The Flavonoid Metabolite 2,4,6-Trihydroxybenzoic Acid Is a CDK Inhibitor and an Anti-Proliferative Agent: A Potential Role in Cancer Prevention
Abstract
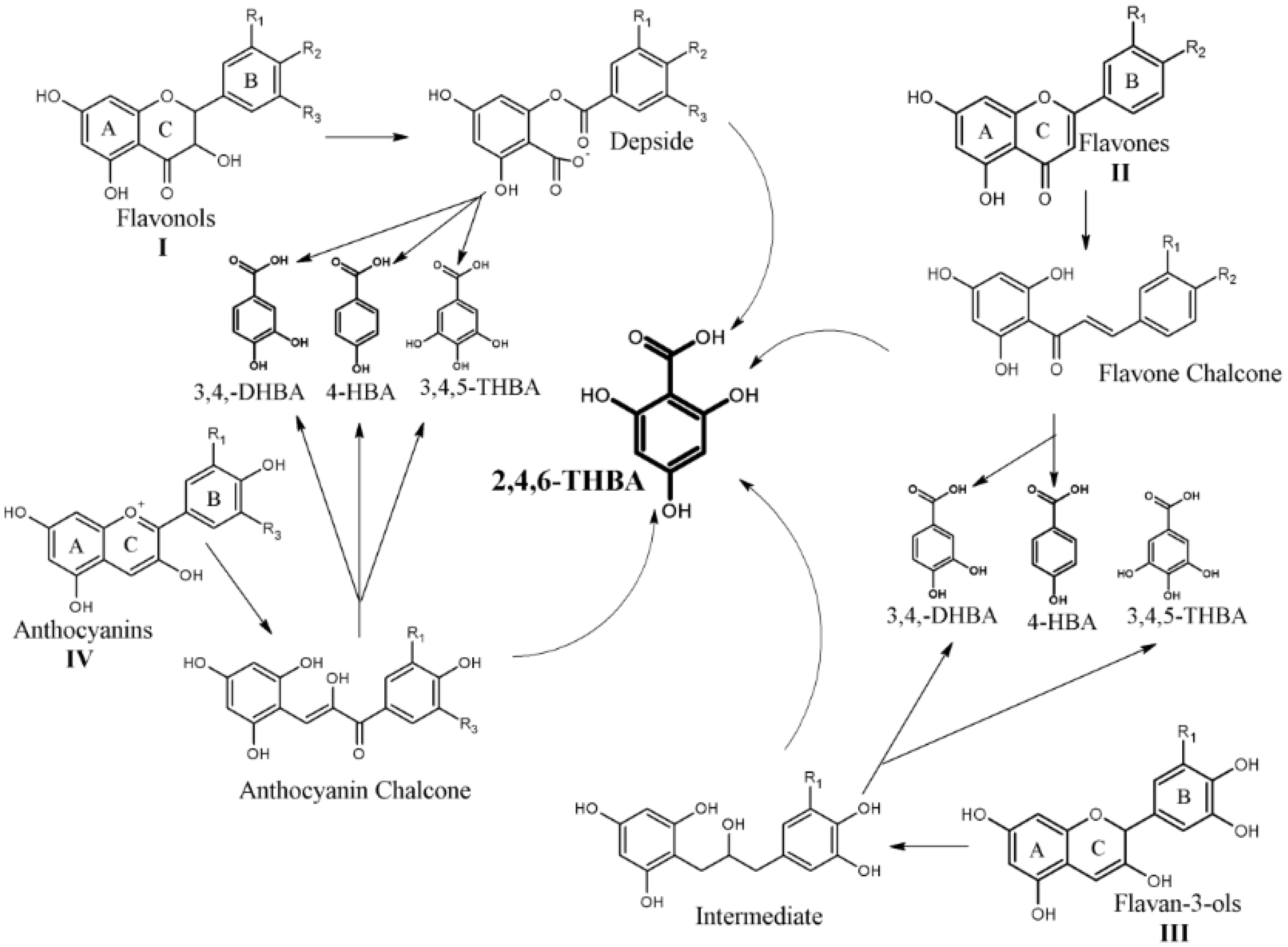
1. Introduction
2. Results
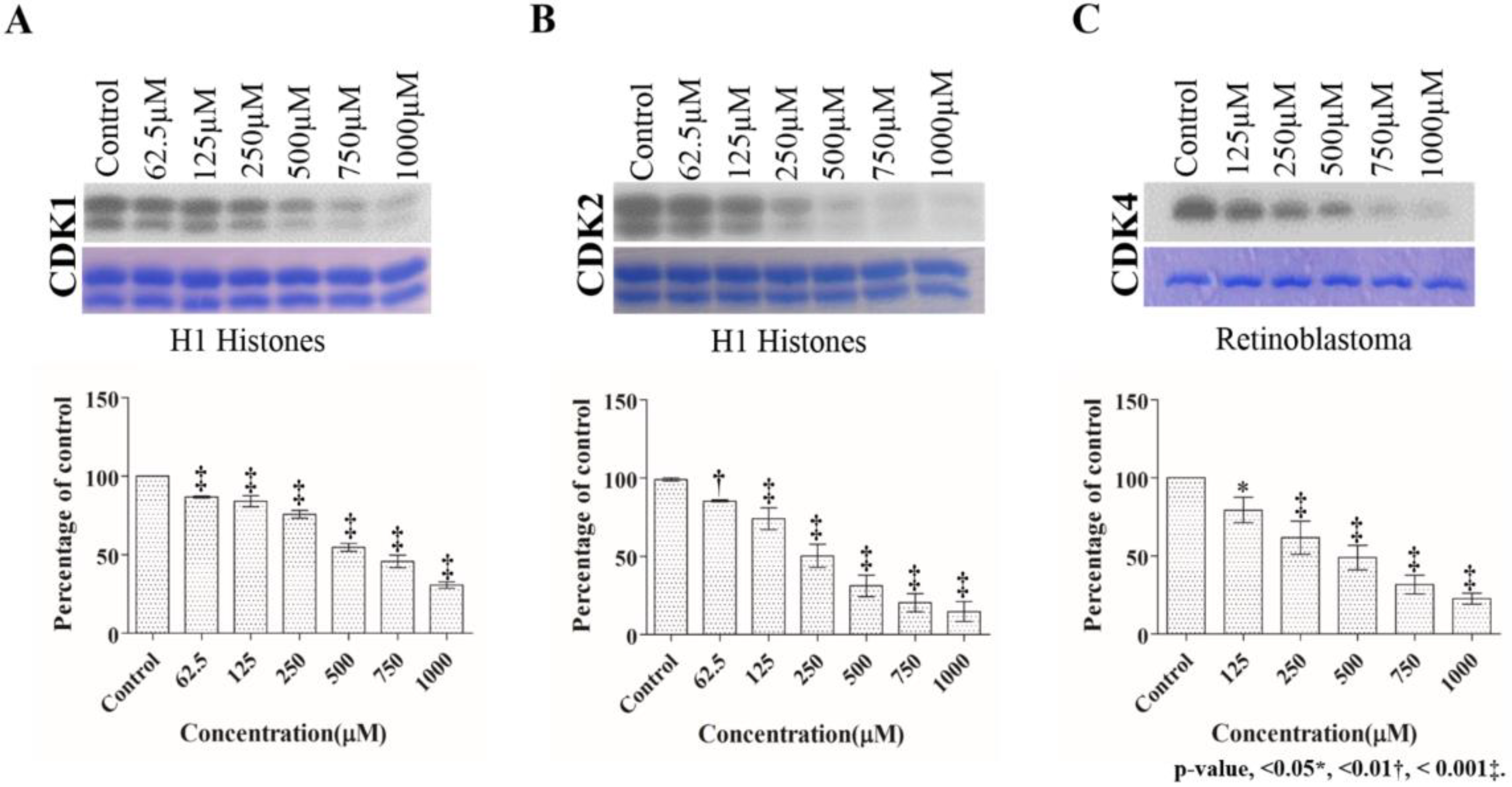
2.1. Dose Dependent Effect of 2,4,6-THBA on CDK 1, 2 and 4 Enzyme Activity
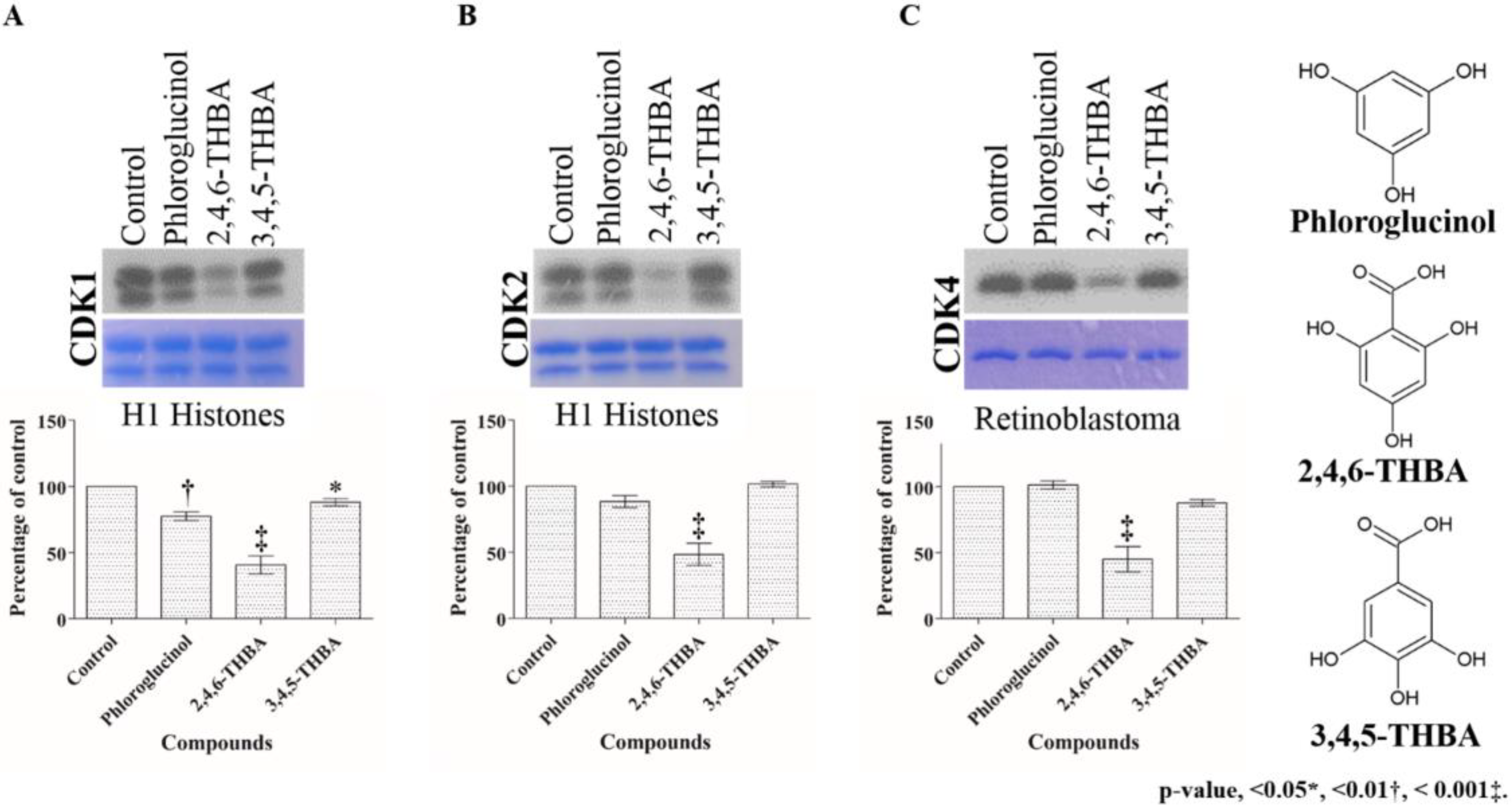
2.2. Comparison of the Effect of 2,4,6-THBA, Phloroglucinol and 3,4,5-THBA on CDK Enzyme Activity
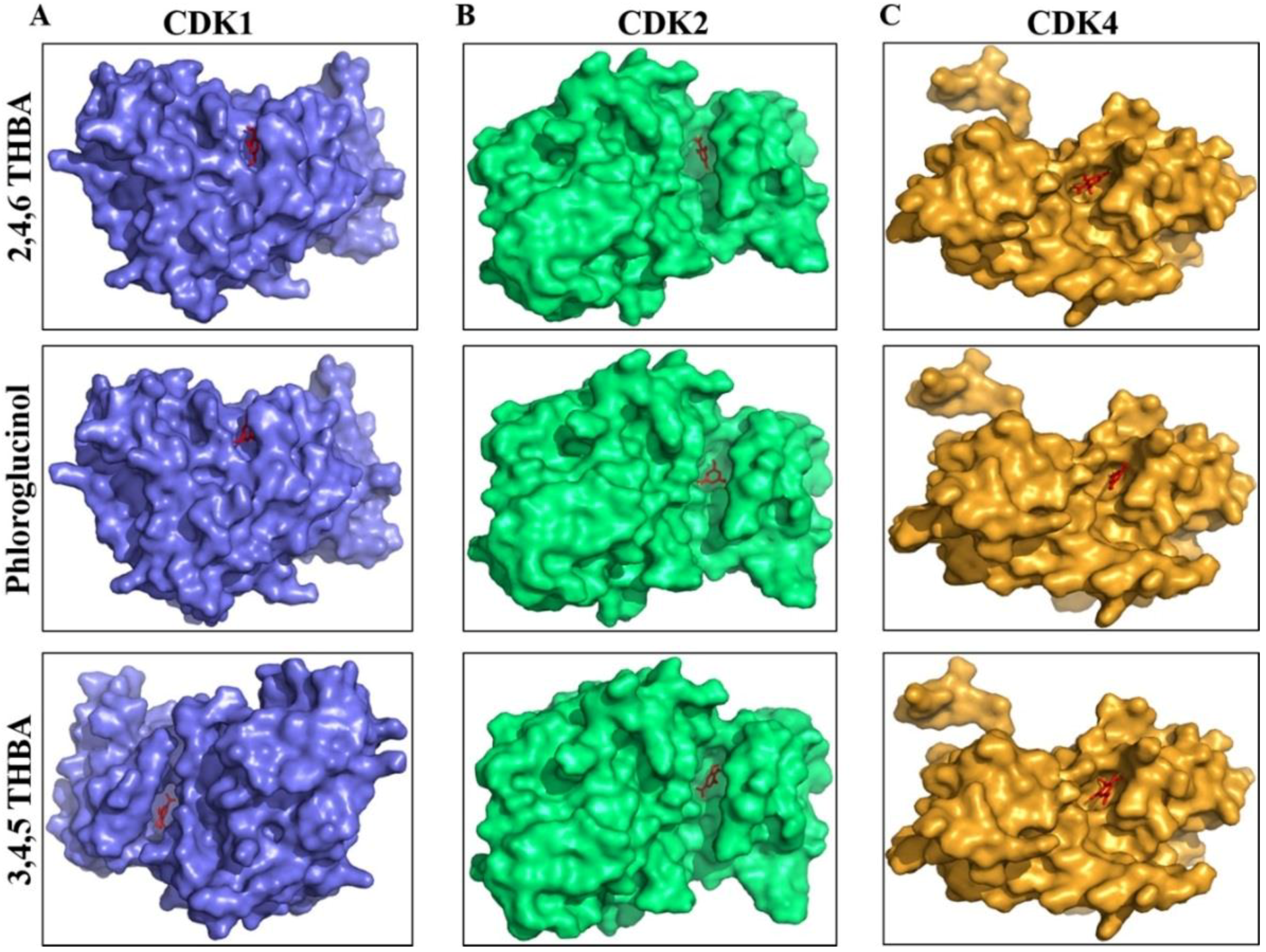
2.3. Molecular Docking Studies Show Potential Interactions of 2,4,6-THBA, Phloroglucinol and 3,4,5-THBA with CDKs
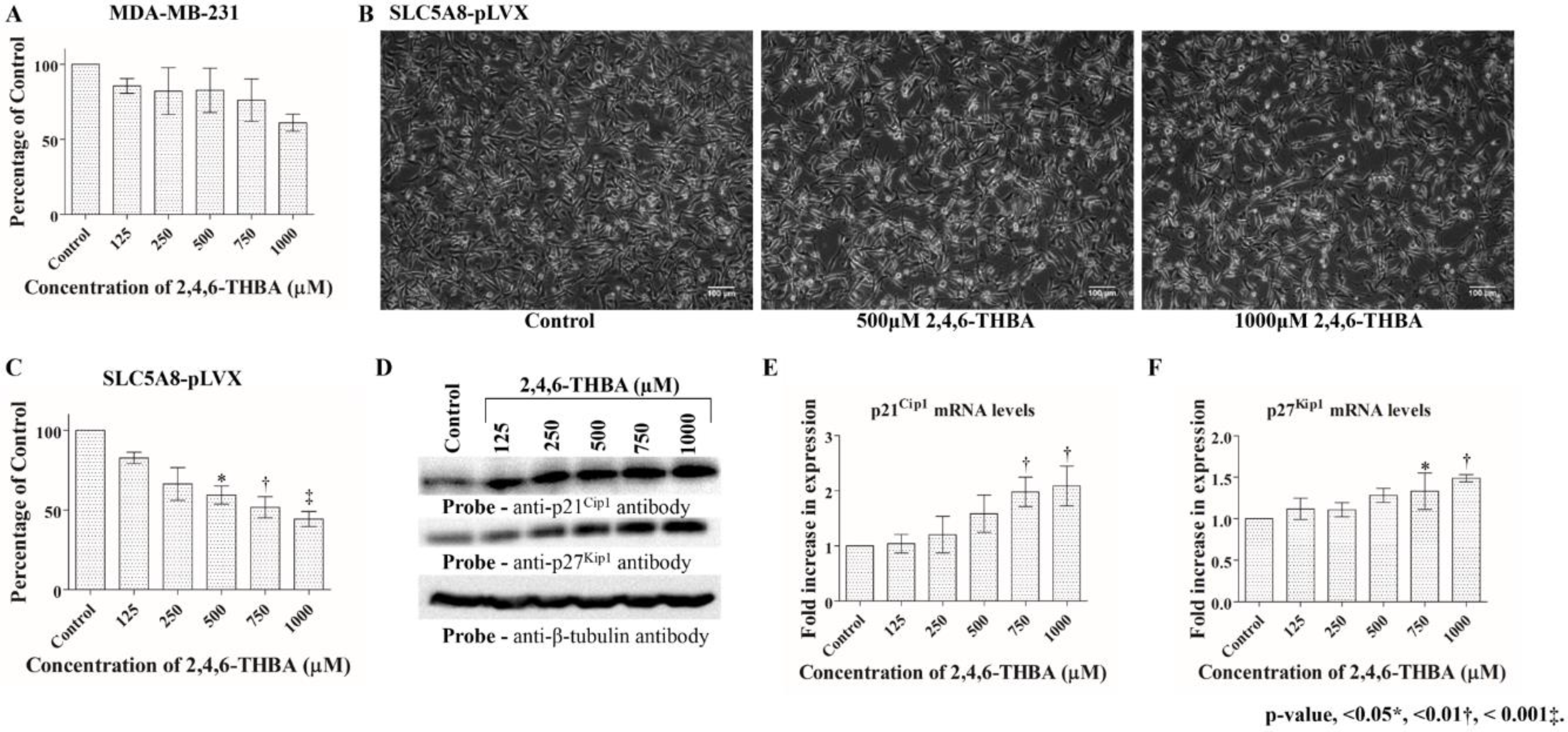
2.4. 2,4,6-THBA Inhibits Cell Proliferation in Cell Lines Expressing Functional SLC5A8
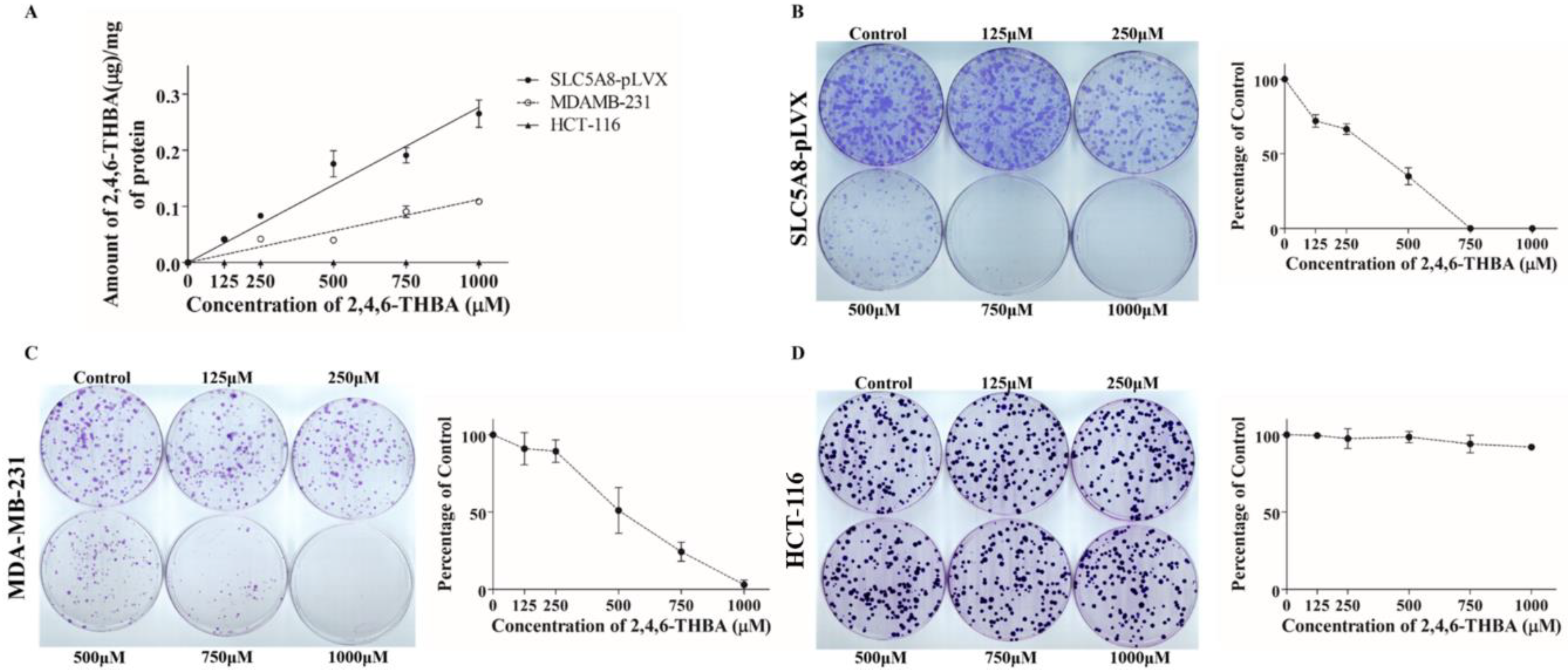
2.5. 2,4,6-THBA Is Taken Up by Cells Expressing Functional SLC5A8
2.6. 2,4,6-THBA Inhibits Colony Formation in Cells Expressing Functional SLC5A8
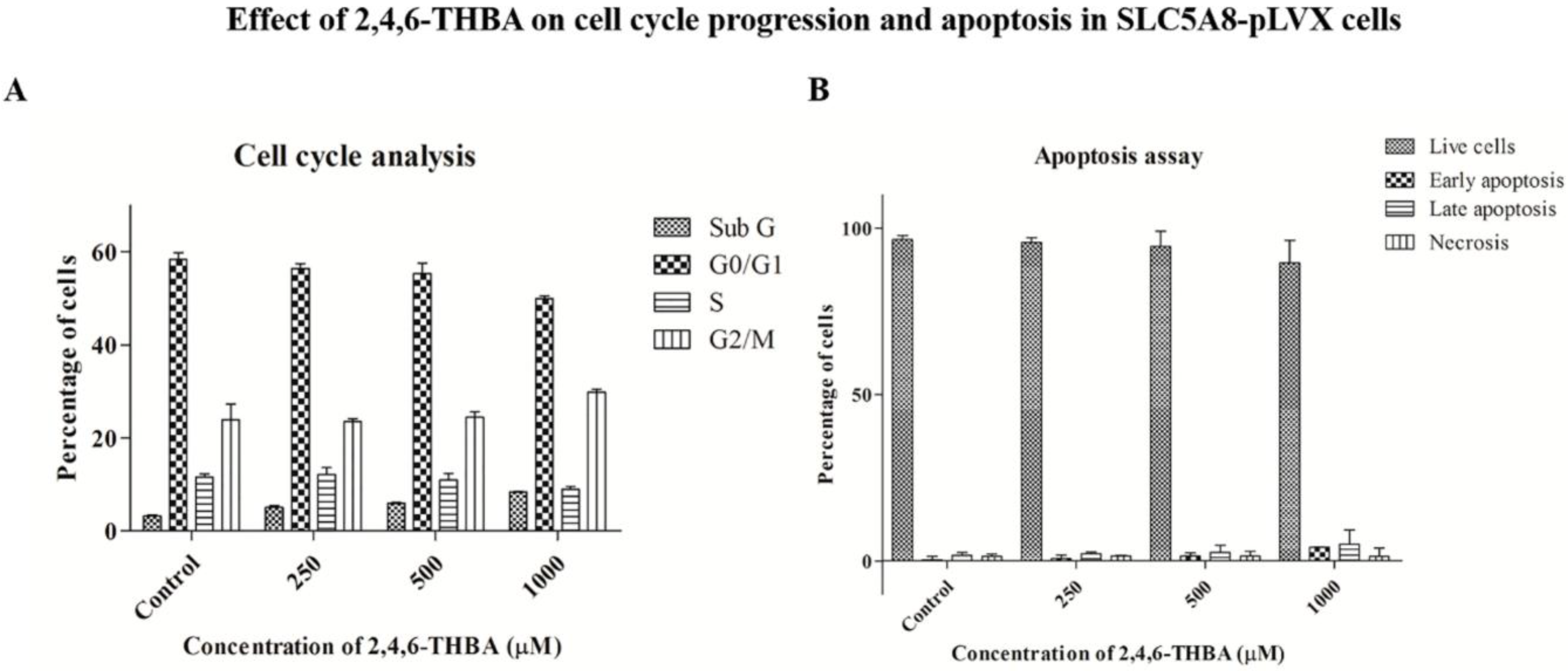
2.7. Cell Cycle Analysis and Apoptosis Assay
2.8. Effect of Other Flavonoid Metabolites (4-HBA; 3,4-DHBA; and 3,4,5-THBA) on Clonal Formation
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. Reagents
4.3. Recombinant Proteins and Antibodies
4.4. Cell Lysate Preparation and Western Blotting
4.5. In Vitro CDK Assay
4.6. Molecular Docking Studies
4.7. HPLC Analysis
4.8. RNA Isolation and qRT-PCR
4.9. Flow Cytometric Analysis
4.10. Clonogenic Assay
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fedewa, S.A.; Ahnen, D.J.; Meester, R.G.S.; Barzi, A.; Jemal, A. Colorectal cancer statistics, 2017. CA Cancer J. Clin. 2017, 67, 177–193. [Google Scholar] [CrossRef]
- Khan, N.; Mukhtar, H. Tea polyphenols for health promotion. Life Sci. 2007, 81, 519–533. [Google Scholar] [CrossRef]
- Wang, L.S.; Stoner, G.D. Anthocyanins and their role in cancer prevention. Cancer Lett. 2008, 269, 281–290. [Google Scholar] [CrossRef]
- Li, D.; Wang, P.; Luo, Y.; Zhao, M.; Chen, F. Health benefits of anthocyanins and molecular mechanisms: Update from recent decade. Crit. Rev. Food Sci. Nutr. 2017, 57, 1729–1741. [Google Scholar] [CrossRef]
- Panche, A.N.; Diwan, A.D.; Chandra, S.R. Flavonoids: An overview. J. Nutr. Sci. 2016, 5, e47. [Google Scholar] [CrossRef] [PubMed]
- Borah, N.; Gunawardana, S.; Torres, H.; McDonnell, S.; Van Slambrouck, S. 5,6,7,3′,4′,5′-Hexamethoxyflavone inhibits growth of triple-negative breast cancer cells via suppression of MAPK and Akt signaling pathways and arresting cell cycle. Int. J. Oncol. 2017, 51, 1685–1693. [Google Scholar] [CrossRef] [PubMed]
- Gao, K.; Xu, A.; Krul, C.; Venema, K.; Liu, Y.; Niu, Y.; Lu, J.; Bensoussan, L.; Seeram, N.P.; Heber, D.; et al. Of the Major Phenolic Acids Formed during Human Microbial Fermentation of Tea, Citrus, and Soy Flavonoid Supplements, only 3,4-Dihydroxyphenylacetic Acid Has Antiproliferative Activity. J. Nutr. 2006, 136, 52–57. [Google Scholar] [CrossRef] [PubMed]
- Hanske, L.; Engst, W.; Loh, G.; Sczesny, S.; Blaut, M.; Braune, A. Contribution of gut bacteria to the metabolism of cyanidin 3-glucoside in human microbiota-associated rats. Br. J. Nutr. 2013, 109, 1433–1441. [Google Scholar] [CrossRef]
- Wang, R.; Zhou, W.; Jiang, X. Reaction Kinetics of Degradation and Epimerization of Epigallocatechin Gallate (EGCG) in Aqueous System over a Wide Temperature Range. J. Agric. Food Chem. 2008, 56, 2694–2701. [Google Scholar] [CrossRef]
- Seeram, N.P.; Bourquin, L.D.; Nair, M.G. Degradation Products of Cyanidin Glycosides from Tart Cherries and Their Bioactivities. J. Agric. Food Chem. 2001, 49, 4924–4929. [Google Scholar] [CrossRef]
- Simons, A.L.; Renouf, M.; Hendrich, S.; Murphy, P.A. Human Gut Microbial Degradation of Flavonoids: Structure−Function Relationships. J. Agric. Food Chem. 2005, 53, 4258–4263. [Google Scholar] [CrossRef]
- Bermudezsoto, M.; Tomasbarberan, F.; Garciaconesa, M. Stability of polyphenols in chokeberry (Aronia melanocarpa) subjected to in vitro gastric and pancreatic digestion. Food Chem. 2007, 102, 865–874. [Google Scholar] [CrossRef]
- Ozdal, T.; Sela, D.A.; Xiao, J.; Boyacioglu, D.; Chen, F.; Capanoglu, E. The Reciprocal Interactions between Polyphenols and Gut Microbiota and Effects on Bioaccessibility. Nutrients 2016, 8, 78. [Google Scholar] [CrossRef] [PubMed]
- Ottaviani, J.I.; Borges, G.; Momma, T.Y.; Spencer, J.P.; Keen, C.L.; Crozier, A.; Schroeter, H. The metabolome of [2-(14)C](-)-epicatechin in humans: Implications for the assessment of efficacy, safety, and mechanisms of action of polyphenolic bioactives. Sci. Rep. 2016, 6, 29034. [Google Scholar] [CrossRef]
- Jenner, A.M.; Rafter, J.; Halliwell, B. Human fecal water content of phenolics: The extent of colonic exposure to aromatic compounds. Free Radic. Biol. Med. 2005, 38, 763–772. [Google Scholar] [CrossRef]
- Brown, S.B.; Rajananda, V.; Holroyd, J.A.; Evans, E.G. A study of the mechanism of quercetin oxygenation by 18O labelling. A comparison of the mechanism with that of haem degradation. Biochem. J. 1982, 205, 239–244. [Google Scholar] [CrossRef]
- Braune, A.; Blaut, M. Bacterial species involved in the conversion of dietary flavonoids in the human gut. Gut Microbes 2016, 7, 216–234. [Google Scholar] [CrossRef] [PubMed]
- Fleschhut, J.; Kratzer, F.; Rechkemmer, G.; Kulling, S.E. Stability and biotransformation of various dietary anthocyanins in vitro. Eur. J. Nutr. 2006, 45, 7–18. [Google Scholar] [CrossRef]
- Takagaki, A.; Nanjo, F. Metabolism of (-)-epigallocatechin gallate by rat intestinal flora. J. Agric. Food Chem. 2010, 58, 1313–1321. [Google Scholar] [CrossRef] [PubMed]
- Schantz, M.; Erk, T.; Richling, E. Metabolism of green tea catechins by the human small intestine. Biotechnol. J. 2010, 5, 1050–1059. [Google Scholar] [CrossRef]
- Woodward, G.; Kroon, P.; Cassidy, A.; Kay, C. Anthocyanin stability and recovery: Implications for the analysis of clinical and experimental samples. J. Agric. Food Chem. 2009, 57, 5271–5278. [Google Scholar] [CrossRef]
- Keppler, K.; Humpf, H.U. Metabolism of anthocyanins and their phenolic degradation products by the intestinal microflora. Bioorg. Med. Chem. 2005, 13, 5195–5205. [Google Scholar] [CrossRef] [PubMed]
- Peiffer, D.S.; Zimmerman, N.P.; Wang, L.S.; Ransom, B.W.; Carmella, S.G.; Kuo, C.T.; Siddiqui, J.; Chen, J.H.; Oshima, K.; Huang, Y.W.; et al. Chemoprevention of esophageal cancer with black raspberries, their component anthocyanins, and a major anthocyanin metabolite, protocatechuic acid. Cancer Prev. Res. 2014, 7, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Raina, K.; Rajamanickam, S.; Deep, G.; Singh, M.; Agarwal, R.; Agarwal, C. Chemopreventive effects of oral gallic acid feeding on tumor growth and progression in TRAMP mice. Mol. Cancer Ther. 2008, 7, 1258–1267. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Singh, A.; Mishra, A. Gallic acid: Molecular rival of cancer. Environ. Toxicol. Pharmacol. 2013, 35, 473–485. [Google Scholar] [CrossRef] [PubMed]
- Forester, S.C.; Choy, Y.Y.; Waterhouse, A.L.; Oteiza, P.I. The anthocyanin metabolites gallic acid, 3-O-methylgallic acid, and 2,4,6-trihydroxybenzaldehyde decrease human colon cancer cell viability by regulating pro-oncogenic signals. Mol. Carcinog. 2014, 53, 432–439. [Google Scholar] [CrossRef]
- Dachineni, R.; Kumar, D.R.; Callegari, E.; Kesharwani, S.S.; Sankaranarayanan, R.; Seefeldt, T.; Tummala, H.; Bhat, G.J. Salicylic acid metabolites and derivatives inhibit CDK activity: Novel insights into aspirin’s chemopreventive effects against colorectal cancer. Int. J. Oncol. 2017, 51, 1661–1673. [Google Scholar] [CrossRef] [PubMed]
- Coxon, C.R.; Anscombe, E.; Harnor, S.J.; Martin, M.P.; Carbain, B.; Golding, B.T.; Hardcastle, I.R.; Harlow, L.K.; Korolchuk, S.; Matheson, C.J.; et al. Cyclin-Dependent Kinase (CDK) Inhibitors: Structure-Activity Relationships and Insights into the CDK-2 Selectivity of 6-Substituted 2-Arylaminopurines. J. Med. Chem. 2017, 60, 1746–1767. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.Y.; McNae, I.; Kontopidis, G.; McClue, S.J.; McInnes, C.; Stewart, K.J.; Wang, S.; Zheleva, D.I.; Marriage, H.; Lane, D.P.; et al. Discovery of a novel family of CDK inhibitors with the program LIDAEUS: Structural basis for ligand-induced disordering of the activation loop. Structure 2003, 11, 399–410. [Google Scholar] [CrossRef]
- Coady, M.J.; Chang, M.H.; Charron, F.M.; Plata, C.; Wallendorff, B.; Sah, J.F.; Markowitz, S.D.; Romero, M.F.; Lapointe, J.Y. The human tumour suppressor gene SLC5A8 expresses a Na+-monocarboxylate cotransporter. J. Physiol. 2004, 557, 719–731. [Google Scholar] [CrossRef]
- Iwanaga, T.; Takebe, K.; Kato, I.; Karaki, S.-I.; Kuwahara, A. Cellular expression of monocarboxylate transporters (MCT) in the digestive tract of the mouse, rat, and humans, with special reference to slc5a8. Biomed. Res. 2006, 27, 243–254. [Google Scholar] [CrossRef] [PubMed]
- Ganapathy, V.; Thangaraju, M.; Gopal, E.; Martin, P.M.; Itagaki, S.; Miyauchi, S.; Prasad, P.D. Sodium-coupled monocarboxylate transporters in normal tissues and in cancer. AAPS J. 2008, 10, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Myeroff, L.; Smiraglia, D.; Romero, M.F.; Pretlow, T.P.; Kasturi, L.; Lutterbaugh, J.; Rerko, R.M.; Casey, G.; Issa, J.P.; et al. SLC5A8, a sodium transporter, is a tumor suppressor gene silenced by methylation in human colon aberrant crypt foci and cancers. Proc. Natl. Acad. Sci. USA 2003, 100, 8412–8417. [Google Scholar] [CrossRef] [PubMed]
- Coothankandaswamy, V.; Elangovan, S.; Singh, N.; Prasad, P.D.; Thangaraju, M.; Ganapathy, V. The plasma membrane transporter SLC5A8 suppresses tumour progression through depletion of survivin without involving its transport function. Biochem. J. 2013, 450, 169–178. [Google Scholar] [CrossRef]
- Babu, E.; Ramachandran, S.; CoothanKandaswamy, V.; Elangovan, S.; Prasad, P.D.; Ganapathy, V.; Thangaraju, M. Role of SLC5A8, a plasma membrane transporter and a tumor suppressor, in the antitumor activity of dichloroacetate. Oncogene 2011, 30, 4026–4037. [Google Scholar] [CrossRef]
- Jeffrey, P.D.; Russo, A.A.; Polyak, K.; Gibbs, E.; Hurwitz, J.; Massague, J.; Pavletich, N.P. Mechanism of CDK activation revealed by the structure of a cyclinA-CDK2 complex. Nature 1995, 376, 313–320. [Google Scholar] [CrossRef]
- Peyressatre, M.; Prevel, C.; Pellerano, M.; Morris, M.C. Targeting cyclin-dependent kinases in human cancers: From small molecules to Peptide inhibitors. Cancers 2015, 7, 179–237. [Google Scholar] [CrossRef]
- Zhang, Y.; Bao, Y.L.; Wu, Y.; Yu, C.L.; Sun, Y.; Li, Y.X. Identification and characterization of the human SLC5A8 gene promoter. Cancer Genet. Cytogenet. 2010, 196, 124–132. [Google Scholar] [CrossRef]
- Scott, S.C.; Lee, S.S.; Abraham, J. Mechanisms of therapeutic CDK4/6 inhibition in breast cancer. Semin. Oncol. 2017, 44, 385–394. [Google Scholar] [CrossRef]
- Siddiqui, I.A.; Sanna, V.; Ahmad, N.; Sechi, M.; Mukhtar, H. Resveratrol nanoformulation for cancer prevention and therapy. Ann. N. Y. Acad. Sci. 2015, 1348, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Wattenberg, L.W. Chemoprevention of Cancer. Cancer Res. 1985, 45, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Nonato, C.F.A.; Leite, D.O.D.; Pereira, R.C.; Boligon, A.A.; Ribeiro-Filho, J.; Rodrigues, F.F.G.; da Costa, J.G.M. Chemical analysis and evaluation of antioxidant and antimicrobial activities of fruit fractions of Mauritia flexuosa L. f. (Arecaceae). PeerJ 2018, 6, e5991. [Google Scholar] [CrossRef] [PubMed]
- Kumar, B.R. Application of HPLC and ESI-MS techniques in the analysis of phenolic acids and flavonoids from green leafy vegetables (GLVs). J. Pharm. Anal. 2017, 7, 349–364. [Google Scholar] [CrossRef] [PubMed]
- Prihantini, A.I.; Tachibana, S.; Itoh, K. Antioxidant Active Compounds from Elaeocarpussylvestris and their Relationship between Structure and Activity. Procedia Environ. Sci. 2015, 28, 758–768. [Google Scholar] [CrossRef]
- Martinez-Lostao, L.; Anel, A.; Pardo, J. How Do Cytotoxic Lymphocytes Kill Cancer Cells? Clin. Cancer Res. 2015, 21, 5047–5056. [Google Scholar] [CrossRef] [PubMed]
- Marcus, A.; Gowen, B.G.; Thompson, T.W.; Iannello, A.; Ardolino, M.; Deng, W.; Wang, L.; Shifrin, N.; Raulet, D.H. Recognition of tumors by the innate immune system and natural killer cells. Adv. Immunol. 2014, 122, 91–128. [Google Scholar] [CrossRef]
- Ai, G.; Dachineni, R.; Muley, P.; Tummala, H.; Bhat, G.J. Aspirin and salicylic acid decrease c-Myc expression in cancer cells: A potential role in chemoprevention. Tumour Biol. 2016, 37, 1727–1738. [Google Scholar] [CrossRef] [PubMed]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J. GROMACS: Fast, flexible, and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef]
- Alfonso, L.F.; Srivenugopal, K.S.; Arumugam, T.V.; Abbruscato, T.J.; Weidanz, J.A.; Bhat, G.J. Aspirin inhibits camptothecin-induced p21CIP1 levels and potentiates apoptosis in human breast cancer cells. Int. J. Oncol. 2009, 34, 597–608. [Google Scholar]
- Zhou, S.; Xia, H.; Xu, H.; Tang, Q.; Nie, Y.; Gong, Q.Y.; Bi, F. ERRalpha suppression enhances the cytotoxicity of the MEK inhibitor trametinib against colon cancer cells. J. Exp. Clin. Cancer Res. 2018, 37, 218. [Google Scholar] [CrossRef] [PubMed]
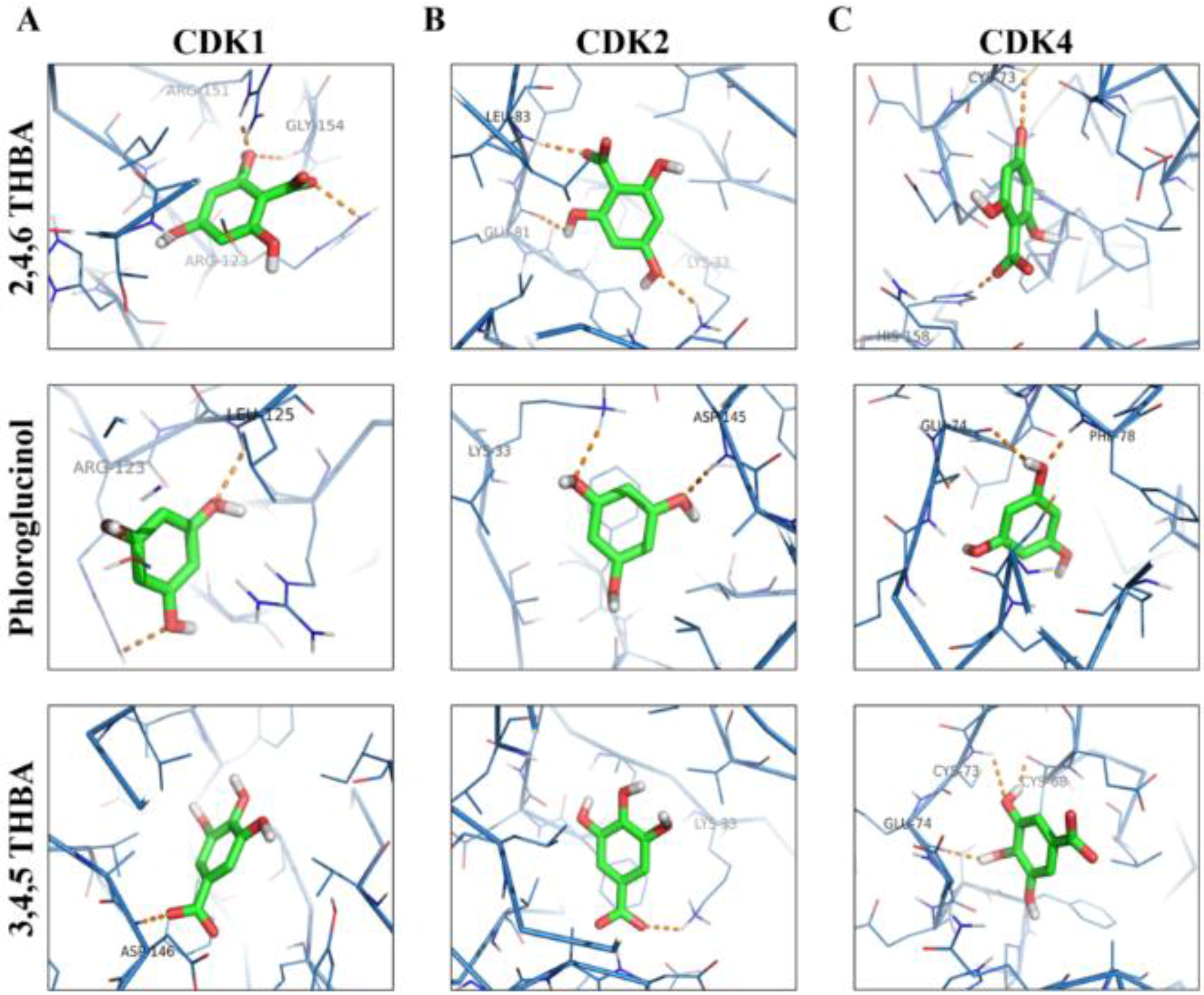
| S.No | Receptor | Ligand | Interacting Amino Acids | Measurement (A°) | No of H Bonds | Energy Value (kCal/mol) |
|---|---|---|---|---|---|---|
| 1. | CDK1 | 2,4,6-THBA | Arg123, Gly154, Arg151 | 2.5, 2.4, 2.0 | 3 | −6.0 |
| 3,4,5-THBA | Asp146 | 2.1 | 1 | −6.1 | ||
| Phloroglucinol | Leu125, Arg123 | 2.4, 2.6 | 2 | −4.9 | ||
| 3,4-DHBA | Asp146 | 2.8 | 1 | −6.3 | ||
| 4-HBA | - | - | 0 | −6.4 | ||
| 2 | CDK2 | 2,4,6-THBA | Leu83, Gln81, Lys33 | 2.2, 2.3, 2.4 | 3 | −5.6 |
| 3,4,5-THBA | Lys33 | 2.5 | 1 | −5.8 | ||
| Phloroglucinol | Asp145, Lys33 | 2.2,2.4 | 2 | −5.2 | ||
| 3,4-DHBA | Leu83(2), Glu81 | 3.2, 3.1, 2.0 | 3 | −5.8 | ||
| 4-HBA | Lys33 | 2.9 | 1 | −5.7 | ||
| 3. | CDK4 | 2,4,6-THBA | His158, Cys73 | 21, 2.8 | 2 | −5.8 |
| 3,4,5-THBA | Glu74, Cys73, Cys68 | 2.2, 2.8, 1.9 | 3 | −5.8 | ||
| Phloroglucinol | Phe78, Glu74 | 2.1, 2.7 | 2 | −5.0 | ||
| 3,4-DHBA | Cys68, Glu75 | 2.5, 2.7 | 2 | −5.7 | ||
| 4-HBA | His158 | 3.2 | 1 | −5.7 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sankaranarayanan, R.; Valiveti, C.K.; Kumar, D.R.; Van slambrouck, S.; Kesharwani, S.S.; Seefeldt, T.; Scaria, J.; Tummala, H.; Bhat, G.J. The Flavonoid Metabolite 2,4,6-Trihydroxybenzoic Acid Is a CDK Inhibitor and an Anti-Proliferative Agent: A Potential Role in Cancer Prevention. Cancers 2019, 11, 427. https://doi.org/10.3390/cancers11030427
Sankaranarayanan R, Valiveti CK, Kumar DR, Van slambrouck S, Kesharwani SS, Seefeldt T, Scaria J, Tummala H, Bhat GJ. The Flavonoid Metabolite 2,4,6-Trihydroxybenzoic Acid Is a CDK Inhibitor and an Anti-Proliferative Agent: A Potential Role in Cancer Prevention. Cancers. 2019; 11(3):427. https://doi.org/10.3390/cancers11030427
Chicago/Turabian StyleSankaranarayanan, Ranjini, Chaitanya K. Valiveti, D. Ramesh Kumar, Severine Van slambrouck, Siddharth S. Kesharwani, Teresa Seefeldt, Joy Scaria, Hemachand Tummala, and G. Jayarama Bhat. 2019. "The Flavonoid Metabolite 2,4,6-Trihydroxybenzoic Acid Is a CDK Inhibitor and an Anti-Proliferative Agent: A Potential Role in Cancer Prevention" Cancers 11, no. 3: 427. https://doi.org/10.3390/cancers11030427
APA StyleSankaranarayanan, R., Valiveti, C. K., Kumar, D. R., Van slambrouck, S., Kesharwani, S. S., Seefeldt, T., Scaria, J., Tummala, H., & Bhat, G. J. (2019). The Flavonoid Metabolite 2,4,6-Trihydroxybenzoic Acid Is a CDK Inhibitor and an Anti-Proliferative Agent: A Potential Role in Cancer Prevention. Cancers, 11(3), 427. https://doi.org/10.3390/cancers11030427






