Methylation Patterns and Chromatin Accessibility in Neuroendocrine Lung Cancer
Abstract
1. Introduction
2. Results
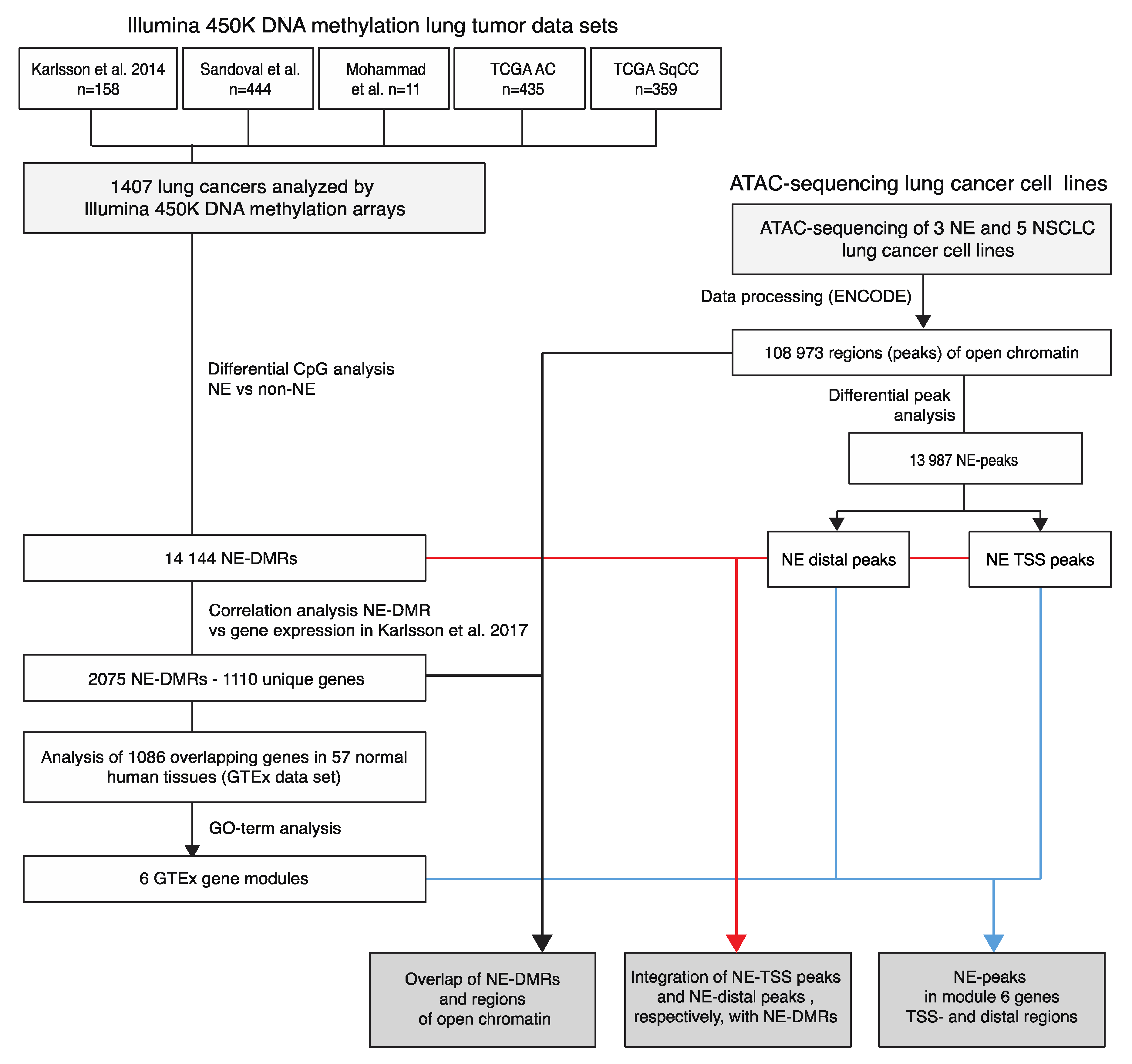
2.1. Differentially Methylated Regions in Neuroendocrine Lung Cancer Correlated with Gene Expression
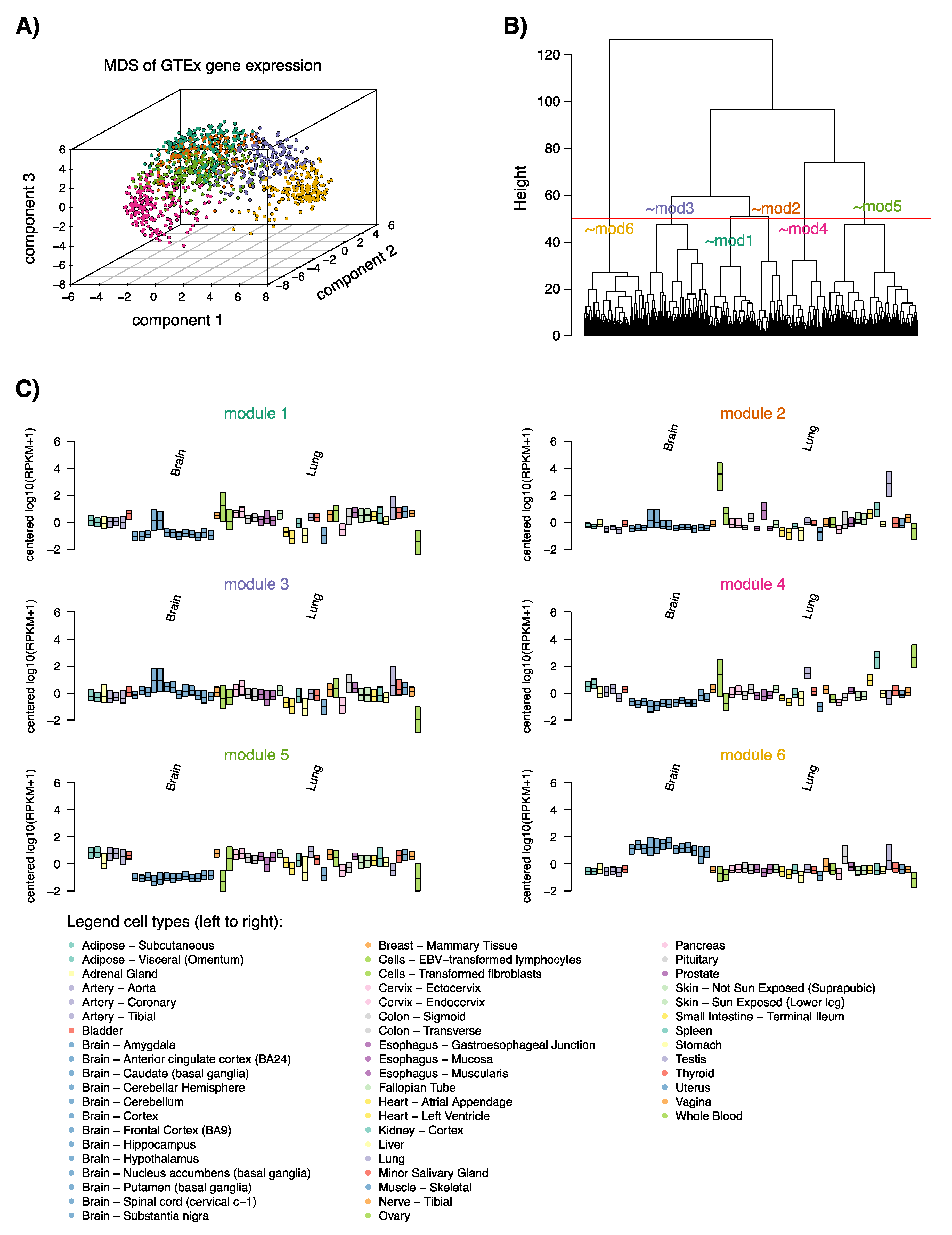
2.2. NE-DMR Genes Expression in Normal Tissues Defines Six Transcriptional Programs
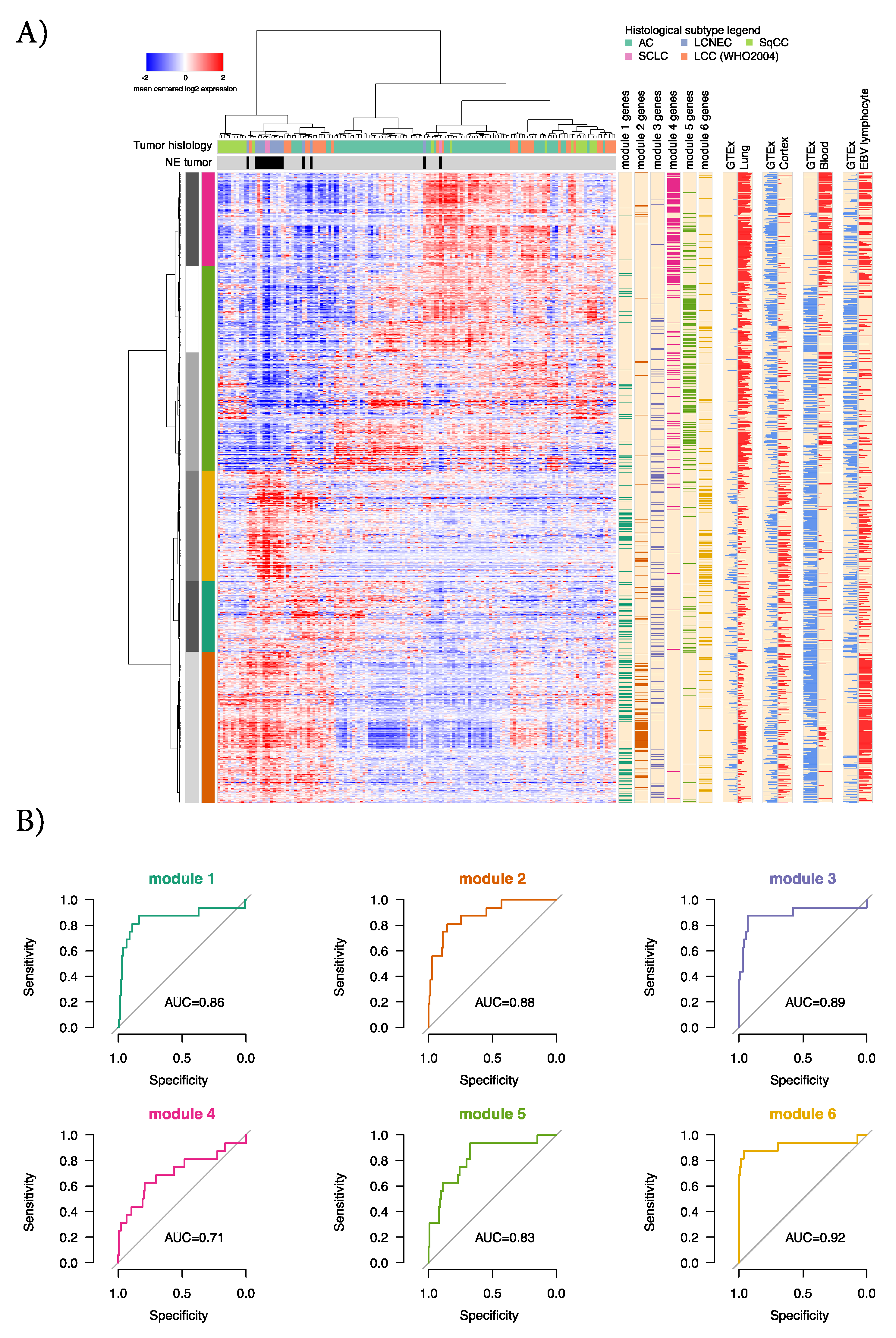
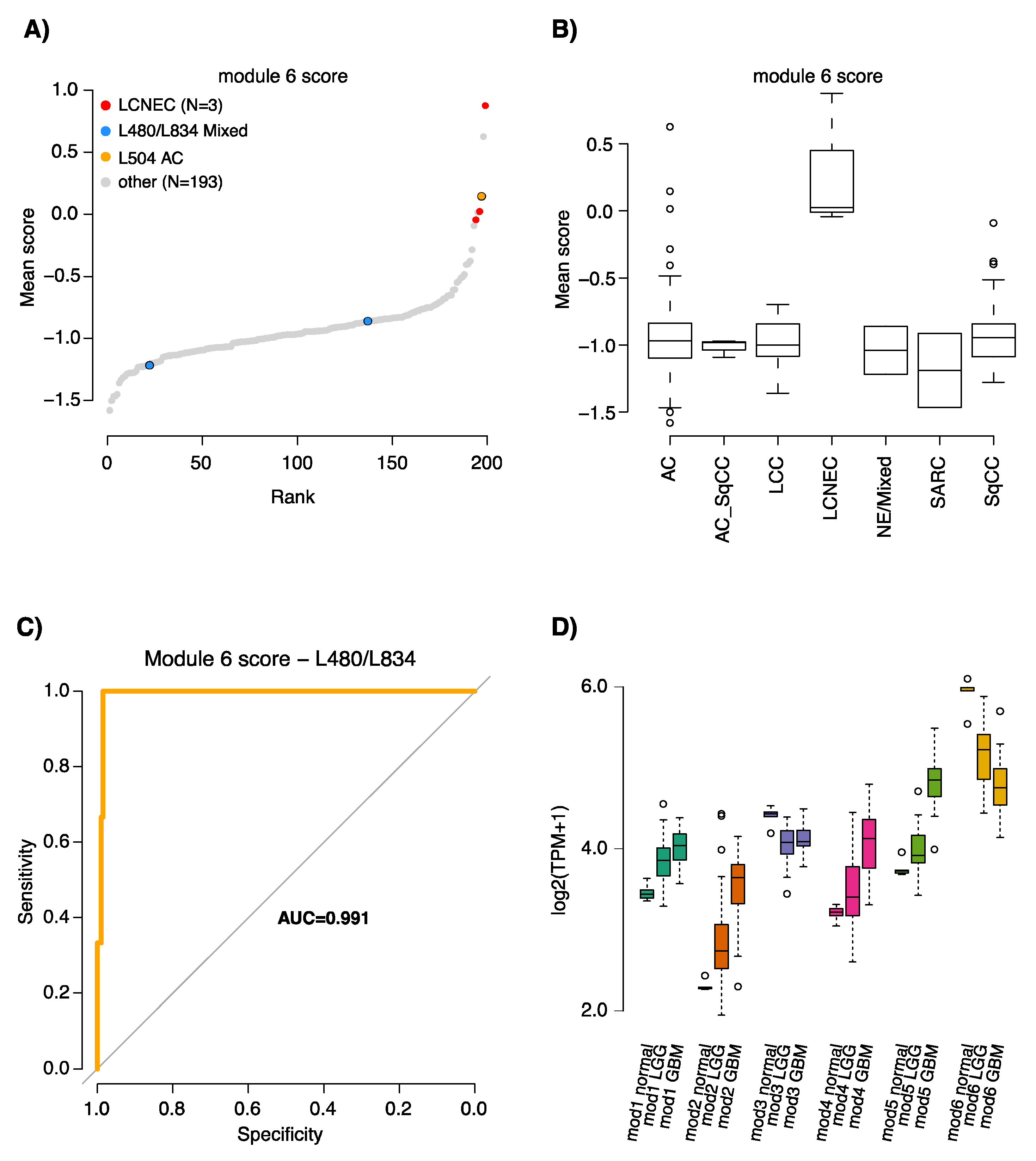
2.3. Characterization of the Six GTEx Modules in Lung Cancer Gene Expression Data
2.4. Assay for Transposase-Accessible Chromatin Sequencing (ATAC-Seq) Identifies Regions of Open Chromatin Associated with NE Histology
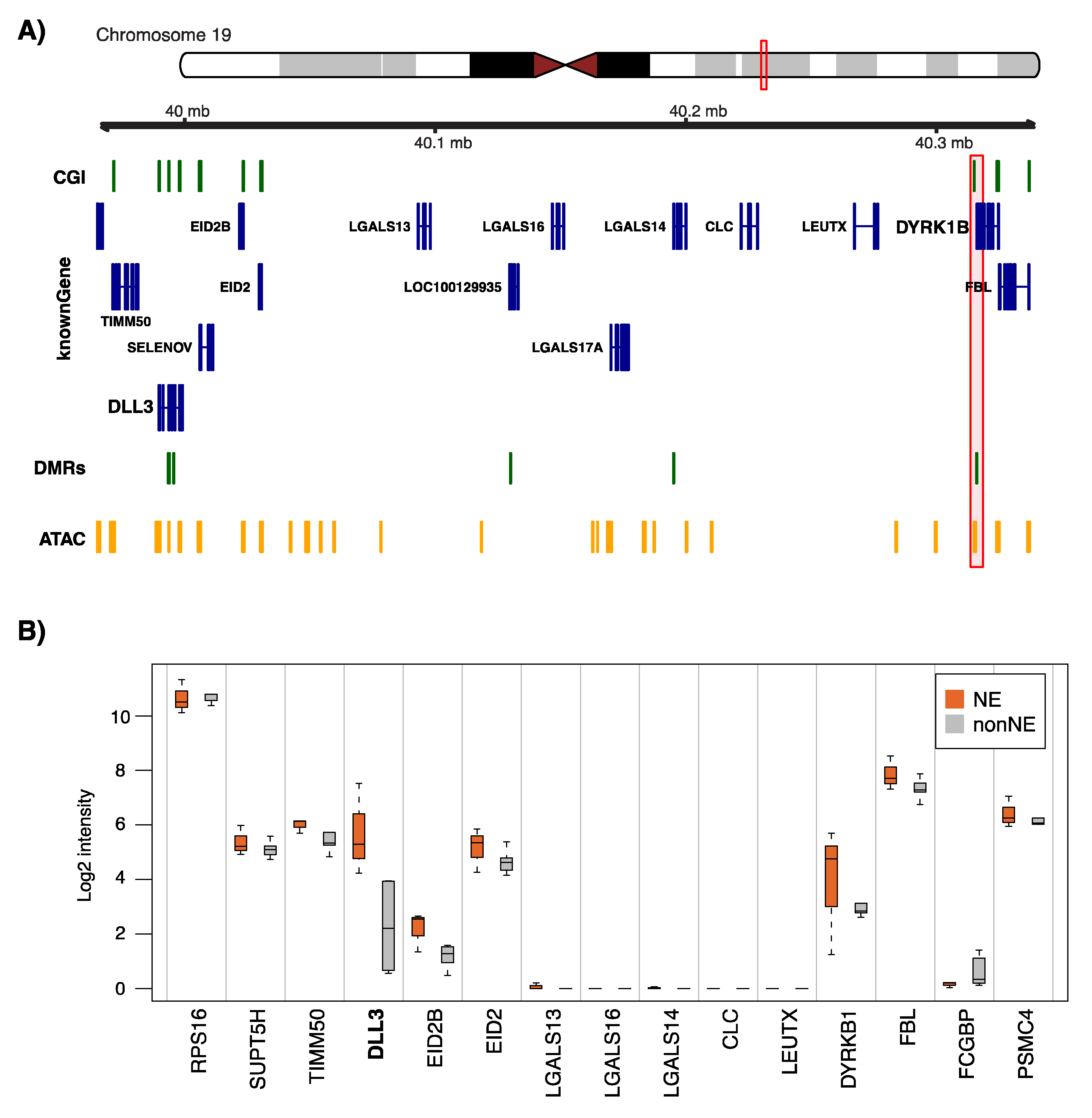
2.5. ATAC-Seq Supports Epigenetic Regulation of NE-DMR Genes
2.6. Assessing the Regulatory Potential of Tumor Derived NE-DMRs in ATAC-Seq Data
3. Discussion
4. Materials and Methods
4.1. Patient Cohorts
4.2. DNA Methylation Analyses
4.3. Gene Expression Analyses
4.4. ATAC- and RNA-Seq of NSCLC and NE Cell Lines
4.5. Data Sharing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Scagliotti, G.V.; Brodowicz, T.; Shepherd, F.A.; Zielinski, C.; Vansteenkiste, J.; Manegold, C.; Simms, L.; Fossella, F.V.; Sugarman, K.; Belani, C.P. Treatment-by-histology interaction analyses in three phase III trials show superiority of pemetrexed in nonsquamous non-small cell lung cancer. J. Thorac. Oncol. 2011, 6, 64–70. [Google Scholar] [CrossRef]
- Scagliotti, G.V.; Parikh, P.M.; von Pawel, J.; Biesma, B.; Vansteenkiste, J.; Manegold, C.; Serwatowski, P.; Gatzemeier, U.; Digumarti, R.; Zukin, M.; et al. Phase III study comparing cisplatin plus gemcitabine with cisplatin plus pemetrexed in chemotherapy-naive patients with advanced-stage non-small-cell lung cancer. J. Clin. Oncol. 2008, 26, 3543–3551. [Google Scholar] [CrossRef] [PubMed]
- Swanton, C.; Govindan, R. Clinical Implications of Genomic Discoveries in Lung Cancer. N. Engl. J. Med. 2016, 374, 1864–1873. [Google Scholar] [CrossRef] [PubMed]
- George, J.; Lim, J.S.; Jang, S.J.; Cun, Y.; Ozretic, L.; Kong, G.; Leenders, F.; Lu, X.; Fernandezcuesta, L.; Bosco, G.; et al. Comprehensive genomic profiles of small cell lung cancer. Nature 2015, 524, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Collisson, E.A.; Campbell, J.D.; Brooks, A.N.; Berger, A.H.; Lee, W.R.; Chmielecki, J.; Beer, D.G.; Cope, L.; Creighton, C.J.; Danilova, L.V.; et al. The Cancer Genome Atlas Network A: Comprehensive molecular profiling of lung adenocarcinoma. Nature 2014, 511, 543–550. [Google Scholar]
- Bhattacharjee, A.; Richards, W.G.; Staunton, J.; Li, C.; Monti, S.; Vasa, P.; Ladd, C.; Beheshti, J.; Bueno, R.; Gillette, M.A.; et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc. Natl. Acad. Sci. USA 2001, 98, 13790–13795. [Google Scholar] [CrossRef] [PubMed]
- Hammerman, P.S.; Voet, D.; Lawrence, M.S.; Voet, D.; Jing, R.; Cibulskis, K.; Sivachenko, A.; Stojanov, P.; Mckenna, A.; Lander, E.S.; et al. Comprehensive genomic characterization of squamous cell lung cancers. Nature 2012, 489, 519–525. [Google Scholar]
- Karlsson, A.; Brunnstrom, H.; Micke, P.; Veerla, S.; Mattsson, J.S.M.; la Fleur, L.; Botling, J.; Jonsson, M.; Reutersward, C.; Planck, M.; et al. Gene Expression Profiling of Large Cell Lung Cancer Links Transcriptional Phenotypes to the New Histological WHO 2015 Classification. J. Thorac. Oncol. 2017, 12, 1257–1267. [Google Scholar] [CrossRef]
- Travis, W.D.; Burke, A.P.; Marx, A.; Nicholson, A.G. (Eds.) WHO Classification of Tumours of the Lung, Pleura, Thymus and Heart, 4th ed.; IARC Press: Lyon, France, 2015. [Google Scholar]
- Rekhtman, N.; Pietanza, M.C.; Hellmann, M.D.; Naidoo, J.; Arora, A.; Won, H.H.; Halpenny, D.; Wang, H.; Tian, S.K.; Litvak, A.; et al. Next-Generation Sequencing of Pulmonary Large Cell Neuroendocrine Carcinoma Reveals Small Cell Carcinoma-like and Non-Small Cell Carcinoma-like Subsets. Clin. Cancer Res. 2016, 22, 3618–3629. [Google Scholar] [CrossRef]
- George, J.; Walter, V.; Peifer, M.; Alexandrov, L.B.; Seidel, D.; Leenders, F.; Maas, L.; Muller, C.; Dahmen, I.; Delhomme, T.M.; et al. Integrative genomic profiling of large-cell neuroendocrine carcinomas reveals distinct subtypes of high-grade neuroendocrine lung tumors. Nat. Commun. 2018, 9, 1048. [Google Scholar] [CrossRef]
- Rossi, G.; Cavazza, A.; Marchioni, A.; Longo, L.; Migaldi, M.; Sartori, G.; Bigiani, N.; Schirosi, L.; Casali, C.; Morandi, U.; et al. Role of chemotherapy and the receptor tyrosine kinases KIT, PDGFRalpha, PDGFRbeta, and Met in large-cell neuroendocrine carcinoma of the lung. J. Clin. Oncol. 2005, 23, 8774–8785. [Google Scholar] [CrossRef]
- Semenova, E.A.; Nagel, R.; Berns, A. Origins, genetic landscape, and emerging therapies of small cell lung cancer. Genes Dev. 2015, 29, 1447–1462. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, A.K.; Jonsson, M.; Lauss, M.; Brunnstrom, H.; Jonsson, P.; Borg, A.; Jonsson, G.; Ringner, M.; Planck, M.; Staaf, J. Genome-wide DNA methylation analysis of lung carcinoma reveals one neuroendocrine and four adenocarcinoma epitypes associated with patient outcome. Clin. Cancer Res. 2014, 20, 6127–6140. [Google Scholar] [CrossRef] [PubMed]
- Sandoval, J.; Mendezgonzalez, J.; Nadal, E.; Chen, G.; Carmona, F.J.; Sayols, S.; Moran, S.; Heyn, H.; Vizoso, M.; Gomez, A.; et al. A Prognostic DNA Methylation Signature for Stage I Non-Small-Cell Lung Cancer. J. Clin. Oncol. 2013, 31, 4140–4147. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Zhang, Y.; Parra, E.R.; Rodriguez, J.; Behrens, C.; Akbani, R.; Lu, Y.S.; Kurie, J.M.; Gibbons, D.L.; Mills, G.B.; et al. Multiplatform-based molecular subtypes of non-small-cell lung cancer. Oncogene 2017, 36, 1384–1393. [Google Scholar] [CrossRef] [PubMed]
- Duruisseaux, M.; Esteller, M. Lung cancer epigenetics: From knowledge to applications. Semin. Cancer Biol. 2017, 51, 116–128. [Google Scholar] [CrossRef]
- Corces, M.R.; Granja, J.M.; Shams, S.; Louie, B.H.; Seoane, J.A.; Zhou, W.; Silva, T.C.; Groeneveld, C.S.; Wong, C.K.; Cho, S.W.; et al. The chromatin accessibility landscape of primary human cancers. Science 2018, 362. [Google Scholar] [CrossRef] [PubMed]
- Djureinovic, D.; Hallstrom, B.M.; Horie, M.; Mattsson, J.S.M.; la Fleur, L.; Fagerberg, L.; Brunnstrom, H.; Lindskog, C.; Madjar, K.; Rahnenfuhrer, J.; et al. Profiling cancer testis antigens in non-small-cell lung cancer. JCI Insight 2016, 1, e86837. [Google Scholar] [CrossRef]
- Karlsson, A.; Cirenajwis, H.; Ericsonlindquist, K.; Brunnstrom, H.; Reutersward, C.; Jonsson, M.; Ortizvillalon, C.; Hussein, A.; Bergman, B.; Vikstrom, A.; et al. A combined gene expression tool for parallel histological prediction and gene fusion detection in non-small cell lung cancer. Sci. Rep. 2019, 9, 5207. [Google Scholar] [CrossRef] [PubMed]
- Hermans, B.C.M.; Derks, J.L.; Thunnissen, E.; van Suylen, R.J.; den Bakker, M.A.; Groen, H.J.M.; Smit, E.F.; Damhuis, R.A.; van den Broek, E.C.; Ruland, A.; et al. DLL3 expression in large cell neuroendocrine carcinoma (LCNEC) and association with molecular subtypes and neuroendocrine profile. Lung Cancer 2019, 138, 102–108. [Google Scholar] [CrossRef]
- Staaf, J.; Isaksson, S.; Karlsson, A.; Jonsson, M.; Johansson, L.I.; Jonsson, P.; Botling, J.; Micke, P.; Baldetorp, B.; Planck, M. Landscape of somatic allelic imbalances and copy number alterations in human lung carcinoma. Int. J. Cancer 2012, 1, 2020–2031. [Google Scholar] [CrossRef] [PubMed]
- Henke, R.M.; Meredith, D.; Borromeo, M.D.; Savage, T.K.; Johnson, J.E. Ascl1 and Neurog2 form novel complexes and regulate Delta-like3 (Dll3) expression in the neural tube. Dev. Biol. 2009, 328, 529–540. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Collins, B.J.; Jin, N.; Watkins, D.N.; Brock, M.V.; Matsui, W.; Nelkin, B.D.; Ball, D.W. Achaete-scute complex homologue 1 regulates tumor-initiating capacity in human small cell lung cancer. Cancer Res. 2009, 69, 845–854. [Google Scholar] [CrossRef] [PubMed]
- Kunnimalaiyaan, M.; Chen, H. Tumor suppressor role of Notch-1 signaling in neuroendocrine tumors. Oncologist 2007, 12, 535–542. [Google Scholar] [CrossRef]
- Mizuno, Y.; Maemura, K.; Tanaka, Y.; Hirata, A.; Futaki, S.; Hamamoto, H.; Taniguchi, K.; Hayashi, M.; Uchiyama, K.; Shibata, M.; et al. Expression of delta-like 3 is downregulated by aberrant DNA methylation and histone modification in hepatocellular carcinoma. Oncol. Rep. 2018, 39, 2209–2216. [Google Scholar] [CrossRef]
- Maemura, K.; Yoshikawa, H.; Yokoyama, K.; Ueno, T.; Kurose, H.; Uchiyama, K.; Otsuki, Y. Delta-like 3 is silenced by methylation and induces apoptosis in human hepatocellular carcinoma. Int. J. Oncol. 2013, 42, 817–822. [Google Scholar] [CrossRef]
- Staaf, O.; Tran, L.; Soderlund, L.; Nodin, B.; Jirstrom, K.; Vidarsdottir, H.; Planck, M.; Mattsson, J.S.M.; Botling, J.; Micke, P.; et al. Diagnostic Value of Insulinoma-Associated Protein 1 (INSM1) and Comparison With Established Neuroendocrine Markers in Pulmonary Cancers: A Comprehensive Study and Review of the Literature. Arch. Pathol. Lab. Med. 2020. [Google Scholar] [CrossRef]
- Kempfer, R.; Pombo, A. Methods for mapping 3D chromosome architecture. Nat. Rev. Genet. 2020, 21, 207–226. [Google Scholar] [CrossRef]
- Mohammad, H.P.; Smitheman, K.N.; Kamat, C.D.; Soong, D.; Federowicz, K.; van Aller, G.S.; Schneck, J.; Carson, J.D.; Liu, Y.; Butticello, M.; et al. A DNA Hypomethylation Signature Predicts Antitumor Activity of LSD1 Inhibitors in SCLC. Cancer Cell 2015, 28, 57–69. [Google Scholar] [CrossRef]
- Bibikova, M.; Barnes, B.; Tsan, C.; Ho, V.T.; Klotzle, B.; Le, J.M.; Delano, D.; Zhang, L.; Schroth, G.P.; Gunderson, K.L.; et al. High density DNA methylation array with single CpG site resolution. Genomics 2011, 98, 288–295. [Google Scholar] [CrossRef]
- Holm, K.; Staaf, J.; Lauss, M.; Aine, M.; Lindgren, D.; Bendahl, P.; Vallonchristersson, J.; Barkardottir, R.B.; Hoglund, M.; Borg, A.; et al. An integrated genomics analysis of epigenetic subtypes in human breast tumors links DNA methylation patterns to chromatin states in normal mammary cells. Breast Cancer Res. 2016, 18, 27. [Google Scholar] [CrossRef] [PubMed]
- Andersson, R.; Gebhard, C.; Miguelescalada, I.; Hoof, I.; Bornholdt, J.; Boyd, M.; Chen, Y.; Zhao, X.; Schmidl, C.; Suzuki, T.; et al. An atlas of active enhancers across human cell types and tissues. Nature 2014, 507, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, M.F.; Huber, W.; Pages, H.; Aboyoun, P.; Carlson, M.R.J.; Gentleman, R.; Morgan, M.T.; Carey, V.J. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 2013, 9, e1003118. [Google Scholar] [CrossRef] [PubMed]
- The R Project for Statistical Computing. Available online: www.r-project.org (accessed on 1 January 2016).
- Mclendon, R.E.; Friedman, A.H.; Bigner, D.D.; van Meir, E.G.; Brat, D.J.; Mastrogianakis, G.M.; Olson, J.J.; Mikkelsen, T.; Lehman, N.L.; Aldape, K.; et al. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008, 455, 1061–1068. [Google Scholar]
- Bergh, J.C. Gene amplification in human lung cancer. The myc family genes and other proto-oncogenes and growth factor genes. Am. Rev. Respir. Dis. 1990, 142, S20–S26. [Google Scholar] [CrossRef]
- Corces, M.R.; Trevino, A.E.; Hamilton, E.G.; Greenside, P.; Sinnottarmstrong, N.A.; Vesuna, S.; Satpathy, A.T.; Rubin, A.J.; Montine, K.S.; Wu, B.; et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 2017, 14, 959–962. [Google Scholar] [CrossRef]
- Saal, L.H.; Vallonchristersson, J.; Hakkinen, J.; Hegardt, C.; Grabau, D.; Winter, C.; Brueffer, C.; Tang, M.E.; Reutersward, C.; Schulz, R.; et al. The Sweden Cancerome Analysis Network—Breast (SCAN-B) Initiative: A large-scale multicenter infrastructure towards implementation of breast cancer genomic analyses in the clinical routine. Genome Med. 2015, 7, 20. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arbajian, E.; Aine, M.; Karlsson, A.; Vallon-Christersson, J.; Brunnström, H.; Davidsson, J.; Mohlin, S.; Planck, M.; Staaf, J. Methylation Patterns and Chromatin Accessibility in Neuroendocrine Lung Cancer. Cancers 2020, 12, 2003. https://doi.org/10.3390/cancers12082003
Arbajian E, Aine M, Karlsson A, Vallon-Christersson J, Brunnström H, Davidsson J, Mohlin S, Planck M, Staaf J. Methylation Patterns and Chromatin Accessibility in Neuroendocrine Lung Cancer. Cancers. 2020; 12(8):2003. https://doi.org/10.3390/cancers12082003
Chicago/Turabian StyleArbajian, Elsa, Mattias Aine, Anna Karlsson, Johan Vallon-Christersson, Hans Brunnström, Josef Davidsson, Sofie Mohlin, Maria Planck, and Johan Staaf. 2020. "Methylation Patterns and Chromatin Accessibility in Neuroendocrine Lung Cancer" Cancers 12, no. 8: 2003. https://doi.org/10.3390/cancers12082003
APA StyleArbajian, E., Aine, M., Karlsson, A., Vallon-Christersson, J., Brunnström, H., Davidsson, J., Mohlin, S., Planck, M., & Staaf, J. (2020). Methylation Patterns and Chromatin Accessibility in Neuroendocrine Lung Cancer. Cancers, 12(8), 2003. https://doi.org/10.3390/cancers12082003





