Hyperspectral Imaging of Head and Neck Squamous Cell Carcinoma for Cancer Margin Detection in Surgical Specimens from 102 Patients Using Deep Learning
Abstract
1. Introduction
2. Materials
2.1. Head and Neck SCC Patient Dataset
2.2. Hyperspectral Imaging
2.3. Fluorescence Imaging
3. Methods
3.1. Histological Ground Truth and Registration
3.2. Intra-Patient Experiments
3.3. Inter-Patient Experiments
3.4. Convolutional Neural Network
3.5. Image Processing and Reconstruction
3.6. Evaluation
4. Results
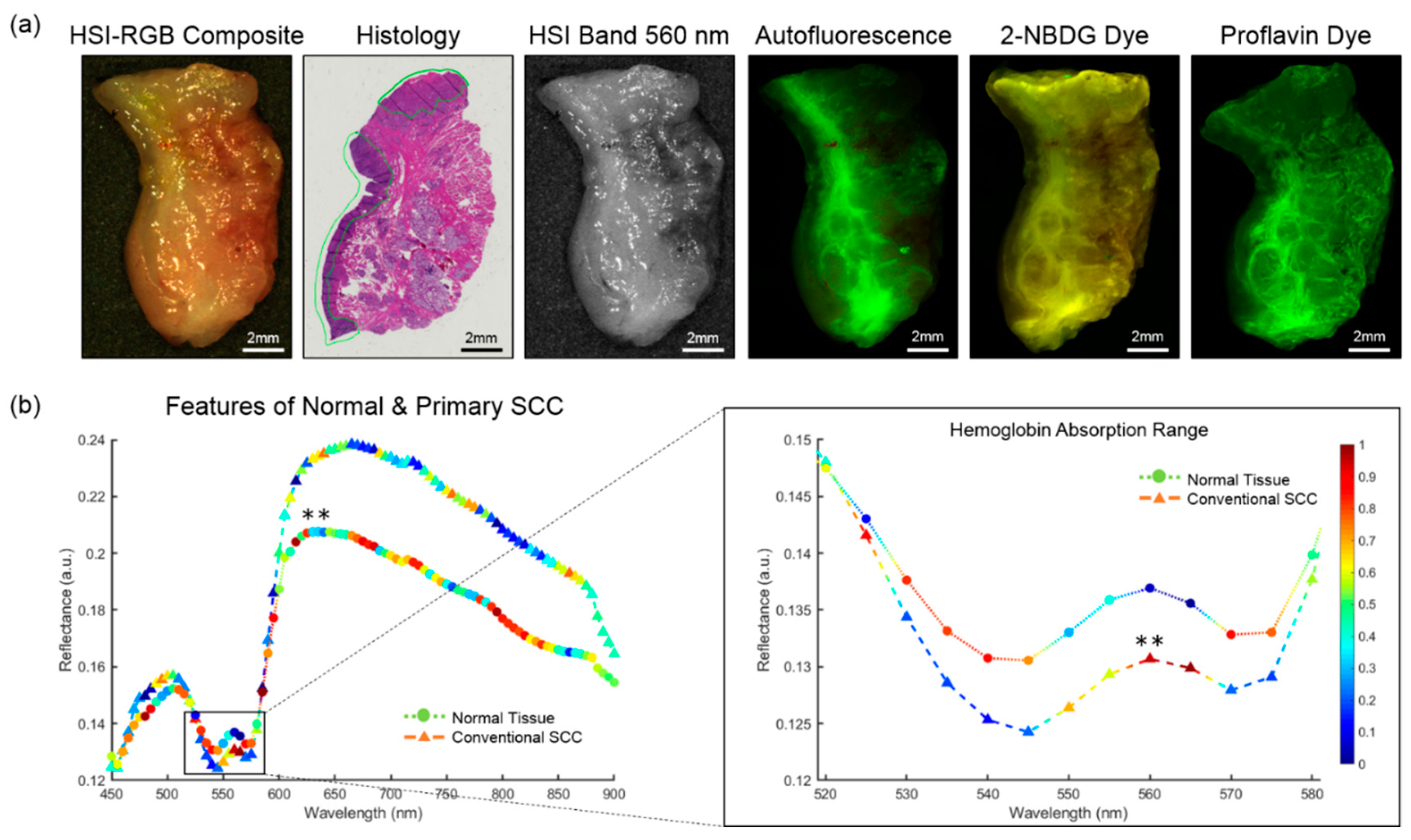
4.1. Surgical Specimens
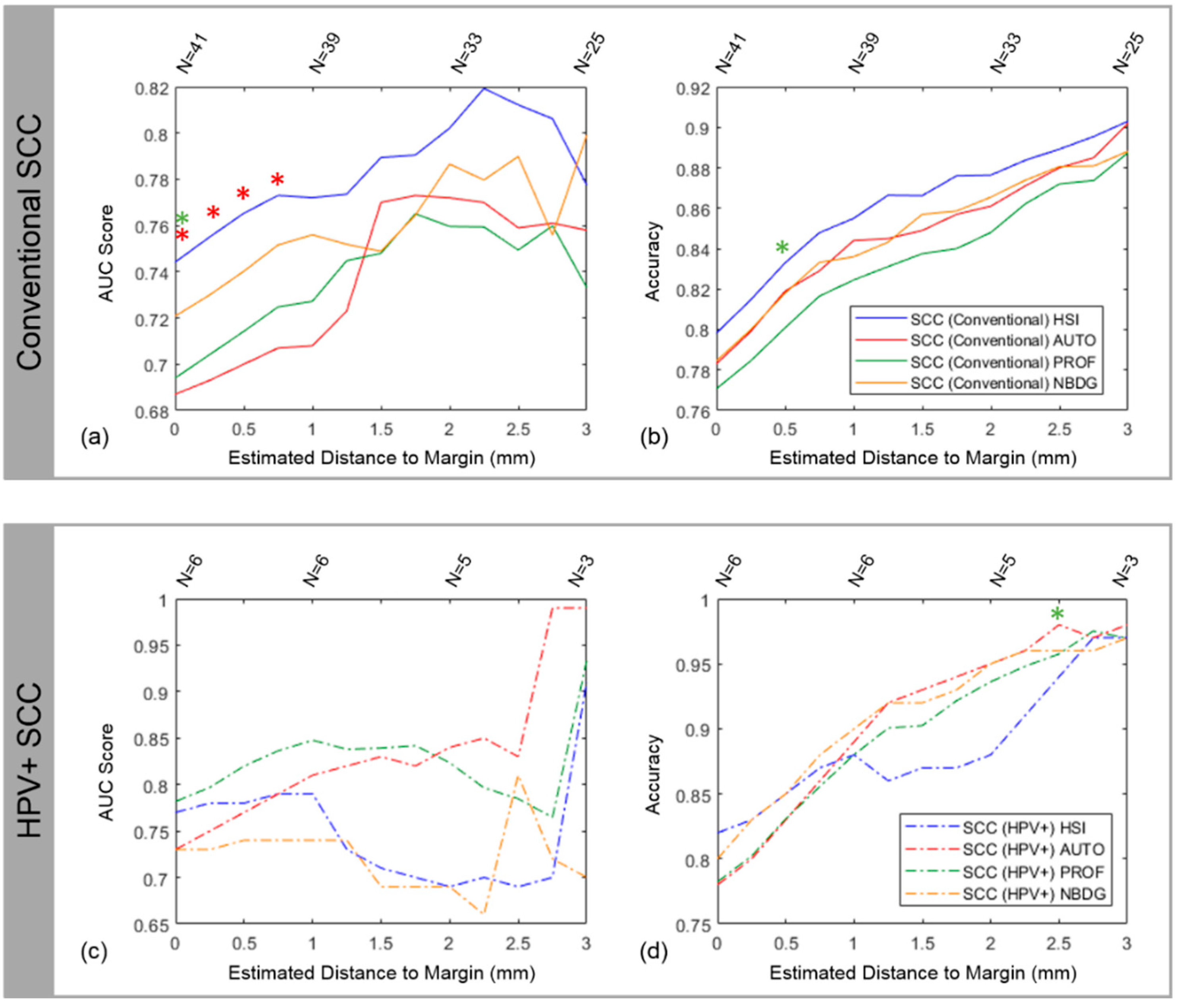
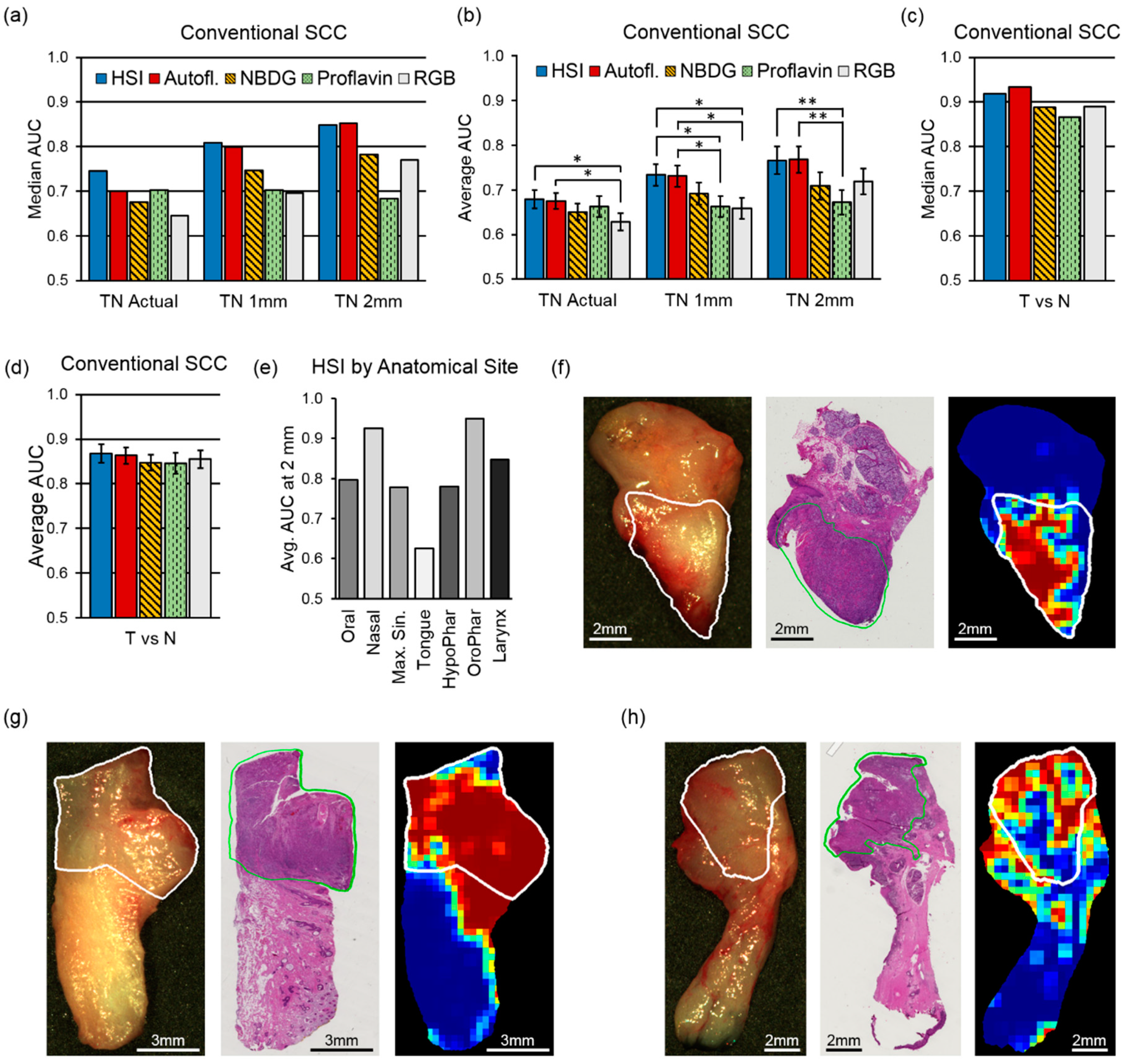
4.2. Intra-patient Experiments of Tumor-Involved Cancer Margins
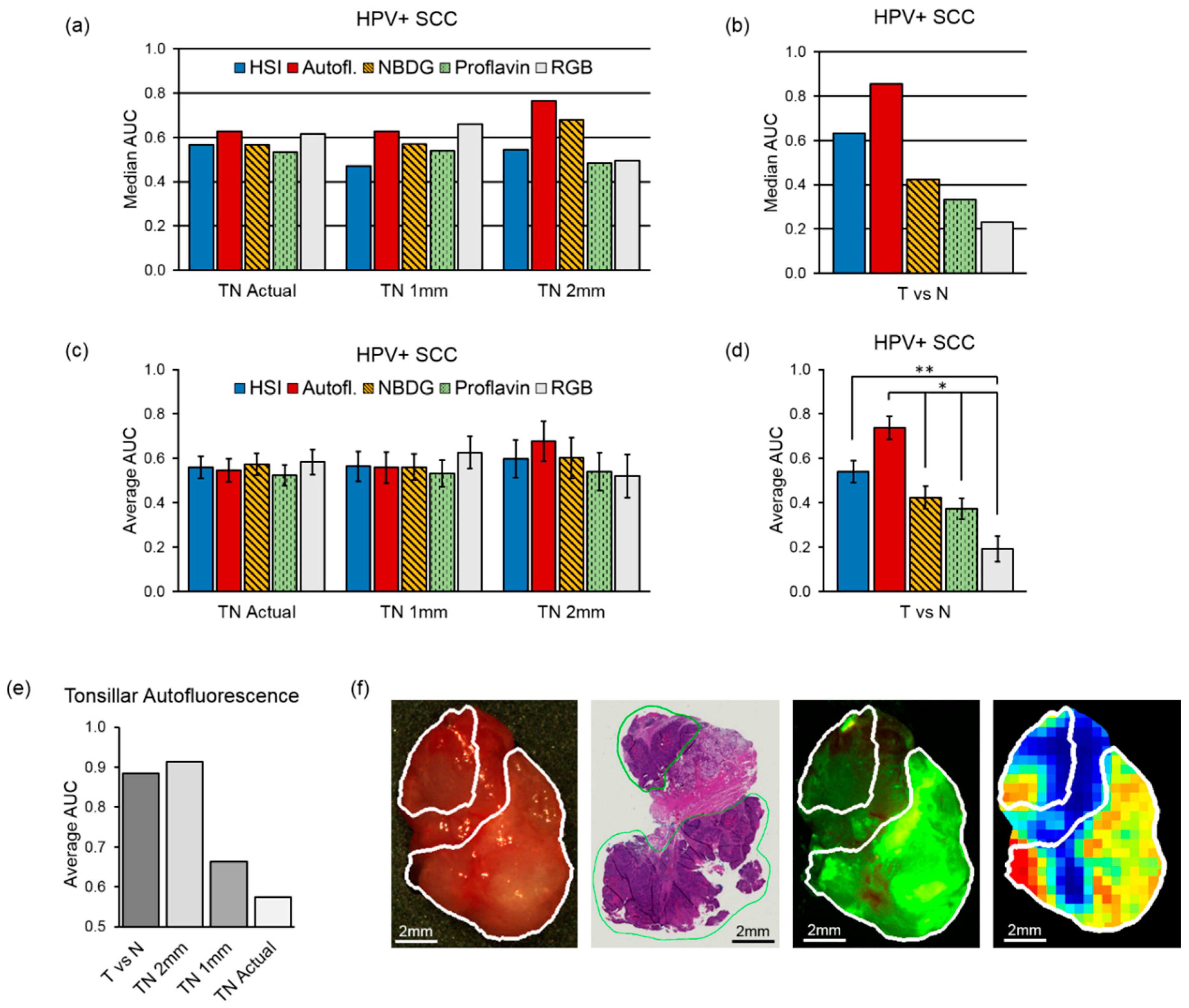
4.3. Inter-Patient Experiments of Tumor vs. Normal
4.4. Inter-patient Experiments of Tumor-Involved Cancer Margins
5. Discussion
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Marur, S.; Forastiere, A.A. Head and Neck Squamous Cell Carcinoma: Update on Epidemiology, Diagnosis, and Treatment. Mayo Clin. Proc. 2016, 91, 386–396. [Google Scholar] [CrossRef] [PubMed]
- Bozec, A.; Culié, D.; Poissonnet, G.; Dassonville, O. Current role of primary surgical treatment in patients with head and neck squamous cell carcinoma. Curr. Opin. Oncol. 2019, 31, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Van Keulen, S.; Nishio, N.; Fakurnejad, S.; Birkeland, A.; Martin, B.A.; Lu, G.; Zhou, Q.; Chirita, S.U.; Forouzanfar, T.; Colevas, A.D.; et al. The Clinical Application of Fluorescence-Guided Surgery in Head and Neck Cancer. J. Nucl. Med. 2019, 60, 758–763. [Google Scholar] [CrossRef] [PubMed]
- Joseph, L.J.; Goodman, M.; Higgins, K.; Pilai, R.; Ramalingam, S.S.; Magliocca, K.; Patel, M.R.; El-Deiry, M.; Wadsworth, J.T.; Owonikoko, T.K.; et al. Racial disparities in squamous cell carcinoma of the oral tongue among women: A SEER data analysis. Oral Oncol. 2015, 51, 586–592. [Google Scholar] [CrossRef] [PubMed]
- Vigneswaran, N.; Williams, M.D. Epidemiological Trends in Head and Neck Cancer and Aids in Diagnosis. Oral Maxillofac. Surg. Clin. North Am. 2014, 26, 123–141. [Google Scholar] [CrossRef] [PubMed]
- Gerstner, A.O.H. Early detection in head and neck cancer—current state and future perspectives. Gms Curr. Top. Otorhinolaryngol. Head Neck Surg. 2010, 7, Doc06. [Google Scholar]
- Yao, M.; Epstein, J.B.; Modi, B.J.; Pytynia, K.B.; Mundt, A.J.; Feldman, L.E. Current surgical treatment of squamous cell carcinoma of the head and neck. Oral Oncol. 2007, 43, 213–223. [Google Scholar] [CrossRef]
- Ringash, J. Survivorship and Quality of Life in Head and Neck Cancer. J. Clin. Oncol. 2015, 33, 3322–3327. [Google Scholar] [CrossRef]
- Baddour, H.M.; Magliocca, K.R.; Chen, A.Y. The importance of margins in head and neck cancer. J. Surg. Oncol. 2016, 113, 248–255. [Google Scholar] [CrossRef]
- Dinardo, L.J.; Lin, J.; Karageorge, L.S.; Powers, C.N. Accuracy, Utility, and Cost of Frozen Section Margins in Head and Neck Cancer Surgery. Laryngoscope 2000, 110, 1773–1776. [Google Scholar] [CrossRef]
- Ribeiro, N.; Godden, D.; Wilson, G.; Butterworth, D.; Woodwards, R. Do frozen sections help achieve adequate surgical margins in the resection of oral carcinoma? Int. J. Oral Maxillofac. Surg. 2003, 32, 152–158. [Google Scholar] [CrossRef]
- Du, E.; Ow, T.J.; Lo, Y.; Gersten, A.; Schiff, B.A.; Tassler, A.B.; Smith, R.V. Refining the utility and role of Frozen section in head and neck squamous cell carcinoma resection. Laryngoscope 2016, 126, 1768–1775. [Google Scholar] [CrossRef]
- Layfield, E.M.; Schmidt, R.L.; Esebua, M.; Layfield, L.J. Frozen Section Evaluation of Margin Status in Primary Squamous Cell Carcinomas of the Head and Neck: A Correlation Study of Frozen Section and Final Diagnoses. Head Neck Pathol. 2018, 12, 175–180. [Google Scholar] [CrossRef]
- Black, C.; Marotti, J.; Zarovnaya, E.; Paydarfar, J. Critical evaluation of frozen section margins in head and neck cancer resections. Cancer 2006, 107, 2792–2800. [Google Scholar] [CrossRef]
- Gao, R.W.; Teraphongphom, N.T.; Berg, N.S.V.D.; Martin, B.A.; Oberhelman, N.J.; Divi, V.; Kaplan, M.J.; Hong, S.S.; Lu, G.; Ertsey, R.; et al. Determination of Tumor Margins with Surgical Specimen Mapping Using Near-Infrared Fluorescence. Cancer Res. 2018, 78, 5144–5154. [Google Scholar] [CrossRef]
- Crowson, M.G.; Ranisau, J.; Eskander, A.; Babier, A.; Xu, B.; Kahmke, R.R.; Chen, J.M.; Chan, T.C.Y. A contemporary review of machine learning in otolaryngology-head and neck surgery. Laryngoscope 2019, 00, 1–7. [Google Scholar] [CrossRef]
- Fei, B.; Lu, G.; Wang, X.; Zhang, H.; Little, J.V.; Patel, M.R.; Griffith, C.C.; El-Diery, M.W.; Chen, A.Y. Label-free reflectance hyperspectral imaging for tumor margin assessment: A pilot study on surgical specimens of cancer patients. J. Biomed. Opt. 2017, 22, 1. [Google Scholar] [CrossRef]
- Lu, G.; Little, J.V.; Wang, X.; Zhang, H.; Patel, M.R.; Griffith, C.C.; El-Deiry, M.W.; Chen, A.Y.; Fei, B. Detection of Head and Neck Cancer in Surgical Specimens Using Quantitative Hyperspectral Imaging. Clin. Cancer Res. 2017, 23, 5426–5436. [Google Scholar] [CrossRef]
- Farah, C.S.; Fox, S.A.; Dalley, A.J. Integrated miRNA-mRNA spatial signature for oral squamous cell carcinoma: A prospective profiling study of Narrow Band Imaging guided resection. Sci. Rep. 2018, 8, 823. [Google Scholar] [CrossRef]
- Farah, C. Narrow Band Imaging-guided resection of oral cavity cancer decreases local recurrence and increases survival. Oral Dis. 2018, 24, 89–97. [Google Scholar] [CrossRef]
- Lu, G.; Fei, B. Medical hyperspectral imaging: A review. J. Biomed. Opt. 2014, 19, 10901. [Google Scholar] [CrossRef]
- Halicek, M.; Fabelo, H.; Ortega, S.; Callico, G.M.; Fei, B. In-Vivo and Ex-Vivo Tissue Analysis through Hyperspectral Imaging Techniques: Revealing the Invisible Features of Cancer. Cancers 2019, 11, 756. [Google Scholar] [CrossRef]
- Shapey, J.; Xie, Y.; Nabavi, E.; Bradford, R.; Saeed, S.R.; Ourselin, S.; Vercauteren, T. Intraoperative multispectral and hyperspectral label-free imaging: A systematic review of in vivo clinical studies. J. Biophotonics 2019, 12, e201800455. [Google Scholar] [CrossRef]
- Fabelo, H.; Halicek, M.; Ortega, S.; Shahedi, M.; Szolna, A.; Piñeiro, J.F.; Sosa, C.; O’Shanahan, A.J.; Bisshopp, S.; Espino, C.; et al. Deep Learning-Based Framework for In Vivo Identification of Glioblastoma Tumor using Hyperspectral Images of Human Brain. Sensors 2019, 19, 920. [Google Scholar] [CrossRef]
- Fabelo, H.; Ortega, S.; Szolna, A.; Bulters, D.; Pineiro, J.F.; Kabwama, S.; J-O’Shanahan, A.; Bulstrode, H.; Bisshopp, S.; Kiran, B.R.; et al. In-Vivo Hyperspectral Human Brain Image Database for Brain Cancer Detection. IEEE Access 2019, 7, 39098–39116. [Google Scholar] [CrossRef]
- Baltussen, E.J.M.; Kok, E.N.D.; De Koning, S.G.B.; Sanders, J.; Aalbers, A.G.J.; Kok, N.F.M.; Beets, G.L.; Flohil, C.C.; Bruin, S.C.; Kuhlmann, K.F.D.; et al. Hyperspectral imaging for tissue classification, a way toward smart laparoscopic colorectal surgery. J. Biomed. Opt. 2019, 24, 016002. [Google Scholar] [CrossRef]
- Leavesley, S.J.; Walters, M.; Lopez, C.; Baker, T.; Favreau, P.F.; Rich, T.C.; Rider, P.F.; Boudreaux, C.W. Hyperspectral imaging fluorescence excitation scanning for colon cancer detection. J. Biomed. Opt. 2016, 21, 104003. [Google Scholar] [CrossRef]
- Halicek, M.; Little, J.V.; Wang, X.; Patel, M.R.; Griffith, C.C.; Chen, A.Y.; Fei, B. Tumor Margin Classification of Head and Neck Cancer Using Hyperspectral Imaging and Convolutional Neural Networks. Proc. SPIE Int. Soc. Opt. Eng. 2018, 10576, 1057605. [Google Scholar]
- Halicek, M.; Lu, G.; Little, J.V.; Wang, X.; Patel, M.; Griffith, C.C.; El-Deiry, M.W.; Chen, A.Y.; Fei, B. Deep convolutional neural networks for classifying head and neck cancer using hyperspectral imaging. J. Biomed. Opt. 2017, 22, 60503. [Google Scholar] [CrossRef]
- Halicek, M.; Little, J.V.; Wang, X.; Chen, A.Y.; Fei, B. Optical biopsy of head and neck cancer using hyperspectral imaging and convolutional neural networks. J. Biomed. Opt. 2019, 24, 036007. [Google Scholar] [CrossRef]
- Manni, F.; Van Der Sommen, F.; Zinger, S.; Kho, E.; De Koning, S.G.B.; Ruers, T.J.M.; Shan, C.; Schleipen, J.; De With, P.H.N. Automated tumor assessment of squamous cell carcinoma on tongue cancer patients with hyperspectral imaging. Proc. SPIE 2019, 10951, 109512K. [Google Scholar]
- Trajanovski, S.; Shan, C.; Weijtmans, P.J.C.; de Koning, S.G.B.; Ruers, T.J.M. Tumor Semantic Segmentation in Hyperspectral Images using Deep Learning. In Proceedings of the 2nd International Conference on Medical Imaging with Deep Learning, London, UK, 8–10 July 2019. [Google Scholar]
- Hellebust, A.; Rosbach, K.; Wu, J.K.; Nguyen, J.; Gillenwater, A.; Vigneswaran, N.; Richards-Kortum, R. Vital-dye-enhanced multimodal imaging of neoplastic progression in a mouse model of oral carcinogenesis. J. Biomed. Opt. 2013, 18, 126017. [Google Scholar] [CrossRef]
- Thekkek, N.; Muldoon, T.; Polydorides, A.D.; Maru, D.M.; Harpaz, N.; Harris, M.T.; Anandasabapathy, S. Vital-dye enhanced fluorescence imaging of GI mucosa: Metaplasia, neoplasia, inflammation. Gastrointest. Endosc. 2012, 75, 877–887. [Google Scholar] [CrossRef]
- Halicek, M.; Shahedi, M.; Little, J.V.; Chen, A.Y.; Myers, L.L.; Sumer, B.D.; Fei, B. Detection of squamous cell carcinoma in digitized histological images from the head and neck using convolutional neural networks. Proc. SPIE 2019, 10956, 109560K. [Google Scholar]
- Halicek, M.; Little, J.V.; Wang, X.; Chen, Z.G.; Patel, M.R.; Griffith, C.C.; El-Deiry, M.W.; Saba, N.F.; Chen, A.Y.; Fei, B. Deformable Registration of Histological Cancer Margins to Gross Hyperspectral Images using Demons. Proc. SPIE Int. Soc. Opt. Eng. 2018, 10581, 105810N. [Google Scholar]
- Halicek, M.; Fabelo, H.; Ortega, S.; Little, J.V.; Wang, X.; Chen, A.Y.; Callico, G.M.; Myers, L.; Sumer, B.D.; Fei, B.; et al. Hyperspectral imaging for head and neck cancer detection: Specular glare and variance of the tumor margin in surgical specimens. J. Med Imaging 2019, 6. in press. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A. Inception-v4, Inception-ResNet and the Impact of Residual Connections on Learning. In Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–7 February 2017. [Google Scholar]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-scale Machine Learning on Heterogeneous Systems. 2015. Available online: https://www.tensorflow.org (accessed on 29 December 2018).
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar]
- Sekar, S.K.V.; Bargigia, I.; Mora, A.D.; Taroni, P.; Ruggeri, A.; Tosi, A.; Pifferi, A.; Farina, A. Diffuse optical characterization of collagen absorption from 500 to 1700 nm. J. Biomed. Opt. 2017, 22, 015006. [Google Scholar] [CrossRef]
- Shi, Z.; Chen, C.; Xiong, Z.; Liu, D.; Wu, F. HSCNN+: Advanced CNN-Based Hyperspectral Recovery from RGB Images. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), Salt Lake City, UT, USA, 18–22 June 2018; Volume 3, p. 5. [Google Scholar]
| Property | Conventional SCC (N = 88) | HPV+ SCC (N = 14) | All SCC (N = 102) | |||
|---|---|---|---|---|---|---|
| Number | % | Number | % | Number | % | |
| Demographics | ||||||
| Mean Age (y.o.) | 64.5 | - | 58.1 | - | 63.6 | - |
| Male | 59 | 67% | 12 | 86% | 71 | 70% |
| Female | 29 | 33% | 2 | 14% | 31 | 30% |
| Tobacco History | 58 | 66% | 5 | 36% | 63 | 62% |
| Primary Location | ||||||
| Oral Cavity | 35 | 40% | 0 | 0% | 35 | 34% |
| Tongue | 19 | 22% | 0 | 0% | 19 | 19% |
| Oropharynx | 2 | 2% | 13 | 93% | 15 | 15% |
| Hypopharynx | 4 | 5% | 0 | 0% | 4 | 4% |
| Larynx | 19 | 22% | 0 | 0% | 19 | 19% |
| Nasal Cavity | 4 | 5% | 0 | 0% | 4 | 4% |
| Maxillary Sinus | 5 | 6% | 0 | 0% | 5 | 5% |
| Unknown | 0 | 0% | 1 | 7% | 1 | 1% |
| Cancer Stage | ||||||
| pT1 | 3 | 3% | 2 | 14% | 5 | 5% |
| pT2 | 7 | 8% | 6 | 43% | 13 | 13% |
| pT3 | 16 | 18% | 2 | 14% | 18 | 18% |
| pT4 | 62 | 70% | 3 | 21% | 65 | 64% |
| Avg. T Size (cm) | 4.4 | - | 3 | - | 4.2 | - |
| N+ | 53 | 60% | 5 | 36% | 58 | 57% |
| Histologic Grade | ||||||
| G1 | 8 | 10% | - | - | 8 | 8% |
| G2 | 60 | 71% | - | - | 60 | 59% |
| G3 | 16 | 19% | - | - | 16 | 16% |
| IPC (Averages) | ||||||
| IPC/Surgery | 2.1 | - | 2.0 | - | 2.1 | - |
| Time/IPC (min) | 41 | - | 42 | - | 41 | - |
| Tissues/Surgery | 3.5 | - | 2.6 | - | 3.4 | - |
| Time/Tissue (min) | 25 | - | 33 | - | 25 | - |
| Imaging Modality | SCC Cohort | 1 mm | 2 mm | 3 mm | |||
|---|---|---|---|---|---|---|---|
| AUC | Accuracy | AUC | Accuracy | AUC | Accuracy | ||
| HSI | SCC, Conventional | 0.77 | 85% | 0.80 | 88% | 0.78 | 90% |
| SCC, HPV+ | 0.79 | 88% | 0.69 | 88% | 0.91 | 97% | |
| Autofluorescence | SCC, Conventional | 0.67 | 85% | 0.72 | 85% | 0.73 | 87% |
| SCC, HPV+ | 0.81 | 89% | 0.84 | 95% | 0.99 | 98% | |
| Proflavin | SCC, Conventional | 0.73 | 82% | 0.76 | 85% | 0.73 | 89% |
| SCC, HPV+ | 0.85 | 88% | 0.82 | 94% | 0.93 | 97% | |
| 2-NBDG | SCC, Conventional | 0.76 | 84% | 0.79 | 87% | 0.80 | 89% |
| SCC, HPV+ | 0.74 | 90% | 0.69 | 95% | 0.70 | 97% | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Halicek, M.; Dormer, J.D.; Little, J.V.; Chen, A.Y.; Myers, L.; Sumer, B.D.; Fei, B. Hyperspectral Imaging of Head and Neck Squamous Cell Carcinoma for Cancer Margin Detection in Surgical Specimens from 102 Patients Using Deep Learning. Cancers 2019, 11, 1367. https://doi.org/10.3390/cancers11091367
Halicek M, Dormer JD, Little JV, Chen AY, Myers L, Sumer BD, Fei B. Hyperspectral Imaging of Head and Neck Squamous Cell Carcinoma for Cancer Margin Detection in Surgical Specimens from 102 Patients Using Deep Learning. Cancers. 2019; 11(9):1367. https://doi.org/10.3390/cancers11091367
Chicago/Turabian StyleHalicek, Martin, James D. Dormer, James V. Little, Amy Y. Chen, Larry Myers, Baran D. Sumer, and Baowei Fei. 2019. "Hyperspectral Imaging of Head and Neck Squamous Cell Carcinoma for Cancer Margin Detection in Surgical Specimens from 102 Patients Using Deep Learning" Cancers 11, no. 9: 1367. https://doi.org/10.3390/cancers11091367
APA StyleHalicek, M., Dormer, J. D., Little, J. V., Chen, A. Y., Myers, L., Sumer, B. D., & Fei, B. (2019). Hyperspectral Imaging of Head and Neck Squamous Cell Carcinoma for Cancer Margin Detection in Surgical Specimens from 102 Patients Using Deep Learning. Cancers, 11(9), 1367. https://doi.org/10.3390/cancers11091367






