Mutual Correlation between Non-Coding RNA and S-Adenosylmethionine in Human Cancer: Roles and Therapeutic Opportunities
Abstract
Simple Summary
Abstract
1. Introduction
2. Insights into the Regulatory Effects of Non-Coding RNAs on the Enzymes Involved in AdoMet Synthesis and Metabolism
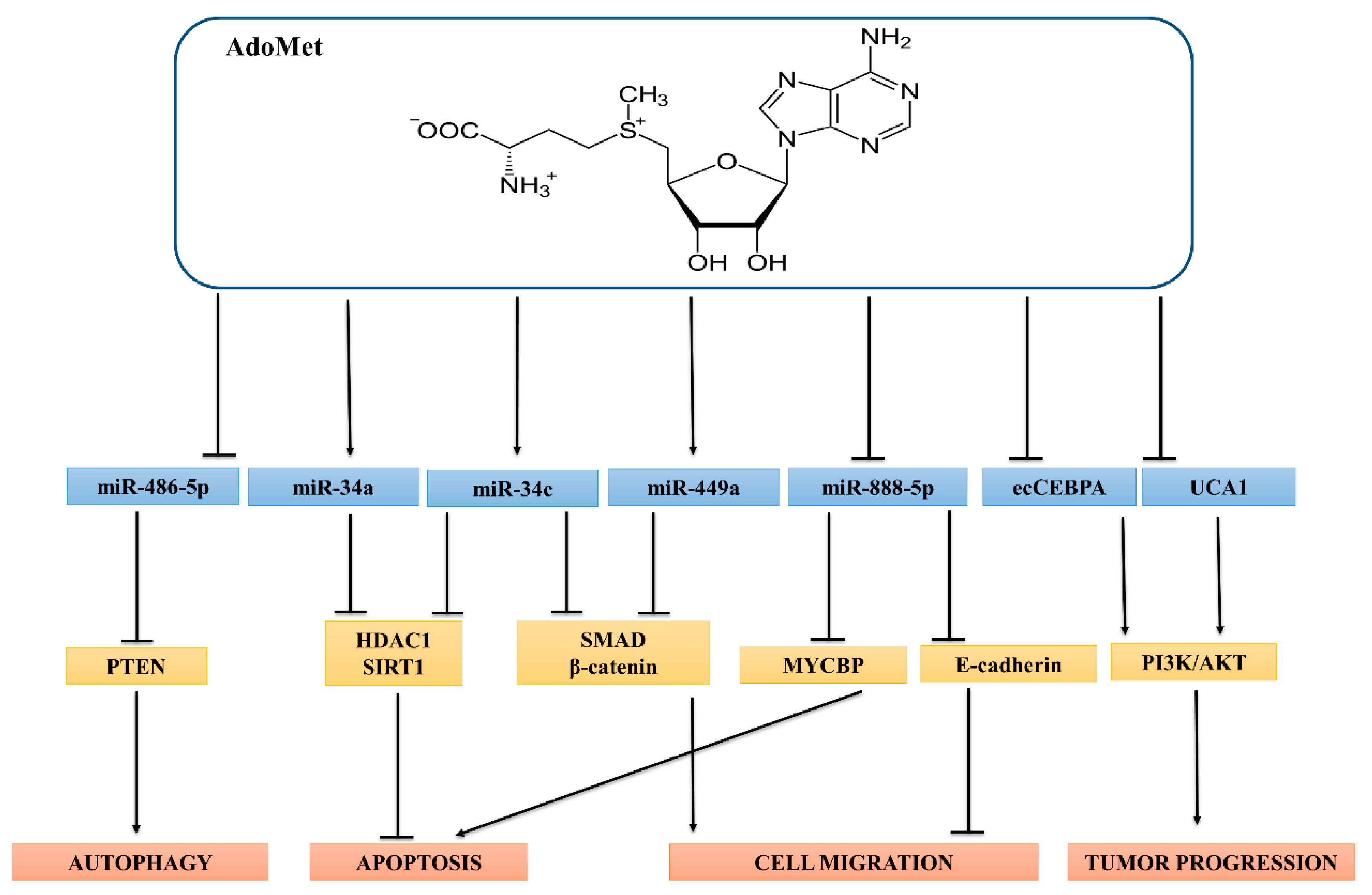
3. AdoMet-Regulated ncRNAs in HUMAN Cancer and Their Role in Chemotherapy
4. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhang, L.; Lu, Q.; Chang, C. Epigenetics in Health and Disease. Adv. Exp. Med. Biol. 2020, 1253, 3–55. [Google Scholar]
- Greenberg, M.; Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 2019, 20, 590–607. [Google Scholar] [CrossRef]
- Sun, Z.; Zhang, Y.; Jia, J.; Fang, Y.; Tang, Y.; Wu, H.; Fang, D. H3K36me3, message from chromatin to DNA damage repair. Cell Biosci. 2020, 10, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Deb, M.; Kar, S.; Sengupta, D.; Shilpi, A.; Parbin, S.; Rath, S.K.; Londhe, V.A.; Patra, S.K. Chromatin dynamics: H3K4 methylation and H3 variant replacement during development and in cancer. Cell. Mol. Life Sci. 2014, 71, 3439–3463. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; He, F.; Zeng, H.; Ling, S.; Chen, A.; Wang, Y.; Yan, X.; Wei, W.; Pang, Y.; Cheng, H.; et al. Identification of functional cooperative mutations of SETD2 in human acute leukemia. Nat. Genet. 2014, 46, 287–293. [Google Scholar] [CrossRef] [PubMed]
- He, P.C.; He, C. m6A RNA methylation: From mechanisms to therapeutic potential. EMBO J. 2021, 40, e105977–e105991. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Chen, C.; Ji, X.; Liu, J.; Zhou, Q.; Wang, G.; Yuan, W.; Kan, Q.; Sun, Z. The interplay between m6A RNA methylation and noncoding RNA in cancer. J. Hematol. Oncol. 2019, 12, 121–135. [Google Scholar] [CrossRef]
- Kim, J.; Lee, G. Metabolic Control of m6A RNA Modification. Metabolites 2021, 11, 80. [Google Scholar] [CrossRef]
- Tian, S.; Lai, J.; Yu, T.; Li, Q.; Chen, Q. Regulation of Gene Expression Associated with the N6-Methyladenosine (m6A) Enzyme System and Its Significance in Cancer. Front. Oncol. 2021, 10, 623634–623647. [Google Scholar] [CrossRef]
- Liu, X.; Wang, P.; Teng, X.; Zhang, Z.; Song, S. Comprehensive Analysis of Expression Regulation for RNA m6A Regulators with Clinical Significance in Human Cancers. Front. Oncol. 2021, 11, 624395–624407. [Google Scholar] [CrossRef]
- Stoccoro, A.; Coppedè, F. Mitochondrial DNA Methylation and Human Diseases. Int. J. Mol. Sci. 2021, 22, 4594. [Google Scholar] [CrossRef]
- Feng, S.; Xiong, L.; Ji, Z.; Cheng, W.; Yang, H. Correlation between increased ND2 expression and demethylated displacementloop of mtDNA in colorectal cancer. Mol. Med. Rep. 2012, 6, 125–130. [Google Scholar] [PubMed]
- Gao, J.; Wen, S.; Zhou, H.; Feng, S. De-methylation of displacement loop of mitochondrial DNA is associated with increased mitochondrial copy number and nicotinamide adenine dinucleotide subunit 2 expression in colorectal cancer. Mol. Med. Rep. 2015, 12, 7033–7038. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Vaghjiani, V.; Jayasekara, W.S.N.; Cain, J.E.; St John, J.C. The degree of mitochondrial DNA methylation in tumor models of glioblastoma and osteosarcoma. Clin. Epigenet. 2018, 10, 157–173. [Google Scholar] [CrossRef]
- Singh, M.; Kumar, V.; Sehrawat, N.; Yadav, M.; Chaudhary, M.; Upadhyay, S.K.; Kumar, S.; Sharma, V.; Kumar, S.; Dilbaghi, N.; et al. Current paradigms in epigenetic anticancer therapeutics and future challenges. Semin. Cancer Biol 2021. [Google Scholar] [CrossRef] [PubMed]
- Cabrera-Licona, A.; Pérez-Añorve, I.X.; Flores-Fortis, M.; Moral-Hernández, O.D.; González-de la Rosa, C.H.; Suárez-Sánchez, R.; Chávez-Saldaña, M.; Aréchaga-Ocampo, E. Deciphering the epigenetic network in cancer radioresistance. Radiother. Oncol. 2021, 159, 48–59. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.C. S-Adenosylmethionine. Int. J. Biochem. Cell Biol. 2000, 32, 391–395. [Google Scholar] [CrossRef]
- Lu, S.C.; Mato, J.M. S-Adenosylmethionine in cell growth, apoptosis and liver cancer. J. Gastroenterol. Hepatol. 2008, 23, S73–S77. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.C.; Mato, J.M. S-adenosylmethionine in liver health, injury, and cancer. Physiol. Rev. 2012, 92, 1515–1542. [Google Scholar] [CrossRef]
- Frau, M.; Feo, F.; Pascale, R.M. Pleiotropic effects of methionine adenosyltransferase deregulation as determinants of liver cancer progression and prognosis. J. Hepatol. 2013, 59, 830–841. [Google Scholar] [CrossRef]
- Papakostas, G.I.; Cassiello, C.F.; Iovieno, N. Folates and S-adenosylmethionine for major depressive disorder. Can. J. Psychiatry Revue Can. Psychiatr. 2012, 57, 406–413. [Google Scholar] [CrossRef] [PubMed]
- Soeken, K.L.; Lee, W.L.; Bausell, R.B.; Agelli, M.; Berman, B.M. Safety and efficacy of S-adenosylmethionine (SAMe) for osteoarthritis. J. Fam. Pract. 2012, 51, 425–430. [Google Scholar]
- Mora, S.I.; García-Román, J.; Gómez-Ñañez, I.; García-Román, R. Chronic liver diseases and the potential use of S-adenosyl-L-methionine as a hepatoprotector. Eur. J. Gastroenterol. Hepatol. 2018, 30, 893–900. [Google Scholar] [CrossRef] [PubMed]
- Karas Kuželički, N. S-Adenosyl Methionine in the therapy of depression and other psychiatric disorders. Drug Dev. Res. 2016, 77, 346–356. [Google Scholar] [CrossRef]
- Furujo, M.; Kinoshita, M.; Nagao, M.; Kubo, T. Methionine adenosyltransferase I/III deficiency: Neurological manifestations and relevance of S-adenosylmethionine. Mol. Genet. Metab. 2012, 107, 253–256. [Google Scholar] [CrossRef]
- Cheng, H.; Gomes-Trolin, C.; Aquilonius, S.M.; Steinberg, A.; Löfberg, C.; Ekblom, J.; Oreland, L. Levels of L-methionine S-adenosyltransferase activity in erythrocytes and concentrations of S-adenosylmethionine and S-adenosylhomocysteine in whole blood of patients with Parkinson′s disease. Exp. Neurol. 1997, 145, 580–585. [Google Scholar] [CrossRef]
- Krzystanek, M.; Pałasz, A.; Krzystanek, E.; Krupka-Matuszczyk, I.; Wiaderkiewicz, R.; Skowronek, R. S-adenosyl L-methionine in CNS diseases. Psychiatr. Pol. 2021, 45, 923–931. [Google Scholar]
- Ilisso, C.P.; Sapio, L.; Delle Cave, D.; Illiano, M.; Spina, A.; Cacciapuoti, G.; Porcelli, M. S-Adenosylmethionine affects ERK1/2 and Stat3 pathways and induces apotosis in osteosarcoma cells. J. Cell. Physiol. 2016, 231, 428–435. [Google Scholar] [CrossRef]
- Parashar, S.; Cheishvili, D.; Arakelian, A.; Hussain, Z.; Tanvir, I.; Khan, H.A.; Szyf, M.; Rabbani, S.A. S-Adenosylmethionine blocks osteosarcoma cells proliferation and invasion in vitro and tumor metastasis in vivo: Therapeutic and diagnostic clinical applications. Cancer Med. 2015, 4, 732–744. [Google Scholar] [CrossRef]
- Cave, D.D.; Desiderio, V.; Mosca, L.; Ilisso, C.P.; Mele, L.; Caraglia, M.; Cacciapuoti, G.; Porcelli, M. S-Adenosylmethionine-mediated apoptosis is potentiated by autophagy inhibition induced by chloroquine in human breast cancer cells. J. Cell. Physiol. 2018, 233, 1370–1383. [Google Scholar] [CrossRef]
- Ilisso, C.P.; Castellano, M.; Zappavigna, S.; Lombardi, A.; Vitale, G.; Dicitore, A.; Dicitore, A.; Cacciapuoti, G.; Caraglia, M.; Porcelli, M. The methyldonor S-adenosylmethionine potentiates doxorubicin effects on apoptosis of hormone-dependent breast cancer cell lines. Endocrine 2015, 50, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Mahmood, N.; Cheishvili, D.; Arakelian, A.; Tanvir, I.; Khan, H.A.; Pépin, A.S.; Szyf, M.; Rabbani, S.A. Methyl donor S-adenosylmethionine (SAM) supplementation attenuates breast cancer growth, invasion, and metastasis in vivo; therapeutic and chemopreventive applications. Oncotarget 2018, 9, 5169–5183. [Google Scholar] [CrossRef] [PubMed]
- Mosca, L.; Pagano, M.; Ilisso, C.P.; Cave, D.D.; Desiderio, V.; Mele, L.; Caraglia, M.; Cacciapuoti, G.; Porcelli, M. AdoMet triggers apoptosis in head and neck squamous cancer by inducing ER stress and potentiates cell sensitivity to cisplatin. J. Cell. Physiol. 2019, 234, 13277–13291. [Google Scholar] [CrossRef] [PubMed]
- Mosca, L.; Minopoli, M.; Pagano, M.; Vitiello, F.; Carriero, M.V.; Cacciapuoti, G.; Porcelli, M. Effects of S-adenosyl-L-methionine on the invasion and migration of head and neck squamous cancer cells and analysis of the underlying mechanisms. Int. J. Oncol. 2020, 56, 1212–1224. [Google Scholar] [CrossRef] [PubMed]
- Pagano, M.; Mosca, L.; Vitiello, F.; Ilisso, C.P.; Coppola, A.; Borzacchiello, L.; Mele, L.; Caruso, F.P.; Ceccarelli, M.; Caraglia, M.; et al. Mi-RNA-888-5p is involved in S-adenosylmethionine antitumor effects in laringea squamous cancer cells. Cancers 2020, 12, 3665. [Google Scholar] [CrossRef]
- Mosca, L.; Pagano, M.; Pecoraro, A.; Borzacchiello, L.; Mele, L.; Cacciapuoti, G.; Porcelli, M.; Russo, G.; Russo, A. S-Adenosyl-L-methionine overcomes uL3-mediated drug resistance in p53 deleted colon cancer cells. Int. J. Mol. Sci. 2020, 22, 103. [Google Scholar] [CrossRef]
- Li, T.W.; Zhang, Q.; Oh, P.; Xia, M.; Chen, H.; Bemanian, S.; Lastra, N.; Circ, M.; Moyer, M.P.; Mato, J.M.; et al. S-Adenosylmethionine and methylthioadenosine inhibit cellular FLICE inhibitory protein expression and induce apoptosis in colon cancer cells. Mol. Pharmacol. 2009, 76, 192–200. [Google Scholar] [CrossRef]
- Zsigrai, S.; Kalmár, A.; Nagy, Z.B.; Barták, B.K.; Valcz, G.; Szigeti, K.A.; Galamb, O.; Dankó, T.; Sebestyén, A.; Barna, G.; et al. S-Adenosylmethionine treatment of colorectal cancer cell lines alters DNA methylation, DNA repair and tumor progression-related gene expression. Cells 2020, 9, 1864. [Google Scholar] [CrossRef]
- Mosca, L.; Vitiello, F.; Coppola, A.; Borzacchiello, L.; Ilisso, C.P.; Pagano, M.; Caraglia, M.; Cacciapuoti, G.; Porcelli, M. Therapeutic potential of the natural compound S-adenosylmethionine as a chemoprotective synergistic agent in breast, and head and neck cancer treatment: Current status of research. Int. J. Mol. Sci. 2020, 21, 8547. [Google Scholar] [CrossRef]
- Yan, L.; Tingting, B.; Linxun, L.; Quangen, G.; Genhai, S.; Lei, Q. S-Adenosylmethionine synergistically enhances the antitumor effect of gemcitabine against pancreatic cancer through JAK2/STAT3 pathway. Naunyn-Schmiedeberg Arch. Pharmacol. 2019, 392, 615–622. [Google Scholar]
- Sun, L.; Zhang, J.; Yang, Q.; Si, Y.; Liu, Y.; Wang, Q.; Han, F.; Huang, Z. Synergistic Effects of SAM and selenium compounds on proliferation, migration and adhesion of HeLa cells. Anticancer Res. 2017, 37, 4433–4441. [Google Scholar]
- Mahmood, N.; Arakelian, A.; Cheishvili, D.; Szyf, M.; Rabbani, S.A. S-adenosylmethionine in combination with decitabine shows enhanced anti-cancer effects in repressing breast cancer growth and metastasis. J. Cell. Mol. Med. 2020, 24, 10322–10337. [Google Scholar] [CrossRef]
- Zhang, P.; Wu, W.; Chen, Q.; Chen, M. Non-Coding RNAs and their integrated networks. J. Integr. Bioinform. 2019, 16, 20190027–20190038. [Google Scholar] [CrossRef]
- Chan, J.J.; Tay, Y. Noncoding RNA: RNA Regulatory Networks in Cancer. Int. J. Mol. Sci. 2018, 19, 1310. [Google Scholar] [CrossRef] [PubMed]
- Slack, F.J.; Chinnaiyan, A.M. The Role of Non-coding RNAs in Oncology. Cell 2019, 179, 1033–1055. [Google Scholar] [CrossRef] [PubMed]
- Ransohoff, J.D.; Wei, Y.; Khavari, P.A. The functions and unique features of long intergenic non-coding RNA. Nat. Rev. Mol. Cell Biol. 2018, 19, 143–157. [Google Scholar] [CrossRef] [PubMed]
- Chi, Y.; Wang, D.; Wang, J.; Yu, W.; Yang, J. Long non-coding RNA in the pathogenesis of cancers. Cells 2019, 8, 1015. [Google Scholar] [CrossRef] [PubMed]
- Statello, L.; Guo, C.J.; Chen, L.L.; Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 2021, 22, 96–118. [Google Scholar] [CrossRef]
- Spitale, R.C.; Tsai, M.C.; Chang, H.Y. RNA templating the epigenome long noncoding RNAs as molecular scaffolds. Epigenetics 2011, 6, 539–543. [Google Scholar] [CrossRef]
- López-Urrutia, E.; Bustamante Montes, L.P.; Ladrón de Guevara Cervantes, D.; Pérez-Plasencia, C.; Campos-Parra, A.D. Crosstalk between long non-coding RNAs, micro-RNAs and mRNAs: Deciphering molecular mechanisms of master regulators in cancer. Front. Oncol. 2019, 9, 669–676. [Google Scholar] [CrossRef]
- Ling, H.; Fabbri, M.; Calin, G.A. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat. Rev. Drug Discov. 2013, 12, 847–865. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Gregory, R.I. MicroRNA biogenesis pathways in cancer. Nat. Rev. Cancer 2015, 15, 321–333. [Google Scholar] [CrossRef] [PubMed]
- Stasio, D.D.; Mosca, L.; Lucchese, A.; Cave, D.D.; Kawasaki, H.; Lombardi, A.; Porcelli, M.; Caraglia, M. Salivary mir-27b expression in oral lichen planuspatients: A series of cases and a narrative review of literature. Curr. Top. Med. Chem. 2019, 19, 2816–2823. [Google Scholar] [CrossRef] [PubMed]
- Grassia, V.; Lombardi, A.; Kawasaki, H.; Ferri, C.; Perillo, L.; Mosca, L.; Delle Cave, D.; Nucci, L.; Porcelli, M.; Caraglia, M. Salivary microRNAs as new molecular markers in cleft lip and palate: A new frontier in molecular medicine. Oncotarget 2018, 9, 18929–18938. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, W.T.; Han, C.; Sun, Y.M.; Chen, T.Q.; Chen, Y.Q. Noncoding RNAs in cancer therapy resistance and targeted drug development. J. Hematol. Oncol. 2019, 12, 55–69. [Google Scholar] [CrossRef] [PubMed]
- Si, W.; Shen, J.; Zheng, H.; Fan, W. The role and mechanisms of action of microRNAs in cancer drug resistance. Clin. Epigenet. 2019, 11, 25–48. [Google Scholar] [CrossRef]
- Mollaei, H.; Safaralizadeh, R.; Rostami, Z. MicroRNA replacement therapy in cancer. J. Cell Physiol. 2019, 234, 12369–12384. [Google Scholar] [CrossRef]
- Lai, X.; Eberhardt, M.; Schmitz, U.; Vera, J. Systems biology-based investigation of cooperating microRNAs as monotherapy or adjuvant therapy in cancer. Nucleic Acids Res. 2019, 47, 7753–7766. [Google Scholar] [CrossRef]
- Murray, B.; Barbier-Torres, L.; Fan, W.; Mato, J.M.; Lu, S.C. Methionine adenosyltransferases in liver cancer. World J. Gastroenterol. 2019, 25, 4300–4319. [Google Scholar] [CrossRef]
- Ramani, K.; Lu, S.C. Methionine adenosyltransferases in liver health and diseases. Liver Res. 2017, 1, 103–111. [Google Scholar] [CrossRef]
- Maldonado, L.Y.; Arsene, D.; Mato, J.M.; Lu, S.C. Methionine adenosyltransferases in cancers: Mechanisms of dysregulation and implications for therapy. Exp. Biol. Med. 2018, 243, 107–117. [Google Scholar] [CrossRef] [PubMed]
- Porcelli, M.; Ilisso, C.P.; Mosca, L.; Cacciapuoti, G. A thermostable archaeal S-adenosylmethionine synthetase: A promising tool to improve the synthesis of adenosylmethionine analogs of biotechnological interest. Bioengineered 2015, 6, 184–186. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yang, H.; Zheng, Y.; Li, T.W.; Peng, H.; Fernandez-Ramos, D.; Martínez-Chantar, M.L.; Rojas, A.L.; Mato, J.M.; Lu, S.C. Methionine adenosyltransferase 2B, HuR, and sirtuin 1 protein cross-talk impacts on the effect of resveratrol on apoptosis and growth in liver cancer cells. J. Boil. Chem. 2013, 288, 23161–23170. [Google Scholar] [CrossRef]
- Peng, H.; Dara, L.; Li, T.W.; Zheng, Y.; Yang, H.; Tomasi, M.L.; Tomasi, I.; Giordano, P.; Mato, J.M.; Lu, S.C. Methionine adenosyltransferase 2B-GIT1 interplay activates MEK1-ERK1/2 to induce growth in human liver and colon cancer. Hepatology 2013, 57, 2299–2313. [Google Scholar] [CrossRef] [PubMed]
- Mato, M.J.; Corrales, F.J.; Lu, S.C.; Avila, M.A. S-Adenosylmethionine: A control switch that regulates liver function. FASEB J. 2002, 16, 15–26. [Google Scholar] [CrossRef]
- Yang, H.; Cho, M.E.; Li, T.W.; Peng, H.; Ko, K.S.; Mato, J.M.; Lu, S.C. MicroRNAs regulate methionine adenosyltransferase 1A expression in hepatocellular carcinoma. J. Clin. Investig. 2013, 123, 285–298. [Google Scholar] [CrossRef]
- Koturbash, I.; Melnyk, S.; James, S.J.; Beland, F.A.; Pogribny, I.P. Role of epigenetic and miR-22 and miR-29b alterations in the downregulation of Mat1a and Mthfr genes in early preneoplastic livers in rats induced by 2-acetylaminofluorene. Mol. Carcinog. 2013, 52, 318–327. [Google Scholar] [CrossRef]
- Pieroth, R.; Paver, S.; Day, S.; Lammersfeld, C. Folate and Its Impact on Cancer Risk. Curr. Nutr. Rep. 2018, 7, 70–84. [Google Scholar] [CrossRef]
- Li, C.; Ni, J.; Liu, Y.X.; Wang, H.; Liang, Z.Q.; Wang, X. Response of MiRNA-22-3p and MiRNA-149-5p to Folate Deficiency and the Differential Regulation of MTHFR Expression in Normal and Cancerous Human Hepatocytes. PLoS ONE 2017, 12, e0168049. [Google Scholar] [CrossRef] [PubMed]
- Tong, D.; Zhang, J.; Wang, X.; Li, Q.; Liu, L.; Lu, A.; Guo, B.; Yang, J.; Ni, L.; Qin, H.; et al. MiR-22, regulated by MeCP2, suppresses gastric cancer cell proliferation by inducing a deficiency in endogenous S-adenosylmethionine. Oncogenesis 2020, 9, 99–116. [Google Scholar] [CrossRef]
- Lo, T.F.; Tsai, W.C.; Chen, S.T. MicroRNA-21-3p, a berberine-induced miRNA, directly down-regulates human methionine adenosyltransferases 2A and 2B and inhibits hepatoma cell growth. PLoS ONE 2013, 30, e75628. [Google Scholar] [CrossRef]
- Ge, X.; Li, W.; Huang, S.; Yin, Z.; Yang, M.; Han, Z.; Han, Z.; Chen, F.; Wang, H.; Lei, P.; et al. Increased miR-21-3p in injured brain microvascular endothelial cells after traumatic brain injury aggravates blood-brain barrier damage by promoting cellular apoptosis and inflammation through targeting MAT2B. J. Neurotrauma 2019, 15, 1291–1305. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Fei, K.; Tian, F.; Gao, C.; Yang, S. Adipose-derived mesenchymal stem cells attenuate ischemic brain injuries in rats by modulating miR-21-3p/MAT2B signaling transduction. Croat. Med. J. 2019, 60, 439–448. [Google Scholar] [CrossRef]
- Tomasi, M.L.; Cossu, C.; Spissu, Y.; Floris, A.; Ryoo, M.; Iglesias-Ara, A.; Wang, Q.; Pandol, S.J.; Bhowmick, N.A.; Seki, E.; et al. S-adenosylmethionine and methylthioadenosine inhibit cancer metastasis by targeting microRNA 34a/b-methionine adenosyltransferase 2A/2B axis. Oncotarget 2017, 12, 78851–78869. [Google Scholar] [CrossRef] [PubMed]
- Simile, M.M.; Peitta, G.; Tomasi, M.L.; Brozzetti, S.; Feo, C.F.; Porcu, A.; Cigliano, A.; Calvisi, D.F.; Feo, F.; Pascale, R.M. MicroRNA-203 impacts on the growth, aggressiveness and prognosis of hepatocellular carcinoma by targeting MAT2A and MAT2B genes. Oncotarget 2019, 19, 2835–2854. [Google Scholar] [CrossRef]
- Zhang, D.; Zhao, T.; Ang, H.S.; Chong, P.; Saiki, R.; Igarashi, K.; Yang, H.; Vardy, L.A. AMD1 is essential for ESC self-renewal and is translationally down-regulated on differentiation to neural precursor cells. Genes Dev. 2012, 26, 461–473. [Google Scholar] [CrossRef]
- Guo, T.; Wang, H.; Liu, P.; Xiao, Y.; Wu, P.; Wang, Y.; Chen, B.; Zhao, Q.; Liu, Z.; Liu, Q. SNHG6 Acts as a Genome-Wide Hypomethylation Trigger via Coupling of miR-1297-Mediated S-Adenosylmethionine-Dependent Positive Feedback Loops. Cancer Res. 2018, 15, 3849–3864. [Google Scholar] [CrossRef]
- Mishra, K.; Kanduri, C. Understanding Long Noncoding RNA and Chromatin Interactions: What We Know So Far. Non-Coding RNA 2019, 5, 54–81. [Google Scholar]
- O’Leary, V.B.; Ovsepian, S.V.; Smida, J.; Atkinson, M.J. Particle—The RNA podium for genomic silencers. J. Cell Physiol. 2019, 234, 19464–19470. [Google Scholar] [CrossRef]
- O’Leary, V.B.; Ovsepian, S.V.; Carrascosa, L.G.; Buske, F.A.; Radulovic, V.; Niyazi, M.; Moertl, S.; Trau, M.; Atkinson, M.J.; Anastasov, N. Particle, a Triplex-Forming Long ncRNA, Regulates Locus-Specific Methylation in Response to Low-Dose Irradiation. Cell Rep. 2015, 11, 474–485. [Google Scholar] [CrossRef] [PubMed]
- O’Leary, V.B.; Hain, S.; Maugg, D.; Smida, J.; Azimzadeh, O.; Tapio, S.; Ovsepian, S.V.; Atkinson, M.J. Long non-coding RNA Particle bridges histone and DNA methylation. Sci. Rep. 2017, 7, 1790–1796. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Gu, B.; Zhao, X.; Zhao, Y.; Huo, S.; Liu, X.; Lu, H. Circular RNA hsa_circ_0007364 increases cervical cancer progression through activating methionine adenosyltransferase II alpha (MAT2A) expression by restraining microRNA-101-5p. Bioengineered 2020, 11, 1269–1279. [Google Scholar] [CrossRef] [PubMed]
- Chu, Y.D.; Lai, H.Y.; Pai, L.M.; Huang, Y.H.; Lin, Y.H.; Liang, K.H.; Yeh, C.T. The methionine salvage pathway-involving ADI1 inhibits hepatoma growth by epigenetically altering genes expression via elevating S-adenosylmethionine. Cell Death Dis. 2019, 10, 240–253. [Google Scholar] [CrossRef] [PubMed]
- Ilisso, C.P.; Delle Cave, D.; Mosca, L.; Pagano, M.; Coppola, A.; Mele, L.; Caraglia, M.; Cacciapuoti, G.; Porcelli, M. S-Adenosylmethionine regulates apoptosis and autophagy in MCF-7 breast cancer cells through the modulation of specific microRNAs. Cancer Cell Int. 2018, 18, 197–209. [Google Scholar] [CrossRef] [PubMed]
- Coppola, A.; Ilisso, C.P.; Stellavato, A.; Schiraldi, C.; Caraglia, M.; Mosca, L.; Cacciapuoti, G.; Porcelli, M. S-Adenosylmethionine Inhibits Cell Growth and Migration of Triple Negative Breast Cancer Cells through Upregulating MiRNA-34c and MiRNA-449a. Int. J. Mol. Sci. 2020, 22, 286. [Google Scholar] [CrossRef] [PubMed]
- Sadek, K.M.; Lebda, M.A.; Nasr, N.E.; Nasr, S.M.; El-Sayed, Y. Role of lncRNAs as prognostic markers of hepatic cancer and potential therapeutic targeting by S-adenosylmethionine via inhibiting PI3K/Akt signaling pathways. Environ. Sci. Pollut. Res. 2018, 25, 20057–20070. [Google Scholar] [CrossRef]
- Yang, X.; Liu, M.; Li, M.; Zhang, S.; Hiju, H.; Sun, J.; Mao, Z.; Zheng, M.; Feng, B. Epigenetic modulations of noncoding RNA: A novel dimension of cancer biology. Mol. Cancer 2020, 19, 64–75. [Google Scholar] [CrossRef]
- Huang, H.; Weng, H.; Chen, J. m6A Modification in Coding and Non-coding RNAs: Roles and Therapeutic Implications in Cancer. Cancer Cell 2020, 37, 270–288. [Google Scholar] [CrossRef]
- Ruszkowska, A. METTL16, Methyltransferase-Like Protein 16: Current Insights into Structure and Function. Int. J. Mol. Sci. 2021, 22, 2176. [Google Scholar] [CrossRef]
- Wang, J.X.; Breaker, R.R. Riboswitches that sense S-adenosylmethionine and S-adenosylhomocysteine. Biochem. Cell Biol. 2008, 86, 157–168. [Google Scholar] [CrossRef]
- Tang, D.J.; Du, X.; Shi, Q.; Zhang, J.L.; He, Y.P.; Chen, Y.M.; Ming, Z.; Wang, D.; Zhong, W.Y.; Liang, Y.W.; et al. A SAM-I riboswitch with the ability to sense and respond to uncharged initiator tRNA. Nat. Commun. 2020, 11, 2794–2804. [Google Scholar] [CrossRef]
- Shima, H.; Matsumoto, M.; Ishigami, Y.; Ebina, M.; Muto, A.; Sato, Y.; Kumagai, S.; Ochiai, K.; Suzuki, T.; Igarashi, K. S-Adenosylmethionine Synthesis Is Regulated by Selective N6-Adenosine Methylation and mRNA Degradation Involving METTL16 and YTHDC1. Cell Rep. 2017, 21, 3354–3363. [Google Scholar] [CrossRef]
- Pendleton, K.E.; Chen, B.; Liu, K.; Hunter, O.V.; Xie, Y.; Tu, B.P.; Conrad, N.K. The U6 snRNA m6A Methyltransferase METTL16 Regulates SAM Synthetase Intron Retention. Cell 2017, 169, 824–835.e14. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, I.; Chatterjee, A. Recent Advances in miRNA Delivery Systems. Methods Protoc. 2021, 4, 10. [Google Scholar] [CrossRef] [PubMed]
- Forterre, A.; Komuro, H.; Aminova, S.; Harada, M. A Comprehensive Review of Cancer MicroRNA Therapeutic Delivery Strategies. Cancers 2020, 12, 1852. [Google Scholar] [CrossRef]
- Magesh, B.; Naidu, P.Y.; Rajarajeswari, G.R. S-adenosyl-l-methionine (SAMe)-loaded nanochitosan particles: Synthesis, characterization and in vitro drug release studies. J. Exp. Nanosci. 2014, 10, 828–843. [Google Scholar] [CrossRef]
- Ergin, A.D.; Bayindir, Z.S.; Ozcelikay, A.T.; Yuksel, A. A novel delivery system for enhancing bioavailability of S-adenosyl-l-methionine: Pectin nanoparticles-in-microparticles and their in vitro-in vivo evaluation. J. Drug Deliv. Sci. Technol. 2020, 61, 102096–102108. [Google Scholar] [CrossRef]
| ncRNAs | Tumor Type | Functions | References |
|---|---|---|---|
| ↑ miR-664 | HCC | ↓ MAT1A | [66] |
| ↑ miR-485-3p | |||
| ↑ miR-495 | |||
| ↑ miR-22 | HCC | ↓ MTHFR | [67,68] |
| ↑ miR-29b | GC | ↓ MAT1A | [70] |
| ↑ miR-149-5p | HCC | ↓ MTHFR | [69] |
| ↑ miR-21-3p | HCC | ↓ MAT2A | [71] |
| BMVECs | ↓ MAT2B | [72] | |
| ↓ miR-21-3p | ADMSCs | ↑ MAT2B | [73] |
| ↑ miR-34a/b | CRC | ↓ MAT2A | [74] |
| ↓ MAT2B | |||
| ↑ miR-203 | HCC | ↓ MAT2A | [75] |
| ↓ MAT2B | |||
| ↓ miR-762 | ESC | ↑ Adm1 | [76] |
| ↑ SNHG6 | HCC | ↓ MAT1A | [77] |
| ↓ miR-1297 | ↑ MAT2A | ||
| ↑ MAT2B | |||
| ↑ PARTICLE | BC | ↓ MAT2A | [81] |
| ↑ has_circ_0007364 | CC | ↓ MAT2A | [82] |
| ↓ miR-101-5p |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mosca, L.; Vitiello, F.; Borzacchiello, L.; Coppola, A.; Tranchese, R.V.; Pagano, M.; Caraglia, M.; Cacciapuoti, G.; Porcelli, M. Mutual Correlation between Non-Coding RNA and S-Adenosylmethionine in Human Cancer: Roles and Therapeutic Opportunities. Cancers 2021, 13, 3264. https://doi.org/10.3390/cancers13133264
Mosca L, Vitiello F, Borzacchiello L, Coppola A, Tranchese RV, Pagano M, Caraglia M, Cacciapuoti G, Porcelli M. Mutual Correlation between Non-Coding RNA and S-Adenosylmethionine in Human Cancer: Roles and Therapeutic Opportunities. Cancers. 2021; 13(13):3264. https://doi.org/10.3390/cancers13133264
Chicago/Turabian StyleMosca, Laura, Francesca Vitiello, Luigi Borzacchiello, Alessandra Coppola, Roberta Veglia Tranchese, Martina Pagano, Michele Caraglia, Giovanna Cacciapuoti, and Marina Porcelli. 2021. "Mutual Correlation between Non-Coding RNA and S-Adenosylmethionine in Human Cancer: Roles and Therapeutic Opportunities" Cancers 13, no. 13: 3264. https://doi.org/10.3390/cancers13133264
APA StyleMosca, L., Vitiello, F., Borzacchiello, L., Coppola, A., Tranchese, R. V., Pagano, M., Caraglia, M., Cacciapuoti, G., & Porcelli, M. (2021). Mutual Correlation between Non-Coding RNA and S-Adenosylmethionine in Human Cancer: Roles and Therapeutic Opportunities. Cancers, 13(13), 3264. https://doi.org/10.3390/cancers13133264







