Discovery of Pancreatic Adenocarcinoma Biomarkers by Untargeted Metabolomics
Abstract
1. Introduction
2. Results
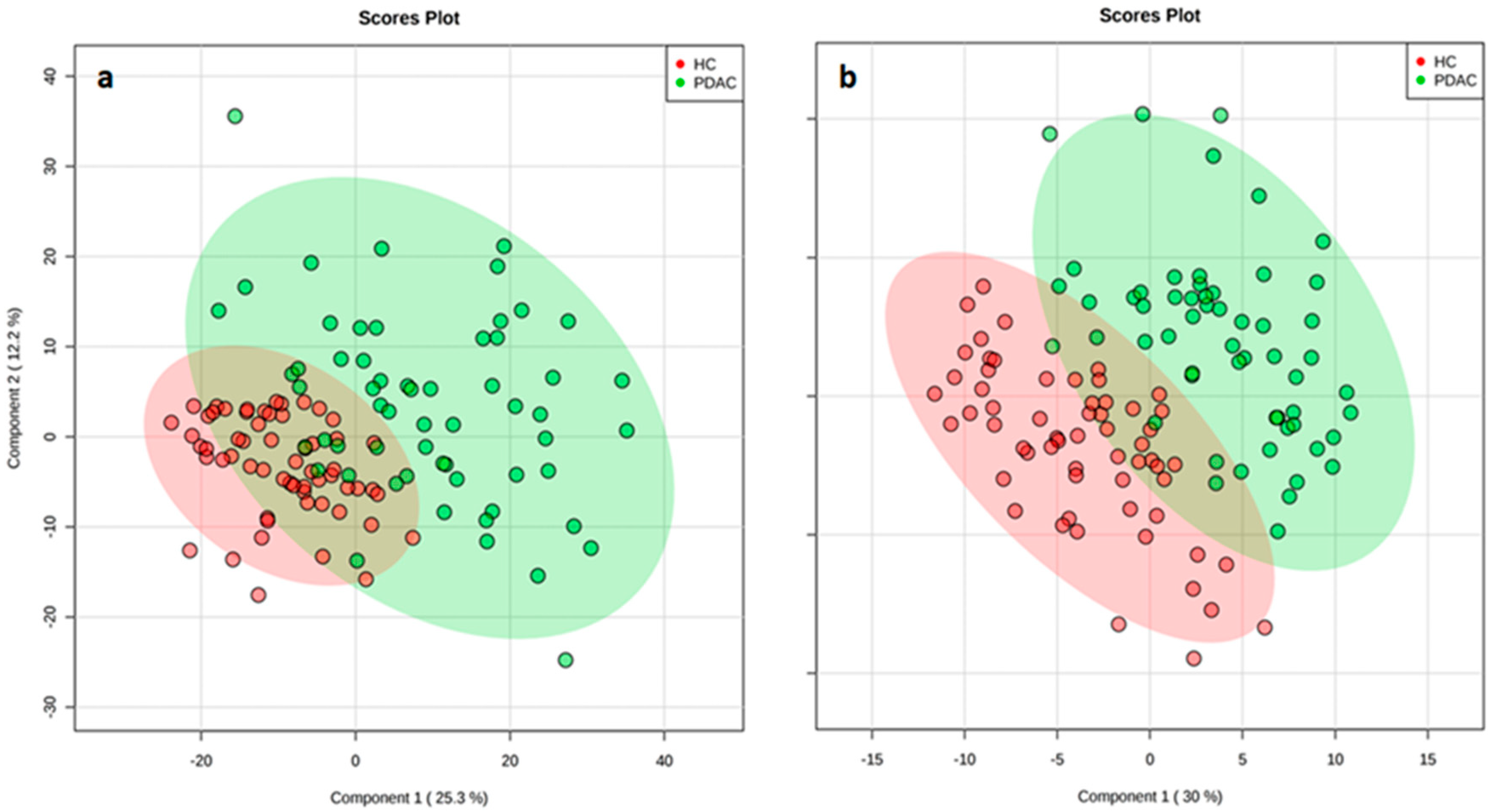
2.1. LC-HRMS Analysis
2.2. Pathway Analysis
2.3. Biomarker Evaluation
3. Discussion
4. Materials and Methods
4.1. Sample Collection
4.2. Metabolite Extraction
4.3. LC-HRMS Analysis
4.4. Data Set Creation
4.5. Normalization and Analytical Validation
4.6. Statistical Analysis
4.7. Biomarker Evaluation
4.8. Biomarker Identification
4.9. Pathway Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Pourshams, A.; Sepanlou, S.G.; Ikuta, K.S.; Bisignano, C.; Safiri, S.; Roshandel, G.; Sharif, M.; Khatibian, M.; Fitzmaurice, C.; Nixon, M.R.; et al. The global, regional, and national burden of pancreatic cancer and its attributable risk factors in 195 countries and territories, 1990–2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol. Hepatol. 2019, 4, 934–947. [Google Scholar] [CrossRef]
- McGuigan, A.; Kelly, P.; Turkington, R.C.; Jones, C.; Coleman, H.G.; McCain, R.S. Pancreatic cancer: A review of clinical diagnosis, epidemiology, treatment and outcomes. World J. Gastroenterol. 2018, 24, 4846–4861. [Google Scholar] [CrossRef]
- Satyananda, V.; Gupta, R.; Hari, D.M.; Yeh, J.; Chen, K.T. Advances in Translational Research and Clinical Care in Pancreatic Cancer: Where Are We Headed? Gastroenterol. Res. Pract. 2019, 2019, 7690528. [Google Scholar] [CrossRef]
- Brunner, M.; Wu, Z.; Krautz, C.; Pilarsky, C.; Grützmann, R.; Weber, G.F. Current Clinical Strategies of Pancreatic Cancer Treatment and Open Molecular Questions. Int. J. Mol. Sci. 2019, 20, 4543. [Google Scholar] [CrossRef]
- Kleeff, J.; Korc, M.; Apte, M.; La Vecchia, C.; Johnson, C.D.; Biankin, A.V.; Neale, R.E.; Tempero, M.; Tuveson, D.A.; Hruban, R.H.; et al. Pancreatic cancer. Nat. Rev. Dis. Primers 2016, 2, 16022. [Google Scholar] [CrossRef]
- Schrimpe-Rutledge, A.C.; Codreanu, S.G.; Sherrod, S.D.; McLean, J.A. Untargeted Metabolomics Strategies-Challenges and Emerging Directions. J. Am. Soc. Mass Spectrom. 2016, 27, 1897–1905. [Google Scholar] [CrossRef]
- Roig, B.; Rodríguez-Balada, M.; Samino, S.; Lam, E.W.; Guaita-Esteruelas, S.; Gomes, A.R.; Correig, X.; Borràs, J.; Yanes, O.; Gumà, J. Metabolomics reveals novel blood plasma biomarkers associated to the BRCA1-mutated phenotype of human breast cancer. Sci. Rep. 2017, 7, 17831. [Google Scholar] [CrossRef]
- Cheung, P.K.; Ma, M.H.; Tse, H.F.; Yeung, K.F.; Tsang, H.F.; Chu, M.K.M.; Kan, C.M.; Cho, W.C.S.; Ng, L.B.W.; Chan, L.W.C. The applications of metabolomics in the molecular diagnostics of cancer. Expert Rev. Mol. Diagn. 2019, 19, 785–793. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.A.; Spellman, D.S. Mass Spectrometry-Based Biomarkers in Drug Development. In Advancements of Mass Spectrometry in Biomedical Research; Woods, A.G., Darie, C.C., Eds.; Springer: New York, NY, USA, 2014; pp. 341–359. [Google Scholar]
- Kamphorst, J.J.; Nofal, M.; Commisso, C.; Hackett, S.R.; Lu, W.; Grabocka, E.; Vander Heiden, M.G.; Miller, G.; Drebin, J.A.; Bar-Sagi, D.; et al. Human pancreatic cancer tumors are nutrient poor and tumor cells actively scavenge extracellular protein. Cancer Res. 2015, 75, 544–553. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Tong, Z. Clinical Application of Metabolomics in Pancreatic Diseases: A Mini-Review. Lab. Med. 2020, 51, 116–121. [Google Scholar] [CrossRef] [PubMed]
- Morin, A.; Letouzé, E.; Gimenez-Roqueplo, A.P.; Favier, J. Oncometabolites-driven tumorigenesis: From genetics to targeted therapy. Int. J. Cancer 2014, 135, 2237–2248. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Nishiumi, S.; Ikeda, A.; Yoshie, T.; Sakai, A.; Matsubara, A.; Izumi, Y.; Tsumura, H.; Tsuda, M.; Nishisaki, H.; et al. A novel serum metabolomics-based diagnostic approach to pancreatic cancer. Cancer Epidemiol. Biomarkers Prev. 2013, 22, 571–579. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Jin, H.; Guo, X.; Yang, Z.; Zhao, L.; Tang, S.; Mo, P.; Wu, K.; Nie, Y.; Pan, Y.; et al. Distinguishing pancreatic cancer from chronic pancreatitis and healthy individuals by (1)H nuclear magnetic resonance-based metabonomic profiles. Clin. Biochem. 2012, 45, 1064–1109. [Google Scholar] [CrossRef] [PubMed]
- Michálková, L.; Horník, Š.; Sýkora, J.; Habartová, L.; Setnička, V. Diagnosis of pancreatic cancer via1H NMR metabolomics of human plasma. Analyst 2018, 143, 5974–5978. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Zhou, J.; Yuan, C.; Zhang, L.; Li, D.; Si, D.; Xiu, D.; Zhong, L. Metabolomics identifies serum and exosomes metabolite markers of pancreatic cancer. Metabolomics 2019, 15, 86. [Google Scholar] [CrossRef] [PubMed]
- Rawla, P.; Sunkara, T.; Gaduputi, V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J. Oncol. 2019, 10, 10–27. [Google Scholar] [CrossRef] [PubMed]
- Rajamani, D.; Bhasin, M.K. Identification of key regulators of pancreatic cancer progression through multidimensional systems-level analysis. Genome Med. 2016, 8, 38. [Google Scholar] [CrossRef]
- Kwon, M.S.; Kim, Y.; Lee, S.; Namkung, J.; Yun, T.; Yi, S.G.; Han, S.; Kang, M.; Kim, S.W.; Jang, J.Y.; et al. Integrative analysis of multi-omics data for identifying multi-markers for diagnosing pancreatic cancer. BMC Genom. 2015, 16, S4. [Google Scholar] [CrossRef]
- Kato, Y.; Maeda, T.; Suzuki, A.; Baba, Y. Cancer metabolism: New insights into classic characteristics. Jpn. Dent. Sci. Rev. 2018, 54, 8–21. [Google Scholar] [CrossRef]
- Schmidt-Hansen, M.; Berendse, S.; Hamilton, W. Symptoms of Pancreatic Cancer in Primary Care: A Systematic Review. Pancreas 2016, 45, 814–818. [Google Scholar] [CrossRef]
- Fest, J.; Vijfhuizen, L.S.; Goeman, J.J.; Veth, O.; Joensuu, A.; Perola, M.; Männistö, S.; Ness-Jensen, E.; Hveem, K.; Haller, T.; et al. Search for Early Pancreatic Cancer Blood Biomarkers in Five European Prospective Population Biobanks Using Metabolomics. Endocrinology 2019, 160, 1731–1742. [Google Scholar] [CrossRef] [PubMed]
- Kamphorst, J.J.; Cross, J.R.; Fan, J.; de Stanchina, E.; Mathew, R.; White, E.P.; Thompson, C.B.; Rabinowitz, J.D. Hypoxic and Ras-transformed cells support growth by scavenging unsaturated fatty acids from lysophospholipids. Proc. Natl. Acad. Sci. USA 2013, 110, 8882–8887. [Google Scholar] [CrossRef] [PubMed]
- Kühn, T.; Floegel, A.; Sookthai, D.; Johnson, T.; Rolle-Kampczyk, U.; Otto, W.; von Bergen, M.; Boeing, H.; Kaaks, R. Higher plasma levels of lysophosphatidylcholine 18:0 are related to a lower risk of common cancers in a prospective metabolomics study. BMC Med. 2016, 14, 13. [Google Scholar] [CrossRef] [PubMed]
- Fahrmann, J.F.; Bantis, L.E.; Capello, M.; Scelo, G.; Dennison, J.B.; Patel, N.; Murage, E.; Vykoukal, J.; Kundnani, D.L.; Foretova, L.; et al. A Plasma-Derived Protein-Metabolite Multiplexed Panel for Early-Stage Pancreatic Cancer. J. Natl. Cancer Inst. 2019, 111, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, S.A.; Akita, H.; Takemasa, I.; Eguchi, H.; Pastural, E.; Nagano, H.; Monden, M.; Doki, Y.; Mori, M.; Jin, W.; et al. Metabolic system alterations in pancreatic cancer patient serum: Potential for early detection. BMC Cancer 2013, 13, 416. [Google Scholar] [CrossRef] [PubMed]
- Akita, H.; Ritchie, S.A.; Takemasa, I.; Eguchi, H.; Pastural, E.; Jin, W.; Yamazaki, Y.; Goodenowe, D.B.; Nagano, H.; Monden, M.; et al. Serum Metabolite Profiling for the Detection of Pancreatic Cancer: Results of a Large Independent Validation Study. Pancreas 2016, 45, 1418–1423. [Google Scholar] [CrossRef]
- Xie, G.; Lu, L.; Qiu, Y.; Ni, Q.; Zhang, W.; Gao, Y.T.; Risch, H.A.; Yu, H.; Jia, W. Plasma metabolite biomarkers for the detection of pancreatic cancer. J. Proteome Res. 2015, 14, 1195–1202. [Google Scholar] [CrossRef]
- Frolkis, A.; Knox, C.; Lim, E.; Jewison, T.; Law, V.; Hau, D.D.; Liu, P.; Gautam, B.; Ly, S.; Guo, A.C.; et al. SMPDB: The Small Molecule Pathway Database. Nucleic Acids Res. 2010, 38, 480–487. [Google Scholar] [CrossRef]
- Hardie, R.A.; van Dam, E.; Cowley, M.; Han, T.L.; Balaban, S.; Pajic, M.; Pinese, M.; Iconomou, M.; Shearer, R.F.; McKenna, J.; et al. Mitochondrial mutations and metabolic adaptation in pancreatic cancer. Cancer Metab. 2017, 5, 2. [Google Scholar] [CrossRef]
- Urayama, S.; Zou, W.; Brooks, K.; Tolstikov, V. Comprehensive mass spectrometry based metabolic profiling of blood plasma reveals potent discriminatory classifiers of pancreatic cancer. Rapid Commun. Mass Spectrom. 2010, 24, 613–620. [Google Scholar] [CrossRef]
- Di Gangi, I.M.; Mazza, T.; Fontana, A.; Copetti, M.; Fusilli, C.; Ippolito, A.; Mattivi, F.; Latiano, A.; Andriulli, A.; Vrhovsek, U.; et al. Metabolomic profile in pancreatic cancer patients: A consensus-based approach to identify highly discriminating metabolites. Oncotarget 2016, 7, 5815–5829. [Google Scholar] [CrossRef] [PubMed]
- Beloribi-Djefaflia, S.; Vasseur, S.; Guillaumond, F. Lipid metabolic reprogramming in cancer cells. Oncogenesis 2016, 5, e189. [Google Scholar] [CrossRef] [PubMed]
- Kraft, M.; Kraft, K.; Gärtner, S.; Mayerle, J.; Simon, P.; Weber, E.; Schütte, K.; Stieler, J.; Koula-Jenik, H.; Holzhauer, P.; et al. L-Carnitine-supplementation in advanced pancreatic cancer (CARPAN)—A randomized multicentre trial. Nutr. J. 2012, 11, 52. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Zhou, J.; Hooi, S.C.; Jiang, Y.M.; Lu, G.D. Fatty acid activation in carcinogenesis and cancer development: Essential roles of long-chain acyl-CoA synthetases. Oncol. Lett. 2018, 16, 1390–1396. [Google Scholar] [CrossRef] [PubMed]
- Feng, H.Y.; Chen, Y.C. Role of bile acids in carcinogenesis of pancreatic cancer: An old topic with new perspective. World J. Gastroenterol. 2016, 22, 7463–7477. [Google Scholar] [CrossRef]
- Lindahl, A.; Heuchel, R.; Forshed, J.; Lehtiö, J.; Löhr, M.; Nordström, A. Discrimination of pancreatic cancer and pancreatitis by LC-MS metabolomics. Metabolomics 2017, 13, 61. [Google Scholar] [CrossRef]
- Muscarella, P.; Boros, L.G.; Fisher, W.E.; Rink, C.; Melvin, W.S. Oral Dehydroepiandrosterone Inhibits the Growth of Human Pancreatic Cancer in Nude Mice. J. Surg. Res. 1998, 79, 154–157. [Google Scholar] [CrossRef]
- Melvin, W.S.; Boros, L.G.; Muscarella, P.; Brandes, J.L.; Johnson, J.A.; Fisher, W.E.; Schirmer, W.J.; Ellison, E.C. Dehydroepiandrosterone-sulfate inhibits pancreatic carcinoma cell proliferation in vitro and in vivo. Surgery 1997, 121, 392–397. [Google Scholar] [CrossRef]
- Chen, M.C.; Hsu, S.L.; Lin, H.; Yang, T.Y. Retinoic acid and cancer treatment. Biomedicine 2014, 4, 22. [Google Scholar] [CrossRef]
- Del Boccio, P.; Perrotti, F.; Rossi, C.; Cicalini, I.; Di Santo, S.; Zucchelli, M.; Sacchetta, P.; Genovesi, D.; Pieragostino, D. Serum lipidomic study reveals potential early biomarkers for predicting response to chemoradiation therapy in advanced rectal cancer: A pilot study. Adv. Radiat. Oncol. 2017, 2, 118–124. [Google Scholar] [CrossRef]
- Lee, J.H.; Yu, S.E.; Kim, K.H.; Yu, M.H.; Jeong, I.H.; Cho, J.Y.; Park, S.J.; Lee, W.J.; Han, S.S.; Kim, T.H.; et al. Individualized metabolic profiling stratifies pancreatic and biliary tract cancer: A useful tool for innovative screening programs and predictive strategies in healthcare. EPMA J. 2018, 9, 287–297. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.Z.; Hennig, R.; Adrian, T.E. Lipoxygenase and cyclooxygenase metabolism: New insights in treatment and chemoprevention of pancreatic cancer. Mol. Cancer 2003, 2, 10. [Google Scholar] [CrossRef] [PubMed]


| Ionization Mode | Total | Monoisotopics | After Organic Solvent (OS) Exclusion | RSD | Evaluated in PCA | R2 | Q2 |
|---|---|---|---|---|---|---|---|
| ESI+ | 1150 | 365 | 145 | 84 | 84 | 0.73 | 0.61 |
| ESI- | 966 | 345 | 152 | 134 | 134 | 0.73 | 0.66 |
| Altered Pathways | P 1 |
|---|---|
| Linoleicacidmetabolism | 5.301 × 10−3 |
| Glycerolipidmetabolism | 5.664 × 10−3 |
| Glycerophospholipidmetabolism | 9.377 × 10−3 |
| Primarybileacidbiosynthesis | 2.190 × 10−2 |
| ESImode | m/z | RT (min) | p(FDR) | PDAC/HC | AUC | Tentativeidentification |
|---|---|---|---|---|---|---|
| + | 646.4145 | 7.65 | 1.65 × 10−15 | ↑ | 0.959 | PS(12:0/15:1) |
| - | 446.3760 | 9.44 | 3.75 × 10−14 | ↓ | 0.902 | TG(22:2/15:0/18:3) |
| - | 627.3741 | 3.53 | 7.46 × 10−15 | ↑ | 0.900 | 4-oxo-Retinoic acid |
| - | 369.1747 | 3.29 | 2.71 × 10−13 | ↓ | 0.878 | Androsterone sulfate |
| - | 476.2792 | 5.08 | 4.65 × 10−12 | ↓ | 0.859 | LysoPE(18:2) |
| - | 311.1396 | 2.62 | 6.27 × 10−17 | ↓ | 0.858 | Phenylalanylphenylalanine |
| + | 430.2939 | 7.58 | 7.08 × 10−10 | ↑ | 0.850 | all-trans-Decaprenyldiphosphate |
| + | 1039.6721 | 10.79 | 1.73 × 10−9 | ↓ | 0.848 | LysoPC(18:2) |
| - | 367.1583 | 3.14 | 1.79 × 10−9 | ↓ | 0.847 | Dehydroepiandrosterone sulfate |
| Characteristic | PDAC Patients | Healthy Controls |
|---|---|---|
| Age (years ± SD) | 61.18 ± 12.17 | 56.16 ± 10.03 |
| Sex | ||
| Male | 32 | 31 |
| Female | 27 | 29 |
| BMI (kg/m2 ± SD) | 25.40 ± 3.58 | 27.17 ± 3.76 |
| Pancreatitis | ||
| Yes | 0 | 0 |
| No | 59 | 60 |
| Diabetes mellitus | ||
| No | 45 | 60 |
| Type I | 0 | 0 |
| Type II | 14 | 0 |
| Type IIIc | 0 | 0 |
| Hypercholesterolemia | ||
| Yes | 15 | 0 |
| No | 44 | 60 |
| Statin and/or fibrate intake | ||
| Yes | 8 | 0 |
| No | 51 | 60 |
| Cachexia | ||
| Yes | 0 | 0 |
| No | 59 | 60 |
| Previous surgeries | ||
| Yes | 0 | 0 |
| No | 59 | 60 |
| Tumor stage | ||
| I | 5 | - |
| II | 6 | - |
| III | 12 | - |
| IV | 36 | - |
| Site of primary tumor | ||
| Head | 36 | - |
| Body | 15 | - |
| Tail | 8 | - |
| Icteric | ||
| Yes | 18 | - |
| No | 41 | - |
| Metastatic | ||
| Yes | 33 | - |
| No | 26 | - |
| Number of metastases | ||
| 1 | 2 | - |
| 2 | 2 | - |
| >2 | 29 | - |
| Metastatic site | ||
| Hepatic | 26 | - |
| Lymph node | 11 | - |
| Lung | 7 | - |
| Osseous | 4 | - |
| Brain | 2 | - |
| Other | 15 | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martín-Blázquez, A.; Jiménez-Luna, C.; Díaz, C.; Martínez-Galán, J.; Prados, J.; Vicente, F.; Melguizo, C.; Genilloud, O.; Pérez del Palacio, J.; Caba, O. Discovery of Pancreatic Adenocarcinoma Biomarkers by Untargeted Metabolomics. Cancers 2020, 12, 1002. https://doi.org/10.3390/cancers12041002
Martín-Blázquez A, Jiménez-Luna C, Díaz C, Martínez-Galán J, Prados J, Vicente F, Melguizo C, Genilloud O, Pérez del Palacio J, Caba O. Discovery of Pancreatic Adenocarcinoma Biomarkers by Untargeted Metabolomics. Cancers. 2020; 12(4):1002. https://doi.org/10.3390/cancers12041002
Chicago/Turabian StyleMartín-Blázquez, Ariadna, Cristina Jiménez-Luna, Caridad Díaz, Joaquina Martínez-Galán, Jose Prados, Francisca Vicente, Consolación Melguizo, Olga Genilloud, José Pérez del Palacio, and Octavio Caba. 2020. "Discovery of Pancreatic Adenocarcinoma Biomarkers by Untargeted Metabolomics" Cancers 12, no. 4: 1002. https://doi.org/10.3390/cancers12041002
APA StyleMartín-Blázquez, A., Jiménez-Luna, C., Díaz, C., Martínez-Galán, J., Prados, J., Vicente, F., Melguizo, C., Genilloud, O., Pérez del Palacio, J., & Caba, O. (2020). Discovery of Pancreatic Adenocarcinoma Biomarkers by Untargeted Metabolomics. Cancers, 12(4), 1002. https://doi.org/10.3390/cancers12041002





