Untwining Anti-Tumor and Immunosuppressive Effects of JAK Inhibitors—A Strategy for Hematological Malignancies?
Simple Summary
Abstract
1. JAK-STAT Signaling in Hematological Malignancies
2. JAK-STAT Signaling in the Immune System
3. The Effects of JAKinibs on the Immune System: Focus on Cytotoxic Lymphocytes
3.1. Ruxolitinib
3.2. Tofacitinib
3.3. Fedratinib
3.4. Momelotinib
3.5. Pacritinib
3.6. Itacitinib
4. When an Immune Cell Becomes Cancerous—Hijacking JAKinibs’ Immunosuppressive Side Effects for Treatment of NK/T-Cell Tumors
5. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schindler, C.; Levy, D.E.; Decker, T. JAK-STAT Signaling: From Interferons to Cytokines. J. Biol. Chem. 2007, 282, 20059–20063. [Google Scholar] [CrossRef] [PubMed]
- O’Shea, J.J.; Plenge, R. JAK and STAT Signaling Molecules in Immunoregulation and Immune-Mediated Disease. Immunity 2012, 36, 542–550. [Google Scholar] [CrossRef]
- Stark, G.R.; Darnell, J.E., Jr. The JAK-STAT Pathway at Twenty. Immunity 2012, 36, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Hammarén, H.M.; Virtanen, A.T.; Raivola, J.; Silvennoinen, O. The regulation of JAKs in cytokine signaling and its breakdown in disease. Cytokine 2019, 118, 48–63. [Google Scholar] [CrossRef]
- Villarino, A.V.; Kanno, Y.; O’Shea, A.V.V.Y.K.J.J. Mechanisms and consequences of Jak–STAT signaling in the immune system. Nat. Immunol. 2017, 18, 374–384. [Google Scholar] [CrossRef] [PubMed]
- Levy, D.E.; Darnell, J.E., Jr. STATs: Transcriptional control and biological impact. Nat. Rev. Mol. Cell Biol. 2002, 3, 651–662. [Google Scholar] [CrossRef]
- Shuai, K.; Liu, B. Regulation of JAK–STAT signalling in the immune system. Nat. Rev. Immunol. 2003, 3, 900–911. [Google Scholar] [CrossRef] [PubMed]
- Kiu, H.; Nicholson, S.E. Biology and significance of the JAK/STAT signalling pathways. Growth Factors 2012, 30, 88–106. [Google Scholar] [CrossRef]
- Morris, R.; Kershaw, N.J.; Babon, J.J. The molecular details of cytokine signaling via the JAK/STAT pathway. Protein Sci. 2018, 27, 1984–2009. [Google Scholar] [CrossRef]
- Villarino, A.; Kanno, Y.; Ferdinand, J.R.; O’Shea, J.J. Mechanisms of Jak/STAT Signaling in Immunity and Disease. J. Immunol. 2015, 194, 21–27. [Google Scholar] [CrossRef]
- McLornan, D.P.; Khan, A.A.; Harrison, C.N. Immunological Consequences of JAK Inhibition: Friend or Foe? Curr. Hematol. Malign- Rep. 2015, 10, 370–379. [Google Scholar] [CrossRef]
- Gadina, M.; Johnson, C.; Schwartz, D.; Bonelli, M.; Hasni, S.; Kanno, Y.; Changelian, P.; Laurence, A.; O’Shea, J.J. Translational and clinical advances in JAK-STAT biology: The present and future of jakinibs. J. Leukoc. Biol. 2018, 104, 499–514. [Google Scholar] [CrossRef]
- Casanova, J.-L.; Holland, S.M.; Notarangelo, L.D. Inborn Errors of Human JAKs and STATs. Immunity 2012, 36, 515–528. [Google Scholar] [CrossRef]
- O’Shea, J.J.; Holland, S.M.; Staudt, L.M. JAKs and STATs in Immunity, Immunodeficiency, and Cancer. New Engl. J. Med. 2013, 368, 161–170. [Google Scholar] [CrossRef]
- Senkevitch, E.; Durum, S. The promise of Janus kinase inhibitors in the treatment of hematological malignancies. Cytokine 2017, 98, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, D.M.; Kanno, Y.; Villarino, A.; Ward, M.; Gadina, M.; O’Shea, J.J. JAK inhibition as a therapeutic strategy for immune and inflammatory diseases. Nat. Rev. Drug Discov. 2017, 16, 843–862. [Google Scholar] [CrossRef]
- Chen, E.; Staudt, L.M.; Green, A.R. Janus Kinase Deregulation in Leukemia and Lymphoma. Immunity 2012, 36, 529–541. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S.J.; Snowden, J.A.; Zeidler, M.; Danson, S. The role of JAK/STAT signalling in the pathogenesis, prognosis and treatment of solid tumours. Br. J. Cancer 2015, 113, 365–371. [Google Scholar] [CrossRef]
- Igelmann, S.; Neubauer, H.A.; Ferbeyre, G. STAT3 and STAT5 Activation in Solid Cancers. Cancers 2019, 11, 1428. [Google Scholar] [CrossRef] [PubMed]
- Saharinen, P.; Takaluoma, K.; Silvennoinen, O. Regulation of the Jak2 Tyrosine Kinase by Its Pseudokinase Domain. Mol. Cell. Biol. 2000, 20, 3387–3395. [Google Scholar] [CrossRef]
- Saharinen, P.; Vihinen, M.; Silvennoinen, O. Autoinhibition of Jak2 Tyrosine Kinase Is Dependent on Specific Regions in Its Pseudokinase Domain. Mol. Biol. Cell 2003, 14, 1448–1459. [Google Scholar] [CrossRef]
- Toms, A.V.; Deshpande, A.; McNally, R.; Jeong, Y.; Rogers, J.M.; Kim, C.U.; Gruner, S.M.; Ficarro, S.B.; A Marto, J.; Sattler, M.; et al. Structure of a pseudokinase-domain switch that controls oncogenic activation of Jak kinases. Nat. Struct. Mol. Biol. 2013, 20, 1221–1223. [Google Scholar] [CrossRef] [PubMed]
- Lupardus, P.J.; Ultsch, M.; Wallweber, H.; Kohli, P.B.; Johnson, A.R.; Eigenbrot, C. Structure of the pseudokinase-kinase domains from protein kinase TYK2 reveals a mechanism for Janus kinase (JAK) autoinhibition. Proc. Natl. Acad. Sci. 2014, 111, 8025–8030. [Google Scholar] [CrossRef] [PubMed]
- Ferrao, R.; Lupardus, P.J. The Janus Kinase (JAK) FERM and SH2 Domains: Bringing Specificity to JAK–Receptor Interactions. Front. Endocrinol. 2017, 8, 71. [Google Scholar] [CrossRef]
- Staerk, J.; Kallin, A.; Demoulin, J.-B.; Vainchenker, W.; Constantinescu, S.N. JAK1 and Tyk2 Activation by the Homologous Polycythemia Vera JAK2 V617F Mutation. J. Biol. Chem. 2005, 280, 41893–41899. [Google Scholar] [CrossRef] [PubMed]
- Baxter, E.J.; Scott, L.M.; Campbell, P.J.; East, C.; Fourouclas, N.; Swanton, S.; Vassiliou, G.; Bench, A.J.; Boyd, E.M.; Curtin, N.; et al. Acquired mutation of the tyrosine kinase JAK2 in human myeloproliferative disorders. Lancet 2005, 365, 1054–1061. [Google Scholar] [CrossRef]
- E James, C.; Ugo, V.; Le Couédic, J.-P.; Staerk, J.; Delhommeau, F.; Lacout, C.; Garçon, L.; Raslova, H.; Berger, R.; Bennaceur-Griscelli, A.; et al. A unique clonal JAK2 mutation leading to constitutive signalling causes polycythaemia vera. Nat. Cell Biol. 2005, 434, 1144–1148. [Google Scholar] [CrossRef] [PubMed]
- Kralovics, R.; Passamonti, F.; Buser, A.S.; Teo, S.-S.; Tiedt, R.; Passweg, J.R.; Tichelli, A.; Cazzola, M.; Skoda, R.C. A Gain-of-Function Mutation ofJAK2in Myeloproliferative Disorders. New Engl. J. Med. 2005, 352, 1779–1790. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Kim, Y.G.; Soung, Y.H.; Han, K.J.; Kim, S.Y.; Rhim, H.; Min, W.S.; Nam, S.W.; Park, W.S.; Yoo, N.J.; et al. The JAK2 V617F mutation in de novo acute myelogenous leukemias. Oncogene 2005, 25, 1434–1436. [Google Scholar] [CrossRef] [PubMed]
- Passamonti, F.; Rumi, E.; Pietra, D.; Della Porta, M.G.; Boveri, E.; Pascutto, C.; Vanelli, L.; Arcaini, L.; Burcheri, S.; Malcovati, L.; et al. Relation between JAK2 (V617F) mutation status, granulocyte activation, and constitutive mobilization of CD34+ cells into peripheral blood in myeloproliferative disorders. Blood 2006, 107, 3676–3682. [Google Scholar] [CrossRef]
- Mascarenhas, J.; Hoffman, R. Ruxolitinib: The First FDA Approved Therapy for the Treatment of Myelofibrosis: Figure 1. Clin. Cancer Res. 2012, 18, 3008–3014. [Google Scholar] [CrossRef] [PubMed]
- Vainchenker, W.; Constantinescu, S. JAK/STAT signaling in hematological malignancies. Oncogene 2012, 32, 2601–2613. [Google Scholar] [CrossRef]
- Raedler, L.A. Jakafi (Ruxolitinib): First FDA-Approved Medication for the Treatment of Patients with Polycythemia Vera. Am. Health Drug benefits 2015, 8, 75–79. [Google Scholar]
- Skoda, R.C.; Duek, A.; Grisouard, J. Pathogenesis of myeloproliferative neoplasms. Exp. Hematol. 2015, 43, 599–608. [Google Scholar] [CrossRef] [PubMed]
- De Noronha, T.R.; Mitne-Neto, M.; Chauffaille, M.D.L. JAK2-mutated acute myeloid leukemia: Comparison of next-generation sequencing (NGS) and single nucleotide polymorphism array (SNPa) findings between two cases. Autops. Case Rep. 2019, 9, e2018084. [Google Scholar] [CrossRef]
- Mead, A.J.; Rugless, M.J.; Jacobsen, S.E.W.; Schuh, A. GermlineJAK2Mutation in a Family with Hereditary Thrombocytosis. New Engl. J. Med. 2012, 366, 967–969. [Google Scholar] [CrossRef] [PubMed]
- Etheridge, S.L.; Cosgrove, M.E.; Sangkhae, V.; Corbo, L.M.; Roh, M.E.; Seeliger, M.A.; Chan, E.L.; Hitchcock, I.S. A novel activating, germline JAK2 mutation, JAK2R564Q, causes familial essential thrombocytosis. Blood 2014, 123, 1059–1068. [Google Scholar] [CrossRef]
- Marty, C.; Saint-Martin, C.; Pecquet, C.; Grosjean, S.; Saliba, J.; Mouton, C.; Leroy, E.; Harutyunyan, A.S.; Abgrall, J.-F.; Favier, R.; et al. Germ-line JAK2 mutations in the kinase domain are responsible for hereditary thrombocytosis and are resistant to JAK2 and HSP90 inhibitors. Blood 2014, 123, 1372–1383. [Google Scholar] [CrossRef] [PubMed]
- Jeong, E.G.; Kim, M.S.; Nam, H.K.; Min, C.K.; Lee, S.; Chung, Y.J.; Yoo, N.J.; Lee, S.H. Somatic Mutations of JAK1 and JAK3 in Acute Leukemias and Solid Cancers. Clin. Cancer Res. 2008, 14, 3716–3721. [Google Scholar] [CrossRef]
- Mullighan, C.G.; Zhang, J.; Harvey, R.C.; Collins-Underwood, J.R.; Schulman, B.A.; Phillips, L.A.; Tasian, S.K.; Loh, M.L.; Su, X.; Liu, W.; et al. JAK mutations in high-risk childhood acute lymphoblastic leukemia. Proc. Natl. Acad. Sci. 2009, 106, 9414–9418. [Google Scholar] [CrossRef]
- Bellanger, D.E.; Jacquemin, V.; Chopin, M.; Pierron, G.; A Bernard, O.; Ghysdael, J.; Stern, M.-H. Recurrent JAK1 and JAK3 somatic mutations in T-cell prolymphocytic leukemia. Leukemia 2014, 28, 417–419. [Google Scholar] [CrossRef] [PubMed]
- Arulogun, S.O.; Choong, H.-L.; Taylor, D.; Ambrosoli, P.; Magor, G.; Irving, I.M.; Keng, T.-B.; Perkins, A.C. JAK1 somatic mutation in a myeloproliferative neoplasm. Haematologica 2017, 102, e324–e327. [Google Scholar] [CrossRef][Green Version]
- Lee, S.; Park, H.Y.; Kang, S.Y.; Kim, S.J.; Hwang, J.; Lee, S.; Kwak, S.H.; Park, K.S.; Yoo, H.Y.; Kim, W.S.; et al. Genetic alterations of JAK/STAT cascade and histone modification in extranodal NK/T-cell lymphoma nasal type. Oncotarget 2015, 6, 17764–17776. [Google Scholar] [CrossRef]
- Ross, J.A.; Kirken, R.A. Transforming Mutations of Jak3 (A573V and M511I) Show Differential Sensitivity to Selective Jak3 Inhibitors. Clin. Cancer Drugs 2016, 3, 131–137. [Google Scholar] [CrossRef]
- Sim, S.H.; Kim, S.; Kim, T.M.; Jeon, Y.K.; Nam, S.J.; Ahn, Y.-O.; Keam, B.; Park, H.H.; Kim, D.-W.; Kim, C.W.; et al. Novel JAK3-Activating Mutations in Extranodal NK/T-Cell Lymphoma, Nasal Type. Am. J. Pathol. 2017, 187, 980–986. [Google Scholar] [CrossRef] [PubMed]
- Kiyoi, H.; Yamaji, S.; Kojima, S.; Naoe, T. JAK3 mutations occur in acute megakaryoblastic leukemia both in Down syndrome children and non-Down syndrome adults. Leukemia 2007, 21, 574–576. [Google Scholar] [CrossRef]
- Malinge, S.; Ragu, C.; Della-Valle, V.; Pisani, D.; Constantinescu, S.N.; Perez, C.; Villeval, J.-L.; Reinhardt, D.; Landman-Parker, J.; Michaux, L.; et al. Activating mutations in human acute megakaryoblastic leukemia. Blood 2008, 112, 4220–4226. [Google Scholar] [CrossRef]
- Yamashita, Y.; Yuan, J.; Suetake, I.; Suzuki, H.; Ishikawa, Y.; Choi, Y.L.; Ueno, T.; Soda, M.; Hamada, T.; Haruta, H.; et al. Array-based genomic resequencing of human leukemia. Oncogene 2010, 29, 3723–3731. [Google Scholar] [CrossRef] [PubMed]
- Bains, T.; Heinrich, M.C.; Loriaux, M.M.; Beadling, C.; Nelson, D.; Warrick, A.; Neff, T.L.; Tyner, J.W.; Dunlap, J.; Corless, C.L.; et al. Newly described activating JAK3 mutations in T-cell acute lymphoblastic leukemia. Leukemia 2012, 26, 2144–2146. [Google Scholar] [CrossRef] [PubMed]
- Koo, G.C.; Tan, S.Y.; Tang, T.; Poon, S.L.; Allen, G.E.; Tan, L.; Chong, S.C.; Ong, W.S.; Tay, K.; Tao, M.; et al. Janus Kinase 3–Activating Mutations Identified in Natural Killer/T-cell Lymphoma. Cancer Discov. 2012, 2, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ding, L.; Holmfeldt, L.; Wu, G.; Heatley, S.L.; Payne-Turner, D.; Easton, J.; Chen, X.; Wang, J.; Rusch, M.; et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 2012, 481, 157–163. [Google Scholar] [CrossRef]
- Bergmann, A.K.; Schneppenheim, S.; Seifert, M.; Betts, M.J.; Haake, A.; Lopez, C.; Penas, E.M.M.; Vater, I.; Jayne, S.; Dyer, M.J.; et al. Recurrent mutation ofJAK3in T-cell prolymphocytic leukemia. Genes, Chromosom. Cancer 2014, 53, 309–316. [Google Scholar] [CrossRef] [PubMed]
- Bouchekioua, A.; Scourzic, L.; De Wever, O.; Zhang, Y.; Cervera, P.; Aline-Fardin, A.; Mercher, T.; Gaulard, P.; Nyga, R.; Jeziorowska, D.; et al. JAK3 deregulation by activating mutations confers invasive growth advantage in extranodal nasal-type natural killer cell lymphoma. Leukemia 2014, 28, 338–348. [Google Scholar] [CrossRef]
- Sanda, T.; Tyner, J.W.; Gutierrez, A.; Ngo, V.N.; Glover, J.; Chang, B.H.; Yost, A.; Ma, W.; Fleischman, A.; Zhou, W.; et al. TYK2–STAT1–BCL2 Pathway Dependence in T-cell Acute Lymphoblastic Leukemia. Cancer Discov. 2013, 3, 564–577. [Google Scholar] [CrossRef] [PubMed]
- Waanders, E.; Scheijen, B.; Jongmans, M.; Venselaar, H.; Van Reijmersdal, S.; Van Dijk, A.; Pastorczak, A.; Weren, R.; Van Der Schoot, C.; Van De Vorst, J.; et al. Germline activating TYK2 mutations in pediatric patients with two primary acute lymphoblastic leukemia occurrences. Leukemia 2017, 31, 821–828. [Google Scholar] [CrossRef]
- Wöss, K.; Simonović, N.; Strobl, B.; Macho-Maschler, S.; Müller, M. TYK2: An Upstream Kinase of STATs in Cancer. Cancers 2019, 11, 1728. [Google Scholar] [CrossRef]
- Scott, L.M. The JAK2 exon 12 mutations: A comprehensive review. Am. J. Hematol. 2011, 86, 668–676. [Google Scholar] [CrossRef] [PubMed]
- O’Shea, J.J.; Schwartz, D.M.; Villarino, A.; Gadina, M.; McInnes, I.B.; Laurence, A. The JAK-STAT Pathway: Impact on Human Disease and Therapeutic Intervention. Annu. Rev. Med. 2015, 66, 311–328. [Google Scholar] [CrossRef]
- Matutes, E. The 2017 WHO update on mature T- and natural killer (NK) cell neoplasms. Int. J. Lab. Hematol. 2018, 40, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Shahmarvand, N.; Nagy, A.; Shahryari, J.; Ohgami, R.S. Mutations in the signal transducer and activator of transcription family of genes in cancer. Cancer Sci. 2018, 109, 926–933. [Google Scholar] [CrossRef]
- De Araujo, E.D.; Orlova, A.; Neubauer, H.A.; Bajusz, D.; Seo, H.-S.; Dhe-Paganon, S.; Keserű, G.M.; Moriggl, R.; Gunning, P.T. Structural Implications of STAT3 and STAT5 SH2 Domain Mutations. Cancers 2019, 11, 1757. [Google Scholar] [CrossRef] [PubMed]
- De Araujo, E.D.; Erdogan, F.; Neubauer, H.A.; Meneksedag-Erol, D.; Manaswiyoungkul, P.; Eram, M.S.; Seo, H.-S.; Qadree, A.K.; Israelian, J.; Orlova, A.; et al. Structural and functional consequences of the STAT5BN642H driver mutation. Nat. Commun. 2019, 10, 1–15. [Google Scholar] [CrossRef]
- Hu, G.; Witzig, T.E.; Gupta, M. A Novel Missense (M206K) STAT3 Mutation in Diffuse Large B Cell Lymphoma Deregulates STAT3 Signaling. PLoS ONE 2013, 8, e67851. [Google Scholar] [CrossRef] [PubMed]
- Haapaniemi, E.M.; Kaustio, M.; Rajala, H.L.M.; Van Adrichem, A.J.; Kainulainen, L.; Glumoff, V.; Doffinger, R.; Kuusanmäki, H.; Heiskanen-Kosma, T.; Trotta, L.; et al. Autoimmunity, hypogammaglobulinemia, lymphoproliferation, and mycobacterial disease in patients with activating mutations in STAT3. Blood 2015, 125, 639–648. [Google Scholar] [CrossRef]
- Andersson, E.; Kuusanmäki, H.; Bortoluzzi, S.; Lagström, S.; Parsons, A.; Rajala, H.; Van Adrichem, A.; Eldfors, S.; Olson, T.; Clemente, M.J.; et al. Activating somatic mutations outside the SH2-domain of STAT3 in LGL leukemia. Leukemia 2015, 30, 1204–1208. [Google Scholar] [CrossRef]
- Ritz, O.; Guiter, C.; Castellano, F.; Dorsch, K.; Melzner, J.; Jais, J.-P.; Dubois, G.; Gaulard, P.; Möller, P.; Leroy, K. Recurrent mutations of the STAT6 DNA binding domain in primary mediastinal B-cell lymphoma. Blood 2009, 114, 1236–1242. [Google Scholar] [CrossRef] [PubMed]
- Yildiz, M.; Li, H.; Bernard, D.; Amin, N.A.; Ouillette, P.; Jones, S.; Saiya-Cork, K.; Parkin, B.; Jacobi, K.; Shedden, K.; et al. Activating STAT6 mutations in follicular lymphoma. Blood 2015, 125, 668–679. [Google Scholar] [CrossRef]
- Morin, R.D.; Assouline, S.; Alcaide, M.; Mohajeri, A.; Johnston, R.L.; Chong, L.; Grewal, J.; Yu, S.; Fornika, D.; Bushell, K.; et al. Genetic Landscapes of Relapsed and Refractory Diffuse Large B-Cell Lymphomas. Clin. Cancer Res. 2016, 22, 2290–2300. [Google Scholar] [CrossRef]
- Tiacci, E.; Ladewig, E.; Schiavoni, G.; Penson, A.; Fortini, E.; Pettirossi, V.; Wang, Y.; Rosseto, A.; Venanzi, A.; Vlasevska, S.; et al. Pervasive mutations of JAK-STAT pathway genes in classical Hodgkin lymphoma. Blood 2018, 131, 2454–2465. [Google Scholar] [CrossRef] [PubMed]
- Levine, R.L.; Wadleigh, M.; Cools, J.; Ebert, B.L.; Wernig, G.; Huntly, B.J.; Boggon, T.J.; Wlodarska, I.; Clark, J.J.; Moore, S.; et al. Activating mutation in the tyrosine kinase JAK2 in polycythemia vera, essential thrombocythemia, and myeloid metaplasia with myelofibrosis. Cancer Cell 2005, 7, 387–397. [Google Scholar] [CrossRef] [PubMed]
- Helbig, G. Classical Philadelphia-negative myeloproliferative neoplasms: Focus on mutations and JAK2 inhibitors. Med Oncol. 2018, 35, 119. [Google Scholar] [CrossRef]
- Pham, H.T.T.; Maurer, B.; Prchal-Murphy, M.; Grausenburger, R.; Grundschober, E.; Javaheri, T.; Nivarthi, H.; Boersma, A.; Kolbe, T.; Elabd, M.; et al. STAT5BN642H is a driver mutation for T cell neoplasia. J. Clin. Investig. 2017, 128, 387–401. [Google Scholar] [CrossRef]
- Klein, K.; Witalisz-Siepracka, A.; Maurer, B.; Prinz, D.; Heller, G.; Leidenfrost, N.; Prchal-Murphy, M.; Suske, T.; Moriggl, R.; Sexl, V. STAT5BN642H drives transformation of NKT cells: A novel mouse model for CD56+ T-LGL leukemia. Leukemia 2019, 33, 2336–2340. [Google Scholar] [CrossRef] [PubMed]
- Blombery, P.; Thompson, E.R.; Jones, K.; Arnau, G.M.; Lade, S.; Markham, J.F.; Li, J.; Deva, A.; Johnstone, R.W.; Khot, A.; et al. Whole exome sequencing reveals activating JAK1 and STAT3 mutations in breast implant-associated anaplastic large cell lymphoma anaplastic large cell lymphoma. Haematologica 2016, 101, e387–e390. [Google Scholar] [CrossRef]
- Li, Q.; Li, B.; Hu, L.; Ning, H.; Jiang, M.; Wang, D.; Liu, T.; Zhang, B.; Chen, H. Identification of a novel functional JAK1 S646P mutation in acute lymphoblastic leukemia. Oncotarget 2017, 8, 34687–34697. [Google Scholar] [CrossRef] [PubMed]
- Flex, E.; Petrangeli, V.; Stella, L.; Chiaretti, S.; Hornakova, T.; Knoops, L.; Ariola, C.; Fodale, V.; Clappier, E.; Paoloni, F.; et al. Somatically acquired JAK1 mutations in adult acute lymphoblastic leukemia. J. Exp. Med. 2008, 205, 751–758. [Google Scholar] [CrossRef] [PubMed]
- Hornakova, T.; Springuel, L.; Devreux, J.; Dusa, A.; Constantinescu, S.N.; Knoops, L.; Renauld, J.-C. Oncogenic JAK1 and JAK2-activating mutations resistant to ATP-competitive inhibitors. Haematologica 2011, 96, 845–853. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Crescenzo, R.; Abate, F.; Lasorsa, E.; Tabbo’, F.; Gaudiano, M.; Chiesa, N.; Di Giacomo, F.; Spaccarotella, E.; Barbarossa, L.; Ercole, E.; et al. Convergent Mutations and Kinase Fusions Lead to Oncogenic STAT3 Activation in Anaplastic Large Cell Lymphoma. Cancer Cell 2015, 27, 516–532. [Google Scholar] [CrossRef] [PubMed]
- Lesmana, H.; Popescu, M.; Lewis, S.; Sahoo, S.S.; Goodings-Harris, C.; Onciu, M.; Choi, J.K.; Takemoto, C.; Nichols, K.E.; Wlodarski, M. Germline Gain-of-Function JAK3 Mutation in Familial Chronic Lymphoproliferative Disorder of NK Cells. Blood 2020, 136, 9–10. [Google Scholar] [CrossRef]
- Klusmann, J.-H.; Reinhardt, D.; Hasle, H.; Kaspers, G.J.; Creutzig, U.; Hählen, K.; Heuvel-Eibrink, M.M.V.D.; Zwaan, C.M. Janus kinase mutations in the development of acute megakaryoblastic leukemia in children with and without Down’s syndrome. Leukemia 2007, 21, 1584–1587. [Google Scholar] [CrossRef][Green Version]
- Elliott, N.E.; Cleveland, S.M.; Grann, V.; Janik, J.; Waldmann, T.A.; Davé, U.P. FERM domain mutations induce gain of function in JAK3 in adult T-cell leukemia/lymphoma. Blood 2011, 118, 3911–3921. [Google Scholar] [CrossRef]
- Degryse, S.; De Bock, C.E.; Cox, L.; Demeyer, S.; Gielen, O.; Mentens, N.; Jacobs, K.; Geerdens, E.; Gianfelici, V.; Hulselmans, G.; et al. JAK3 mutants transform hematopoietic cells through JAK1 activation, causing T-cell acute lymphoblastic leukemia in a mouse model. Blood 2014, 124, 3092–3100. [Google Scholar] [CrossRef] [PubMed]
- Jerez, A.; Clemente, M.J.; Makishima, H.; Koskela, H.; Leblanc, F.; Ng, K.P.; Olson, T.; Przychodzen, B.; Afable, M.; Gomez-Segui, I.; et al. STAT3 mutations unify the pathogenesis of chronic lymphoproliferative disorders of NK cells and T-cell large granular lymphocyte leukemia. Blood 2012, 120, 3048–3057. [Google Scholar] [CrossRef]
- Koskela, H.L.; Eldfors, S.; Ellonen, P.; Van Adrichem, A.J.; Kuusanmäki, H.; Andersson, E.I.; Lagström, S.; Clemente, M.J.; Olson, T.; Jalkanen, S.E.; et al. SomaticSTAT3Mutations in Large Granular Lymphocytic Leukemia. New Engl. J. Med. 2012, 366, 1905–1913. [Google Scholar] [CrossRef] [PubMed]
- Fasan, A.; Kern, W.; Grossmann, V.; Haferlach, C.; Schnittger, S.; Haferlach, T. STAT3 mutations are highly specific for large granular lymphocytic leukemia. Leukemia 2012, 27, 1598–1600. [Google Scholar] [CrossRef]
- Küçük, C.; Jiang, B.; Hu, X.; Zhang, W.; Chan, J.K.C.; Xiao, W.; Lack, N.; Alkan, C.; Williams, J.C.; Avery, K.N.; et al. Activating mutations of STAT5B and STAT3 in lymphomas derived from γδ-T or NK cells. Nat. Commun. 2015, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Bilori, B.; Thota, S.; Clemente, M.J.; Patel, B.; Jerez, A.; Ii, M.A.; Maciejewski, J.P. Tofacitinib as a novel salvage therapy for refractory T-cell large granular lymphocytic leukemia. Leukemia 2015, 29, 2427–2429. [Google Scholar] [CrossRef]
- Kuusanmäki, H.; Dufva, O.; Parri, E.; Van Adrichem, A.J.; Rajala, H.; Majumder, M.M.; Yadav, B.; Parsons, A.; Chan, W.C.; Wennerberg, K.; et al. Drug sensitivity profiling identifies potential therapies for lymphoproliferative disorders with overactive JAK/STAT3 signaling. Oncotarget 2017, 8, 97516–97527. [Google Scholar] [CrossRef]
- Nicolae, A.; Xi, L.; Pittaluga, S.; Abdullaev, Z.; Pack, S.D.; Chen, J.; A Waldmann, T.; Jaffe, E.S.; Raffeld, M. Frequent STAT5B mutations in γδ hepatosplenic T-cell lymphomas. Leukemia 2014, 28, 2244–2248. [Google Scholar] [CrossRef]
- Babushok, D.V.; Perdigones, N.; Perin, J.C.; Olson, T.S.; Ye, W.; Roth, J.J.; Lind, C.; Cattier, C.; Li, Y.; Hartung, H.; et al. Emergence of clonal hematopoiesis in the majority of patients with acquired aplastic anemia. Cancer Genet. 2015, 208, 115–128. [Google Scholar] [CrossRef]
- Jiang, L.; Gu, Z.-H.; Yan, Z.-X.; Zhao, X.; Xie, Y.-Y.; Zhang, Z.-G.; Pan, C.-M.; Hu, Y.; Cai, C.-P.; Dong, Y.; et al. Exome sequencing identifies somatic mutations of DDX3X in natural killer/T-cell lymphoma. Nat. Genet. 2015, 47, 1061–1066. [Google Scholar] [CrossRef]
- Kiel, M.J.; Velusamy, T.; Rolland, D.; Sahasrabuddhe, A.; Chung, F.; Bailey, N.G.; Schrader, A.; Li, B.; Li, J.Z.; Ozel, A.B.; et al. Integrated genomic sequencing reveals mutational landscape of T-cell prolymphocytic leukemia. Blood 2014, 124, 1460–1472. [Google Scholar] [CrossRef]
- Kiel, M.J.; Sahasrabuddhe, A.A.; Rolland, D.C.M.; Velusamy, T.; Chung, F.; Schaller, M.; Bailey, N.G.; Betz, B.L.; Miranda, R.N.; Porcu, P.; et al. Genomic analyses reveal recurrent mutations in epigenetic modifiers and the JAK–STAT pathway in Sézary syndrome. Nat. Commun. 2015, 6, 8470. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Wen, L.; Wu, L.; Wang, Q.; Yao, H.; Wang, Q.; Ma, L.; Chen, S. Rare occurrence of a STAT5B N642H mutation in adult T-cell acute lymphoblastic leukemia. Cancer Genet. 2015, 208, 52–53. [Google Scholar] [CrossRef]
- Andersson, E.I.; Tanahashi, T.; Sekiguchi, N.; Gasparini, V.R.; Bortoluzzi, S.; Kawakami, T.; Matsuda, K.; Mitsui, T.; Eldfors, S.; Bortoluzzi, S.; et al. High incidence of activating STAT5B mutations in CD4-positive T-cell large granular lymphocyte leukemia. Blood 2016, 128, 2465–2468. [Google Scholar] [CrossRef]
- Nairismägi, M.-L.; Tan, J.; Lim, J.Q.; Nagarajan, S.; Ng, C.C.Y.; Rajasegaran, V.; Huang, D.; Lim, W.K.; Laurensia, Y.; Wijaya, G.C.; et al. JAK-STAT and G-protein-coupled receptor signaling pathways are frequently altered in epitheliotropic intestinal T-cell lymphoma. Leukemia 2016, 30, 1311–1319. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.-M.; Zhao, S.; Liu, W.-P.; Zhang, W.-Y.; Li, G.-D.; Küçük, C.; Hu, X.-Z.; Chan, W.C.; Tang, Y.; Ding, W.-S.; et al. Clinicopathologic Characterization of Aggressive Natural Killer Cell Leukemia Involving Different Tissue Sites. Am. J. Surg. Pathol. 2016, 40, 836–846. [Google Scholar] [CrossRef] [PubMed]
- Teramo, A.; Barilà, G.; Calabretto, G.; Ercolin, C.; Lamy, T.; Moignet, A.; Roussel, M.; Pastoret, C.; Leoncin, M.; Gattazzo, C.; et al. STAT3 mutation impacts biological and clinical features of T-LGL leukemia. Oncotarget 2017, 8, 61876–61889. [Google Scholar] [CrossRef]
- Ma, C.A.; Xi, L.; Cauff, B.; DeZure, A.; Freeman, A.F.; Hambleton, S.; Kleiner, G.; Leahy, T.R.; O’Sullivan, M.; Makiya, M.; et al. Somatic STAT5b gain-of-function mutations in early onset nonclonal eosinophilia, urticaria, dermatitis, and diarrhea. Blood 2017, 129, 650–653. [Google Scholar] [CrossRef] [PubMed]
- Dufva, O.; Kankainen, M.; Kelkka, T.; Sekiguchi, N.; Awad, S.A.; Eldfors, S.; Yadav, B.; Kuusanmäki, H.; Malani, D.; I Andersson, E.; et al. Aggressive natural killer-cell leukemia mutational landscape and drug profiling highlight JAK-STAT signaling as therapeutic target. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Luo, Q.; Shen, J.; Yang, Y.; Tang, H.; Shi, M.; Liu, J.; Liu, Z.; Shi, X.; Yi, Y. CSF3RT618I,ASXL1G942 fs andSTAT5BN642H trimutation co-contribute to a rare chronic neutrophilic leukaemia manifested by rapidly progressive leucocytosis, severe infections, persistent fever and deep venous thrombosis. Br. J. Haematol. 2016, 180, 892–894. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Liu, D.; Wang, N.; Ling, S.; Tang, Y.; Wu, J.; Hao, L.; Luo, H.; Hu, X.; Sheng, L.; et al. Integrated genomic analysis identifies deregulated JAK/STAT-MYC-biosynthesis axis in aggressive NK-cell leukemia. Cell Res. 2017, 28, 172–186. [Google Scholar] [CrossRef]
- Song, T.L.; Nairismägi, M.-L.; Laurensia, Y.; Lim, J.-Q.; Tan, J.; Li, Z.-M.; Pang, W.-L.; Kizhakeyil, A.; Wijaya, G.-C.; Huang, D.; et al. Oncogenic activation of the STAT3 pathway drives PD-L1 expression in natural killer/T-cell lymphoma. Blood 2018, 132, 1146–1158. [Google Scholar] [CrossRef]
- Cross, N.C.P.; Hoade, Y.; Tapper, W.J.; Carreno-Tarragona, G.; Fanelli, T.; Jawhar, M.; Naumann, N.; Pieniak, I.; Lübke, J.; Ali, S.; et al. Recurrent activating STAT5B N642H mutation in myeloid neoplasms with eosinophilia. Leukemia 2018, 33, 415–425. [Google Scholar] [CrossRef]
- Govaerts, I.; Jacobs, K.; Vandepoel, R.; Cools, J. JAK/STAT Pathway Mutations in T-ALL, Including the STAT5B N642H Mutation, are Sensitive to JAK1/JAK3 Inhibitors. HemaSphere 2019, 3, e313. [Google Scholar] [CrossRef] [PubMed]
- Rajala, H.L.M.; Porkka, K.; Maciejewski, J.P.; Loughran, T.P.; Mustjoki, S. Uncovering the pathogenesis of large granular lymphocytic leukemia—novel STAT3 and STAT5b mutations. Ann. Med. 2014, 46, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Rajala, H.L.M.; Eldfors, S.; Kuusanmäki, H.; Van Adrichem, A.J.; Olson, T.; Lagström, S.; Andersson, E.I.; Jerez, A.; Clemente, M.J.; Yan, Y.; et al. Discovery of somatic STAT5b mutations in large granular lymphocytic leukemia. Blood 2013, 121, 4541–4550. [Google Scholar] [CrossRef]
- Bandapalli, O.R.; Schuessele, S.; Kunz, J.B.; Rausch, T.; Stütz, A.M.; Tal, N.; Geron, I.; Gershman, N.; Izraeli, S.; Eilers, J.; et al. The activating STAT5B N642H mutation is a common abnormality in pediatric T-cell acute lymphoblastic leukemia and confers a higher risk of relapse. Haematologica 2014, 99, e188–e192. [Google Scholar] [CrossRef]
- Kontro, M.; Kuusanmäki, H.; Eldfors, S.; Burmeister, T.; Andersson, E.I.; Bruserud, O.; Brümmendorf, T.H.; Edgren, H.; Gjertsen, B.T.; Italaremes, M.; et al. Novel activating STAT5B mutations as putative drivers of T-cell acute lymphoblastic leukemia. Leukemia 2014, 28, 1738–1742. [Google Scholar] [CrossRef]
- Lorenzini, T.; Dotta, L.; Giacomelli, M.; Vairo, D.; Badolato, R. STAT mutations as program switchers: Turning primary immunodeficiencies into autoimmune diseases. J. Leukoc. Biol. 2016, 101, 29–38. [Google Scholar] [CrossRef]
- Kanai, T.; Jenks, J.; Nadeau, K.C. The STAT5b Pathway Defect and Autoimmunity. Front. Immunol. 2012, 3, 234. [Google Scholar] [CrossRef] [PubMed]
- Karjalainen, A.; Shoebridge, S.; Krunic, M.; Simonović, N.; Tebb, G.; Macho-Maschler, S.; Strobl, B.; Müller, M. TYK2 in Tumor Immunosurveillance. Cancers 2020, 12, 150. [Google Scholar] [CrossRef]
- Hambleton, S.; Goodbourn, S.; Young, D.F.; Dickinson, P.; Mohamad, S.M.B.; Valappil, M.; McGovern, N.; Cant, A.J.; Hackett, S.J.; Ghazal, P.; et al. STAT2 deficiency and susceptibility to viral illness in humans. Proc. Natl. Acad. Sci. 2013, 110, 3053–3058. [Google Scholar] [CrossRef]
- Freij, B.J.; Hanrath, A.T.; Chen, R.; Hambleton, S.; Duncan, C.J.A. Life-Threatening Influenza, Hemophagocytic Lymphohistiocytosis and Probable Vaccine-Strain Varicella in a Novel Case of Homozygous STAT2 Deficiency. Front. Immunol. 2021, 11, 11. [Google Scholar] [CrossRef] [PubMed]
- Schimke, L.F.; Hibbard, J.; Martinez-Barricarte, R.; Khan, T.A.; Cavalcante, R.D.S.; Junior, E.B.D.O.; França, T.T.; Iqbal, A.; Yamamoto, G.; Arslanian, C.; et al. Paracoccidioidomycosis Associated With a Heterozygous STAT4 Mutation and Impaired IFN-γ Immunity. J. Infect. Dis. 2017, 216, 1623–1634. [Google Scholar] [CrossRef] [PubMed]
- Powell, D.A.; Shubitz, L.F.; Butkiewicz, C.D.; Moale, H.; Trinh, H.T.; Doetschman, T.; Hsu, A.P.; Holland, S.M.; Galgiani, J.N.; Frelinger, J.A. Modeling a Human STAT4 Mutation That Predisposes to Disseminated Coccidioidomycosis in Mice. J. Immunol. 2020, 204 (Suppl. 1). [Google Scholar]
- Okada, S.; Asano, T.; Moriya, K.; Boisson-Dupuis, S.; Kobayashi, M.; Casanova, J.-L.; Puel, A. Human STAT1 Gain-of-Function Heterozygous Mutations: Chronic Mucocutaneous Candidiasis and Type I Interferonopathy. J. Clin. Immunol. 2020, 40, 1065–1081. [Google Scholar] [CrossRef]
- Kleppe, M.; Spitzer, M.H.; Li, S.; Hill, C.; Dong, L.; Papalexi, E.; De Groote, S.; Bowman, R.L.; Keller, M.; Koppikar, P.; et al. Jak1 Integrates Cytokine Sensing to Regulate Hematopoietic Stem Cell Function and Stress Hematopoiesis. Cell Stem Cell 2017, 21, 489–501.e7. [Google Scholar] [CrossRef] [PubMed]
- Witalisz-Siepracka, A.; Klein, K.; Prinz, D.; Leidenfrost, N.; Schabbauer, G.; Dohnal, A.; Sexl, V. Loss of JAK1 Drives Innate Immune Deficiency. Front. Immunol. 2019, 9, 10. [Google Scholar] [CrossRef]
- Park, S.O.; Wamsley, H.L.; Bae, K.; Hu, Z.; Li, X.; Choe, S.-W.; Slayton, W.B.; Oh, S.P.; Wagner, K.-U.; Sayeski, P.P. Conditional Deletion of Jak2 Reveals an Essential Role in Hematopoiesis throughout Mouse Ontogeny: Implications for Jak2 Inhibition in Humans. PLoS ONE 2013, 8, e59675. [Google Scholar] [CrossRef] [PubMed]
- Betts, B.C.; Bastian, D.; Iamsawat, S.; Nguyen, H.; Heinrichs, J.L.; Wu, Y.; Daenthanasanmak, A.; Veerapathran, A.; O’Mahony, A.; Walton, K.; et al. Targeting JAK2 reduces GVHD and xenograft rejection through regulation of T cell differentiation. Proc. Natl. Acad. Sci. 2018, 115, 1582–1587. [Google Scholar] [CrossRef]
- Simonović, N.; Witalisz-Siepracka, A.; Meissl, K.; Lassnig, C.; Reichart, U.; Kolbe, T.; Farlik, M.; Bock, C.; Sexl, V.; Müller, M.; et al. NK Cells Require Cell-Extrinsic and -Intrinsic TYK2 for Full Functionality in Tumor Surveillance and Antibacterial Immunity. J. Immunol. 2019, 202, 1724–1734. [Google Scholar] [CrossRef]
- Prchal-Murphy, M.; Witalisz-Siepracka, A.; Bednarik, K.T.; Putz, E.M.; Gotthardt, D.; Meissl, K.; Sexl, V.; Müller, M.; Strobl, B. In vivotumor surveillance by NK cells requires TYK2 but not TYK2 kinase activity. OncoImmunology 2015, 4, e1047579. [Google Scholar] [CrossRef]
- Stoiber, D.; Kovacic, B.; Schuster, C.; Schellack, C.; Karaghiosoff, M.; Kreibich, R.; Weisz, E.; Artwohl, M.; Kleine, O.C.; Muller, M.; et al. TYK2 is a key regulator of the surveillance of B lymphoid tumors. J. Clin. Investig. 2004, 114, 1650–1658. [Google Scholar] [CrossRef] [PubMed]
- Eletto, D.; Burns, S.O.; Angulo, I.; Plagnol, V.; Gilmour, K.C.; Henriquez, F.; Curtis, J.; Gaspar, M.; Nowak, K.; Daza-Cajigal, V.; et al. Biallelic JAK1 mutations in immunodeficient patient with mycobacterial infection. Nat. Commun. 2016, 7, 13992. [Google Scholar] [CrossRef] [PubMed]
- Rodig, S.J.; A Meraz, M.; White, J.; A Lampe, P.; Riley, J.K.; Arthur, C.D.; King, K.L.; Sheehan, K.C.; Yin, L.; Pennica, D.; et al. Disruption of the Jak1 Gene Demonstrates Obligatory and Nonredundant Roles of the Jaks in Cytokine-Induced Biologic Responses. Cell 1998, 93, 373–383. [Google Scholar] [CrossRef]
- Macchi, P.; Villa, A.; Giliani, S.; Sacco, M.G.; Frattini, A.; Porta, F.; Ugazio, A.G.; Johnston, J.A.; Candotti, F.; O’Sheai, J.J.; et al. Mutations of Jak-3 gene in patients with autosomal severe combined immune deficiency (SCID). Nat. Cell Biol. 1995, 377, 65–68. [Google Scholar] [CrossRef]
- Russell, S.M.; Tayebi, N.; Nakajima, H.; Riedy, M.C.; Roberts, J.L.; Aman, M.J.; Migone, T.-S.; Noguchi, M.; Markert, M.L.; Buckley, R.H.; et al. Mutation of Jak3 in a Patient with SCID: Essential Role of Jak3 in Lymphoid Development. Science 1995, 270, 797–800. [Google Scholar] [CrossRef] [PubMed]
- Frucht, D.M.; Gadina, M.; Jagadeesh, G.J.; Aksentijevich, I.; Takada, K.; Bleesing, J.; Nelson, J.; Muul, L.M.; Perham, G.; Morgan, G.; et al. Unexpected and variable phenotypes in a family with JAK3 deficiency. Genes Immun. 2001, 2, 422–432. [Google Scholar] [CrossRef] [PubMed]
- Notarangelo, L.D.; Giliani, S.; Mella, P.; Schumacher, R.F.; Mazza, C.; Savoldi, G.; Rodriguez-Perez, C.; Badolato, R.; Mazzolari, E.; Porta, F.; et al. Combined Immunodeficiencies Due to Defects in Signal Transduction: Defects of the γc-JAK3 Signaling Pathway as a Model. Immunobiology 2000, 202, 106–119. [Google Scholar] [CrossRef]
- Robinette, M.L.; Cella, M.; Telliez, J.B.; Ulland, T.K.; Barrow, A.D.; Capuder, K.; Gilfillan, S.; Lin, L.-L.; Notarangelo, L.D.; Colonna, M. Jak3 deficiency blocks innate lymphoid cell development. Mucosal Immunol. 2018, 11, 50–60. [Google Scholar] [CrossRef]
- Baird, A.M.; Thomis, D.C.; Berg, L.J. T cell development and activation in Jak3-deficient mice. J. Leukoc. Biol. 1998, 63, 669–677. [Google Scholar] [CrossRef]
- Thomis, D.C.; Gurniak, C.B.; Tivol, E.; Sharpe, A.H.; Berg, L.J. Defects in B Lymphocyte Maturation and T Lymphocyte Activation in Mice Lacking Jak3. Science 1995, 270, 794–797. [Google Scholar] [CrossRef]
- Kreins, A.Y.; Ciancanelli, M.J.; Okada, S.; Kong, X.-F.; Ramírez-Alejo, N.; Kilic, S.S.; El Baghdadi, J.; Nonoyama, S.; Mahdaviani, S.A.; Ailal, F.; et al. Human TYK2 deficiency: Mycobacterial and viral infections without hyper-IgE syndrome. J. Exp. Med. 2015, 212, 1641–1662. [Google Scholar] [CrossRef] [PubMed]
- Minegishi, Y.; Saito, M.; Morio, T.; Watanabe, K.; Agematsu, K.; Tsuchiya, S.; Takada, H.; Hara, T.; Kawamura, N.; Ariga, T.; et al. Human Tyrosine Kinase 2 Deficiency Reveals Its Requisite Roles in Multiple Cytokine Signals Involved in Innate and Acquired Immunity. Immunity 2006, 25, 745–755. [Google Scholar] [CrossRef] [PubMed]
- Sarrafzadeh, S.A.; Mahloojirad, M.; Casanova, J.-L.; Badalzadeh, M.; Bustamante, J.; Boisson-Dupuis, S.; Pourpak, Z.; Nourizadeh, M.; Moin, M. A New Patient with Inherited TYK2 Deficiency. J. Clin. Immunol. 2020, 40, 232–235. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, S.; Kaiser-Labusch, P.; Bank, J.; Ammann, S.; Kolb-Kokocinski, A.; Edelbusch, C.; Omran, H.; Ehl, S. Tyrosine kinase 2 is not limiting human antiviral type III interferon responses. Eur. J. Immunol. 2016, 46, 2639–2649. [Google Scholar] [CrossRef] [PubMed]
- Oyamada, A.; Ikebe, H.; Itsumi, M.; Saiwai, H.; Okada, S.; Shimoda, K.; Iwakura, Y.; Nakayama, K.I.; Iwamoto, Y.; Yoshikai, Y.; et al. Tyrosine Kinase 2 Plays Critical Roles in the Pathogenic CD4 T Cell Responses for the Development of Experimental Autoimmune Encephalomyelitis. J. Immunol. 2009, 183, 7539–7546. [Google Scholar] [CrossRef]
- Chapgier, A.; Boisson-Dupuis, S.; Jouanguy, E.; Vogt, G.; Feinberg, J.; Prochnicka-Chalufour, A.; Casrouge, A.; Yang, K.; Soudais, C.; Fieschi, C.; et al. Novel STAT1 Alleles in Otherwise Healthy Patients with Mycobacterial Disease. PLoS Genet. 2006, 2, e131. [Google Scholar] [CrossRef] [PubMed]
- Boisson-Dupuis, S.; Jouanguy, E.; Al-Hajjar, S.; Fieschi, C.; Al-Mohsen, I.Z.; Al-Jumaah, S.; Yang, K.; Chapgier, A.; Eidenschenk, C.; Eid, P.; et al. Impaired response to interferon-α/β and lethal viral disease in human STAT1 deficiency. Nat. Genet. 2003, 33, 388–391. [Google Scholar] [CrossRef] [PubMed]
- Boisson-Dupuis, S.; Dargemont, C.; Fieschi, C.; Thomassin, N.; Rosenzweig, S.; Harris, J.; Holland, S.M.; Schreiber, R.D.; Casanova, J.-L. Impairment of Mycobacterial But Not Viral Immunity by a Germline Human STAT1 Mutation. Science 2001, 293, 300–303. [Google Scholar] [CrossRef]
- Chapgier, A.; Kong, X.-F.; Boisson-Dupuis, S.; Jouanguy, E.; Averbuch, D.; Feinberg, J.; Zhang, S.-Y.; Bustamante, J.; Vogt, G.; Lejeune, J.; et al. A partial form of recessive STAT1 deficiency in humans. J. Clin. Investig. 2009, 119, 1502–1514. [Google Scholar] [CrossRef]
- Lee, C.-K.; Rao, D.T.; Gertner, R.; Gimeno, R.; Frey, A.B.; Levy, D.E. Distinct Requirements for IFNs and STAT1 in NK Cell Function. J. Immunol. 2000, 165, 3571–3577. [Google Scholar] [CrossRef] [PubMed]
- Putz, E.M.; Gotthardt, D.; Hoermann, G.; Csiszar, A.; Wirth, S.; Berger, A.; Straka, E.; Rigler, D.; Wallner, B.; Jamieson, A.; et al. CDK8-Mediated STAT1-S727 Phosphorylation Restrains NK Cell Cytotoxicity and Tumor Surveillance. Cell Rep. 2013, 4, 437–444. [Google Scholar] [CrossRef] [PubMed]
- Semper, C.; Leitner, N.R.; Lassnig, C.; Parrini, M.; Mahlakõiv, T.; Rammerstorfer, M.; Lorenz, K.; Rigler, D.; Müller, S.; Kolbe, T.; et al. STAT1 Is Not Dominant Negative and Is Capable of Contributing to Gamma Interferon-Dependent Innate Immunity. Mol. Cell. Biol. 2014, 34, 2235–2248. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, R.; Wu, W.; Wang, A.; Wan, Z.; Van De Veerdonk, F.L.; Li, R. New and recurrentSTAT1mutations in seven Chinese patients with chronic mucocutaneous candidiasis. Int. J. Dermatol. 2016, 56, e30–e33. [Google Scholar] [CrossRef]
- Toubiana, J.; Okada, S.; Hiller, J.; Oleastro, M.; Gomez, M.L.; Becerra, J.C.A.; Ouachã©E-Chardin, M.; Fouyssac, F.; Girisha, K.M.; Etzioni, A.; et al. Heterozygous STAT1 gain-of-function mutations underlie an unexpectedly broad clinical phenotype. Blood 2016, 127, 3154–3164. [Google Scholar] [CrossRef]
- Ifrim, D.C.; Quintin, J.; Meerstein-Kessel, L.; Plantinga, T.; Joosten, L.A.B.; Van Der Meer, J.W.M.; Van De Veerdonk, F.L.; Netea, M.G. Defective trained immunity in patients with STAT-1-dependent chronic mucocutaneaous candidiasis. Clin. Exp. Immunol. 2015, 181, 434–440. [Google Scholar] [CrossRef]
- Zheng, J.; Van De Veerdonk, F.L.; Crossland, K.L.; Smeekens, S.P.; Chan, C.M.; Al Shehri, T.; Abinun, M.; Gennery, A.R.; Mann, J.; Lendrem, D.W.; et al. Gain-of-function STAT1 mutations impair STAT3 activity in patients with chronic mucocutaneous candidiasis (CMC). Eur. J. Immunol. 2015, 45, 2834–2846. [Google Scholar] [CrossRef]
- Tamaura, M.; Satoh-Takayama, N.; Tsumura, M.; Sasaki, T.; Goda, S.; Kageyama, T.; Hayakawa, S.; Kimura, S.; Asano, T.; Nakayama, M.; et al. Human gain-of-function STAT1 mutation disturbs IL-17 immunity in mice. Int. Immunol. 2019, 32, 259–272. [Google Scholar] [CrossRef] [PubMed]
- Duncan, C.J.A.; Thompson, B.J.; Chen, R.; Rice, G.I.; Gothe, F.; Young, D.F.; Lovell, S.C.; Shuttleworth, V.G.; Brocklebank, V.; Corner, B.; et al. Severe type I interferonopathy and unrestrained interferon signaling due to a homozygous germline mutation inSTAT2. Sci. Immunol. 2019, 4, eaav7501. [Google Scholar] [CrossRef] [PubMed]
- Park, C.; Li, S.; Cha, E.; Schindler, C. Immune Response in Stat2 Knockout Mice. Immunity 2000, 13, 795–804. [Google Scholar] [CrossRef]
- Pelham, S.J.; Lenthall, H.C.; Deenick, E.K.; Tangye, S.G. Elucidating the effects of disease-causing mutations on STAT3 function in autosomal-dominant hyper-IgE syndrome. J. Allergy Clin. Immunol. 2016, 138, 1210–1213.e5. [Google Scholar] [CrossRef]
- De Beaucoudrey, L.; Puel, A.; Filipe-Santos, O.; Cobat, A.; Ghandil, P.; Chrabieh, M.; Feinberg, J.; Von Bernuth, H.; Samarina, A.; Jannière, L.; et al. Mutations in STAT3 and IL12RB1 impair the development of human IL-17–producing T cells. J. Exp. Med. 2008, 205, 1543–1550. [Google Scholar] [CrossRef]
- Takeda, K.; Noguchi, K.; Shi, W.; Tanaka, T.; Matsumoto, M.; Yoshida, N.; Kishimoto, T.; Akira, S. Targeted disruption of the mouse Stat3 gene leads to early embryonic lethality. Proc. Natl. Acad. Sci. 1997, 94, 3801–3804. [Google Scholar] [CrossRef]
- Flanagan, S.E.; Haapaniemi, E.; Russell, M.A.; Caswell, R.C.; Allen, H.L.; De Franco, E.; McDonald, T.J.; Rajala, H.L.M.; Ramelius, A.; Barton, J.; et al. Activating germline mutations in STAT3 cause early-onset multi-organ autoimmune disease. Nat. Genet. 2014, 46, 812–814. [Google Scholar] [CrossRef] [PubMed]
- Chitnis, T.; Najafian, N.; Benou, C.; Salama, A.D.; Grusby, M.J.; Sayegh, M.H.; Khoury, S.J. Effect of targeted disruption of STAT4 and STAT6 on the induction of experimental autoimmune encephalomyelitis. J. Clin. Investig. 2001, 108, 739–747. [Google Scholar] [CrossRef]
- Hwa, V.; Little, B.; Adiyaman, P.; Kofoed, E.M.; Pratt, K.L.; Ocal, G.; Berberoglu, M.; Rosenfeld, R.G. Severe Growth Hormone Insensitivity Resulting from Total Absence of Signal Transducer and Activator of Transcription 5b. J. Clin. Endocrinol. Metab. 2005, 90, 4260–4266. [Google Scholar] [CrossRef] [PubMed]
- Kofoed, E.M.; Hwa, V.; Little, B.; Woods, K.A.; Buckway, C.K.; Tsubaki, J.; Pratt, K.L.; Bezrodnik, L.; Jasper, H.; Tepper, A.; et al. Growth Hormone Insensitivity Associated with aSTAT5bMutation. New Engl. J. Med. 2003, 349, 1139–1147. [Google Scholar] [CrossRef] [PubMed]
- Bernasconi, A.; Marino, R.; Ribas, A.; Rossi, J.; Ciaccio, M.; Oleastro, M.; Ornani, A.; Paz, R.; Rivarola, M.A.; Zelazko, M.; et al. Characterization of Immunodeficiency in a Patient With Growth Hormone Insensitivity Secondary to a Novel STAT5b Gene Mutation. Pediatrics 2006, 118, e1584–e1592. [Google Scholar] [CrossRef] [PubMed]
- Vargas-Hernández, A.; Witalisz-Siepracka, A.; Prchal-Murphy, M.; Klein, K.; Mahapatra, S.; Al-Herz, W.; Mace, E.M.; Carisey, A.; Orange, J.S.; Sexl, V.; et al. Human signal transducer and activator of transcription 5b (STAT5b) mutation causes dysregulated human natural killer cell maturation and impaired lytic function. J. Allergy Clin. Immunol. 2020, 145, 345–357.e9. [Google Scholar] [CrossRef]
- Imada, K.; Bloom, E.T.; Nakajima, H.; Horvath-Arcidiacono, J.A.; Udy, G.B.; Davey, H.W.; Leonard, W.J. Stat5b Is Essential for Natural Killer Cell–mediated Proliferation and Cytolytic Activity. J. Exp. Med. 1998, 188, 2067–2074. [Google Scholar] [CrossRef] [PubMed]
- Villarino, A.V.; Sciumè, G.; Davis, F.P.; Iwata, S.; Zitti, B.; Robinson, G.W.; Hennighausen, L.; Kanno, Y.; O’Shea, J.J. Subset- and tissue-defined STAT5 thresholds control homeostasis and function of innate lymphoid cells. J. Exp. Med. 2017, 214, 2999–3014. [Google Scholar] [CrossRef] [PubMed]
- Villarino, A.; Laurence, A.; Robinson, G.W.; Bonelli, M.; Dema, B.; Afzali, B.; Shih, H.-Y.; Sun, H.-W.; Brooks, S.R.; Hennighausen, L.; et al. Signal transducer and activator of transcription 5 (STAT5) paralog dose governs T cell effector and regulatory functions. eLife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Quintás-Cardama, A.; Verstovsek, S. New JAK2 inhibitors for myeloproliferative neoplasms. Expert Opin. Investig. Drugs 2011, 20, 961–972. [Google Scholar] [CrossRef]
- Kleppe, M.; Koche, R.; Zou, L.; van Galen, P.; Hill, C.; Dong, L.; De Groote, S.; Papalexi, E.; Somasundara, A.V.H.; Cordner, K.; et al. Dual Targeting of Oncogenic Activation and Inflammatory Signaling Increases Therapeutic Efficacy in Myeloproliferative Neoplasms. Cancer Cell 2018, 33, 29–43.e7. [Google Scholar] [CrossRef] [PubMed]
- Mondet, J.; Hussein, K.; Mossuz, P. Circulating Cytokine Levels as Markers of Inflammation in Philadelphia Negative Myeloproliferative Neoplasms: Diagnostic and Prognostic Interest. Mediat. Inflamm. 2015, 2015, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Kleppe, M.; Kwak, M.; Koppikar, P.; Riester, M.; Keller, M.; Bastian, L.; Hricik, T.; Bhagwat, N.; McKenney, A.S.; Papalexi, E.; et al. JAK–STAT Pathway Activation in Malignant and Nonmalignant Cells Contributes to MPN Pathogenesis and Therapeutic Response. Cancer Discov. 2015, 5, 316–331. [Google Scholar] [CrossRef]
- Przepiorka, D.; Luo, L.; Subramaniam, S.; Qiu, J.; Gudi, R.; Cunningham, L.C.; Nie, L.; Leong, R.; Ma, L.; Sheth, C.; et al. FDA Approval Summary: Ruxolitinib for Treatment of Steroid-Refractory Acute Graft-Versus-Host Disease. Oncol. 2019, 25, e328–e334. [Google Scholar] [CrossRef]
- Harrison, C.N.; on Behalf of the COMFORT-II Investigators; Vannucchi, A.M.; Kiladjian, J.-J.; Al-Ali, H.K.; Gisslinger, H.; Knoops, L.; Cervantes, F.; Jones, M.M.; Sun, K.; et al. Long-term findings from COMFORT-II, a phase 3 study of ruxolitinib vs best available therapy for myelofibrosis. Leukemia 2016, 30, 1701–1707. [Google Scholar] [CrossRef]
- Manduzio, P. Ruxolitinib in myelofibrosis: To be or not to be an immune disruptor. Ther. Clin. Risk Manag. 2017, 13, 169–177. [Google Scholar] [CrossRef]
- Tremblay, D.; King, A.; Li, L.; Moshier, E.; Coltoff, A.; Koshy, A.; Kremyanskaya, M.; Hoffman, R.; Mauro, M.J.; Rampal, R.K.; et al. Risk factors for infections and secondary malignancies in patients with a myeloproliferative neoplasm treated with ruxolitinib: A dual-center, propensity score-matched analysis. Leuk. Lymphoma 2019, 61, 660–667. [Google Scholar] [CrossRef]
- Porpaczy, E.; Tripolt, S.; Hoelbl-Kovacic, A.; Gisslinger, B.; Bago-Horvath, Z.; Casanova-Hevia, E.; Clappier, E.; Decker, T.; Fajmann, S.; Fux, D.A.; et al. Aggressive B-cell lymphomas in patients with myelofibrosis receiving JAK1/2 inhibitor therapy. Blood 2018, 132, 694–706. [Google Scholar] [CrossRef]
- Rumi, E.; Zibellini, S.; Boveri, E.; Cavalloni, C.; Riboni, R.; Casetti, I.C.; Ciboddo, M.; Trotti, C.; Favaron, C.; Pietra, D.; et al. Ruxolitinib treatment and risk of B-cell lymphomas in myeloproliferative neoplasms. Am. J. Hematol. 2019, 94, E185–E188. [Google Scholar] [CrossRef] [PubMed]
- Mora, B.; Rumi, E.; Guglielmelli, P.; Barraco, D.; Maffioli, M.; Rambaldi, A.; Caramella, M.; Komrokji, R.; Gotlib, J.; Kiladjian, J.J.; et al. Second primary malignancies in postpolycythemia vera and postessential thrombocythemia myelofibrosis: A study on 2233 patients. Cancer Med. 2019, 8, 4089–4092. [Google Scholar] [CrossRef] [PubMed]
- Maffioli, M.; Giorgino, T.; Mora, B.; Iurlo, A.; Elli, E.; Finazzi, M.C.; Caramella, M.; Rumi, E.; Carraro, M.C.; Polverelli, N.; et al. Second primary malignancies in ruxolitinib-treated myelofibrosis: Real-world evidence from 219 consecutive patients. Blood Adv. 2019, 3, 3196–3200. [Google Scholar] [CrossRef]
- Smolen, J.S.; Genovese, M.C.; Takeuchi, T.; Hyslop, D.L.; Macias, W.L.; Rooney, T.; Chen, L.; Dickson, C.L.; Camp, J.R.; Cardillo, T.E.; et al. Safety Profile of Baricitinib in Patients with Active Rheumatoid Arthritis with over 2 Years Median Time in Treatment. J. Rheumatol. 2018, 46, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Schönberg, K.; Rudolph, J.; Vonnahme, M.; Yajnanarayana, S.P.; Cornez, I.; Hejazi, M.; Manser, A.R.; Uhrberg, M.; Verbeek, W.; Koschmieder, S.; et al. JAK Inhibition Impairs NK Cell Function in Myeloproliferative Neoplasms. Cancer Res. 2015, 75, 2187–2199. [Google Scholar] [CrossRef]
- Curran, S.A.; Shyer, J.A.; Angelo, E.T.S.; Talbot, L.R.; Sharma, S.; Chung, D.J.; Heller, G.; Hsu, K.C.; Betts, B.C.; Young, J.W. Human Dendritic Cells Mitigate NK-Cell Dysfunction Mediated by Nonselective JAK1/2 Blockade. Cancer Immunol. Res. 2017, 5, 52–60. [Google Scholar] [CrossRef]
- Lucas, M.; Schachterle, W.; Oberle, K.; Aichele, P.; Diefenbach, A. Dendritic Cells Prime Natural Killer Cells by trans-Presenting Interleukin 15. Immunity 2007, 26, 503–517. [Google Scholar] [CrossRef]
- Heine, A.; Held, S.A.E.; Daecke, S.N.; Wallner, S.; Yajnanarayana, S.P.; Kurts, C.; Wolf, D.; Brossart, P. The JAK-inhibitor ruxolitinib impairs dendritic cell function in vitro and in vivo. Blood 2013, 122, 1192–1202. [Google Scholar] [CrossRef]
- Rudolph, J.; Heine, A.; Quast, T.; Kolanus, W.; Trebicka, J.; Brossart, P.; Wolf, D. The JAK inhibitor ruxolitinib impairs dendritic cell migration via off-target inhibition of ROCK. Leukemia 2016, 30, 2119–2123. [Google Scholar] [CrossRef]
- Yajnanarayana, S.P.; Stübig, T.; Cornez, I.; Alchalby, H.; Schönberg, K.; Rudolph, J.; Triviai, I.; Wolschke, C.; Heine, A.; Brossart, P.; et al. JAK1/2 inhibition impairs T cell functionin vitroand in patients with myeloproliferative neoplasms. Br. J. Haematol. 2015, 169, 824–833. [Google Scholar] [CrossRef] [PubMed]
- Keohane, C.; Kordasti, S.Y.; Seidl, T.; Abellan, P.P.; Thomas, N.S.B.; Harrison, C.N.; McLornan, D.P.; Mufti, G.J. JAK inhibition induces silencing of T Helper cytokine secretion and a profound reduction in T regulatory cells. Br. J. Haematol. 2015, 171, 60–73. [Google Scholar] [CrossRef]
- Massa, M.L.; Rosti, V.; Campanelli, R.; Fois, G.; Barosi, G. Rapid and long-lasting decrease of T-regulatory cells in patients with myelofibrosis treated with ruxolitinib. Leukemia 2014, 28, 449–451. [Google Scholar] [CrossRef] [PubMed]
- Spoerl, S.; Mathew, N.R.; Bscheider, M.; Schmitt-Graeff, A.; Chen, S.; Mueller, T.; Verbeek, M.; Fischer, J.; Otten, V.; Schmickl, M.; et al. Activity of therapeutic JAK 1/2 blockade in graft-versus-host disease. Blood 2014, 123, 3832–3842. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, A.L.; Bjørn, M.E.; Riley, C.H.; Holmstrøm, M.; Andersen, M.H.; Svane, I.M.; Mikkelsen, S.U.; Skov, V.; Kjaer, L.; Hasselbalch, H.C.; et al. B-cell frequencies and immunoregulatory phenotypes in myeloproliferative neoplasms: Influence of ruxolitinib, interferon-α2, or combination treatment. Eur. J. Haematol. 2019, 103, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Fleischmann, R.; Kremer, J.; Cush, J.; Schulze-Koops, H.; Connell, C.A.; Bradley, J.D.; Gruben, D.; Wallenstein, G.V.; Zwillich, S.H.; Kanik, K.S. Placebo-Controlled Trial of Tofacitinib Monotherapy in Rheumatoid Arthritis. New Engl. J. Med. 2012, 367, 495–507. [Google Scholar] [CrossRef] [PubMed]
- Kontzias, A.; Kotlyar, A.; Laurence, A.; Changelian, P.; O’Shea, J.J. Jakinibs: A new class of kinase inhibitors in cancer and autoimmune disease. Curr. Opin. Pharmacol. 2012, 12, 464–470. [Google Scholar] [CrossRef]
- Degryse, S.; Cools, J. JAK kinase inhibitors for the treatment of acute lymphoblastic leukemia. J. Hematol. Oncol. 2015, 8, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Ghoreschi, K.; Jesson, M.I.; Li, X.; Lee, J.L.; Ghosh, S.; Alsup, J.W.; Warner, J.D.; Tanaka, M.; Steward-Tharp, S.M.; Gadina, M.; et al. Modulation of Innate and Adaptive Immune Responses by Tofacitinib (CP-690,550). J. Immunol. 2011, 186, 4234–4243. [Google Scholar] [CrossRef] [PubMed]
- Migita, K.; Miyashita, T.; Izumi, Y.; Koga, T.; Komori, A.; Maeda, Y.; Jiuchi, Y.; Aiba, Y.; Yamasaki, S.; Kawakami, A.; et al. Inhibitory effects of the JAK inhibitor CP690,550 on human CD4+ T lymphocyte cytokine production. BMC Immunol. 2011, 12, 51. [Google Scholar] [CrossRef]
- Sonomoto, K.; Yamaoka, K.; Kubo, S.; Hirata, S.; Fukuyo, S.; Maeshima, K.; Suzuki, K.; Saito, K.; Tanaka, Y. Effects of tofacitinib on lymphocytes in rheumatoid arthritis: Relation to efficacy and infectious adverse events. Rheumatology 2014, 53, 914–918. [Google Scholar] [CrossRef]
- A Hodge, J.; Kawabata, T.T.; Krishnaswami, S.; Clark, J.D.; Telliez, J.-B.; E Dowty, M.; Menon, S.; Lamba, M.; Zwillich, S. The mechanism of action of tofacitinib - an oral Janus kinase inhibitor for the treatment of rheumatoid arthritis. Clin. Exp. Rheumatol. 2016, 34, 318–328. [Google Scholar] [PubMed]
- Conklyn, M.; Andresen, C.; Changelian, P.; Kudlacz, E. The JAK3 inhibitor CP-690550 selectively reduces NK and CD8+ cell numbers in cynomolgus monkey blood following chronic oral dosing. J. Leukoc. Biol. 2004, 76, 1248–1255. [Google Scholar] [CrossRef] [PubMed]
- Kudlacz, E.; Perry, B.; Sawyer, P.; Conklyn, M.; McCurdy, S.; Brissette, W.; And, M.F.; Changelian, P. The Novel JAK-3 Inhibitor CP-690550 Is a Potent Immunosuppressive Agent in Various Murine Models. Arab. Archaeol. Epigr. 2004, 4, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Shimaoka, H.; Takeno, S.; Maki, K.; Sasaki, T.; Hasegawa, S.; Yamashita, Y. A cytokine signal inhibitor for rheumatoid arthritis enhances cancer metastasis via depletion of NK cells in an experimental lung metastasis mouse model of colon cancer. Oncol. Lett. 2017, 14, 3019–3027. [Google Scholar] [CrossRef]
- Vian, L.; Le, M.T.; Gazaniga, N.; Kieltyka, J.; Liu, C.; Pietropaolo, G.; Dell’Orso, S.; Brooks, S.R.; Furumoto, Y.; Thomas, C.J.; et al. JAK Inhibition Differentially Affects NK Cell and ILC1 Homeostasis. Front. Immunol. 2019, 10, 2972. [Google Scholar] [CrossRef]
- Rochman, Y.; Spolski, R.; Leonard, W.J. New insights into the regulation of T cells by γc family cytokines. Nat. Rev. Immunol. 2009, 9, 480–490. [Google Scholar] [CrossRef]
- Meazza, R.; Azzarone, B.; Orengo, A.M.; Ferrini, S. Role of Common-Gamma Chain Cytokines in NK Cell Development and Function: Perspectives for Immunotherapy. J. Biomed. Biotechnol. 2011, 2011, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Marçais, A.; Viel, S.; Grau, M.; Henry, T.; Marvel, J.; Walzer, T. Regulation of Mouse NK Cell Development and Function by Cytokines. Front. Immunol. 2013, 4, 450. [Google Scholar] [CrossRef] [PubMed]
- Changelian, P.S.; Flanagan, M.E.; Ball, D.J.; Kent, C.R.; Magnuson, K.S.; Martin, W.H.; Rizzuti, B.J.; Sawyer, P.S.; Perry, B.D.; Brissette, W.H.; et al. Prevention of Organ Allograft Rejection by a Specific Janus Kinase 3 Inhibitor. Science 2003, 302, 875–878. [Google Scholar] [CrossRef]
- Van Vollenhoven, R.; Tanaka, Y.; Lamba, M.; Collinge, M.; Hendrikx, T.; Hirose, T.; Toyoizumi, S.; Hazra, A.; Krishnaswami, S. THU0178 Relationship Between NK Cell Count and Important Safety Events in Rheumatoid Arthritis Patients Treated with Tofacitinib. Ann. Rheum. Dis. 2015, 74, 258.3–259. [Google Scholar] [CrossRef]
- Van Vollenhoven, R.; Choy, E.; Lee, E.; Hazra, A.; Anisfeld, A.; Lazariciu, I.; Biswas, P.; Lamba, M.; Menon, S.; Hodge, J.; et al. THU0199 Tofacitinib, An Oral Janus Kinase Inhibitor, in The Treatment of Rheumatoid Arthritis: Changes in Lymphocytes and Lymphocyte Subset Counts and Reversibility after Up To 8 Years of Tofacitinib Treatment. Ann. Rheum. Dis. 2016, 75, 258. [Google Scholar] [CrossRef]
- Van Vollenhoven, R.; Lee, E.B.; Strengholt, S.; Mojcik, C.; Valdez, H.; Krishnaswami, S.; Biswas, P.; Lazariciu, I.; Hazra, A.; Clark, J.D.; et al. Evaluation of the Short-, Mid-, and Long-Term Effects of Tofacitinib on Lymphocytes in Patients With Rheumatoid Arthritis. Arthritis Rheumatol. 2019, 71, 685–695. [Google Scholar] [CrossRef]
- Weinhold, K.J.; Bukowski, J.F.; Brennan, T.V.; Noveck, R.J.; Staats, J.S.; Lin, L.; Stempora, L.; Hammond, C.; Wouters, A.; Mojcik, C.F.; et al. Reversibility of peripheral blood leukocyte phenotypic and functional changes after exposure to and withdrawal from tofacitinib, a Janus kinase inhibitor, in healthy volunteers. Clin. Immunol. 2018, 191, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Angelini, J.; Talotta, R.; Roncato, R.; Fornasier, G.; Barbiero, G.; Cin, L.D.; Brancati, S.; Scaglione, F. JAK-Inhibitors for the Treatment of Rheumatoid Arthritis: A Focus on the Present and an Outlook on the Future. Biomolecules 2020, 10, 1002. [Google Scholar] [CrossRef]
- Nocturne, G.; Pascaud, J.; Ly, B.; Tahmasebi, F.; Mariette, X. JAK inhibitors alter NK cell functions and may impair immunosurveillance against lymphomagenesis. Cell. Mol. Immunol. 2020, 17, 552–553. [Google Scholar] [CrossRef]
- Kubo, S.; Yamaoka, K.; Kondo, M.; Yamagata, K.; Zhao, J.; Iwata, S.; Tanaka, Y. The JAK inhibitor, tofacitinib, reduces the T cell stimulatory capacity of human monocyte-derived dendritic cells. Ann. Rheum. Dis. 2013, 73, 2192–2198. [Google Scholar] [CrossRef] [PubMed]
- Sewgobind, V.D.K.D.; Quaedackers, M.E.; Van Der Laan, L.J.W.; Kraaijeveld, R.; Korevaar, S.S.; Chan, G.; Weimar, W.; Baan, C.C. The Jak Inhibitor CP-690,550 Preserves the Function of CD4+CD25brightFoxP3+ Regulatory T Cells and Inhibits Effector T Cells. Arab. Archaeol. Epigr. 2010, 10, 1785–1795. [Google Scholar] [CrossRef]
- Meyer, A.; Wittekind, P.S.; Kotschenreuther, K.; Schiller, J.; Von Tresckow, J.; Haak, T.H.; Kofler, D.M. Regulatory T cell frequencies in patients with rheumatoid arthritis are increased by conventional and biological DMARDs but not by JAK inhibitors. Ann. Rheum. Dis. 2019. [Google Scholar] [CrossRef]
- Rizzi, M.; Lorenzetti, R.; Fischer, K.; Staniek, J.; Janowska, I.; Troilo, A.; Strohmeier, V.; Erlacher, M.; Kunze, M.; Bannert, B.; et al. Impact of tofacitinib treatment on human B-cells in vitro and in vivo. J. Autoimmun. 2017, 77, 55–66. [Google Scholar] [CrossRef]
- Mariette, X.; Chen, C.; Biswas, P.; Kwok, K.; Boy, M.G. Lymphoma in the Tofacitinib Rheumatoid Arthritis Clinical Development Program. Arthritis Rheum. 2018, 70, 685–694. [Google Scholar] [CrossRef] [PubMed]
- PFIZER press release. Available online: https://www.pfizer.com/news/press-release/press-release-detail/pfizer-shares-co-primary-endpoint-results-post-marketing (accessed on 1 April 2021).
- Talpaz, M.; Kiladjian, J.-J. Fedratinib, a newly approved treatment for patients with myeloproliferative neoplasm-associated myelofibrosis. Leukemia 2021, 35, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.S.; Kim, M.J.; Kim, D.O.; Byun, J.-E.; Huy, H.; Song, H.Y.; Park, Y.-J.; Kim, T.-D.; Yoon, S.R.; Choi, E.-J.; et al. Suppressor of Cytokine Signaling 2 Negatively Regulates NK Cell Differentiation by Inhibiting JAK2 Activity. Sci. Rep. 2017, 7, 46153. [Google Scholar] [CrossRef] [PubMed]
- Betts, B.C.; Abdel-Wahab, O.; Curran, S.A.; Angelo, E.T.S.; Koppikar, P.; Heller, G.; Levine, R.L.; Young, J.W. Janus kinase-2 inhibition induces durable tolerance to alloantigen by human dendritic cell–stimulated T cells yet preserves immunity to recall antigen. Blood 2011, 118, 5330–5339. [Google Scholar] [CrossRef] [PubMed]
- Mesa, R.A.; Kiladjian, J.-J.; Catalano, J.V.; Devos, T.; Egyed, M.; Hellmann, A.; McLornan, D.; Shimoda, K.; Winton, E.F.; Deng, W.; et al. SIMPLIFY-1: A Phase III Randomized Trial of Momelotinib Versus Ruxolitinib in Janus Kinase Inhibitor–Naïve Patients With Myelofibrosis. J. Clin. Oncol. 2017, 35, 3844–3850. [Google Scholar] [CrossRef]
- Patel, A.A.; Odenike, O. The Next Generation of JAK Inhibitors: An Update on Fedratinib, Momelotonib, and Pacritinib. Curr. Hematol. Malign- Rep. 2020, 15, 409–418. [Google Scholar] [CrossRef]
- Singer, J.W.; Al-Fayoumi, S.; Taylor, J.; Velichko, S.; O’Mahony, A. Comparative phenotypic profiling of the JAK2 inhibitors ruxolitinib, fedratinib, momelotinib, and pacritinib reveals distinct mechanistic signatures. PLoS ONE 2019, 14, e0222944. [Google Scholar] [CrossRef]
- Singer, J.W.; Al-Fayoumi, S.; Ma, H.; Komrokji, R.S.; Mesa, R.; Verstovsek, S. Comprehensive kinase profile of pacritinib, a nonmyelosuppressive Janus kinase 2 inhibitor. J. Exp. Pharmacol. 2016, ume 8, 11–19. [Google Scholar] [CrossRef]
- Hosseini, M.M.; Kurtz, S.E.; Abdelhamed, S.; Mahmood, S.; Davare, M.A.; Kaempf, A.; Elferich, J.; McDermott, J.E.; Liu, T.; Payne, S.H.; et al. Inhibition of interleukin-1 receptor-associated kinase-1 is a therapeutic strategy for acute myeloid leukemia subtypes. Leukemia 2018, 32, 2374–2387. [Google Scholar] [CrossRef]
- Pidala, J.; Walton, K.; Elmariah, H.; Kim, J.; Mishra, A.; Bejanyan, N.; Nishihori, T.; Khimani, F.; Perez, L.; Faramand, R.G.; et al. Pacritinib Combined with Sirolimus and Low-Dose Tacrolimus for GVHD Prevention after Allogeneic Hematopoietic Cell Transplantation: Preclinical and Phase I Trial Results. Clin. Cancer Res. 2021, 27, 2712–2722. [Google Scholar] [CrossRef] [PubMed]
- Covington, M.; He, X.; Scuron, M.; Li, J.; Collins, R.; Juvekar, A.; Shin, N.; Favata, M.; Gallagher, K.; Sarah, S.; et al. Preclinical characterization of itacitinib (INCB039110), a novel selective inhibitor of JAK1, for the treatment of inflammatory diseases. Eur. J. Pharmacol. 2020, 885, 173505. [Google Scholar] [CrossRef]
- Murray, P.J. The JAK-STAT Signaling Pathway: Input and Output Integration. J. Immunol. 2007, 178, 2623–2629. [Google Scholar] [CrossRef] [PubMed]
- Huarte, E.; O’Connor, R.S.; Peel, M.T.; Nunez-Cruz, S.; Leferovich, J.; Juvekar, A.; Yang, Y.-O.; Truong, L.; Huang, T.; Naim, A.; et al. Itacitinib (INCB039110), a JAK1 Inhibitor, Reduces Cytokines Associated with Cytokine Release Syndrome Induced by CAR T-cell Therapy. Clin. Cancer Res. 2020, 26, 6299–6309. [Google Scholar] [CrossRef]
- Juvekar, A.; Ruggeri, B.; Condon, S.; Borkowski, A.; Huber, R.; Smith, P. Itacitinib, a JAK1 Selective Inhibitor Preserves Graft-Versus-Leukemia (GVL), Enhances Survival and Is Highly Efficacious in a MHC-Mismatched Mouse Model of Acute GvHD. Blood 2018, 132, 4522. [Google Scholar] [CrossRef]
- Schroeder, M.A.; Khoury, H.J.; Jagasia, M.; Ali, H.; Schiller, G.J.; Staser, K.; Choi, J.; Gehrs, L.; Arbushites, M.C.; Yan, Y.; et al. A phase 1 trial of itacitinib, a selective JAK1 inhibitor, in patients with acute graft-versus-host disease. Blood Adv. 2020, 4, 1656–1669. [Google Scholar] [CrossRef] [PubMed]
- Luchi, M.; Fidelus-Gort, R.; Douglas, D.; Zhang, H.; Flores, R.; Newton, R.; Scherle, P.; Yeleswaram, S.; Chen, X.; Sandor, V.A. Randomized, Dose-Ranging, Placebo-Controlled, 84-Day Study Of INCB039110, a Selective Janus Kinase-1 Inhibitor, In Pa-tients With Active Rheumatoid Arthritis - ACR Meeting Abstracts. Arthritis Rheum 2013, 65 (Suppl. 10), 1797. [Google Scholar]
- Norman, P. Selective JAK inhibitors in development for rheumatoid arthritis. Expert Opin. Investig. Drugs 2014, 23, 1067–1077. [Google Scholar] [CrossRef] [PubMed]
- Bissonnette, R.; Luchi, M.; Fidelus-Gort, R.; Jackson, S.; Zhang, H.; Flores, R.; Newton, R.; Scherle, P.; Yeleswaram, S.; Chen, X.; et al. A randomized, double-blind, placebo-controlled, dose-escalation study of the safety and efficacy of INCB039110, an oral janus kinase 1 inhibitor, in patients with stable, chronic plaque psoriasis. J. Dermatol. Treat. 2015, 27, 332–338. [Google Scholar] [CrossRef]
- Banerjee, S.; Biehl, A.; Gadina, M.; Hasni, S.; Schwartz, D.M. JAK–STAT Signaling as a Target for Inflammatory and Autoimmune Diseases: Current and Future Prospects. Drugs 2017, 77, 521–546. [Google Scholar] [CrossRef]
- Zhang, M.; Griner, L.A.M.; Ju, W.; Duveau, D.Y.; Guha, R.; Petrus, M.N.; Wen, B.; Maeda, M.; Shinn, P.; Ferrer, M.; et al. Selective targeting of JAK/STAT signaling is potentiated by Bcl-xL blockade in IL-2–dependent adult T-cell leukemia. Proc. Natl. Acad. Sci. 2015, 112, 12480–12485. [Google Scholar] [CrossRef] [PubMed]
- Waldmann, T.A. JAK/STAT pathway directed therapy of T-cell leukemia/lymphoma: Inspired by functional and structural genomics. Mol. Cell. Endocrinol. 2017, 451, 66–70. [Google Scholar] [CrossRef] [PubMed]
- Senkevitch, E.; Li, W.; Hixon, J.A.; Andrews, C.; Cramer, S.D.; Pauly, G.; Back, T.; Czarra, K.; Durum, S.K. Inhibiting Janus Kinase 1 and BCL-2 to treat T cell acute lymphoblastic leukemia with IL7-Rα mutations. Oncotarget 2018, 9, 22605–22617. [Google Scholar] [CrossRef]
- Shouse, G.; Nikolaenko, L. Targeting the JAK/STAT Pathway in T Cell Lymphoproliferative Disorders. Curr. Hematol. Malign- Rep. 2019, 14, 570–576. [Google Scholar] [CrossRef]
- Hee, Y.T.; Yan, J.; Nizetic, D.; Chng, W.-J. LEE011 and ruxolitinib: A synergistic drug combination for natural killer/T-cell lymphoma (NKTCL). Oncotarget 2018, 9, 31832–31841. [Google Scholar] [CrossRef][Green Version]
- Mondéjar, R.; Pérez, C.; Onaindía, A.; Martínez, N.; González-Rincón, J.; Pisonero, H.; Vaque, J.P.; Cereceda, L.; Santibañez, M.; Sánchez-Beato, M.; et al. Molecular basis of targeted therapy in T/NK-cell lymphoma/leukemia: A comprehensive genomic and immunohistochemical analysis of a panel of 33 cell lines. PLoS ONE 2017, 12, e0177524. [Google Scholar] [CrossRef] [PubMed]
- Neste, E.V.D.; André, M.; Gastinne, T.; Stamatoullas, A.; Haioun, C.; Belhabri, A.; Reman, O.; Casasnovas, O.; Ghesquieres, H.; Verhoef, G.; et al. A phase II study of the oral JAK1/JAK2 inhibitor ruxolitinib in advanced relapsed/refractory Hodgkin lymphoma. Haematologica 2018, 103, 840–848. [Google Scholar] [CrossRef] [PubMed]
- Moskowitz, A.J.; Ghione, P.; Jacobsen, E.D.; Ruan, J.; Schatz, J.H.; Noor, S.; Myskowski, P.; Hancock, A.H.; Davey, M.T.; Obadi, O.; et al. Final Results of a Phase II Biomarker-Driven Study of Ruxolitinib in Relapsed and Refractory T-Cell Lymphoma. Blood 2019, 134, 4019. [Google Scholar] [CrossRef]
- Mulvey, E.; Ruan, J. Biomarker-driven management strategies for peripheral T cell lymphoma. J. Hematol. Oncol. 2020, 13, 1–20. [Google Scholar] [CrossRef]
- Karagianni, F.; Piperi, C.; Mpakou, V.; Spathis, A.; Foukas, P.G.; Dalamaga, M.; Pappa, V.; Papadavid, E. Ruxolitinib with resminostat exert synergistic antitumor effects in Cutaneous T-cell Lymphoma. PLoS ONE 2021, 16, e0248298. [Google Scholar] [CrossRef]
- Braun, T.; Von Jan, J.; Wahnschaffe, L.; Herling, M. Advances and Perspectives in the Treatment of T-PLL. Curr. Hematol. Malign- Rep. 2020, 15, 113–124. [Google Scholar] [CrossRef] [PubMed]
- Ando, S.; Kawada, J.-I.; Watanabe, T.; Suzuki, M.; Sato, Y.; Torii, Y.; Asai, M.; Goshima, F.; Murata, T.; Shimizu, N.; et al. Tofacitinib induces G1 cell-cycle arrest and inhibits tumor growth in Epstein-Barr virus-associated T and natural killer cell lymphoma cells. Oncotarget 2016, 7, 76793–76805. [Google Scholar] [CrossRef]
- Wei, B.M.; Koshy, N.; Van Besien, K.; Inghirami, G.; Horwitz, S.M. Refractory T-Cell Prolymphocytic Leukemia with JAK3 Mutation: In Vitro and Clinical Synergy of Tofacitinib and Ruxolitinib. Blood 2015, 126, 5486. [Google Scholar] [CrossRef]
- Gomez-Arteaga, A.; Margolskee, E.; Wei, M.T.; Van Besien, K.; Inghirami, G.; Horwitz, S. Combined use of tofacitinib (pan-JAK inhibitor) and ruxolitinib (a JAK1/2 inhibitor) for refractory T-cell prolymphocytic leukemia (T-PLL) with a JAK3 mutation. Leuk. Lymphoma 2019, 60, 1626–1631. [Google Scholar] [CrossRef]
- Lindahl, L.M.; Fredholm, S.; Joseph, C.; Nielsen, B.S.; Jønson, L.; Willerslev-Olsen, A.; Gluud, M.; Blümel, E.; Petersen, D.L.; Sibbesen, N.; et al. STAT5 induces miR-21 expression in cutaneous T cell lymphoma. Oncotarget 2016, 7, 45730–45744. [Google Scholar] [CrossRef]
- Cabannes, A.; Schmidt, A.; Brissot, E.; Balsat, M.; Maury, S.; Isnard, F.; Chevallier, P.; Cacheux, V.; Cluzeau, T.; Graux, C.; et al. The Combination of Venetoclax and Tofacitinib Induced Hematological Responses in Patients with Relapse/ Refractory T-ALL with BCL2 Expression and Surface IL7R Expression or IL7R-Pathway Mutations (On behalf of the GRAALL). Blood 2019, 134, 1339. [Google Scholar] [CrossRef]
- Wong, J.; Wall, M.; Corboy, G.P.; Taubenheim, N.; Gregory, G.P.; Opat, S.; Shortt, J. Failure of tofacitinib to achieve an objective response in a DDX3X-MLLT10 T-lymphoblastic leukemia with activating JAK3 mutations. Mol. Case Stud. 2020, 6, a004994. [Google Scholar] [CrossRef]
- Zhang, R.; Shah, M.V.; Loughran, T.P. The root of many evils: Indolent large granular lymphocyte leukaemia and associated disorders. Hematol. Oncol. 2009, 28, 105–117. [Google Scholar] [CrossRef]
- Lamy, T.; Loughran, J.T.P. How I treat LGL leukemia. Blood 2011, 117, 2764–2774. [Google Scholar] [CrossRef] [PubMed]
- Shah, M.V.; Hook, C.C.; Call, T.G.; Go, R.S. A population-based study of large granular lymphocyte leukemia. Blood Cancer J. 2016, 6, e455. [Google Scholar] [CrossRef] [PubMed]
- Lamy, T.; Moignet, A.; Loughran, T.P. LGL leukemia: From pathogenesis to treatment. Blood 2017, 129, 1082–1094. [Google Scholar] [CrossRef] [PubMed]
- Firestein, G.S. Evolving concepts of rheumatoid arthritis. Nat. Cell Biol. 2003, 423, 356–361. [Google Scholar] [CrossRef]
- Liu, X.; Loughran, T.P. The spectrum of large granular lymphocyte leukemia and Feltyʼs syndrome. Curr. Opin. Hematol. 2011, 18, 254–259. [Google Scholar] [CrossRef]
- Bockorny, B.; Dasanu, C.A. Autoimmune Manifestations in Large Granular Lymphocyte Leukemia. Clin. Lymphoma Myeloma Leuk. 2012, 12, 400–405. [Google Scholar] [CrossRef]
- Poullot, E.; Zambello, R.; Leblanc, F.; Bareau, B.; De March, E.; Roussel, M.; Boulland, M.L.; Houot, R.; Renault, A.; Fest, T.; et al. Chronic natural killer lymphoproliferative disorders: Characteristics of an international cohort of 70 patients. Ann. Oncol. 2014, 25, 2030–2035. [Google Scholar] [CrossRef]
- Moignet, A.; Lamy, T. Latest Advances in the Diagnosis and Treatment of Large Granular Lymphocytic Leukemia. Am. Soc. Clin. Oncol. Educ. Book 2018, 38, 616–625. [Google Scholar] [CrossRef]
- Hu, M.; Xu, C.; Yang, C.; Zuo, H.; Chen, C.; Zhang, D.; Shi, G.; Wang, W.; Shi, J.; Zhang, T. Discovery and evaluation of ZT55, a novel highly-selective tyrosine kinase inhibitor of JAK2V617F against myeloproliferative neoplasms. J. Exp. Clin. Cancer Res. 2019, 38, 1–12. [Google Scholar] [CrossRef]
- Betts, B.C.; Young, J.W. Less Can Be More When Targeting Interleukin-6-Mediated Cytokine Release Syndrome in Coronavirus Disease 2019. Crit. Care Explor. 2020, 2, e0138. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhu, S.; Liu, W.; Ming, J.; Wang, X.; Hu, X. Ruxolitinib-based combinations in the treatment of myelofibrosis: Worth looking forward to. Ann. Hematol. 2020, 99, 1161–1176. [Google Scholar] [CrossRef] [PubMed]
 ), the same signaling pathway drives MPNs (myeloproliferative neoplasms; left side of the receptor). Right side: In a physiological situation, common gamma chain-dependent cytokines (e.g., IL-2/7/15) signal via JAK1/JAK3 pairs to induce lymphopoiesis and regulate T/B/NK-cell function (right side of the receptor). In a cancerous situation (indicated by
), the same signaling pathway drives MPNs (myeloproliferative neoplasms; left side of the receptor). Right side: In a physiological situation, common gamma chain-dependent cytokines (e.g., IL-2/7/15) signal via JAK1/JAK3 pairs to induce lymphopoiesis and regulate T/B/NK-cell function (right side of the receptor). In a cancerous situation (indicated by  ), the same signaling pathway drives leukemias and lymphomas originating from T/B/NK cells (left side of the receptor). The consequences of JAKinib treatment on physiological and pathological effects of the pathways are indicated by colors (see legend).
), the same signaling pathway drives leukemias and lymphomas originating from T/B/NK cells (left side of the receptor). The consequences of JAKinib treatment on physiological and pathological effects of the pathways are indicated by colors (see legend).
 ), the same signaling pathway drives MPNs (myeloproliferative neoplasms; left side of the receptor). Right side: In a physiological situation, common gamma chain-dependent cytokines (e.g., IL-2/7/15) signal via JAK1/JAK3 pairs to induce lymphopoiesis and regulate T/B/NK-cell function (right side of the receptor). In a cancerous situation (indicated by
), the same signaling pathway drives MPNs (myeloproliferative neoplasms; left side of the receptor). Right side: In a physiological situation, common gamma chain-dependent cytokines (e.g., IL-2/7/15) signal via JAK1/JAK3 pairs to induce lymphopoiesis and regulate T/B/NK-cell function (right side of the receptor). In a cancerous situation (indicated by  ), the same signaling pathway drives leukemias and lymphomas originating from T/B/NK cells (left side of the receptor). The consequences of JAKinib treatment on physiological and pathological effects of the pathways are indicated by colors (see legend).
), the same signaling pathway drives leukemias and lymphomas originating from T/B/NK cells (left side of the receptor). The consequences of JAKinib treatment on physiological and pathological effects of the pathways are indicated by colors (see legend).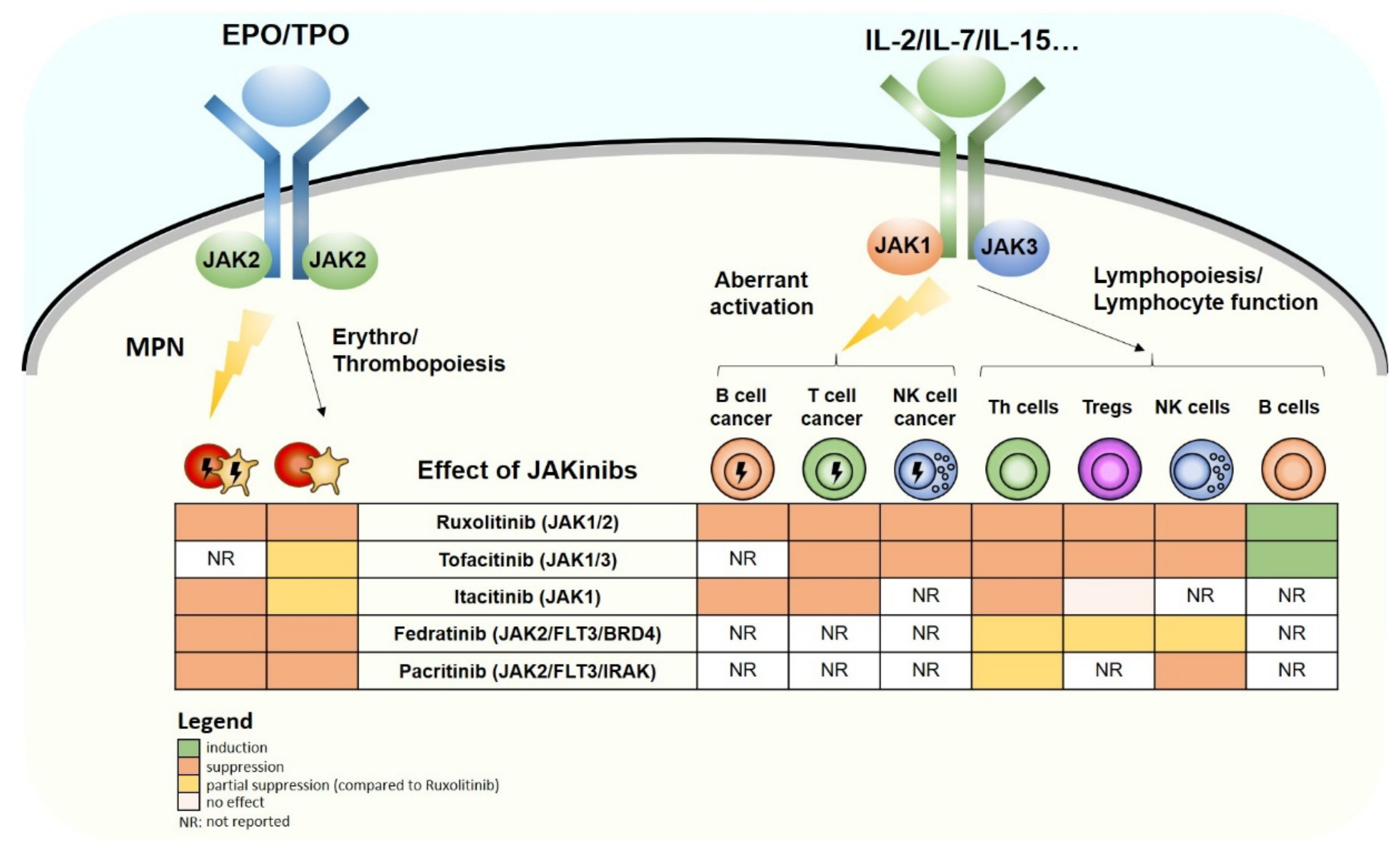
| JAK/ STAT | Type of mutations | Hematological malignancies 1 | Examples for JAKinib sensitivity | References |
|---|---|---|---|---|
| JAK1 | somatic GOF(e.g., JAK1S646F, JAK1S646P, JAK1V658I) | ALL (B-ALL, ETP-ALL, adult T-ALL), T-PLL, BIA-ALCL, ALK- ALCL, AML, MPN-unclassifiable, CMML | Ba/F3 cells expressing JAK1S646F, JAK1S646P, or JAK1V658I are sensitive to JAKinibs, including Ruxolitinib. | [4,15,39,40,41,42,51,74,75,76,77,78] |
| JAK2 | germline GOF(e.g., JAK2R564Q, JAK2V617I) | hereditary essential thrombocytosis | Ba/F3-MPL cells expressing JAK2R564Q are more sensitive to Ruxolitinib than JAK2V617F-expressers. | [36,37,38] |
| somatic GOF(e.g., JAK2V617F) | MPN (PV > ET, PMF), AML, pediatric, and DS-ALL | Ruxolitinib is approved for treatment of MPN. | [26,27,28,29,30,31,32,33,34,35] | |
| JAK3 | germline GOF(JAK3Q507P) | familial CLPD-NK | [79] | |
| somatic GOF(e.g., JAK3M511I, JAK3A572V, JAK3A573V) | ALL (ETP-ALL, T-ALL), ATLL, T-PLL, AML, (DS-)AMKL, NKTCL | Ba/F3 cells expressing JAK3M511I or JAK3A573V are sensitive to Tofacitinib. JAK3A573V mutant NKTCL, and JAK3M511I mutant T-ALL-like disease models are responsive to Tofacitinib treatment in vivo. | [4,15,41,43,44,45,46,47,48,49,50,51,52,53,80,81,82] | |
| TYK2 | germline GOF(TYK2G716V, TYK2P760L) | pediatric ALL (B-ALL, T-ALL) | [55,56] | |
| somatic GOF(e.g., TYK2E957D) | T-ALL | Ba/F3 cells expressing TYK2E957D are sensitive to JAK inhibitor I. | [54,56] | |
| STAT3 | germline GOF(e.g., STAT3K392R) | pediatric LGLL | [64] | |
| somatic GOF(e.g., STAT3Y640F, STAT3D661Y/I/V/H) | T-LGLL, NK-LGLL, ALK-ALCL, HSTL, DLBCL NOS, NKTCL, CLPD-NK, ANKL, Sezary syndrome | Tofacitinib could be a promising salvage therapy for refractory T-LGLL patients with or without STAT3 mutations. | [60,61,63,65,83,84,85,86,87,88] | |
| STAT5B | somatic GOF(e.g., STAT5BN642H, STAT5BY665H/F) | NKTCL, ANKL, NK-LGLL, T-LGLL, T-PLL, T-ALL, MEITL, HSTL, PCTL, Sezary Syndrome, PTCL-NOS, AML, AAA, CNL, Eosinophilia | STAT5BN642H-driven CD8+ T-cell disease and CD56+ T-LGL (NKT) leukemia models are sensitive to Ruxolitinib. STAT5BN642H T-ALL is sensitive to JAK1/JAK3 inhibitors. | [59,60,61,62,72,73,86,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109] |
| STAT6 | somatic GOF(e.g., STAT6D419H/G/A) | CHL, FL, RR-DLBCL, PMBCL | STAT6D419-missense mutated PMBCL cell lines are sensitive to JAK2 inhibitors. | [66,67,68,69] |
| NCT number | JAKinib | Phase | Disease(s) 2 |
|---|---|---|---|
| NCT02723994 | Ruxolitinib (JAK1/2) | 2 | ALL |
| NCT03571321 | 1 | ALL (Ph-like) | |
| NCT03874052 | 1 | AML | |
| NCT03286530 | 2 | AML | |
| NCT04055844 | 2 | AML, MDS | |
| NCT03654768 | 2 | CML | |
| NCT03610971 | 2 | CML (chronic phase) | |
| NCT03722407 | 2 | CMML | |
| NCT03801434 | 2 | Eosinphilic syndromes | |
| NCT04669210 | 2 | GvHD, HSCT complications, ALL, AML | |
| NCT02613598 | 1 | (Non)Hodgkin Lymphoma | |
| NCT03681561 | 1/2 | Hodgkin Lymphoma | |
| NCT03017820 | 1 | Leukemia/lymphoma | |
| NCT03878199 | 1/2 | MPN | |
| NCT04281498 | 2 | MPN | |
| NCT04041050 | 1 | MPN | |
| NCT02158858 | 1/2 | MPN, MDS | |
| NCT03558607 | 1/2 | sAML | |
| NCT01712659 | 1/2 | T-cell Leukemia | |
| NCT03613428 | 1/2 | T-cell Leukemia | |
| NCT03117751 | 2/3 | T-cell Leukemia/Lymphoma | |
| NCT02974647 | 2 | T/NK lymphoma | |
| NCT04282187 | Ruxolitinib (JAK1/2) or Fedratinib (JAK2) | 2 | MPN, AML |
| NCT04282187 | Fedratinib (JAK2) | MPN, AML | |
| NCT03598959 | Tofacitinib (JAK1/3) | 2 | T/NK lymphoma |
| NCT04640025 | Itacitinib (JAK1) | 2 | MF |
| NCT01633372/NCT04629508 | 2 | MPN | |
| NCT03144687 | 2 | MPN | |
| NCT04061421 | 1/2 | MDS/MPN | |
| NCT03697408 | 1/2 | classical HL | |
| NCT02760485 | 1/2 | Relapsed or Refractory DLBCL | |
| NCT01905813 | 1 | B-cell Malignancies (previously treated) | |
| NCT02018861/NCT04509700 | 1/2 | B-cell Malignancies (previously treated) | |
| NCT03989466 | 1 | (recurrent) T-PLL | |
| NCT04173494 | Momelotinib (JAK1/2) | 3 | MPN (pMF, PV) |
| NCT03645824 | Patricinib (JAK2) | 2 | MF |
| NCT03165734 | 3 | MF (primary and secondary) | |
| NCT02891603 | 1/2 | GvHD |
| JAK/ STAT | Type of mutations | Immune phenotype of patients 3 | Immune phenotype of knockout/-in mice | References |
|---|---|---|---|---|
| JAK1 | LOF (e.g., JAK1P733L; JAK1P832S) | Immunodeficiency (early onset cancer and recurrent mycobacterial infections) | Perinatally lethal; severe reduction of pre–B cells, and mature T and B lymphocytes | [125,126] |
| JAK3 | LOF (e.g., JAK3Y100C; JAK3D169E) | autosomal recessive T-B+NK- SCID (null mutations), broader range of clinical immunosuppressive phenotypes | Defective T, B, ILC (incl. NK) cell development | [80,127,128,129,130,131,132,133] |
| TYK2 | LOF (e.g., TYK2I684S) | Mycobacterial and viral infections | Impaired T and NK-mediated anti-viral, anti-bacterial, and anti-tumor responses | [123,124,134,135,136,137,138] |
| STAT1 | LOF (e.g., STAT1K201L; STAT1K211R) | Complete deficiency: mycobacteria, virus infection; dysfunctional NK cells; partial deficiency: mycobacteria but no virus infection | Impaired responses to Type I and Type II IFN, increased susceptibility to infections, impaired NK cells | [139,140,141,142,143,144,145] |
| GOF (e.g., STAT1Y170N; STAT1C174R) | viral, bacterial infections, combined immunodeficiency (reduced memory B, Th17 cells, impaired NK cells); autoinflammation, organ-specific autoimmune disorders | Impaired IL-17 immunity | [146,147,148,149,150] | |
| STAT2 | LOF (e.g., STAT2c.1836C4A) | primary immunodeficiency (viral infections) | Impaired response to Type I IFN and susceptibility to viral infections | [113,114,151,152] |
| STAT3 | LOF (e.g., STAT3Y657N) | AD-HIES, primary immunodeficiency (susceptibilities to infections, impaired Th17 and B cells) | Embryonically lethal | [110,153,154,155] |
| GOF (e.g., STAT3Q344H) | Immune deficiency (reduced memory B cells, NK cells, pDCs); various organ autoimmunity | - | [64,110,156] | |
| STAT4 | LOF (e.g., STAT4E651V) | Fungal infections | Inhibited Th1 differentiation | [115,116,157] |
| STAT5B | LOF (e.g., STAT5BA630P) | combined immunodeficiency (Treg deficiency, reduced T cells and NK cells) serve viral infections; autoimmune symptoms | Impaired NK and T cells | [158,159,160,161,162,163,164] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Klein, K.; Stoiber, D.; Sexl, V.; Witalisz-Siepracka, A. Untwining Anti-Tumor and Immunosuppressive Effects of JAK Inhibitors—A Strategy for Hematological Malignancies? Cancers 2021, 13, 2611. https://doi.org/10.3390/cancers13112611
Klein K, Stoiber D, Sexl V, Witalisz-Siepracka A. Untwining Anti-Tumor and Immunosuppressive Effects of JAK Inhibitors—A Strategy for Hematological Malignancies? Cancers. 2021; 13(11):2611. https://doi.org/10.3390/cancers13112611
Chicago/Turabian StyleKlein, Klara, Dagmar Stoiber, Veronika Sexl, and Agnieszka Witalisz-Siepracka. 2021. "Untwining Anti-Tumor and Immunosuppressive Effects of JAK Inhibitors—A Strategy for Hematological Malignancies?" Cancers 13, no. 11: 2611. https://doi.org/10.3390/cancers13112611
APA StyleKlein, K., Stoiber, D., Sexl, V., & Witalisz-Siepracka, A. (2021). Untwining Anti-Tumor and Immunosuppressive Effects of JAK Inhibitors—A Strategy for Hematological Malignancies? Cancers, 13(11), 2611. https://doi.org/10.3390/cancers13112611






