, due to the lack of rigorously formulated pharmacokinetic/pharmacodynamic (PK/PD) modules as part of the overall whole-patient model, it is difficult to obtain quantitative predictions of patient response to the therapies.
In order to properly account for the patient-scale multi-organ system response to anti-tumor immune checkpoint therapy and the tissue–cellular scale heterogeneity, hybrid models are desirable in that they can potentially take advantage of the strengths of both QSP and ABM models [
45]. Hybrid models integrating ABM simulation with ODE-based models have already been used in other studies in the context of tumor development and drug testing. In addition to ABM simulations, these models use differential equations to recapitulate sub-cellular dynamics of signal transduction [
46,
47] or transport of anti-tumor drugs [
48]. Software packages have been created to facilitate development of hybrid models involving ABM for human disease studies, especially cancers. These packages, such as PhysiCell [
49], CompuCell [
50] and Hybrid Automata Library (HAL) [
51], integrate ABM simulation of the tissue-scale multicellular system with PDE/ODE solvers to handle chemical environment and molecular scale sub-cellular interactions. However, so far, these hybrid models have not incorporated a full-fledged QSP model to allow quantitative assessment of the efficacy of pharmaceutical agents against tumor development in the human body.
In the present study, based on the QSP-IO platform [
52], we formulate a general framework for a hybrid, fully coupled ABM-QSP model for immuno-oncology, which we refer to as spQSP-IO, where “sp” stands for spatial, and initially apply it to non-small-cell lung cancer (NSCLC). Lung cancer accounts for approximately 20% of cancer-related deaths worldwide, and more than 85% of lung cancer cases are classified as NSCLC [
53]. Cancer immunotherapy is becoming adopted as a new treatment modality for NSCLC, giving hope to patients with advanced-stage disease. In order to better harness the potential of immune checkpoint therapies, progress is needed in areas including identifying predictive biomarkers for patients’ responsiveness and finding combination therapies that synergize with immune checkpoint inhibitors to improve treatment efficacy [
54,
55,
56,
57,
58,
59,
60]. The spQSP-IO model is an extension of a previously published QSP model for NSCLC [
20] by representing a fraction of the tumor compartment dynamics with an ABM module to track the spatial heterogeneity in the tumor microenvironment. We aim at demonstrating how our spQSP-IO platform uses in silico simulations to facilitate testing the efficacy of immunotherapy agents and the discovery of biomarkers from simulated histopathology.
3. Results
3.1. Studying Tumor Spatial Characteristics Using spQSP-IO
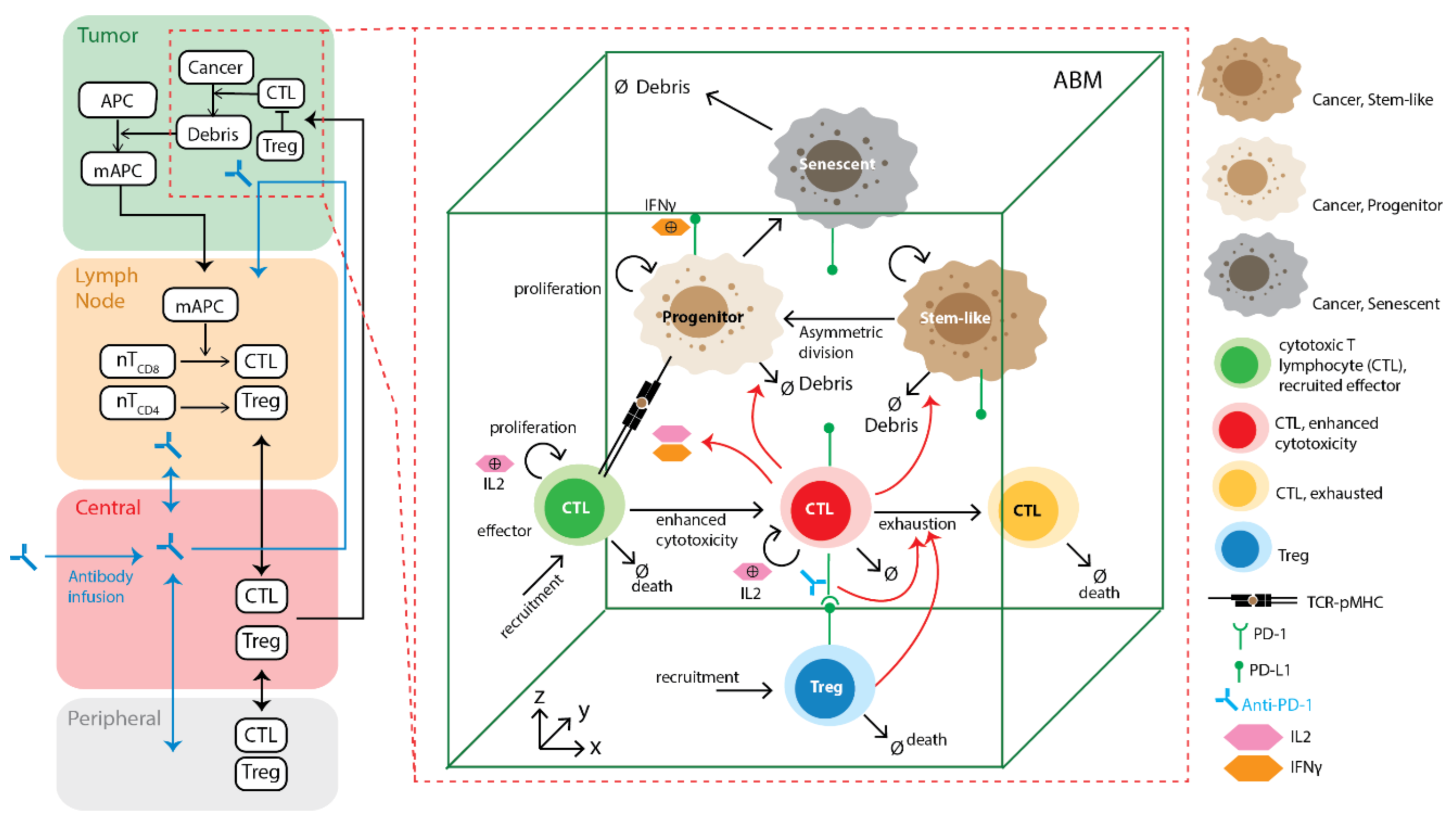
The spatially resolved ABM module of spQSP-IO allows us to study the impact on tumor dynamics from mechanisms that are stochastic and spatial in nature. In the current version of the ABM implemented within the spQSP-IO platform, we expand the cancer cell growth mechanism from our previous ABM model by including cancer stem-like cells (CSC), progenitor cells and senescent cells. The rules governing their proliferation are explained in the methods section. To assess how migration and proliferation of CSCs affect the spatial characteristics of the tumor, we performed simulations in which two parameters are varied: (i) one controlling the asymmetric division probability which determines the probability that a progenitor cell and a CSC are produced as daughter cells when a CSC divides, as opposed to two CSCs, and (ii) the probability of CSC movement at each time point.
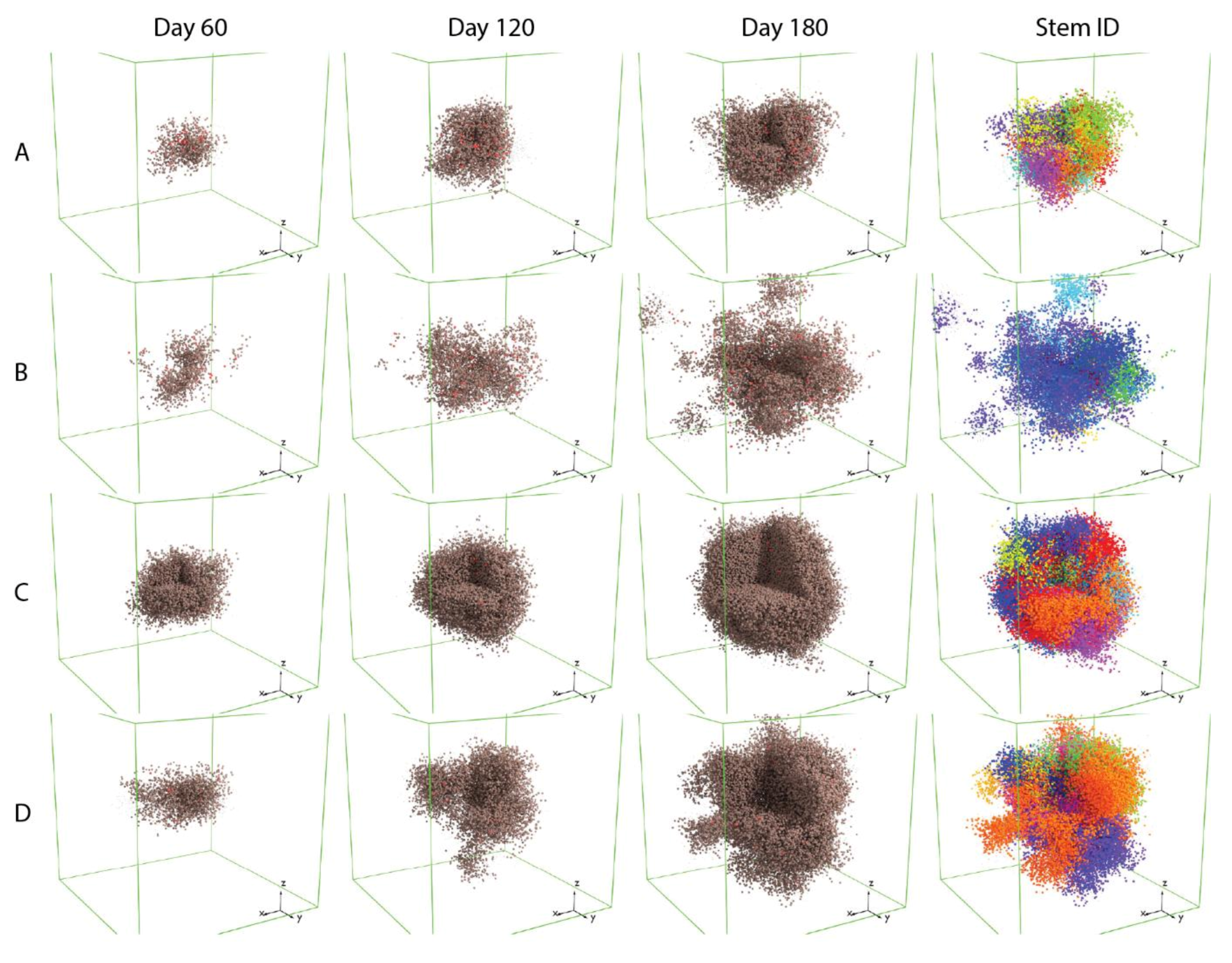
Figure 3 shows that these two mechanisms both contribute to the variability of the morphology of simulated tumors; the box volumes are 3 × 3 × 3 mm.
Simulations of tumor with CSC that have a low asymmetric division probability (Rows A and B) tend to grow more slowly compared to those with higher asymmetric division probability. This could result from spatial limitation imposed on the migration and proliferation of CSCs. When the asymmetric division probability is low, the tumor will consist of a higher fraction of CSCs, which are more likely to be surrounded by other cancer cell subtypes, making it difficult to find open neighboring voxels to move or proliferate into. Tumors in which CSCs have higher migration rates (Rows B and D), or higher movement probabilities, have more protrusions compared to those with CSCs that have a lower migration rate (Rows A and C). When we visualize cancer progenitor cells using colors based on their origin of CSC (sub-clones, or Stem ID), it appears that each of these protrusions is mainly composed of cells from the same sub-clone, indicating that the irregular shape of tumor is driven by single CSCs, and that the migration of these CSCs is leading the formation of protrusions.
Additionally, we observe that in simulations with a low asymmetric division probability and a high migration rate (Row B), the protrusions become detached from the mass of the tumor. This could be the result of two factors: first, because the rate of symmetric division of CSC is determined by tumor growth rate from the QSP parameter and is fixed across the four scenarios, the lower asymmetric division probability results in fewer progenitor cell created from CSC division and lower cancer cell density in the tumor, allowing CSC to migrate more freely and travel longer distances; secondly, the appearance of detachment can be a result of fewer cancer cells between the CSC core of the protrusion and the main tumor body. These observations can have clinical implications: reducing CSC proliferation might not be enough to improve the outcome of cancer. With the CSCs’ ability to produce progenitor daughter cells limited and their potential to divide symmetrically into stem-like cells unhindered, the morphology of tumor could become even more invasive. This could increase the difficulty of surgically removing the tumor, or even potentiate metastasis. A more detailed discussion regarding the role of maximum number of divisions for cancer cells in tumor morphology is included in
Supplementary Materials, Text S2 and Figures S1 and S2.
3.2. Simulating Patient Response to Nivolumab Treatment with spQSP-IO
With the incorporation of QSP model in the spQSP-IO platform, we can simulate tumor response to immunotherapies targeting immune checkpoint molecules with increased level of predictive power. The QSP model provides whole patient-level systems dynamics, including pharmacokinetics of drug, generation of cytotoxic and regulatory T lymphocytes in the tumor-draining lymph nodes, as well as transport of tumor neoantigens and immune cells through lymphatic and blood circulations. To verify the simulation result in the hybrid model, we replicated simulations performed in [
20] (shown in
Supplementary Materials, Figure S3). The overall dynamics at the whole patient scale closely resembles stand-alone version of the QSP model. In addition to the whole patient-scale dynamics captured by the QSP model, the ABM provides us with a spatial dimension to investigate how tumor grows and responds to treatment such as immune checkpoint inhibition.
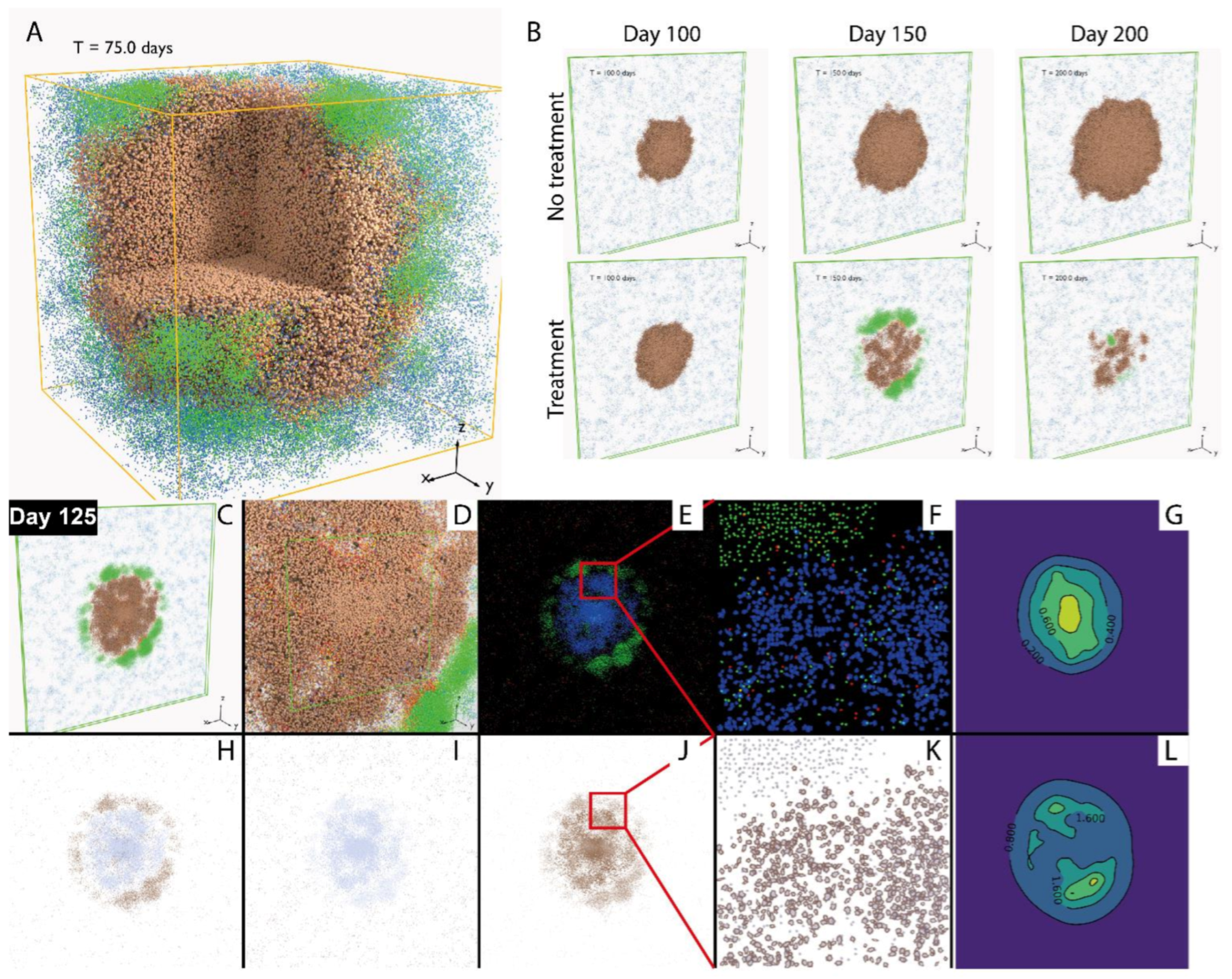
In
Figure 4A, the simulation includes one single solid tumor in a volume of 3 × 3 × 3 mm. The simulated tumor grows from the center of the volume, and as cancer cells die, tumor neoantigens are transported to tumor-draining lymph nodes by antigen presenting cells (APC), prompting the priming of naïve T cells and their differentiation and clonal expansion. In this process, cytotoxic and regulatory T cells are generated.
These lymphocytes travel through blood and are recruited to the tumor volume. Upon encountering the cancer cells, cytotoxic T cells further proliferate and exhibit an enhanced level of activity by producing IL-2 and IFNγ, while regulatory T cells negatively modulate their activity by inducing their exhaustion. By visualizing the cells in 3D, we can observe the spatial distribution of different types of cells in the tumor. In this case, since the entire tumor is represented in the spatial model, the weight of QSP tumor is set to 0.
In order to make more direct comparison with multiplex digital pathology image data which are becoming more available in cancer, we performed simulation in a volume in the shape of a flattened box of size 10 × 10 × 0.2 mm. This shape is chosen due to the computational constraint on the domain of simulation (2,500,000 voxels), while representing a larger region in the
x–
y directions.
Figure 4B illustrates how the simulated tumor appears in 3D, without treatment (upper row) and in response to anti-PD-1 treatment. A time-lapse movie displaying the dynamics of the tumor generated using a series of snapshots over the course of 200 days is included in the
Supplementary Materials (Supplement_section.mp4). In order to create results which are more visually comparable to immunohistochemistry and multiplex immunofluorescence, we generated 2D snapshots using a slice of the 3D simulation (
Figure 4C,D) mimicking the effect of those imaging techniques shown in
Figure 4E,F,H–K. Additionally, the concentrations of IL-2 and IFNγ are visualized in
Figure 4G,L. By visualizing the simulated tumor in 2D, virtual tissue slides are created, allowing qualitative and quantitative comparison with pathology data from real patients of various forms, including whole-slide images, biopsies and tissue microarray (TMA) data. Such comparisons could use methods of spatial statistics and machine learning performed on both pathology samples and simulated images.
3.3. Parameter Sensitivity Analysis in spQSP-IO
QSP models, such as the one we incorporated in our hybrid model, can have a large number of equations and parameters. On top of that, spQSP-IO model has an ABM module with additional parameters governing rules of interactions between factors of higher granularity. Some of these parameters are directly calculated from the parameters defined in the QSP model or adopted from other experimental or modeling literature; other parameters cannot be accurately determined due to naturally arising variability among the population and a lack of direct measurement. In those cases, we only define physiologically plausible ranges for their values, and assess the uncertainty they introduce to the system and the sensitivity of key model output to these mechanisms.
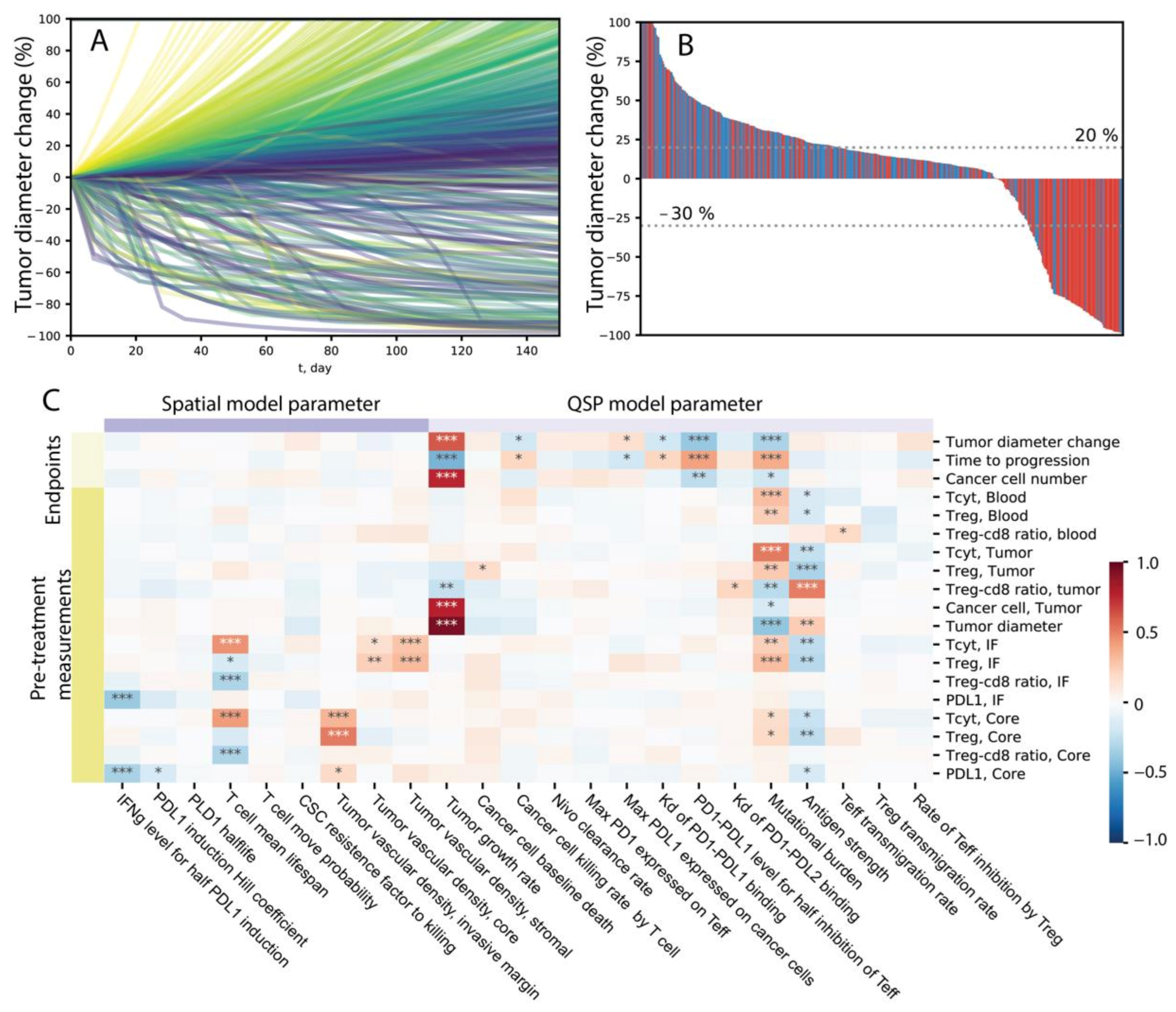
To more efficiently cover the high-dimensional parameter space, we use LHS to sample the parameters varied for sensitivity analysis. The batch simulation performed here can be considered as a virtual clinical trial, where single simulations produced using each parameter combination represent the dynamics of one virtual patient in response to anti-PD1 treatment. We plotted the relative tumor diameter change (compared to the tumor diameter before the first infusion time point) in the form of a spider plot in
Figure 5A.
Here, each line represents one patient, and the color corresponds to the value of tumor vascular density in the invasive front volume of the ABM, which is one of the many parameters we are varying among the virtual cohort. A range of dynamics is produced in these simulations, including progressive disease cases where tumor diameter increases to more than 20% over time, stable disease cases where the tumor diameter change remain between −30% and 20%, and partial or complete responses where tumor shrinks more than 30%. In some cases, the tumor diameter increases initially but ends up with tumor responding to treatment, creating a pseudo-progression scenario. The temporal dynamics of other variables from 100 randomly selected cases are shown in
Supplementary Figures S4–S7. In
Figure 5B, the best response of each patient is calculated using the minimum value of relative diameter change starting from 8 weeks after the first infusion and displayed in a waterfall plot. The colors correspond to the value of tumor mutational burden in each patient, where red indicates that patient has a mutational burden higher than the median value within the virtual cohort. We can see that 19% of patients responded to the treatment (PR/CR), and among the responders, the majority has higher than median tumor mutational burden.
We calculated PRCC to evaluate global sensitivity of various model outputs to parameters included in the analysis. Model outputs include direct measurement of species in the QSP and ABM modules, as well as simulated clinical endpoints such as relative diameter change, which are composite variables derived as functions of multiple variables. The results are shown in the heatmap in
Figure 5C, where the darkness of red/blue indicates the level of positive/negative correlation and the number of asterisks indicates the level of significance. From
Figure 5C, we can see that various mechanisms are correlated with larger tumor diameter reduction after treatment, including high mutational burden, higher PD-L1/PD-1 bond threshold for T cell exhaustion, lower PD-L1/PD-1 affinity, lower maximum PD-L1 expression level on cancer cells, higher affinity of the antibody, lower maximum PD-1 expression on T cells, lower antibody clearance rate, lower tumor growth rate, and higher T cell lifespan. Additionally, higher PD-L1/PD-1 bond threshold for T cell exhaustion, lower affinity between PD-L1 and PD-1 binding, and higher vascular density in the invasive front are correlated with better treatment outcomes in that it takes longer time for the tumor to reach the designation of progressive disease. Based on these sensitivity analysis results, it is possible to identify mechanisms that could synergize with anti-PD-1 treatment to achieve higher efficacy when modulated by therapeutic interventions.
3.4. Assessment of Impact of Tumor Vascular Density and Its Heterogeneity Using spQSP-IO
One aspect of the tumor–immune interaction that we are able to explore using the spQSP-IO platform in addition to non-spatial QSP model is that with the ABM, we can assess how various mechanisms affect the system behavior in a spatially heterogeneous manner. To illustrate this, we employed a version of the model with two volumes representing tumor invasive front and tumor core, respectively and varied the vascular density parameter in these two volumes. This parameter is chosen based on observations from experimental and clinical data indicating that there are differences in tumor vasculature and blood perfusion among these regions [
63,
64,
65]. In the simulations, we tested four different scenarios, with vascular density increased by 10× in tumor core, on the stromal side of the invasive front, on the tumor side of the invasive front, and one negative control where vascular density is kept the same in all ABM regions. Each parameter combination is simulated with 10 replications to account for the stochasticity from the ABM.
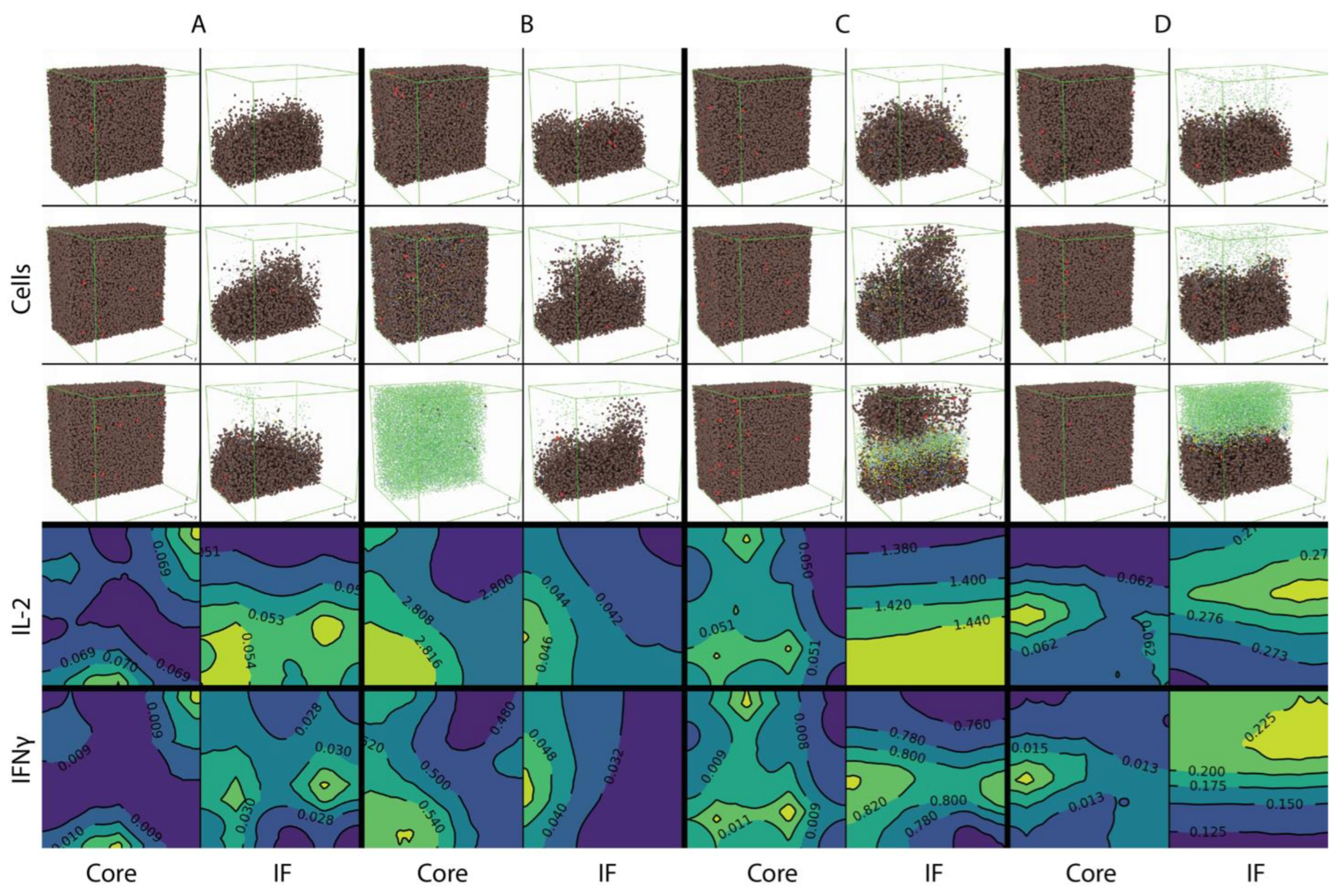
The 3D distribution of cells and cytokine concentration of the four cases are shown in
Figure 6.
Group A, B, C, D correspond to the four cases, respectively: baseline vascular density; increased vascular density in tumor core; increased vascular density in the tumor side of the invasive front; and increased vascular density in the stromal side of the invasive front. The spatial distributions of cells in each case are shown at three time points: pretreatment (day 20 of simulation, or 8 days before first dose), early treatment (day 50 of simulation, or 22 days after first dose) and after approximately half a year (day 200, or 172 days into treatment) of nivolumab treatment. Cells in the front half of the volume are not visualized to make the section through the middle of the volume is visible.
Figure 6 shows that with the baseline vascular density (A), both effector CD8+ and Tregs are recruited at low rates and remain sparsely distributed through the course of treatment. When the vascular density is increased in the tumor core (B), effector CD8+ T cells are able to kill most of the cancer cells over the course of treatment, while the cancer cell distribution at the invasive front remains intact. When the vascular density is increased in the tumor side at the invasive front (C), T cells are able to penetrate the boundary of the tumor and break the barrier between tumor and stromal tissues by killing cancer cells. When vascular density is increased at the stromal side of the invasive front (D), T cell density is much higher compared with the other cases after treatment starts; however, T cells mostly remain outside of the tumor and are not able to penetrate the invasive front.
The pretreatment cytokine concentrations are also shown. Here, a cross section at the location y = 0.5 mm is taken and the contours of cytokine concentration in that plane are plotted. IL-2 concentration is low in compartments where vascular density is at baseline; when vascular density is increased, IL-2 concentration also increases, especially when T cells are recruited closer to cancer cells (case B tumor core and case C invasive front), in which case T cells have higher chance of being further activated by tumor neoantigens. The release of IFNγ by CD8+ T cells is similarly affected by location of T cell entry point; however, its concentration is higher at the invasive front as shown in case B, which has higher IFNγ concentration compared to the core of tumor from case A.
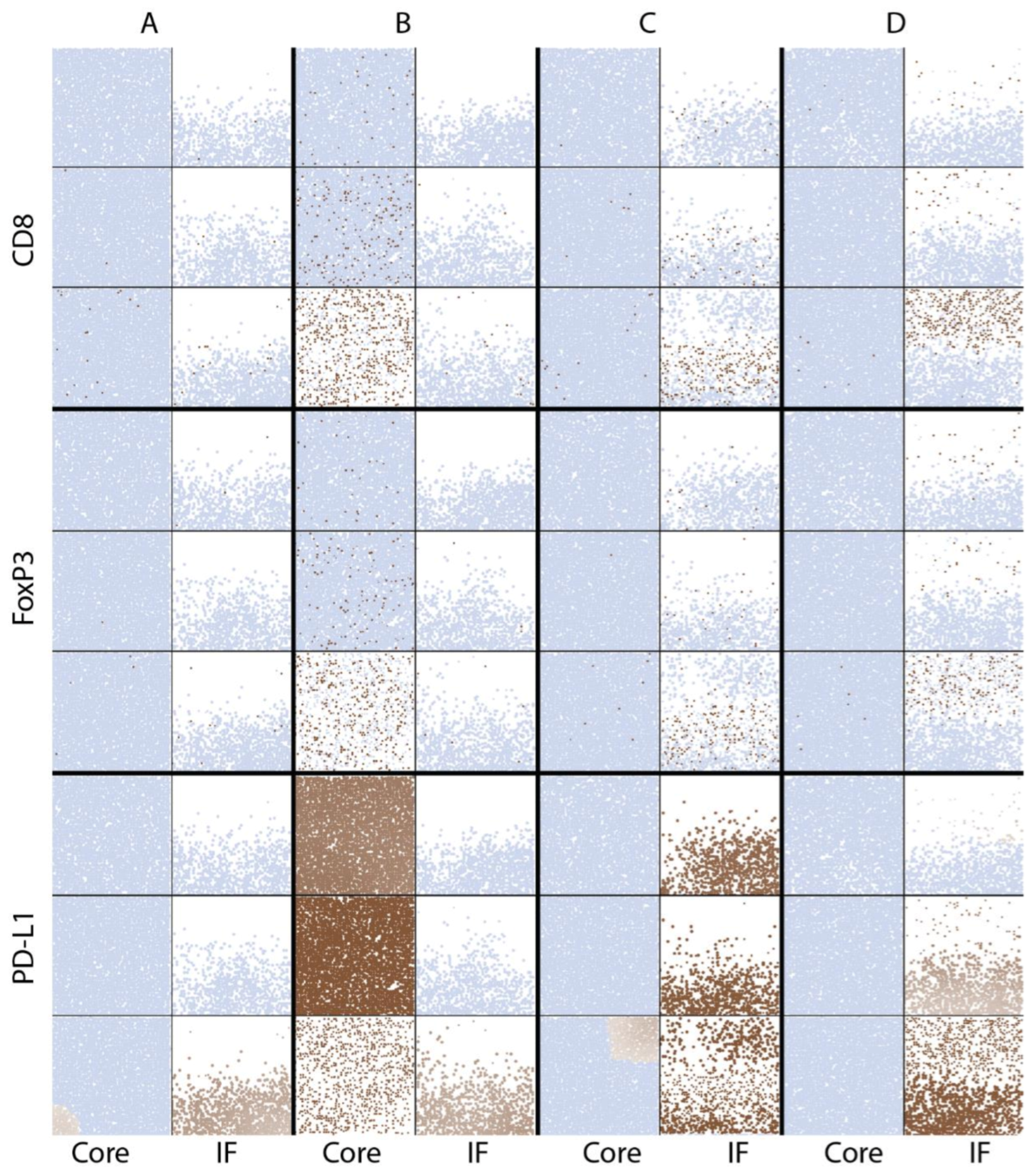
The cell distributions are also visualized in simulated immunohistochemistry, IHC, in
Figure 7.
CD8 and FoxP3 label the location of cytotoxic and regulatory T cells, respectively; their distribution is similar in each case. In the baseline vascular density case (A), T cells are sparsely distributed in both tumor core and the invasive front. When vascular density is increased in the tumor core (B), T cell densities increase over time until cancer cells are cleared in that volume. When tumor vascular density increases within the tumor at the invasive front (C), T cells and cancer cells are better mixed and killing result in a more irregular boundary compared to the compartmentalized distribution when vascular density increases outside of the tumor (D).
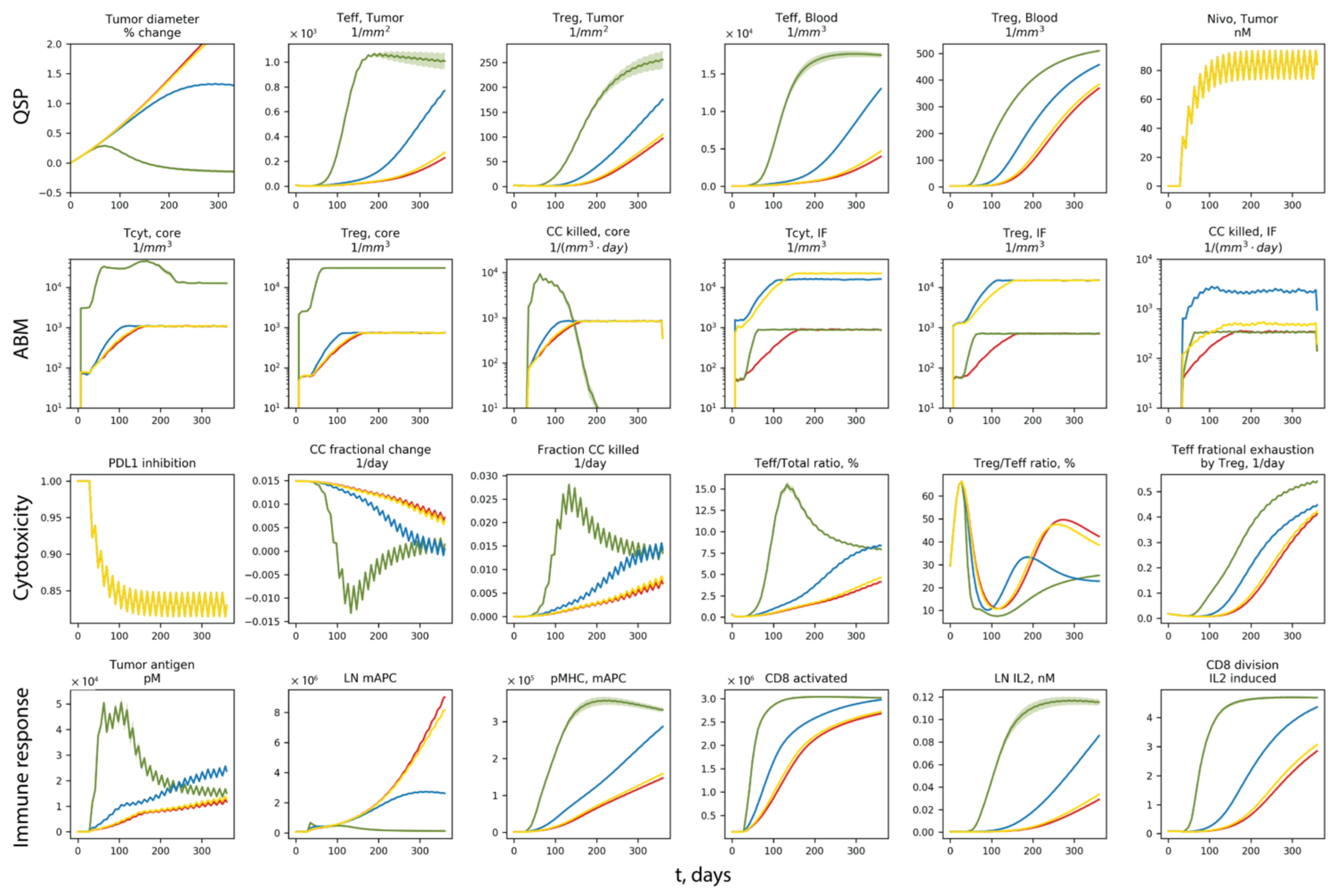
To further investigate how the differences in vascular densities contribute to different patient-scale response to nivolumab treatment, the temporal dynamics of the system are shown in
Figure 8.
Red, green, blue and yellow represent the same cases of A, B, C and D as in
Figure 6 and
Figure 7. For each case, the solid line is the average of 10 replications, and the band represents the standard error of the mean. In
Figure 8, Row 1 panels are dynamics from the QSP model; Row 2 panels are dynamics in the ABM volume of tumor core and the invasive front; Row 3 and 4 are dynamics related to anti-cancer cytotoxicity and immune response, respectively. From Row 1, we can see that tumor is responding to nivolumab treatment when the density of tumor vasculature increases in the core; when vascular density increases in the tumor side of the invasive front, tumor growth is reduced, but still progresses; increasing the vascular density in the stromal tissue at the invasive front is not effective. The responsiveness coincides with the rapid increase in effector CD8+ T cell (Teff) and Tregs in case B (green), especially Teff density in the tumor compartment. Since all QSP model parameters are the same in case A–D, the differences in the ABM are driving the diverging behaviors. From Row 2, case B (green) has increased Teff recruitment to the tumor, and a much higher cancer cell killing rate in the tumor core. Although the killing rate in case B drops when cancer cells are cleared in that volume after several months of treatment, the Teff concentration in the blood remains at high levels. Despite the lack of direct increase in the vascular density in the QSP model, the increase in T cell blood concentration results in higher recruitment to the tumor compartment, which leads to higher Teff ratio in the tumor and a boost to cancer cell killing (with the presence of nivolumab). The boost in cancer cell killing in both the ABM and QSP contributes to a boost in Teff priming and expansion in the LN by providing more tumor neoantigens, creating a positive feedback between tumor neoantigen supply and anti-cancer effector CD8+ T cell generation.
3.5. Identifying Biomarker for Different Simulated Clinical Endpoints Using spQSP-IO
We observed that parameter input to the spQSP model, such as the vascular density in the invasive front of tumor, can affect tissue pathology characteristics (
Figure 7) as well as trajectories of tumor development at the same time. Because model parameter values are not always directly measurable in patients, it is of substantial clinical value if the correlation between measurements of patient samples and responsiveness to treatment can be identified, since those biomarkers found to correlate with certain treatment outcomes can have predictive values to guide cancer treatment or help design clinical trials with improved inclusion criteria.
To illustrate how data from virtual clinical trials based on spQSP-IO platform can be facilitated by biomarker analysis, we chose simulated clinical endpoints of three different types: first, a continuous endpoint where we use tumor diameter at the end of treatment. Second, a categorical endpoint where we use responsiveness to the treatment. Using RECIST criteria, we categorize virtual patients into responder (PR/CR, relative diameter change ≤ −30%) and non-responder (SD/PD, relative diameter change > −30%). Third, a time-to-event, or survival type endpoint where we use time-to-progression. For each endpoint, we use the corresponding multivariate regression model to perform variable selection procedure from a pool of potential pretreatment biomarkers. The biomarkers which significantly contribute to the multivariate regression are shown in
Figure 9,
Figure 10 and
Figure 11, respectively for the three endpoints we chose. For each of the selected pretreatment measurement, and parameters that ranked high from the PRCC analysis to each endpoint (
Figure 5), we dichotomized the virtual cohort into two subgroups using the median value of the biomarker, and analyzed the efficacy in each subgroup.
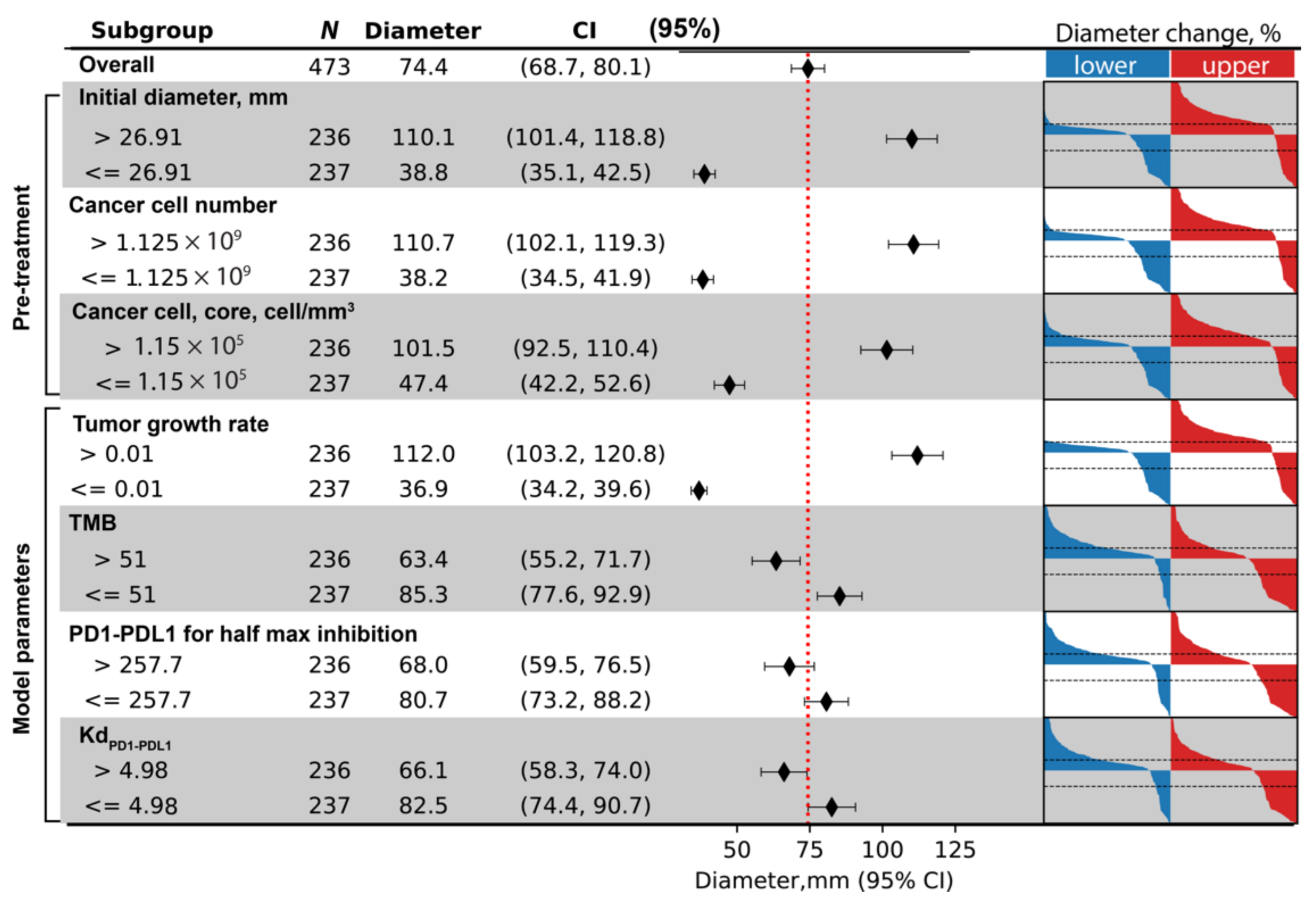
For tumor diameter at the end of treatment, three measurements are found to contribute significantly to its prediction: initial tumor size, total cancer cell number, and the density of cancer cells in tumor core. For each of the selected biomarkers, the average diameter among the two subgroups, as well as 95% confidence intervals (calculated based on t-distribution) are calculated and shown in the forest plot, with the vertical dashed red line indicating overall average tumor diameter). On the right-hand side of the figure, waterfall plot is generated for both subgroups, with blue and red representing the lower and upper half of virtual patients with regard to each biomarker. The three pretreatment measurements indicate that tumors of larger size and higher density of cancer cells correlate with larger post-treatment tumor diameter. In addition to the three pretreatment measurements, we also included parameters ranked highest for sensitivity of tumor diameter change: tumor growth rate; tumor mutational burden; half maximum inhibition concentration of PD1–PD-L1 for Teff inhibition; and Kd of PD1–PD-L1 binding. The results are shown in
Figure 9.
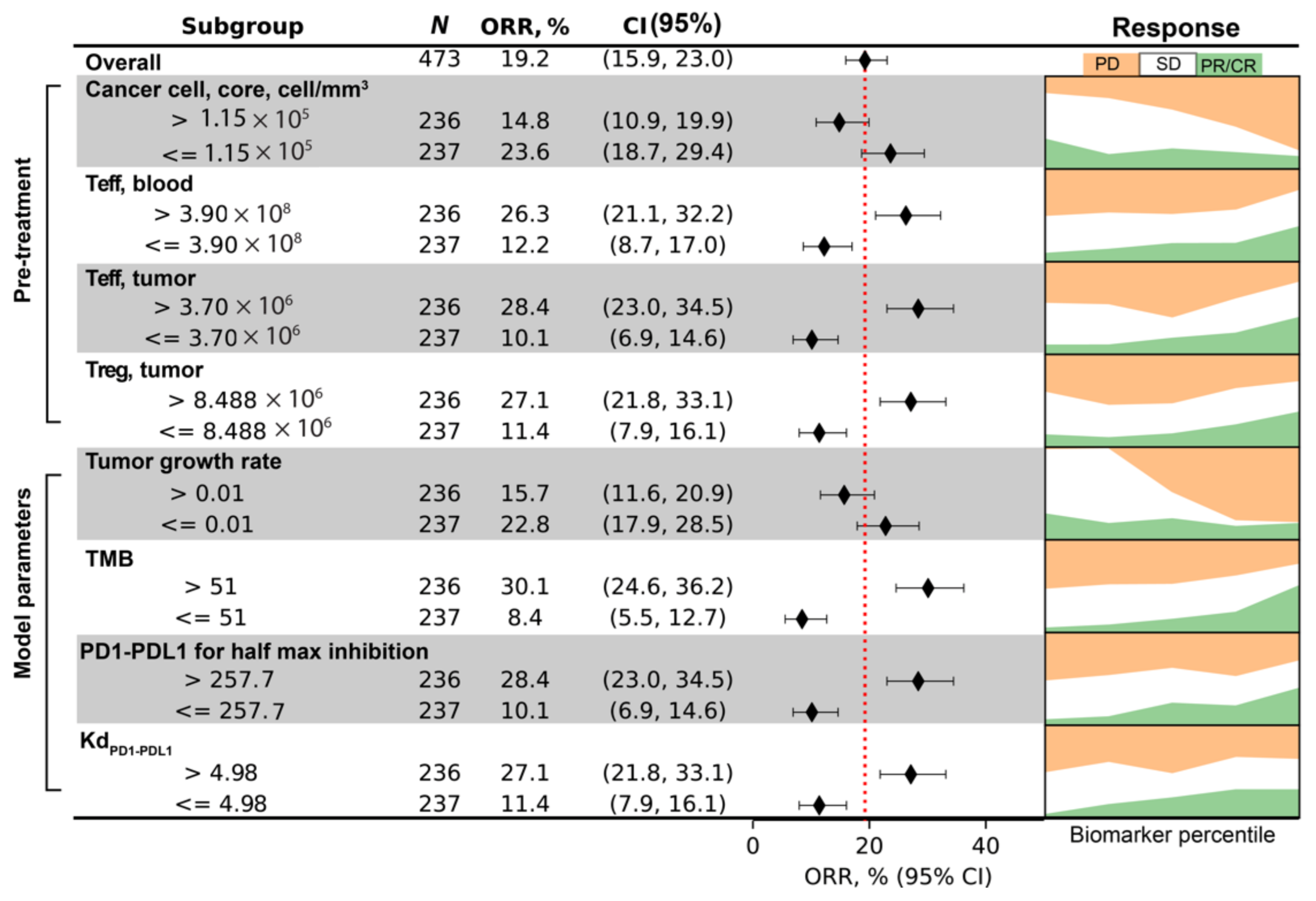
For the endpoint of responder/non-responder binary designation, four biomarkers are found to significantly contribute to prediction of responsiveness of virtual patients: cancer cell density in the tumor core; blood concentration of Teff; tumor Teff and Treg cell numbers. To examine how the values of these biomarkers correlate with responsiveness, we dichotomize the virtual cohort into two subgroups, and compute the objective response rate (ORR) and its 95% confidence intervals of each subgroup to generate the forest plot (
Figure 10). The binomial proportion confidence interval is calculated using Wilson score interval. The dashed red line indicates ORR of the overall virtual cohort, which is about 19%. On the right-hand side, the fraction of virtual patients in each RECIST response category after treatment is plotted against the percentile of the biomarker values. The results indicate that tumors with higher cancer cell density in tumor core are less likely to respond to anti-PD1 treatment, while higher number of Teff and Treg correlates with responsiveness. We also included the four parameters correlated with tumor diameter change, and results are shown in
Figure 10.
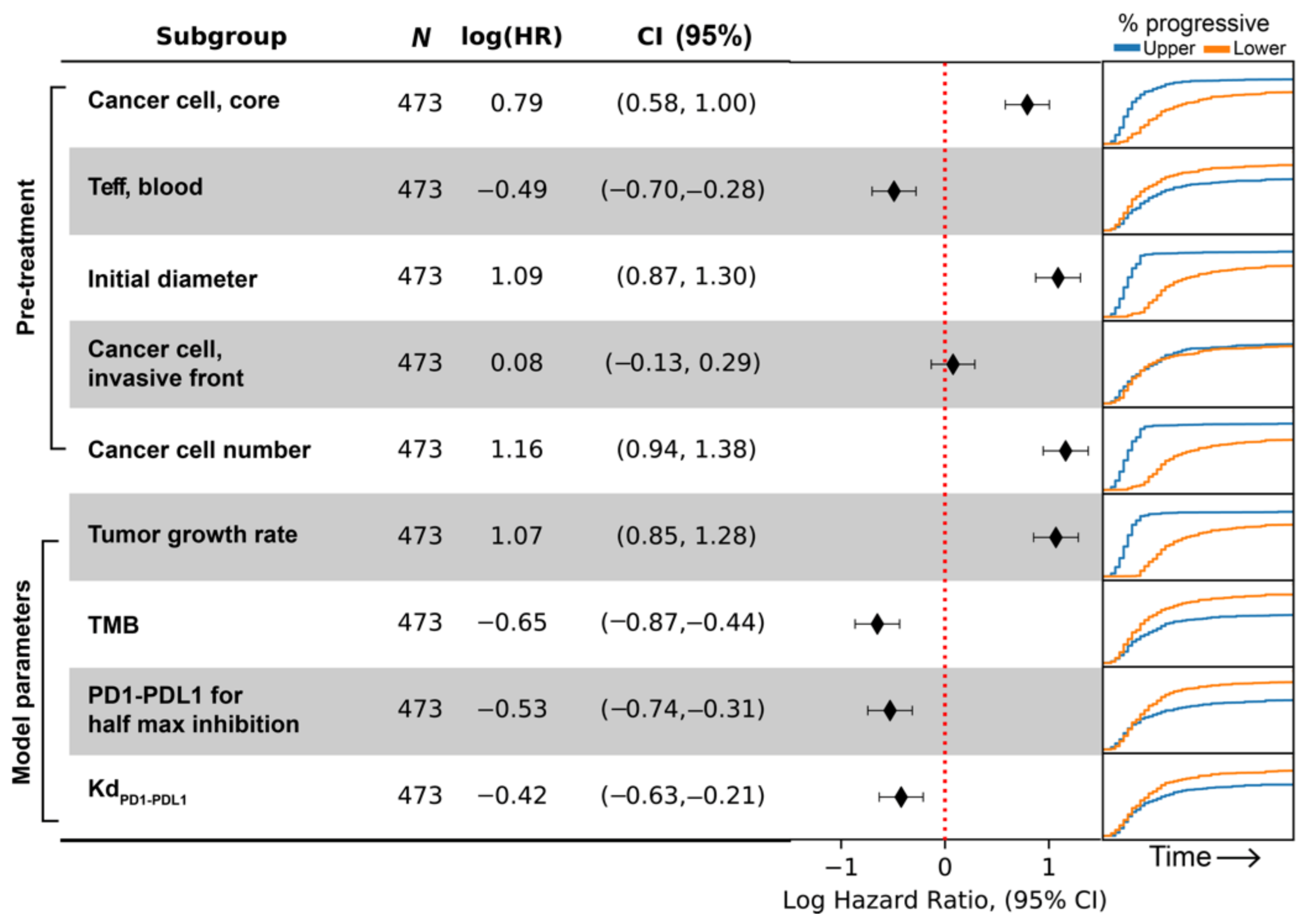
For the endpoint concerning time-to-progression, the biomarkers selected are pretreatment cancer cell density in tumor core, blood Teff count, initial tumor diameter, cancer cell density in the invasive front, and total cancer cell count. Similar to the previous two endpoints, we divide the virtual cohort into two subgroups with median value of each biomarker, and calculate hazard ratio and its confidence intervals by fitting a single variable (i.e., subgroup designation) Cox proportional hazard model to time-to-progression data. The results are shown as forest plot in
Figure 11, with positive log-hazard ratio indicating an increased risk of progression in the subgroup with higher value for the corresponding biomarker. Additionally, the Kaplan–Meier graph showing the percentage of virtual patients with progressive disease over time is shown on the right-hand side of the forest plot. In
Figure 11, the results suggest that patients with more overall cancer cells and larger initial tumor size are likely to progress earlier, whereas pretreatment Teff in the blood is associated with lower progression.
Despite being selected by the multivariate regression model in the resampling process, cancer cell densities in the invasive front do not seem to divide the overall population into two subgroups with different rate of progression. The reason could be that this covariate only works in combination with other biomarkers, and they add predictive power to certain strata determined by these other biomarkers.
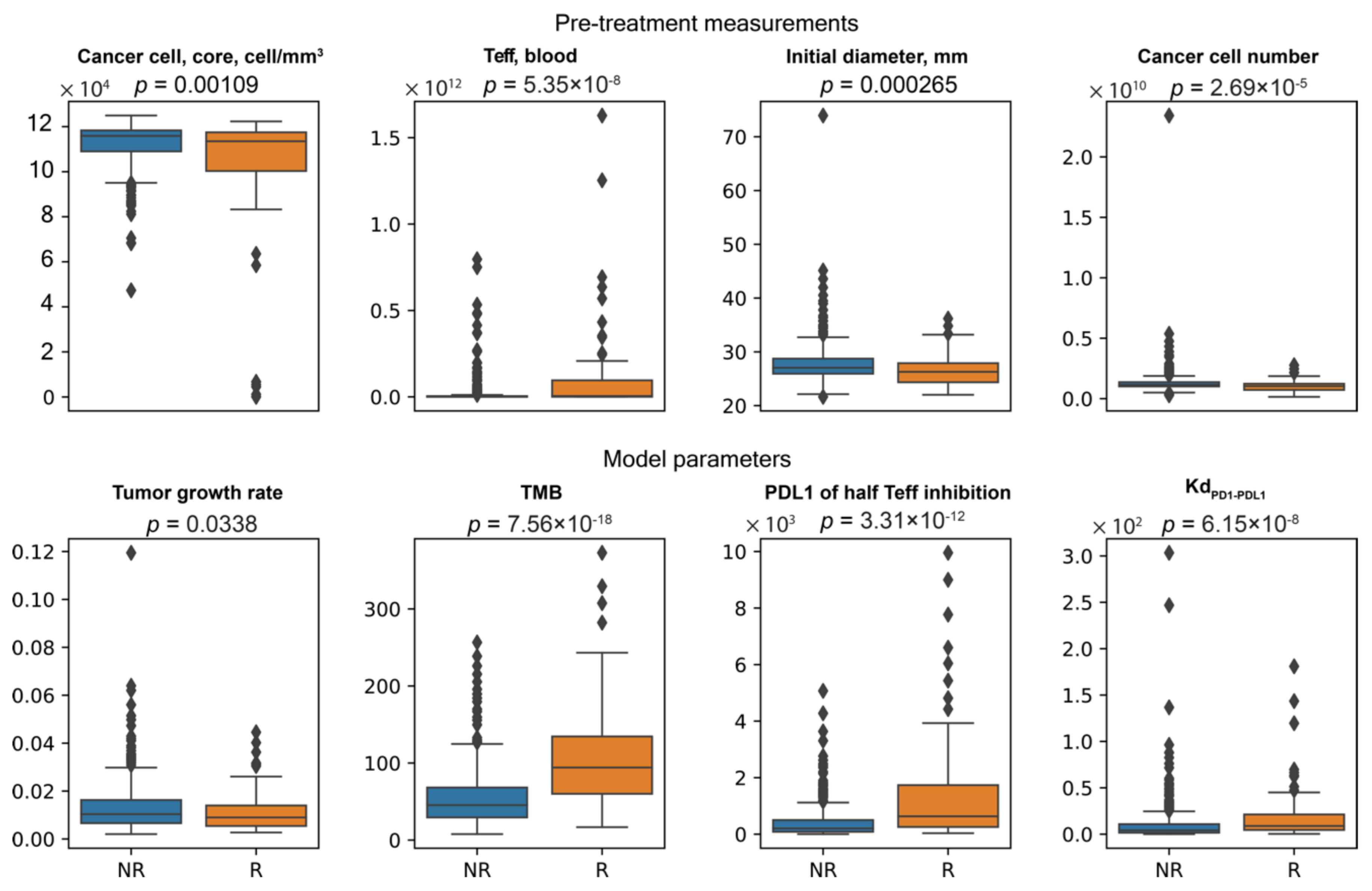
Since all three clinical endpoints could be potentially important for determining whether or not a patient is likely to benefit from the treatment, we created a list including biomarkers selected from the three individual analyses shown in
Figure 9,
Figure 10 and
Figure 11, and tested whether or not their values are significantly different in responder and non-responder groups using the non-parametric Mann–Whitney
U test (
Figure 12). The result shows that except for the model parameter tumor growth rate (which has a
p-value of 0.03, indicating only weak evidence of significance), the values of the biomarkers differ significantly in responder sub-population of the cohort compared to those in the non-responder sub-population. The difference of significance regarding the correlations between tumor growth rate and responsiveness observed in the sensitivity analysis (
Figure 5C) and biomarker analysis (
Figure 12) could be attributed to the method used in analysis: in the sensitivity analysis, the correlation is calculated based on the rank of tumor size change, whereas in the biomarker analysis, the binary designation of responder/non-responder leaves out the information about the degree of progression or recess each patient has. This finding suggests that the intensity of these selected pretreatment measurements and mechanisms can potentially serve as predictive biomarkers when the decision is made regarding whether or not anti-PD-1 therapy is likely to benefit the patient. Note that the analyses presented in this study serve as an illustration of the capabilities of the spQSP-IO platform, rather than a systematic comparison with clinical and experimental data. Such comparisons and quantitative validation of the model remain to be explored in future studies.
4. Discussion
In this study, we present a computational model platform, spQSP-IO, which incorporates both a QSP model with its capacity to capture the whole patient-scale system dynamics and an ABM with its spatial resolution to account for spatial tumor heterogeneity. Using this platform, we constructed a hybrid model of non-small-cell lung cancer, with both QSP and ABM components derived from previous published models. We demonstrate the functionality of the model in several different ways. With the spatial and lineage granularity explicitly represented in the ABM, we are able to interrogate the in silico system for the role of cancer stem-like cell in driving the formation of protrusions in tumor shape, and how morphology of tumor is affected by the division and migration properties of these cells. With anti-cancer immune response and immune checkpoint inhibitor PK/PD, modeled with the QSP model, its simulations are able to recapitulate the complex interactions in the tumor microenvironment, especially tumor growth in the context of anti-tumor immunosurveillance and its response to immunotherapy. These spatial outputs from model simulations can be compared with image-based pathology data with cell types and selected molecular markers visualized (e.g., with immunohistochemistry and immunofluorescence), allowing additional dimensions of model validation and potentially more accurate prediction of responsiveness of individual patient to proposed treatment strategies. By allowing variability in parameter values and initial conditions to represent a cohort of simulated patients, virtual clinical trials can be carried out using models derived from this platform. The simulation results from virtual clinical trials can also facilitate the discovery of biomarkers measured from pretreatment samples of patient which are correlated with various treatment endpoints.
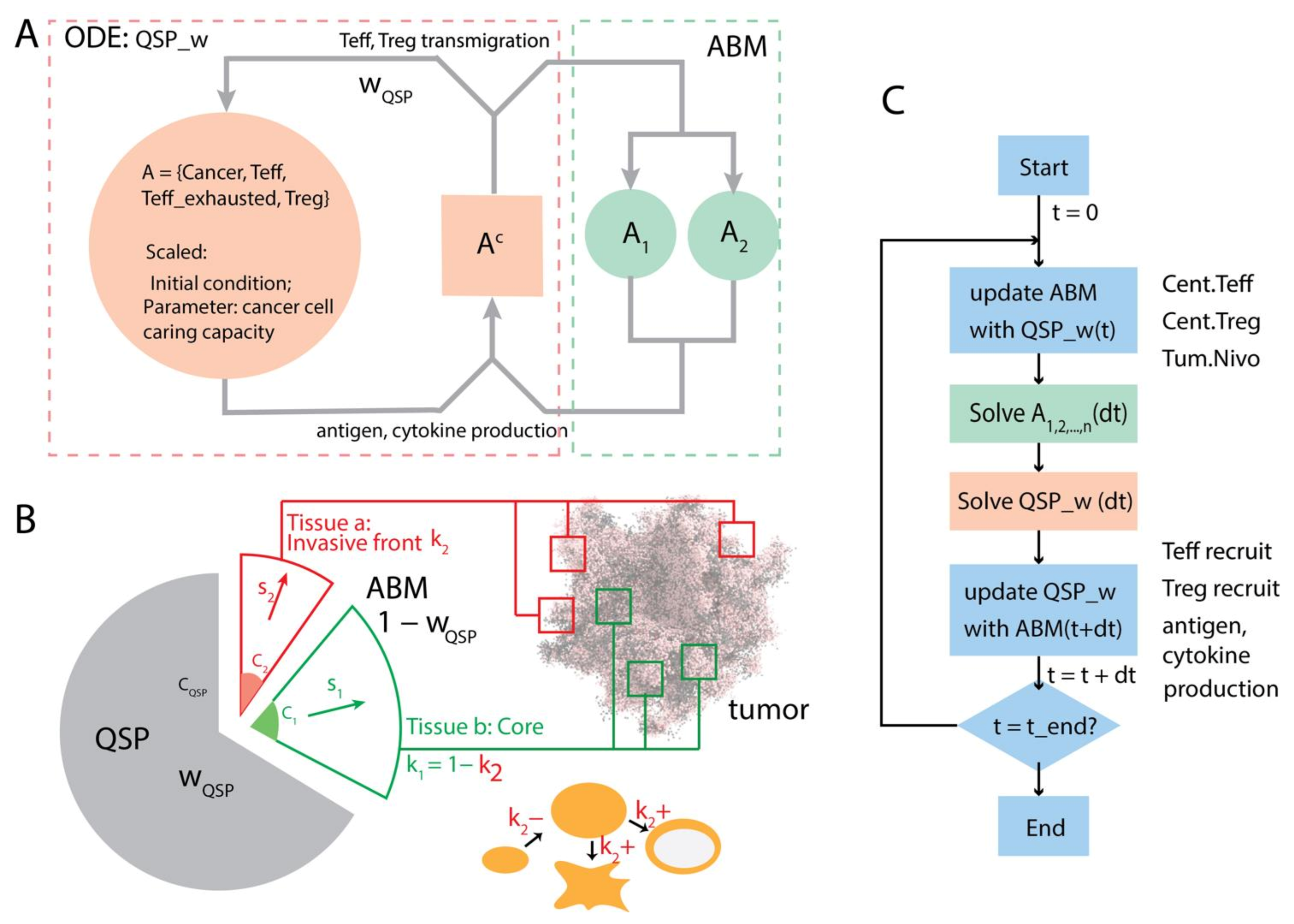
In order to take advantage of both QSP and ABM modeling and capture whole patient-scale dynamics as well as spatially resolved characteristics of the tumor at the same time, we coupled these two model types together, with a proportion of the tumor represented using ABM while the remaining part of tumor and the rest of the system represented with ODE-based QSP module. The coupling is achieved through transport of cellular and molecular components through blood and lymphatic circulation: tumor neoantigens resulting from cancer cell death in the tumor ABM are transported to the LN, and immune cells produced in the LN in response to tumor neoantigens are transported to tumor vasculature and recruited to the tumor. Due to limits in computational resources, it is unfeasible to simulate a substantial portion of a large tumor—those that could be more than ten centimeters in diameter and include more than hundreds of billions of cells—in full detail, particularly because the ultimate goal is to use the model to simulate tumor growth in many virtual patients as part of a virtual clinical trial. However, if the tumor volume represented in the ABM is small, the impact of dynamics in the spatial model—some may not be fully captured by the non-spatial QSP model—will be underrepresented in the full hybrid model.
Here, we have two design choices. In the first option, the ABM simulation only accounts for a fraction of tumor equal to the volume it explicitly represents. In this scenario, one assumes that the overall dynamics is mostly driven by the QSP model and the behavior of the spatial ABM is dependent on input from the QSP model. The shortfall of this option is that the output from the ABM and its contribution to the overall behavior of the system become insignificant. In other words, any mechanisms specific to the spatial model, which could potentially have a substantial impact on the emergence of heterogeneity and various other characteristics of the tumor, will not be detectable during parameter sensitivity analysis performed based on the patient-scale outputs. In a second option, the simulated volume, along with the exchange happening at the interface of the spatial and non-spatial models, can be scaled up to represent a fraction of the tumor which is larger than the volume under simulation. In this scenario, which is what we chose to implement in the current version of the platform, the spatial simulation account for a proportion of the tumor, where the proportion is controlled with a weight parameter. If such condition is satisfied that the dynamics of the tumor compartment represented by ABM and QSP model are equivalent, the overall system behavior should not change when this weight parameter is varied. The actual simulated volume is then scaled up to match the weight assigned to the ABM simulation. This method also has its limitations: on the one hand, the stochasticity generated in the relatively small simulated volume could be amplified when scaled up along with the variables tracked in the volume, resulting in deviation in simulated results compared to the situation where a larger volume is simulated; on the other hand, the limited volume may also not fully recapitulate the level of spatial heterogeneity of a larger volume. To address this issue, with the total simulated volume as a constraint, one can leverage the available resource to two dimensions at the cost of the third dimension, or use sampling-based method to simulate multiple stochastic realizations of smaller ROIs from different part of the tumor.
Correspondingly, in this study, we explored two different methods of representing the tumor volume in ABM. First, we use a flattened box to represent a volume with relatively large area (10 × 10 mm) but shallow depth (0.2 mm) in an attempt to better capture the spatial characteristics in two dimensions at the cost of the third. Secondly, similar to the concept of stratified sampling, we use two separate volumes to track the invasive front and core of the tumor, respectively. Yet, both methods have their limitations: for the flattened box volume case, 1 cm2 area is comparable to whole-slide image data, but still only large enough to host a section of a small-sized tumor, and the small depth used in the simulation might skew the result toward a 2D simulation. In the case of the multi-volume simulations, in order to keep the invasive front volume at the invasive front of the tumor, the window is shifted towards the normal tissue when cancer cell number increases so that the volume is not filled with the growing tumor, and towards the tumor side when cancer cell number drops to avoid eliminating cancer cells at the invasive front entirely. In the latter case, new cells are introduced to populate the opening space created after shifting, and the arrangement of these cells can bring bias to the simulation. To strike a balance between better capturing the spatial heterogeneity and maintaining adequate computational performance, improvement in the coupling method of the ABM and QSP modules need to be made in future developments of such hybrid models.
In addition to the aforementioned limitations of using a smaller domain for simulation of spatial tumor dynamics, another potential source of inconsistency lies in the differences between rules governing the ABM tumor behavior and the equations governing the species in the tumor compartment of the QSP model. During model construction, when considering which cell types and mechanisms are included, the QSP model is used as the reference for the ABM design. In this process, the major components of both models are well aligned, and parameters in the ABM is converted from the corresponding QSP parameters whenever possible. However, to better utilize the spatial resolution and higher granularity of the ABM, some rules in the ABM are defined in more details compared to the QSP equations. For example, because individual cancer cells are tracked, we are able to define different subtypes of cancer cells, including stem-like cells, progenitor cells and senescent cells, with information such as lineage and number of divisions preserved during the simulation. Such differences between the ABM and QSP model could potentially result in discrepancies between their respective simulation outcomes, which may create difficulties in interpreting the results and making predictions with the model. However, this also creates an opportunity to evaluate the impact of those mechanisms specific to the spatial model on tumor growth and responsiveness to treatment. If certain mechanisms turn out to be important to the overall dynamics and are not sufficiently accounted for in the QSP model, adjustments should be made to the non-spatial model so that such discrepancies are removed from the hybrid model.
One important feature of spQSP-IO is that it has a spatially resolved representation of tumor compartment, which allows the model output to be compared with pathology imaging data (digital pathology) collected from patients as well as spatial transcriptomics, potentially enhancing the model prediction power and making it patient-specific. Such comparisons can be carried out at different stages of model development: the imaging data, such as pretreatment biopsies, can be used as a guide for setting up initial conditions for simulations and serve as direct inputs, or they can be used during the model validation stage and compared with emerging spatial patterns from the simulation. Additionally, the comparison can be performed in different manners: qualitative, quantitative, or exact mapping. Exact mapping is only applicable for setting up initial conditions for spatial simulation, and even in those cases, because the pathology images are 2D, extrapolation techniques would be required for setting up cell distributions in a 3D domain. Quantitative comparison between imaging and simulations is the most desirable method. In previous digital pathology studies, researchers have discussed methods of using spatial statistical analysis to quantitatively describe characteristics and heterogeneity of the tumor microenvironment. These quantification methods can range from simple density of different types of cell and variability of local density, spatial point process model-based analysis of clustering pattern, to spatial relationships between multiple cell types such as degree of colocalization and Shannon’s spatial entropy [
36,
38]. When combined, these measurements can serve as reference points for spatial simulations, where the same quantification methods can be applied to the simulation output, allowing a direct and quantitative comparison of spatial characteristics without losing generality of variations in the exact spatial arrangement of cells. The flexibility of the setup of the simulation domain also makes it more intuitive when comparing spQSP-IO model to imaging data of different varieties: depending on the type of data available, whether they are whole-slide images, tissue microarrays or biopsies, the number, shape and size of the ABM tumor simulation can be adjusted so that they are more in line with the actual data.
As a platform based on mechanistic models, when applied to virtual clinical trials, three categories of variables involved in the simulations bear particular significance: first, variables and their combinations that can help interpret patient clinical endpoints, such as tumor diameter or responsiveness to treatment; secondly, variables which mechanistically drive the dynamics towards different outcomes, such as rate parameters governing various biological processes; and thirdly, variables which can potentially be measured in patients and may help predict trial endpoints, such as blood concentration of cells and cytokines, or density of certain cells and concentrations of molecular species in the tumor. The second and the third category can have overlaps; for example, tumor mutational burden is a parameter input to the model and is also a measurable biomarker. Using parameter sensitivity analysis, one can identify correlation between the first two categories of variables. In sensitivity analysis, a subset of parameters is chosen and varied in a physiologically plausible range to perform batch simulations, and correlation is assessed to determine how the endpoint readouts are determined by the mechanisms controlled by these parameters. This analysis is useful in identifying potential targets for therapeutic intervention, since the terms involving these parameters usually represent rates of biological processes, and changes in the parameter value correspond to modulation of those processes. In this study, we use PRCC to assess the correlation between parameter values and model outcomes. This method is limited in that it may not be very effective in detecting non-monotonic relationships. To allow more flexibility in such analysis, future studies can explore other sensitivity techniques which can identify non-monotonic relationships between parameter input and simulation outcomes to better understand the role of the corresponding mechanisms.
Assessment of relationships between variables from the first and the third category, i.e., between treatment endpoints and biomarkers, also has tremendous clinical value. In this study, we demonstrated the methods of performing biomarker analysis on different types of outcomes, including continuous, categorical, and time-to-event endpoints. However, in those analyses, we did not take into account possible nonlinear effect of biomarkers or interactions between them. Additionally, during the example of subgroup analysis, we use median value of each parameter/biomarker to dichotomize the virtual cohort, which can result in suboptimal grouping. In future studies, biomarker and subgroup analysis can be further strengthened by using statistical models with nonlinear and interaction terms, and also more sophisticated subgroup identification methods [
72].
With the challenge produced by a combinatorial explosion between the large number of potential immune checkpoint targets and possible dosing regimens, the cost of conducting such clinical trials could become forbidding. To alleviate this issue, we have developed spQSP-IO as a platform intended to perform virtual clinical trials to explore a wide range of possible treatment combinations on large scale virtual cohorts. With future improvements made in the aforementioned aspects, including more robust coupling methods of non-spatial and spatial components, quantitative validation with digital pathology data as well as spatial transcriptomics, and more advanced biomarker analysis, models developed with spQSP-IO platform can be adopted as an extension to current QSP models and help improve and refine design of clinical trials of immunotherapy and their combination with other anti-cancer treatment agents.