MiR-215-5p Reduces Liver Metastasis in an Experimental Model of Colorectal Cancer through Regulation of ECM-Receptor Interactions and Focal Adhesion
Abstract
:Simple Summary
Abstract
1. Introduction
2. Results
2.1. Selection of Stably Transfected Clones
2.2. MiR-215-5p Inhibits the Clonogenic Potential, Arrests the Cell Cycle, and Induce Apoptosis of CRC Cells
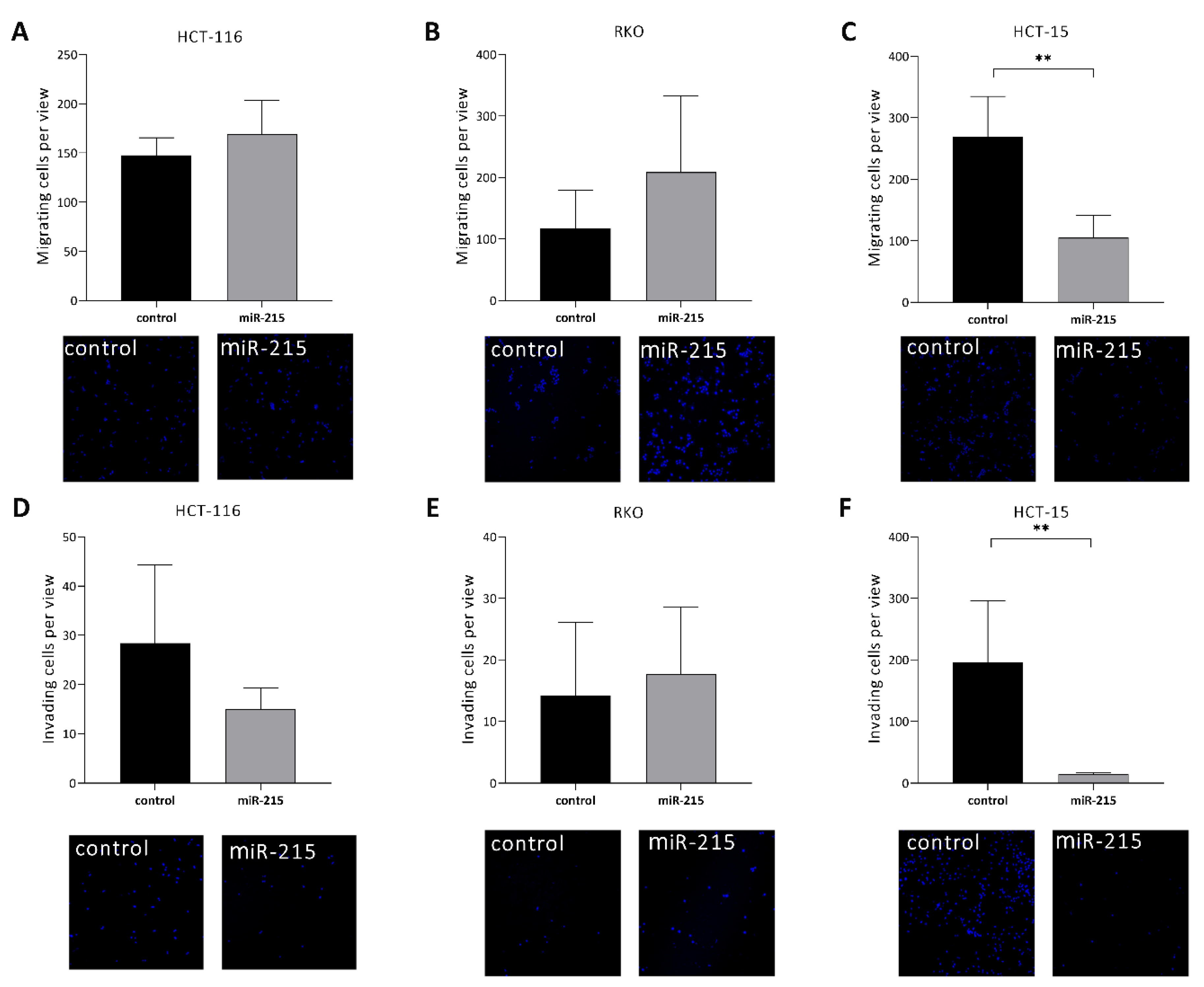
2.3. MiR-215-5p Inhibits Cell Migration/Invasion of HCT-15 Cells In Vitro
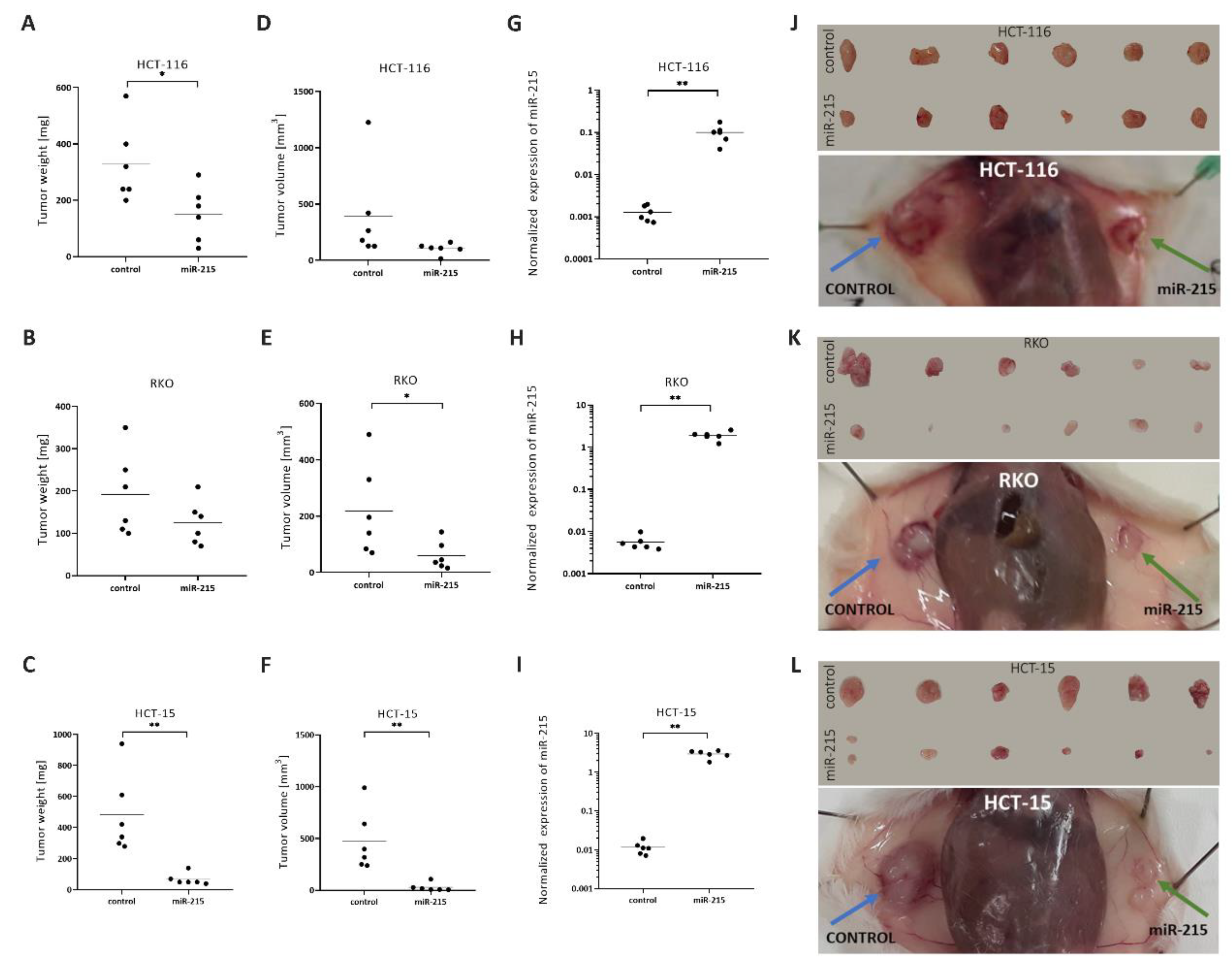
2.4. MiR-215-5p Inhibits CRC Cells Tumor Growth In Vivo
2.5. MiR-215-5p Inhibits the Metastatic Potential of CRC Cells In Vivo
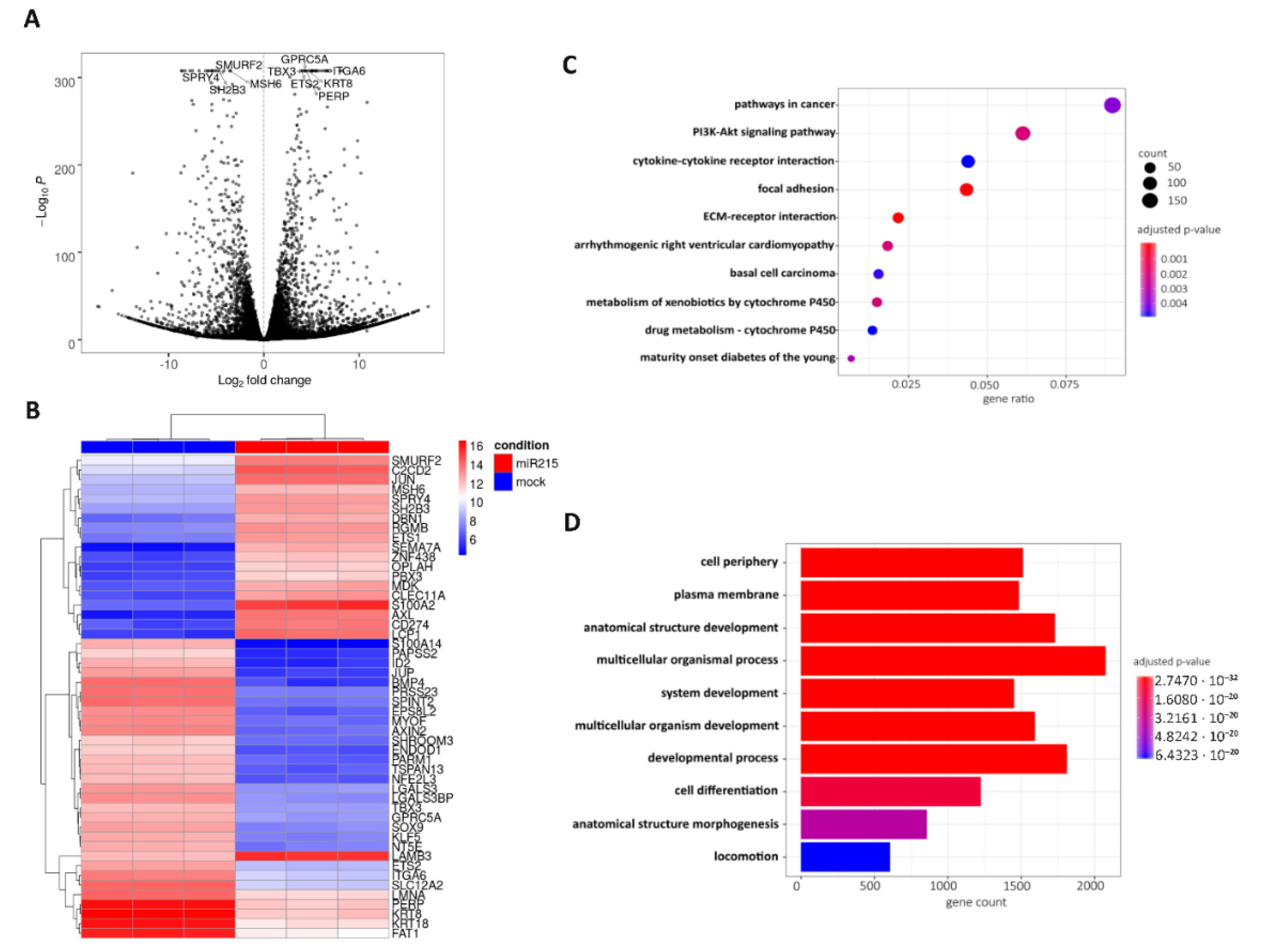
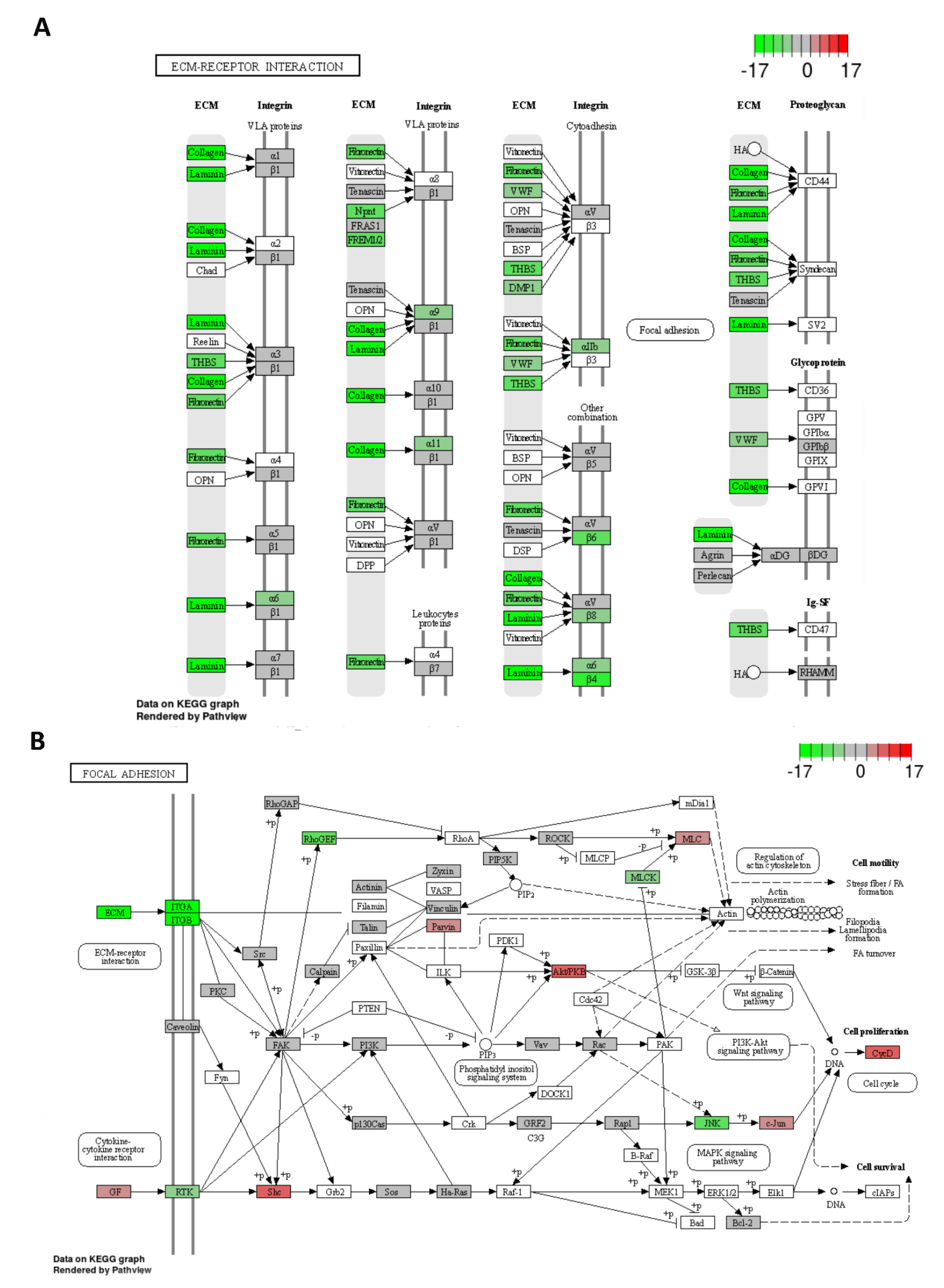
2.6. Possible Molecular Mechanism of Metastatic Process Regulated by miR-215-5p via Involvement of ECM-Receptor Interactions and Focal Adhesion Pathways
2.7. Validation of Potential miR-215-5p Targets
3. Discussion
4. Materials and Methods
4.1. Cell Lines and Cell Culture
4.2. Patient Samples
4.3. Stable Transfection of miR-215
4.4. Colony Forming Aassay
4.5. Cell Cycle, Apoptosis and Cell Death analysis
4.6. Transwell Migration and Invasiveness Assay
4.7. Animal Models
4.8. Tumorigenicity Assay
4.9. Intra-Splenic Metastatic Model
4.10. RNA Isolation
4.11. Reverse Transcription and Quantitative Real-Time PCR
4.12. Transcriptome Profiling
4.13. Statistical Analysis
4.14. Processing of RNA-Seq Data
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
List of Abbreviations
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- A Issa, I.; Noureddine, M. Colorectal cancer screening: An updated review of the available options. World J. Gastroenterol. 2017, 23, 5086–5096. [Google Scholar] [CrossRef]
- Engstrand, J.; Nilsson, H.; Strömberg, C.; Jonas, E.; Freedman, J. Colorectal cancer liver metastases—A population-based study on incidence, management and survival. BMC Cancer 2018, 18, 78. [Google Scholar] [CrossRef] [PubMed]
- Balchen, V.; Simon, K. Colorectal cancer development and advances in screening. Clin. Interv. Aging 2016, 11, 967–976. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Molinari, C.; Marisi, G.; Passardi, A.; Matteucci, L.; De Maio, G.; Ulivi, P. Heterogeneity in Colorectal Cancer: A Challenge for Personalized Medicine? Int. J. Mol. Sci. 2018, 19, 3733. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lai, E.C. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat. Genet. 2002, 30, 363–364. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Lan, Z.; Xiong, X.; Ao, H.; Feng, Y.; Gu, H.; Yu, M.; Cui, Q. The Dual Role of MicroRNAs in Colorectal Cancer Progression. Int. J. Mol. Sci. 2018, 19, 2791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khella, H.; Bakhet, M.; Allo, G.; Jewett, M.; Girgis, A.; Latif, A.; Von Both, I.; Bjarnason, G.; Yousef, G.M. miR-192, miR-194 and miR-215: A convergent microRNA network suppressing tumor progression in renal cell carcinoma. Carcinogenesis 2013, 34, 2231–2239. [Google Scholar] [CrossRef]
- Braun, C.J.; Zhang, X.; Savelyeva, I.; Wolff, S.; Moll, U.M.; Schepeler, T.; Ørntoft, T.F.; Andersen, C.L.; Dobbelstein, M. p53—Responsive MicroRNAs 192 and 215 Are Capable of Inducing Cell Cycle Arrest. Cancer Res. 2008, 68, 10094–10104. [Google Scholar] [CrossRef] [Green Version]
- Jones, M.F.; Hara, T.; Francis, P.; Li, X.L.; Bilke, S.; Zhu, Y.; Pineda, M.; Subramanian, M.; Bodmer, W.; Lal, A. The CDX1-microRNA-215 axis regulates colorectal cancer stem cell differentiation. Proc. Natl. Acad. Sci. USA 2015, 112, E1550–E1558. [Google Scholar] [CrossRef] [Green Version]
- Lu, H.; Lei, X.; Liu, J.; Klaassen, C. Regulation of hepatic microRNA expression by hepatocyte nuclear factor 4 alpha. World J. Hepatol. 2017, 9, 191–208. [Google Scholar] [CrossRef]
- Mechtler, P.; Singhal, R.; Kichina, J.V.; Bard, J.E.; Buck, M.J.; Kandel, E.S. MicroRNA analysis suggests an additional level of feedback regulation in the NF-κB signaling cascade. Oncotarget 2015, 6, 17097–17106. [Google Scholar] [CrossRef]
- Hu, J.; Sun, T.; Wang, H.; Chen, Z.; Wang, S.; Yuan, L.; Liu, T.; Li, H.-R.; Wang, P.; Feng, Y.; et al. MiR-215 Is Induced Post-transcriptionally via HIF-Drosha Complex and Mediates Glioma-Initiating Cell Adaptation to Hypoxia by Targeting KDM1B. Cancer Cell 2016, 29, 49–60. [Google Scholar] [CrossRef] [Green Version]
- Chen, D.-L.; Lu, Y.-X.; Zhang, J.-X.; Wei, X.-L.; Wang, F.-H.; Zeng, Z.-L.; Pan, Z.-Z.; Yuan, Y.-F.; Pelicano, H.; Chiao, P.J.; et al. Long non-coding RNA UICLM promotes colorectal cancer liver metastasis by acting as a ceRNA for microRNA-215 to regulate ZEB2 expression. Theranostics 2017, 7, 4836–4849. [Google Scholar] [CrossRef]
- Kong, X.; Duan, Y.; Sang, Y.; Li, Y.; Zhang, H.; Liang, Y.; Liu, Y.; Zhang, N.; Yang, Q. LncRNA-CDC6 promotes breast cancer progression and function as ceRNA to target CDC6 by sponging microRNA-215. J. Cell. Physiol. 2019, 234, 9105–9117. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, J.; Chen, X.; Xu, X.; Cao, G.; Li, H.; Wu, T. LncRNA FTX sponges miR-215 and inhibits phosphorylation of vimentin for promoting colorectal cancer progression. Gene Ther. 2018, 25, 321–330. [Google Scholar] [CrossRef]
- Vychytilova-Faltejskova, P.; Slaby, O. MicroRNA-215: From biology to theranostic applications. Mol. Asp. Med. 2019, 70, 72–89. [Google Scholar] [CrossRef]
- Necela, B.M.; Carr, J.M.; Asmann, Y.W.; Thompson, E.A. Differential Expression of MicroRNAs in Tumors from Chronically Inflamed or Genetic (APCMin/+) Models of Colon Cancer. PLoS ONE 2011, 6, e18501. [Google Scholar] [CrossRef] [Green Version]
- Faltejskova, P.; Svoboda, M.; Srutova, K.; Mlcochova, J.; Besse, A.; Nekvindova, J.; Radova, L.; Fabian, P.; Slaba, K.; Kiss, I.; et al. Identification and functional screening of microRNAs highly deregulated in colorectal cancer. J. Cell. Mol. Med. 2012, 16, 2655–2666. [Google Scholar] [CrossRef]
- Vychytilova-Faltejskova, P.; Merhautova, J.; Machackova, T.; Gutierrez-Garcia, I.; Garcia-Solano, J.; Radova, L.; Brchnelova, D.; Slaba, K.; Svoboda, M.; Halamkova, J.; et al. MiR-215-5p is a tumor suppressor in colorectal cancer targeting EGFR ligand epiregulin and its transcriptional inducer HOXB9. Oncogenesis 2017, 6, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Li, S.; Gao, J.; Gu, J.; Yuan, J.; Hua, D.; Shen, L. MicroRNA-215 inhibits relapse of colorectal cancer patients following radical surgery. Med. Oncol. 2013, 30, 549. [Google Scholar] [CrossRef]
- Hou, Y.; Zhen, J.; Xu, X.; Zhen, K.; Zhu, B.; Pan, R.; Zhao, C. miR-215 functions as a tumor suppressor and directly targets ZEB2 in human non-small cell lung cancer. Oncol. Lett. 2015, 10, 1985–1992. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, Y.; Shen, H.; Zhou, Y.; Yang, Z.; Hu, T. MicroRNA-215 suppresses the proliferation, migration and invasion of non-small cell lung carcinoma cells through the downregulation of matrix metalloproteinase-16 expression. Exp. Ther. Med. 2018, 15, 3239–3246. [Google Scholar] [CrossRef] [Green Version]
- Gao, J.B.; Zhu, M.N.; Zhu, X.L. miRNA-215-5p suppresses the aggressiveness of breast cancer cells by targeting Sox9. FEBS Open Bio. 2019, 9, 1957–1967. [Google Scholar] [CrossRef] [Green Version]
- Yao, J.; Zhang, P.; Li, J.; Xu, W. MicroRNA-215 acts as a tumor suppressor in breast cancer by targeting AKT serine/threonine kinase 1. Oncol. Lett. 2017, 14, 1097–1104. [Google Scholar] [CrossRef] [Green Version]
- Singh, A.; Bhattacharyya, N.; Srivastava, A.; Pruett, N.; Ripley, R.T.; Schrump, D.S.; Hoang, C.D. MicroRNA-215-5p Treatment Suppresses Mesothelioma Progression via the MDM2-p53-Signaling Axis. Mol. Ther. 2019, 27, 1665–1680. [Google Scholar] [CrossRef]
- Ren, Y.; Shang, J.; Li, J.; Liu, W.; Zhang, Z.; Yuan, J.; Yang, M. The long noncoding RNA PCAT-1 links the microRNA miR-215 to oncogene CRKL-mediated signaling in hepatocellular carcinoma. J. Biol. Chem. 2017, 292, 17939–17949. [Google Scholar] [CrossRef] [Green Version]
- Wei, Y.; Sun, J.; Li, X. MicroRNA-215 enhances invasion and migration by targeting retinoblastoma tumor suppressor gene 1 in high-grade glioma. Biotechnol. Lett. 2016, 39, 197–205. [Google Scholar] [CrossRef]
- Chen, Z.; Liu, K.; Li, L.; Chen, Y.; Du, S. miR-215 promotes cell migration and invasion of gastric cancer by targeting Retinoblastoma tumor suppressor gene 1. Pathol. Res. Pract. 2017, 213, 889–894. [Google Scholar] [CrossRef]
- Li, N.; Zhang, Q.-Y.; Zou, J.-L.; Li, Z.-W.; Tian, T.-T.; Dong, B.; Liu, X.-J.; Ge, S.; Zhu, Y.; Gao, J.; et al. miR-215 promotes malignant progression of gastric cancer by targeting RUNX1. Oncotarget 2016, 7, 4817–4828. [Google Scholar] [CrossRef] [Green Version]
- Mamdouh, S.; Khorshed, F.E.; Aboushousha, T.; Hamdy, H.; Diab, A.; Seleem, M.; Saber, M. Evaluation of Mir-224, Mir-215 and Mir-143 as Serum Biomarkers for HCV Associated Hepatocellular Carcinoma. Asian Pac. J. Cancer Prev. 2017, 18, 3167–3171. [Google Scholar] [PubMed]
- Karaayvaz, M.; Pal, T.; Song, B.; Zhang, C.; Georgakopoulos, P.; Mehmood, S.; Burke, S.; Shroyer, K.; Ju, J. Prognostic Significance of miR-215 in Colon Cancer. Clin. Color. Cancer 2011, 10, 340–347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Agostini, A.; Brunetti, M.; Davidson, B.; Tropé, C.G.; Eriksson, A.G.Z.; Heim, S.; Panagopoulos, I.; Micci, F. The microRNA miR-192/215 family is upregulated in mucinous ovarian carcinomas. Sci. Rep. 2018, 8, 11069. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Ding, Y.; Yao, J.; Wei, Z.; Jin, H.; Chen, C.; Feng, J.; Ying, R. miR-215 Inhibits Colorectal Cancer Cell Migration and Invasion via Targeting Stearoyl-CoA Desaturase. Comput. Math. Methods Med. 2020, 2020, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Mauvoisin, D.; Charfi, C.; Lounis, M.A.; Rassart, E.; Mounier, C. Decreasing stearoyl-CoA desaturase-1 expression inhibits βcatenin signaling in breast cancer cells. Cancer Sci. 2012, 104, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Jin, Y.; Xu, T.; Zhou, S.; Cui, M. MicroRNA-215 targets NOB1 and inhibits growth and invasion of epithelial ovarian cancer. Am. J. Transl. Res. 2017, 9, 466–477. [Google Scholar]
- Tong, Y.-Q.; Liu, B.; Zheng, H.-Y.; Gu, J.; Liu, H.; Li, F.; Tan, B.-H.; Hartman, M.; Song, C.; Li, Y. MiR-215, an activator of the CTNNBIP1/β-catenin pathway, is a marker of poor prognosis in human glioma. Oncotarget 2015, 6, 25024–25033. [Google Scholar] [CrossRef] [Green Version]
- Mu, J.; Pang, Q.; Guo, Y.-H.; Chen, J.-G.; Zeng, W.; Huang, Y.-J.; Zhang, J.; Feng, B. Functional Implications of MicroRNA-215 in TGF-β1-Induced Phenotypic Transition of Mesangial Cells by Targeting CTNNBIP1. PLoS ONE 2013, 8, e58622. [Google Scholar] [CrossRef] [Green Version]
- Senanayake, U.; Das, S.; Vesely, P.; Alzoughbi, W.; Fröhlich, L.F.; Chowdhury, P.; Leuschner, I.; Hoefler, G.; Guertl, B. miR-192, miR-194, miR-215, miR-200c and miR-141 are downregulated and their common target ACVR2B is strongly expressed in renal childhood neoplasms. Carcinogenesis 2012, 33, 1014–1021. [Google Scholar] [CrossRef] [Green Version]
- A Blair, S.; Kane, S.V.; Clayburgh, D.R.; Turner, J.R. Epithelial myosin light chain kinase expression and activity are upregulated in inflammatory bowel disease. Lab. Investig. 2006, 86, 191–201. [Google Scholar] [CrossRef] [Green Version]
- E Axelrad, J.; Lichtiger, S.; Yajnik, V. Inflammatory bowel disease and cancer: The role of inflammation, immunosuppression, and cancer treatment. World J. Gastroenterol. 2016, 22, 4794–4801. [Google Scholar] [CrossRef] [PubMed]
- Fazal, F.; Gu, L.; Ihnatovych, I.; Han, Y.; Hu, W.; Antic, N.; Carreira, F.; Blomquist, J.; Hope, T.; Ucker, D.; et al. Inhibiting Myosin Light Chain Kinase Induces Apoptosis In Vitro and In Vivo. Mol Cell Biol. 2005, 25, 6259–6266. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patergnani, S.; Marchi, S.; Rimessi, A.; Bonora, M.; Giorgi, C.; Mehta, K.D.; Pinton, P. PRKCB/protein kinase C, beta and the mitochondrial axis as key regulators of autophagy. Autophagy 2013, 9, 1367–1385. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shin, B.K.; Kim, C.Y.; Jung, W.Y.; Lee, H.J.; Kim, H.K.; Kim, A. Proteomic analysis reveals overexpression of moesin and cytokeratin 17 proteins in colorectal carcinoma. Oncol. Rep. 2011, 27, 608–620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, S.; Qiu, J.; Liu, L.; Su, C.; Qi, L.; Huang, C.; Chen, X.; Zhang, Y.; Ye, Y.; Ding, Y.; et al. CREB5 promotes invasiveness and metastasis in colorectal cancer by directly activating MET. J. Exp. Clin. Cancer Res. 2020, 39, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Kumaradevan, S.; Lee, S.Y.; Richards, S.; Lyle, C.; Zhao, Q.; Tapan, U.; Jiangliu, Y.; Ghumman, S.; Walker, J.; Belghasem, M.; et al. c-Cbl Expression Correlates with Human Colorectal Cancer Survival and Its Wnt/β-Catenin Suppressor Function Is Regulated by Tyr371 Phosphorylation. Am. J. Pathol. 2018, 188, 1921–1933. [Google Scholar] [CrossRef] [Green Version]
- Melo, F.D.S.E.; Kurtova, A.V.; Harnoss, J.M.; Kljavin, N.; Hoeck, J.D.; Hung, J.; Anderson, J.E.; Storm, E.E.; Modrusan, Z.; Koeppen, J.; et al. A distinct role for Lgr5+ stem cells in primary and metastatic colon cancer. Nat. Cell Biol. 2017, 543, 676–680. [Google Scholar] [CrossRef]
- Barker, N.; Van Es, J.H.; Kuipers, J.; Kujala, P.; Born, M.V.D.; Cozijnsen, M.; Haegebarth, A.; Korving, J.; Begthel, H.; Peters, P.J.; et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nat. Cell Biol. 2007, 449, 1003–1007. [Google Scholar] [CrossRef]
- Ullmann, P.; Nurmik, M.; Schmitz, M.; Rodriguez, F.; Weiler, J.; Qureshi-Baig, K.; Felten, P.; Nazarov, P.V.; Nicot, N.; Zuegel, N.; et al. Tumor suppressor miR-215 counteracts hypoxia-induced colon cancer stem cell activity. Cancer Lett. 2019, 450, 32–41. [Google Scholar] [CrossRef]
- Babraham Bioinformatics. FastQC version 0.11.2. 2014. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 10 January 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Okonechnikov, K.; Conesa, A.; García-Alcalde, F. Qualimap 2: Advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 2016, 32, 292–294. [Google Scholar] [CrossRef] [PubMed]
- Wingett, S.W.; Andrews, S. FastQ Screen: A tool for multi-genome mapping and quality control. F1000Research 2018, 7, 1338. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. ClusterProfiler: An R Package for Comparing Biological Themes Among Gene Clusters. OMICS A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Kanehisa, M. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [Green Version]
- The Gene Ontology Resource: 20 years and still Going strong. Nucleic Acids Res. 2019, 47, D330–D338. [CrossRef] [Green Version]



| KEGG ID | Number of Genes | Adjusted p-Value | Description |
|---|---|---|---|
| hsa04512 | 45 | 1.9955 × 10−5 | ECM-receptor interaction |
| hsa04510 | 90 | 2.7430 × 10−5 | Focal adhesion |
| hsa05412 | 38 | 1.9617 × 10−3 | Arrhythmogenic right ventricular cardiomyopathy |
| hsa04151 | 127 | 2.0126 × 10−3 | PI3K-Akt signaling pathway |
| hsa00980 | 31 | 2.0813 × 10−3 | Metabolism of xenobiotics by cytochrome P450 |
| hsa04950 | 14 | 3.3039 × 10−3 | Maturity onset diabetes of the young |
| hsa05200 | 186 | 3.9291 × 10−3 | Pathways in cancer |
| hsa05217 | 32 | 4.6798 × 10−3 | Basal cell carcinoma |
| hsa00982 | 28 | 4.7884 × 10−3 | Drug metabolism - cytochrome P450 |
| hsa04060 | 91 | 4.7884 × 10−3 | Cytokine-cytokine receptor interaction |
| Top 10 GO Molecular Functions Terms | ||||
| GO ID | Number of Genes | Background Genes | Adjusted p-Value | Term |
| GO:0005509 | 235 | 544 | 3.5724 × 10−7 | Calcium ion binding |
| GO:0019838 | 58 | 110 | 1.0470 × 10−3 | Growth factor binding |
| GO:0003779 | 143 | 336 | 1.0470 × 10−3 | Actin binding |
| GO:0005201 | 65 | 129 | 1.0470 × 10−3 | Extracellular matrix structural constituent |
| GO:0098772 | 464 | 1282 | 1.2080 × 10−3 | Molecular function regulator |
| GO:0015267 | 122 | 282 | 1.2080 × 10−3 | Channel activity |
| GO:0022803 | 122 | 282 | 1.2080 × 10−3 | Passive transmembrane transporter activity |
| GO:0004714 | 28 | 44 | 1.6820 × 10−3 | Transmembrane receptor protein tyrosine kinase activity |
| GO:0005102 | 417 | 1150 | 2.3635 × 10−3 | Signaling receptor binding |
| GO:0019199 | 34 | 59 | 3.0812 × 10−3 | Transmembrane receptor protein kinase activity |
| Top 10 GO biological processes terms | ||||
| GO ID | Number of Genes | Background Genes | Adjusted p-Value | Term |
| GO:0048856 | 1729 | 4694 | 3.4494 × 10−25 | Anatomical structure development |
| GO:0032501 | 2074 | 5804 | 2.8832 × 10−24 | Multicellular organismal process |
| GO:0048731 | 1451 | 3876 | 3.4175 × 10−23 | System development |
| GO:0007275 | 1593 | 4320 | 6.2510 × 10−23 | Multicellular organism development |
| GO:0032502 | 1811 | 5012 | 2.1700 × 10−22 | Developmental process |
| GO:0030154 | 1225 | 3234 | 1.0946 × 10−20 | Cell differentiation |
| GO:0009653 | 858 | 2152 | 3.8805 × 10−20 | Anatomical structure morphogenesis |
| GO:0040011 | 607 | 1436 | 6.4324 × 10−20 | Locomotion |
| GO:0051239 | 1005 | 2592 | 9.4443 × 10−20 | Regulation of multicellular organismal process |
| GO:0048869 | 1266 | 3380 | 1.3098 × 10−19 | Cellular developmental process |
| Top 10 GO cellular components terms | ||||
| GO ID | Number of Genes | Background Genes | Adjusted p-Value | Term |
| GO:0071944 | 1512 | 3934 | 2.7470 × 10−32 | Cell periphery |
| GO:0005886 | 1484 | 3853 | 2.7470 × 10−32 | Plasma membrane |
| GO:0044459 | 773 | 1916 | 4.0527 × 10−20 | Plasma membrane part |
| GO:0044425 | 1723 | 4911 | 8.8179 × 10−15 | Membrane part |
| GO:0031224 | 1351 | 3755 | 1.7922 × 10−14 | Intrinsic component of membrane |
| GO:0098590 | 412 | 976 | 2.1436 × 10−13 | Plasma membrane region |
| GO:0016021 | 1300 | 3640 | 1.2325 × 10−12 | Integral component of membrane |
| GO:0005576 | 1188 | 3298 | 1.7219 × 10−12 | Extracellular region |
| GO:0016020 | 2454 | 7347 | 5.7937 × 10−12 | Membrane |
| GO:0005911 | 177 | 373 | 4.0630 × 10−10 | Cell-cell junction |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Machackova, T.; Vychytilova-Faltejskova, P.; Souckova, K.; Trachtova, K.; Brchnelova, D.; Svoboda, M.; Kiss, I.; Prochazka, V.; Kala, Z.; Slaby, O. MiR-215-5p Reduces Liver Metastasis in an Experimental Model of Colorectal Cancer through Regulation of ECM-Receptor Interactions and Focal Adhesion. Cancers 2020, 12, 3518. https://doi.org/10.3390/cancers12123518
Machackova T, Vychytilova-Faltejskova P, Souckova K, Trachtova K, Brchnelova D, Svoboda M, Kiss I, Prochazka V, Kala Z, Slaby O. MiR-215-5p Reduces Liver Metastasis in an Experimental Model of Colorectal Cancer through Regulation of ECM-Receptor Interactions and Focal Adhesion. Cancers. 2020; 12(12):3518. https://doi.org/10.3390/cancers12123518
Chicago/Turabian StyleMachackova, Tana, Petra Vychytilova-Faltejskova, Kamila Souckova, Karolina Trachtova, Dominika Brchnelova, Marek Svoboda, Igor Kiss, Vladimir Prochazka, Zdenek Kala, and Ondrej Slaby. 2020. "MiR-215-5p Reduces Liver Metastasis in an Experimental Model of Colorectal Cancer through Regulation of ECM-Receptor Interactions and Focal Adhesion" Cancers 12, no. 12: 3518. https://doi.org/10.3390/cancers12123518






