Stratifying Brain Tumour Histological Sub-Types: The Application of ATR-FTIR Serum Spectroscopy in Secondary Care
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Preparation
2.2. Spectral Collection
2.3. Spectral Analysis
3. Results
3.1. Brain Tumour vs. Healthy Control
3.1.1. Principal Component Analysis
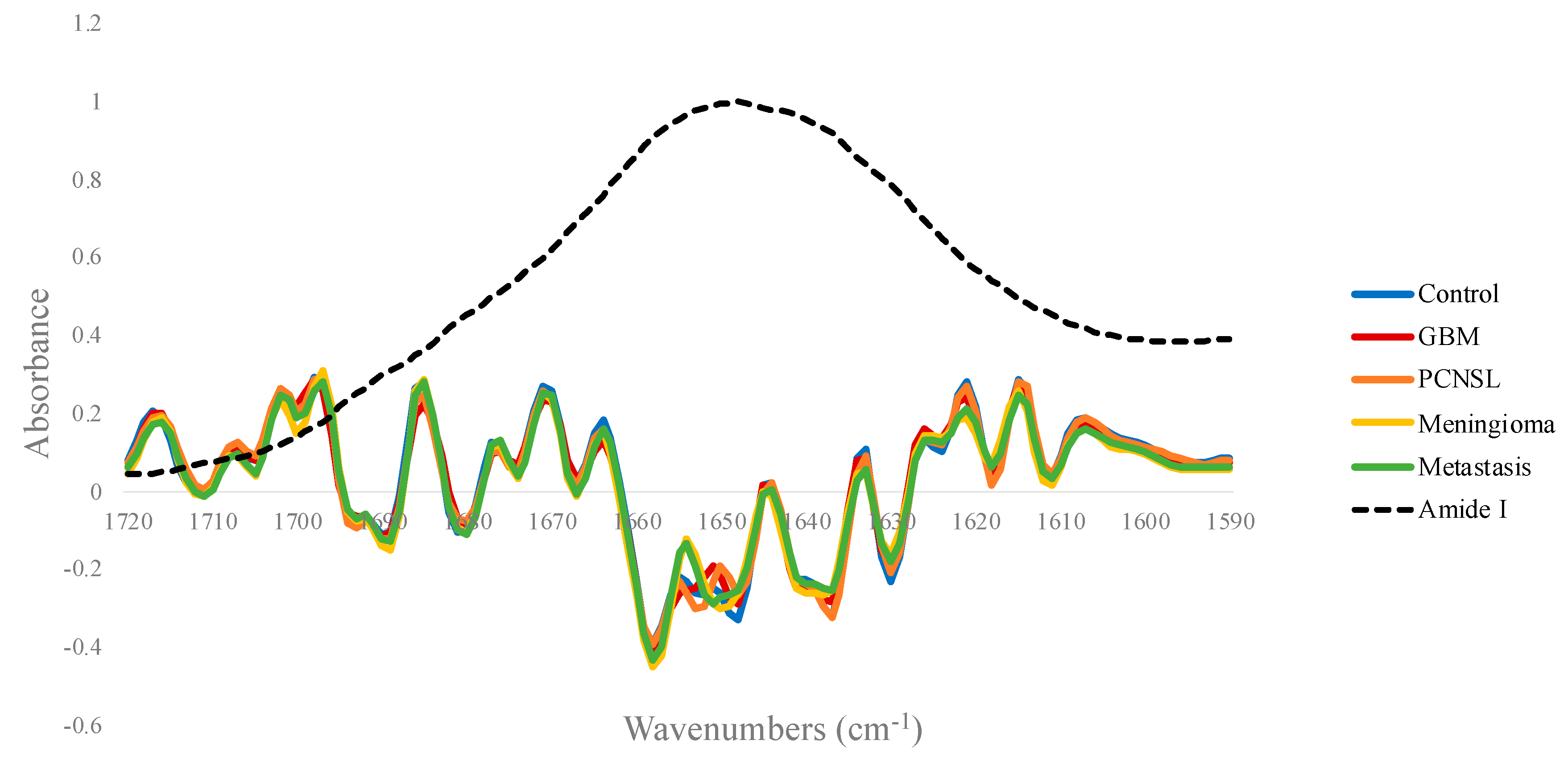
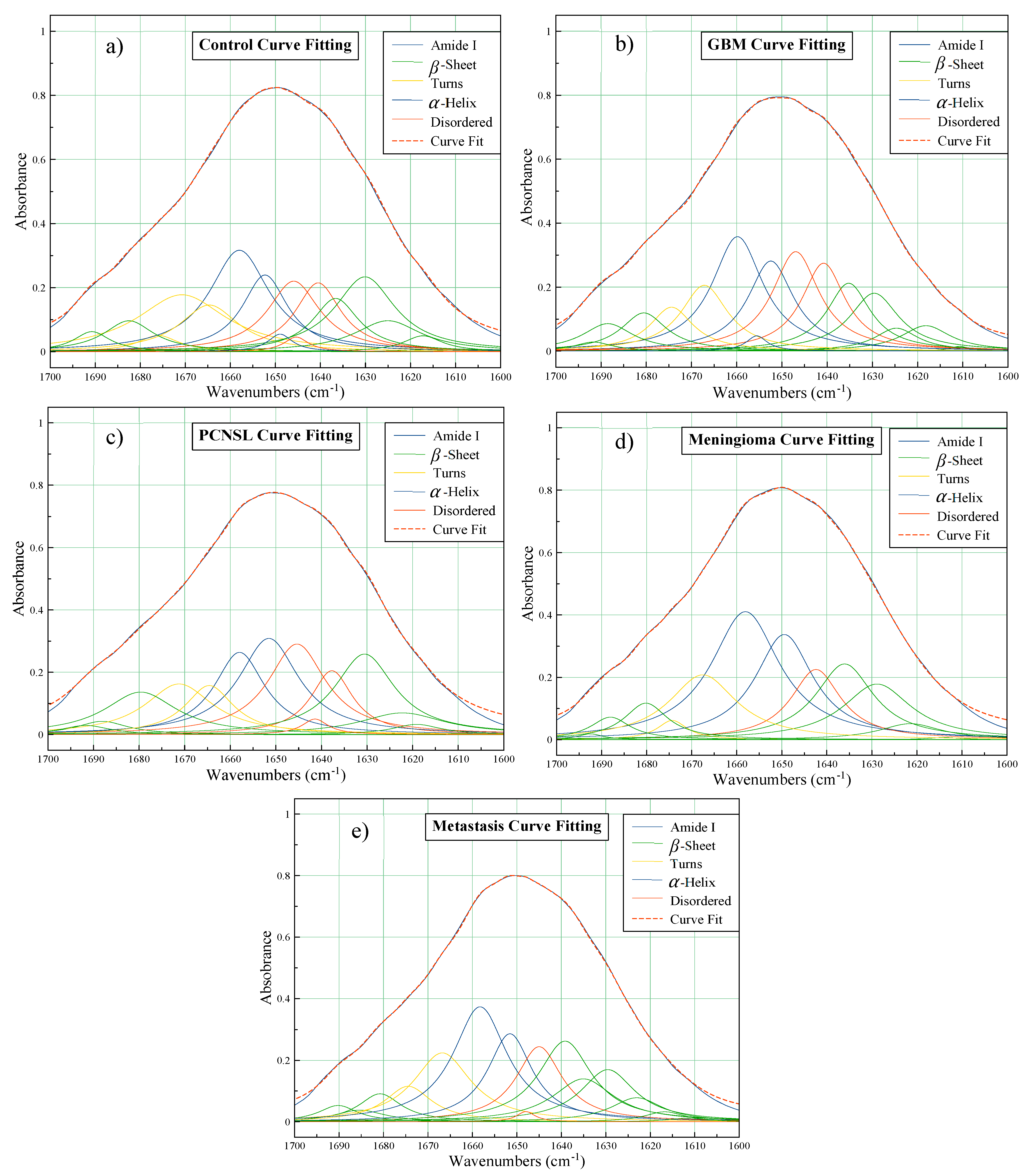
3.1.2. Amide I Deconvolution
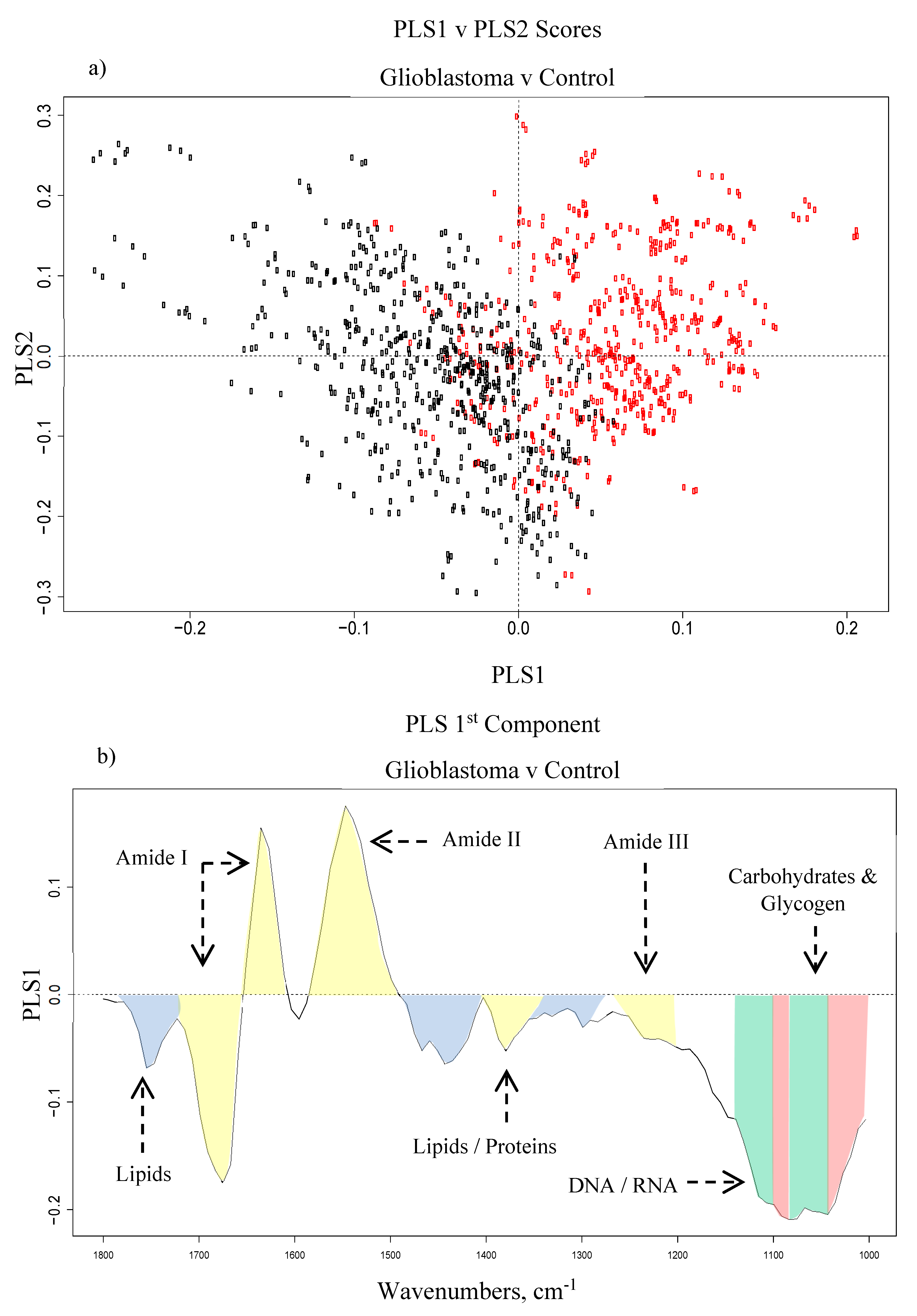
3.1.3. Partial Least Squares-Discriminant Analysis
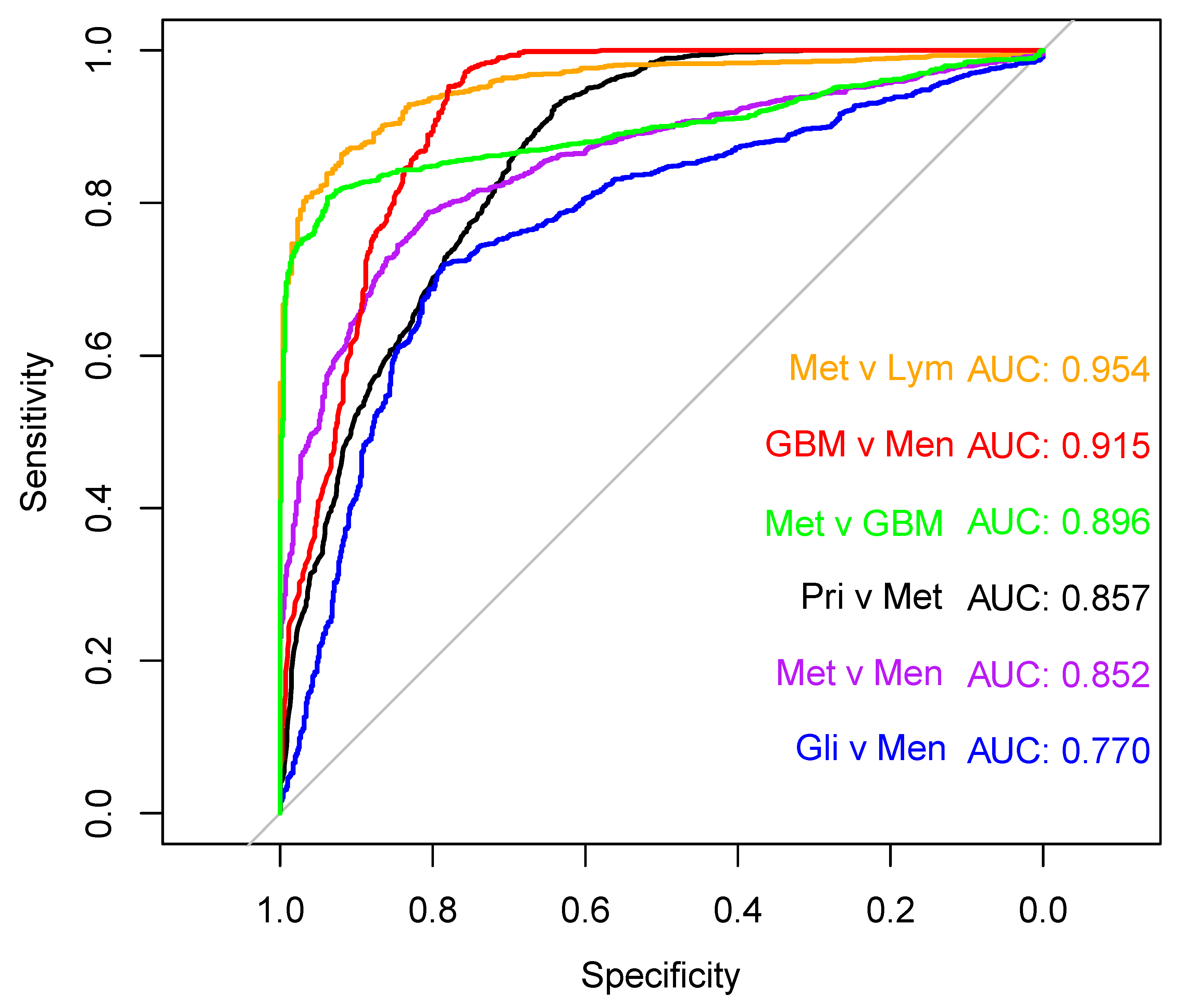
3.2. Brain Tumour Differentiation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Burnet, N.G.; Jefferies, S.J.; Benson, R.J.; Hunt, D.P.; Treasure, F.P. ‘Years of life lost (YLL) from cancer is an important measure of population burden—And should be considered when allocating research funds’. Br. J. Cancer 2005, 92, 241–245. [Google Scholar] [CrossRef]
- Brain Tumour Research. ‘Report on National Research Funding’. 2016. Available online: https://www.braintumourresearch.org/docs/default-source/default-document-library/public-affairs-and-campaigning-documents/brain-tumour-research—report-on-national-research-funding—2016.pdf (accessed on 10 September 2017).
- Patel, A.P.; Fisher, J.L.; Nichols, E.; Abd-Allah, F.; Abdela, J.; Abdelalim, A.; Abraha, H.N.; Agius, D.; Alahdab, F.; Alam, T.; et al. Global, regional, and national burden of brain and other CNS cancer, 1990–2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2019, 18, 376–393. [Google Scholar] [CrossRef]
- Brain Tumour Research. ‘Brain Tumour Research’. Available online: https://www.braintumourresearch.org/campaigning/stark-facts?gclid=CjwKCAiAp5nyBRABEiwApTwjXkb2HvKCz7rHEiQBR4swacKw6zZK3X3d6Fj6W0p8TGOfm_Ab6VNhURoCplgQAvD_BwE (accessed on 10 September 2017).
- Yan, P.-F.; Yan, L.; Zhang, Z.; Salim, A.; Wang, L.; Hu, T.T.; Zhao, H.Y. Accuracy of conventional MRI for preoperative diagnosis of intracranial tumors: A retrospective cohort study of 762 cases. Int. J. Surg. 2016, 36, 109–117. [Google Scholar] [CrossRef]
- Pope, W.B.; Brandal, G. Conventional and advanced magnetic resonance imaging in patients with high-grade glioma. Q. J. Nucl. Med. Mol. Imaging 2018, 62, 239–253. [Google Scholar] [CrossRef]
- Giannini, C.; Dogan, A.; Salomão, D.R. CNS Lymphoma: A Practical Diagnostic Approach. J. Neuropathol. Exp. Neurol. 2014, 73, 478–494. [Google Scholar] [CrossRef] [PubMed]
- Infusino, I.; Panteghini, M. Serum albumin: Accuracy and clinical use. Clin. Chim. Acta 2013, 419, 15–18. [Google Scholar] [CrossRef] [PubMed]
- Stuart, B. Biological Applications of Infrared Spectroscopy; John Wiley & Sons: Chichester, UK; New York, NY, USA, 1997. [Google Scholar]
- Baker, M.J.; Trevisan, J.; Bassan, P.; Bhargava, R.; Butler, H.J.; Dorling, K.M.; Fielden, P.R.; Fogarty, S.W.; Fullwood, N.J.; Heys, K.A.; et al. Using Fourier transform IR spectroscopy to analyze biological materials. Nat. Protoc. 2014, 9, 1771–1791. [Google Scholar] [CrossRef] [PubMed]
- Bellisola, G.; Sorio, C. Infrared spectroscopy and microscopy in cancer research and diagnosis. Am. J. Cancer Res. 2012, 2, 1. [Google Scholar] [PubMed]
- Butler, H.J.; Brennan, P.M.; Cameron, J.M.; Finlayson, D.; Hegarty, M.G.; Jenkinson, M.D.; Palmer, D.S.; Smith, B.R.; Baker, M.J. Development of high-throughput ATR-FTIR technology for rapid triage of brain cancer. Nat. Commun. 2019, 10, 4501. [Google Scholar] [CrossRef] [PubMed]
- Cameron, J.M.; Butler, H.J.; Smith, B.R.; Hegarty, M.G.; Jenkinson, M.D.; Syed, K.; Brennan, P.M.; Ashton, K.; Dawson, T.; Palmer, D.S.; et al. Developing infrared spectroscopic detection for stratifying brain tumour patients: Glioblastoma multiforme vs. lymphoma. Analyst 2019, 144, 6736–6750. [Google Scholar] [CrossRef]
- Campos, S.; Davey, P.; Hird, A.; Pressnail, B.; Bilbao, J.; Aviv, R.I.; Symons, S.; Pirouzmand, F.; Sinclair, E.; Culleton, S.; et al. Brain metastasis from an unknown primary, or primary brain tumour? A diagnostic dilemma. Curr. Oncol. 2009, 16, 62. [Google Scholar] [PubMed]
- Cameron, J.M.; Butler, H.J.; Palmer, D.S.; Baker, M.J. Biofluid spectroscopic disease diagnostics: A review on the processes and spectral impact of drying. J. Biophotonics 2018, 11, e201700299. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.R.; Baker, M.J.; Palmer, D.S. PRFFECT: A versatile tool for spectroscopists. Chemom. Intell. Lab. Syst. 2018, 172, 33–42. [Google Scholar] [CrossRef]
- Butler, H.J.; Smith, B.R.; Fritzsch, R.; Radhakrishnan, P.; Palmer, D.S.; Baker, M.J. Optimised spectral pre-processing for discrimination of biofluids via ATR-FTIR spectroscopy. Analyst 2018, 143, 6121–6134. [Google Scholar] [CrossRef]
- Abdi, H.; Williams, L.J. Principal component analysis: Principal component analysis. WIREs Comput. Stat. 2010, 2, 433–459. [Google Scholar] [CrossRef]
- Karthikeyan, S.; Easwaran, R. Analysis of a curve fitting model in the amide region applied to the muscle tissues of an edible fish: Labeo rohita fingerlings. JBPC 2013, 13, 125–130. [Google Scholar] [CrossRef]
- Sarver, R.W.; Krueger, W.C. Protein secondary structure from fourier transform infrared spectroscopy: A data base analysis. Anal. Biochem. 1991, 194, 89–100. [Google Scholar] [CrossRef]
- Breiman. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Ballabio, D.; Consonni, V. Classification tools in chemistry. Part 1: Linear models. PLS-DA. Anal. Methods 2013, 5, 3790. [Google Scholar] [CrossRef]
- Huang, S.; Cai, N.; Pacheco, P.P.; Narrandes, S.; Wang, Y.; Xu, W. Applications of Support Vector Machine (SVM) Learning in Cancer Genomics. Cancer Genom. Proteom. 2018, 15, 41–51. [Google Scholar] [CrossRef]
- Krafft, C.; Shapoval, L.; Sobottka, S.B.; Schackert, G.; Salzer, R. Identification of Primary Tumors of Brain Metastases by Infrared Spectroscopic Imaging and Linear Discriminant Analysis. Technol. Cancer Res. Treat. 2006, 5, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Glassford, S.E.; Byrne, B.; Kazarian, S.G. Recent applications of ATR FTIR spectroscopy and imaging to proteins. Biochim. Biophys. Acta (BBA) Proteins Proteom. 2013, 1834, 2849–2858. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.; Yu, S. Fourier Transform Infrared Spectroscopic Analysis of Protein Secondary Structures. Acta Biochim. Biophys. Sin. 2007, 39, 549–559. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Secondary Structure: Polypeptide Chains Can Fold Into Regular Structures Such as the Alpha Helix, the Beta Sheet, and Turns and Loops. In Biochemistry, 5th ed.; W.H. Freeman: New York, NY, USA, 2002. [Google Scholar]
- Barth. Infrared spectroscopy of proteins. Biochim. Biophys. Acta (BBA) Bioenerg. 2007, 1767, 1073–1101. [Google Scholar] [CrossRef] [PubMed]
- Gray, E.; Butler, H.J.; Board, R.; Brennan, P.M.; Chalmers, A.J.; Dawson, T.; Goodden, J.; Hamilton, W.; Hegarty, M.G.; James, A.; et al. Health economic evaluation of a serum-based blood test for brain tumour diagnosis: Exploration of two clinical scenarios. BMJ Open 2018, 8, e017593. [Google Scholar] [CrossRef] [PubMed]
- Reynés, G.; Vila, V.; Martín, M.; Parada, A.; Fleitas, T.; Reganon, E.; Martínez-Sales, V. Circulating markers of angiogenesis, inflammation, and coagulation in patients with glioblastoma. J. Neurooncol. 2011, 102, 35–41. [Google Scholar] [CrossRef]
- Hormigo, A.; Gu, B.; Karimi, S.; Riedel, E.; Panageas, K.S.; Edgar, M.A.; Tanwar, M.K.; Rao, J.S.; Fleisher, M.; DeAngelis, L.M.; et al. YKL-40 and Matrix Metalloproteinase-9 as Potential Serum Biomarkers for Patients with High-Grade Gliomas. Clin. Cancer Res. 2006, 12, 5698–5704. [Google Scholar] [CrossRef]
- Iwamoto, F.M.; Hottinger, A.F.; Karimi, S.; Riedel, E.; Dantis, J.; Jahdi, M.; Panageas, K.S.; Lassman, A.B.; Abrey, L.E.; Fleisher, M.; et al. Serum YKL-40 is a marker of prognosis and disease status in high-grade gliomas. Neuro-Oncology 2011, 13, 1244–1251. [Google Scholar] [CrossRef]
- Albulescu, R.; Codrici, E.; Popescu, I.D.; Mihai, S.; Necula, L.G.; Petrescu, D.; Teodoru, M.; Tanase, C.P. Cytokine Patterns in Brain Tumour Progression. Mediat. Inflamm. 2013, 2013, 1–7. [Google Scholar] [CrossRef]
- Lippitz, B.E.; Harris, R.A. Cytokine patterns in cancer patients: A review of the correlation between interleukin 6 and prognosis. OncoImmunology 2016, 5, e1093722. [Google Scholar] [CrossRef]
- Ghimire, H.; Venkataramani, M.; Bian, Z.; Liu, Y.; Perera, A.G.U. ATR-FTIR spectral discrimination between normal and tumorous mouse models of lymphoma and melanoma from serum samples. Sci. Rep. 2017, 7, 16993. [Google Scholar] [CrossRef] [PubMed]
- Surowka, D.; Adamek, D.; Szczerbowska-Boruchowska, M. The combination of artificial neural networks and synchrotron radiation-based infrared micro-spectroscopy for a study on the protein composition of human glial tumors. Analyst 2015, 140, 2428–2438. [Google Scholar] [CrossRef] [PubMed]
- Petricoin, E.F.; Belluco, C.; Araujo, R.P.; Liotta, L.A. The blood peptidome: A higher dimension of information content for cancer biomarker discovery. Nat. Rev. Cancer 2006, 6, 961–967. [Google Scholar] [CrossRef] [PubMed]
- Petrich, W. Mid-infrared and Raman spectroscopy for medical diagnostics. Appl. Spectrosc. Rev. 2001, 36, 181–237. [Google Scholar] [CrossRef]
- Byrne, H.J. Vibrational Spectroscopy: Disease Diagnostics and Beyond. In Optical Spectroscopy and Computational Methods in Biology and Medicine; Springer: New York, NY, USA, 2013. [Google Scholar]
- Haber, D.A.; Velculescu, V.E. Blood-Based Analyses of Cancer: Circulating Tumor Cells and Circulating Tumor DNA. Cancer Discov. 2014, 4, 650–661. [Google Scholar] [CrossRef]
- Zhang, L.; Liang, Y.; Li, S.; Zeng, F.; Meng, Y.; Chen, Z.; Liu, S.; Tao, Y.; Yu, F. The interplay of circulating tumor DNA and chromatin modification, therapeutic resistance, and metastasis. Mol. Cancer 2019, 18, 36. [Google Scholar] [CrossRef]
- Bark, J.M.; Kulasinghe, A.; Chua, B.; Day, B.W.; Punyadeera, C. Circulating biomarkers in patients with glioblastoma. Br. J. Cancer 2020, 122, 295–305. [Google Scholar] [CrossRef]
- Boire, A.; Brandsma, D.; Brastianos, P.K.; Le Rhun, E.; Ahluwalia, M.; Junck, L.; Glantz, M.; Groves, M.D.; Lee, E.Q.; Lin, N.; et al. Liquid biopsy in central nervous system metastases: A RANO review and proposals for clinical applications. Neuro-Oncology 2019, 21, 571–584. [Google Scholar] [CrossRef]
- Mariner, P.D.; Korst, A.; Karimpour-Fard, A.; Stauffer, B.L.; Miyamoto, S.D.; Sucharov, C.C. Improved Detection of Circulating miRNAs in Serum and Plasma Following Rapid Heat/Freeze Cycling. MIRNA 2018, 7, 138–147. [Google Scholar] [CrossRef]
- Xiao, D.-D.; Yan, P.-F.; Wang, Y.-X.; Osman, M.S.; Zhao, H.-Y. Glioblastoma and primary central nervous system lymphoma: Preoperative differentiation by using MRI-based 3D texture analysis. Clin. Neurol. Neurosurg. 2018, 173, 84–90. [Google Scholar] [CrossRef]
| Tumour Type Against Healthy Control (n = 87) | No. of Patients | Sampling | Sensitivity (%) | Specificity (%) | Balanced Accuracy (%) | |||
|---|---|---|---|---|---|---|---|---|
| Mean | SD | Mean | SD | Mean | SD | |||
| GBM | 96 | No | 95.5 | 4.3 | 94.9 | 4.2 | 95.2 | 2.9 |
| PCNSL | 41 | Up | 92.2 | 6.9 | 96.7 | 3.5 | 94.4 | 3.9 |
| Meningioma | 111 | Up | 94.7 | 3.7 | 98.4 | 2.2 | 96.6 | 2.0 |
| Metastasis | 210 | Up | 95.9 | 2.6 | 95.0 | 4.2 | 95.4 | 2.3 |
| Approximate Wavenumbers (cm−1) | Tentative Biological Assignments | Vibrational Modes |
|---|---|---|
| 1012 | Carbohydrate | C-O stretch |
| 1030 | Glycogen | C-O and C-C stretch, C-OH deformation |
| 1045 | DNA and RNA | symmetric stretch |
| 1050 | Carbohydrate/Glycogen | C-O-C stretching and bending |
| 1050–1100 | DNA and RNA | Symmetric stretch |
| 1240–1310 | Amide III of Proteins | N-H in plane bend, C-N stretch |
| 1245 | Phosphodiesters | Asymmetric stretch |
| 1340 | Phospholipids | CH2 wagging |
| 1400 | Lipids/Proteins | CH3 bending |
| 1470 | Lipids | CH2 scissoring |
| 1500–1600 | Amide II of Proteins | N-H bending, C-N stretching |
| 1600–1700 | Amide I of Proteins | C=O and C-N stretch, N-H bending |
| 1750 | Lipids | C=O stretching |
| Classification (Positive Class v Negative Class) | No. of Patients (Positive Class/ Negative Class) | Model + Sampling | Sensitivity (%) | Specificity (%) | Balanced Accuracy (%) | |||
|---|---|---|---|---|---|---|---|---|
| Mean | SD | Mean | SD | Mean | SD | |||
| Primary v Metastasis | 303/210 | RF + up | 90.9 | 3.1 | 66.4 | 5.5 | 78.8 | 2.8 |
| Glioma v Meningioma | 192/111 | SVM + down | 70.9 | 5.5 | 81.8 | 6.2 | 76.3 | 4.4 |
| GBM v Meningioma | 96/111 | RF + no | 94.4 | 5.1 | 83.4 | 5.6 | 88.9 | 3.0 |
| Metastasis v GBM | 210/96 | SVM + down | 84.3 | 3.8 | 96.2 | 3.4 | 90.3 | 2.6 |
| Metastasis v PCNSL | 210/41 | PLS-DA + smote | 91.5 | 3.1 | 91.1 | 9.2 | 91.3 | 4.6 |
| Metastasis v Meningioma | 210/111 | PLS-DA + up | 71.3 | 6.2 | 86.1 | 5.5 | 78.7 | 3.6 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cameron, J.M.; Rinaldi, C.; Butler, H.J.; Hegarty, M.G.; Brennan, P.M.; Jenkinson, M.D.; Syed, K.; Ashton, K.M.; Dawson, T.P.; Palmer, D.S.; et al. Stratifying Brain Tumour Histological Sub-Types: The Application of ATR-FTIR Serum Spectroscopy in Secondary Care. Cancers 2020, 12, 1710. https://doi.org/10.3390/cancers12071710
Cameron JM, Rinaldi C, Butler HJ, Hegarty MG, Brennan PM, Jenkinson MD, Syed K, Ashton KM, Dawson TP, Palmer DS, et al. Stratifying Brain Tumour Histological Sub-Types: The Application of ATR-FTIR Serum Spectroscopy in Secondary Care. Cancers. 2020; 12(7):1710. https://doi.org/10.3390/cancers12071710
Chicago/Turabian StyleCameron, James M., Christopher Rinaldi, Holly J. Butler, Mark G Hegarty, Paul M. Brennan, Michael D. Jenkinson, Khaja Syed, Katherine M. Ashton, Timothy P. Dawson, David S. Palmer, and et al. 2020. "Stratifying Brain Tumour Histological Sub-Types: The Application of ATR-FTIR Serum Spectroscopy in Secondary Care" Cancers 12, no. 7: 1710. https://doi.org/10.3390/cancers12071710
APA StyleCameron, J. M., Rinaldi, C., Butler, H. J., Hegarty, M. G., Brennan, P. M., Jenkinson, M. D., Syed, K., Ashton, K. M., Dawson, T. P., Palmer, D. S., & Baker, M. J. (2020). Stratifying Brain Tumour Histological Sub-Types: The Application of ATR-FTIR Serum Spectroscopy in Secondary Care. Cancers, 12(7), 1710. https://doi.org/10.3390/cancers12071710






