Multigene Panel Testing Increases the Number of Loci Associated with Gastric Cancer Predisposition
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Patients’ Selection
- I: Two or more GC cases regardless of age, at least one confirmed DGC;
- II: One case of DGC <40 years;
- III: Personal or family history of DGC and LBC, one diagnosed <50 years;
- IV: Bilateral LBC or family history of two or more cases of LBC <50 years;
- V: GC ≤ 60 years with a family history of colorectal cancer;
- VI: Two or more GC cases ≤ 60 years in first-degree relatives;
- VII: Several gastric polyps ≤ 60 years with a family history of at least two GC cases.
4.2. Sample Collection and DNA Extraction
4.3. Multigene Panel (MGP) Testing
4.4. Data Analysis and Variant Calling
4.5. Additional Molecular Analyses
4.6. Confirmation of Variants
4.7. Variant Classification
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Lauren, P. The two histological main types of gastric carcinoma: Diffuse and so-called intestinal-type carcinoma. An attempt at a histo-clinical classification. Acta Pathol. Microbiol. Scand. 1965, 64, 31–49. [Google Scholar] [CrossRef] [PubMed]
- Zali, H.; Rezaei-Tavirani, M.; Azodi, M. Gastric cancer: Prevention, risk factors and treatment. Gastroenterol. Hepatol. from bed to bench Gastroenterol Hepatol Bed Bench 2011, 4, 175–185. [Google Scholar] [PubMed]
- Petrovchich, I.; Ford, J.M. Genetic predisposition to gastric cancer. Semin. Oncol. 2016, 43, 554–559. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, C.; Pinheiro, H.; Figueiredo, J.; Seruca, R.; Carneiro, F. Familial gastric cancer: Genetic susceptibility, pathology, and implications for management. Lancet. Oncol. 2015, 16, e60–e70. [Google Scholar] [CrossRef]
- Berx, G.; Staes, K.; van Hengel, J.; Molemans, F.; Bussemakers, M.J.; van Bokhoven, A.; van Roy, F. Cloning and characterization of the human invasion suppressor gene E-cadherin (CDH1). Genomics 1995, 26, 281–289. [Google Scholar] [CrossRef]
- Guilford, P.; Hopkins, J.; Harraway, J.; McLeod, M.; McLeod, N.; Harawira, P.; Taite, H.; Scoular, R.; Miller, A.; Reeve, A.E. E-cadherin germline mutations in familial gastric cancer. Nature 1998, 392, 402–405. [Google Scholar] [CrossRef]
- Guilford, P.; Humar, B.; Blair, V. Hereditary diffuse gastric cancer: Translation of CDH1 germline mutations into clinical practice. Gastric Cancer 2010, 13, 1–10. [Google Scholar] [CrossRef]
- Hansford, S.; Kaurah, P.; Li-Chang, H.; Woo, M.; Senz, J.; Pinheiro, H.; Schrader, K.A.; Schaeffer, D.F.; Shumansky, K.; Zogopoulos, G.; et al. Hereditary Diffuse Gastric Cancer Syndrome: CDH1 Mutations and Beyond. JAMA Oncol. 2015, 1, 23–32. [Google Scholar] [CrossRef]
- Caldas, C.; Carneiro, F.; Lynch, H.T.; Yokota, J.; Wiesner, G.L.; Powell, S.M.; Lewis, F.R.; Huntsman, D.G.; Pharoah, P.D.; Jankowski, J.A.; et al. Familial gastric cancer: Overview and guidelines for management. J. Med. Genet. 1999, 36, 873–880. [Google Scholar]
- Brooks-Wilson, A.R.; Kaurah, P.; Suriano, G.; Leach, S.; Senz, J.; Grehan, N.; Butterfield, Y.S.N.; Jeyes, J.; Schinas, J.; Bacani, J.; et al. Germline E-cadherin mutations in hereditary diffuse gastric cancer: Assessment of 42 new families and review of genetic screening criteria. J. Med. Genet. 2004, 41, 508–517. [Google Scholar] [CrossRef] [PubMed]
- Fitzgerald, R.C.; Hardwick, R.; Huntsman, D.; Carneiro, F.; Guilford, P.; Blair, V.; Chung, D.C.; Norton, J.; Ragunath, K.; Van Krieken, J.H.; et al. Hereditary diffuse gastric cancer: Updated consensus guidelines for clinical management and directions for future research. J. Med. Genet. 2010, 47, 436–444. [Google Scholar] [CrossRef] [PubMed]
- van der Post, R.S.; Vogelaar, I.P.; Carneiro, F.; Guilford, P.; Huntsman, D.; Hoogerbrugge, N.; Caldas, C.; Schreiber, K.E.C.; Hardwick, R.H.; Ausems, M.G.E.M.; et al. Hereditary diffuse gastric cancer: Updated clinical guidelines with an emphasis on germline CDH1 mutation carriers. J. Med. Genet. 2015, 52, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, C.; Senz, J.; Kaurah, P.; Pinheiro, H.; Sanges, R.; Haegert, A.; Corso, G.; Schouten, J.; Fitzgerald, R.; Vogelsang, H.; et al. Germline CDH1 deletions in hereditary diffuse gastric cancer families. Hum. Mol. Genet. 2009, 18, 1545–1555. [Google Scholar] [CrossRef] [PubMed]
- Molinaro, V.; Pensotti, V.; Marabelli, M.; Feroce, I.; Barile, M.; Pozzi, S.; Laghi, L.; Serrano, D.; Bernard, L.; Bonanni, B.; et al. Complementary molecular approaches reveal heterogeneous CDH1 germline defects in Italian patients with hereditary diffuse gastric cancer (HDGC) syndrome. Genes Chromosomes Cancer 2014, 53, 432–445. [Google Scholar] [CrossRef] [PubMed]
- Worthley, D.L.; Phillips, K.D.; Wayte, N.; Schrader, K.A.; Healey, S.; Kaurah, P.; Shulkes, A.; Grimpen, F.; Clouston, A.; Moore, D.; et al. Gastric adenocarcinoma and proximal polyposis of the stomach (GAPPS): A new autosomal dominant syndrome. Gut 2012, 61, 774–779. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Woods, S.L.; Healey, S.; Beesley, J.; Chen, X.; Lee, J.S.; Sivakumaran, H.; Wayte, N.; Nones, K.; Waterfall, J.J.; et al. Point Mutations in Exon 1B of APC Reveal Gastric Adenocarcinoma and Proximal Polyposis of the Stomach as a Familial Adenomatous Polyposis Variant. Am. J. Hum. Genet. 2016, 98, 830–842. [Google Scholar] [CrossRef] [PubMed]
- Majewski, I.J.; Kluijt, I.; Cats, A.; Scerri, T.S.; de Jong, D.; Kluin, R.J.C.; Hansford, S.; Hogervorst, F.B.L.; Bosma, A.J.; Hofland, I.; et al. An α-E-catenin (CTNNA1) mutation in hereditary diffuse gastric cancer. J. Pathol. 2013, 229, 621–629. [Google Scholar] [CrossRef] [PubMed]
- Gaston, D.; Hansford, S.; Oliveira, C.; Nightingale, M.; Pinheiro, H.; Macgillivray, C.; Kaurah, P.; Rideout, A.L.; Steele, P.; Soares, G.; et al. Germline mutations in MAP3K6 are associated with familial gastric cancer. PLoS Genet. 2014, 10, e1004669. [Google Scholar] [CrossRef]
- Vogelaar, I.P.; Ligtenberg, M.J.L.; van der Post, R.S.; de Voer, R.M.; Kets, C.M.; Jansen, T.J.G.; Jacobs, L.; Schreibelt, G.; International Gastric Cancer Genetics Group; de Vries, I.J.M.; et al. Recurrent candidiasis and early-onset gastric cancer in a patient with a genetically defined partial MYD88 defect. Fam. Cancer 2016, 15, 289–296. [Google Scholar] [CrossRef]
- Sahasrabudhe, R.; Lott, P.; Bohorquez, M.; Toal, T.; Estrada, A.P.; Suarez, J.J.; Brea-Fernández, A.; Cameselle-Teijeiro, J.; Pinto, C.; Ramos, I.; et al. Germline Mutations in PALB2, BRCA1, and RAD51C, Which Regulate DNA Recombination Repair, in Patients With Gastric Cancer. Gastroenterology 2017, 152, 983–986.e6. [Google Scholar] [CrossRef] [PubMed]
- Vogelaar, I.P.; van der Post, R.S.; van Krieken, J.H.J.; Spruijt, L.; van Zelst-Stams, W.A.; Kets, C.M.; Lubinski, J.; Jakubowska, A.; Teodorczyk, U.; Aalfs, C.M.; et al. Unraveling genetic predisposition to familial or early onset gastric cancer using germline whole-exome sequencing. Eur. J. Hum. Genet. 2017, 25, 1246–1252. [Google Scholar] [CrossRef] [PubMed]
- Weren, R.D.A.; van der Post, R.S.; Vogelaar, I.P.; van Krieken, J.H.; Spruijt, L.; Lubinski, J.; Jakubowska, A.; Teodorczyk, U.; Aalfs, C.M.; van Hest, L.P.; et al. Role of germline aberrations affecting CTNNA1, MAP3K6 and MYD88 in gastric cancer susceptibility. J. Med. Genet. 2018, 55, 669–674. [Google Scholar] [CrossRef] [PubMed]
- Fewings, E.; Larionov, A.; Redman, J.; Goldgraben, M.A.; Scarth, J.; Richardson, S.; Brewer, C.; Davidson, R.; Ellis, I.; Evans, D.G.; et al. Germline pathogenic variants in PALB2 and other cancer-predisposing genes in families with hereditary diffuse gastric cancer without CDH1 mutation: A whole-exome sequencing study. Lancet. Gastroenterol. Hepatol. 2018, 3, 489–498. [Google Scholar] [CrossRef]
- Slavin, T.; Neuhausen, S.L.; Rybak, C.; Solomon, I.; Nehoray, B.; Blazer, K.; Niell-Swiller, M.; Adamson, A.W.; Yuan, Y.-C.; Yang, K.; et al. Genetic Gastric Cancer Susceptibility in the International Clinical Cancer Genomics Community Research Network. Cancer Genet. 2017, 216–217, 111–119. [Google Scholar] [CrossRef]
- Lee, K.; Krempely, K.; Roberts, M.E.; Anderson, M.J.; Carneiro, F.; Chao, E.; Dixon, K.; Figueiredo, J.; Ghosh, R.; Huntsman, D.; et al. Specifications of the ACMG/AMP variant curation guidelines for the analysis of germline CDH1 sequence variants. Hum. Mutat. 2018, 39, 1553–1568. [Google Scholar] [CrossRef] [PubMed]
- More, H.; Humar, B.; Weber, W.; Ward, R.; Christian, A.; Lintott, C.; Graziano, F.; Ruzzo, A.-M.; Acosta, E.; Boman, B.; et al. Identification of seven novel germline mutations in the human E-cadherin (CDH1) gene. Hum. Mutat. 2007, 28, 203. [Google Scholar] [CrossRef]
- Lynch, H.T.; Aldoss, I.; Lynch, J.F. The identification and management of hereditary diffuse gastric cancer in a large Jordanian family. Fam. Cancer 2011, 10, 667–672. [Google Scholar] [CrossRef]
- Benusiglio, P.R.; Malka, D.; Rouleau, E.; De Pauw, A.; Buecher, B.; Noguès, C.; Fourme, E.; Colas, C.; Coulet, F.; Warcoin, M.; et al. CDH1 germline mutations and the hereditary diffuse gastric and lobular breast cancer syndrome: A multicentre study. J. Med. Genet. 2013, 50, 486–489. [Google Scholar] [CrossRef]
- ClinVar–NCBI–NIH. Available online: https://www.ncbi.nlm.nih.gov/clinvar (accessed on 30 July 2019).
- Berx, G.; Cleton-Jansen, A.M.; Nollet, F.; de Leeuw, W.J.; van de Vijver, M.; Cornelisse, C.; van Roy, F. E-cadherin is a tumour/invasion suppressor gene mutated in human lobular breast cancers. EMBO J. 1995, 14, 6107–6115. [Google Scholar] [CrossRef]
- Jonsson, B.-A.; Bergh, A.; Stattin, P.; Emmanuelsson, M.; Grönberg, H. Germline mutations in E-cadherin do not explain association of hereditary prostate cancer, gastric cancer and breast cancer. Int. J. cancer 2002, 98, 838–843. [Google Scholar] [CrossRef] [PubMed]
- Suriano, G.; Yew, S.; Ferreira, P.; Senz, J.; Kaurah, P.; Ford, J.M.; Longacre, T.A.; Norton, J.A.; Chun, N.; Young, S.; et al. Characterization of a recurrent germ line mutation of the E-cadherin gene: Implications for genetic testing and clinical management. Clin. Cancer Res. 2005, 11, 5401–5409. [Google Scholar] [CrossRef] [PubMed]
- Rogers, W.M.; Dobo, E.; Norton, J.A.; Van Dam, J.; Jeffrey, R.B.; Huntsman, D.G.; Kingham, K.; Chun, N.; Ford, J.M.; Longacre, T.A. Risk-reducing total gastrectomy for germline mutations in E-cadherin (CDH1): Pathologic findings with clinical implications. Am. J. Surg. Pathol. 2008, 32, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Chung, J.-W.; Jeong, T.-D.; Park, Y.-S.; Lee, J.H.; Ahn, J.Y.; Kim, D.H.; Choi, K.D.; Lee, W.; Song, H.J.; et al. Searching for E-cadherin gene mutations in early onset diffuse gastric cancer and hereditary diffuse gastric cancer in Korean patients. Fam. Cancer 2013, 12, 503–507. [Google Scholar] [CrossRef] [PubMed]
- Frebourg, T.; Oliveira, C.; Hochain, P.; Karam, R.; Manouvrier, S.; Graziadio, C.; Vekemans, M.; Hartmann, A.; Baert-Desurmont, S.; Alexandre, C.; et al. Cleft lip/palate and CDH1/E-cadherin mutations in families with hereditary diffuse gastric cancer. J. Med. Genet. 2006, 43, 138–142. [Google Scholar] [CrossRef] [PubMed]
- Kaurah, P.; MacMillan, A.; Boyd, N.; Senz, J.; De Luca, A.; Chun, N.; Suriano, G.; Zaor, S.; Van Manen, L.; Gilpin, C.; et al. Founder and recurrent CDH1 mutations in families with hereditary diffuse gastric cancer. JAMA 2007, 297, 2360–2372. [Google Scholar] [CrossRef]
- Wijnen, J.; van der Klift, H.; Vasen, H.; Khan, P.M.; Menko, F.; Tops, C.; Meijers Heijboer, H.; Lindhout, D.; Møller, P.; Fodde, R. MSH2 genomic deletions are a frequent cause of HNPCC. Nat. Genet. 1998, 20, 326. [Google Scholar] [CrossRef]
- Stepp, S.E.; Dufourcq-Lagelouse, R.; Le Deist, F.; Bhawan, S.; Certain, S.; Mathew, P.A.; Henter, J.I.; Bennett, M.; Fischer, A.; de Saint Basile, G.; et al. Perforin gene defects in familial hemophagocytic lymphohistiocytosis. Science 1999, 286, 1957–1959. [Google Scholar] [CrossRef] [PubMed]
- Lakin, N.D.; Weber, P.; Stankovic, T.; Rottinghaus, S.T.; Taylor, A.M.; Jackson, S.P. Analysis of the ATM protein in wild-type and ataxia telangiectasia cells. Oncogene 1996, 13, 2707–2716. [Google Scholar]
- Saviozzi, S.; Saluto, A.; Piane, M.; Prudente, S.; Migone, N.; DeMarchi, M.; Brusco, A.; Chessa, L. Six novel ATM mutations in Italian patients with classical ataxia-telangiectasia. Hum. Mutat. 2003, 21, 450. [Google Scholar] [CrossRef]
- Meindl, A. German Consortium for Hereditary Breast and Ovarian Cancer Comprehensive analysis of 989 patients with breast or ovarian cancer provides BRCA1 and BRCA2 mutation profiles and frequencies for the German population. Int. J. Cancer 2002, 97, 472–480. [Google Scholar] [PubMed]
- Casadei, S.; Norquist, B.M.; Walsh, T.; Stray, S.; Mandell, J.B.; Lee, M.K.; Stamatoyannopoulos, J.A.; King, M.-C. Contribution of inherited mutations in the BRCA2-interacting protein PALB2 to familial breast cancer. Cancer Res. 2011, 71, 2222–2229. [Google Scholar] [CrossRef]
- Kluska, A.; Balabas, A.; Paziewska, A.; Kulecka, M.; Nowakowska, D.; Mikula, M.; Ostrowski, J. New recurrent BRCA1/2 mutations in Polish patients with familial breast/ovarian cancer detected by next generation sequencing. BMC Med. Genomics 2015, 8, 19. [Google Scholar] [CrossRef] [PubMed]
- Charlton, A.; Blair, V.; Shaw, D.; Parry, S.; Guilford, P.; Martin, I.G. Hereditary diffuse gastric cancer: Predominance of multiple foci of signet ring cell carcinoma in distal stomach and transitional zone. Gut 2004, 53, 814–820. [Google Scholar] [CrossRef] [PubMed]
- Rocha, J.P.; Gullo, I.; Wen, X.; Devezas, V.; Baptista, M.; Oliveira, C.; Carneiro, F. Pathological features of total gastrectomy specimens from asymptomatic hereditary diffuse gastric cancer patients and implications for clinical management. Histopathology 2018, 73, 878–886. [Google Scholar] [CrossRef] [PubMed]
- Mi, E.Z.E.Z.; Mi, E.Z.E.Z.; di Pietro, M.; O’Donovan, M.; Hardwick, R.H.; Richardson, S.; Ziauddeen, H.; Fletcher, P.C.; Caldas, C.; Tischkowitz, M.; et al. Comparative study of endoscopic surveillance in hereditary diffuse gastric cancer according to CDH1 mutation status. Gastrointest. Endosc. 2018, 87, 408–418. [Google Scholar] [CrossRef]
- Kohlmann, W.; Gruber, S.B. Lynch Syndrome. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Stephens, K., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 2004; [updated 2018]. [Google Scholar]
- Larsen Haidle, J.; Howe, J.R. Juvenile Polyposis Syndrome. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Stephens, K., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 2003; [updated 2017]. [Google Scholar]
- Aarnio, M.; Salovaara, R.; Aaltonen, L.A.; Mecklin, J.P.; Järvinen, H.J. Features of gastric cancer in hereditary non-polyposis colorectal cancer syndrome. Int. J. Cancer 1997, 74, 551–555. [Google Scholar] [CrossRef]
- Bonadona, V.; Bonaïti, B.; Olschwang, S.; Grandjouan, S.; Huiart, L.; Longy, M.; Guimbaud, R.; Buecher, B.; Bignon, Y.-J.; Caron, O.; et al. Cancer risks associated with germline mutations in MLH1, MSH2, and MSH6 genes in Lynch syndrome. JAMA 2011, 305, 2304–2310. [Google Scholar] [CrossRef]
- Giardiello, F.M.; Allen, J.I.; Axilbund, J.E.; Boland, C.R.; Burke, C.A.; Burt, R.W.; Church, J.M.; Dominitz, J.A.; Johnson, D.A.; Kaltenbach, T.; et al. Guidelines on genetic evaluation and management of Lynch syndrome: A consensus statement by the US Multi-society Task Force on colorectal cancer. Am. J. Gastroenterol. 2014, 109, 1159–1179. [Google Scholar] [CrossRef]
- Brosens, L.A.A.; van Hattem, A.; Hylind, L.M.; Iacobuzio-Donahue, C.; Romans, K.E.; Axilbund, J.; Cruz-Correa, M.; Tersmette, A.C.; Offerhaus, G.J.A.; Giardiello, F.M. Risk of colorectal cancer in juvenile polyposis. Gut 2007, 56, 965–967. [Google Scholar] [CrossRef]
- Latchford, A.R.; Neale, K.; Phillips, R.K.S.; Clark, S.K. Juvenile polyposis syndrome: A study of genotype, phenotype, and long-term outcome. Dis. Colon Rectum 2012, 55, 1038–1043. [Google Scholar] [CrossRef] [PubMed]
- Aytac, E.; Sulu, B.; Heald, B.; O’Malley, M.; LaGuardia, L.; Remzi, F.H.; Kalady, M.F.; Burke, C.A.; Church, J.M. Genotype-defined cancer risk in juvenile polyposis syndrome. Br. J. Surg. 2015, 102, 114–118. [Google Scholar] [CrossRef] [PubMed]
- Adar, T.; Rodgers, L.H.; Shannon, K.M.; Yoshida, M.; Ma, T.; Mattia, A.; Lauwers, G.Y.; Iafrate, A.J.; Hartford, N.M.; Oliva, E.; et al. Universal screening of both endometrial and colon cancers increases the detection of Lynch syndrome. Cancer 2018, 124, 3145–3153. [Google Scholar] [CrossRef] [PubMed]
- Ciambotti, B.; Mussolin, L.; D’Amore, E.S.G.; Pillon, M.; Sieni, E.; Coniglio, M.L.; Ros, M.D.; Cetica, V.; Aricò, M.; Rosolen, A. Monoallelic mutations of the perforin gene may represent a predisposing factor to childhood anaplastic large cell lymphoma. J. Pediatr. Hematol. Oncol. 2014, 36, e359–e365. [Google Scholar] [CrossRef] [PubMed]
- Cunniff, C.; Bassetti, J.A.; Ellis, N.A. Bloom’s Syndrome: Clinical Spectrum, Molecular Pathogenesis, and Cancer Predisposition. Mol. Syndromol. 2017, 8, 4–23. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.; Ranzani, G.N.; Amadori, D.; Herlea, V.; Matei, I.; Barbanti-Brodano, G.; Negrini, M. Somatic frameshift mutations in the Bloom syndrome BLM gene are frequent in sporadic gastric carcinomas with microsatellite mutator phenotype. BMC Genet. 2001, 2, 14. [Google Scholar] [CrossRef]
- El Abed, R.; Bourdon, V.; Voskoboinik, I.; Omri, H.; Youssef, Y.B.; Laatiri, M.A.; Huiart, L.; Eisinger, F.; Rabayrol, L.; Frenay, M.; et al. Molecular study of the perforin gene in familial hematological malignancies. Hered. Cancer Clin. Pract. 2011, 9, 9. [Google Scholar] [CrossRef]
- Tedaldi, G.; Tebaldi, M.; Zampiga, V.; Danesi, R.; Arcangeli, V.; Ravegnani, M.; Cangini, I.; Pirini, F.; Petracci, E.; Rocca, A.; et al. Multiple-gene panel analysis in a case series of 255 women with hereditary breast and ovarian cancer. Oncotarget 2017, 8, 47064–47075. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef] [PubMed]
- Plon, S.E.; Eccles, D.M.; Easton, D.; Foulkes, W.D.; Genuardi, M.; Greenblatt, M.S.; Hogervorst, F.B.L.; Hoogerbrugge, N.; Spurdle, A.B.; Tavtigian, S.V.; et al. Sequence variant classification and reporting: Recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum. Mutat. 2008, 29, 1282–1291. [Google Scholar] [CrossRef] [PubMed]
- Global Variome Shared LOVD-CDH1. Available online: https://databases.lovd.nl/shared/genes/CDH1 (accessed on 30 July 2019).
- dbSNP–NCBI–NIH. Available online: https://www.ncbi.nlm.nih.gov/snp (accessed on 30 July 2019).
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Henikoff, S.; Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 2009, 4, 1073–1081. [Google Scholar] [CrossRef] [PubMed]
| AIP | ALK | APC | ATM | BAP1 | BLM | BMPR1A | BRCA1 | BRCA2 | BRIP1 |
| BUB1B | CDC73 | CDH1 | CDK4 | CDKN1C | CDKN2A | CEBPA | CEP57 | CHEK2 | CYLD |
| DDB2 | DICER1 | DIS3L2 | EGFR | EPCAM | ERCC2 | ERCC3 | ERCC4 | ERCC5 | EXT1 |
| EXT2 | EZH2 | FANCA | FANCB | FANCC | FANCD2 | FANCE | FANCF | FANCG | FANCI |
| FANCL | FANCM | FH | FLCN | GATA2 | GPC3 | HNF1A | HRAS | KIT | MAX |
| MEN1 | MET | MLH1 | MSH2 | MSH6 | MUTYH | NBN | NF1 | NF2 | NSD1 |
| PALB2 | PHOX2B | PMS1 | PMS2 | PRF1 | PRKAR1A | PTCH1 | PTEN | RAD51C | RAD51D |
| RB1 | RECQL4 | RET | RHBDF2 | RUNX1 | SBDS | SDHAF2 | SDHB | SDHC | SDHD |
| SLX4 | SMAD4 | SMARCB1 | STK11 | SUFU | TMEM127 | TP53 | TSC1 | TSC2 | VHL |
| WRN | WT1 | XPA | XPC |
| Patient ID | Sex | Selection Criteria | Cancer | Age at Diagnosis | Gene | Exon | CDNA | Protein | IARC Class | DbSNP | ClinVar | Literature |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
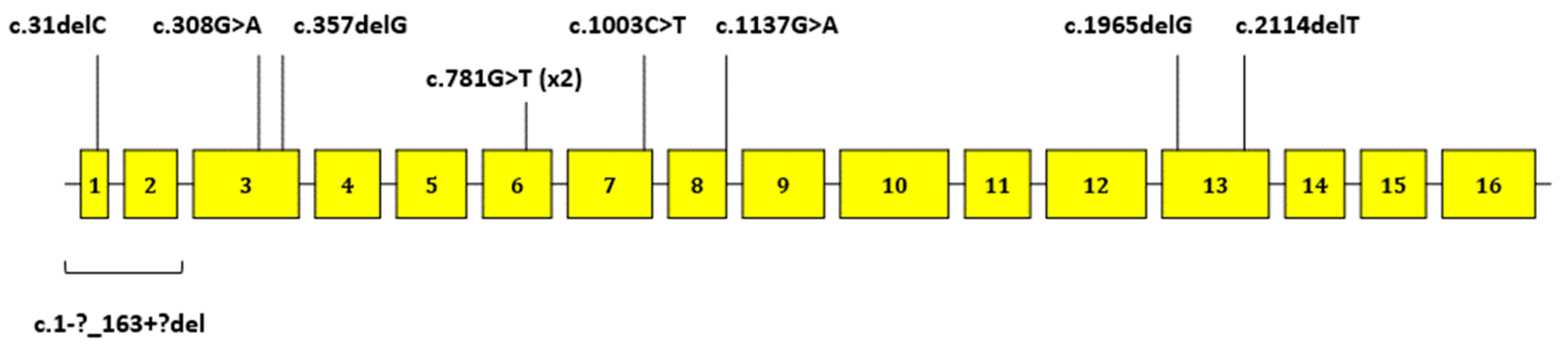
| BM112 | F | II | DGC | 37 | CDH1 | 1–2 | c.1-?_163+?del | p.? | 5 | − | − | Oliveira C. et al. 2009 [14] |
| BM73 | F | III | LBC | 52 | CDH1 | 1 | c.31delC | p.(Leu11Cysfs*45) | 4 | − | − | − |
| BM37 | F | II | DGC | 37 | CDH1 | 3 | c.308G>A | p.Trp103* | 5 | − | pathogenic | − |
| BM100 | M | I | DGC | 58 | CDH1 | 3 | c.360delG | p.(His121Thrfs*94) | 4 | − | − | − |
| BM81 | F | II | DGC | 18 | CDH1 | 6 | c.781G>T | p.Glu261* | 5 | rs121964873 | pathogenic | Berx G. et al. 1995 [31] |
| BM115 | F | II | DGC | 31 | CDH1 | 6 | c.781G>T | p.Glu261* | 5 | rs121964873 | pathogenic | Berx G. et al. 1995 [31] |
| BM60 | M | II | DGC | 39 | CDH1 | 7 | c.1003C>T | p.Arg335* | 5 | rs587780784 | pathogenic | Jonsson B.A. et al. 2002 [32] |
| BM119 | M | II | DGC | 33 | CDH1 | 8 | c.1137G>A | p.Thr379= | 4–5 | rs587783050 | pathogenic/likely pathogenic | Frebourg T. et al. 2006 [36] |
| BM74 | M | I | DGC | 59 | CDH1 | 13 | c.1965delG | p.(Met656Trpfs*3) | 4 | − | − | − |
| BM45 | M | I | DGC | 47 | CDH1 | 13 | c.2114delT | p.(Leu705Cysfs*17) | 4 | − | − | − |
| Patient ID. | Sex | Selection Criteria | Cancer(s) | Age at Diagnosis | Gene | Exon | CDNA | Protein | IARC Class | DbSNP | ClinVar | Literature |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BM10 | M | V | IGC | 57 | MSH2 | 3 | c.367-?_645+?del | p.? | 5 | − | pathogenic | Wijnen J. et al. 1998 [38] |
| BM90 | M | I | DGC | 73 | PMS2 | 13 | c.2182_2183delAC | p.(Thr728Serfs*7) | 4 | − | − | − |
| BM89 | F | I | DGC | 65 | PRF1 | 3 | c.1122G > A | p.Trp374* | 5 | rs104894176 | pathogenic | Stepp S.E. et al. 1999 [39] |
| BM46 | M | I + VI | IGC | 54 | ATM | 10 | c.1564_1565delGA | p.Glu522Ilefs*43 | 5 | rs587779817 | pathogenic | Lakin N.D. et al. 1996 [40] |
| BM76 | F | II | DGC | 32 | ATM | 14 | c.2192dupA | p.(Tyr731*) | 4 | − | − | Saviozzi S. et al. 2003 [41] |
| BM38 | M | VI | IGC | 60 | BRCA2 | 11 | c.6037A > T | p.Lys2013* | 5 | rs80358840 | pathogenic | Meindl A. et al. 2002 [42] |
| BM24 | M | VII | GP | 52 | BMPR1A | 3 | c.34G > T | p.(Gly12*) | 4 | − | − | − |
| BM47 | F | I | LBC, DGC | 50, 54 | BLM | 11 | c.2395delT | p.(Cys799Valfs*16) | 4 | − | − | − |
| A530 | F | IV | LBC, LBC | 62, 66 | PALB2 | 4 | c.535C > T | p.(Gln179*) | 4 | − | − | − |
| BM126 | F | IV | LBC | 62 | PALB2 | 7 | c.2718G > A | p.Trp906* | 4–5 | rs180177122 | pathogenic/likely pathogenic | Casadei S. et al. 2011 [43] |
| BM110 | F | I | DGC | 47 | BRCA1 | 7 | c.406delA | p.Arg136Aspfs*27 | 5 | rs886040196 | pathogenic | Kluska A. et al. 2015 [44] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tedaldi, G.; Pirini, F.; Tebaldi, M.; Zampiga, V.; Cangini, I.; Danesi, R.; Arcangeli, V.; Ravegnani, M.; Abou Khouzam, R.; Molinari, C.; et al. Multigene Panel Testing Increases the Number of Loci Associated with Gastric Cancer Predisposition. Cancers 2019, 11, 1340. https://doi.org/10.3390/cancers11091340
Tedaldi G, Pirini F, Tebaldi M, Zampiga V, Cangini I, Danesi R, Arcangeli V, Ravegnani M, Abou Khouzam R, Molinari C, et al. Multigene Panel Testing Increases the Number of Loci Associated with Gastric Cancer Predisposition. Cancers. 2019; 11(9):1340. https://doi.org/10.3390/cancers11091340
Chicago/Turabian StyleTedaldi, Gianluca, Francesca Pirini, Michela Tebaldi, Valentina Zampiga, Ilaria Cangini, Rita Danesi, Valentina Arcangeli, Mila Ravegnani, Raefa Abou Khouzam, Chiara Molinari, and et al. 2019. "Multigene Panel Testing Increases the Number of Loci Associated with Gastric Cancer Predisposition" Cancers 11, no. 9: 1340. https://doi.org/10.3390/cancers11091340
APA StyleTedaldi, G., Pirini, F., Tebaldi, M., Zampiga, V., Cangini, I., Danesi, R., Arcangeli, V., Ravegnani, M., Abou Khouzam, R., Molinari, C., Oliveira, C., Morgagni, P., Saragoni, L., Bencivenga, M., Ulivi, P., Amadori, D., Martinelli, G., Falcini, F., Ranzani, G. N., & Calistri, D. (2019). Multigene Panel Testing Increases the Number of Loci Associated with Gastric Cancer Predisposition. Cancers, 11(9), 1340. https://doi.org/10.3390/cancers11091340








