Targeted Next-Generation Sequencing of 117 Routine Clinical Samples Provides Further Insights into the Molecular Landscape of Uveal Melanoma
Abstract
1. Introduction
2. Results
2.1. Patient and Tumor Demographics
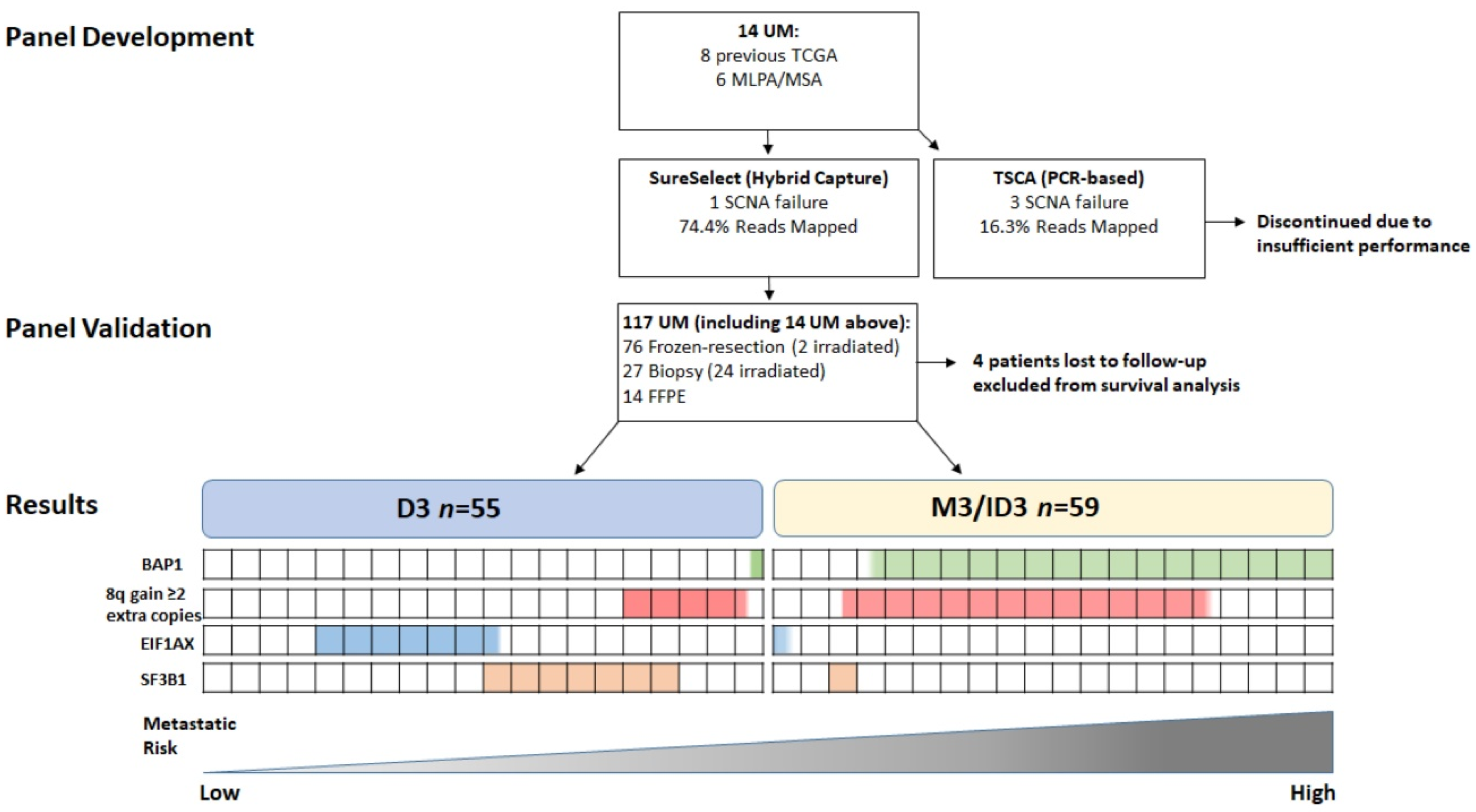
2.2. Panel Comparison (14 Samples)
2.3. Mutation Frequency
2.4. SCNA Analysis and Comparison with MLPA/MSA
2.5. Cox Regression
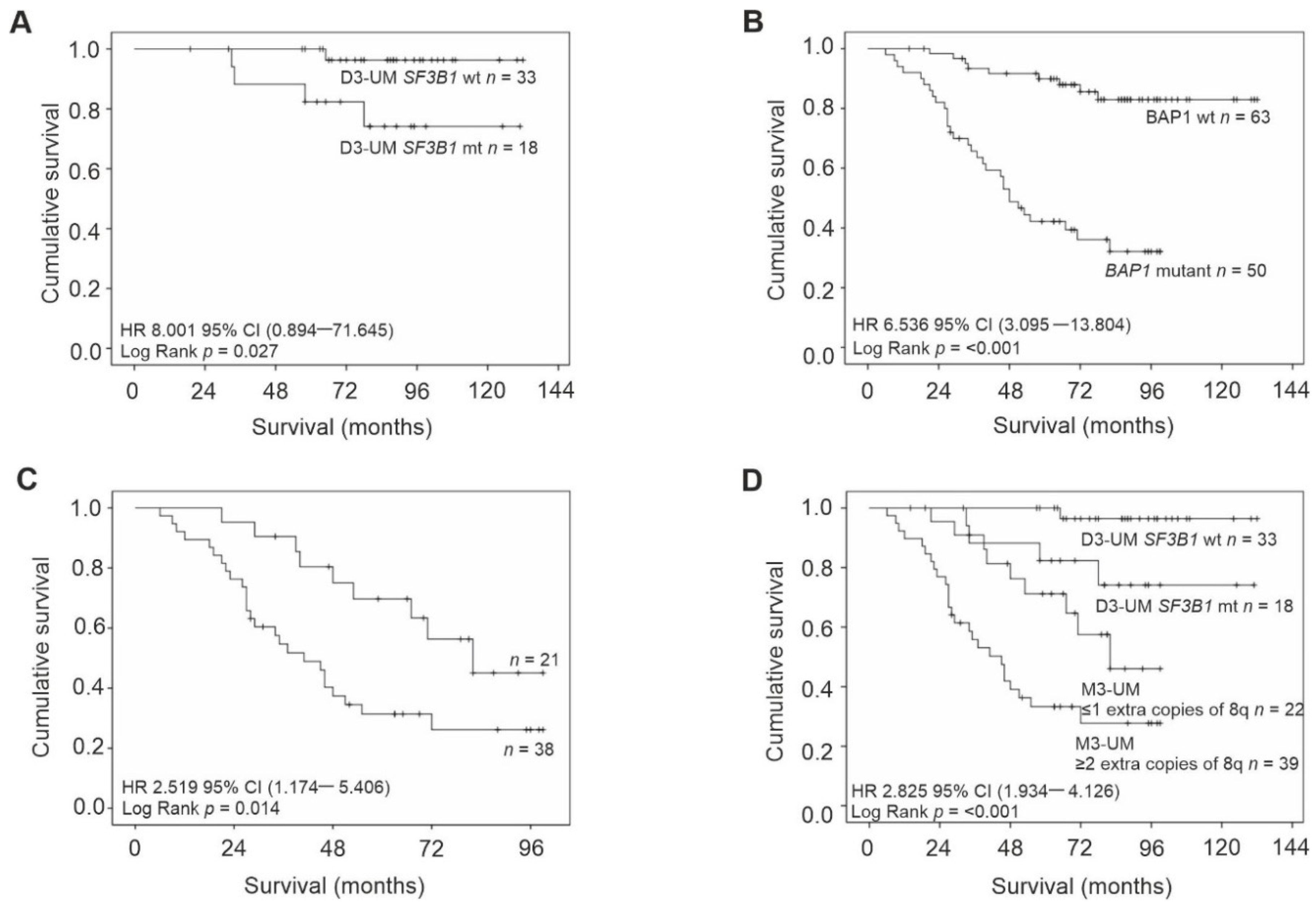
2.6. Survival
2.7. BAP1 IHC
2.8. SF3B1 Mutations in M3 UM
3. Discussion
3.1. Enrichment Comparison
3.2. Comparison with Previous MLPA
3.3. Irradiated Samples
3.4. BAP1 Mutations
3.5. SF3B1 Mutations
3.6. EIF1AX Mutations
3.7. Initiating Mutations
3.8. Other Mutations
4. Materials and Methods
4.1. Patients
4.2. Specimen Characteristics
4.3. Study Design
4.4. Assay Methods
4.4.1. Morphological/Histological Studies
4.4.2. DNA Extraction and Quantification
4.4.3. Chromosomal SCNA Analysis
4.4.4. Next-Generation Sequencing
4.4.5. Sanger Sequencing
4.5. NGS Data Analysis
4.6. Statistical Analysis Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Virgili, G.; Gatta, G.; Ciccolallo, L.; Capocaccia, R.; Biggeri, A.; Crocetti, E.; Lutz, J.M.; Paci, E. Incidence of uveal melanoma in Europe. Ophthalmology 2007, 114, 2309–2315. [Google Scholar] [CrossRef] [PubMed]
- Yonekawa, Y.; Kim, I.K. Epidemiology and management of uveal melanoma. Hematol./Oncol. Clin. N. Am. 2012, 26, 1169–1184. [Google Scholar] [CrossRef] [PubMed]
- Damato, B. Progress in the management of patients with uveal melanoma. the 2012 Ashton Lecture. Eye 2012, 26, 1157–1172. [Google Scholar] [CrossRef] [PubMed]
- Carvajal, R.D.; Schwartz, G.K.; Tezel, T.; Marr, B.; Francis, J.H.; Nathan, P.D. Metastatic disease from uveal melanoma: Treatment options and future prospects. Br. J. Ophthalmol. 2017, 101, 38–44. [Google Scholar] [CrossRef]
- Damato, B.; Eleuteri, A.; Taktak, A.F.G.; Coupland, S.E. Estimating prognosis for survival after treatment of choroidal melanoma. Prog. IN Retin. Eye Res. 2011, 30, 285–295. [Google Scholar] [CrossRef]
- DeParis, S.W.; Taktak, A.; Eleuteri, A.; Enanoria, W.; Heimann, H.; Coupland, S.E.; Damato, B. External Validation of the Liverpool Uveal Melanoma Prognosticator Online. Investig. Ophthalmol. Vis. Sci. 2016, 57, 6116–6122. [Google Scholar] [CrossRef]
- Gomez, D.; Wetherill, C.; Cheong, J.; Jones, L.; Marshall, E.; Damato, B.; Coupland, S.E.; Ghaneh, P.; Poston, G.J.; Malik, H.Z.; et al. The Liverpool uveal melanoma liver metastases pathway: Outcome following liver resection. J. Surg. Oncol. 2014, 109, 542–547. [Google Scholar] [CrossRef]
- Prescher, G.; Bornfeld, N. Prognostic implications of monosomy 3 in uveal melanoma. Lancet 1996, 347, 1222. [Google Scholar]
- Horsman, D.E.; Rootman, J.; White, V.A.; Sroka, H. Monosomy 3 and isochromosome 8q in a uveal melanoma. Cancer Genet. Cytogenet. 1990, 45, 249–253. [Google Scholar] [CrossRef]
- Sisley, K.; Cottam, D.W.; Rennie, I.G.; Parsons, M.A.; Potter, A.M.; Potter, C.W.; Rees, R.C. Non-random abnormalities of chromosomes 3, 6, and 8 associated with posterior uveal melanoma. Genes Chromosomes Cancer 1992, 5, 197–200. [Google Scholar] [CrossRef]
- Robertson, A.G.; Shih, J.; Yau, C.; Gibb, E.A.; Oba, J.; Mungall, K.L.; Hess, J.M.; Uzunangelov, V.; Walter, V.; Danilova, L.; et al. Integrative Analysis Identifies Four Molecular and Clinical Subsets in Uveal Melanoma. Cancer Cell 2017, 32, 204–220. [Google Scholar] [CrossRef]
- Kilic, E.; Naus, N.C.; van Gils, W.; Klaver, C.C.; van Til, M.E.; Verbiest, M.M.; Stijnen, T.; Mooy, C.M.; Paridaens, D.; Beverloo, H.B.; et al. Concurrent loss of chromosome arm 1p and chromosome 3 predicts a decreased disease-free survival in uveal melanoma patients. Investig. Ophthalmol. Vis. Sci. 2005, 46, 2253–2257. [Google Scholar] [CrossRef] [PubMed]
- Damato, B.; Dopierala, J.; Klaasen, A.; van Dijk, M.; Sibbring, J.; Coupland, S.E. Multiplex ligation-dependent probe amplification of uveal melanoma: Correlation with metastatic death. Investig. Ophthalmol. Vis. Sci. 2009, 50, 3048–3055. [Google Scholar] [CrossRef] [PubMed]
- Royer-Bertrand, B.; Torsello, M.; Rimoldi, D.; El Zaoui, I.; Cisarova, K.; Pescini-Gobert, R.; Raynaud, F.; Zografos, L.; Schalenbourg, A.; Speiser, D.; et al. Comprehensive Genetic Landscape of Uveal Melanoma by Whole-Genome Sequencing. Am. J. Hum. Genet. 2016, 99, 1190–1198. [Google Scholar] [CrossRef] [PubMed]
- Van Raamsdonk, C.D.; Bezrookove, V.; Green, G.; Bauer, J.; Gaugler, L.; O’Brien, J.M.; Simpson, E.M.; Barsh, G.S.; Bastian, B.C. Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature 2009, 457, 599–602. [Google Scholar] [CrossRef]
- Van Raamsdonk, C.D.; Griewank, K.G.; Crosby, M.B.; Garrido, M.C.; Vemula, S.; Wiesner, T.; Obenauf, A.C.; Wackernagel, W.; Green, G.; Bouvier, N.; et al. Mutations in GNA11 in uveal melanoma. N. Engl. J. Med. 2010, 363, 2191–2199. [Google Scholar] [CrossRef]
- Moore, A.R.; Ceraudo, E.; Sher, J.J.; Guan, Y.; Shoushtari, A.N.; Chang, M.T.; Zhang, J.Q.; Walczak, E.G.; Kazmi, M.A.; Taylor, B.S.; et al. Recurrent activating mutations of G-protein-coupled receptor CYSLTR2 in uveal melanoma. Nat. Genet. 2016, 48, 675–680. [Google Scholar] [CrossRef]
- Johansson, P.; Aoude, L.G.; Wadt, K.; Glasson, W.J.; Warrier, S.K.; Hewitt, A.W.; Kiilgaard, J.F.; Heegaard, S.; Isaacs, T.; Franchina, M.; et al. Deep sequencing of uveal melanoma identifies a recurrent mutation in PLCB4. Oncotarget 2015, 7, 4624–4631. [Google Scholar] [CrossRef]
- Harbour, J.W.; Onken, M.D.; Worley, L.A.; Matatall, K.A.; Roberson, E.D.O.; Duan, S.; Cao, L.; Council, M.L.; Helms, C.; Bowcock, A.M. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science 2010, 330, 1410–1413. [Google Scholar] [CrossRef]
- Martin, M.; Maßhöfer, L.; Temming, P.; Rahmann, S.; Metz, C.; Bornfeld, N.; van de Nes, J.; Klein-Hitpass, L.; Hinnebusch, A.G.; Horsthemke, B.; et al. Exome sequencing identifies recurrent somatic mutations in EIF1AX and SF3B1 in uveal melanoma with disomy 3. Nat. Genet. 2013, 45, 933–936. [Google Scholar] [CrossRef]
- Farquhar, N.; Thornton, S.; Coupland, S.E.; Coulson, J.M.; Sacco, J.J.; Krishna, Y.; Heimann, H.; Taktak, A.; Cebulla, C.M.; Abdel-Rahman, M.H.; et al. Patterns of BAP1 protein expression provide insights into prognostic significance and the biology of uveal melanoma. J. Pathol. Clin. Res. 2017, 4, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Yavuzyigitoglu, S.; Koopmans, A.E.; Verdijk, R.M.; Vaarwater, J.; Eussen, B.; van Bodegom, A.; Paridaens, D.; Kilic, E.; de Klein, A. Uveal Melanomas with SF3B1 Mutations: A Distinct Subclass Associated with Late-Onset Metastases. Ophthalmology 2016, 123, 1118–1128. [Google Scholar] [CrossRef]
- Dono, M.; Angelini, G.; Cecconi, M.; Amaro, A.; Esposito, A.I.; Mirisola, V.; Maric, I.; Lanza, F.; Nasciuti, F.; Viaggi, S.; et al. Mutation frequencies of GNAQ, GNA11, BAP1, SF3B1, EIF1AX and TERT in uveal melanoma: Detection of an activating mutation in the TERT gene promoter in a single case of uveal melanoma. Br. J. Cancer 2014, 110, 1058–1065. [Google Scholar] [CrossRef]
- Reiman, A.; Kikuchi, H.; Scocchia, D.; Smith, P.; Tsang, Y.W.; Snead, D.; Cree, I.A. Validation of an NGS mutation detection panel for melanoma. BMC Cancer 2017, 17, 150. [Google Scholar] [CrossRef]
- Smit, K.N.; van Poppelen, N.M.; Vaarwater, J.; Verdijk, R.; van Marion, R.; Kalirai, H.; Coupland, S.E.; Thornton, S.; Farquhar, N.; Dubbink, H.J.; et al. Combined mutation and copy-number variation detection by targeted next-generation sequencing in uveal melanoma. Mod. Pathol. Off. J. USA Can. Acad. Pathol. Inc. 2018, 31, 763–771. [Google Scholar] [CrossRef] [PubMed]
- Afshar, A.R.; Damato, B.E.; Stewart, J.M.; Zablotska, L.B.; Roy, R.; Olshen, A.B.; Joseph, N.M.; Bastian, B.C. Next-Generation Sequencing of Uveal Melanoma for Detection of Genetic Alterations Predicting Metastasis. Transl. Vis. Sci. Technol. 2019, 8, 18. [Google Scholar] [CrossRef] [PubMed]
- Shain, A.H.; Bagger, M.M.; Yu, R.; Chang, D.; Liu, S.; Vemula, S.; Weier, J.F.; Wadt, K.; Heegaard, S.; Bastian, B.C.; et al. The genetic evolution of metastatic uveal melanoma. Nat. Genet. 2019, 51, 1123–1130. [Google Scholar] [CrossRef] [PubMed]
- Koopmans, A.E.; Verdijk, R.M.; Brouwer, R.W.; van den Bosch, T.P.; van den Berg, M.M.; Vaarwater, J.; Kockx, C.E.; Paridaens, D.; Naus, N.C.; Nellist, M.; et al. Clinical significance of immunohistochemistry for detection of BAP1 mutations in uveal melanoma. Mod. Pathol. Off. J. USA Can. Acad. Pathol. Inc. 2014, 27, 1321–1330. [Google Scholar] [CrossRef]
- Cunha Rola, A.; Taktak, A.; Eleuteri, A.; Kalirai, H.; Heimann, H.; Hussain, R.; Bonnett, L.J.; Hill, C.J.; Traynor, M.; Jager, M.J.; et al. Multicenter External Validation of the Liverpool Uveal Melanoma Prognosticator Online: An OOG Collaborative Study. Cancers 2020, 12, 477. [Google Scholar] [CrossRef]
- Kozarewa, I.; Armisen, J.; Gardner, A.F.; Slatko, B.E.; Hendrickson, C.L. Overview of Target Enrichment Strategies. Curr. Protoc. Mol. Biol. 2015, 112, 7–21. [Google Scholar] [CrossRef]
- Mamedov, T.G.; Pienaar, E.; Whitney, S.E.; TerMaat, J.R.; Carvill, G.; Goliath, R.; Subramanian, A.; Viljoen, H.J. A fundamental study of the PCR amplification of GC-rich DNA templates. Comput. Biol. Chem. 2008, 32, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Nolan, T. Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J. Biomol. Tech. JBT 2004, 15, 155–166. [Google Scholar] [PubMed]
- Hung, S.S.; Meissner, B.; Chavez, E.A.; Ben-Neriah, S.; Ennishi, D.; Jones, M.R.; Shulha, H.P.; Chan, F.C.; Boyle, M.; Kridel, R.; et al. Assessment of Capture and Amplicon-Based Approaches for the Development of a Targeted Next-Generation Sequencing Pipeline to Personalize Lymphoma Management. J. Mol. Diagn. JMD 2018, 20, 203–214. [Google Scholar] [CrossRef] [PubMed]
- Samorodnitsky, E.; Jewell, B.M.; Hagopian, R.; Miya, J.; Wing, M.R.; Lyon, E.; Damodaran, S.; Bhatt, D.; Reeser, J.W.; Datta, J.; et al. Evaluation of Hybridization Capture Versus Amplicon-Based Methods for Whole-Exome Sequencing. Hum. Mutat. 2015, 36, 903–914. [Google Scholar] [CrossRef]
- Versluis, M.; de Lange, M.J.; van Pelt, S.I.; Ruivenkamp, C.A.; Kroes, W.G.; Cao, J.; Jager, M.J.; Luyten, G.P.; van der Velden, P.A. Digital PCR validates 8q dosage as prognostic tool in uveal melanoma. PLoS ONE 2015, 10, e0116371. [Google Scholar] [CrossRef]
- Coupland, S.E.; Kalirai, H.; Ho, V.; Thornton, S.; Damato, B.E.; Heimann, H. Concordant chromosome 3 results in paired choroidal melanoma biopsies and subsequent tumour resection specimens. Br. J. Ophthalmol. 2015, 99, 1444–1450. [Google Scholar] [CrossRef]
- Hussain, R.N.; Kalirai, H.; Groenewald, C.; Kacperek, A.; Errington, R.D.; Coupland, S.E.; Heimann, H.; Damato, B. Prognostic Biopsy of Choroidal Melanoma after Proton Beam Radiation Therapy. Ophthalmology 2016, 123, 2264–2265. [Google Scholar] [CrossRef]
- Thornton, S.; Coupland, S.E.; Heimann, H.; Hussain, R.; Groenewald, C.; Kacperek, A.; Damato, B.; Taktak, A.; Eleuteri, A.; Kalirai, H. Effects of plaque brachytherapy and proton beam radiotherapy on prognostic testing: A comparison of uveal melanoma genotyped by microsatellite analysis. Br. J. Ophthalmol. 2020. [Google Scholar] [CrossRef]
- van de Nes, J.A.; Nelles, J.; Kreis, S.; Metz, C.H.; Hager, T.; Lohmann, D.R.; Zeschnigk, M. Comparing the Prognostic Value of BAP1 Mutation Pattern, Chromosome 3 Status, and BAP1 Immunohistochemistry in Uveal Melanoma. Am. J. Surg. Pathol. 2016, 40, 796–805. [Google Scholar] [CrossRef]
- Izumiyama, T.; Minoshima, S.; Yoshida, T.; Shimizu, N. A novel big protein TPRBK possessing 25 units of TPR motif is essential for the progress of mitosis and cytokinesis. Gene 2012, 511, 202–217. [Google Scholar] [CrossRef]
- Tang, M.-R.; Wang, Y.-X.; Guo, S.; Han, S.-Y.; Wang, D. CSMD1 exhibits antitumor activity in A375 melanoma cells through activation of the Smad pathway. Apoptosis 2012, 17, 927–937. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Quesnelle, K.M.; Sparano, A.; Rao, S.; Park, M.S.; Cohen, M.A.; Wang, Y.; Samanta, M.; Kumar, M.S.; Aziz, M.U.; et al. Characterization CSMD1 in a large set of primary lung, head and neck, breast and skin cancer tissues. Cancer Biol. Ther. 2009, 8, 907–916. [Google Scholar] [CrossRef] [PubMed]
- Machleidt, T.; Geller, P.; Schwandner, R.; Scherer, G.; Kronke, M. Caspase 7-induced cleavage of kinectin in apoptotic cells. FEBS Lett. 1998, 436, 51–54. [Google Scholar] [CrossRef]
- Panier, S.; Boulton, S.J. Double-strand break repair: 53BP1 comes into focus. Nat. Rev. Mol. Cell Biol. 2014, 15, 7–18. [Google Scholar] [CrossRef] [PubMed]
- van Poppelen, N.M.; Drabarek, W.; Smit, K.N.; Vaarwater, J.; Brands, T.; Paridaens, D.; Kilic, E.; de Klein, A. SRSF2 Mutations in Uveal Melanoma: A Preference for In-Frame Deletions? Cancers 2019, 11, 1200. [Google Scholar] [CrossRef]
- Lake, S.L.; Kalirai, H.; Dopierala, J.; Damato, B.E.; Coupland, S.E. Comparison of formalin-fixed and snap-frozen samples analyzed by multiplex ligation-dependent probe amplification for prognostic testing in uveal melanoma. Investig. Ophthalmol. Vis. Sci. 2012, 53, 2647–2652. [Google Scholar] [CrossRef]
- Dopierala, J.; Lake, S.L.; Coupland, S.E.; Damato, B.E.; Taktak, A.F.G. Genetic heterogeneity in uveal melanoma assessed by multiplex ligation-dependent probe amplification. Investig. Ophthalmol. Vis. Sci. 2010, 51, 4898–4905. [Google Scholar] [CrossRef]
- Thomas, S.; Putter, C.; Weber, S.; Bornfeld, N.; Lohmann, D.R.; Zeschnigk, M. Prognostic significance of chromosome 3 alterations determined by microsatellite analysis in uveal melanoma: A long-term follow-up study. Br. J. Cancer 2012, 106, 1171–1176. [Google Scholar] [CrossRef]
- Caines, R.; Eleuteri, A.; Kalirai, H.; Fisher, A.C.; Heimann, H.; Damato, B.E.; Coupland, S.E.; Taktak, A.F. Cluster analysis of multiplex ligation-dependent probe amplification data in choroidal melanoma. Mol. Vis. 2015, 21, 1–11. [Google Scholar]
| Variable | Value (% or Range) |
|---|---|
| Age at PM (years) | |
| Median | 64 (16–87) |
| Gender | |
| Female | 54 (47%) |
| Male | 63 (53%) |
| Survival | |
| Alive | 62 (53%) |
| Death from MUM | 40 (34%) |
| Death other causes | 11 (10%) |
| Lost to follow-up | 4 (3%) |
| Median (months) | 65 (0–132) |
| Largest basal diameter (mm) | |
| Median | 15.0 (4–22) |
| Ultrasound height (mm) | |
| Median | 7.5 (1–15.7) |
| Ciliary body involvement | |
| Yes | 36 (31%) |
| No | 81 (69%) |
| Extra-ocular extension | |
| Yes | 9 (8%) |
| No | 108 (92%) |
| Epithelioid cells | |
| Yes | 50 (43%) |
| No | 67 (57%) |
| Closed loops present | |
| Yes | 47 (40%) |
| No | 43 (37%) |
| Not assessed | 27 (23%) |
| Necrosis | |
| Yes | 21 (17%) |
| No | 68 (59%) |
| Not assessed | 28 (24%) |
| Mitotic count per 40 high power field | |
| Median | 5 (1–72) |
| Primary Management | |
| Enucleation | 78/117 (66%) |
| Local Resection | 12/117 (10%) |
| Endoresection | 1/117 (1%) |
| Proton Beam RXT | 10/117 (9%) |
| Ruthenium Plaque RXT | 16/117 (14%) |
| Variable | Sig. | Hazard Ratio (HR) | 95.0% CI for HR | |
|---|---|---|---|---|
| Lower | Upper | |||
| Age at PM | 0.605 | 1.006 | 0.983 | 1.031 |
| LBD | ≤0.001 | 1.229 | 1.107 | 1.365 |
| UH | ≤0.001 | 1.198 | 1.086 | 1.322 |
| CBI | 0.003 | 2.602 | 1.396 | 4.849 |
| EOE | 0.183 | 2.024 | 0.717 | 5.715 |
| Epithelioid | 0.001 | 4.552 | 1.910 | 10.850 |
| Chr 3 | ≤0.001 | 9.236 | 3.602 | 23.683 |
| Extra copies 8 | 0.018 | 2.519 | 1.174 | 5.406 |
| SF3B1 | 0.131 | 0.486 | 0.190 | 1.241 |
| BAP1 | ≤0.001 | 6.536 | 3.095 | 13.804 |
| EIF1AX | 0.029 | 0.269 | 0.830 | 0.875 |
| Variable | Sig. | Hazard Ratio (HR) | 95.0% CI for HR | |
|---|---|---|---|---|
| Lower | Upper | |||
| UH | 0.016 | 1.124 | 1.022 | 1.235 |
| Chr3 | ≤0.001 | 5.949 | 2.226 | 15.898 |
| Epithelioid | 0.059 | 2.375 | 0.969 | 5.825 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thornton, S.; Coupland, S.E.; Olohan, L.; Sibbring, J.S.; Kenny, J.G.; Hertz-Fowler, C.; Liu, X.; Haldenby, S.; Heimann, H.; Hussain, R.; et al. Targeted Next-Generation Sequencing of 117 Routine Clinical Samples Provides Further Insights into the Molecular Landscape of Uveal Melanoma. Cancers 2020, 12, 1039. https://doi.org/10.3390/cancers12041039
Thornton S, Coupland SE, Olohan L, Sibbring JS, Kenny JG, Hertz-Fowler C, Liu X, Haldenby S, Heimann H, Hussain R, et al. Targeted Next-Generation Sequencing of 117 Routine Clinical Samples Provides Further Insights into the Molecular Landscape of Uveal Melanoma. Cancers. 2020; 12(4):1039. https://doi.org/10.3390/cancers12041039
Chicago/Turabian StyleThornton, Sophie, Sarah E. Coupland, Lisa Olohan, Julie S. Sibbring, John G. Kenny, Christiane Hertz-Fowler, Xuan Liu, Sam Haldenby, Heinrich Heimann, Rumana Hussain, and et al. 2020. "Targeted Next-Generation Sequencing of 117 Routine Clinical Samples Provides Further Insights into the Molecular Landscape of Uveal Melanoma" Cancers 12, no. 4: 1039. https://doi.org/10.3390/cancers12041039
APA StyleThornton, S., Coupland, S. E., Olohan, L., Sibbring, J. S., Kenny, J. G., Hertz-Fowler, C., Liu, X., Haldenby, S., Heimann, H., Hussain, R., Kipling, N., Taktak, A., & Kalirai, H. (2020). Targeted Next-Generation Sequencing of 117 Routine Clinical Samples Provides Further Insights into the Molecular Landscape of Uveal Melanoma. Cancers, 12(4), 1039. https://doi.org/10.3390/cancers12041039





