FUT9-Driven Programming of Colon Cancer Cells towards a Stem Cell-Like State
Simple Summary
Abstract
1. Introduction
2. Results
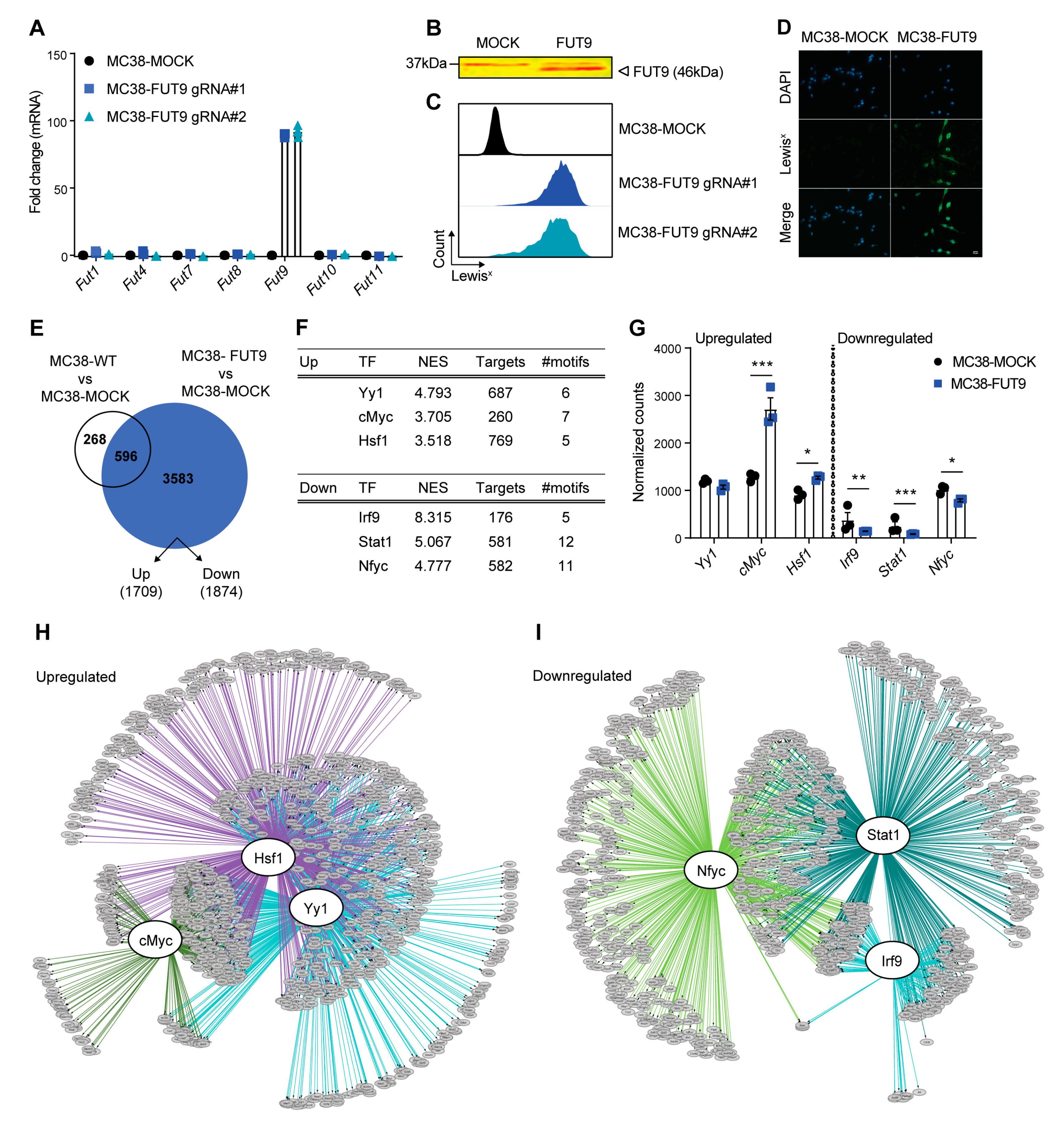
2.1. Identification of Gene Regulatory Networks Linking FUT9 Expression to Stemness in Glyco-Engineered Colon Cancer Cells
2.2. FUT9 Neo-Expression Results in an Enhanced Stem Cell-Like Transcriptional Profile and Phenotype
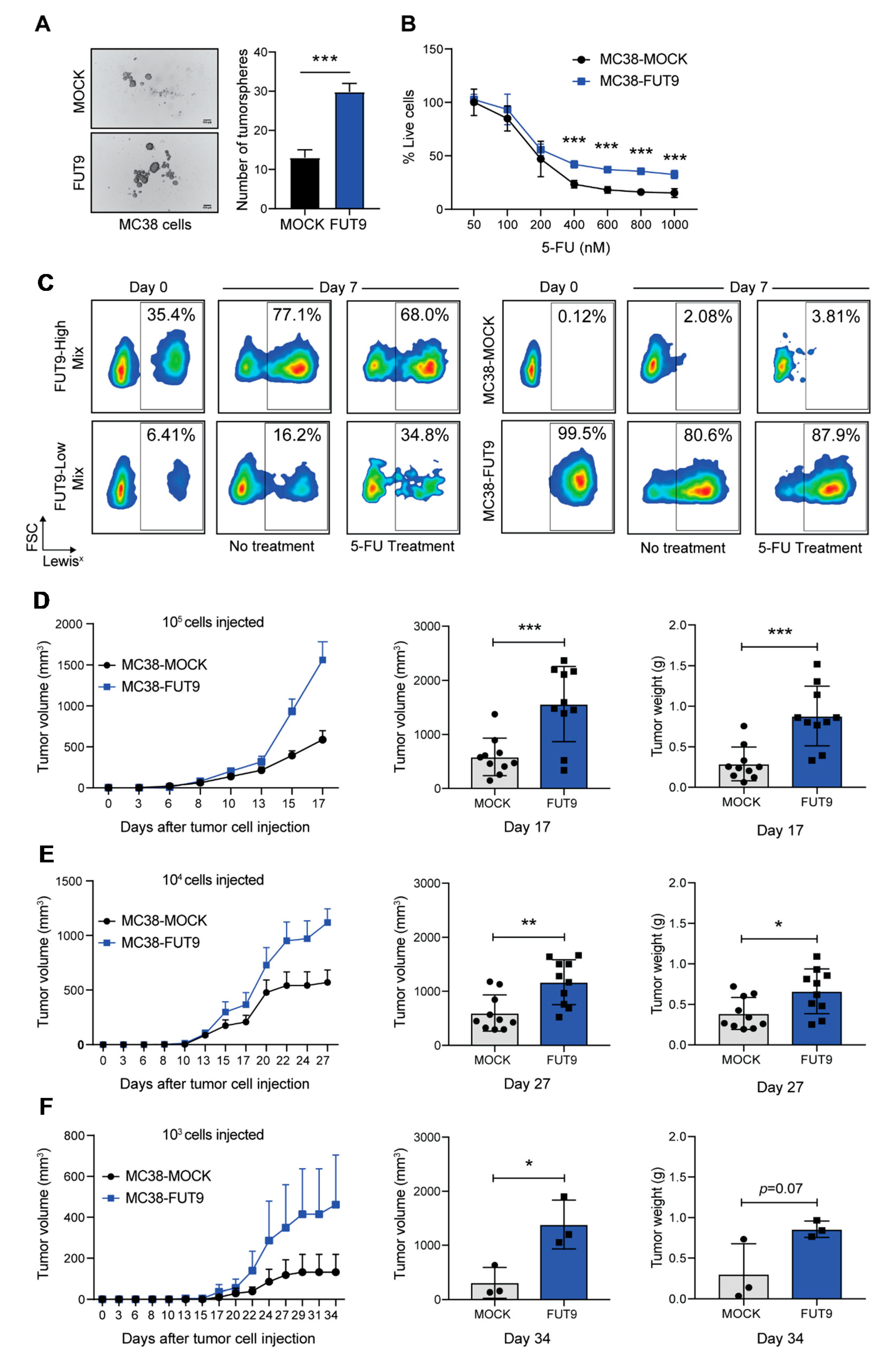
2.3. MC38-FUT9-Expressing Cells Display Functional Properties of Cancer Stem-Like Cells
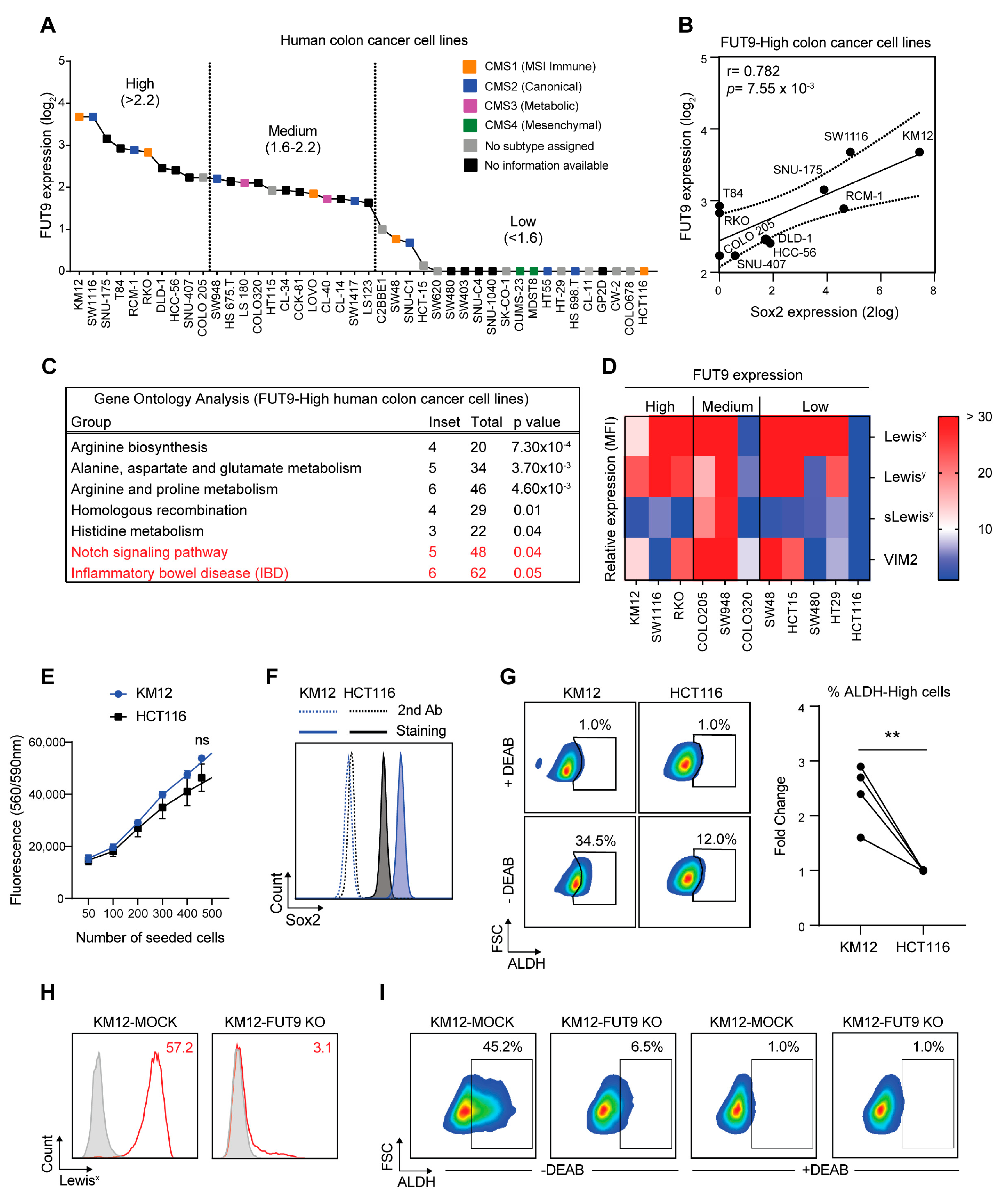
2.4. FUT9 Expression in Human Colon Cancer Cell Lines is Correlated with High Sox2 Expression and CSC-Like Features
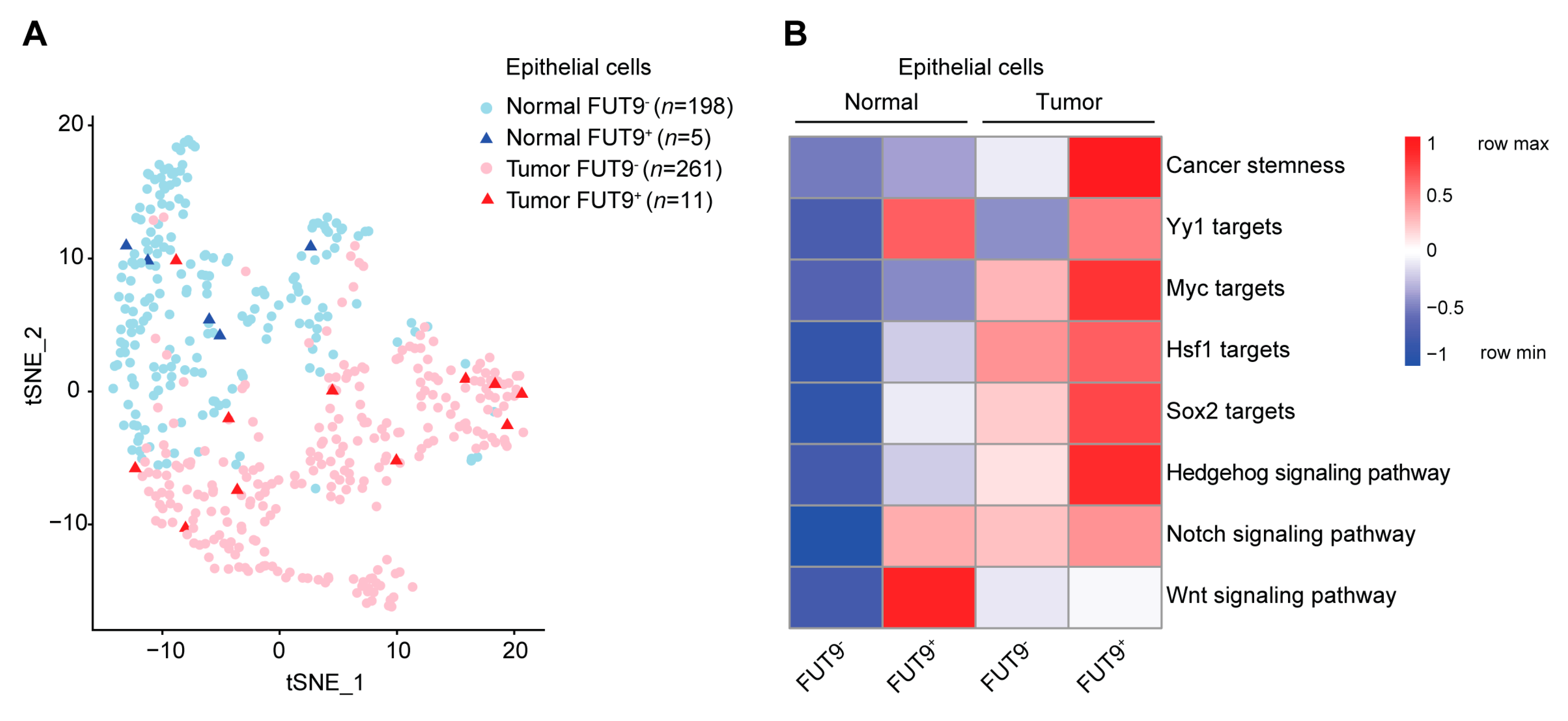
2.5. FUT9-Associated Transcriptional Programming of Cancer Stemness in Primary CRC Tumor Cells
3. Discussion
4. Materials and Methods
4.1. Mice
4.2. Cell Culture
4.3. Generation of Glyco-Engineered Colon Cancer Cell Lines
4.4. qRT-PCR Analysis
4.5. Immunoblotting
4.6. Flow Cytometry
4.7. Immunofluorescence
4.8. mRNA Library Preparation, RNA-Sequencing, Alignment and Differential Expression Analysis
4.9. Comparative Analysis of Gene Sets
4.10. Motif Enrichment Analysis
4.11. Gene Ontology Term Enrichment and Gene Signature Analysis
4.12. ALDH Activity
4.13. Tumorsphere Formation
4.14. Drug Resistance Analysis
4.15. Cancer Stem Cell (CSC)-Enrichment Assay
4.16. Cell Cycle Analysis
4.17. Cell Proliferation Assay
4.18. Public Databases
4.19. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Pineros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2019, 144, 1941–1953. [Google Scholar] [CrossRef] [PubMed]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reynies, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef] [PubMed]
- Tsikitis, V.L.; Malireddy, K.; Green, E.A.; Christensen, B.; Whelan, R.; Hyder, J.; Marcello, P.; Larach, S.; Lauter, D.; Sargent, D.J.; et al. Postoperative surveillance recommendations for early stage colon cancer based on results from the clinical outcomes of surgical therapy trial. J. Clin. Oncol. 2009, 27, 3671–3676. [Google Scholar] [CrossRef] [PubMed]
- Douillard, J.Y.; Cunningham, D.; Roth, A.D.; Navarro, M.; James, R.D.; Karasek, P.; Jandik, P.; Iveson, T.; Carmichael, J.; Alakl, M.; et al. Irinotecan combined with fluorouracil compared with fluorouracil alone as first-line treatment for metastatic colorectal cancer: A multicentre randomised trial. Lancet 2000, 355, 1041–1047. [Google Scholar] [CrossRef]
- Giacchetti, S.; Perpoint, B.; Zidani, R.; Le Bail, N.; Faggiuolo, R.; Focan, C.; Chollet, P.; Llory, J.F.; Letourneau, Y.; Coudert, B.; et al. Phase III multicenter randomized trial of oxaliplatin added to chronomodulated fluorouracil-leucovorin as first-line treatment of metastatic colorectal cancer. J. Clin. Oncol. 2000, 18, 136–147. [Google Scholar] [CrossRef]
- Syn, N.L.; Teng, M.W.L.; Mok, T.S.K.; Soo, R.A. De-novo and acquired resistance to immune checkpoint targeting. Lancet Oncol. 2017, 18, e731–e741. [Google Scholar] [CrossRef]
- Yaffee, P.; Osipov, A.; Tan, C.; Tuli, R.; Hendifar, A. Review of systemic therapies for locally advanced and metastatic rectal cancer. J. Gastrointest. Oncol. 2015, 6, 185–200. [Google Scholar] [CrossRef]
- Ganesh, K.; Stadler, Z.K.; Cercek, A.; Mendelsohn, R.B.; Shia, J.; Segal, N.H.; Diaz, L.A., Jr. Immunotherapy in colorectal cancer: Rationale, challenges and potential. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 361–375. [Google Scholar] [CrossRef]
- Ychou, M.; Rivoire, M.; Thezenas, S.; Quenet, F.; Delpero, J.R.; Rebischung, C.; Letoublon, C.; Guimbaud, R.; Francois, E.; Ducreux, M.; et al. A randomized phase II trial of three intensified chemotherapy regimens in first-line treatment of colorectal cancer patients with initially unresectable or not optimally resectable liver metastases. The METHEP trial. Ann. Surg. Oncol. 2013, 20, 4289–4297. [Google Scholar] [CrossRef]
- Zhou, Y.; Xia, L.; Wang, H.; Oyang, L.; Su, M.; Liu, Q.; Lin, J.; Tan, S.; Tian, Y.; Liao, Q.; et al. Cancer stem cells in progression of colorectal cancer. Oncotarget 2018, 9, 33403–33415. [Google Scholar] [CrossRef]
- Clevers, H. The cancer stem cell: Premises, promises and challenges. Nat. Med. 2011, 17, 313–319. [Google Scholar] [CrossRef]
- Nassar, D.; Blanpain, C. Cancer Stem Cells: Basic Concepts and Therapeutic Implications. Annu. Rev. Pathol. 2016, 11, 47–76. [Google Scholar] [CrossRef]
- Lenos, K.J.; Miedema, D.M.; Lodestijn, S.C.; Nijman, L.E.; van den Bosch, T.; Romero Ros, X.; Lourenco, F.C.; Lecca, M.C.; van der Heijden, M.; van Neerven, S.M.; et al. Stem cell functionality is microenvironmentally defined during tumour expansion and therapy response in colon cancer. Nat. Cell Biol. 2018, 20, 1193–1202. [Google Scholar] [CrossRef]
- Batlle, E.; Clevers, H. Cancer stem cells revisited. Nat. Med. 2017, 23, 1124–1134. [Google Scholar] [CrossRef] [PubMed]
- Mallard, B.W.; Tiralongo, J. Cancer stem cell marker glycosylation: Nature, function and significance. Glycoconj. J. 2017, 34, 441–452. [Google Scholar] [CrossRef] [PubMed]
- Barkeer, S.; Chugh, S.; Batra, S.K.; Ponnusamy, M.P. Glycosylation of Cancer Stem Cells: Function in Stemness, Tumorigenesis, and Metastasis. Neoplasia 2018, 20, 813–825. [Google Scholar] [CrossRef] [PubMed]
- Varki, A. Biological roles of glycans. Glycobiology 2017, 27, 3–49. [Google Scholar] [CrossRef]
- Pinho, S.S.; Reis, C.A. Glycosylation in cancer: Mechanisms and clinical implications. Nat. Rev. Cancer 2015, 15, 540–555. [Google Scholar] [CrossRef]
- Blanas, A.; Sahasrabudhe, N.M.; Rodriguez, E.; van Kooyk, Y.; van Vliet, S.J. Fucosylated Antigens in Cancer: An Alliance toward Tumor Progression, Metastasis, and Resistance to Chemotherapy. Front. Oncol. 2018, 8, 39. [Google Scholar] [CrossRef]
- Saldova, R.; Struwe, W.B.; Wynne, K.; Elia, G.; Duffy, M.J.; Rudd, P.M. Exploring the glycosylation of serum CA125. Int. J. Mol. Sci. 2013, 14, 15636–15654. [Google Scholar] [CrossRef]
- Scara, S.; Bottoni, P.; Scatena, R. CA 19-9: Biochemical and Clinical Aspects. Adv. Exp. Med. Biol. 2015, 867, 247–260. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Zhan, T.; Deng, Z.; Li, Q.; Liu, Y.; Yang, S.; Ji, D.; Li, Y. Glycan analysis of colorectal cancer samples reveals stage-dependent changes in CEA glycosylation patterns. Clin. Proteom. 2018, 15, 9. [Google Scholar] [CrossRef]
- Dirkse, A.; Golebiewska, A.; Buder, T.; Nazarov, P.V.; Muller, A.; Poovathingal, S.; Brons, N.H.C.; Leite, S.; Sauvageot, N.; Sarkisjan, D.; et al. Stem cell-associated heterogeneity in Glioblastoma results from intrinsic tumor plasticity shaped by the microenvironment. Nat. Commun. 2019, 10, 1787. [Google Scholar] [CrossRef]
- Friedman, G.K.; Moore, B.P.; Nan, L.; Kelly, V.M.; Etminan, T.; Langford, C.P.; Xu, H.; Han, X.; Markert, J.M.; Beierle, E.A.; et al. Pediatric medulloblastoma xenografts including molecular subgroup 3 and CD133+ and CD15+ cells are sensitive to killing by oncolytic herpes simplex viruses. Neuro. Oncol. 2016, 18, 227–235. [Google Scholar] [CrossRef]
- Mondal, N.; Dykstra, B.; Lee, J.; Ashline, D.J.; Reinhold, V.N.; Rossi, D.J.; Sackstein, R. Distinct human alpha(1,3)-fucosyltransferases drive Lewis-X/sialyl Lewis-X assembly in human cells. J. Biol. Chem. 2018, 293, 7300–7314. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, M.; Kudo, T.; Iwasaki, H.; Ikehara, Y.; Nishihara, S.; Nakagawa, S.; Sasaki, K.; Shiina, T.; Inoko, H.; Saitou, N.; et al. Alpha1,3-fucosyltransferase IX (Fuc-TIX) is very highly conserved between human and mouse; molecular cloning, characterization and tissue distribution of human Fuc-TIX. FEBS Lett. 1999, 452, 237–242. [Google Scholar] [CrossRef]
- Cailleau-Thomas, A.; Coullin, P.; Candelier, J.J.; Balanzino, L.; Mennesson, B.; Oriol, R.; Mollicone, R. FUT4 and FUT9 genes are expressed early in human embryogenesis. Glycobiology 2000, 10, 789–802. [Google Scholar] [CrossRef] [PubMed]
- Stadlmann, J.; Taubenschmid, J.; Wenzel, D.; Gattinger, A.; Durnberger, G.; Dusberger, F.; Elling, U.; Mach, L.; Mechtler, K.; Penninger, J.M. Comparative glycoproteomics of stem cells identifies new players in ricin toxicity. Nature 2017, 549, 538–542. [Google Scholar] [CrossRef]
- Noro, E.; Togayachi, A.; Sato, T.; Tomioka, A.; Fujita, M.; Sukegawa, M.; Suzuki, N.; Kaji, H.; Narimatsu, H. Large-Scale Identification of N-Glycan Glycoproteins Carrying Lewis x and Site-Specific N-Glycan Alterations in Fut9 Knockout Mice. J. Proteome Res. 2015, 14, 3823–3834. [Google Scholar] [CrossRef]
- Gouveia, R.; Schaffer, L.; Papp, S.; Grammel, N.; Kandzia, S.; Head, S.R.; Kleene, R.; Schachner, M.; Conradt, H.S.; Costa, J. Expression of glycogenes in differentiating human NT2N neurons. Downregulation of fucosyltransferase 9 leads to decreased Lewis(x) levels and impaired neurite outgrowth. Biochim. Biophys. Acta 2012, 1820, 2007–2019. [Google Scholar] [CrossRef]
- Auslander, N.; Cunningham, C.E.; Toosi, B.M.; McEwen, E.J.; Yizhak, K.; Vizeacoumar, F.S.; Parameswaran, S.; Gonen, N.; Freywald, T.; Bhanumathy, K.K.; et al. An integrated computational and experimental study uncovers FUT9 as a metabolic driver of colorectal cancer. Mol. Syst. Biol. 2017, 13, 956. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Li, L.; Starr, T.K.; Subramanian, S. Tumor location impacts immune response in mouse models of colon cancer. Oncotarget 2017, 8, 54775–54787. [Google Scholar] [CrossRef] [PubMed]
- Blanas, A.; Cornelissen, L.A.M.; Kotsias, M.; van der Horst, J.C.; van de Vrugt, H.J.; Kalay, H.; Spencer, D.I.R.; Kozak, R.P.; van Vliet, S.J. Transcriptional activation of fucosyltransferase (FUT) genes using the CRISPR-dCas9-VPR technology reveals potent N-glycome alterations in colorectal cancer cells. Glycobiology 2019, 29, 137–150. [Google Scholar] [CrossRef] [PubMed]
- Janky, R.; Verfaillie, A.; Imrichova, H.; Van de Sande, B.; Standaert, L.; Christiaens, V.; Hulselmans, G.; Herten, K.; Naval Sanchez, M.; Potier, D.; et al. iRegulon: From a gene list to a gene regulatory network using large motif and track collections. PLoS Comput. Biol. 2014, 10, e1003731. [Google Scholar] [CrossRef] [PubMed]
- Varum, S.; Baggiolini, A.; Zurkirchen, L.; Atak, Z.K.; Cantu, C.; Marzorati, E.; Bossart, R.; Wouters, J.; Hausel, J.; Tuncer, E.; et al. Yin Yang 1 Orchestrates a Metabolic Program Required for Both Neural Crest Development and Melanoma Formation. Cell Stem. Cell 2019, 24, 637–653. [Google Scholar] [CrossRef] [PubMed]
- Sarvagalla, S.; Kolapalli, S.P.; Vallabhapurapu, S. The Two Sides of YY1 in Cancer: A Friend and a Foe. Front. Oncol. 2019, 9, 1230. [Google Scholar] [CrossRef]
- Elbadawy, M.; Usui, T.; Yamawaki, H.; Sasaki, K. Emerging Roles of C-Myc in Cancer Stem Cell-Related Signaling and Resistance to Cancer Chemotherapy: A Potential Therapeutic Target Against Colorectal Cancer. Int. J. Mol. Sci. 2019, 20, 2340. [Google Scholar] [CrossRef]
- Carpenter, R.L.; Gokmen-Polar, Y. HSF1 as a Cancer Biomarker and Therapeutic Target. Curr. Cancer Drug Targets 2019, 19, 515–524. [Google Scholar] [CrossRef]
- Michalska, A.; Blaszczyk, K.; Wesoly, J.; Bluyssen, H.A.R. A Positive Feedback Amplifier Circuit that Regulates Interferon (IFN)-Stimulated Gene Expression and Controls Type I and Type II IFN Responses. Front. Immunol. 2018, 9, 1135. [Google Scholar] [CrossRef]
- Doherty, M.R.; Cheon, H.; Junk, D.J.; Vinayak, S.; Varadan, V.; Telli, M.L.; Ford, J.M.; Stark, G.R.; Jackson, M.W. Interferon-beta represses cancer stem cell properties in triple-negative breast cancer. Proc. Natl. Acad. Sci. USA 2017, 114, 13792–13797. [Google Scholar] [CrossRef]
- Ohsugi, T.; Yamaguchi, K.; Zhu, C.; Ikenoue, T.; Takane, K.; Shinozaki, M.; Tsurita, G.; Yano, H.; Furukawa, Y. Anti-apoptotic effect by the suppression of IRF1 as a downstream of Wnt/beta-catenin signaling in colorectal cancer cells. Oncogene 2019, 38, 6051–6064. [Google Scholar] [CrossRef] [PubMed]
- Hong, M.; Zhang, Z.; Chen, Q.; Lu, Y.; Zhang, J.; Lin, C.; Zhang, F.; Zhang, W.; Li, X.; Zhang, W.; et al. IRF1 inhibits the proliferation and metastasis of colorectal cancer by suppressing the RAS-RAC1 pathway. Cancer Manag. Res. 2019, 11, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Zhang, J.; Liu, H.; Wan, L.; Zhang, H.; Huang, Q.; Xu, E.; Lai, M. IL-13/STAT6 signaling plays a critical role in the epithelial-mesenchymal transition of colorectal cancer cells. Oncotarget 2016, 7, 61183–61198. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Oron, E.; Nelson, B.; Razis, S.; Ivanova, N. Distinct lineage specification roles for NANOG, OCT4, and SOX2 in human embryonic stem cells. Cell Stem Cell 2012, 10, 440–454. [Google Scholar] [CrossRef] [PubMed]
- Velychko, S.; Adachi, K.; Kim, K.P.; Hou, Y.; MacCarthy, C.M.; Wu, G.; Scholer, H.R. Excluding Oct4 from Yamanaka Cocktail Unleashes the Developmental Potential of iPSCs. Cell Stem Cell 2019, 25, 737–753. [Google Scholar] [CrossRef]
- Abdul Khalek, F.J.; Gallicano, G.I.; Mishra, L. Colon cancer stem cells. Gastrointest. Cancer Res. 2010, (Suppl. 1), S16–S23. [Google Scholar]
- Vassalli, G. Aldehyde Dehydrogenases: Not Just Markers, but Functional Regulators of Stem Cells. Stem. Cells Int. 2019, 2019, 3904645. [Google Scholar] [CrossRef]
- Chaicharoenaudomrung, N.; Kunhorm, P.; Noisa, P. Three-dimensional cell culture systems as an in vitro platform for cancer and stem cell modeling. World J. Stem Cells 2019, 11, 1065–1083. [Google Scholar] [CrossRef]
- Blanas, A. Amsterdam Infection & Immunity Institute, Cancer Center Amsterdam, Department of Molecular Cell Biology and Immunology, Vrije Universiteit Amsterdam, Amsterdam UMC, 1081 HZ, Amsterdam, The Netherlands. MuBase data mining and RNAseq analysis reveals that MC38-WT and MC38-glyco-engineered cells, respectively, only express CD44 and not the stem cell-associated molecules CD133, Lgr5, CD24, EpCAM and ABCG2, 2020.
- Linnekamp, J.F.; Hooff, S.R.V.; Prasetyanti, P.R.; Kandimalla, R.; Buikhuisen, J.Y.; Fessler, E.; Ramesh, P.; Lee, K.; Bochove, G.G.W.; de Jong, J.H.; et al. Consensus molecular subtypes of colorectal cancer are recapitulated in in vitro and in vivo models. Cell Death Differ. 2018, 25, 616–633. [Google Scholar] [CrossRef]
- Blanas, A. Amsterdam Infection & Immunity Institute, Cancer Center Amsterdam, Department of Molecular Cell Biology and Immunology, Vrije Universiteit Amsterdam, Amsterdam UMC, 1081 HZ, Amsterdam, The Netherlands. No significant correlation exists between the expression of Sox2 and FUT9 in the human cell lines expressing medium or low FUT9 levels, 2020.
- Muchero, W.; Ehlers, J.D.; Close, T.J.; Roberts, P.A. Mapping QTL for drought stress-induced premature senescence and maturity in cowpea [Vigna unguiculata (L.) Walp.]. Theor. Appl. Genet. 2009, 118, 849–863. [Google Scholar] [CrossRef]
- Moriwaki, K.; Noda, K.; Furukawa, Y.; Ohshima, K.; Uchiyama, A.; Nakagawa, T.; Taniguchi, N.; Daigo, Y.; Nakamura, Y.; Hayashi, N.; et al. Deficiency of GMDS leads to escape from NK cell-mediated tumor surveillance through modulation of TRAIL signaling. Gastroenterology 2009, 137, 188–198. [Google Scholar] [CrossRef] [PubMed]
- DeBose-Boyd, R.A.; Nyame, A.K.; Smith, D.F.; Cummings, R.D. alpha1,4-Fucosyltransferase activity in human serum and saliva. Arch. Biochem. Biophys. 1996, 335, 109–117. [Google Scholar] [CrossRef] [PubMed]
- Holst, S.; Deuss, A.J.; van Pelt, G.W.; van Vliet, S.J.; Garcia-Vallejo, J.J.; Koeleman, C.A.; Deelder, A.M.; Mesker, W.E.; Tollenaar, R.A.; Rombouts, Y.; et al. N-glycosylation Profiling of Colorectal Cancer Cell Lines Reveals Association of Fucosylation with Differentiation and Caudal Type Homebox 1 (CDX1)/Villin mRNA Expression. Mol. Cell. Proteom. 2016, 15, 124–140. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Courtois, E.T.; Sengupta, D.; Tan, Y.; Chen, K.H.; Goh, J.J.L.; Kong, S.L.; Chua, C.; Hon, L.K.; Tan, W.S.; et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 2017, 49, 708–718. [Google Scholar] [CrossRef] [PubMed]
- Blanas, A.; Kok, M.d. Amsterdam Infection & Immunity Institute, Cancer Center Amsterdam, Department of Molecular Cell Biology and Immunology, Vrije Universiteit Amsterdam, Amsterdam UMC, 1081 HZ, Amsterdam, The Netherlands. Single-cell transcriptomic data derived from 11 colorectal cancer primary tumors reveals that B cells express low levels of FUT9, 2020.
- Miranda, A.; Hamilton, P.T.; Zhang, A.W.; Pattnaik, S.; Becht, E.; Mezheyeuski, A.; Bruun, J.; Micke, P.; de Reynies, A.; Nelson, B.H. Cancer stemness, intratumoral heterogeneity, and immune response across cancers. Proc. Natl. Acad. Sci. USA 2019, 116, 9020–9029. [Google Scholar] [CrossRef] [PubMed]
- Affar el, B.; Gay, F.; Shi, Y.; Liu, H.; Huarte, M.; Wu, S.; Collins, T.; Li, E.; Shi, Y. Essential dosage-dependent functions of the transcription factor yin yang 1 in late embryonic development and cell cycle progression. Mol. Cell. Biol. 2006, 26, 3565–3581. [Google Scholar] [CrossRef]
- Schaefer, C.F.; Anthony, K.; Krupa, S.; Buchoff, J.; Day, M.; Hannay, T.; Buetow, K.H. PID: The Pathway Interaction Database. Nucleic. Acids Res. 2009, 37, D674–D679. [Google Scholar] [CrossRef]
- Ben-Porath, I.; Thomson, M.W.; Carey, V.J.; Ge, R.; Bell, G.W.; Regev, A.; Weinberg, R.A. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat. Genet. 2008, 40, 499–507. [Google Scholar] [CrossRef]
- Wei, S.; Chen, J.; Huang, Y.; Sun, Q.; Wang, H.; Liang, X.; Hu, Z.; Li, X. Identification of hub genes and construction of transcriptional regulatory network for the progression of colon adenocarcinoma hub genes and TF regulatory network of colon adenocarcinoma. J. Cell. Physiol. 2020, 235, 2037–2048. [Google Scholar] [CrossRef]
- Uddin, M.N.; Li, M.; Wang, X. Identification of Transcriptional Signatures of Colon Tumor Stroma by a Meta-Analysis. J. Oncol. 2019, 2019, 8752862. [Google Scholar] [CrossRef]
- Poli, V.; Fagnocchi, L.; Fasciani, A.; Cherubini, A.; Mazzoleni, S.; Ferrillo, S.; Miluzio, A.; Gaudioso, G.; Vaira, V.; Turdo, A.; et al. MYC-driven epigenetic reprogramming favors the onset of tumorigenesis by inducing a stem cell-like state. Nat. Commun. 2018, 9, 1024. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Labbadia, J.; Morimoto, R.I. Rethinking HSF1 in Stress, Development, and Organismal Health. Trends Cell Biol. 2017, 27, 895–905. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.; Qian, M.; Lu, L.; Chen, Y.; Zhang, X.; Wu, Q.; Liu, Y.; Bian, Z.; Yang, Y.; Guo, S.; et al. O-GlcNAcylation of YY1 stimulates tumorigenesis in colorectal cancer cells by targeting SLC22A15 and AANAT. Carcinogenesis 2019, 40, 1121–1131. [Google Scholar] [CrossRef]
- Wang, G.Z.; Goff, S.P. Regulation of Yin Yang 1 by Tyrosine Phosphorylation. J. Biol. Chem. 2015, 290, 21890–21900. [Google Scholar] [CrossRef] [PubMed]
- Hadjimichael, C.; Chanoumidou, K.; Papadopoulou, N.; Arampatzi, P.; Papamatheakis, J.; Kretsovali, A. Common stemness regulators of embryonic and cancer stem cells. World J. Stem Cells 2015, 7, 1150–1184. [Google Scholar] [CrossRef] [PubMed]
- Takeda, K.; Mizushima, T.; Yokoyama, Y.; Hirose, H.; Wu, X.; Qian, Y.; Ikehata, K.; Miyoshi, N.; Takahashi, H.; Haraguchi, N.; et al. Sox2 is associated with cancer stem-like properties in colorectal cancer. Sci. Rep. 2018, 8, 17639. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, T.; Lengerke, C. SOX2 protein biochemistry in stemness, reprogramming, and cancer: The PI3K/AKT/SOX2 axis and beyond. Oncogene 2020, 39, 278–292. [Google Scholar] [CrossRef]
- He, L.; Zhu, H.; Zhou, S.; Wu, T.; Wu, H.; Yang, H.; Mao, H.; SekharKathera, C.; Janardhan, A.; Edick, A.M.; et al. Wnt pathway is involved in 5-FU drug resistance of colorectal cancer cells. Exp. Mol. Med. 2018, 50, 101. [Google Scholar] [CrossRef]
- Regan, J.L.; Schumacher, D.; Staudte, S.; Steffen, A.; Haybaeck, J.; Keilholz, U.; Schweiger, C.; Golob-Schwarzl, N.; Mumberg, D.; Henderson, D.; et al. Non-Canonical Hedgehog Signaling Is a Positive Regulator of the WNT Pathway and Is Required for the Survival of Colon Cancer Stem Cells. Cell Rep. 2017, 21, 2813–2828. [Google Scholar] [CrossRef]
- Akbarzadeh, M.; Maroufi, N.F.; Tazehkand, A.P.; Akbarzadeh, M.; Bastani, S.; Safdari, R.; Farzane, A.; Fattahi, A.; Nejabati, H.R.; Nouri, M.; et al. Current approaches in identification and isolation of cancer stem cells. J. Cell. Physiol. 2019, 234, 14759–14772. [Google Scholar] [CrossRef]
- Son, M.J.; Woolard, K.; Nam, D.H.; Lee, J.; Fine, H.A. SSEA-1 is an enrichment marker for tumor-initiating cells in human glioblastoma. Cell Stem Cell 2009, 4, 440–452. [Google Scholar] [CrossRef]
- Swindall, A.F.; Londono-Joshi, A.I.; Schultz, M.J.; Fineberg, N.; Buchsbaum, D.J.; Bellis, S.L. ST6Gal-I protein expression is upregulated in human epithelial tumors and correlates with stem cell markers in normal tissues and colon cancer cell lines. Cancer Res. 2013, 73, 2368–2378. [Google Scholar] [CrossRef]
- Schultz, M.J.; Holdbrooks, A.T.; Chakraborty, A.; Grizzle, W.E.; Landen, C.N.; Buchsbaum, D.J.; Conner, M.G.; Arend, R.C.; Yoon, K.J.; Klug, C.A.; et al. The Tumor-Associated Glycosyltransferase ST6Gal-I Regulates Stem Cell Transcription Factors and Confers a Cancer Stem Cell Phenotype. Cancer Res. 2016, 76, 3978–3988. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Nagy, T.; Pierce, M. Post-translational glycoprotein modifications regulate colon cancer stem cells and colon adenoma progression in Apcmin/+ mice through altered Wnt receptor signaling. J. Biol. Chem. 2014, 289, 31534–31549. [Google Scholar] [CrossRef] [PubMed]
- Che, M.I.; Huang, J.; Hung, J.S.; Lin, Y.C.; Huang, M.J.; Lai, H.S.; Hsu, W.M.; Liang, J.T.; Huang, M.C. beta1, 4-N-acetylgalactosaminyltransferase III modulates cancer stemness through EGFR signaling pathway in colon cancer cells. Oncotarget 2014, 5, 3673–3684. [Google Scholar] [CrossRef] [PubMed]
- Very, N.; Lefebvre, T.; El Yazidi-Belkoura, I. Drug resistance related to aberrant glycosylation in colorectal cancer. Oncotarget 2018, 9, 1380–1402. [Google Scholar] [CrossRef] [PubMed]
- Schulz, A.; Meyer, F.; Dubrovska, A.; Borgmann, K. Cancer Stem Cells and Radioresistance: DNA Repair and Beyond. Cancers 2019, 11, 862. [Google Scholar] [CrossRef] [PubMed]
- Setubal Destro Rodrigues, M.F.; Gammon, L.; Rahman, M.M.; Biddle, A.; Nunes, F.D.; Mackenzie, I.C. Effects of Cetuximab and Erlotinib on the behaviour of cancer stem cells in head and neck squamous cell carcinoma. Oncotarget 2018, 9, 13488–13500. [Google Scholar] [CrossRef] [PubMed]
- Huyghe, N.; Baldin, P.; Van den Eynde, M. Immunotherapy with immune checkpoint inhibitors in colorectal cancer: What is the future beyond deficient mismatch-repair tumours? Gastroenterol. Rep. 2020, 8, 11–24. [Google Scholar] [CrossRef] [PubMed]
- Codd, A.S.; Kanaseki, T.; Torigo, T.; Tabi, Z. Cancer stem cells as targets for immunotherapy. Immunology 2018, 153, 304–314. [Google Scholar] [CrossRef]
- Wei, F.; Zhang, T.; Deng, S.C.; Wei, J.C.; Yang, P.; Wang, Q.; Chen, Z.P.; Li, W.L.; Chen, H.C.; Hu, H.; et al. PD-L1 promotes colorectal cancer stem cell expansion by activating HMGA1-dependent signaling pathways. Cancer Lett. 2019, 450, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Tallerico, R.; Todaro, M.; Di Franco, S.; Maccalli, C.; Garofalo, C.; Sottile, R.; Palmieri, C.; Tirinato, L.; Pangigadde, P.N.; La Rocca, R.; et al. Human NK cells selective targeting of colon cancer-initiating cells: A role for natural cytotoxicity receptors and MHC class I molecules. J. Immunol. 2013, 190, 2381–2390. [Google Scholar] [CrossRef] [PubMed]
- Budhwani, M.; Mazzieri, R.; Dolcetti, R. Plasticity of Type I Interferon-Mediated Responses in Cancer Therapy: From Anti-tumor Immunity to Resistance. Front. Oncol. 2018, 8, 322. [Google Scholar] [CrossRef] [PubMed]
- Ran, F.A.; Hsu, P.D.; Wright, J.; Agarwala, V.; Scott, D.A.; Zhang, F. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 2013, 8, 2281–2308. [Google Scholar] [CrossRef]
- Cornelissen, L.A.M.; Blanas, A.; van der Horst, J.C.; Kruijssen, L.; Zaal, A.; O’Toole, T.; Wiercx, L.; van Kooyk, Y.; van Vliet, S.J. Disruption of sialic acid metabolism drives tumor growth by augmenting CD8(+) T cell apoptosis. Int. J. Cancer 2019, 144, 2290–2302. [Google Scholar] [CrossRef]
- Zaal, A.; Li, R.J.E.; Lubbers, J.; Bruijns, S.C.M.; Kalay, H.; van Kooyk, Y.; van Vliet, S.J. Activation of the C-Type Lectin MGL by Terminal GalNAc Ligands Reduces the Glycolytic Activity of Human Dendritic Cells. Front. Immunol. 2020, 11, 305. [Google Scholar] [CrossRef]
- Cornelissen, L.A.M.; Blanas, A.; Zaal, A.; van der Horst, J.C.; Kruijssen, L.J.W.; O’Toole, T.; van Kooyk, Y.; van Vliet, S.J. Tn Antigen Expression Contributes to an Immune Suppressive Microenvironment and Drives Tumor Growth in Colorectal Cancer. Front. Oncol. 2020, 10, 1622. [Google Scholar] [CrossRef]
- Butler, A.; Hoffman, P.; Smibert, P.; Papalexi, E.; Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 2018, 36, 411–420. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Blanas, A.; Zaal, A.; van der Haar Àvila, I.; Kempers, M.; Kruijssen, L.; de Kok, M.; Popovic, M.A.; van der Horst, J.C.; J. van Vliet, S. FUT9-Driven Programming of Colon Cancer Cells towards a Stem Cell-Like State. Cancers 2020, 12, 2580. https://doi.org/10.3390/cancers12092580
Blanas A, Zaal A, van der Haar Àvila I, Kempers M, Kruijssen L, de Kok M, Popovic MA, van der Horst JC, J. van Vliet S. FUT9-Driven Programming of Colon Cancer Cells towards a Stem Cell-Like State. Cancers. 2020; 12(9):2580. https://doi.org/10.3390/cancers12092580
Chicago/Turabian StyleBlanas, Athanasios, Anouk Zaal, Irene van der Haar Àvila, Maxime Kempers, Laura Kruijssen, Mike de Kok, Marko A. Popovic, Joost C. van der Horst, and Sandra J. van Vliet. 2020. "FUT9-Driven Programming of Colon Cancer Cells towards a Stem Cell-Like State" Cancers 12, no. 9: 2580. https://doi.org/10.3390/cancers12092580
APA StyleBlanas, A., Zaal, A., van der Haar Àvila, I., Kempers, M., Kruijssen, L., de Kok, M., Popovic, M. A., van der Horst, J. C., & J. van Vliet, S. (2020). FUT9-Driven Programming of Colon Cancer Cells towards a Stem Cell-Like State. Cancers, 12(9), 2580. https://doi.org/10.3390/cancers12092580








