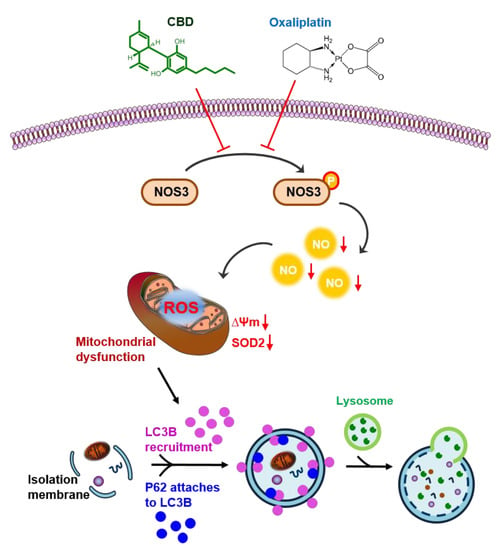
Cannabidiol Overcomes Oxaliplatin Resistance by Enhancing NOS3- and SOD2-Induced Autophagy in Human Colorectal Cancer Cells
Abstract
1. Introduction
2. Results
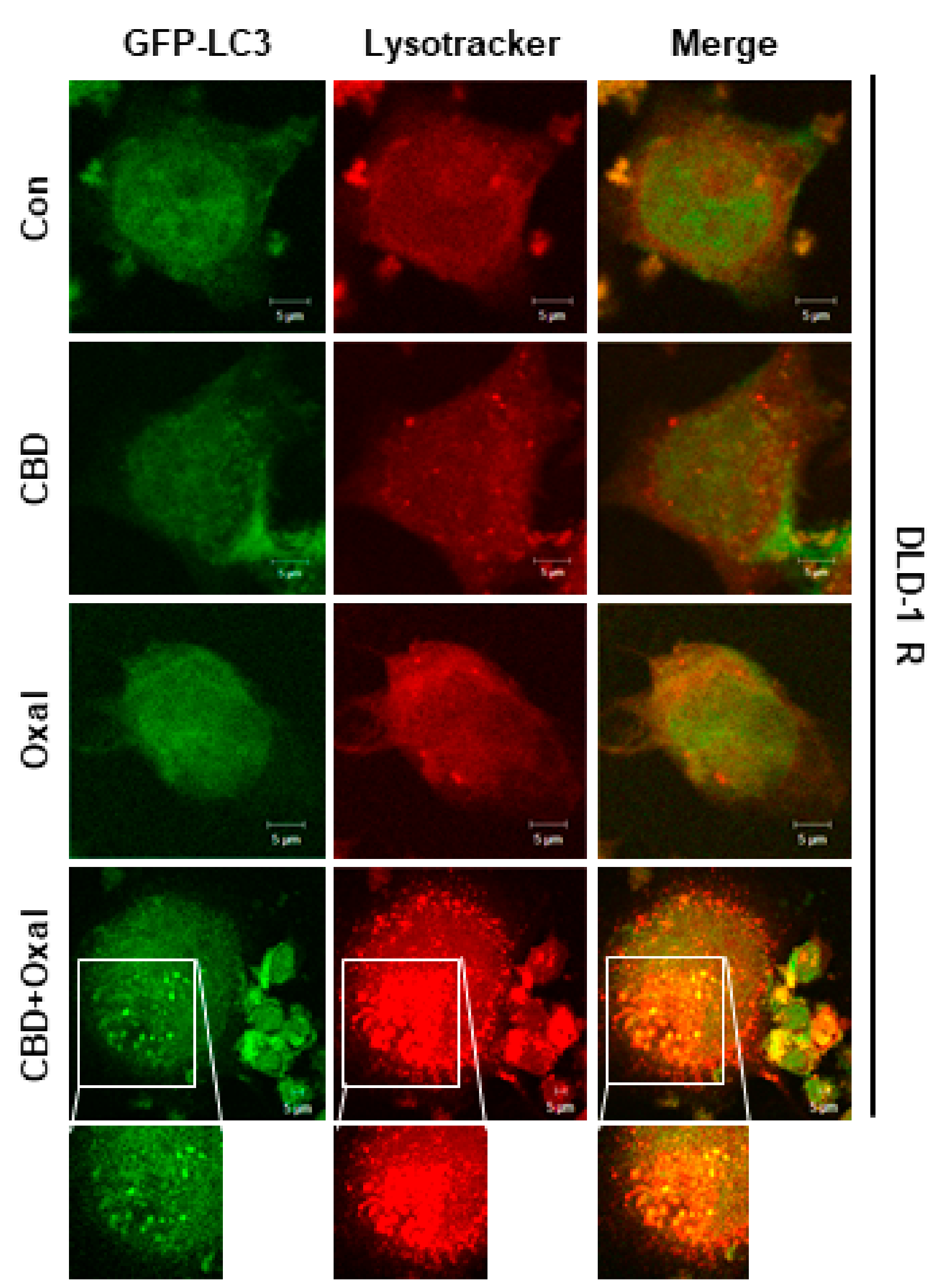
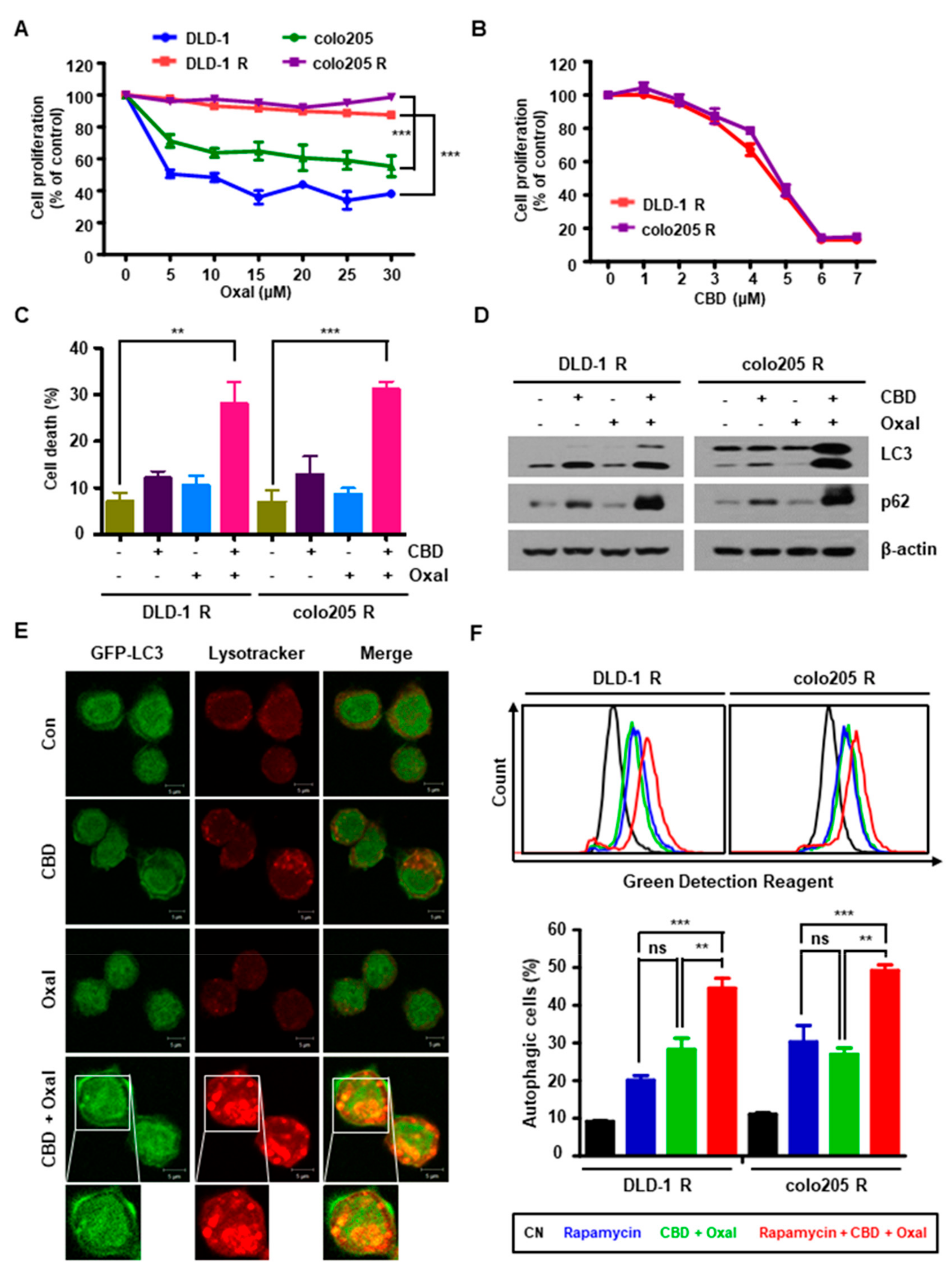
2.1. CBD and Oxaliplatin Reduce Proliferation and Induce Autophagic Death of Oxaliplatin-Resistant CRC Cells
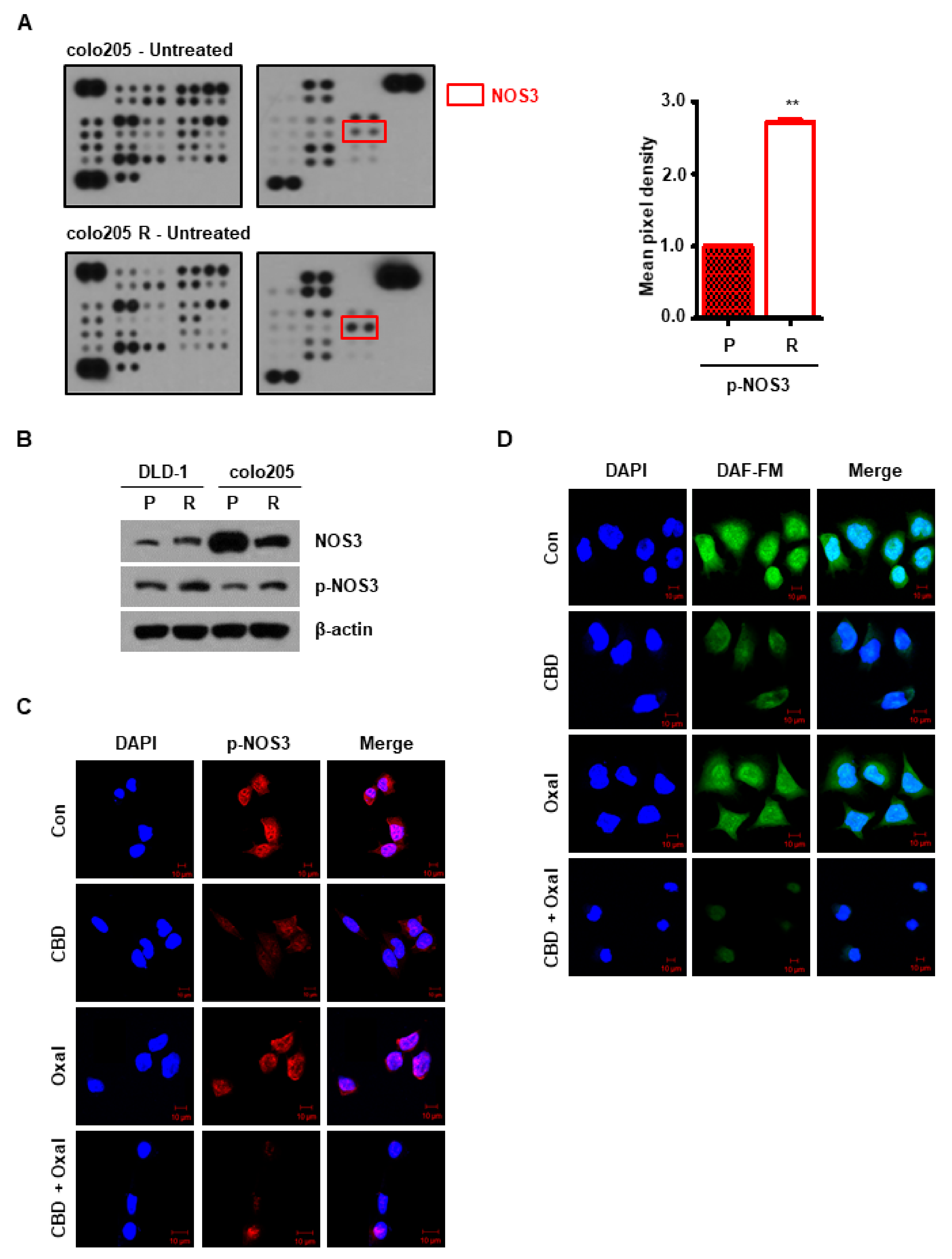
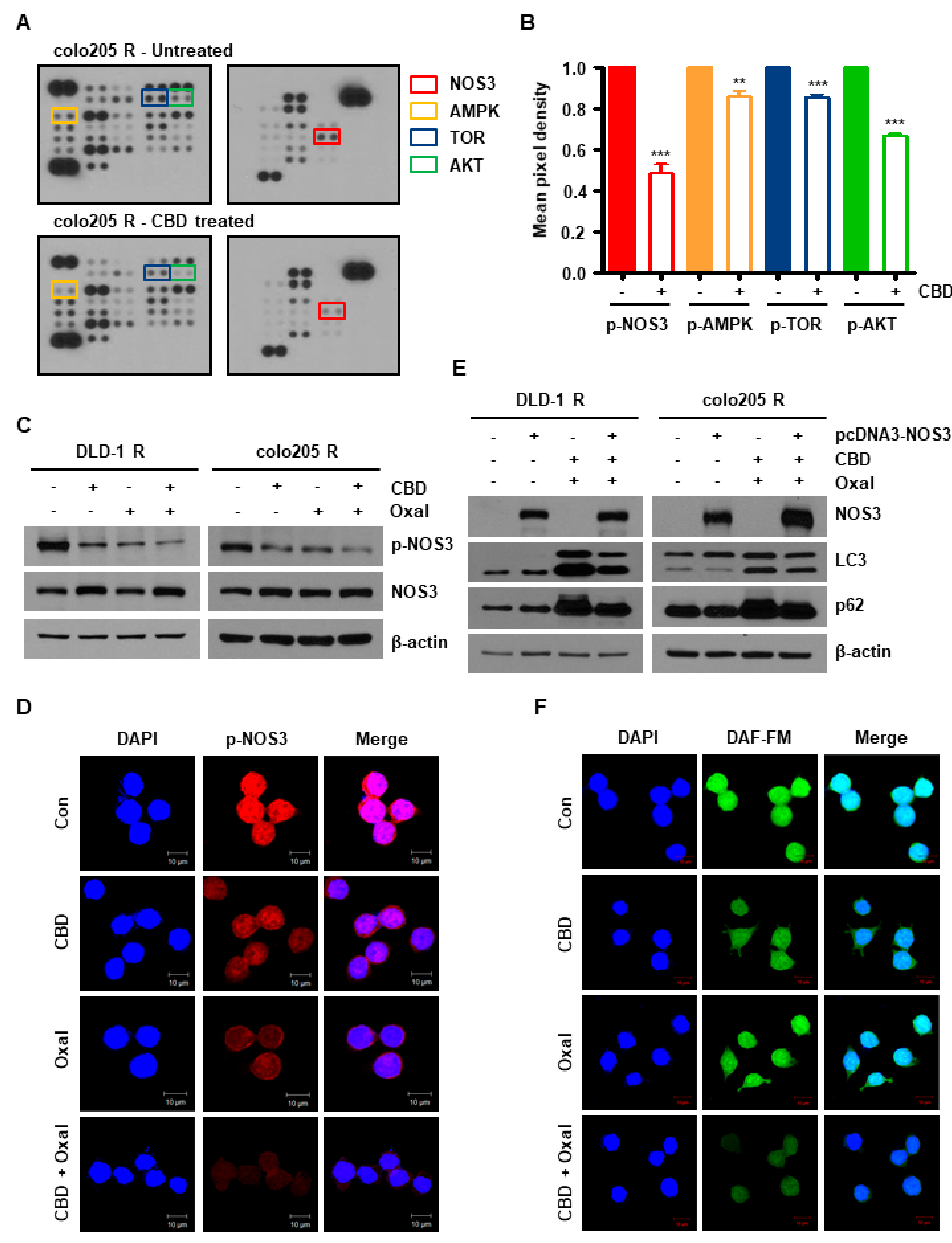
2.2. NOS3 Is Associated with Oxaliplatin Resistance and Decreased NOS3 Activity Induces Autophagy
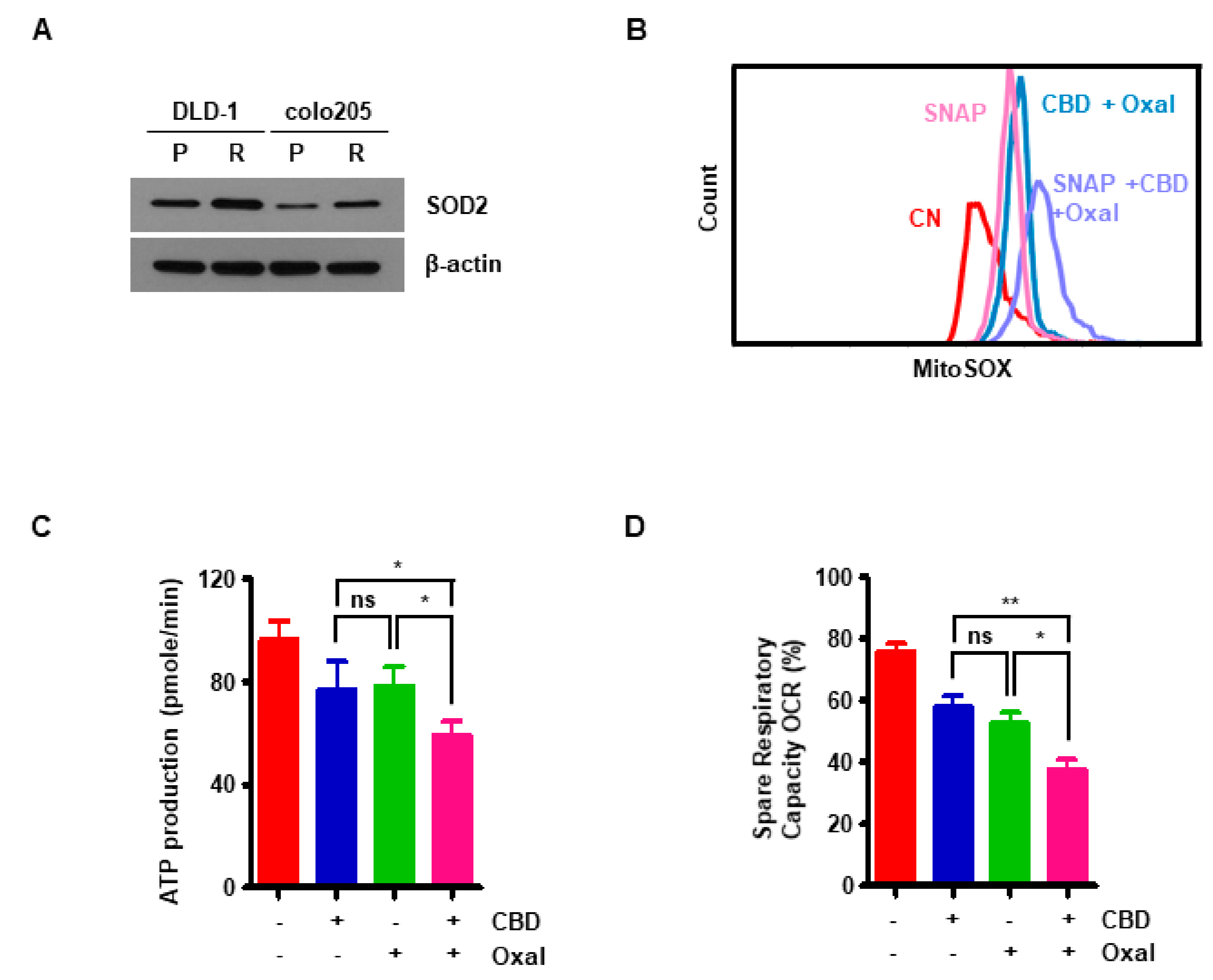
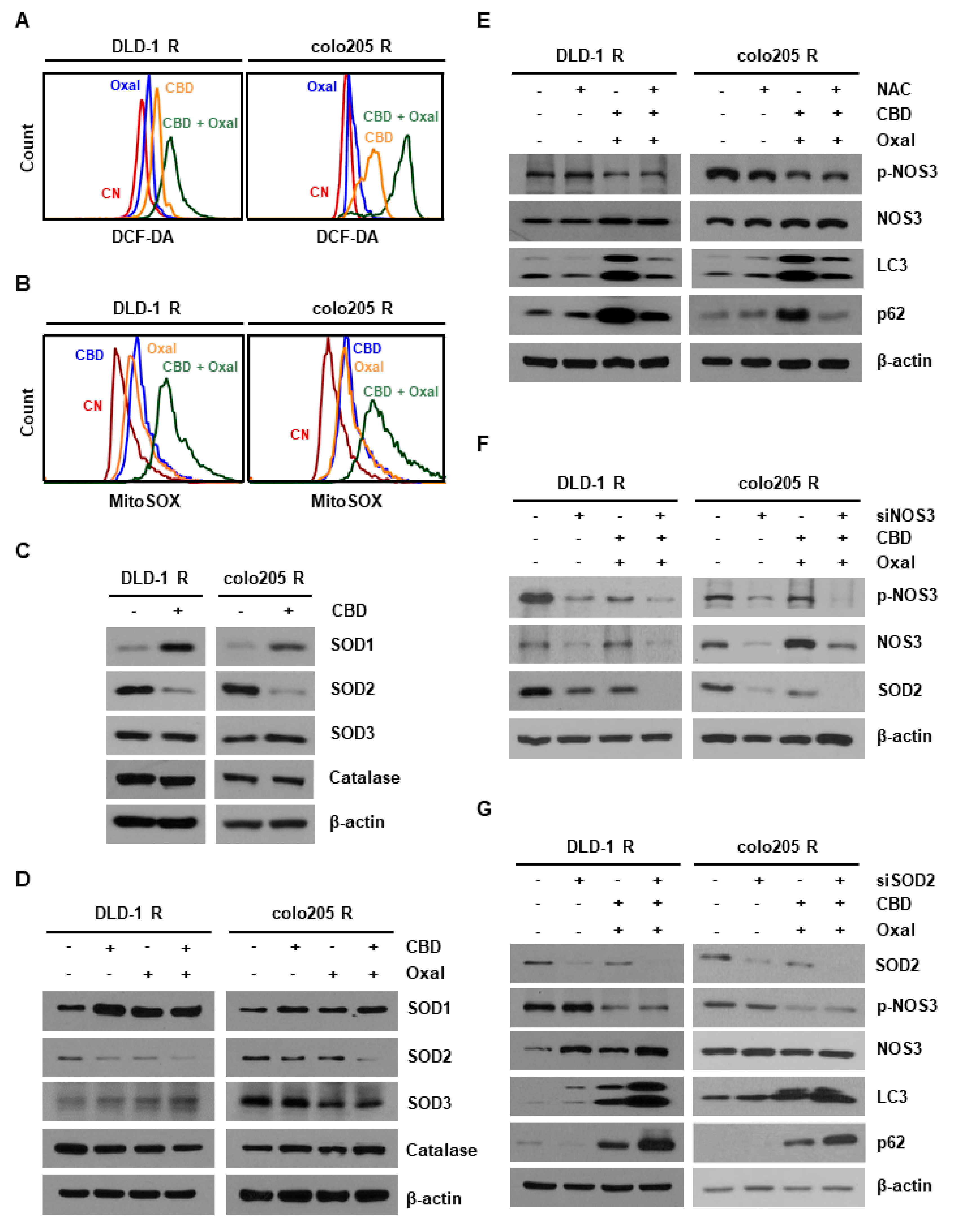
2.3. Reduction of Antioxidant SOD2 by Oxaliplatin and CBD Induces Reactive Oxygen Species (ROS) Overproduction
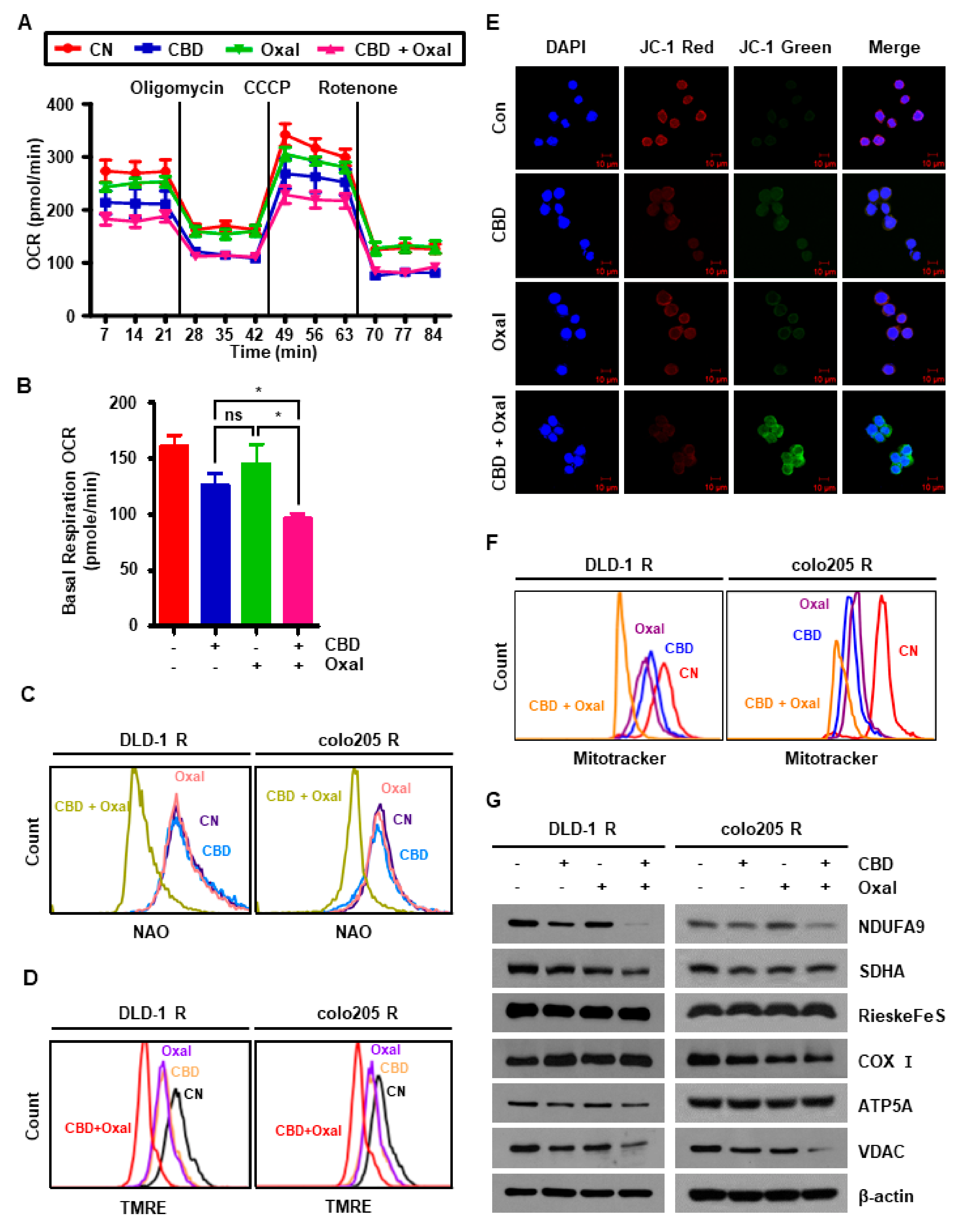
2.4. CBD and Oxaliplatin Treatment Leads to Mitochondrial Dysfunction
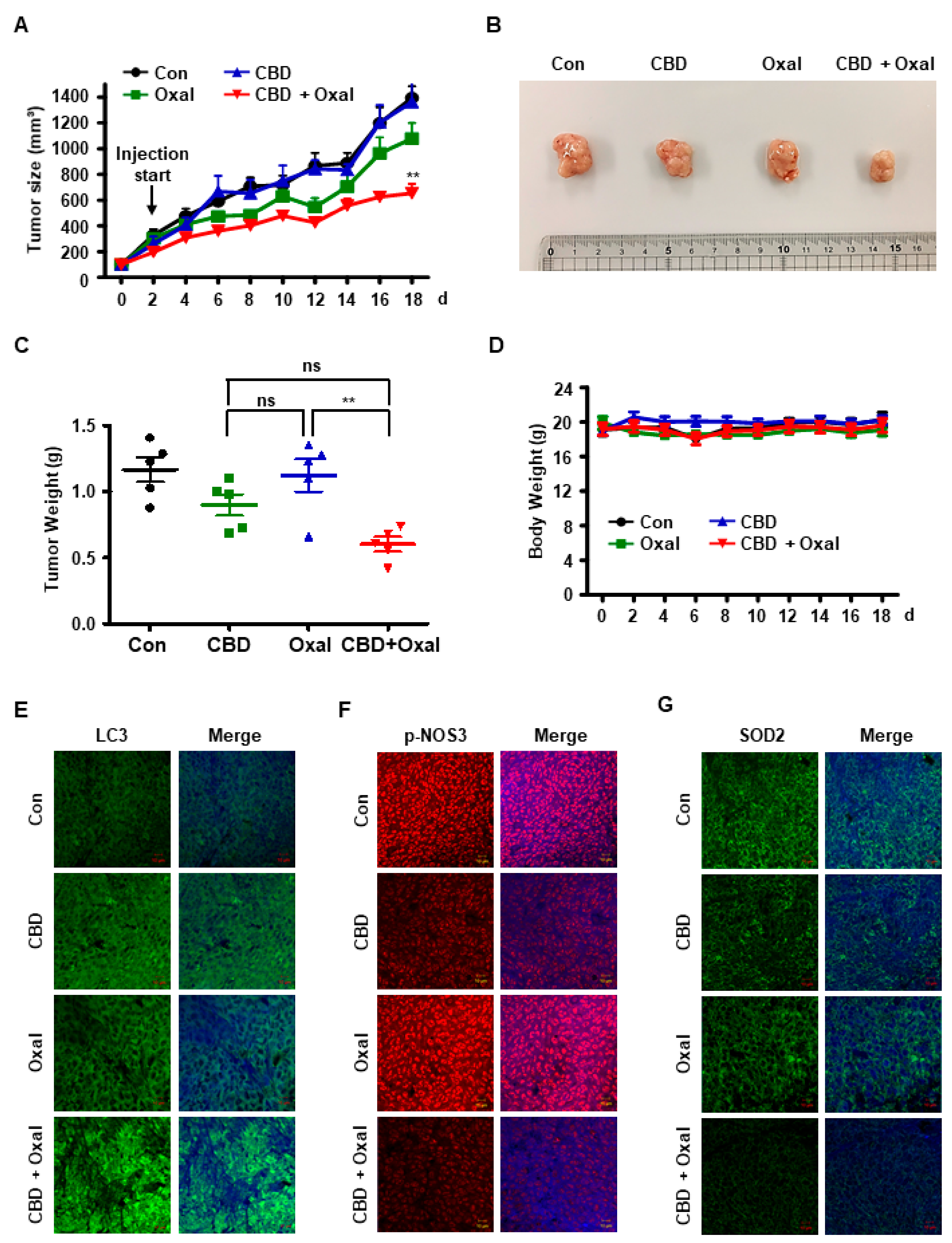
2.5. CBD and Oxaliplatin Inhibit Tumor Growth In Vivo by Decreasing Phospho-NOS3 and SOD2 Levels and Inducing Subsequent Autophagy
3. Discussion
4. Materials and Methods
4.1. Cell Lines and Cell Culture
4.2. Establishment of Oxaliplatin-Resistant Cell Lines
4.3. Reagents and Antibodies
4.4. Cell Proliferation Assay
4.5. Immunoblotting
4.6. Green Fluorescent Protein (GFP)-LC3 Puncta
4.7. Autophagic Activity
4.8. Analysis of Cell Death
4.9. Human Phospho-Kinase Array
4.10. Immunofluorescence Staining
4.11. Measurement of ROS
4.12. Transfection
4.13. OCR
4.14. Determination of Mitochondrial Function
4.15. In Vivo Tumor Xenograft Model
4.16. Immunohistochemistry (IHC)
4.17. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Goodwin, R.A.; Asmis, T.R. Overview of systemic therapy for colorectal cancer. Clin. Colon Rectal Surg. 2009, 22, 251–256. [Google Scholar] [CrossRef]
- Bendell, J.C.; Bekaii-Saab, T.S.; Cohn, A.L.; Hurwitz, H.I.; Kozloff, M.; Tezcan, H.; Roach, N.; Mun, Y.; Fish, S.; Flick, E.D.; et al. Treatment patterns and clinical outcomes in patients with metastatic colorectal cancer initially treated with FOLFOX-bevacizumab or FOLFIRI-bevacizumab: Results from ARIES, a bevacizumab observational cohort study. Oncologist 2012, 17, 1486–1495. [Google Scholar] [CrossRef] [PubMed]
- Hua, Y.; Zhu, Y.; Zhang, J.; Zhu, Z.; Ning, Z.; Chen, H.; Liu, L.; Chen, Z.; Meng, Z. miR-122 Targets X-Linked Inhibitor of Apoptosis Protein to Sensitize Oxaliplatin-Resistant Colorectal Cancer Cells to Oxaliplatin-Mediated Cytotoxicity. Cell. Physiol. Biochem. 2018, 51, 2148–2159. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ghareeb, W.M.; Zhang, Y.; Yu, Q.; Lu, X.; Huang, Y.; Huang, S.; Sun, Y.; Chi, P. Hypermethylated and downregulated MEIS2 are involved in stemness properties and oxaliplatin-based chemotherapy resistance of colorectal cancer. J. Cell. Physiol. 2019. [Google Scholar] [CrossRef]
- Hammond, W.A.; Swaika, A.; Mody, K. Pharmacologic resistance in colorectal cancer: A review. Ther. Adv. Med. Oncol. 2016, 8, 57–84. [Google Scholar] [CrossRef]
- Liang, H.; Xu, Y.; Zhang, Q.; Yang, Y.; Mou, Y.; Gao, Y.; Chen, R.; Chen, C.; Dai, P. MiR-483-3p regulates oxaliplatin resistance by targeting FAM171B in human colorectal cancer cells. Artif. Cells Nanomed. Biotechnol. 2019, 47, 725–736. [Google Scholar] [CrossRef]
- Kim, N.; Jeong, S.; Jing, K.; Shin, S.; Kim, S.; Heo, J.Y.; Kweon, G.R.; Park, S.K.; Wu, T.; Park, J.I.; et al. Docosahexaenoic Acid Induces Cell Death in Human Non-Small Cell Lung Cancer Cells by Repressing mTOR via AMPK Activation and PI3K/Akt Inhibition. BioMed Res. Int. 2015, 2015, 239764. [Google Scholar] [CrossRef]
- Skarkova, V.; Kralova, V.; Vitovcova, B.; Rudolf, E. Selected Aspects of Chemoresistance Mechanisms in Colorectal Carcinoma-A Focus on Epithelial-to-Mesenchymal Transition, Autophagy, and Apoptosis. Cells 2019, 8, 234. [Google Scholar] [CrossRef]
- Forstermann, U.; Sessa, W.C. Nitric oxide synthases: Regulation and function. Eur. Heart J. 2012, 33, 829–837. [Google Scholar] [CrossRef]
- Jadeski, L.C.; Hum, K.O.; Chakraborty, C.; Lala, P.K. Nitric oxide promotes murine mammary tumour growth and metastasis by stimulating tumour cell migration, invasiveness and angiogenesis. Int. J. Cancer 2000, 86, 30–39. [Google Scholar] [CrossRef]
- Penarando, J.; Lopez-Sanchez, L.M.; Mena, R.; Guil-Luna, S.; Conde, F.; Hernandez, V.; Toledano, M.; Gudino, V.; Raponi, M.; Billard, C.; et al. A role for endothelial nitric oxide synthase in intestinal stem cell proliferation and mesenchymal colorectal cancer. BMC Biol. 2018, 16, 3. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.P.; Gu, M.; Agarwal, R. Silibinin inhibits colorectal cancer growth by inhibiting tumor cell proliferation and angiogenesis. Cancer Res. 2008, 68, 2043–2050. [Google Scholar] [CrossRef] [PubMed]
- Jeong, S.; Yun, H.K.; Jeong, Y.A.; Jo, M.J.; Kang, S.H.; Kim, J.L.; Kim, D.Y.; Park, S.H.; Kim, B.R.; Na, Y.J.; et al. Cannabidiol-induced apoptosis is mediated by activation of Noxa in human colorectal cancer cells. Cancer Lett. 2019, 447, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Kogan, N.M.; Mechoulam, R. Cannabinoids in health and disease. Dialogues Clin. Neurosci. 2007, 9, 413–430. [Google Scholar] [PubMed]
- Lopez-Valero, I.; Torres, S.; Salazar-Roa, M.; Garcia-Taboada, E.; Hernandez-Tiedra, S.; Guzman, M.; Sepulveda, J.M.; Velasco, G.; Lorente, M. Optimization of a preclinical therapy of cannabinoids in combination with temozolomide against glioma. Biochem. Pharmacol. 2018, 157, 275–284. [Google Scholar] [CrossRef]
- Schlafli, A.M.; Berezowska, S.; Adams, O.; Langer, R.; Tschan, M.P. Reliable LC3 and p62 autophagy marker detection in formalin fixed paraffin embedded human tissue by immunohistochemistry. Eur. J. Histochem. 2015, 59, 2481. [Google Scholar] [CrossRef]
- Barouch, L.A.; Harrison, R.W.; Skaf, M.W.; Rosas, G.O.; Cappola, T.P.; Kobeissi, Z.A.; Hobai, I.A.; Lemmon, C.A.; Burnett, A.L.; O’Rourke, B.; et al. Nitric oxide regulates the heart by spatial confinement of nitric oxide synthase isoforms. Nature 2002, 416, 337–339. [Google Scholar] [CrossRef]
- Takimoto, E.; Champion, H.C.; Li, M.; Ren, S.; Rodriguez, E.R.; Tavazzi, B.; Lazzarino, G.; Paolocci, N.; Gabrielson, K.L.; Wang, Y.; et al. Oxidant stress from nitric oxide synthase-3 uncoupling stimulates cardiac pathologic remodeling from chronic pressure load. J. Clin. Invest. 2005, 115, 1221–1231. [Google Scholar] [CrossRef]
- Vannini, F.; Kashfi, K.; Nath, N. The dual role of iNOS in cancer. Redox Biol. 2015, 6, 334–343. [Google Scholar] [CrossRef]
- Tan, S.; Peng, X.; Peng, W.; Zhao, Y.; Wei, Y. Enhancement of oxaliplatin-induced cell apoptosis and tumor suppression by 3-methyladenine in colon cancer. Oncology Lett. 2015, 9, 2056–2062. [Google Scholar] [CrossRef] [PubMed]
- Hector, S.; Bolanowska-Higdon, W.; Zdanowicz, J.; Hitt, S.; Pendyala, L. In vitro studies on the mechanisms of oxaliplatin resistance. Cancer Chemother. Pharmacol. 2001, 48, 398–406. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, S. Platinum-DNA interactions and subsequent cellular processes controlling sensitivity to anticancer platinum complexes. Chem. Biodivers. 2010, 7, 543–566. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.Y.; Sales, K.M.; Fuller, B.; Seifalian, A.M.; Winslet, M.C. Apoptosis and colorectal cancer: Implications for therapy. Trends. Mol. Med. 2009, 15, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Stuehr, D.J. Enzymes of the L-arginine to nitric oxide pathway. J. Nutr. 2004, 134, 2748S–2751S; discussion 2765S–2767S. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Ma, C.; Shi, X.; Wen, Z.; Li, D.; Sun, M.; Ding, H. Effect of nitric oxide synthase on multiple drug resistance is related to Wnt signaling in non-small cell lung cancer. Oncol. Rep. 2014, 32, 1703–1708. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Xie, Z.X. Nitric oxide synthase expression and multiple drug resistance in patients with acute leukemia. Zhong Nan Da Xue Xue Bao Yi Xue Ban 2004, 29, 25–27. [Google Scholar] [PubMed]
- Bartz, R.R.; Suliman, H.B.; Piantadosi, C.A. Redox mechanisms of cardiomyocyte mitochondrial protection. Front. Physiol. 2015, 6, 291. [Google Scholar] [CrossRef]
- Cimolai, M.C.; Alvarez, S.; Bode, C.; Bugger, H. Mitochondrial Mechanisms in Septic Cardiomyopathy. Int. J. Mol. Sci. 2015, 16, 17763–17778. [Google Scholar] [CrossRef]
- Valerio, A.; Nisoli, E. Nitric oxide, interorganelle communication, and energy flow: A novel route to slow aging. Front. Cell Dev. Biol. 2015, 3, 6. [Google Scholar] [CrossRef]
- Heymann, D. Autophagy: A protective mechanism in response to stress and inflammation. Curr. Opin. Investig. Drugs. 2006, 7, 443–450. [Google Scholar] [PubMed]
- Das, G.; Shravage, B.V.; Baehrecke, E.H. Regulation and function of autophagy during cell survival and cell death. Cold Spring Harb. Perspect. Biol. 2012, 4. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Rehman, S.K.; Zhang, W.; Wen, A.; Yao, L.; Zhang, J. Autophagy is a therapeutic target in anticancer drug resistance. Biochim. Biophys. Acta 2010, 1806, 220–229. [Google Scholar] [CrossRef] [PubMed]
- Yan, M.M.; Ni, J.D.; Song, D.; Ding, M.; Huang, J. Interplay between unfolded protein response and autophagy promotes tumor drug resistance. Oncol. Lett. 2015, 10, 1959–1969. [Google Scholar] [CrossRef] [PubMed]
- Areti, A.; Komirishetty, P.; Akuthota, M.; Malik, R.A.; Kumar, A. Melatonin prevents mitochondrial dysfunction and promotes neuroprotection by inducing autophagy during oxaliplatin-evoked peripheral neuropathy. J. Pineal Res. 2017, 62. [Google Scholar] [CrossRef] [PubMed]
- Kidwell, K.M.; Yothers, G.; Ganz, P.A.; Land, S.R.; Ko, C.Y.; Cecchini, R.S.; Kopec, J.A.; Wolmark, N. Long-term neurotoxicity effects of oxaliplatin added to fluorouracil and leucovorin as adjuvant therapy for colon cancer: Results from National Surgical Adjuvant Breast and Bowel Project trials C-07 and LTS-01. Cancer 2012, 118, 5614–5622. [Google Scholar] [CrossRef] [PubMed]
- Hamelink, C.; Hampson, A.; Wink, D.A.; Eiden, L.E.; Eskay, R.L. Comparison of cannabidiol, antioxidants, and diuretics in reversing binge ethanol-induced neurotoxicity. J. Pharmacol. Exp. Ther. 2005, 314, 780–788. [Google Scholar] [CrossRef] [PubMed]
- Kozela, E.; Pietr, M.; Juknat, A.; Rimmerman, N.; Levy, R.; Vogel, Z. Cannabinoids Delta(9)-tetrahydrocannabinol and cannabidiol differentially inhibit the lipopolysaccharide-activated NF-kappaB and interferon-beta/STAT proinflammatory pathways in BV-2 microglial cells. J. Biol. Chem. 2010, 285, 1616–1626. [Google Scholar] [CrossRef]
- Schonhofen, P.; de Medeiros, L.M.; Bristot, I.J.; Lopes, F.M.; De Bastiani, M.A.; Kapczinski, F.; Crippa, J.A.; Castro, M.A.; Parsons, R.B.; Klamt, F. Cannabidiol Exposure During Neuronal Differentiation Sensitizes Cells Against Redox-Active Neurotoxins. Mol. Neurobiol. 2015, 52, 26–37. [Google Scholar] [CrossRef]
- Kim, H.J.; Jo, M.J.; Kim, B.R.; Kim, J.L.; Jeong, Y.A.; Na, Y.J.; Park, S.H.; Lee, S.Y.; Lee, D.H.; Lee, H.S.; et al. Reactive oxygen species modulator-1 (Romo1) predicts unfavorable prognosis in colorectal cancer patients. PLoS ONE 2017, 12, e0176834. [Google Scholar] [CrossRef]
- Kim, B.R.; Jeong, Y.A.; Jo, M.J.; Park, S.H.; Na, Y.J.; Kim, J.L.; Jeong, S.; Yun, H.K.; Kang, S.H.; Lee, D.H.; et al. Genipin enhances the therapeutic effects of oxaliplatin by upregulating BIM in colorectal cancer. Mol. Cancer Ther. 2019. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jeong, S.; Kim, B.G.; Kim, D.Y.; Kim, B.R.; Kim, J.L.; Park, S.H.; Na, Y.J.; Jo, M.J.; Yun, H.K.; Jeong, Y.A.; et al. Cannabidiol Overcomes Oxaliplatin Resistance by Enhancing NOS3- and SOD2-Induced Autophagy in Human Colorectal Cancer Cells. Cancers 2019, 11, 781. https://doi.org/10.3390/cancers11060781
Jeong S, Kim BG, Kim DY, Kim BR, Kim JL, Park SH, Na YJ, Jo MJ, Yun HK, Jeong YA, et al. Cannabidiol Overcomes Oxaliplatin Resistance by Enhancing NOS3- and SOD2-Induced Autophagy in Human Colorectal Cancer Cells. Cancers. 2019; 11(6):781. https://doi.org/10.3390/cancers11060781
Chicago/Turabian StyleJeong, Soyeon, Bu Gyeom Kim, Dae Yeong Kim, Bo Ram Kim, Jung Lim Kim, Seong Hye Park, Yoo Jin Na, Min Jee Jo, Hye Kyeong Yun, Yoon A. Jeong, and et al. 2019. "Cannabidiol Overcomes Oxaliplatin Resistance by Enhancing NOS3- and SOD2-Induced Autophagy in Human Colorectal Cancer Cells" Cancers 11, no. 6: 781. https://doi.org/10.3390/cancers11060781
APA StyleJeong, S., Kim, B. G., Kim, D. Y., Kim, B. R., Kim, J. L., Park, S. H., Na, Y. J., Jo, M. J., Yun, H. K., Jeong, Y. A., Kim, H. J., Lee, S. I., Kim, H. D., Kim, D. H., Oh, S. C., & Lee, D.-H. (2019). Cannabidiol Overcomes Oxaliplatin Resistance by Enhancing NOS3- and SOD2-Induced Autophagy in Human Colorectal Cancer Cells. Cancers, 11(6), 781. https://doi.org/10.3390/cancers11060781