Pan-Cancer Analysis of Alternative Lengthening of Telomere Activity
Abstract
1. Introduction
2. Results
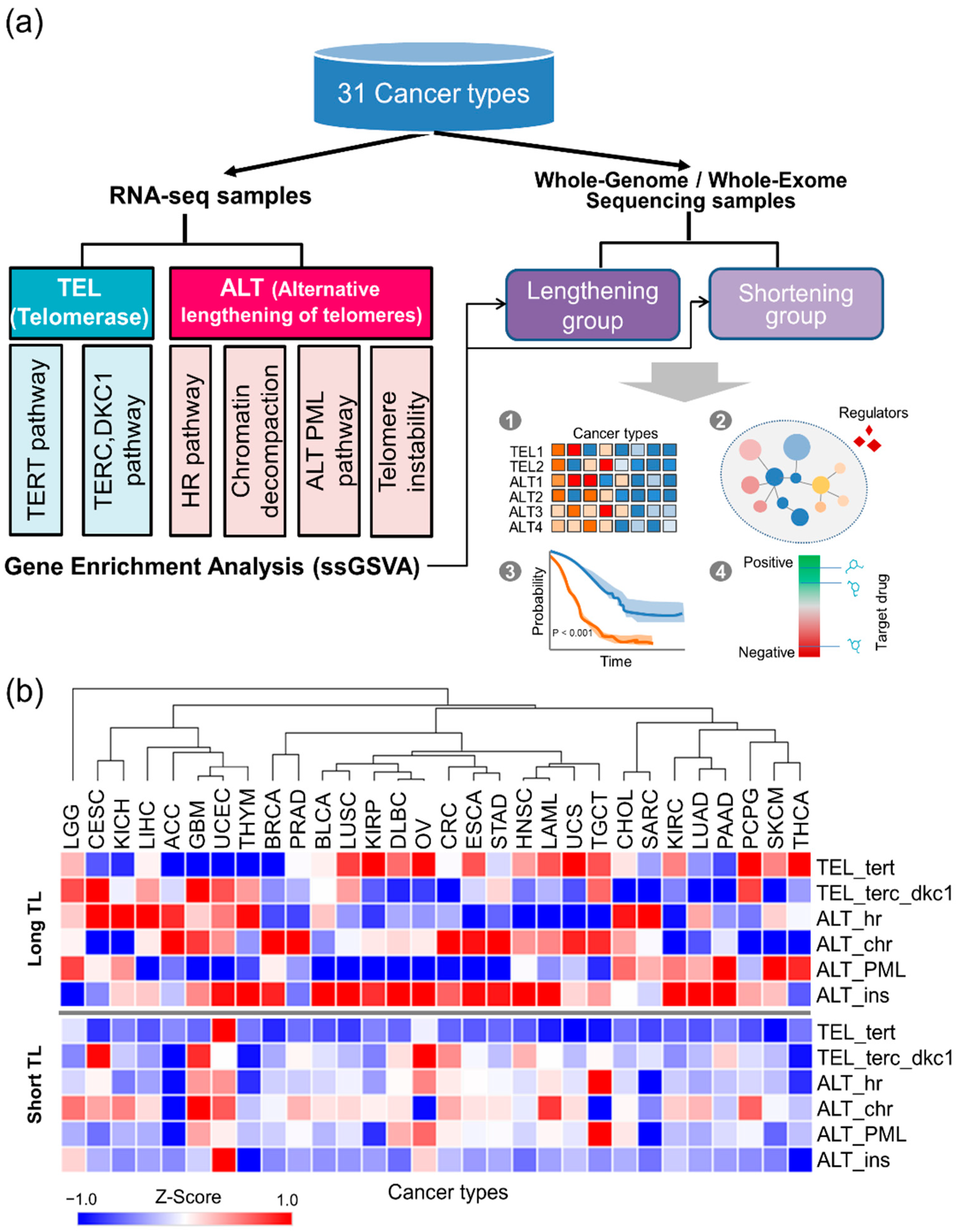
2.1. TMM Activities Across 31 Cancer Types
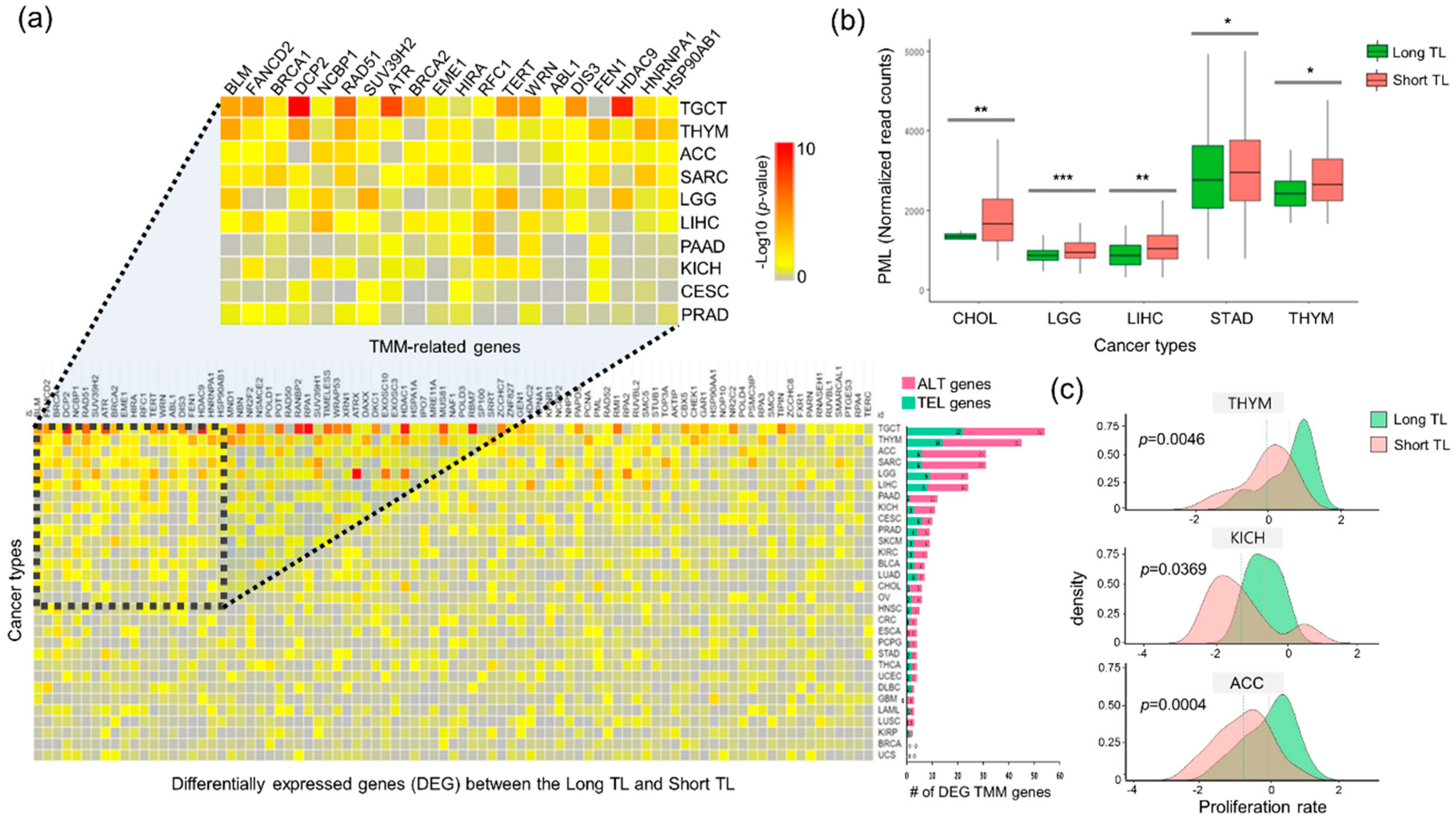
2.2. Differential Gene Expression of the TMMs Reveals Different Cancer Hallmarks
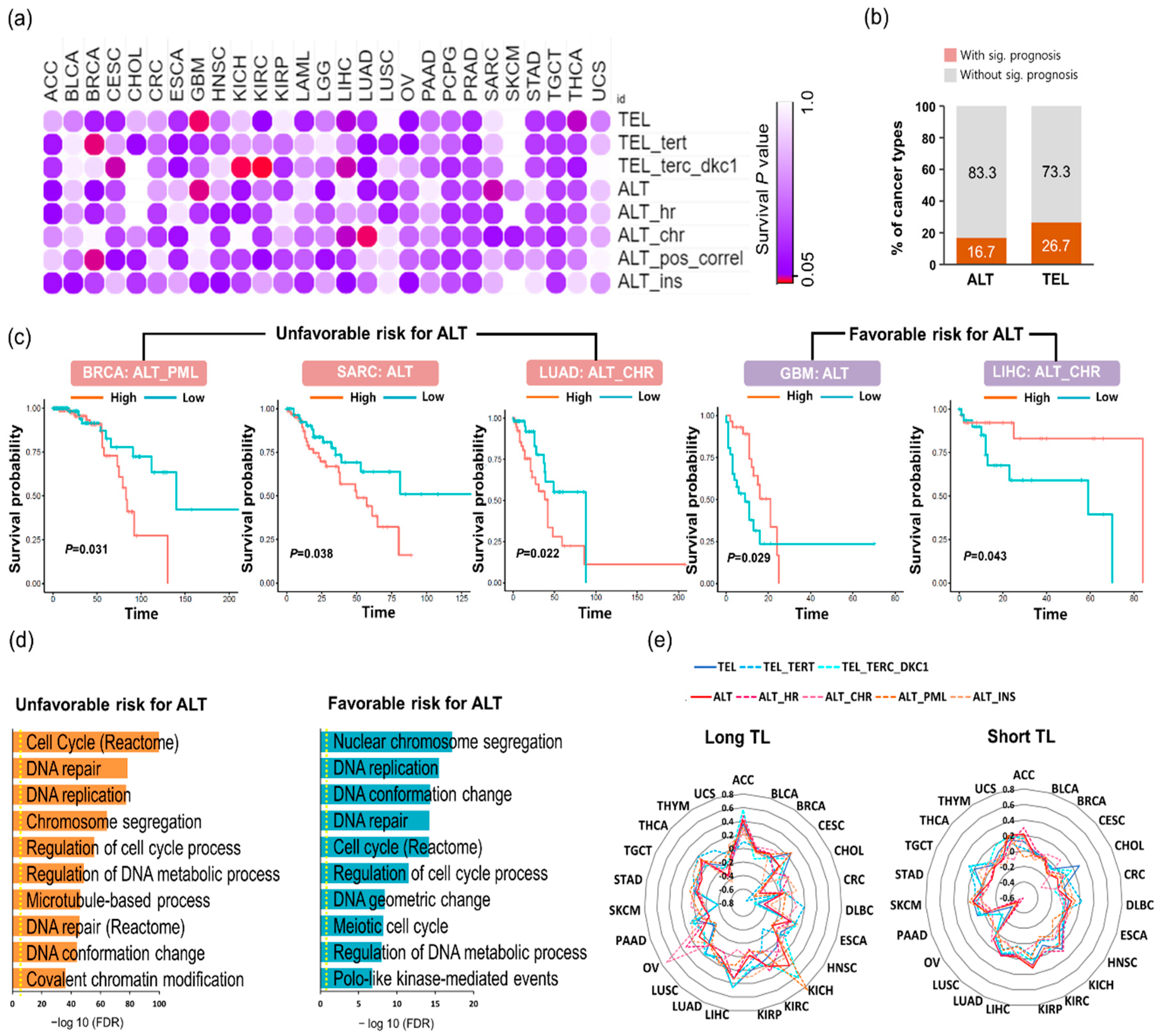
2.3. Distinct TMM Pathways are Correlated with Prognosis in A Cancer Type-Specific Manner
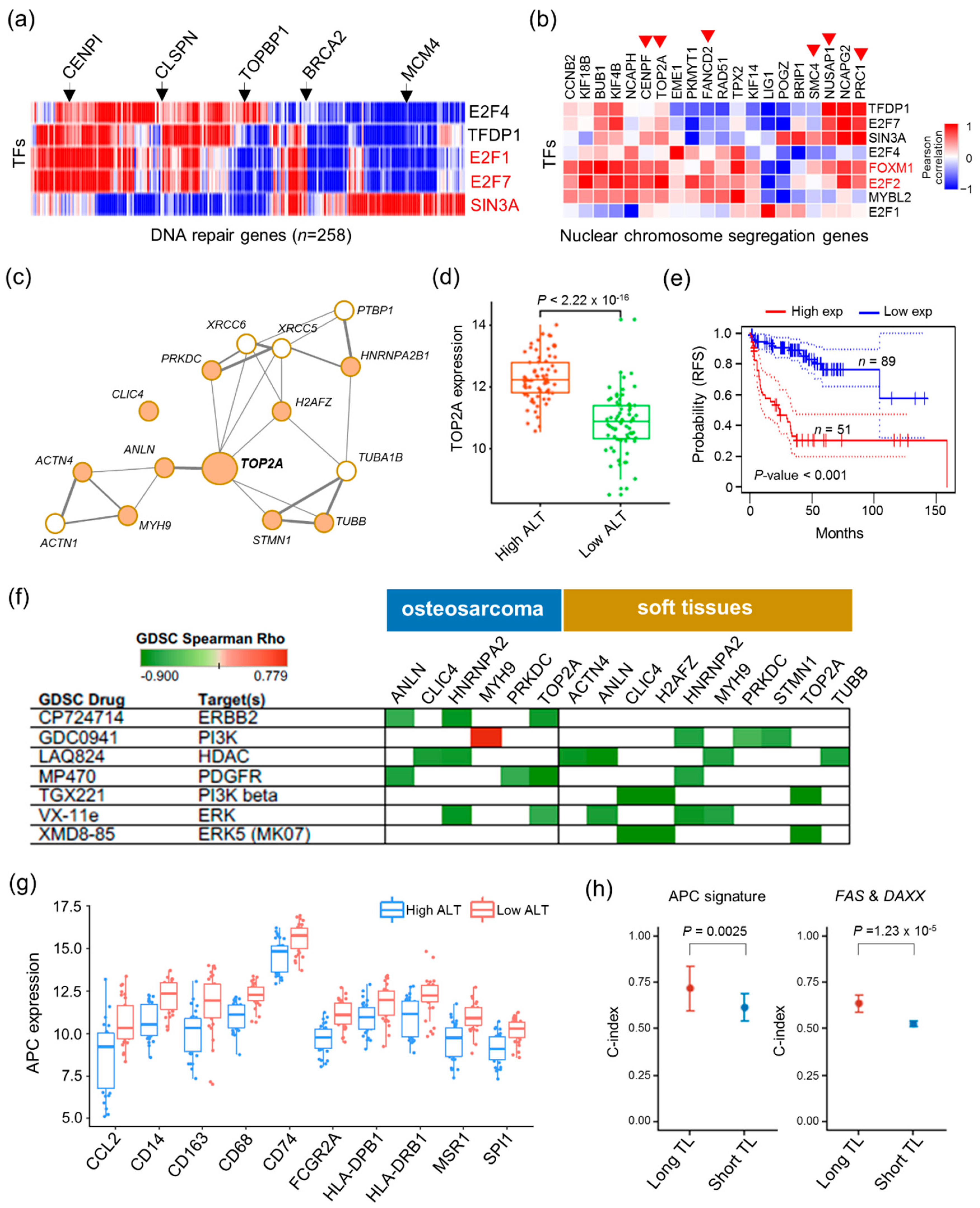
2.4. Vulnerabilities of ALT Activity for Cancer Therapy
3. Discussion
4. Materials and Methods
4.1. Data Sets
4.2. TMM Signature Analysis
4.3. Differential Expression Gene Analysis of Cancer Types
4.4. Gene Ontology and Correlation Analysis
4.5. Transcription Factor Analysis Protein Association Network
4.6. Survival Probability Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Greider, C.W. Telomerase and telomere-length regulation: Lessons from small eukaryotes to mammals. Cold Spring Harb. Symp. Quant. Biol. 1993, 58, 719–723. [Google Scholar] [CrossRef] [PubMed]
- Denchi, E.L.; de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 2007, 448, 1068–1071. [Google Scholar] [CrossRef] [PubMed]
- Hayflick, L. The Limited in Vitro Lifetime of Human Diploid Cell Strains. Exp. Cell Res. 1965, 37, 614–636. [Google Scholar] [CrossRef]
- Counter, C.M.; Avilion, A.A.; LeFeuvre, C.E.; Stewart, N.G.; Greider, C.W.; Harley, C.B.; Bacchetti, S. Telomere shortening associated with chromosome instability is arrested in immortal cells which express telomerase activity. EMBO J. 1992, 11, 1921–1929. [Google Scholar] [CrossRef]
- Gaspar, T.B.; Sa, A.; Lopes, J.M.; Sobrinho-Simoes, M.; Soares, P.; Vinagre, J. Telomere Maintenance Mechanisms in Cancer. Genes 2018, 9, 241. [Google Scholar] [CrossRef]
- Dilley, R.L.; Greenberg, R.A. ALTernative Telomere Maintenance and Cancer. Trends Cancer 2015, 1, 145–156. [Google Scholar] [CrossRef]
- Reddel, R.R.; Bryan, T.M.; Murnane, J.P. Immortalized cells with no detectable telomerase activity. A review. Biochemistry 1997, 62, 1254–1262. [Google Scholar]
- Heaphy, C.M.; Subhawong, A.P.; Hong, S.M.; Goggins, M.G.; Montgomery, E.A.; Gabrielson, E.; Netto, G.J.; Epstein, J.I.; Lotan, T.L.; Westra, W.H.; et al. Prevalence of the alternative lengthening of telomeres telomere maintenance mechanism in human cancer subtypes. Am. J. Pathol. 2011, 179, 1608–1615. [Google Scholar] [CrossRef]
- Dunham, M.A.; Neumann, A.A.; Fasching, C.L.; Reddel, R.R. Telomere maintenance by recombination in human cells. Nat. Genet. 2000, 26, 447–450. [Google Scholar] [CrossRef]
- Pickett, H.A.; Cesare, A.J.; Johnston, R.L.; Neumann, A.A.; Reddel, R.R. Control of telomere length by a trimming mechanism that involves generation of t-circles. EMBO J. 2009, 28, 799–809. [Google Scholar] [CrossRef]
- Bryan, T.M.; Englezou, A.; Gupta, J.; Bacchetti, S.; Reddel, R.R. Telomere elongation in immortal human cells without detectable telomerase activity. EMBO J. 1995, 14, 4240–4248. [Google Scholar] [CrossRef] [PubMed]
- Nabetani, A.; Ishikawa, F. Unusual telomeric DNAs in human telomerase-negative immortalized cells. Mol. Cell. Biol. 2009, 29, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Bechter, O.E.; Zou, Y.; Walker, W.; Wright, W.E.; Shay, J.W. Telomeric recombination in mismatch repair deficient human colon cancer cells after telomerase inhibition. Cancer Res. 2004, 64, 3444–3451. [Google Scholar] [CrossRef] [PubMed]
- Yeager, T.R.; Neumann, A.A.; Englezou, A.; Huschtscha, L.I.; Noble, J.R.; Reddel, R.R. Telomerase-negative immortalized human cells contain a novel type of promyelocytic leukemia (PML) body. Cancer Res. 1999, 59, 4175–4179. [Google Scholar] [PubMed]
- Lovejoy, C.A.; Li, W.; Reisenweber, S.; Thongthip, S.; Bruno, J.; de Lange, T.; De, S.; Petrini, J.H.; Sung, P.A.; Jasin, M.; et al. Loss of ATRX, genome instability, and an altered DNA damage response are hallmarks of the alternative lengthening of telomeres pathway. PLoS Genet. 2012, 8, e1002772. [Google Scholar] [CrossRef]
- Schwartzentruber, J.; Korshunov, A.; Liu, X.Y.; Jones, D.T.; Pfaff, E.; Jacob, K.; Sturm, D.; Fontebasso, A.M.; Quang, D.A.; Tonjes, M.; et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 2012, 482, 226–231. [Google Scholar] [CrossRef]
- Lawlor, R.T.; Veronese, N.; Pea, A.; Nottegar, A.; Smith, L.; Pilati, C.; Demurtas, J.; Fassan, M.; Cheng, L.; Luchini, C. Alternative lengthening of telomeres (ALT) influences survival in soft tissue sarcomas: A systematic review with meta-analysis. BMC Cancer 2019, 19, 232. [Google Scholar] [CrossRef]
- Hu, J.; Hwang, S.S.; Liesa, M.; Gan, B.; Sahin, E.; Jaskelioff, M.; Ding, Z.; Ying, H.; Boutin, A.T.; Zhang, H.; et al. Antitelomerase therapy provokes ALT and mitochondrial adaptive mechanisms in cancer. Cell 2012, 148, 651–663. [Google Scholar] [CrossRef]
- Nersisyan, L.; Hopp, L.; Loeffler-Wirth, H.; Galle, J.; Loeffler, M.; Arakelyan, A.; Binder, H. Telomere Length Maintenance and Its Transcriptional Regulation in Lynch Syndrome and Sporadic Colorectal Carcinoma. Front. Oncol. 2019, 9, 1172. [Google Scholar] [CrossRef]
- Hanzelmann, S.; Castelo, R.; Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013, 14, 7. [Google Scholar] [CrossRef]
- Barthel, F.P.; Wei, W.; Tang, M.; Martinez-Ledesma, E.; Hu, X.; Amin, S.B.; Akdemir, K.C.; Seth, S.; Song, X.; Wang, Q.; et al. Systematic analysis of telomere length and somatic alterations in 31 cancer types. Nat. Genet. 2017, 49, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Cerone, M.A.; Londono-Vallejo, J.A.; Bacchetti, S. Telomere maintenance by telomerase and by recombination can coexist in human cells. Hum. Mol. Genet. 2001, 10, 1945–1952. [Google Scholar] [CrossRef] [PubMed]
- Ford, L.P.; Zou, Y.; Pongracz, K.; Gryaznov, S.M.; Shay, J.W.; Wright, W.E. Telomerase can inhibit the recombination-based pathway of telomere maintenance in human cells. J. Biol. Chem. 2001, 276, 32198–32203. [Google Scholar] [CrossRef] [PubMed]
- Grobelny, J.V.; Kulp-McEliece, M.; Broccoli, D. Effects of reconstitution of telomerase activity on telomere maintenance by the alternative lengthening of telomeres (ALT) pathway. Hum. Mol. Genet. 2001, 10, 1953–1961. [Google Scholar] [CrossRef]
- Perrem, K.; Colgin, L.M.; Neumann, A.A.; Yeager, T.R.; Reddel, R.R. Coexistence of alternative lengthening of telomeres and telomerase in hTERT-transfected GM847 cells. Mol. Cell. Biol. 2001, 21, 3862–3875. [Google Scholar] [CrossRef]
- Stewart, S.A.; Hahn, W.C.; O’Connor, B.F.; Banner, E.N.; Lundberg, A.S.; Modha, P.; Mizuno, H.; Brooks, M.W.; Fleming, M.; Zimonjic, D.B.; et al. Telomerase contributes to tumorigenesis by a telomere length-independent mechanism. Proc. Natl. Acad. Sci. USA 2002, 99, 12606–12611. [Google Scholar] [CrossRef]
- Hoang, S.M.; O’Sullivan, R.J. Alternative Lengthening of Telomeres: Building Bridges to Connect Chromosome Ends. Trends Cancer 2020, 6, 247–260. [Google Scholar] [CrossRef]
- Root, H.; Larsen, A.; Komosa, M.; Al-Azri, F.; Li, R.; Bazett-Jones, D.P.; Stephen Meyn, M. FANCD2 limits BLM-dependent telomere instability in the alternative lengthening of telomeres pathway. Hum. Mol. Genet. 2016, 25, 3255–3268. [Google Scholar] [CrossRef]
- Liberzon, A.; Birger, C.; Thorvaldsdottir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef]
- Aviv, A.; Anderson, J.J.; Shay, J.W. Mutations, Cancer and the Telomere Length Paradox. Trends Cancer 2017, 3, 253–258. [Google Scholar] [CrossRef]
- Guan, D.; Kao, H.Y. The function, regulation and therapeutic implications of the tumor suppressor protein, PML. Cell Biosci. 2015, 5, 60. [Google Scholar] [CrossRef] [PubMed]
- Marchesini, M.; Matocci, R.; Tasselli, L.; Cambiaghi, V.; Orleth, A.; Furia, L.; Marinelli, C.; Lombardi, S.; Sammarelli, G.; Aversa, F.; et al. PML is required for telomere stability in non-neoplastic human cells. Oncogene 2016, 35, 1811–1821. [Google Scholar] [CrossRef] [PubMed]
- Gong, P.; Wang, H.; Zhang, J.; Fu, Y.; Zhu, Z.; Wang, J.; Yin, Y.; Wang, H.; Zhou, Z.; Yang, J.; et al. Telomere Maintenance-Associated PML Is a Potential Specific Therapeutic Target of Human Colorectal Cancer. Transl. Oncol. 2019, 12, 1164–1176. [Google Scholar] [CrossRef] [PubMed]
- Holt, S.E.; Shay, J.W. Role of telomerase in cellular proliferation and cancer. J. Cell. Physiol. 1999, 180, 10–18. [Google Scholar] [CrossRef]
- Belair, C.D.; Yeager, T.R.; Lopez, P.M.; Reznikoff, C.A. Telomerase activity: A biomarker of cell proliferation, not malignant transformation. Proc. Natl. Acad. Sci. USA 1997, 94, 13677–13682. [Google Scholar] [CrossRef]
- McDonald, K.L.; McDonnell, J.; Muntoni, A.; Henson, J.D.; Hegi, M.E.; von Deimling, A.; Wheeler, H.R.; Cook, R.J.; Biggs, M.T.; Little, N.S.; et al. Presence of alternative lengthening of telomeres mechanism in patients with glioblastoma identifies a less aggressive tumor type with longer survival. J. Neuropathol. Exp. Neurol. 2010, 69, 729–736. [Google Scholar] [CrossRef]
- Hakin-Smith, V.; Jellinek, D.A.; Levy, D.; Carroll, T.; Teo, M.; Timperley, W.R.; McKay, M.J.; Reddel, R.R.; Royds, J.A. Alternative lengthening of telomeres and survival in patients with glioblastoma multiforme. Lancet 2003, 361, 836–838. [Google Scholar] [CrossRef]
- Clark, G.M.; Osborne, C.K.; Levitt, D.; Wu, F.; Kim, N.W. Telomerase activity and survival of patients with node-positive breast cancer. J. Natl. Cancer Inst. 1997, 89, 1874–1881. [Google Scholar] [CrossRef]
- Poremba, C.; Heine, B.; Diallo, R.; Heinecke, A.; Wai, D.; Schaefer, K.L.; Braun, Y.; Schuck, A.; Lanvers, C.; Bankfalvi, A.; et al. Telomerase as a prognostic marker in breast cancer: High-throughput tissue microarray analysis of hTERT and hTR. J. Pathol. 2002, 198, 181–189. [Google Scholar] [CrossRef]
- Sakurai, S.; Fukayama, M.; Kaizaki, Y.; Saito, K.; Kanazawa, K.; Kitamura, M.; Iwasaki, Y.; Hishima, T.; Hayashi, Y.; Koike, M. Telomerase activity in gastrointestinal stromal tumors. Cancer 1998, 83, 2060–2066. [Google Scholar] [CrossRef]
- Bertorelle, R.; Briarava, M.; Rampazzo, E.; Biasini, L.; Agostini, M.; Maretto, I.; Lonardi, S.; Friso, M.L.; Mescoli, C.; Zagonel, V.; et al. Telomerase is an independent prognostic marker of overall survival in patients with colorectal cancer. Br. J. Cancer 2013, 108, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Arita, H.; Narita, Y.; Fukushima, S.; Tateishi, K.; Matsushita, Y.; Yoshida, A.; Miyakita, Y.; Ohno, M.; Collins, V.P.; Kawahara, N.; et al. Upregulating mutations in the TERT promoter commonly occur in adult malignant gliomas and are strongly associated with total 1p19q loss. Acta Neuropathol. 2013, 126, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Pekmezci, M.; Rice, T.; Molinaro, A.M.; Walsh, K.M.; Decker, P.A.; Hansen, H.; Sicotte, H.; Kollmeyer, T.M.; McCoy, L.S.; Sarkar, G.; et al. Adult infiltrating gliomas with WHO 2016 integrated diagnosis: Additional prognostic roles of ATRX and TERT. Acta Neuropathol. 2017, 133, 1001–1016. [Google Scholar] [CrossRef] [PubMed]
- Jeong, D.E.; Woo, S.R.; Nam, H.; Nam, D.H.; Lee, J.H.; Joo, K.M. Preclinical and clinical implications of TERT promoter mutation in glioblastoma multiforme. Oncol. Lett. 2017, 14, 8213–8219. [Google Scholar] [CrossRef] [PubMed]
- Flynn, R.L.; Cox, K.E.; Jeitany, M.; Wakimoto, H.; Bryll, A.R.; Ganem, N.J.; Bersani, F.; Pineda, J.R.; Suva, M.L.; Benes, C.H.; et al. Alternative lengthening of telomeres renders cancer cells hypersensitive to ATR inhibitors. Science 2015, 347, 273–277. [Google Scholar] [CrossRef]
- Deeg, K.I.; Chung, I.; Bauer, C.; Rippe, K. Cancer Cells with Alternative Lengthening of Telomeres Do Not Display a General Hypersensitivity to ATR Inhibition. Front. Oncol. 2016, 6, 186. [Google Scholar] [CrossRef]
- Qin, Y.; Conley, A.P.; Grimm, E.A.; Roszik, J. A tool for discovering drug sensitivity and gene expression associations in cancer cells. PLoS ONE 2017, 12, e0176763. [Google Scholar] [CrossRef]
- Hung, N.A.; Eiholzer, R.A.; Kirs, S.; Zhou, J.; Ward-Hartstonge, K.; Wiles, A.K.; Frampton, C.M.; Taha, A.; Royds, J.A.; Slatter, T.L. Telomere profiles and tumor-associated macrophages with different immune signatures affect prognosis in glioblastoma. Mod. Pathol. 2016, 29, 212–226. [Google Scholar] [CrossRef]
- De Bonis, P.; Albanese, A.; Lofrese, G.; de Waure, C.; Mangiola, A.; Pettorini, B.L.; Pompucci, A.; Balducci, M.; Fiorentino, A.; Lauriola, L.; et al. Postoperative infection may influence survival in patients with glioblastoma: Simply a myth? Neurosurgery 2011, 69, 864–868. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Crowe, D.L.; Nguyen, D.C.; Tsang, K.J.; Kyo, S. E2F-1 represses transcription of the human telomerase reverse transcriptase gene. Nucleic Acids Res. 2001, 29, 2789–2794. [Google Scholar] [CrossRef] [PubMed]
- Udayakumar, T.; Shareef, M.M.; Diaz, D.A.; Ahmed, M.M.; Pollack, A. The E2F1/Rb and p53/MDM2 pathways in DNA repair and apoptosis: Understanding the crosstalk to develop novel strategies for prostate cancer radiotherapy. Semin. Radiat. Oncol. 2010, 20, 258–266. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.Z.; Tsai, S.Y.; Leone, G. Emerging roles of E2Fs in cancer: An exit from cell cycle control. Nat. Rev. Cancer 2009, 9, 785–797. [Google Scholar] [CrossRef] [PubMed]
- Crosby, M.E.; Almasan, A. Opposing roles of E2Fs in cell proliferation and death. Cancer Biol. Ther. 2004, 3, 1208–1211. [Google Scholar] [CrossRef]
- Vernier, M.; Bourdeau, V.; Gaumont-Leclerc, M.F.; Moiseeva, O.; Begin, V.; Saad, F.; Mes-Masson, A.M.; Ferbeyre, G. Regulation of E2Fs and senescence by PML nuclear bodies. Genes Dev. 2011, 25, 41–50. [Google Scholar] [CrossRef]
- Ding, Z.; Mangino, M.; Aviv, A.; Spector, T.; Durbin, R.; Consortium, U.K. Estimating telomere length from whole genome sequence data. Nucleic Acids Res. 2014, 42, e75. [Google Scholar] [CrossRef]
- Liu, J.; Lichtenberg, T.; Hoadley, K.A.; Poisson, L.M.; Lazar, A.J.; Cherniack, A.D.; Kovatich, A.J.; Benz, C.C.; Levine, D.A.; Lee, A.V.; et al. An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell 2018, 173, 400–416.e11. [Google Scholar] [CrossRef]
- Thorsson, V.; Gibbs, D.L.; Brown, S.D.; Wolf, D.; Bortone, D.S.; Ou Yang, T.H.; Porta-Pardo, E.; Gao, G.F.; Plaisier, C.L.; Eddy, J.A.; et al. The Immune Landscape of Cancer. Immunity 2018, 48, 812–830.e14. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1523. [Google Scholar] [CrossRef]
- Janky, R.; Verfaillie, A.; Imrichova, H.; Van de Sande, B.; Standaert, L.; Christiaens, V.; Hulselmans, G.; Herten, K.; Naval Sanchez, M.; Potier, D.; et al. iRegulon: From a gene list to a gene regulatory network using large motif and track collections. PLoS Comput. Biol. 2014, 10, e1003731. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Durisova, M.; Dedik, L. SURVIVAL—An integrated software package for survival curve estimation and statistical comparison of survival rates of two groups of patients or experimental animals. Methods Find Exp. Clin. Pharmacol. 1993, 15, 535–540. [Google Scholar] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sung, J.-Y.; Lim, H.-W.; Joung, J.-G.; Park, W.-Y. Pan-Cancer Analysis of Alternative Lengthening of Telomere Activity. Cancers 2020, 12, 2207. https://doi.org/10.3390/cancers12082207
Sung J-Y, Lim H-W, Joung J-G, Park W-Y. Pan-Cancer Analysis of Alternative Lengthening of Telomere Activity. Cancers. 2020; 12(8):2207. https://doi.org/10.3390/cancers12082207
Chicago/Turabian StyleSung, Ji-Yong, Hee-Woong Lim, Je-Gun Joung, and Woong-Yang Park. 2020. "Pan-Cancer Analysis of Alternative Lengthening of Telomere Activity" Cancers 12, no. 8: 2207. https://doi.org/10.3390/cancers12082207
APA StyleSung, J.-Y., Lim, H.-W., Joung, J.-G., & Park, W.-Y. (2020). Pan-Cancer Analysis of Alternative Lengthening of Telomere Activity. Cancers, 12(8), 2207. https://doi.org/10.3390/cancers12082207





