Organoids and Colorectal Cancer
Abstract
Simple Summary
Abstract
1. Introduction
1.1. Organoid Definition
1.2. Colorectal Cancer (CRC)
1.3. The Short History of Intestinal Organoids
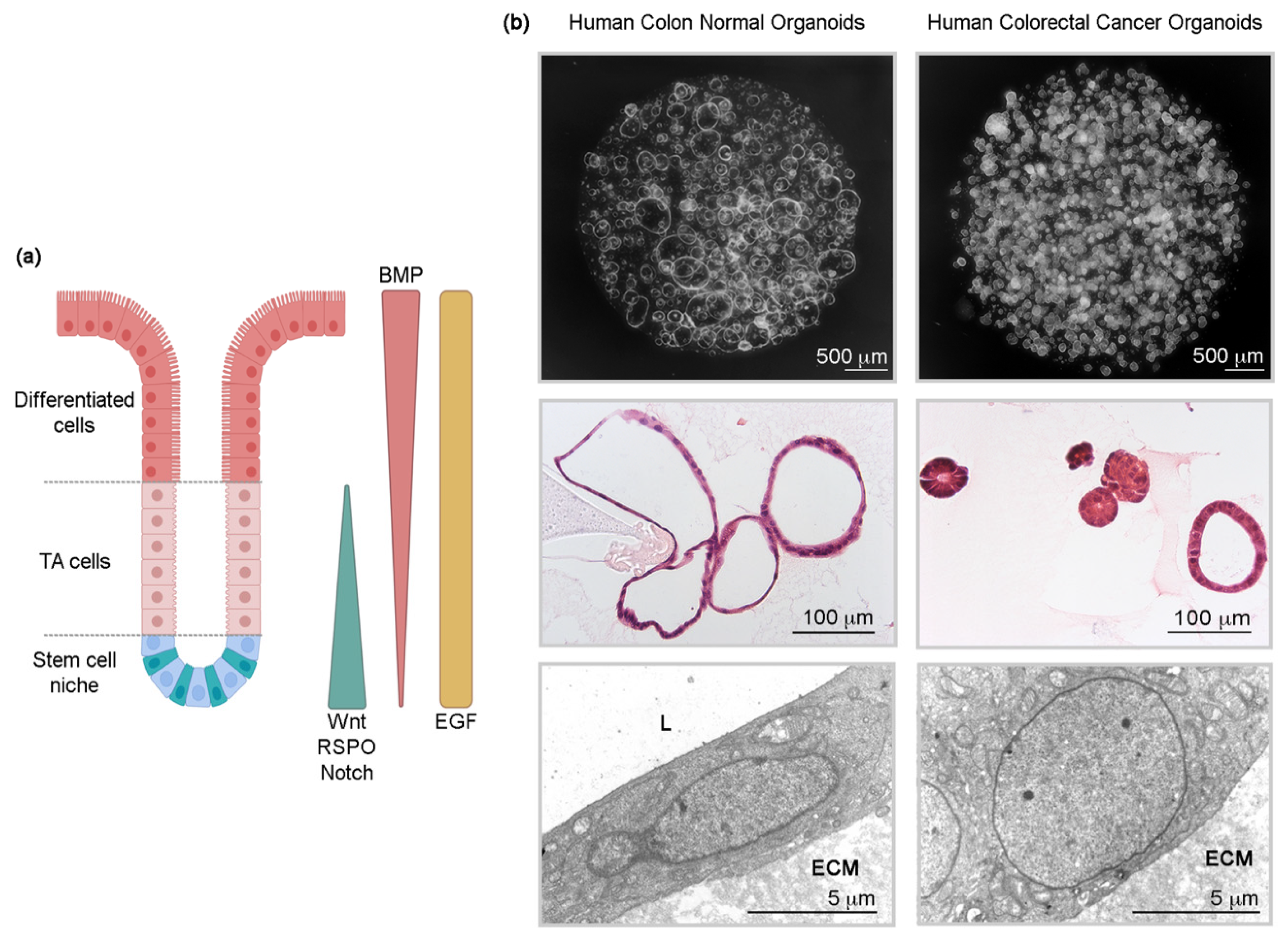
2. Normal and Tumor Intestinal Organoid Culture
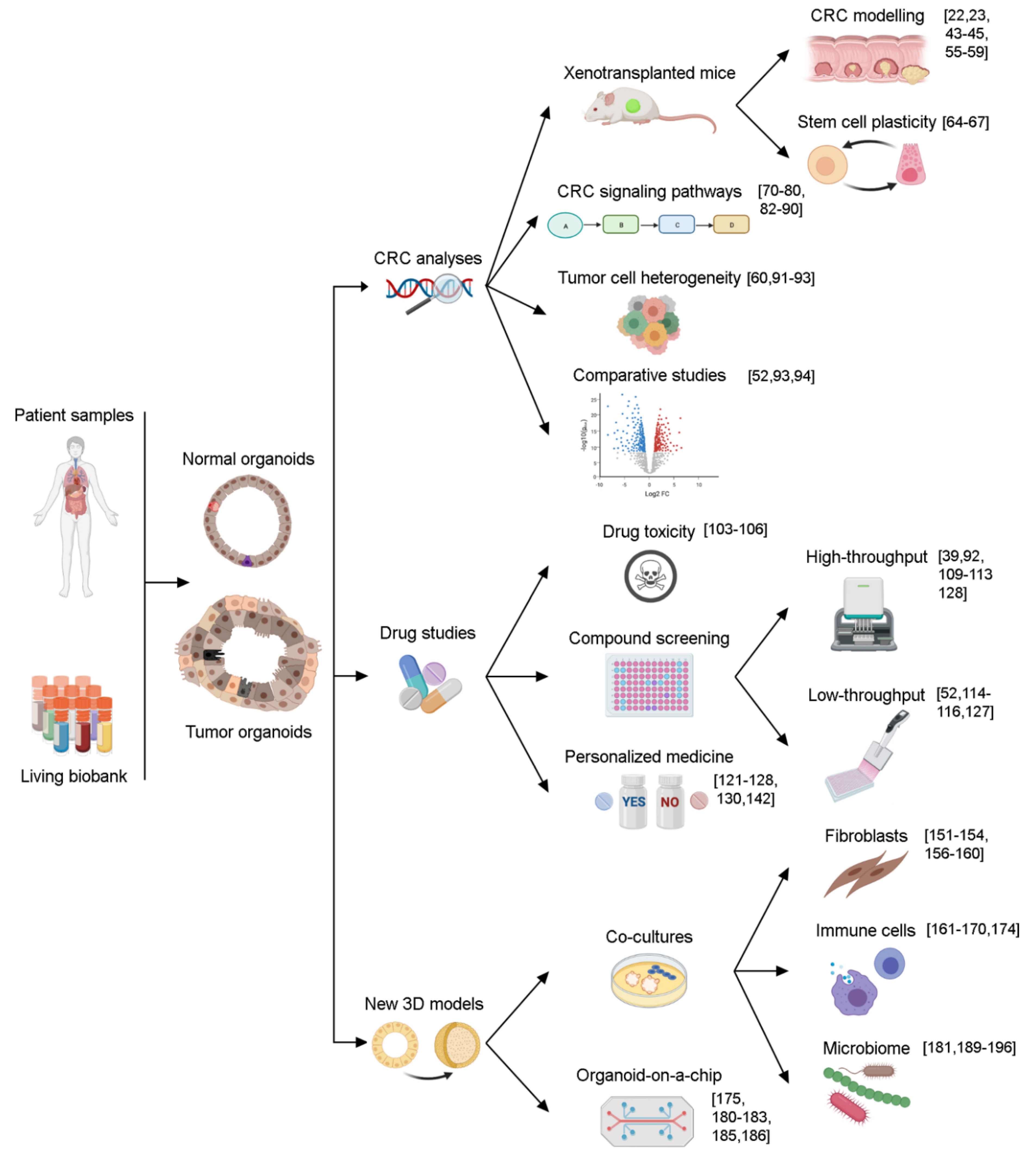
3. Organoids as a Model to Study CRC
3.1. Organoids for CRC Modelling and Cell Plasticity Analyses
3.2. Organoids for Molecular and Genetic CRC Studies
4. Organoids as a Platform for Screening Assays
4.1. Toxicity Studies
4.2. Drug Discovery
4.3. Organoids for Personalized Treatment of CRC Patients. Early-Onset CRC and Colorectal Carcinomatosis
5. Organoid Limitations and New 3D Models
5.1. Co-Culture with Fibroblasts
5.2. Co-Culture with Immune Cells
5.3. Organoid-on-a-Chip
6. Organoids to Study the Intestinal Microbiome and Host-Pathogen Interactions
7. Perspectives and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Schutgens, F.; Clevers, H. Human Organoids: Tools for Understanding Biology and Treating Diseases. Annu. Rev. Pathol. 2020, 15, 211–234. [Google Scholar] [CrossRef]
- Kim, J.; Koo, B.K.; Knoblich, J.A. Human organoids: Model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 2020, 21, 571–584. [Google Scholar] [CrossRef]
- Fujii, M.; Sato, T. Somatic cell-derived organoids as prototypes of human epithelial tissues and diseases. Nat. Mater. 2021, 20, 156–169. [Google Scholar] [CrossRef]
- Bock, C.; Boutros, M.; Camp, J.G.; Clarke, L.; Clevers, H.; Knoblich, J.A.; Liberali, P.; Regev, A.; Rios, A.C.; Stegle, O.; et al. The Organoid Cell Atlas. Nat. Biotechnol. 2021, 39, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Nusse, R.; Clevers, H. Wnt/beta-Catenin Signaling, Disease, and Emerging Therapeutic Modalities. Cell 2017, 169, 985–999. [Google Scholar] [CrossRef] [PubMed]
- Huch, M.; Knoblich, J.A.; Lutolf, M.P.; Martinez-Arias, A. The hope and the hype of organoid research. Development 2017, 144, 938–941. [Google Scholar] [CrossRef]
- Cancer Genome Atlas, N. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Yaeger, R.; Chatila, W.K.; Lipsyc, M.D.; Hechtman, J.F.; Cercek, A.; Sanchez-Vega, F.; Jayakumaran, G.; Middha, S.; Zehir, A.; Donoghue, M.T.A.; et al. Clinical Sequencing Defines the Genomic Landscape of Metastatic Colorectal Cancer. Cancer Cell 2018, 33, 125–136.e123. [Google Scholar] [CrossRef] [PubMed]
- Sievers, C.K.; Grady, W.M.; Halberg, R.B.; Pickhardt, P.J. New insights into the earliest stages of colorectal tumorigenesis. Expert Rev. Gastroenterol. Hepatol. 2017, 11, 723–729. [Google Scholar] [CrossRef] [PubMed]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reynies, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef]
- Dienstmann, R.; Vermeulen, L.; Guinney, J.; Kopetz, S.; Tejpar, S.; Tabernero, J. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat. Rev. Cancer 2017, 17, 79–92. [Google Scholar] [CrossRef]
- Sanz-Pamplona, R.; Cordero, D.; Berenguer, A.; Lejbkowicz, F.; Rennert, H.; Salazar, R.; Biondo, S.; Sanjuan, X.; Pujana, M.A.; Rozek, L.; et al. Gene expression differences between colon and rectum tumors. Clin. Cancer Res. 2011, 17, 7303–7312. [Google Scholar] [CrossRef]
- Wei, C.; Chen, J.; Pande, M.; Lynch, P.M.; Frazier, M.L. A pilot study comparing protein expression in different segments of the normal colon and rectum and in normal colon versus adenoma in patients with Lynch syndrome. J. Cancer Res. Clin. Oncol. 2013, 139, 1241–1250. [Google Scholar] [CrossRef][Green Version]
- Barker, N.; van Es, J.H.; Kuipers, J.; Kujala, P.; van den Born, M.; Cozijnsen, M.; Haegebarth, A.; Korving, J.; Begthel, H.; Peters, P.J.; et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 2007, 449, 1003–1007. [Google Scholar] [CrossRef]
- De Lau, W.; Barker, N.; Low, T.Y.; Koo, B.K.; Li, V.S.; Teunissen, H.; Kujala, P.; Haegebarth, A.; Peters, P.J.; van de Wetering, M.; et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 2011, 476, 293–297. [Google Scholar] [CrossRef] [PubMed]
- Schuijers, J.; Clevers, H. Adult mammalian stem cells: The role of Wnt, Lgr5 and R-spondins. EMBO J. 2012, 31, 2685–2696. [Google Scholar] [CrossRef] [PubMed]
- De Lau, W.; Peng, W.C.; Gros, P.; Clevers, H. The R-spondin/Lgr5/Rnf43 module: Regulator of Wnt signal strength. Genes Dev. 2014, 28, 305–316. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Vries, R.G.; Snippert, H.J.; van de Wetering, M.; Barker, N.; Stange, D.E.; van Es, J.H.; Abo, A.; Kujala, P.; Peters, P.J.; et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265. [Google Scholar] [CrossRef]
- Sato, T.; Stange, D.E.; Ferrante, M.; Vries, R.G.; Van Es, J.H.; Van den Brink, S.; Van Houdt, W.J.; Pronk, A.; Van Gorp, J.; Siersema, P.D.; et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 2011, 141, 1762–1772. [Google Scholar] [CrossRef]
- Jung, P.; Sato, T.; Merlos-Suarez, A.; Barriga, F.M.; Iglesias, M.; Rossell, D.; Auer, H.; Gallardo, M.; Blasco, M.A.; Sancho, E.; et al. Isolation and in vitro expansion of human colonic stem cells. Nat. Med. 2011, 17, 1225–1227. [Google Scholar] [CrossRef]
- Leung, C.; Tan, S.H.; Barker, N. Recent Advances in Lgr5(+) Stem Cell Research. Trends Cell Biol. 2018, 28, 380–391. [Google Scholar] [CrossRef]
- Barker, N.; Ridgway, R.A.; van Es, J.H.; van de Wetering, M.; Begthel, H.; van den Born, M.; Danenberg, E.; Clarke, A.R.; Sansom, O.J.; Clevers, H. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 2009, 457, 608–611. [Google Scholar] [CrossRef]
- Schepers, A.G.; Snippert, H.J.; Stange, D.E.; van den Born, M.; van Es, J.H.; van de Wetering, M.; Clevers, H. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science 2012, 337, 730–735. [Google Scholar] [CrossRef] [PubMed]
- Fujii, M.; Matano, M.; Toshimitsu, K.; Takano, A.; Mikami, Y.; Nishikori, S.; Sugimoto, S.; Sato, T. Human Intestinal Organoids Maintain Self-Renewal Capacity and Cellular Diversity in Niche-Inspired Culture Condition. Cell Stem Cell 2018, 23, 787–793.e786. [Google Scholar] [CrossRef]
- Fujii, M.; Matano, M.; Nanki, K.; Sato, T. Efficient genetic engineering of human intestinal organoids using electroporation. Nat. Protoc. 2015, 10, 1474–1485. [Google Scholar] [CrossRef] [PubMed]
- Fujii, M.; Shimokawa, M.; Date, S.; Takano, A.; Matano, M.; Nanki, K.; Ohta, Y.; Toshimitsu, K.; Nakazato, Y.; Kawasaki, K.; et al. A Colorectal Tumor Organoid Library Demonstrates Progressive Loss of Niche Factor Requirements during Tumorigenesis. Cell Stem Cell 2016, 18, 827–838. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, K.; Toshimitsu, K.; Matano, M.; Fujita, M.; Fujii, M.; Togasaki, K.; Ebisudani, T.; Shimokawa, M.; Takano, A.; Takahashi, S.; et al. An Organoid Biobank of Neuroendocrine Neoplasms Enables Genotype-Phenotype Mapping. Cell 2020, 183, 1420–1435.e21. [Google Scholar] [CrossRef] [PubMed]
- Beumer, J.; Clevers, H. Cell fate specification and differentiation in the adult mammalian intestine. Nat. Rev. Mol. Cell Biol. 2021, 22, 39–53. [Google Scholar] [CrossRef]
- Sato, T.; van Es, J.H.; Snippert, H.J.; Stange, D.E.; Vries, R.G.; van den Born, M.; Barker, N.; Shroyer, N.F.; van de Wetering, M.; Clevers, H. Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature 2011, 469, 415–418. [Google Scholar] [CrossRef] [PubMed]
- Vermeulen, L.; De Sousa, E.M.F.; van der Heijden, M.; Cameron, K.; de Jong, J.H.; Borovski, T.; Tuynman, J.B.; Todaro, M.; Merz, C.; Rodermond, H.; et al. Wnt activity defines colon cancer stem cells and is regulated by the microenvironment. Nat. Cell Biol. 2010, 12, 468–476. [Google Scholar] [CrossRef]
- Sasaki, N.; Sachs, N.; Wiebrands, K.; Ellenbroek, S.I.; Fumagalli, A.; Lyubimova, A.; Begthel, H.; van den Born, M.; van Es, J.H.; Karthaus, W.R.; et al. Reg4+ deep crypt secretory cells function as epithelial niche for Lgr5+ stem cells in colon. Proc. Natl. Acad. Sci. USA 2016, 113, E5399–E5407. [Google Scholar] [CrossRef]
- Mihara, E.; Hirai, H.; Yamamoto, H.; Tamura-Kawakami, K.; Matano, M.; Kikuchi, A.; Sato, T.; Takagi, J. Active and water-soluble form of lipidated Wnt protein is maintained by a serum glycoprotein afamin/alpha-albumin. Elife 2016, 5. [Google Scholar] [CrossRef]
- Naschberger, A.; Orry, A.; Lechner, S.; Bowler, M.W.; Nurizzo, D.; Novokmet, M.; Keller, M.A.; Oemer, G.; Seppi, D.; Haslbeck, M.; et al. Structural Evidence for a Role of the Multi-functional Human Glycoprotein Afamin in Wnt Transport. Structure 2017, 25, 1907–1915.e1905. [Google Scholar] [CrossRef]
- Haramis, A.P.; Begthel, H.; van den Born, M.; van Es, J.; Jonkheer, S.; Offerhaus, G.J.; Clevers, H. De novo crypt formation and juvenile polyposis on BMP inhibition in mouse intestine. Science 2004, 303, 1684–1686. [Google Scholar] [CrossRef] [PubMed]
- Dignass, A.U.; Sturm, A. Peptide growth factors in the intestine. Eur. J. Gastroenterol. Hepatol. 2001, 13, 763–770. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, C.; Obermeier, F.; Artinger, M.; Hausmann, M.; Falk, W.; Schoelmerich, J.; Rogler, G.; Grossmann, J. Cell-cell contacts prevent anoikis in primary human colonic epithelial cells. Gastroenterology 2007, 132, 587–600. [Google Scholar] [CrossRef]
- Kretzschmar, K.; Clevers, H. Organoids: Modeling Development and the Stem Cell Niche in a Dish. Dev. Cell 2016, 38, 590–600. [Google Scholar] [CrossRef] [PubMed]
- Fujii, S.; Suzuki, K.; Kawamoto, A.; Ishibashi, F.; Nakata, T.; Murano, T.; Ito, G.; Shimizu, H.; Mizutani, T.; Oshima, S.; et al. PGE2 is a direct and robust mediator of anion/fluid secretion by human intestinal epithelial cells. Sci. Rep. 2016, 6, 36795. [Google Scholar] [CrossRef]
- Yin, X.; Farin, H.F.; van Es, J.H.; Clevers, H.; Langer, R.; Karp, J.M. Niche-independent high-purity cultures of Lgr5+ intestinal stem cells and their progeny. Nat. Methods 2014, 11, 106–112. [Google Scholar] [CrossRef]
- Langlands, A.J.; Carroll, T.D.; Chen, Y.; Nathke, I. Chir99021 and Valproic acid reduce the proliferative advantage of Apc mutant cells. Cell Death Dis. 2018, 9, 255. [Google Scholar] [CrossRef] [PubMed]
- Van de Wetering, M.; Francies, H.E.; Francis, J.M.; Bounova, G.; Iorio, F.; Pronk, A.; van Houdt, W.; van Gorp, J.; Taylor-Weiner, A.; Kester, L.; et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 2015, 161, 933–945. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Larsson, P.; Ljuslinder, I.; Ohlund, D.; Myte, R.; Lofgren-Burstrom, A.; Zingmark, C.; Ling, A.; Edin, S.; Palmqvist, R. Ex Vivo Organoid Cultures Reveal the Importance of the Tumor Microenvironment for Maintenance of Colorectal Cancer Stem Cells. Cancers 2020, 12, 923. [Google Scholar] [CrossRef] [PubMed]
- Matano, M.; Date, S.; Shimokawa, M.; Takano, A.; Fujii, M.; Ohta, Y.; Watanabe, T.; Kanai, T.; Sato, T. Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nat. Med. 2015, 21, 256–262. [Google Scholar] [CrossRef] [PubMed]
- Drost, J.; van Jaarsveld, R.H.; Ponsioen, B.; Zimberlin, C.; van Boxtel, R.; Buijs, A.; Sachs, N.; Overmeer, R.M.; Offerhaus, G.J.; Begthel, H.; et al. Sequential cancer mutations in cultured human intestinal stem cells. Nature 2015, 521, 43–47. [Google Scholar] [CrossRef]
- Fumagalli, A.; Oost, K.C.; Kester, L.; Morgner, J.; Bornes, L.; Bruens, L.; Spaargaren, L.; Azkanaz, M.; Schelfhorst, T.; Beerling, E.; et al. Plasticity of Lgr5-Negative Cancer Cells Drives Metastasis in Colorectal Cancer. Cell Stem Cell 2020, 26, 569–578.e567. [Google Scholar] [CrossRef]
- Behjati, S.; Huch, M.; van Boxtel, R.; Karthaus, W.; Wedge, D.C.; Tamuri, A.U.; Martincorena, I.; Petljak, M.; Alexandrov, L.B.; Gundem, G.; et al. Genome sequencing of normal cells reveals developmental lineages and mutational processes. Nature 2014, 513, 422–425. [Google Scholar] [CrossRef]
- Wang, X.; Yamamoto, Y.; Wilson, L.H.; Zhang, T.; Howitt, B.E.; Farrow, M.A.; Kern, F.; Ning, G.; Hong, Y.; Khor, C.C.; et al. Cloning and variation of ground state intestinal stem cells. Nature 2015, 522, 173–178. [Google Scholar] [CrossRef]
- Lugli, N.; Dionellis, V.S.; Ordonez-Moran, P.; Kamileri, I.; Sotiriou, S.K.; Huelsken, J.; Halazonetis, T.D. Enhanced Rate of Acquisition of Point Mutations in Mouse Intestinal Adenomas Compared to Normal Tissue. Cell Rep. 2017, 19, 2185–2192. [Google Scholar] [CrossRef][Green Version]
- Lewis, S.K.; Nachun, D.; Martin, M.G.; Horvath, S.; Coppola, G.; Jones, D.L. DNA Methylation Analysis Validates Organoids as a Viable Model for Studying Human Intestinal Aging. Cell Mol. Gastroenterol. Hepatol. 2020, 9, 527–541. [Google Scholar] [CrossRef]
- Weeber, F.; van de Wetering, M.; Hoogstraat, M.; Dijkstra, K.K.; Krijgsman, O.; Kuilman, T.; Gadellaa-van Hooijdonk, C.G.; van der Velden, D.L.; Peeper, D.S.; Cuppen, E.P.; et al. Preserved genetic diversity in organoids cultured from biopsies of human colorectal cancer metastases. Proc. Natl. Acad. Sci. USA 2015, 112, 13308–13311. [Google Scholar] [CrossRef]
- Buzzelli, J.N.; Ouaret, D.; Brown, G.; Allen, P.D.; Muschel, R.J. Colorectal cancer liver metastases organoids retain characteristics of original tumor and acquire chemotherapy resistance. Stem Cell Res. 2018, 27, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Barral, A.; Costales-Carrera, A.; Buira, S.P.; Jung, P.; Ferrer-Mayorga, G.; Larriba, M.J.; Bustamante-Madrid, P.; Dominguez, O.; Real, F.X.; Guerra-Pastrian, L.; et al. Vitamin D differentially regulates colon stem cells in patient-derived normal and tumor organoids. FEBS J. 2020, 287, 53–72. [Google Scholar] [CrossRef] [PubMed]
- Fujii, M.; Clevers, H.; Sato, T. Modeling Human Digestive Diseases With CRISPR-Cas9-Modified Organoids. Gastroenterology 2019, 156, 562–576. [Google Scholar] [CrossRef] [PubMed]
- Hendriks, D.; Artegiani, B.; Hu, H.; Chuva de Sousa Lopes, S.; Clevers, H. Establishment of human fetal hepatocyte organoids and CRISPR-Cas9-based gene knockin and knockout in organoid cultures from human liver. Nat. Protoc. 2021, 16, 182–217. [Google Scholar] [CrossRef] [PubMed]
- Fumagalli, A.; Drost, J.; Suijkerbuijk, S.J.; van Boxtel, R.; de Ligt, J.; Offerhaus, G.J.; Begthel, H.; Beerling, E.; Tan, E.H.; Sansom, O.J.; et al. Genetic dissection of colorectal cancer progression by orthotopic transplantation of engineered cancer organoids. Proc. Natl. Acad. Sci. USA 2017, 114, E2357–E2364. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, K.; Fujii, M.; Sugimoto, S.; Ishikawa, K.; Matano, M.; Ohta, Y.; Toshimitsu, K.; Takahashi, S.; Hosoe, N.; Sekine, S.; et al. Chromosome Engineering of Human Colon-Derived Organoids to Develop a Model of Traditional Serrated Adenoma. Gastroenterology 2020, 158, 638–651.e638. [Google Scholar] [CrossRef] [PubMed]
- Roper, J.; Tammela, T.; Cetinbas, N.M.; Akkad, A.; Roghanian, A.; Rickelt, S.; Almeqdadi, M.; Wu, K.; Oberli, M.A.; Sanchez-Rivera, F.J.; et al. In Vivo genome editing and organoid transplantation models of colorectal cancer and metastasis. Nat. Biotechnol. 2017, 35, 569–576. [Google Scholar] [CrossRef]
- O’Rourke, K.P.; Loizou, E.; Livshits, G.; Schatoff, E.M.; Baslan, T.; Manchado, E.; Simon, J.; Romesser, P.B.; Leach, B.; Han, T.; et al. Transplantation of engineered organoids enables rapid generation of metastatic mouse models of colorectal cancer. Nat. Biotechnol. 2017, 35, 577–582. [Google Scholar] [CrossRef] [PubMed]
- Tauriello, D.V.F.; Palomo-Ponce, S.; Stork, D.; Berenguer-Llergo, A.; Badia-Ramentol, J.; Iglesias, M.; Sevillano, M.; Ibiza, S.; Canellas, A.; Hernando-Momblona, X.; et al. TGFbeta drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 2018, 554, 538–543. [Google Scholar] [CrossRef]
- Cortina, C.; Turon, G.; Stork, D.; Hernando-Momblona, X.; Sevillano, M.; Aguilera, M.; Tosi, S.; Merlos-Suarez, A.; Stephan-Otto Attolini, C.; Sancho, E.; et al. A genome editing approach to study cancer stem cells in human tumors. EMBO Mol. Med. 2017, 9, 869–879. [Google Scholar] [CrossRef]
- Vermeulen, L.; Snippert, H.J. Stem cell dynamics in homeostasis and cancer of the intestine. Nat. Rev. Cancer 2014, 14, 468–480. [Google Scholar] [CrossRef] [PubMed]
- Batlle, E.; Clevers, H. Cancer stem cells revisited. Nat. Med. 2017, 23, 1124–1134. [Google Scholar] [CrossRef] [PubMed]
- Shivdasani, R.A.; Clevers, H.; de Sauvage, F.J. Tissue regeneration: Reserve or reverse? Science 2021, 371, 784–786. [Google Scholar] [CrossRef] [PubMed]
- De Sousa e Melo, F.; Kurtova, A.V.; Harnoss, J.M.; Kljavin, N.; Hoeck, J.D.; Hung, J.; Anderson, J.E.; Storm, E.E.; Modrusan, Z.; Koeppen, H.; et al. A distinct role for Lgr5(+) stem cells in primary and metastatic colon cancer. Nature 2017, 543, 676–680. [Google Scholar] [CrossRef] [PubMed]
- Shimokawa, M.; Ohta, Y.; Nishikori, S.; Matano, M.; Takano, A.; Fujii, M.; Date, S.; Sugimoto, S.; Kanai, T.; Sato, T. Visualization and targeting of LGR5(+) human colon cancer stem cells. Nature 2017, 545, 187–192. [Google Scholar] [CrossRef]
- Dessen, P.; Huelsken, J.; Ordonez-Moran, P. Specific Gene Expression in Lgr5(+) Stem Cells by Using Cre-Lox Recombination. Methods Mol. Biol. 2020, 2171, 249–255. [Google Scholar] [CrossRef]
- Oost, K.C.; van Voorthuijsen, L.; Fumagalli, A.; Lindeboom, R.G.H.; Sprangers, J.; Omerzu, M.; Rodriguez-Colman, M.J.; Heinz, M.C.; Verlaan-Klink, I.; Maurice, M.M.; et al. Specific Labeling of Stem Cell Activity in Human Colorectal Organoids Using an ASCL2-Responsive Minigene. Cell Rep. 2018, 22, 1600–1614. [Google Scholar] [CrossRef]
- Polakis, P. Wnt signaling in cancer. Cold Spring Harb. Perspect Biol. 2012, 4. [Google Scholar] [CrossRef]
- Merenda, A.; Fenderico, N.; Maurice, M.M. Wnt Signaling in 3D: Recent Advances in the Applications of Intestinal Organoids. Trends Cell Biol. 2020, 30, 60–73. [Google Scholar] [CrossRef] [PubMed]
- Michels, B.E.; Mosa, M.H.; Grebbin, B.M.; Yepes, D.; Darvishi, T.; Hausmann, J.; Urlaub, H.; Zeuzem, S.; Kvasnicka, H.M.; Oellerich, T.; et al. Human colon organoids reveal distinct physiologic and oncogenic Wnt responses. J. Exp. Med. 2019, 216, 704–720. [Google Scholar] [CrossRef]
- Spit, M.; Fenderico, N.; Jordens, I.; Radaszkiewicz, T.; Lindeboom, R.G.; Bugter, J.M.; Cristobal, A.; Ootes, L.; van Osch, M.; Janssen, E.; et al. RNF43 truncations trap CK1 to drive niche-independent self-renewal in cancer. EMBO J. 2020, 39, e103932. [Google Scholar] [CrossRef]
- Tsukiyama, T.; Zou, J.; Kim, J.; Ogamino, S.; Shino, Y.; Masuda, T.; Merenda, A.; Matsumoto, M.; Fujioka, Y.; Hirose, T.; et al. A phospho-switch controls RNF43-mediated degradation of Wnt receptors to suppress tumorigenesis. Nat. Commun. 2020, 11, 4586. [Google Scholar] [CrossRef]
- Giebel, N.; de Jaime-Soguero, A.; Garcia Del Arco, A.; Landry, J.J.M.; Tietje, M.; Villacorta, L.; Benes, V.; Fernandez-Saiz, V.; Acebron, S.P. USP42 protects ZNRF3/RNF43 from R-spondin-dependent clearance and inhibits Wnt signalling. EMBO Rep. 2021, e51415. [Google Scholar] [CrossRef]
- Adam, R.S.; van Neerven, S.M.; Pleguezuelos-Manzano, C.; Simmini, S.; Leveille, N.; de Groot, N.E.; Holding, A.N.; Markowetz, F.; Vermeulen, L. Intestinal region-specific Wnt signalling profiles reveal interrelation between cell identity and oncogenic pathway activity in cancer development. Cancer Cell Int. 2020, 20, 578. [Google Scholar] [CrossRef] [PubMed]
- Gan, T.; Stevens, A.T.; Xiong, X.; Wen, Y.A.; Farmer, T.N.; Li, A.T.; Stevens, P.D.; Golshani, S.; Weiss, H.L.; Mark Evers, B.; et al. Inhibition of protein tyrosine phosphatase receptor type F suppresses Wnt signaling in colorectal cancer. Oncogene 2020, 39, 6789–6801. [Google Scholar] [CrossRef] [PubMed]
- Tran, T.Q.; Hanse, E.A.; Habowski, A.N.; Li, H.; Gabra, M.B.I.; Yang, Y.; Lowman, X.H.; Ooi, A.M.; Liao, S.Y.; Edwards, R.A.; et al. alpha-Ketoglutarate attenuates Wnt signaling and drives differentiation in colorectal cancer. Nat. Cancer 2020, 1, 345–358. [Google Scholar] [CrossRef]
- Dietinger, V.; Garcia de Durango, C.R.; Wiechmann, S.; Boos, S.L.; Michl, M.; Neumann, J.; Hermeking, H.; Kuster, B.; Jung, P. Wnt-driven LARGE2 mediates laminin-adhesive O-glycosylation in human colonic epithelial cells and colorectal cancer. Cell Commun. Signal. 2020, 18, 102. [Google Scholar] [CrossRef] [PubMed]
- Gbenedio, O.M.; Bonnans, C.; Grun, D.; Wang, C.Y.; Hatch, A.J.; Mahoney, M.R.; Barras, D.; Matli, M.; Miao, Y.; Garcia, K.C.; et al. RasGRP1 is a potential biomarker to stratify anti-EGFR therapy response in colorectal cancer. JCI Insight 2019, 5. [Google Scholar] [CrossRef]
- Cheung, P.; Xiol, J.; Dill, M.T.; Yuan, W.C.; Panero, R.; Roper, J.; Osorio, F.G.; Maglic, D.; Li, Q.; Gurung, B.; et al. Regenerative Reprogramming of the Intestinal Stem Cell State via Hippo Signaling Suppresses Metastatic Colorectal Cancer. Cell Stem Cell 2020, 27, 590–604.e599. [Google Scholar] [CrossRef]
- Joosten, S.P.J.; Zeilstra, J.; van Andel, H.; Mijnals, R.C.; Zaunbrecher, J.; Duivenvoorden, A.A.M.; van de Wetering, M.; Clevers, H.; Spaargaren, M.; Pals, S.T. MET Signaling Mediates Intestinal Crypt-Villus Development, Regeneration, and Adenoma Formation and Is Promoted by Stem Cell CD44 Isoforms. Gastroenterology 2017, 153, 1040–1053.e1044. [Google Scholar] [CrossRef]
- Joosten, S.P.J.; Spaargaren, M.; Clevers, H.; Pals, S.T. Hepatocyte growth factor/MET and CD44 in colorectal cancer: Partners in tumorigenesis and therapy resistance. Biochim. Biophys. Acta Rev. Cancer 2020, 1874, 188437. [Google Scholar] [CrossRef] [PubMed]
- Regan, J.L.; Schumacher, D.; Staudte, S.; Steffen, A.; Haybaeck, J.; Keilholz, U.; Schweiger, C.; Golob-Schwarzl, N.; Mumberg, D.; Henderson, D.; et al. Non-Canonical Hedgehog Signaling Is a Positive Regulator of the WNT Pathway and Is Required for the Survival of Colon Cancer Stem Cells. Cell Rep. 2017, 21, 2813–2828. [Google Scholar] [CrossRef] [PubMed]
- Ponsioen, B.; Post, J.B.; Buissant des Amorie, J.R.; Laskaris, D.; van Ineveld, R.L.; Kersten, S.; Bertotti, A.; Sassi, F.; Sipieter, F.; Cappe, B.; et al. Quantifying single-cell ERK dynamics in colorectal cancer organoids reveals EGFR as an amplifier of oncogenic MAPK pathway signalling. Nat. Cell Biol. 2021, 23, 377–390. [Google Scholar] [CrossRef]
- Fessler, E.; Drost, J.; van Hooff, S.R.; Linnekamp, J.F.; Wang, X.; Jansen, M.; De Sousa, E.M.F.; Prasetyanti, P.R.; JE, I.J.; Franitza, M.; et al. TGFbeta signaling directs serrated adenomas to the mesenchymal colorectal cancer subtype. EMBO Mol. Med. 2016, 8, 745–760. [Google Scholar] [CrossRef] [PubMed]
- Ringel, T.; Frey, N.; Ringnalda, F.; Janjuha, S.; Cherkaoui, S.; Butz, S.; Srivatsa, S.; Pirkl, M.; Russo, G.; Villiger, L.; et al. Genome-Scale CRISPR Screening in Human Intestinal Organoids Identifies Drivers of TGF-beta Resistance. Cell Stem Cell 2020, 26, 431–440.e438. [Google Scholar] [CrossRef]
- Michels, B.E.; Mosa, M.H.; Streibl, B.I.; Zhan, T.; Menche, C.; Abou-El-Ardat, K.; Darvishi, T.; Czlonka, E.; Wagner, S.; Winter, J.; et al. Pooled In Vitro and In Vivo CRISPR-Cas9 Screening Identifies Tumor Suppressors in Human Colon Organoids. Cell Stem Cell 2020, 26, 782–792.e787. [Google Scholar] [CrossRef]
- Kraiczy, J.; Nayak, K.M.; Howell, K.J.; Ross, A.; Forbester, J.; Salvestrini, C.; Mustata, R.; Perkins, S.; Andersson-Rolf, A.; Leenen, E.; et al. DNA methylation defines regional identity of human intestinal epithelial organoids and undergoes dynamic changes during development. Gut 2019, 68, 49–61. [Google Scholar] [CrossRef]
- Drost, J.; Clevers, H. Translational applications of adult stem cell-derived organoids. Development 2017, 144, 968–975. [Google Scholar] [CrossRef]
- Colomer, C.; Margalef, P.; Gonzalez, J.; Vert, A.; Bigas, A.; Espinosa, L. IKKalpha is required in the intestinal epithelial cells for tumour stemness. Br. J. Cancer 2018, 118, 839–846. [Google Scholar] [CrossRef]
- Marruecos, L.; Bertran, J.; Guillen, Y.; Gonzalez, J.; Batlle, R.; Lopez-Arribillaga, E.; Garrido, M.; Ruiz-Herguido, C.; Lisiero, D.; Gonzalez-Farre, M.; et al. IkappaBalpha deficiency imposes a fetal phenotype to intestinal stem cells. EMBO Rep. 2020, 21, e49708. [Google Scholar] [CrossRef]
- Leung, M.L.; Davis, A.; Gao, R.; Casasent, A.; Wang, Y.; Sei, E.; Vilar, E.; Maru, D.; Kopetz, S.; Navin, N.E. Single-cell DNA sequencing reveals a late-dissemination model in metastatic colorectal cancer. Genome Res. 2017, 27, 1287–1299. [Google Scholar] [CrossRef]
- Roerink, S.F.; Sasaki, N.; Lee-Six, H.; Young, M.D.; Alexandrov, L.B.; Behjati, S.; Mitchell, T.J.; Grossmann, S.; Lightfoot, H.; Egan, D.A.; et al. Intra-tumour diversification in colorectal cancer at the single-cell level. Nature 2018, 556, 457–462. [Google Scholar] [CrossRef]
- Okamoto, T.; duVerle, D.; Yaginuma, K.; Natsume, Y.; Yamanaka, H.; Kusama, D.; Fukuda, M.; Yamamoto, M.; Perraudeau, F.; Srivastava, U.; et al. Comparative Analysis of Patient-Matched PDOs Revealed a Reduction in OLFM4-Associated Clusters in Metastatic Lesions in Colorectal Cancer. Stem Cell Rep. 2021. [Google Scholar] [CrossRef] [PubMed]
- Costales-Carrera, A.; Fernandez-Barral, A.; Bustamante-Madrid, P.; Dominguez, O.; Guerra-Pastrian, L.; Cantero, R.; Del Peso, L.; Burgos, A.; Barbachano, A.; Munoz, A. Comparative Study of Organoids from Patient-Derived Normal and Tumor Colon and Rectal Tissue. Cancers 2020, 12, 2302. [Google Scholar] [CrossRef]
- Granat, L.M.; Kambhampati, O.; Klosek, S.; Niedzwecki, B.; Parsa, K.; Zhang, D. The promises and challenges of patient-derived tumor organoids in drug development and precision oncology. Anim. Model. Exp. Med. 2019, 2, 150–161. [Google Scholar] [CrossRef]
- Aboulkheyr Es, H.; Montazeri, L.; Aref, A.R.; Vosough, M.; Baharvand, H. Personalized Cancer Medicine: An Organoid Approach. Trends Biotechnol. 2018, 36, 358–371. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Izpisua Belmonte, J.C. Organoids—Preclinical Models of Human Disease. N. Engl. J. Med. 2019, 380, 569–579. [Google Scholar] [CrossRef]
- Ooft, S.N.; Weeber, F.; Dijkstra, K.K.; McLean, C.M.; Kaing, S.; van Werkhoven, E.; Schipper, L.; Hoes, L.; Vis, D.J.; van de Haar, J.; et al. Patient-derived organoids can predict response to chemotherapy in metastatic colorectal cancer patients. Sci. Transl. Med. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Pauli, C.; Hopkins, B.D.; Prandi, D.; Shaw, R.; Fedrizzi, T.; Sboner, A.; Sailer, V.; Augello, M.; Puca, L.; Rosati, R.; et al. Personalized In Vitro and In Vivo Cancer Models to Guide Precision Medicine. Cancer Discov. 2017, 7, 462–477. [Google Scholar] [CrossRef]
- Marshall, L.J.; Triunfol, M.; Seidle, T. Patient-Derived Xenograft vs. Organoids: A Preliminary Analysis of Cancer Research Output, Funding and Human Health Impact in 2014–2019. Animals 2020, 10, 1923. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Huang, J.; Ning, B.; Liu, Z.; Chen, S.; Zhao, W. Drug Discovery via Human-Derived Stem Cell Organoids. Front. Pharmacol. 2016, 7, 334. [Google Scholar] [CrossRef] [PubMed]
- Clotworthy, M.; Archibald, K. Advances in the development and use of human tissue-based techniques for drug toxicity testing. Expert Opin Drug Metab. Toxicol. 2013, 9, 1155–1169. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.; Rettenmeier, E.; Paszek, M.; Yueh, M.F.; Tukey, R.H.; Trottier, J.; Barbier, O.; Chen, S. Crypt Organoid Culture as an in Vitro Model in Drug Metabolism and Cytotoxicity Studies. Drug Metab. Dispos. 2017, 45, 748–754. [Google Scholar] [CrossRef]
- Christensen, S.; Van der Roest, B.; Besselink, N.; Janssen, R.; Boymans, S.; Martens, J.W.M.; Yaspo, M.L.; Priestley, P.; Kuijk, E.; Cuppen, E.; et al. 5-Fluorouracil treatment induces characteristic T>G mutations in human cancer. Nat. Commun. 2019, 10, 4571. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.H.; Ro, E.J.; Yoon, J.S.; Mizutani, T.; Kang, D.W.; Park, J.C.; Il Kim, T.; Clevers, H.; Choi, K.Y. 5-FU promotes stemness of colorectal cancer via p53-mediated WNT/beta-catenin pathway activation. Nat. Commun. 2020, 11, 5321. [Google Scholar] [CrossRef] [PubMed]
- Engel, R.M.; Chan, W.H.; Nickless, D.; Hlavca, S.; Richards, E.; Kerr, G.; Oliva, K.; McMurrick, P.J.; Jarde, T.; Abud, H.E. Patient-Derived Colorectal Cancer Organoids Upregulate Revival Stem Cell Marker Genes following Chemotherapeutic Treatment. J. Clin. Med. 2020, 9, 128. [Google Scholar] [CrossRef]
- Rae, C.; Amato, F.; Braconi, C. Patient-Derived Organoids as a Model for Cancer Drug Discovery. Int. J. Mol. Sci. 2021, 22, 3483. [Google Scholar] [CrossRef]
- Ashley, N.; Jones, M.; Ouaret, D.; Wilding, J.; Bodmer, W.F. Rapidly derived colorectal cancer cultures recapitulate parental cancer characteristics and enable personalized therapeutic assays. J. Pathol. 2014, 234, 34–45. [Google Scholar] [CrossRef]
- Boehnke, K.; Iversen, P.W.; Schumacher, D.; Lallena, M.J.; Haro, R.; Amat, J.; Haybaeck, J.; Liebs, S.; Lange, M.; Schafer, R.; et al. Assay Establishment and Validation of a High-Throughput Screening Platform for Three-Dimensional Patient-Derived Colon Cancer Organoid Cultures. J. Biomol. Screen. 2016, 21, 931–941. [Google Scholar] [CrossRef]
- Verissimo, C.S.; Overmeer, R.M.; Ponsioen, B.; Drost, J.; Mertens, S.; Verlaan-Klink, I.; Gerwen, B.V.; van der Ven, M.; Wetering, M.V.; Egan, D.A.; et al. Targeting mutant RAS in patient-derived colorectal cancer organoids by combinatorial drug screening. Elife 2016, 5. [Google Scholar] [CrossRef]
- Schutte, M.; Risch, T.; Abdavi-Azar, N.; Boehnke, K.; Schumacher, D.; Keil, M.; Yildiriman, R.; Jandrasits, C.; Borodina, T.; Amstislavskiy, V.; et al. Molecular dissection of colorectal cancer in pre-clinical models identifies biomarkers predicting sensitivity to EGFR inhibitors. Nat. Commun. 2017, 8, 14262. [Google Scholar] [CrossRef] [PubMed]
- Lukonin, I.; Serra, D.; Challet Meylan, L.; Volkmann, K.; Baaten, J.; Zhao, R.; Meeusen, S.; Colman, K.; Maurer, F.; Stadler, M.B.; et al. Phenotypic landscape of intestinal organoid regeneration. Nature 2020, 586, 275–280. [Google Scholar] [CrossRef]
- Norkin, M.; Ordonez-Moran, P.; Huelsken, J. High-content, targeted RNA-seq screening in organoids for drug discovery in colorectal cancer. Cell Rep. 2021, 35, 109026. [Google Scholar] [CrossRef] [PubMed]
- Costales-Carrera, A.; Fernandez-Barral, A.; Bustamante-Madrid, P.; Guerra, L.; Cantero, R.; Barbachano, A.; Munoz, A. Plocabulin Displays Strong Cytotoxic Activity in a Personalized Colon Cancer Patient-Derived 3D Organoid Assay. Mar. Drugs 2019, 17, 648. [Google Scholar] [CrossRef]
- Cai, T.; Qi, Y.; Jergens, A.; Wannemuehler, M.; Barrett, T.A.; Wang, Q. Effects of six common dietary nutrients on murine intestinal organoid growth. PLoS ONE 2018, 13, e0191517. [Google Scholar] [CrossRef]
- Marchelletta, R.; Yu, J.; Moon, C.; Kim, M.M. Evaluation of Compound Activity in Primary Human Intestinal Organoids Using Gene Expression and Histology. Curr Protoc. Pharmacol. 2019, 85, e54. [Google Scholar] [CrossRef]
- Gonzalez-Sancho, J.M.; Larriba, M.J.; Munoz, A. Wnt and Vitamin D at the Crossroads in Solid Cancer. Cancers 2020, 12, 3434. [Google Scholar] [CrossRef] [PubMed]
- Schwank, G.; Koo, B.K.; Sasselli, V.; Dekkers, J.F.; Heo, I.; Demircan, T.; Sasaki, N.; Boymans, S.; Cuppen, E.; van der Ent, C.K.; et al. Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell 2013, 13, 653–658. [Google Scholar] [CrossRef] [PubMed]
- Dekkers, J.F.; Wiegerinck, C.L.; de Jonge, H.R.; Bronsveld, I.; Janssens, H.M.; de Winter-de Groot, K.M.; Brandsma, A.M.; de Jong, N.W.; Bijvelds, M.J.; Scholte, B.J.; et al. A functional CFTR assay using primary cystic fibrosis intestinal organoids. Nat. Med. 2013, 19, 939–945. [Google Scholar] [CrossRef] [PubMed]
- DeHaan, R.K.; Sarvestani, S.K.; Huang, E.H. Organoid Models of Colorectal Pathology: Do They Hold the Key to Personalized Medicine? A Systematic Review. Dis. Colon Rectum. 2020, 63, 1559–1569. [Google Scholar] [CrossRef]
- Vlachogiannis, G.; Hedayat, S.; Vatsiou, A.; Jamin, Y.; Fernandez-Mateos, J.; Khan, K.; Lampis, A.; Eason, K.; Huntingford, I.; Burke, R.; et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 2018, 359, 920–926. [Google Scholar] [CrossRef] [PubMed]
- Ganesh, K.; Wu, C.; O’Rourke, K.P.; Szeglin, B.C.; Zheng, Y.; Sauve, C.G.; Adileh, M.; Wasserman, I.; Marco, M.R.; Kim, A.S.; et al. A rectal cancer organoid platform to study individual responses to chemoradiation. Nat. Med. 2019, 25, 1607–1614. [Google Scholar] [CrossRef] [PubMed]
- Pasch, C.A.; Favreau, P.F.; Yueh, A.E.; Babiarz, C.P.; Gillette, A.A.; Sharick, J.T.; Karim, M.R.; Nickel, K.P.; DeZeeuw, A.K.; Sprackling, C.M.; et al. Patient-Derived Cancer Organoid Cultures to Predict Sensitivity to Chemotherapy and Radiation. Clin. Cancer Res. 2019, 25, 5376–5387. [Google Scholar] [CrossRef] [PubMed]
- Driehuis, E.; Kolders, S.; Spelier, S.; Lohmussaar, K.; Willems, S.M.; Devriese, L.A.; de Bree, R.; de Ruiter, E.J.; Korving, J.; Begthel, H.; et al. Oral Mucosal Organoids as a Potential Platform for Personalized Cancer Therapy. Cancer Discov. 2019, 9, 852–871. [Google Scholar] [CrossRef] [PubMed]
- Driehuis, E.; van Hoeck, A.; Moore, K.; Kolders, S.; Francies, H.E.; Gulersonmez, M.C.; Stigter, E.C.A.; Burgering, B.; Geurts, V.; Gracanin, A.; et al. Pancreatic cancer organoids recapitulate disease and allow personalized drug screening. Proc. Natl. Acad. Sci. USA 2019. [Google Scholar] [CrossRef]
- Yao, Y.; Xu, X.; Yang, L.; Zhu, J.; Wan, J.; Shen, L.; Xia, F.; Fu, G.; Deng, Y.; Pan, M.; et al. Patient-Derived Organoids Predict Chemoradiation Responses of Locally Advanced Rectal Cancer. Cell Stem Cell 2020, 26, 17–26.e16. [Google Scholar] [CrossRef]
- Bruun, J.; Kryeziu, K.; Eide, P.W.; Moosavi, S.H.; Eilertsen, I.A.; Langerud, J.; Rosok, B.; Totland, M.Z.; Brunsell, T.H.; Pellinen, T.; et al. Patient-Derived Organoids from Multiple Colorectal Cancer Liver Metastases Reveal Moderate Intra-patient Pharmacotranscriptomic Heterogeneity. Clin. Cancer Res. 2020, 26, 4107–4119. [Google Scholar] [CrossRef]
- Driehuis, E.; Kretzschmar, K.; Clevers, H. Establishment of patient-derived cancer organoids for drug-screening applications. Nat. Protoc. 2020, 15, 3380–3409. [Google Scholar] [CrossRef]
- Crespo, M.; Vilar, E.; Tsai, S.Y.; Chang, K.; Amin, S.; Srinivasan, T.; Zhang, T.; Pipalia, N.H.; Chen, H.J.; Witherspoon, M.; et al. Colonic organoids derived from human induced pluripotent stem cells for modeling colorectal cancer and drug testing. Nat. Med. 2017, 23, 878–884. [Google Scholar] [CrossRef]
- Ooft, S.N.; Weeber, F.; Schipper, L.; Dijkstra, K.K.; McLean, C.M.; Kaing, S.; van de Haar, J.; Prevoo, W.; van Werkhoven, E.; Snaebjornsson, P.; et al. Prospective experimental treatment of colorectal cancer patients based on organoid drug responses. ESMO Open 2021, 6, 100103. [Google Scholar] [CrossRef]
- Arnold, D.; Lueza, B.; Douillard, J.Y.; Peeters, M.; Lenz, H.J.; Venook, A.; Heinemann, V.; Van Cutsem, E.; Pignon, J.P.; Tabernero, J.; et al. Prognostic and predictive value of primary tumour side in patients with RAS wild-type metastatic colorectal cancer treated with chemotherapy and EGFR directed antibodies in six randomized trials. Ann. Oncol. 2017, 28, 1713–1729. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Li, L.; Zhang, X.; Gao, G.; Shen, L.; Hu, J.; Yang, M.; Liu, B.; Qian, X. FOLFOX plus anti-epidermal growth factor receptor (EGFR) monoclonal antibody (mAb) is an effective first-line treatment for patients with RAS-wild left-sided metastatic colorectal cancer: A meta-analysis. Medicine 2018, 97, e0097. [Google Scholar] [CrossRef]
- Kucukoner, M.; Oztekin, E.; Akdeniz, N.; Morkuzu, S.; Yerlikaya, H.; Urakci, Z.; Kaplan, M.A.; Isikdogan, A. Prognostic importance of tumor location and anti-EGFR therapy in patients with K-RAS wild type metastatic colorectal cancer. J. BUON 2019, 24, 1501–1506. [Google Scholar]
- Sahin, S.; Karatas, F. The impact of primary tumor localization on survival and treatment outcomes in patients with metastatic colorectal cancer-a multicenter study. J. BUON 2019, 24, 479–487. [Google Scholar]
- Yan, H.H.N.; Siu, H.C.; Ho, S.L.; Yue, S.S.K.; Gao, Y.; Tsui, W.Y.; Chan, D.; Chan, A.S.; Wong, J.W.H.; Man, A.H.Y.; et al. Organoid cultures of early-onset colorectal cancers reveal distinct and rare genetic profiles. Gut 2020, 69, 2165–2179. [Google Scholar] [CrossRef]
- Bozzi, F.; Mogavero, A.; Varinelli, L.; Belfiore, A.; Manenti, G.; Caccia, C.; Volpi, C.C.; Beznoussenko, G.V.; Milione, M.; Leoni, V.; et al. MIF/CD74 axis is a target for novel therapies in colon carcinomatosis. J. Exp. Clin. Cancer Res. 2017, 36, 16. [Google Scholar] [CrossRef]
- Roy, P.; Canet-Jourdan, C.; Annereau, M.; Zajac, O.; Gelli, M.; Broutin, S.; Mercier, L.; Paci, A.; Lemare, F.; Ducreux, M.; et al. Organoids as preclinical models to improve intraperitoneal chemotherapy effectiveness for colorectal cancer patients with peritoneal metastases: Preclinical models to improve HIPEC. Int. J. Pharm. 2017, 531, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Ubink, I.; Bolhaqueiro, A.C.F.; Elias, S.G.; Raats, D.A.E.; Constantinides, A.; Peters, N.A.; Wassenaar, E.C.E.; de Hingh, I.; Rovers, K.P.; van Grevenstein, W.M.U.; et al. Organoids from colorectal peritoneal metastases as a platform for improving hyperthermic intraperitoneal chemotherapy. Br. J. Surg. 2019, 106, 1404–1414. [Google Scholar] [CrossRef] [PubMed]
- Votanopoulos, K.I.; Mazzocchi, A.; Sivakumar, H.; Forsythe, S.; Aleman, J.; Levine, E.A.; Skardal, A. Appendiceal Cancer Patient-Specific Tumor Organoid Model for Predicting Chemotherapy Efficacy Prior to Initiation of Treatment: A Feasibility Study. Ann. Surg. Oncol. 2019, 26, 139–147. [Google Scholar] [CrossRef]
- Phan, N.; Hong, J.J.; Tofig, B.; Mapua, M.; Elashoff, D.; Moatamed, N.A.; Huang, J.; Memarzadeh, S.; Damoiseaux, R.; Soragni, A. A simple high-throughput approach identifies actionable drug sensitivities in patient-derived tumor organoids. Commun. Biol. 2019, 2, 78. [Google Scholar] [CrossRef] [PubMed]
- Forsythe, S.D.; Sasikumar, S.; Moaven, O.; Sivakumar, H.; Shen, P.; Levine, E.A.; Soker, S.; Skardal, A.; Votanopoulos, K.I. Personalized Identification of Optimal HIPEC Perfusion Protocol in Patient-Derived Tumor Organoid Platform. Ann. Surg. Oncol. 2020, 27, 4950–4960. [Google Scholar] [CrossRef] [PubMed]
- Narasimhan, V.; Wright, J.A.; Churchill, M.; Wang, T.; Rosati, R.; Lannagan, T.R.M.; Vrbanac, L.; Richardson, A.B.; Kobayashi, H.; Price, T.; et al. Medium-throughput Drug Screening of Patient-derived Organoids from Colorectal Peritoneal Metastases to Direct Personalized Therapy. Clin. Cancer Res. 2020, 26, 3662–3670. [Google Scholar] [CrossRef]
- Gjorevski, N.; Sachs, N.; Manfrin, A.; Giger, S.; Bragina, M.E.; Ordonez-Moran, P.; Clevers, H.; Lutolf, M.P. Designer matrices for intestinal stem cell and organoid culture. Nature 2016, 539, 560–564. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Acuna, R.; Quiros, M.; Farkas, A.E.; Dedhia, P.H.; Huang, S.; Siuda, D.; Garcia-Hernandez, V.; Miller, A.J.; Spence, J.R.; Nusrat, A.; et al. Synthetic hydrogels for human intestinal organoid generation and colonic wound repair. Nat. Cell Biol. 2017, 19, 1326–1335. [Google Scholar] [CrossRef]
- Hernandez-Gordillo, V.; Kassis, T.; Lampejo, A.; Choi, G.; Gamboa, M.E.; Gnecco, J.S.; Brown, A.; Breault, D.T.; Carrier, R.; Griffith, L.G. Fully synthetic matrices for in vitro culture of primary human intestinal enteroids and endometrial organoids. Biomaterials 2020, 254, 120125. [Google Scholar] [CrossRef]
- Tayler, I.M.; Stowers, R.S. Engineering Hydrogels for Personalized Disease Modeling and Regenerative Medicine. Acta Biomater. 2021. [Google Scholar] [CrossRef]
- AlMusawi, S.; Ahmed, M.; Nateri, A.S. Understanding cell-cell communication and signaling in the colorectal cancer microenvironment. Clin. Transl. Med. 2021, 11, e308. [Google Scholar] [CrossRef]
- Ootani, A.; Li, X.; Sangiorgi, E.; Ho, Q.T.; Ueno, H.; Toda, S.; Sugihara, H.; Fujimoto, K.; Weissman, I.L.; Capecchi, M.R.; et al. Sustained in vitro intestinal epithelial culture within a Wnt-dependent stem cell niche. Nat. Med. 2009, 15, 701–706. [Google Scholar] [CrossRef]
- Lannagan, T.R.; Jackstadt, R.; Leedham, S.J.; Sansom, O.J. Advances in colon cancer research: In Vitro and animal models. Curr Opin Genet. Dev. 2021, 66, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Reidy, E.; Leonard, N.A.; Treacy, O.; Ryan, A.E. A 3D View of Colorectal Cancer Models in Predicting Therapeutic Responses and Resistance. Cancers 2021, 13, 227. [Google Scholar] [CrossRef]
- Pastula, A.; Middelhoff, M.; Brandtner, A.; Tobiasch, M.; Hohl, B.; Nuber, A.H.; Demir, I.E.; Neupert, S.; Kollmann, P.; Mazzuoli-Weber, G.; et al. Three-Dimensional Gastrointestinal Organoid Culture in Combination with Nerves or Fibroblasts: A Method to Characterize the Gastrointestinal Stem Cell Niche. Stem Cells Int. 2016, 2016, 3710836. [Google Scholar] [CrossRef]
- Kabiri, Z.; Greicius, G.; Madan, B.; Biechele, S.; Zhong, Z.; Zaribafzadeh, H.; Aliyev, J.; Wu, Y.; Bunte, R.; Williams, B.O.; et al. Stroma provides an intestinal stem cell niche in the absence of epithelial Wnts. Development 2014, 141, 2206–2215. [Google Scholar] [CrossRef]
- Hirokawa, Y.; Yip, K.H.; Tan, C.W.; Burgess, A.W. Colonic myofibroblast cell line stimulates colonoid formation. Am. J. Physiol. Gastrointest Liver Physiol. 2014, 306, G547–G556. [Google Scholar] [CrossRef]
- Yin, Y.; Zhou, D. Organoid and Enteroid Modeling of Salmonella Infection. Front. Cell Infect. Microbiol. 2018, 8, 102. [Google Scholar] [CrossRef]
- McCarthy, N.; Kraiczy, J.; Shivdasani, R.A. Cellular and molecular architecture of the intestinal stem cell niche. Nat. Cell Biol. 2020, 22, 1033–1041. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, N.; Manieri, E.; Storm, E.E.; Saadatpour, A.; Luoma, A.M.; Kapoor, V.N.; Madha, S.; Gaynor, L.T.; Cox, C.; Keerthivasan, S.; et al. Distinct Mesenchymal Cell Populations Generate the Essential Intestinal BMP Signaling Gradient. Cell Stem Cell 2020, 26, 391–402. [Google Scholar] [CrossRef] [PubMed]
- Szvicsek, Z.; Oszvald, A.; Szabo, L.; Sandor, G.O.; Kelemen, A.; Soos, A.A.; Paloczi, K.; Harsanyi, L.; Tolgyes, T.; Dede, K.; et al. Extracellular vesicle release from intestinal organoids is modulated by Apc mutation and other colorectal cancer progression factors. Cell Mol. Life Sci. 2019, 76, 2463–2476. [Google Scholar] [CrossRef] [PubMed]
- Oszvald, A.; Szvicsek, Z.; Sandor, G.O.; Kelemen, A.; Soos, A.A.; Paloczi, K.; Bursics, A.; Dede, K.; Tolgyes, T.; Buzas, E.I.; et al. Extracellular vesicles transmit epithelial growth factor activity in the intestinal stem cell niche. Stem Cells 2020, 38, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Mosa, M.H.; Michels, B.E.; Menche, C.; Nicolas, A.M.; Darvishi, T.; Greten, F.R.; Farin, H.F. A Wnt-Induced Phenotypic Switch in Cancer-Associated Fibroblasts Inhibits EMT in Colorectal Cancer. Cancer Res. 2020, 80, 5569–5582. [Google Scholar] [CrossRef]
- Qin, X.; Sufi, J.; Vlckova, P.; Kyriakidou, P.; Acton, S.E.; Li, V.S.W.; Nitz, M.; Tape, C.J. Cell-type-specific signaling networks in heterocellular organoids. Nat. Methods 2020, 17, 335–342. [Google Scholar] [CrossRef] [PubMed]
- Noel, G.; Baetz, N.W.; Staab, J.F.; Donowitz, M.; Kovbasnjuk, O.; Pasetti, M.F.; Zachos, N.C. A primary human macrophage-enteroid co-culture model to investigate mucosal gut physiology and host-pathogen interactions. Sci. Rep. 2017, 7, 45270. [Google Scholar] [CrossRef]
- Rogoz, A.; Reis, B.S.; Karssemeijer, R.A.; Mucida, D. A 3-D enteroid-based model to study T-cell and epithelial cell interaction. J. Immunol. Methods 2015, 421, 89–95. [Google Scholar] [CrossRef]
- Nozaki, K.; Mochizuki, W.; Matsumoto, Y.; Matsumoto, T.; Fukuda, M.; Mizutani, T.; Watanabe, M.; Nakamura, T. Co-culture with intestinal epithelial organoids allows efficient expansion and motility analysis of intraepithelial lymphocytes. J. Gastroenterol. 2016, 51, 206–213. [Google Scholar] [CrossRef]
- Neal, J.T.; Li, X.; Zhu, J.; Giangarra, V.; Grzeskowiak, C.L.; Ju, J.; Liu, I.H.; Chiou, S.H.; Salahudeen, A.A.; Smith, A.R.; et al. Organoid Modeling of the Tumor Immune Microenvironment. Cell 2018, 175, 1972–1988.e1916. [Google Scholar] [CrossRef]
- Jenkins, R.W.; Aref, A.R.; Lizotte, P.H.; Ivanova, E.; Stinson, S.; Zhou, C.W.; Bowden, M.; Deng, J.; Liu, H.; Miao, D.; et al. Ex Vivo Profiling of PD-1 Blockade Using Organotypic Tumor Spheroids. Cancer Discov. 2018, 8, 196–215. [Google Scholar] [CrossRef]
- Dijkstra, K.K.; Cattaneo, C.M.; Weeber, F.; Chalabi, M.; van de Haar, J.; Fanchi, L.F.; Slagter, M.; van der Velden, D.L.; Kaing, S.; Kelderman, S.; et al. Generation of Tumor-Reactive T Cells by Co-culture of Peripheral Blood Lymphocytes and Tumor Organoids. Cell 2018, 174, 1586–1598.e1512. [Google Scholar] [CrossRef] [PubMed]
- Cattaneo, C.M.; Dijkstra, K.K.; Fanchi, L.F.; Kelderman, S.; Kaing, S.; van Rooij, N.; van den Brink, S.; Schumacher, T.N.; Voest, E.E. Tumor organoid-T-cell coculture systems. Nat. Protoc. 2020, 15, 15–39. [Google Scholar] [CrossRef]
- Schnalzger, T.E.; de Groot, M.H.; Zhang, C.; Mosa, M.H.; Michels, B.E.; Roder, J.; Darvishi, T.; Wels, W.S.; Farin, H.F. 3D model for CAR-mediated cytotoxicity using patient-derived colorectal cancer organoids. EMBO J. 2019, 38. [Google Scholar] [CrossRef] [PubMed]
- Bar-Ephraim, Y.E.; Kretzschmar, K.; Clevers, H. Organoids in immunological research. Nat. Rev. Immunol. 2020, 20, 279–293. [Google Scholar] [CrossRef] [PubMed]
- Allaire, J.M.; Poon, A.; Crowley, S.M.; Han, X.; Sharafian, Z.; Moore, N.; Stahl, M.; Bressler, B.; Lavoie, P.M.; Jacobson, K.; et al. Interleukin-37 regulates innate immune signaling in human and mouse colonic organoids. Sci. Rep. 2021, 11, 8206. [Google Scholar] [CrossRef] [PubMed]
- Dotti, I.; Mora-Buch, R.; Ferrer-Picon, E.; Planell, N.; Jung, P.; Masamunt, M.C.; Leal, R.F.; Martin de Carpi, J.; Llach, J.; Ordas, I.; et al. Alterations in the epithelial stem cell compartment could contribute to permanent changes in the mucosa of patients with ulcerative colitis. Gut 2017, 66, 2069–2079. [Google Scholar] [CrossRef]
- D’Aldebert, E.; Quaranta, M.; Sebert, M.; Bonnet, D.; Kirzin, S.; Portier, G.; Duffas, J.P.; Chabot, S.; Lluel, P.; Allart, S.; et al. Characterization of Human Colon Organoids From Inflammatory Bowel Disease Patients. Front. Cell Dev. Biol. 2020, 8, 363. [Google Scholar] [CrossRef] [PubMed]
- Li, V.S.W. Modelling intestinal inflammation and infection using ‘mini-gut’ organoids. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 89–90. [Google Scholar] [CrossRef]
- Cook, L.; Stahl, M.; Han, X.; Nazli, A.; MacDonald, K.N.; Wong, M.Q.; Tsai, K.; Dizzell, S.; Jacobson, K.; Bressler, B.; et al. Suppressive and Gut-Reparative Functions of Human Type 1 T Regulatory Cells. Gastroenterology 2019, 157, 1584–1598. [Google Scholar] [CrossRef]
- Kasendra, M.; Tovaglieri, A.; Sontheimer-Phelps, A.; Jalili-Firoozinezhad, S.; Bein, A.; Chalkiadaki, A.; Scholl, W.; Zhang, C.; Rickner, H.; Richmond, C.A.; et al. Development of a primary human Small Intestine-on-a-Chip. using biopsy-derived organoids. Sci. Rep. 2018, 8, 2871. [Google Scholar] [CrossRef] [PubMed]
- Brassard, J.A.; Lutolf, M.P. Engineering Stem Cell Self-organization to Build Better Organoids. Cell Stem Cell 2019, 24, 860–876. [Google Scholar] [CrossRef] [PubMed]
- Bein, A.; Shin, W.; Jalili-Firoozinezhad, S.; Park, M.H.; Sontheimer-Phelps, A.; Tovaglieri, A.; Chalkiadaki, A.; Kim, H.J.; Ingber, D.E. Microfluidic Organ-on-a-Chip. Models of Human Intestine. Cell Mol. Gastroenterol. Hepatol. 2018, 5, 659–668. [Google Scholar] [CrossRef] [PubMed]
- Takebe, T.; Zhang, B.; Radisic, M. Synergistic Engineering: Organoids Meet Organs-on-a-Chip. Cell Stem Cell 2017, 21, 297–300. [Google Scholar] [CrossRef]
- Park, S.E.; Georgescu, A.; Huh, D. Organoids-on-a-chip. Science 2019, 364, 960–965. [Google Scholar] [CrossRef]
- Sontheimer-Phelps, A.; Chou, D.B.; Tovaglieri, A.; Ferrante, T.C.; Duckworth, T.; Fadel, C.; Frismantas, V.; Sutherland, A.D.; Jalili-Firoozinezhad, S.; Kasendra, M.; et al. Human Colon-on-a-Chip. Enables Continuous In Vitro Analysis of Colon Mucus Layer Accumulation and Physiology. Cell Mol. Gastroenterol. Hepatol. 2020, 9, 507–526. [Google Scholar] [CrossRef]
- Nikolaev, M.; Mitrofanova, O.; Broguiere, N.; Geraldo, S.; Dutta, D.; Tabata, Y.; Elci, B.; Brandenberg, N.; Kolotuev, I.; Gjorevski, N.; et al. Homeostatic mini-intestines through scaffold-guided organoid morphogenesis. Nature 2020, 585, 574–578. [Google Scholar] [CrossRef]
- Verhulsel, M.; Simon, A.; Bernheim-Dennery, M.; Gannavarapu, V.R.; Geremie, L.; Ferraro, D.; Krndija, D.; Talini, L.; Viovy, J.L.; Vignjevic, D.M.; et al. Developing an advanced gut on chip model enabling the study of epithelial cell/fibroblast interactions. Lab. Chip. 2021, 21, 365–377. [Google Scholar] [CrossRef]
- Rajasekar, S.; Lin, D.S.Y.; Abdul, L.; Liu, A.; Sotra, A.; Zhang, F.; Zhang, B. IFlowPlate-A Customized 384-Well Plate for the Culture of Perfusable Vascularized Colon Organoids. Adv. Mater. 2020, 32, e2002974. [Google Scholar] [CrossRef]
- Skardal, A.; Shupe, T.; Atala, A. Organoid-on-a-chip and body-on-a-chip systems for drug screening and disease modeling. Drug Discov. Today 2016, 21, 1399–1411. [Google Scholar] [CrossRef] [PubMed]
- Pinho, D.; Santos, D.; Vila, A.; Carvalho, S. Establishment of Colorectal Cancer Organoids in Microfluidic-Based System. Micromachines 2021, 12, 497. [Google Scholar] [CrossRef]
- Carvalho, M.R.; Barata, D.; Teixeira, L.M.; Giselbrecht, S.; Reis, R.L.; Oliveira, J.M.; Truckenmuller, R.; Habibovic, P. Colorectal tumor-on-a-chip system: A 3D tool for precision onco-nanomedicine. Sci. Adv. 2019, 5, eaaw1317. [Google Scholar] [CrossRef] [PubMed]
- Oke, S.; Martin, A. Insights into the role of the intestinal microbiota in colon cancer. Therap Adv. Gastroenterol. 2017, 10, 417–428. [Google Scholar] [CrossRef]
- Allen, J.; Sears, C.L. Impact of the gut microbiome on the genome and epigenome of colon epithelial cells: Contributions to colorectal cancer development. Genome Med. 2019, 11, 11. [Google Scholar] [CrossRef]
- Co, J.Y.; Margalef-Catala, M.; Li, X.; Mah, A.T.; Kuo, C.J.; Monack, D.M.; Amieva, M.R. Controlling Epithelial Polarity: A Human Enteroid Model for Host-Pathogen Interactions. Cell Rep. 2019, 26, 2509–2520.e2504. [Google Scholar] [CrossRef] [PubMed]
- Nalluri, H.; Subramanian, S.; Staley, C. Intestinal organoids: A model to study the role of microbiota in the colonic tumor microenvironment. Future Microbiol. 2020, 15, 1583–1594. [Google Scholar] [CrossRef]
- Moysidou, C.M.; Owens, R.M. Advances in modelling the human microbiome-gut-brain axis in vitro. Biochem. Soc. Trans. 2021, 49, 187–201. [Google Scholar] [CrossRef] [PubMed]
- Gazzaniga, F.S.; Camacho, D.M.; Wu, M.; Silva Palazzo, M.F.; Dinis, A.L.M.; Grafton, F.N.; Cartwright, M.J.; Super, M.; Kasper, D.L.; Ingber, D.E. Harnessing Colon Chip. Technology to Identify Commensal Bacteria That Promote Host Tolerance to Infection. Front. Cell Infect. Microbiol. 2021, 11, 638014. [Google Scholar] [CrossRef] [PubMed]
- Pleguezuelos-Manzano, C.; Puschhof, J.; Rosendahl Huber, A.; van Hoeck, A.; Wood, H.M.; Nomburg, J.; Gurjao, C.; Manders, F.; Dalmasso, G.; Stege, P.B.; et al. Mutational signature in colorectal cancer caused by genotoxic pks(+) E. coli. Nature 2020, 580, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, T. Recent progress in organoid culture to model intestinal epithelial barrier functions. Int. Immunol. 2019, 31, 13–21. [Google Scholar] [CrossRef]
- Geurts, M.H.; van der Vaart, J.; Beumer, J.; Clevers, H. The Organoid Platform: Promises and Challenges as Tools in the Fight against COVID-19. Stem Cell Rep. 2021, 16, 412–418. [Google Scholar] [CrossRef]
- Zhao, X.; Li, C.; Liu, X.; Chiu, M.C.; Wang, D.; Wei, Y.; Chu, H.; Cai, J.P.; Hau-Yee Chan, I.; Kak-Yuen Wong, K.; et al. Human Intestinal Organoids Recapitulate Enteric Infections of Enterovirus and Coronavirus. Stem Cell Rep. 2021, 16, 493–504. [Google Scholar] [CrossRef]
| Reagent | Function | Human | Mouse | |
|---|---|---|---|---|
| Normal | Tumor | Normal | ||
| Advanced DMEM/F12 | Basal medium | + | + | + |
| HEPES | pH buffer | + | + | + |
| Glutamax | L-glutamine supplier | + | + | + |
| B27 supplement | Vitamins/cofactors mix | + | + | + |
| N2 supplement * | Vitamins/cofactors mix | +/− | +/− | − |
| N-acetyl-L-cysteine | Antioxidant | + | + | + |
| Antibiotics | To avoid bacterial contamination | + | + | + |
| Wnt3A-cond. medium | Stemness/Wnt signaling activator | + | + #/− | + |
| RSPO | Stemness/Wnt signaling enhancer | + | + #/− | + |
| Nicotinamide * | Stemness/cystic phenotype | +/− | +/− | − |
| PGE2 * | Wnt activator/cystic phenotype | +/− | +/− | − |
| EGF | Mitogen | + | + | + |
| Gastrin | Mitogen | + | + | − |
| FGF-2 | Mitogen | + ¥/− | + ¥/− | − |
| IGF-1 | Mitogen | + ¥/− | + ¥/− | − |
| Noggin | BMP inhibitor | + | + | + |
| Y-27632 † | ROCK/anoikis inhibitor | + | + | + |
| SB202190 | p38MAPK inhibitor | +/− ¥ | + #/− ¥ | − |
| A 83-01/LY2157299 | TGF-β inhibitor | + | + | − |
| Features | Cell Lines | PDX | PDTO |
|---|---|---|---|
| Initiation success rate | Low | Moderate | High |
| Possibility of expansion | Very high | Moderate | High |
| Time consuming | Low | High | Moderate |
| Cost | Low | Very high | High |
| Expertise | Low | High | High |
| Ease of maintenance | High | Moderate | Moderate |
| Amenability for genetic manipulation | High | Low | High |
| Ease of downstream assays | High | Low | High |
| High-throughput drug screening | Very high | Low | High |
| Low-throughput drug screening | Very high | Moderate | Very high |
| Modelling early stage CRC | No | No | Yes |
| Autologous normal controls | No | No | Yes |
| Isogenic control by genome editing | Yes | No | Yes |
| Patient-specific | No | Yes | Yes |
| Save animal testing | Yes | No | Yes |
| Recapitulation of tissue features | Low | High | High |
| Need of specialized facilities | Low | High | Moderate |
| Cell-matrix interaction | No | Yes | Yes |
| Lack of microenvironment | Yes | No * | Yes |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barbáchano, A.; Fernández-Barral, A.; Bustamante-Madrid, P.; Prieto, I.; Rodríguez-Salas, N.; Larriba, M.J.; Muñoz, A. Organoids and Colorectal Cancer. Cancers 2021, 13, 2657. https://doi.org/10.3390/cancers13112657
Barbáchano A, Fernández-Barral A, Bustamante-Madrid P, Prieto I, Rodríguez-Salas N, Larriba MJ, Muñoz A. Organoids and Colorectal Cancer. Cancers. 2021; 13(11):2657. https://doi.org/10.3390/cancers13112657
Chicago/Turabian StyleBarbáchano, Antonio, Asunción Fernández-Barral, Pilar Bustamante-Madrid, Isabel Prieto, Nuria Rodríguez-Salas, María Jesús Larriba, and Alberto Muñoz. 2021. "Organoids and Colorectal Cancer" Cancers 13, no. 11: 2657. https://doi.org/10.3390/cancers13112657
APA StyleBarbáchano, A., Fernández-Barral, A., Bustamante-Madrid, P., Prieto, I., Rodríguez-Salas, N., Larriba, M. J., & Muñoz, A. (2021). Organoids and Colorectal Cancer. Cancers, 13(11), 2657. https://doi.org/10.3390/cancers13112657







