Homeobox Genes and Hepatocellular Carcinoma
Abstract
1. Epidemiology
2. Etiologies
3. Treatments
4. Homeobox Genes
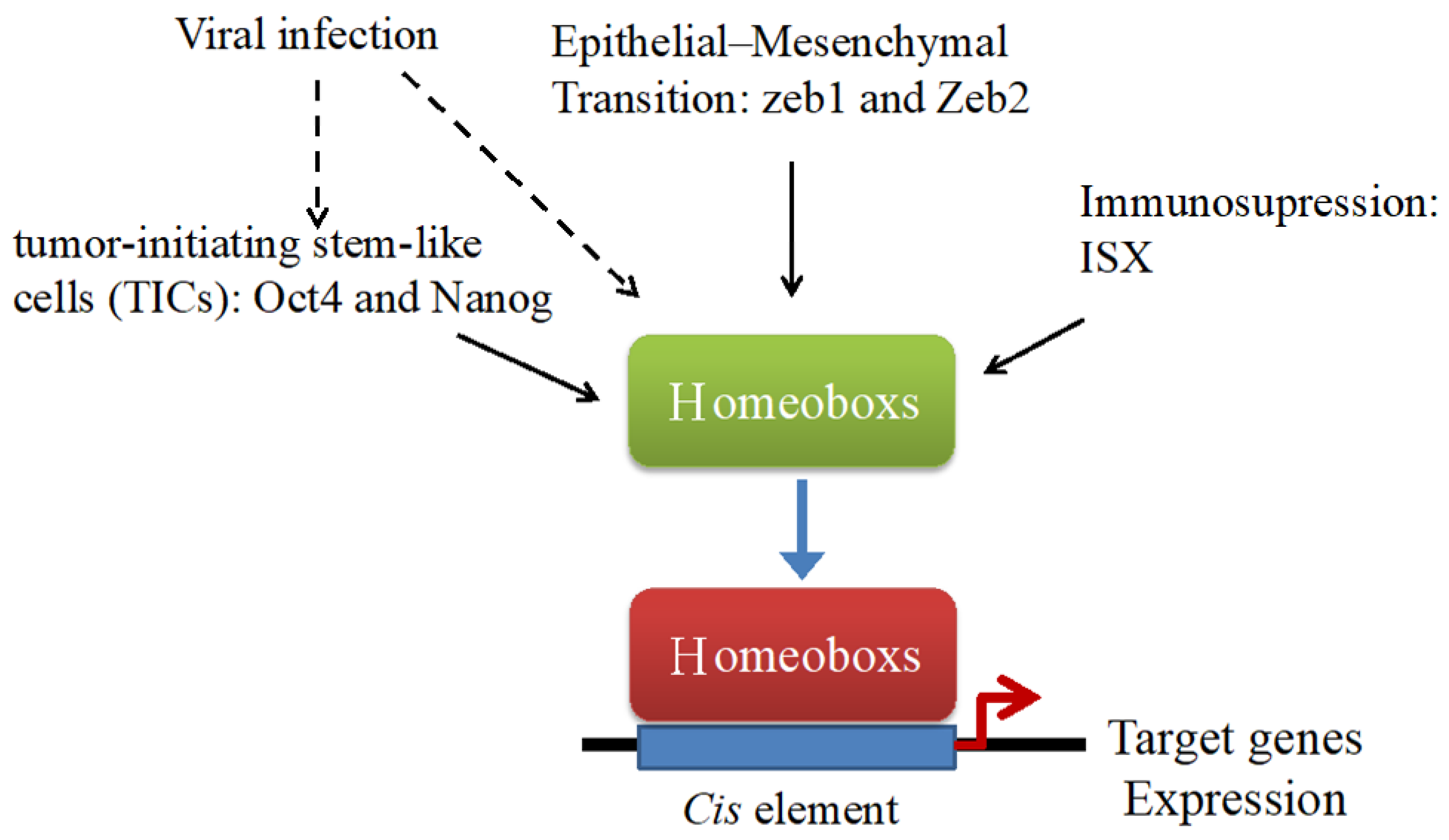
5. Homeobox Genes (Oct4 and Nanog) in TICs of HCC
6. Homeobox in EMT of HCC
7. Homeobox Genes and Immunotolerance in HCC
8. Homeobox Genes in HBV- and HCV-Associated HCC
9. Conclusions
Funding
Conflicts of Interest
Abbreviations
| Hepatocellular carcinoma (HCC) | transforming growth factor-β (TGF-β) |
| nonalcoholic fatty liver disease (NAFLD) | reactive oxygen species (ROS) |
| hepatitis B virus (HBV) | Receptor for activated C kinase1 (RACK1) |
| hepatitis C virus (HCV) | Myeloid cell leukemia-1 (Mcl-1) |
| HBV X protein (HBx) | Transforming acidic coiled-coil protein 3 (TACC3) |
| Aflatoxin B1 (AFB1) | GATA transcription factor 5 (GATA5) |
| tumor necrosis factor-α (TNF-α) | Level of Atonal homolog 8 (ATOH8) |
| interleukin 6 (IL-6) | basic-helix-loop-helix (bHLH) |
| programmed death-ligand 1 (PD-L1) | Zinc finger E-box binding Hox 1 (ZEB1) |
| programmed cell death 1 (PD-1) | MYC-associated zinc finger protein (MAZ) |
| tumor-initiating stem-like cells (TICs) | microRNAs (miRNAs) |
| mesenchymal transition (EMT) | long non-coding RNAs (lncRNAs) |
| epithelial cell adhesion molecule (EpCAM) | Long non-coding RNA activated by TGF-β (lncRNA-ATB) |
| octamer-binding transcription factor 4 (Oct4) | intestine-specific homeobox transcription factor (ISX) |
| insulin-like growth factor (IGF) 2 | aryl hydrocarbon receptor (AhR) |
| IGF 1 receptor (IGF1R) | Toll-like receptor 4 (TLR4) |
| vascular endothelial growth factor (VEGF) | Damage-associated molecular patterns (DAMPs) |
References
- Braillon, A. Hepatocellular carcinoma. Lancet 2012, 380, 469. [Google Scholar] [CrossRef]
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef] [PubMed]
- Torre, L.A.; Bray, F.; Siegel, R.L.; Ferlay, J.; Lortet-Tieulent, J.; Jemal, A. Global cancer statistics, 2012. CA Cancer J. Clin. 2015, 65, 87–108. [Google Scholar] [CrossRef] [PubMed]
- El-Serag, H.B. Epidemiology of viral hepatitis and hepatocellular carcinoma. Gastroenterology 2012, 142, 1264–1273.e1. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.W.; Ting, Y.W.; Chan, W.K. Epidemiology of non-alcoholic fatty liver disease-related hepatocellular carcinoma and its implications. JGH Open 2018, 2, 235–241. [Google Scholar] [CrossRef] [PubMed]
- Blumberg, B.S. Australia antigen and the biology of hepatitis B. Science 1977, 197, 17–25. [Google Scholar]
- Kuo, T.C.; Chao, C.C. Hepatitis B virus X protein prevents apoptosis of hepatocellular carcinoma cells by upregulating SATB1 and HURP expression. Biochem. Pharmacol. 2010, 80, 1093–1102. [Google Scholar] [CrossRef]
- Disease, G.B.D.; Injury, I.; Prevalence, C. Global, regional, and national incidence, prevalence, and years lived with disability for 310 diseases and injuries, 1990–2015: A systematic analysis for the Global Burden of Disease Study 2015. Lancet 2016, 388, 1545–1602. [Google Scholar] [CrossRef]
- Hoshida, Y.; Fuchs, B.C.; Bardeesy, N.; Baumert, T.F.; Chung, R.T. Pathogenesis and prevention of hepatitis C virus-induced hepatocellular carcinoma. J. Hepatol. 2014, 61, S79–S90. [Google Scholar] [CrossRef] [PubMed]
- Kalaitzakis, E.; Gunnarsdottir, S.A.; Josefsson, A.; Bjornsson, E. Increased risk for malignant neoplasms among patients with cirrhosis. Clin. Gastroenterol. Hepatol. 2011, 9, 168–174. [Google Scholar] [CrossRef]
- Martin, J.; Dufour, J.F. Tumor suppressor and hepatocellular carcinoma. World J. Gastroenterol. 2008, 14, 1720–1733. [Google Scholar] [CrossRef]
- Caldwell, S.H.; Oelsner, D.H.; Iezzoni, J.C.; Hespenheide, E.E.; Battle, E.H.; Driscoll, C.J. Cryptogenic cirrhosis: Clinical characterization and risk factors for underlying disease. Hepatology 1999, 29, 664–669. [Google Scholar] [CrossRef]
- Llovet, J.M.; Bru, C.; Bruix, J. Prognosis of hepatocellular carcinoma: The BCLC staging classification. Semin. Liver Dis. 1999, 19, 329–338. [Google Scholar] [CrossRef]
- Llovet, J.M.; Fuster, J.; Bruix, J. Intention-to-treat analysis of surgical treatment for early hepatocellular carcinoma: Resection versus transplantation. Hepatology 1999, 30, 1434–1440. [Google Scholar] [CrossRef] [PubMed]
- Bruix, J.; Sherman, M.; American Association for the Study of Liver, D. Management of hepatocellular carcinoma: An update. Hepatology 2011, 53, 1020–1022. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, N.; Yu, Y.; Walsh, W.R.; Yang, J.L. Molecular targeted therapies for cancer: Sorafenib mono-therapy and its combination with other therapies (review). Oncol. Rep. 2012, 27, 1303–1311. [Google Scholar] [CrossRef]
- Wilhelm, S.M.; Adnane, L.; Newell, P.; Villanueva, A.; Llovet, J.M.; Lynch, M. Preclinical overview of sorafenib, a multikinase inhibitor that targets both Raf and VEGF and PDGF receptor tyrosine kinase signaling. Mol. Cancer Ther. 2008, 7, 3129–3140. [Google Scholar] [CrossRef] [PubMed]
- Llovet, J.M.; Ricci, S.; Mazzaferro, V.; Hilgard, P.; Gane, E.; Blanc, J.F.; de Oliveira, A.C.; Santoro, A.; Raoul, J.L.; Forner, A.; et al. Sorafenib in advanced hepatocellular carcinoma. N. Engl. J. Med. 2008, 359, 378–390. [Google Scholar] [CrossRef]
- Kalogeridi, M.A.; Zygogianni, A.; Kyrgias, G.; Kouvaris, J.; Chatziioannou, S.; Kelekis, N.; Kouloulias, V. Role of radiotherapy in the management of hepatocellular carcinoma: A systematic review. World J. Hepatol. 2015, 7, 101–112. [Google Scholar] [CrossRef]
- Han, Y.; Chen, Z.; Yang, Y.; Jiang, Z.; Gu, Y.; Liu, Y.; Lin, C.; Pan, Z.; Yu, Y.; Jiang, M.; et al. Human CD14+ CTLA-4+ regulatory dendritic cells suppress T-cell response by cytotoxic T-lymphocyte antigen-4-dependent IL-10 and indoleamine-2,3-dioxygenase production in hepatocellular carcinoma. Hepatology 2014, 59, 567–579. [Google Scholar] [CrossRef]
- Shi, F.; Shi, M.; Zeng, Z.; Qi, R.Z.; Liu, Z.W.; Zhang, J.Y.; Yang, Y.P.; Tien, P.; Wang, F.S. PD-1 and PD-L1 upregulation promotes CD8(+) T-cell apoptosis and postoperative recurrence in hepatocellular carcinoma patients. Int. J. Cancer 2011, 128, 887–896. [Google Scholar] [CrossRef]
- El-Khoueiry, A.B.; Sangro, B.; Yau, T.; Crocenzi, T.S.; Kudo, M.; Hsu, C.; Kim, T.Y.; Choo, S.P.; Trojan, J.; Welling, T.H.R.; et al. Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): An open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet 2017, 389, 2492–2502. [Google Scholar] [CrossRef]
- Holland, P.W. Evolution of homeobox genes. Wiley Interdiscip. Rev. Dev. Biol. 2013, 2, 31–45. [Google Scholar] [CrossRef]
- Lewis, E.B. A gene complex controlling segmentation in Drosophila. Nature 1978, 276, 565–570. [Google Scholar] [CrossRef]
- Nam, J.; Nei, M. Evolutionary change of the numbers of homeobox genes in bilateral animals. Mol. Biol. Evol. 2005, 22, 2386–2394. [Google Scholar] [CrossRef]
- Holland, P.W.; Booth, H.A.; Bruford, E.A. Classification and nomenclature of all human homeobox genes. BMC Biol. 2007, 5, 47. [Google Scholar] [CrossRef] [PubMed]
- Noyes, M.B.; Christensen, R.G.; Wakabayashi, A.; Stormo, G.D.; Brodsky, M.H.; Wolfe, S.A. Analysis of homeodomain specificities allows the family-wide prediction of preferred recognition sites. Cell 2008, 133, 1277–1289. [Google Scholar] [CrossRef]
- Young, R.A. Control of the embryonic stem cell state. Cell 2011, 144, 940–954. [Google Scholar] [CrossRef] [PubMed]
- Quinonez, S.C.; Innis, J.W. Human HOX gene disorders. Mol. Genet. Metab. 2014, 111, 4–15. [Google Scholar] [CrossRef] [PubMed]
- Shah, N.; Sukumar, S. The Hox genes and their roles in oncogenesis. Nat. Rev. Cancer 2010, 10, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Sun, X.; He, X. Overexpression of Aristaless-Like Homeobox-4 Inhibits Proliferation, Invasion, and EMT in Hepatocellular Carcinoma Cells. Oncol. Res. 2017, 25, 11–18. [Google Scholar] [CrossRef]
- Wang, G.; Liu, J.; Cai, Y.; Chen, J.; Xie, W.; Kong, X.; Huang, W.; Guo, H.; Zhao, X.; Lu, Y.; et al. Loss of Barx1 promotes hepatocellular carcinoma metastasis through up-regulating MGAT5 and MMP9 expression and indicates poor prognosis. Oncotarget 2017, 8, 71867–71880. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, J.X.; Huang, L.L.; He, L.J.; Liao, Y.J.; Lai, Y.R.; Deng, H.X.; Tian, X.P.; Kung, H.F.; Xie, D.; et al. Low expression of BARX2 in human primary hepatocellular carcinoma correlates with metastasis and predicts poor prognosis. Hepatol. Res. 2015, 45, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Yang, Y.; Wang, M.C.; Yuan, S.X.; Tian, T.; Han, J.; Ni, J.S.; Wang, J.; Xing, H.; Zhou, W.P. Low CDX1 expression predicts a poor prognosis for hepatocellular carcinoma patients after hepatectomy. Surg. Oncol. 2016, 25, 171–177. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Jiang, W.; Wu, L.; Chang, R.; Wu, K.; Wang, Z. miR-301a is a candidate oncogene that targets the homeobox gene Gax in human hepatocellular carcinoma. Dig. Dis. Sci. 2012, 57, 1171–1180. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; You, P.; Zhao, J.P.; Zhang, S.L.; Song, S.H.; Fu, Z.R.; Ye, L.W.; Zi, X.Y.; Xie, D.F.; Zhu, M.H.; et al. A potential role for the homeoprotein Hhex in hepatocellular carcinoma progression. Med. Oncol. 2012, 29, 1059–1067. [Google Scholar] [CrossRef]
- Li, Q.; Ding, C.; Chen, C.; Zhang, Z.; Xiao, H.; Xie, F.; Lei, L.; Chen, Y.; Mao, B.; Jiang, M.; et al. miR-224 promotion of cell migration and invasion by targeting Homeobox D 10 gene in human hepatocellular carcinoma. J. Gastroenterol. Hepatol. 2014, 29, 835–842. [Google Scholar] [CrossRef]
- Qu, L.; Deng, B.; Zeng, Y.; Cao, Z. Decreased expression of the Nkx2.8 gene correlates with tumor progression and a poor prognosis in HCC cancer. Cancer Cell Int. 2014, 14, 28. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Liu, Z.; Ge, C.; Chen, C.; Zhao, F.; Li, H.; Chen, T.; Yao, M.; Li, J. NK3 homeobox 1 (NKX3.1) up-regulates forkhead box O1 expression in hepatocellular carcinoma and thereby suppresses tumor proliferation and invasion. J. Biol. Chem. 2017, 292, 19146–19159. [Google Scholar] [CrossRef]
- Hirata, H.; Sugimachi, K.; Takahashi, Y.; Ueda, M.; Sakimura, S.; Uchi, R.; Kurashige, J.; Takano, Y.; Nanbara, S.; Komatsu, H.; et al. Downregulation of PRRX1 Confers Cancer Stem Cell-Like Properties and Predicts Poor Prognosis in Hepatocellular Carcinoma. Ann. Surg. Oncol. 2015, 22, S1402–S1409. [Google Scholar] [CrossRef]
- Fan, M.; Shen, J.; Liu, H.; Wen, Z.; Yang, J.; Yang, P.; Liu, K.; Chang, Y.; Duan, J.; Lu, K. Downregulation of PRRX1 via the p53-dependent signaling pathway predicts poor prognosis in hepatocellular carcinoma. Oncol. Rep. 2017, 38, 1083–1090. [Google Scholar] [CrossRef]
- Zhu, R.; Wong, K.F.; Lee, N.P.; Lee, K.F.; Luk, J.M. HNF1alpha and CDX2 transcriptional factors bind to cadherin-17 (CDH17) gene promoter and modulate its expression in hepatocellular carcinoma. J. Cell. Biochem. 2010, 111, 618–626. [Google Scholar] [CrossRef]
- Liu, J.; Cui, X.; Qu, L.; Hua, L.; Wu, M.; Shen, Z.; Lu, C.; Ni, R. Overexpression of DLX2 is associated with poor prognosis and sorafenib resistance in hepatocellular carcinoma. Exp. Mol. Pathol. 2016, 101, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.H.; Xu, X.P.; Sun, C.Y.; Yu, Z.J. Regulation of the oncogenic function of distal-less 4 by microRNA-122 in hepatocellular carcinoma. Mol. Med. Rep. 2015, 12, 1375–1380. [Google Scholar] [CrossRef][Green Version]
- Xue, T.C.; Ge, N.L.; Zhang, L.; Cui, J.F.; Chen, R.X.; You, Y.; Ye, S.L.; Ren, Z.G. Goosecoid promotes the metastasis of hepatocellular carcinoma by modulating the epithelial-mesenchymal transition. PLoS ONE 2014, 9, e109695. [Google Scholar] [CrossRef] [PubMed]
- Erokhin, V.V.; Lepekha, L.N.; Erokhina, M.V.; Bocharova, I.V.; Kurynina, A.V.; Onishchenko, G.E. Selective effects of pulmonary surfactant on various subpopulations of alveolar macrophages in the model of experimental tuberculosis. Vestn. Ross. Akad. Med. Nauk. 2012, 22–28. [Google Scholar]
- Pan, T.T.; Jia, W.D.; Yao, Q.Y.; Sun, Q.K.; Ren, W.H.; Huang, M.; Ma, J.; Li, J.S.; Ma, J.L.; Yu, J.H.; et al. Overexpression of HOXA13 as a potential marker for diagnosis and poor prognosis of hepatocellular carcinoma. Tohoku J. Exp. Med. 2014, 234, 209–219. [Google Scholar] [CrossRef]
- Quagliata, L.; Quintavalle, C.; Lanzafame, M.; Matter, M.S.; Novello, C.; di Tommaso, L.; Pressiani, T.; Rimassa, L.; Tornillo, L.; Roncalli, M.; et al. High expression of HOXA13 correlates with poorly differentiated hepatocellular carcinomas and modulates sorafenib response in in vitro models. Lab. Investig. 2018, 98, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, H.; Iguchi, T.; Masuda, T.; Ueda, M.; Kidogami, S.; Ogawa, Y.; Nambara, S.; Sato, K.; Hu, Q.; Saito, T.; et al. HOXB7 Expression is a Novel Biomarker for Long-term Prognosis After Resection of Hepatocellular Carcinoma. Anticancer Res. 2016, 36, 2767–2773. [Google Scholar]
- Huan, H.B.; Yang, D.P.; Wen, X.D.; Chen, X.J.; Zhang, L.; Wu, L.L.; Bie, P.; Xia, F. HOXB7 accelerates the malignant progression of hepatocellular carcinoma by promoting stemness and epithelial-mesenchymal transition. J. Exp. Clin. Cancer Res. 2017, 36, 86. [Google Scholar] [CrossRef]
- Wang, W.M.; Xu, Y.; Wang, Y.H.; Sun, H.X.; Sun, Y.F.; He, Y.F.; Zhu, Q.F.; Hu, B.; Zhang, X.; Xia, J.L.; et al. HOXB7 promotes tumor progression via bFGF-induced activation of MAPK/ERK pathway and indicated poor prognosis in hepatocellular carcinoma. Oncotarget 2017, 8, 47121–47135. [Google Scholar] [CrossRef]
- Sha, L.; Dong, L.; Lv, L.; Bai, L.; Ji, X. HOXB9 promotes epithelial-to-mesenchymal transition via transforming growth factor-beta1 pathway in hepatocellular carcinoma cells. Clin. Exp. Med. 2015, 15, 55–64. [Google Scholar] [CrossRef]
- Lv, X.; Li, L.; Lv, L.; Qu, X.; Jin, S.; Li, K.; Deng, X.; Cheng, L.; He, H.; Dong, L. HOXD9 promotes epithelial-mesenchymal transition and cancer metastasis by ZEB1 regulation in hepatocellular carcinoma. J. Exp. Clin. Cancer Res. 2015, 34, 133. [Google Scholar] [CrossRef]
- Wang, P.; Zhuang, C.; Huang, D.; Xu, K. Downregulation of miR-377 contributes to IRX3 deregulation in hepatocellular carcinoma. Oncol. Rep. 2016, 36, 247–252. [Google Scholar] [CrossRef]
- Wang, S.N.; Wang, L.T.; Sun, D.P.; Chai, C.Y.; Hsi, E.; Kuo, H.T.; Yokoyama, K.K.; Hsu, S.H. Intestine-specific homeobox (ISX) upregulates E2F1 expression and related oncogenic activities in HCC. Oncotarget 2016, 7, 36924–36939. [Google Scholar] [CrossRef]
- Wang, L.T.; Chiou, S.S.; Chai, C.Y.; Hsi, E.; Yokoyama, K.K.; Wang, S.N.; Huang, S.K.; Hsu, S.H. Intestine-Specific Homeobox Gene ISX Integrates IL6 Signaling, Tryptophan Catabolism, and Immune Suppression. Cancer Res. 2017, 77, 4065–4077. [Google Scholar] [CrossRef]
- Hsu, S.H.; Wang, L.T.; Lee, K.T.; Chen, Y.L.; Liu, K.Y.; Suen, J.L.; Chai, C.Y.; Wang, S.N. Proinflammatory homeobox gene, ISX, regulates tumor growth and survival in hepatocellular carcinoma. Cancer Res. 2013, 73, 508–518. [Google Scholar] [CrossRef]
- Dang, H.; Ding, W.; Emerson, D.; Rountree, C.B. Snail1 induces epithelial-to-mesenchymal transition and tumor initiating stem cell characteristics. BMC Cancer 2011, 11, 396. [Google Scholar] [CrossRef]
- Jiang, L.; Shan, J.; Shen, J.; Wang, Y.; Yan, P.; Liu, L.; Zhao, W.; Xu, Y.; Zhu, W.; Su, L.; et al. Androgen/androgen receptor axis maintains and promotes cancer cell stemness through direct activation of Nanog transcription in hepatocellular carcinoma. Oncotarget 2016, 7, 36814–36828. [Google Scholar] [CrossRef]
- Xu, G.; Ye, J.; Liu, X.J.; Zhang, N.P.; Zhao, Y.M.; Fan, J.; Liu, X.P.; Wu, J. Activation of pluripotent genes in hepatic progenitor cells in the transition of nonalcoholic steatohepatitis to pre-malignant lesions. Lab. Investig. 2017, 97, 1201–1217. [Google Scholar] [CrossRef]
- Yin, X.; Zhang, B.H.; Zheng, S.S.; Gao, D.M.; Qiu, S.J.; Wu, W.Z.; Ren, Z.G. Coexpression of gene Oct4 and Nanog initiates stem cell characteristics in hepatocellular carcinoma and promotes epithelial-mesenchymal transition through activation of Stat3/Snail signaling. J. Hematol. Oncol. 2015, 8, 23. [Google Scholar] [CrossRef]
- Shan, J.; Shen, J.; Liu, L.; Xia, F.; Xu, C.; Duan, G.; Xu, Y.; Ma, Q.; Yang, Z.; Zhang, Q.; et al. Nanog regulates self-renewal of cancer stem cells through the insulin-like growth factor pathway in human hepatocellular carcinoma. Hepatology 2012, 56, 1004–1014. [Google Scholar] [CrossRef]
- Sun, C.; Sun, L.; Jiang, K.; Gao, D.M.; Kang, X.N.; Wang, C.; Zhang, S.; Huang, S.; Qin, X.; Li, Y.; et al. NANOG promotes liver cancer cell invasion by inducing epithelial-mesenchymal transition through NODAL/SMAD3 signaling pathway. Int. J. Biochem. Cell Biol. 2013, 45, 1099–1108. [Google Scholar] [CrossRef]
- Zhao, X.; Sun, B.; Sun, D.; Liu, T.; Che, N.; Gu, Q.; Dong, X.; Li, R.; Liu, Y.; Li, J. Slug promotes hepatocellular cancer cell progression by increasing sox2 and nanog expression. Oncol. Rep. 2015, 33, 149–156. [Google Scholar] [CrossRef]
- Uthaya Kumar, D.B.; Chen, C.L.; Liu, J.C.; Feldman, D.E.; Sher, L.S.; French, S.; DiNorcia, J.; French, S.W.; Naini, B.V.; Junrungsee, S.; et al. TLR4 Signaling via NANOG Cooperates with STAT3 to Activate Twist1 and Promote Formation of Tumor-Initiating Stem-Like Cells in Livers of Mice. Gastroenterology 2016, 150, 707–719. [Google Scholar] [CrossRef]
- Chen, C.L.; Uthaya Kumar, D.B.; Punj, V.; Xu, J.; Sher, L.; Tahara, S.M.; Hess, S.; Machida, K. NANOG Metabolically Reprograms Tumor-Initiating Stem-like Cells through Tumorigenic Changes in Oxidative Phosphorylation and Fatty Acid Metabolism. Cell Metab. 2016, 23, 206–219. [Google Scholar] [CrossRef]
- Cai, W.; Wang, Z.; Wei, C.; Wu, M.; Zheng, W.; Zhang, H.; Liu, C.; Liu, L. Prognostic evaluation of NANOG and OCT4 expression for posttransplantation hepatocellular carcinoma recurrence. J. Cell. Biochem. 2018. [Google Scholar] [CrossRef]
- Li, L.; Tang, J.; Zhang, B.; Yang, W.; LiuGao, M.; Wang, R.; Tan, Y.; Fan, J.; Chang, Y.; Fu, J.; et al. Epigenetic modification of MiR-429 promotes liver tumour-initiating cell properties by targeting Rb binding protein 4. Gut 2015, 64, 156–167. [Google Scholar] [CrossRef]
- Chang, T.S.; Wu, Y.C.; Chi, C.C.; Su, W.C.; Chang, P.J.; Lee, K.F.; Tung, T.H.; Wang, J.; Liu, J.J.; Tung, S.Y.; et al. Activation of IL6/IGFIR confers poor prognosis of HBV-related hepatocellular carcinoma through induction of OCT4/NANOG expression. Clin. Cancer Res. 2015, 21, 201–210. [Google Scholar] [CrossRef]
- Han, S.Y.; Han, H.B.; Tian, X.Y.; Sun, H.; Xue, D.; Zhao, C.; Jiang, S.T.; He, X.R.; Zheng, W.X.; Wang, J.; et al. MicroRNA-33a-3p suppresses cell migration and invasion by directly targeting PBX3 in human hepatocellular carcinoma. Oncotarget 2016, 7, 42461–42473. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, J.B.; Qin, Y.; Wang, W.; Wei, L.; Teng, Y.; Guo, L.; Zhang, B.; Lin, Z.; Liu, J.; et al. PROX1 promotes hepatocellular carcinoma metastasis by way of up-regulating hypoxia-inducible factor 1alpha expression and protein stability. Hepatology 2013, 58, 692–705. [Google Scholar] [CrossRef]
- Dudas, J.; Mansuroglu, T.; Moriconi, F.; Haller, F.; Wilting, J.; Lorf, T.; Fuzesi, L.; Ramadori, G. Altered regulation of Prox1-gene-expression in liver tumors. BMC Cancer 2008, 8, 92. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ye, X.; Zhang, J.B.; Ouyang, H.; Shen, Z.; Wu, Y.; Wang, W.; Wu, J.; Tao, S.; Yang, X.; et al. PROX1 promotes hepatocellular carcinoma proliferation and sorafenib resistance by enhancing beta-catenin expression and nuclear translocation. Oncogene 2015, 34, 5524–5535. [Google Scholar] [CrossRef]
- Yang, T.; Zhang, H.; Cai, S.Y.; Shen, Y.N.; Yuan, S.X.; Yang, G.S.; Wu, M.C.; Lu, J.H.; Shen, F. Elevated SHOX2 expression is associated with tumor recurrence of hepatocellular carcinoma. Ann. Surg. Oncol. 2013, 20, S644–S649. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.; Zhou, X.; Liu, S.; Jin, T.; Piao, Y.; Liu, C.; Lin, Z. Overexpression of sineoculis homeobox homolog 1 predicts poor prognosis of hepatocellular carcinoma. Int. J. Clin. Exp. Pathol. 2014, 7, 3018–3027. [Google Scholar]
- Huang, M.T.; Wei, P.L.; Liu, J.J.; Liu, D.Z.; Huey-Chun, H.; An, J.; Wu, C.C.; Wu, C.H.; Ho, Y.S.; Yang, Y.Y.; et al. Knockdown of thrombomodulin enhances HCC cell migration through increase of ZEB1 and decrease of E-cadherin gene expression. Ann. Surg. Oncol. 2010, 17, 3379–3385. [Google Scholar] [CrossRef]
- Liu, T.A.; Jan, Y.J.; Ko, B.S.; Liang, S.M.; Chen, S.C.; Wang, J.; Hsu, C.; Wu, Y.M.; Liou, J.Y. 14-3-3epsilon overexpression contributes to epithelial-mesenchymal transition of hepatocellular carcinoma. PLoS ONE 2013, 8, e57968. [Google Scholar] [CrossRef]
- Hashiguchi, M.; Ueno, S.; Sakoda, M.; Iino, S.; Hiwatashi, K.; Minami, K.; Ando, K.; Mataki, Y.; Maemura, K.; Shinchi, H.; et al. Clinical implication of ZEB-1 and E-cadherin expression in hepatocellular carcinoma (HCC). BMC Cancer 2013, 13, 572. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.H.; Yang, F.; Wang, F.; Ma, J.Z.; Guo, Y.J.; Tao, Q.F.; Liu, F.; Pan, W.; Wang, T.T.; Zhou, C.C.; et al. A long noncoding RNA activated by TGF-beta promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell 2014, 25, 666–681. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.M.; Lee, W.H.; Lee, C.G.; An, J.; Kim, E.S.; Kim, S.H.; Lee, S.K.; Lee, C.H.; Dhanasekaran, D.N.; Moon, A.; et al. Galpha12 gep oncogene deregulation of p53-responsive microRNAs promotes epithelial-mesenchymal transition of hepatocellular carcinoma. Oncogene 2015, 34, 2910–2921. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Zhang, Y.; Zheng, X.; Tu, X.; Li, H.; Chen, J.; Zang, Y.; Zhang, J. Loss of MicroRNA-101 Promotes Epithelial to Mesenchymal Transition in Hepatocytes. J. Cell. Physiol. 2015, 230, 2706–2717. [Google Scholar] [CrossRef]
- Li, T.; Xie, J.; Shen, C.; Cheng, D.; Shi, Y.; Wu, Z.; Deng, X.; Chen, H.; Shen, B.; Peng, C.; et al. Amplification of Long Noncoding RNA ZFAS1 Promotes Metastasis in Hepatocellular Carcinoma. Cancer Res. 2015, 75, 3181–3191. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Xie, J.; Shen, C.; Cheng, D.; Shi, Y.; Wu, Z.; Deng, X.; Chen, H.; Shen, B.; Peng, C.; et al. Upregulation of long noncoding RNA ZEB1-AS1 promotes tumor metastasis and predicts poor prognosis in hepatocellular carcinoma. Oncogene 2016, 35, 1575–1584. [Google Scholar] [CrossRef] [PubMed]
- Chang, L.; Yuan, Y.; Li, C.; Guo, T.; Qi, H.; Xiao, Y.; Dong, X.; Liu, Z.; Liu, Q. Upregulation of SNHG6 regulates ZEB1 expression by competitively binding miR-101-3p and interacting with UPF1 in hepatocellular carcinoma. Cancer Lett. 2016, 383, 183–194. [Google Scholar] [CrossRef]
- Luo, W.; Zhu, X.; Liu, W.; Ren, Y.; Bei, C.; Qin, L.; Miao, X.; Tang, F.; Tang, G.; Tan, S. MYC associated zinc finger protein promotes the invasion and metastasis of hepatocellular carcinoma by inducing epithelial mesenchymal transition. Oncotarget 2016, 7, 86420–86432. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Yao, H.; Wang, K.; Liu, X. Long Non-Coding RNA MALAT1 Regulates ZEB1 Expression by Sponging miR-143-3p and Promotes Hepatocellular Carcinoma Progression. J. Cell. Biochem. 2017, 118, 4836–4843. [Google Scholar] [CrossRef]
- Fu, X.; Zhu, Y.; Zheng, B.; Zou, Y.; Wang, C.; Wu, P.; Wang, J.; Chen, H.; Du, P.; Liang, B.; et al. KIFC1, a novel potential prognostic factor and therapeutic target in hepatocellular carcinoma. Int. J. Oncol. 2018, 52, 1912–1922. [Google Scholar] [CrossRef]
- He, C.; Liu, Z.; Jin, L.; Zhang, F.; Peng, X.; Xiao, Y.; Wang, X.; Lyu, Q.; Cai, X. lncRNA TUG1-Mediated Mir-142-3p Downregulation Contributes to Metastasis and the Epithelial-to-Mesenchymal Transition of Hepatocellular Carcinoma by Targeting ZEB1. Cell. Physiol. Biochem. 2018, 48, 1928–1941. [Google Scholar] [CrossRef] [PubMed]
- Qiu, B.; Wei, W.; Zhu, J.; Fu, G.; Lu, D. EMT induced by loss of LKB1 promotes migration and invasion of liver cancer cells through ZEB1-induced YAP signaling. Oncol. Lett. 2018, 16, 6465–6471. [Google Scholar] [CrossRef]
- Kim, T.; Veronese, A.; Pichiorri, F.; Lee, T.J.; Jeon, Y.J.; Volinia, S.; Pineau, P.; Marchio, A.; Palatini, J.; Suh, S.S.; et al. p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J. Exp. Med. 2011, 208, 875–883. [Google Scholar] [CrossRef] [PubMed]
- Cai, M.Y.; Luo, R.Z.; Chen, J.W.; Pei, X.Q.; Lu, J.B.; Hou, J.H.; Yun, J.P. Overexpression of ZEB2 in peritumoral liver tissue correlates with favorable survival after curative resection of hepatocellular carcinoma. PLoS ONE 2012, 7, e32838. [Google Scholar] [CrossRef]
- Xia, L.; Huang, W.; Tian, D.; Zhang, L.; Qi, X.; Chen, Z.; Shang, X.; Nie, Y.; Wu, K. Forkhead box Q1 promotes hepatocellular carcinoma metastasis by transactivating ZEB2 and VersicanV1 expression. Hepatology 2014, 59, 958–973. [Google Scholar] [CrossRef] [PubMed]
- Pang, X.; Huang, K.; Zhang, Q.; Zhang, Y.; Niu, J. miR-154 targeting ZEB2 in hepatocellular carcinoma functions as a potential tumor suppressor. Oncol. Rep. 2015, 34, 3272–3279. [Google Scholar] [CrossRef] [PubMed]
- Qiu, G.; Lin, Y.; Zhang, H.; Wu, D. miR-139-5p inhibits epithelial-mesenchymal transition, migration and invasion of hepatocellular carcinoma cells by targeting ZEB1 and ZEB2. Biochem. Biophys. Res. Commun. 2015, 463, 315–321. [Google Scholar] [CrossRef]
- Lan, T.; Chang, L.; Wu, L.; Yuan, Y. Downregulation of ZEB2-AS1 decreased tumor growth and metastasis in hepatocellular carcinoma. Mol. Med. Rep. 2016, 14, 4606–4612. [Google Scholar] [CrossRef]
- Gao, P.T.; Cheng, J.W.; Gong, Z.J.; Hu, B.; Sun, Y.F.; Cao, Y.; Qiu, S.J.; Zhou, J.; Fan, J.; Yang, X.R. Low SLC29A1 expression is associated with poor prognosis in patients with hepatocellular carcinoma. Am. J. Cancer Res. 2017, 7, 2465–2477. [Google Scholar] [PubMed]
- Jiang, G.; Wen, L.; Deng, W.; Jian, Z.; Zheng, H. Regulatory role of miR-211-5p in hepatocellular carcinoma metastasis by targeting ZEB2. Biomed. Pharmacother. 2017, 90, 806–812. [Google Scholar] [CrossRef]
- O’Brien, C.A.; Pollett, A.; Gallinger, S.; Dick, J.E. A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 2007, 445, 106–110. [Google Scholar] [CrossRef]
- Ricci-Vitiani, L.; Lombardi, D.G.; Pilozzi, E.; Biffoni, M.; Todaro, M.; Peschle, C.; De Maria, R. Identification and expansion of human colon-cancer-initiating cells. Nature 2007, 445, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Rycaj, K.; Tang, D.G. Cell-of-Origin of Cancer versus Cancer Stem Cells: Assays and Interpretations. Cancer Res. 2015, 75, 4003–4011. [Google Scholar] [CrossRef]
- Visvader, J.E.; Lindeman, G.J. Cancer stem cells in solid tumours: Accumulating evidence and unresolved questions. Nat. Rev. Cancer 2008, 8, 755–768. [Google Scholar] [CrossRef]
- Chen, C.L.; Tsukamoto, H.; Liu, J.C.; Kashiwabara, C.; Feldman, D.; Sher, L.; Dooley, S.; French, S.W.; Mishra, L.; Petrovic, L.; et al. Reciprocal regulation by TLR4 and TGF-beta in tumor-initiating stem-like cells. J. Clin. Investig. 2013, 123, 2832–2849. [Google Scholar] [CrossRef]
- Ma, S.; Chan, K.W.; Hu, L.; Lee, T.K.; Wo, J.Y.; Ng, I.O.; Zheng, B.J.; Guan, X.Y. Identification and characterization of tumorigenic liver cancer stem/progenitor cells. Gastroenterology 2007, 132, 2542–2556. [Google Scholar] [CrossRef]
- Pan, G.J.; Chang, Z.Y.; Scholer, H.R.; Pei, D. Stem cell pluripotency and transcription factor Oct4. Cell Res. 2002, 12, 321–329. [Google Scholar] [CrossRef]
- Torres-Padilla, M.E.; Chambers, I. Transcription factor heterogeneity in pluripotent stem cells: A stochastic advantage. Development 2014, 141, 2173–2181. [Google Scholar] [CrossRef]
- Wang, Y.J.; Herlyn, M. The emerging roles of Oct4 in tumor-initiating cells. Am. J. Physiol.-Cell Physiol. 2015, 309, C709–C718. [Google Scholar] [CrossRef]
- Gawlik-Rzemieniewska, N.; Bednarek, I. The role of NANOG transcriptional factor in the development of malignant phenotype of cancer cells. Cancer Biol. Ther. 2016, 17, 1–10. [Google Scholar] [CrossRef]
- Cao, J.; Zhao, M.; Liu, J.; Zhang, X.; Pei, Y.; Wang, J.; Yang, X.; Shen, B.; Zhang, J. RACK1 Promotes Self-Renewal and Chemoresistance of Cancer Stem Cells in Human Hepatocellular Carcinoma through Stabilizing Nanog. Theranostics 2019, 9, 811–828. [Google Scholar] [CrossRef]
- Zhang, H.; Li, G.; Chen, G.; Zhang, Y.; Pan, J.; Tang, H.; Li, J.; Guo, W.; Zhang, S. Targeting Mcl-1 inhibits survival and self-renewal of hepatocellular cancer stem-like cells. Clin. Res. Hepatol. Gastroenterol. 2018. [Google Scholar] [CrossRef]
- Zhou, D.S.; Wang, H.B.; Zhou, Z.G.; Zhang, Y.J.; Zhong, Q.; Xu, L.; Huang, Y.H.; Yeung, S.C.; Chen, M.S.; Zeng, M.S. TACC3 promotes stemness and is a potential therapeutic target in hepatocellular carcinoma. Oncotarget 2015, 6, 24163–24177. [Google Scholar] [CrossRef][Green Version]
- Feng, H.; Zhu, M.; Zhang, R.; Wang, Q.; Li, W.; Dong, X.; Chen, Y.; Lu, Y.; Liu, K.; Lin, B.; et al. GATA5 inhibits hepatocellular carcinoma cells malignant behaviours by blocking expression of reprogramming genes. J. Cell. Mol. Med. 2019, 23, 2536–2548. [Google Scholar] [CrossRef]
- Song, Y.; Pan, G.; Chen, L.; Ma, S.; Zeng, T.; Man Chan, T.H.; Li, L.; Lian, Q.; Chow, R.; Cai, X.; et al. Loss of ATOH8 Increases Stem Cell Features of Hepatocellular Carcinoma Cells. Gastroenterology 2015, 149, 1068–1081.e5. [Google Scholar] [CrossRef]
- Acloque, H.; Adams, M.S.; Fishwick, K.; Bronner-Fraser, M.; Nieto, M.A. Epithelial-mesenchymal transitions: The importance of changing cell state in development and disease. J. Clin. Invest. 2009, 119, 1438–1449. [Google Scholar] [CrossRef]
- Van Zijl, F.; Zulehner, G.; Petz, M.; Schneller, D.; Kornauth, C.; Hau, M.; Machat, G.; Grubinger, M.; Huber, H.; Mikulits, W. Epithelial-mesenchymal transition in hepatocellular carcinoma. Future Oncol. 2009, 5, 1169–1179. [Google Scholar] [CrossRef]
- Giannelli, G.; Koudelkova, P.; Dituri, F.; Mikulits, W. Role of epithelial to mesenchymal transition in hepatocellular carcinoma. J. Hepatol. 2016, 65, 798–808. [Google Scholar] [CrossRef] [PubMed]
- Peinado, H.; Olmeda, D.; Cano, A. Snail, Zeb and bHLH factors in tumour progression: An alliance against the epithelial phenotype? Nat. Rev. Cancer 2007, 7, 415–428. [Google Scholar] [CrossRef]
- Eger, A.; Aigner, K.; Sonderegger, S.; Dampier, B.; Oehler, S.; Schreiber, M.; Berx, G.; Cano, A.; Beug, H.; Foisner, R. DeltaEF1 is a transcriptional repressor of E-cadherin and regulates epithelial plasticity in breast cancer cells. Oncogene 2005, 24, 2375–2385. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Sawada, J.; Sui, G.; Affar el, B.; Whetstine, J.R.; Lan, F.; Ogawa, H.; Luke, M.P.; Nakatani, Y.; Shi, Y. Coordinated histone modifications mediated by a CtBP co-repressor complex. Nature 2003, 422, 735–738. [Google Scholar] [CrossRef]
- Sanchez-Tillo, E.; Lazaro, A.; Torrent, R.; Cuatrecasas, M.; Vaquero, E.C.; Castells, A.; Engel, P.; Postigo, A. ZEB1 represses E-cadherin and induces an EMT by recruiting the SWI/SNF chromatin-remodeling protein BRG1. Oncogene 2010, 29, 3490–3500. [Google Scholar] [CrossRef]
- Abba, M.L.; Patil, N.; Leupold, J.H.; Allgayer, H. MicroRNA Regulation of Epithelial to Mesenchymal Transition. J. Clin. Med. 2016, 5, 8. [Google Scholar] [CrossRef]
- Xu, Q.; Deng, F.; Qin, Y.; Zhao, Z.; Wu, Z.; Xing, Z.; Ji, A.; Wang, Q.J. Long non-coding RNA regulation of epithelial-mesenchymal transition in cancer metastasis. Cell Death Dis. 2016, 7, e2254. [Google Scholar] [CrossRef]
- Wei, W.; Pelechano, V.; Jarvelin, A.I.; Steinmetz, L.M. Functional consequences of bidirectional promoters. Trends Genet. 2011, 27, 267–276. [Google Scholar] [CrossRef]
- Jenne, C.N.; Kubes, P. Immune surveillance by the liver. Nat. Immunol. 2013, 14, 996–1006. [Google Scholar] [CrossRef]
- Notas, G.; Kisseleva, T.; Brenner, D. NK and NKT cells in liver injury and fibrosis. Clin. Immunol. 2009, 130, 16–26. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.W.; Harmon, C.; O’Farrelly, C. Liver immunology and its role in inflammation and homeostasis. Cell. Mol. Immunol. 2016, 13, 267–276. [Google Scholar] [CrossRef]
- Protzer, U.; Maini, M.K.; Knolle, P.A. Living in the liver: Hepatic infections. Nat. Rev. Immunol. 2012, 12, 201–213. [Google Scholar] [CrossRef] [PubMed]
- Berasain, C.; Castillo, J.; Perugorria, M.J.; Latasa, M.U.; Prieto, J.; Avila, M.A. Inflammation and liver cancer: New molecular links. Ann. N. Y. Acad. Sci. 2009, 1155, 206–221. [Google Scholar] [CrossRef]
- Nakamoto, Y.; Guidotti, L.G.; Kuhlen, C.V.; Fowler, P.; Chisari, F.V. Immune pathogenesis of hepatocellular carcinoma. J. Exp. Med. 1998, 188, 341–350. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Chesnokov, V.; Yokoyama, K.K.; Carr, B.I.; Itakura, K. Expression of the Hoxa-13 gene correlates to hepatitis B and C virus associated HCC. Biochem. Biophys. Res. Commun. 2001, 281, 1041–1044. [Google Scholar] [CrossRef] [PubMed]
- Shimoda, M.; Takahashi, M.; Yoshimoto, T.; Kono, T.; Ikai, I.; Kubo, H. A homeobox protein, prox1, is involved in the differentiation, proliferation, and prognosis in hepatocellular carcinoma. Clin. Cancer Res. 2006, 12, 6005–6011. [Google Scholar] [CrossRef] [PubMed]
- Goto, T.; Elbahrawy, A.; Furuyama, K.; Horiguchi, M.; Hosokawa, S.; Aoyama, Y.; Tsuboi, K.; Sakikubo, M.; Hirata, K.; Masui, T.; et al. Liver-specific Prox1 inactivation causes hepatic injury and glucose intolerance in mice. FEBS Lett. 2017, 591, 624–635. [Google Scholar] [CrossRef]
- Chang, T.M.; Hung, W.C. Transcriptional repression of TWIST1 gene by Prospero-related homeobox 1 inhibits invasiveness of hepatocellular carcinoma cells. FEBS Lett. 2012, 586, 3746–3752. [Google Scholar] [CrossRef]
- Chang, T.M.; Hung, W.C. The homeobox transcription factor Prox1 inhibits proliferation of hepatocellular carcinoma cells by inducing p53-dependent senescence-like phenotype. Cancer Biol. Ther. 2013, 14, 222–229. [Google Scholar] [CrossRef]
- Qin, J.; Zhai, J.; Hong, R.; Shan, S.; Kong, Y.; Wen, Y.; Wang, Y.; Liu, J.; Xie, Y. Prospero-related homeobox protein (Prox1) inhibits hepatitis B virus replication through repressing multiple cis regulatory elements. J. Gen. Virol. 2009, 90, 1246–1255. [Google Scholar] [CrossRef] [PubMed]
- Ching, R.H.H.; Sze, K.M.F.; Lau, E.Y.T.; Chiu, Y.T.; Lee, J.M.F.; Ng, I.O.L.; Lee, T.K.W. C-terminal truncated hepatitis B virus X protein regulates tumorigenicity, self-renewal and drug resistance via STAT3/Nanog signaling pathway. Oncotarget 2017, 8, 23507–23516. [Google Scholar] [CrossRef] [PubMed]
- Machida, K.; Tsukamoto, H.; Mkrtchyan, H.; Duan, L.; Dynnyk, A.; Liu, H.M.; Asahina, K.; Govindarajan, S.; Ray, R.; Ou, J.H.; et al. Toll-like receptor 4 mediates synergism between alcohol and HCV in hepatic oncogenesis involving stem cell marker Nanog. Proc. Natl. Acad. Sci. USA 2009, 106, 1548–1553. [Google Scholar] [CrossRef]
| Homeobox Gene | Experiment Model | Function in HCC | Ref. |
|---|---|---|---|
| aristaless-like homeobox 4 (ALX4) | cell lines: Huh7, HepG2, and HCCLM3 | Overexpression of ALX4 inhibits the proliferation, invasion, and EMT. | [31] |
| BARX homeobox 1 (BARX1) | cell lines: HepG2, Huh7, Hep3B, SMMC7721, MHCC97L, MHCC97H, HCCLM3, and HCCLM6 mouse model HCC tissues | 1. Low expression of BARX1 correlates with poor prognosis. | [32] |
| 2. BARX1 suppresses invasion and metastasis by inhibiting MGAT5 and MMP9 transcription. | |||
| BARX homeobox 2 (BARX2) | HCC tissues | Low expression of BARX2 is correlated with tumor metastasis. | [33] |
| caudal-type homeobox 1 (CDX1) | HCC tissues | Low expression of CDX1 is associated with poor prognosis. | [34] |
| growth arrest-specific homeobox (Gax) | cell lines: HepG2, Huh7, and HCCLM3 HCC tissues | Gax expression inhibits NF-kappa B signal, and its expression negatively regulated by miR-301a. | [35] |
| hematopoietically expressed homeobox protein (Hhex) | cell line: Hepa1-6 mouse model HCC tissues | 1. Overexpression of Hhex resulted in decreases expression of c-Jun and Bcl2, and increases expression of P53 and Rb. | [36] |
| 2. Hhex expression attenuates tumorigenicity in nude mice. | |||
| homeobox D10 (HOXD10) | cell lines: MHCC97H, MHCC97L, and HepG2 | HOXD10 is downregulated by miR-224 repression that causes cell migration and invasion. | [37] |
| NK2 homeobox 8 (NKX2.8) | cell lines: PLC and Hep3B HCC tissues | NKX2.8 expression is downregulated in HCC, and low NKX2.8 expression is negatively correlated with poor survival in patients. | [38] |
| NK3 homeobox 1 (NKX3.1) | cell lines: SMMC-7721, Li7, Huh7, HCCLM3 MHCC-97L, HCCLY10, PLC5, and SK-Hep1 mouse model HCC tissues | NKX3.1 suppresses tumor proliferation and invasion by up-regulating Foxo1 expression. | [39] |
| paired related homeobox 1 (PRRX1) | cell lines: Huh7, Hep3B, HepG2, SMMC7721, and PLC5 HCC tissues | 1. Hepatic cancer-stem cell properties are disrupted by PRRX1 overexpression. | [40,41] |
| 2. PRRX1 overexpression induces HCC apoptosis via the p53-signaling. |
| Homeobox Gene | Experiment Model | Function in HCC | Ref. |
|---|---|---|---|
| caudal-related homeobox 2 (CDX2) | cell lines: MHCC97L and Hep3B | CDX2 binds to CDH17 promoter and modulates its expression. | [42] |
| distal-less homeobox 2 (DLX2) | cell lines: Huh7, HepG2, Hep3B, SMMC7721, MHCC97H, and MHCC97L HCC tissues | 1. Overexpression of DLX2 in HCC tissues is an indicator of poor prognosis. | [43] |
| 2. DLX2 increases sorafenib resistance by promoting the ERK pathway and EMT. | |||
| distal-less homeobox 4 (DLX4) | cell line: Hep3B HCC tissues | 1. DLX4 is up-regulated in HCC tissues. | [44] |
| 2. miR-122 binds 3′UTR of DLX4 for down-regulated its expression. | |||
| goosecoid (GSC) | cell lines: MHCC97L, MHCC97H, HCCLM3, SMMC7721, Hep3B, and HepG2 HCC tissues | GSC expression is associated with metastasis and EMT in patients. | [45] |
| homeobox HB9 (HLXB9) | cell lines: HLE, HLF, Huh7, HepG2, and Hep3B HCC tissues | HLXB9 upregulation is observed in poorly differentiated HCC with a pseudoglandular pattern. | [46] |
| homeobox A13 (HOXA13) | cell lines: SNU-449 and HepG2 HCC tissues | 1. High HOXA13 expression is positively correlated with tumor size, microvascular invasion, pathological grade, tumor capsula status, AFP level, metastasis, and microvessel density. | [47,48] |
| 2. Overexpression of HOXA13 increases colony formation on soft agar and migration, and reduces sensitivity to sorafenib. | |||
| homeobox B7 (HOXB7) | cell lines: SMMC-7721, MHCC97L, MHCC97H, HCCLM3, PLC, HepG2, and Huh7 mouse model HCC tissues | 1. High HOXB7 expression is associated with larger tumor size and a higher rate of biliary invasion. | [49,50,51] |
| 2. HOXB7 promotes c-Myc and Slug expression through AKT activation, resulting in HCC progression. | |||
| 3. HOXB7 promotes proliferation, migration, and invasion through activation of the MAPK/ERK axis. | |||
| homeobox B9 (HOXB9) | cell lines: BEL-7402, BEL-7404, BEL-7405, HepG2, Hep3B, and SNU475 HCC tissues | HOXB9 regulates TGF-β1 and ZEB1 signaling to promote EMT and cancer metastasis. | [52,53] |
| iroquois homeobox3 (IRX3) | cell lines: HepG2 and SMMC7721 | IRX3 induces proliferation, migration, and invasion, but its expression is repressed by miR-377. | [54] |
| intestine-specific homeobox (ISX) | cell lines: HepG2 and Huh7 mouse model HCC tissues | 1. ISX is a regulator in HCC progression as a prognostic and therapeutic target in HCC. | [55,56,57] |
| 2. Cyclin D1 and E2F1 are downstream target genes of ISX in HCC. | |||
| 3. ISX involves kynurenine–AHR axis and immunosuppression effect of PD-L1 and CTLA-4 for immune escape by HCC. | |||
| NANOG | cell lines: Huh7, MHCC97L, HepG2, and SMMC7221 mouse model HCC tissues | 1. NANOG expression is required for TICs of HCC. | [58,59,60,61,62,63,64,65,66,67] |
| 2. Nanog maintains TICs through the insulin-like growth factor pathway in HCC. | |||
| 3. Nanog promotes EMT through Stat3-dependent Snail activation. | |||
| 4. HCV-NS5A induces TLR4–NANOG axis, promoting the formation of liver TICs. | |||
| POU class 5 homeobox 1 (POU5F1 or OCT4) | cell lines mouse model HCC tissues | OCT4 expression is required for TICs. | [67,68,69] |
| pre-B-Cell leukemia homeobox 3 (PBX3) | cell lines: HepG2 Huh7, QGY-7701, and BEL-7402 Chick model HCC tissues | miR-33a-3p suppresses the cell growth, spreading, and invasion by inhibiting PBX3 expression. | [70] |
| prospero-related homeobox 1 (PROX1) | cell lines: Hep3B, Huh7, HepG2, BEL-7402, QGY7701, QGY7703, SMCC7721, and MHCC97H mouse model HCC tissues | 1. PROX1 is required for hepatocyte migration. | [71,72,73] |
| 2. High PROX1 expression is associated with poor survival and tumor recurrence of HCC. | |||
| 3. PROX1 promotes HCC metastasis by induction and stabilization of HIF1a. | |||
| 4. MAZ contributes to Prox1 isoform expressions in HCC. | |||
| 5. PROX1 positively regulate HCC proliferation and sorafenib resistance by enhancing β-catenin signaling. | |||
| short stature homeobox 2 (SHOX2) | cell lines: HepG2, Huh7, and SMMC772 HCC tissues | SHOX2 gene is associated with poor prognosis | [74] |
| sineoculis homeobox homolog 1 (SIX1) | cell lines: HepG2 HCC tissues | The expression status of SIX1 is associated with the five-year survival rate duration of patients with early stage (I-II) of HCC, but not the advanced stage (III–IV) of HCC | [75] |
| zinc finger E-box binding homeobox 1/2 (ZEB1/2) | cell lines: Huh-7, HepG2, SMMC7721, Hep3B, SNU449, MHCC97H, HCCLM3, BEL-7402 QGY-7701, PLC5, and SK-Hep1 mouse model HCC tissues | ZEB1/2 is a transcription factor as a hub that promotes tumor invasion and metastasis by inducing EMT (detail in the text) | [76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, K.-Y.; Wang, L.-T.; Hsu, S.-H.; Wang, S.-N. Homeobox Genes and Hepatocellular Carcinoma. Cancers 2019, 11, 621. https://doi.org/10.3390/cancers11050621
Liu K-Y, Wang L-T, Hsu S-H, Wang S-N. Homeobox Genes and Hepatocellular Carcinoma. Cancers. 2019; 11(5):621. https://doi.org/10.3390/cancers11050621
Chicago/Turabian StyleLiu, Kwei-Yan, Li-Ting Wang, Shih-Hsien Hsu, and Shen-Nien Wang. 2019. "Homeobox Genes and Hepatocellular Carcinoma" Cancers 11, no. 5: 621. https://doi.org/10.3390/cancers11050621
APA StyleLiu, K.-Y., Wang, L.-T., Hsu, S.-H., & Wang, S.-N. (2019). Homeobox Genes and Hepatocellular Carcinoma. Cancers, 11(5), 621. https://doi.org/10.3390/cancers11050621





