FBXW7 in Cancer: What Has Been Unraveled Thus Far?
Abstract
:1. Introduction
2. FBXW7
3. Substrates of FBXW7
4. Regulation of FBXW7
4.1. p53
4.2. C/EBP-δ
4.3. Pin1
4.4. Other Regulators
5. Genetic and Epigenetic Alterations of FBXW7 in Cancer
6. Deregulation of FBXW7 in Different Types of Cancer
6.1. Brain Cancer
6.2. Breast Cancer
6.3. Colorectal Cancer (CRC)
6.4. Esophageal Cancer
6.5. Gastric Cancer (GC)
6.6. Gynecological Cancers
6.7. Hepatocellular Carcinoma (HCC)
6.8. Leukemia
6.9. Lung Cancer
6.10. Pancreatic Cancer
6.11. Prostate Cancer
6.12. Renal Cancer
6.13. Skin Cancer
6.14. Other Cancers
7. Role of FBXW7 in Cancer Cell Chemosensitization
8. Conclusions and Future Perspective
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ABC-DLBCL | Activated B-cell like diffuse large B-cell lymphoma |
| API-1 | 4-amino-5,8-dihydro-5-oxo-8-β-D-ribofuranosyl-pyrido[2,3-d]pyrimidine-6-carboxamide |
| ATL | T-cell leukemia |
| BMSCs | Bone marrow-derived stromal cells |
| C/EBP-δ | CCAAT/enhancer-binding protein-δ |
| circRNAs | Circular RNAs |
| CML | Chronic myeloid leukemia |
| CPD | Cdc4 phosphodegron |
| CRC | Colorectal cancer |
| CSCs | Cancer stem cells |
| Cul1 | Cullin1 |
| DMBA | 7,12-dimethylbenz[a]anthracene |
| DNMT1 | DNA methyltransferase 1 |
| EGLN2 | Egl-9 Family Hypoxia-Inducible Factor 2 |
| EMT | Epithelial–mesenchymal transition |
| ENO1 | Enolase 1 |
| ENT1 | Equilibrative nucleoside transporter 1 |
| EOGC | Early-onset of gastric cancers |
| ERK | Extracellular signal-regulated kinase |
| ESCC | Esophageal squamous cell carcinoma |
| ESCs | Embryonic stem cells |
| EZH2 | Enhancer of zeste homolog 2 polycomb repressive complex 2 |
| FAM83D | Family with sequence similarity 83, member D |
| FBXL | F-box coupled with LRRs |
| FBXO | F-box with no motifs |
| FBXW | F-box coupled with WD repeats |
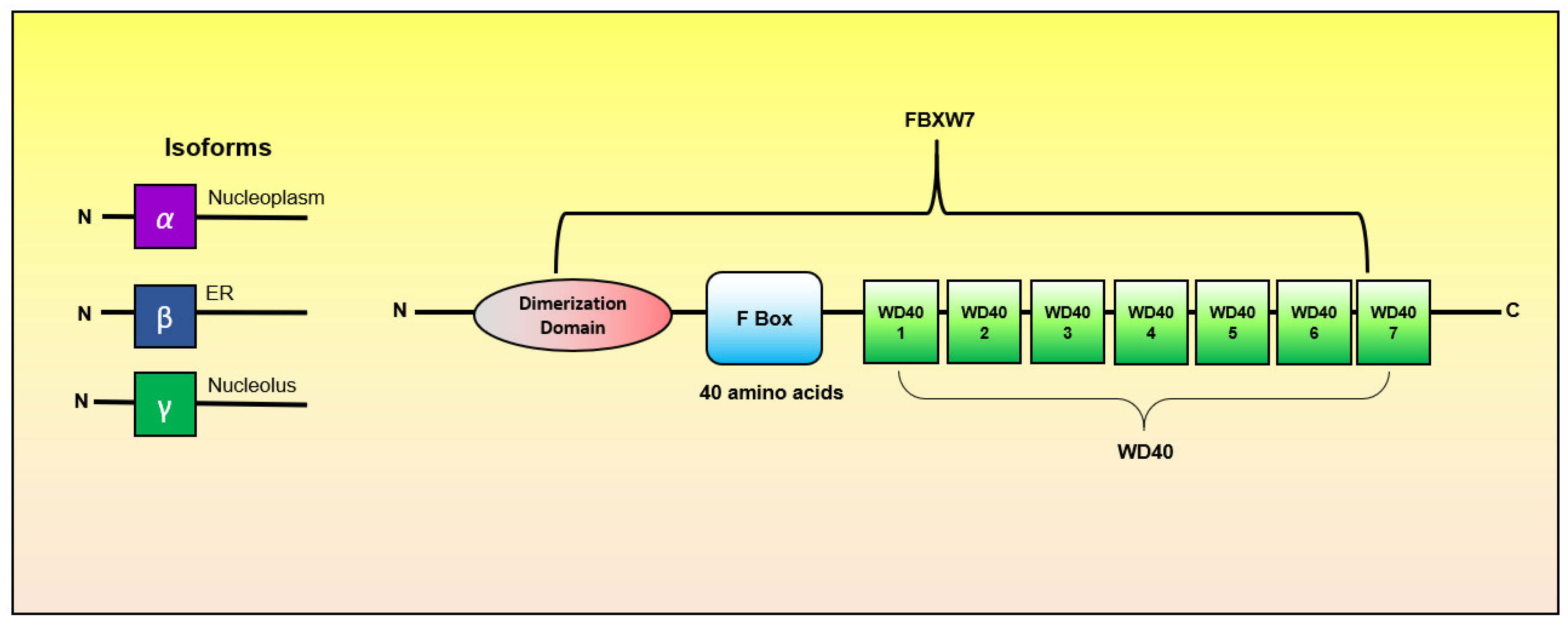
| FBXW7 | F-box with 7 tandem WD40 repeats |
| GC | Gastric cancer |
| GSI | Gamma-secretase inhibitors |
| GSK3 | Glycogen synthase kinase 3 |
| HDAC | Histone deacetylase |
| Hes-5 | Hairy and Enhancer-of-split homologues 5 |
| HIF-1α | Hypoxia-inducible factor-1α |
| HSF1 | Heat-shock factor 1 |
| Hsp90 | Heat shock protein 90 |
| IHCC | Intrahepatic cholangiocarcinoma |
| KLFs | Kruppel-like factors |
| LICs | Leukemia-initiating cells |
| LRR | Leucine-rich repeats |
| MALAT1 | Metastasis-associated lung adenocarcinoma transcript 1 |
| MAPK | Mitogen-activated protein kinase |
| MCL-1 | Myeloid cell leukemia 1 |
| MED13 | Mediator 13 |
| miR-10b | microRNA-10b |
| MiRNAs | MicroRNAs |
| MMP | Metalloproteinase |
| Mo-MDSCs | Monocytic myeloid-derived suppressor cells |
| MTDH | Metadherin |
| mTOR | Mammalian target of rapamycin |
| NF1 | Neurofibromatosis type 1 |
| NF-ΚB | Nuclear factor kappa B |
| NPC | Nasopharyngeal carcinoma |
| NRF1 | Nuclear factor E2-related factor 1 |
| NSC | Neural stem cell |
| NSCLC | Non-small cell lung cancer |
| PC | Pancreatic cancer |
| PDAC | Pancreatic ductal adenocarcinoma |
| Pin1 | Peptidyl-prolyl cis-trans isomerase NIMA-interacting1 |
| PLK1 | Polo-like kinase 1 |
| RCC | Renal cell carcinoma |
| RhoA | Ras homologue gene family member A |
| RhoGDI | RHO guanosine diphosphate dissociation inhibitor; |
| Rictor | Rapamycin-insensitive companion of mTOR |
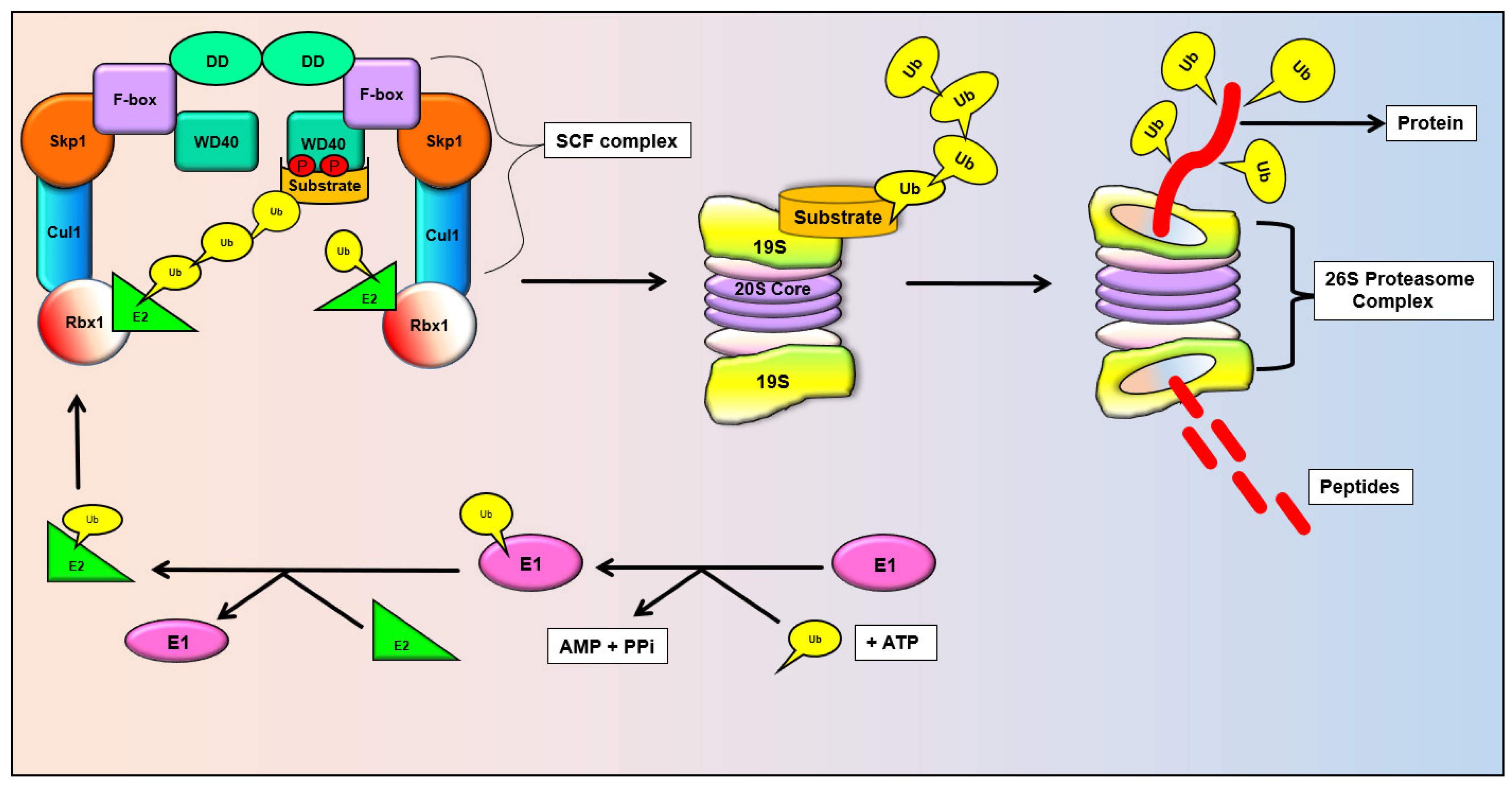
| SCF | Skp1-Cullin1-F-box |
| SCNC | Small cell neuroendocrine carcinoma |
| SINE | Specific inhibitors of nuclear export |
| Skp1 | S phase kinase-associated protein 1 |
| STAT3 | Signal transducer and activator of transcription 3 |
| T-ALL | T-cell acute lymphoblastic leukemia |
| TAMs | Tumor-associated macrophages |
| TINCR | Terminal differentiation-induced lncRNA |
| TNBC | Triple-negative breast cancer |
| TPA | 12-O-tetradecanoylphorbol-13-acetate |
| Ubc | Ubiquitin-conjugating enzyme |
| UPS | Ubiquitin-proteasome system |
| WT | Wilms’ tumor |
| YAP | Yes-associated protein |
| ZNF322A | Zinc-finger 322A |
References
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef] [PubMed]
- Zhao, E.; Maj, T.; Kryczek, I.; Li, W.; Wu, K.; Zhao, L.; Wei, S.; Crespo, J.; Wan, S.; Vatan, L.; et al. Cancer mediates effector T cell dysfunction by targeting microRNAs and EZH2 via glycolysis restriction. Nat. Immunol. 2016, 17, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Shabnam, B.; Padmavathi, G.; Banik, K.; Girisa, S.; Monisha, J.; Sethi, G.; Fan, L.; Wang, L.; Mao, X.; Kunnumakkara, A.B. Sorcin a Potential Molecular Target for Cancer Therapy. Transl. Oncol. 2018, 11, 1379–1389. [Google Scholar] [CrossRef] [PubMed]
- Ranaware, A.M.; Banik, K.; Deshpande, V.; Padmavathi, G.; Roy, N.K.; Sethi, G.; Fan, L.; Kumar, A.P.; Kunnumakkara, A.B. Magnolol: A Neolignan from the Magnolia Family for the Prevention and Treatment of Cancer. Int. J. Mol. Sci. 2018, 19, 2362. [Google Scholar] [CrossRef] [PubMed]
- Banik, K.; Harsha, C.; Bordoloi, D.; Lalduhsaki Sailo, B.; Sethi, G.; Leong, H.C.; Arfuso, F.; Mishra, S.; Wang, L.; Kumar, A.P.; et al. Therapeutic potential of gambogic acid, a caged xanthone, to target cancer. Cancer Lett. 2018, 416, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; He, C.; Wang, J. Regulation mechanism of Fbxw7-related signaling pathways (Review). Oncol. Rep. 2015, 34, 2215–2224. [Google Scholar] [CrossRef] [PubMed]
- Diaz, V.M.; de Herreros, A.G. F-box proteins: Keeping the epithelial-to-mesenchymal transition (EMT) in check. Semin. Cancer Biol. 2016, 36, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Zheng, N.; Zhou, Q.; Wang, Z.; Wei, W. Recent advances in SCF ubiquitin ligase complex: Clinical implications. Biochim. Biophys. Acta 2016, 1866, 12–22. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Cao, J.; Liu, G.; Huo, J.R. Function and mechanism of F-box proteins in gastric cancer (Review). Int. J. Oncol. 2015, 47, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Zheng, N.; Schulman, B.A.; Song, L.; Miller, J.J.; Jeffrey, P.D.; Wang, P.; Chu, C.; Koepp, D.M.; Elledge, S.J.; Pagano, M.; et al. Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 2002, 416, 703–709. [Google Scholar] [CrossRef] [PubMed]
- Wei, D.; Sun, Y. Small RING Finger Proteins RBX1 and RBX2 of SCF E3 Ubiquitin Ligases: The Role in Cancer and as Cancer Targets. Genes Cancer 2010, 1, 700–707. [Google Scholar] [CrossRef] [PubMed]
- Kipreos, E.T.; Pagano, M. The F-box protein family. Genome Biol. 2000, 1. [Google Scholar] [CrossRef] [PubMed]
- Davis, R.J.; Welcker, M.; Clurman, B.E. Tumor suppression by the Fbw7 ubiquitin ligase: Mechanisms and opportunities. Cancer Cell 2014, 26, 455–464. [Google Scholar] [CrossRef] [PubMed]
- Bailey, M.L.; Singh, T.; Mero, P.; Moffat, J.; Hieter, P. Dependence of Human Colorectal Cells Lacking the FBW7 Tumor Suppressor on the Spindle Assembly Checkpoint. Genetics 2015, 201, 885–895. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Chen, G.; Martinka, M.; Ho, V.; Li, G. Prognostic significance of Fbw7 in human melanoma and its role in cell migration. J. Investig. Dermatol. 2013, 133, 1794–1802. [Google Scholar] [CrossRef] [PubMed]
- Takeishi, S.; Nakayama, K.I. Role of Fbxw7 in the maintenance of normal stem cells and cancer-initiating cells. Br. J. Cancer 2014, 111, 1054–1059. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.M.; Wei, W.; Sun, Y. Role of SKP1-CUL1-F-box-protein (SCF) E3 ubiquitin ligases in skin cancer. J. Genet. Genom. 2013, 40, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Tang, J.; Men, W.; Ren, K. FBXW7-mediated degradation of CCDC6 is impaired by ATM during DNA damage response in lung cancer cells. FEBS Lett. 2012, 586, 4257–4263. [Google Scholar] [CrossRef] [PubMed]
- Akhoondi, S.; Lindstrom, L.; Widschwendter, M.; Corcoran, M.; Bergh, J.; Spruck, C.; Grander, D.; Sangfelt, O. Inactivation of FBXW7/hCDC4-beta expression by promoter hypermethylation is associated with favorable prognosis in primary breast cancer. Breast Cancer Res. 2010, 12, R105. [Google Scholar] [CrossRef] [PubMed]
- Spruck, C.H.; Strohmaier, H.; Sangfelt, O.; Muller, H.M.; Hubalek, M.; Muller-Holzner, E.; Marth, C.; Widschwendter, M.; Reed, S.I. hCDC4 gene mutations in endometrial cancer. Cancer Res. 2002, 62, 4535–4539. [Google Scholar] [PubMed]
- Busino, L.; Millman, S.E.; Scotto, L.; Kyratsous, C.A.; Basrur, V.; O’Connor, O.; Hoffmann, A.; Elenitoba-Johnson, K.S.; Pagano, M. Fbxw7alpha- and GSK3-mediated degradation of p100 is a pro-survival mechanism in multiple myeloma. Nat. Cell Biol. 2012, 14, 375–385. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Swartz, K.L.; Siu, K.T.; Bhattacharyya, M.; Minella, A.C. Fbw7-dependent cyclin E regulation ensures terminal maturation of bone marrow erythroid cells by restraining oxidative metabolism. Oncogene 2014, 33, 3161–3171. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, S.; Oike, Y.; Onoyama, I.; Iwama, A.; Arai, F.; Takubo, K.; Mashimo, Y.; Oguro, H.; Nitta, E.; Ito, K.; et al. Fbxw7 acts as a critical fail-safe against premature loss of hematopoietic stem cells and development of T-ALL. Genes Dev. 2008, 22, 986–991. [Google Scholar] [CrossRef] [PubMed]
- Kuiper, R.P.; Vreede, L.; Venkatachalam, R.; Ricketts, C.; Kamping, E.; Verwiel, E.; Govaerts, L.; Debiec-Rychter, M.; Lerut, E.; van Erp, F.; et al. The tumor suppressor gene FBXW7 is disrupted by a constitutional t(3;4)(q21;q31) in a patient with renal cell cancer. Cancer Genet. Cytogenet. 2009, 195, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Yokobori, T.; Mimori, K.; Iwatsuki, M.; Ishii, H.; Onoyama, I.; Fukagawa, T.; Kuwano, H.; Nakayama, K.I.; Mori, M. p53-Altered FBXW7 expression determines poor prognosis in gastric cancer cases. Cancer Res. 2009, 69, 3788–3794. [Google Scholar] [CrossRef] [PubMed]
- Jardim, D.L.; Wheler, J.J.; Hess, K.; Tsimberidou, A.M.; Zinner, R.; Janku, F.; Subbiah, V.; Naing, A.; Piha-Paul, S.A.; Westin, S.N.; et al. FBXW7 mutations in patients with advanced cancers: Clinical and molecular characteristics and outcomes with mTOR inhibitors. PLoS ONE 2014, 9, e89388. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Xiao, J.; Hu, K.; Wang, G.; Li, M.; Zhang, J.; Cheng, G. FBXW7 acts as an independent prognostic marker and inhibits tumor growth in human osteosarcoma. Int. J. Mol. Sci. 2015, 16, 2294–2306. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ye, X.; Liu, Y.; Wei, W.; Wang, Z. Aberrant regulation of FBW7 in cancer. Oncotarget 2014, 5, 2000–2015. [Google Scholar] [CrossRef] [PubMed]
- Perez-Losada, J.; Wu, D.; DelRosario, R.; Balmain, A.; Mao, J.H. Allele-specific deletions in mouse tumors identify Fbxw7 as germline modifier of tumor susceptibility. PLoS ONE 2012, 7, e31301. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Zhou, Y.; Liu, D.; Huo, J. F-box proteins involved in cancer-associated drug resistance. Oncol. Lett. 2018, 15, 8891–8900. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Rodriguez-Barrueco, R.; Yu, J.; Do, C.; Silva, J.M.; Gautier, J. MYC is a critical target of FBXW7. Oncotarget 2015, 6, 3292–3305. [Google Scholar] [CrossRef] [PubMed]
- Sancho, R.; Blake, S.M.; Tendeng, C.; Clurman, B.E.; Lewis, J.; Behrens, A. Fbw7 repression by hes5 creates a feedback loop that modulates Notch-mediated intestinal and neural stem cell fate decisions. PLoS Biol. 2013, 11, e1001586. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, V.; Bandapalli, O.R.; Rajkumar, T.; Sagar, T.G.; Karunakaran, N. NOTCH1 and FBXW7 mutations favor better outcome in pediatric South Indian T-cell acute lymphoblastic leukemia. J. Pediatr. Hematol. Oncol. 2015, 37, e23–e30. [Google Scholar] [CrossRef] [PubMed]
- Yumimoto, K.; Nakayama, K.I. Fbxw7 suppresses cancer metastasis by inhibiting niche formation. Oncoimmunology 2015, 4, e1022308. [Google Scholar] [CrossRef] [PubMed]
- Sethi, G.; Tergaonkar, V. Potential pharmacological control of the NF-kappaB pathway. Trends Pharmacol. Sci. 2009, 30, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Sethi, G. Targeting transcription factor NF-kappaB to overcome chemoresistance and radioresistance in cancer therapy. Biochim. Biophys. Acta 2010, 1805, 167–180. [Google Scholar] [CrossRef] [PubMed]
- Sethi, G.; Shanmugam, M.K.; Ramachandran, L.; Kumar, A.P.; Tergaonkar, V. Multifaceted link between cancer and inflammation. Biosci. Rep. 2012, 32, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Zhang, J.; Arfuso, F.; Chinnathambi, A.; Zayed, M.E.; Alharbi, S.A.; Kumar, A.P.; Ahn, K.S.; Sethi, G. NF-kappaB in cancer therapy. Arch. Toxicol. 2015, 89, 711–731. [Google Scholar] [CrossRef] [PubMed]
- Chai, E.Z.; Siveen, K.S.; Shanmugam, M.K.; Arfuso, F.; Sethi, G. Analysis of the intricate relationship between chronic inflammation and cancer. Biochem. J. 2015, 468, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Ahn, K.S.; Sethi, G.; Aggarwal, B.B. Nuclear factor-kappa B: From clone to clinic. Curr. Mol. Med. 2007, 7, 619–637. [Google Scholar] [CrossRef] [PubMed]
- Puar, Y.R.; Shanmugam, M.K.; Fan, L.; Arfuso, F.; Sethi, G.; Tergaonkar, V. Evidence for the Involvement of the Master Transcription Factor NF-kappaB in Cancer Initiation and Progression. Biomedicines 2018, 6, 82. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, M.K.; Ahn, K.S.; Lee, J.H.; Kannaiyan, R.; Mustafa, N.; Manu, K.A.; Siveen, K.S.; Sethi, G.; Chng, W.J.; Kumar, A.P. Celastrol Attenuates the Invasion and Migration and Augments the Anticancer Effects of Bortezomib in a Xenograft Mouse Model of Multiple Myeloma. Front. Pharmacol. 2018, 9, 365. [Google Scholar] [CrossRef] [PubMed]
- Manu, K.A.; Shanmugam, M.K.; Ramachandran, L.; Li, F.; Siveen, K.S.; Chinnathambi, A.; Zayed, M.E.; Alharbi, S.A.; Arfuso, F.; Kumar, A.P.; et al. Isorhamnetin augments the anti-tumor effect of capeciatbine through the negative regulation of NF-kappaB signaling cascade in gastric cancer. Cancer Lett. 2015, 363, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Shanmugam, M.K.; Siveen, K.S.; Wang, F.; Ong, T.H.; Loo, S.Y.; Swamy, M.M.; Mandal, S.; Kumar, A.P.; Goh, B.C.; et al. Garcinol sensitizes human head and neck carcinoma to cisplatin in a xenograft mouse model despite downregulation of proliferative biomarkers. Oncotarget 2015, 6, 5147–5163. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, M.K.; Ahn, K.S.; Hsu, A.; Woo, C.C.; Yuan, Y.; Tan, K.H.B.; Chinnathambi, A.; Alahmadi, T.A.; Alharbi, S.A.; Koh, A.P.F.; et al. Thymoquinone Inhibits Bone Metastasis of Breast Cancer Cells Through Abrogation of the CXCR4 Signaling Axis. Front. Pharmacol. 2018, 9, 1294. [Google Scholar] [CrossRef] [PubMed]
- Mohan, C.D.; Bharathkumar, H.; Dukanya, D.; Rangappa, S.; Shanmugam, M.K.; Chinnathambi, A.; Alharbi, S.A.; Alahmadi, T.A.; Bhattacharjee, A.; Lobie, P.E.; et al. N-Substituted Pyrido-1,4-Oxazin-3-Ones Induce Apoptosis of Hepatocellular Carcinoma Cells by Targeting NF-kappaB Signaling Pathway. Front. Pharmacol. 2018, 9, 1125. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Ahn, K.S.; Shanmugam, M.K.; Wang, H.; Shen, H.; Arfuso, F.; Chinnathambi, A.; Alharbi, S.A.; Chang, Y.; Sethi, G.; et al. Oleuropein induces apoptosis via abrogating NF-kappaB activation cascade in estrogen receptor-negative breast cancer cells. J. Cell. Biochem. 2019, 120, 4504–4513. [Google Scholar] [CrossRef] [PubMed]
- Ningegowda, R.; Shivananju, N.S.; Rajendran, P.; Rangappa, K.S.; Chinnathambi, A.; Li, F.; Achar, R.R.; Shanmugam, M.K.; Bist, P.; Alharbi, S.A.; et al. A novel 4,6-disubstituted-1,2,4-triazolo-1,3,4-thiadiazole derivative inhibits tumor cell invasion and potentiates the apoptotic effect of TNFalpha by abrogating NF-kappaB activation cascade. Apoptosis 2017, 22, 145–157. [Google Scholar] [CrossRef] [PubMed]
- Siveen, K.S.; Mustafa, N.; Li, F.; Kannaiyan, R.; Ahn, K.S.; Kumar, A.P.; Chng, W.J.; Sethi, G. Thymoquinone overcomes chemoresistance and enhances the anticancer effects of bortezomib through abrogation of NF-kappaB regulated gene products in multiple myeloma xenograft mouse model. Oncotarget 2014, 5, 634–648. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.; Cho, S.K.; Kim, K.D.; Nam, D.; Chung, W.S.; Jang, H.J.; Lee, S.G.; Shim, B.S.; Sethi, G.; Ahn, K.S. beta-Caryophyllene oxide potentiates TNFalpha-induced apoptosis and inhibits invasion through down-modulation of NF-kappaB-regulated gene products. Apoptosis 2014, 19, 708–718. [Google Scholar] [CrossRef] [PubMed]
- Manu, K.A.; Shanmugam, M.K.; Li, F.; Chen, L.; Siveen, K.S.; Ahn, K.S.; Kumar, A.P.; Sethi, G. Simvastatin sensitizes human gastric cancer xenograft in nude mice to capecitabine by suppressing nuclear factor-kappa B-regulated gene products. J. Mol. Med. 2014, 92, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Shanmugam, M.K.; Chen, L.; Chatterjee, S.; Basha, J.; Kumar, A.P.; Kundu, T.K.; Sethi, G. Garcinol, a polyisoprenylated benzophenone modulates multiple proinflammatory signaling cascades leading to the suppression of growth and survival of head and neck carcinoma. Cancer Prev. Res. 2013, 6, 843–854. [Google Scholar] [CrossRef] [PubMed]
- Manu, K.A.; Shanmugam, M.K.; Ong, T.H.; Subramaniam, A.; Siveen, K.S.; Perumal, E.; Samy, R.P.; Bist, P.; Lim, L.H.; Kumar, A.P.; et al. Emodin suppresses migration and invasion through the modulation of CXCR4 expression in an orthotopic model of human hepatocellular carcinoma. PLoS ONE 2013, 8, e57015. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, M.K.; Ong, T.H.; Kumar, A.P.; Lun, C.K.; Ho, P.C.; Wong, P.T.; Hui, K.M.; Sethi, G. Ursolic acid inhibits the initiation, progression of prostate cancer and prolongs the survival of TRAMP mice by modulating pro-inflammatory pathways. PLoS ONE 2012, 7, e32476. [Google Scholar] [CrossRef] [PubMed]
- Manu, K.A.; Shanmugam, M.K.; Ramachandran, L.; Li, F.; Fong, C.W.; Kumar, A.P.; Tan, P.; Sethi, G. First evidence that gamma-tocotrienol inhibits the growth of human gastric cancer and chemosensitizes it to capecitabine in a xenograft mouse model through the modulation of NF-kappaB pathway. Clin. Cancer Res. 2012, 18, 2220–2229. [Google Scholar] [CrossRef] [PubMed]
- Manu, K.A.; Shanmugam, M.K.; Rajendran, P.; Li, F.; Ramachandran, L.; Hay, H.S.; Kannaiyan, R.; Swamy, S.N.; Vali, S.; Kapoor, S.; et al. Plumbagin inhibits invasion and migration of breast and gastric cancer cells by downregulating the expression of chemokine receptor CXCR4. Mol. Cancer 2011, 10, 107. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, M.K.; Rajendran, P.; Li, F.; Nema, T.; Vali, S.; Abbasi, T.; Kapoor, S.; Sharma, A.; Kumar, A.P.; Ho, P.C.; et al. Ursolic acid inhibits multiple cell survival pathways leading to suppression of growth of prostate cancer xenograft in nude mice. J. Mol. Med. 2011, 89, 713–727. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, M.K.; Manu, K.A.; Ong, T.H.; Ramachandran, L.; Surana, R.; Bist, P.; Lim, L.H.; Kumar, A.P.; Hui, K.M.; Sethi, G. Inhibition of CXCR4/CXCL12 signaling axis by ursolic acid leads to suppression of metastasis in transgenic adenocarcinoma of mouse prostate model. Int. J. Cancer 2011, 129, 1552–1563. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Tergaonkar, V.; Krishna, S.; Androphy, E.J. Human papillomavirus type 16 E6-enhanced susceptibility of L929 cells to tumor necrosis factor alpha correlates with increased accumulation of reactive oxygen species. J. Boil. Chem. 1999, 274, 24819–24827. [Google Scholar] [CrossRef]
- Akincilar, S.C.; Low, K.C.; Liu, C.Y.; Yan, T.D.; Oji, A.; Ikawa, M.; Li, S.; Tergaonkar, V. Quantitative assessment of telomerase components in cancer cell lines. FEBS Lett. 2015, 589, 974–984. [Google Scholar] [CrossRef] [PubMed]
- Khattar, E.; Kumar, P.; Liu, C.Y.; Akincilar, S.C.; Raju, A.; Lakshmanan, M.; Maury, J.J.; Qiang, Y.; Li, S.; Tan, E.Y.; et al. Telomerase reverse transcriptase promotes cancer cell proliferation by augmenting tRNA expression. J. Clin. Investig. 2016, 126, 4045–4060. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Cheng, H.S.; Chng, W.J.; Tergaonkar, V. Activation of mutant TERT promoter by RAS-ERK signaling is a key step in malignant progression of BRAF-mutant human melanomas. Proc. Natl. Acad. Sci. USA 2016, 113, 14402–14407. [Google Scholar] [CrossRef] [PubMed]
- Arabi, A.; Ullah, K.; Branca, R.M.; Johansson, J.; Bandarra, D.; Haneklaus, M.; Fu, J.; Aries, I.; Nilsson, P.; Den Boer, M.L.; et al. Proteomic screen reveals Fbw7 as a modulator of the NF-kappaB pathway. Nat. Commun. 2012, 3, 976. [Google Scholar] [CrossRef] [PubMed]
- Fukushima, H.; Matsumoto, A.; Inuzuka, H.; Zhai, B.; Lau, A.W.; Wan, L.; Gao, D.; Shaik, S.; Yuan, M.; Gygi, S.P.; et al. SCF(Fbw7) modulates the NFkB signaling pathway by targeting NFkB2 for ubiquitination and destruction. Cell Rep. 2012, 1, 434–443. [Google Scholar] [CrossRef] [PubMed]
- Kourtis, N.; Moubarak, R.S.; Aranda-Orgilles, B.; Lui, K.; Aydin, I.T.; Trimarchi, T.; Darvishian, F.; Salvaggio, C.; Zhong, J.; Bhatt, K.; et al. FBXW7 modulates cellular stress response and metastatic potential through HSF1 post-translational modification. Nat. Cell Biol. 2015, 17, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Hao, J.; Zheng, Y.; Jing, D.; Shen, Y.; Wang, J.; Zhao, Z. Role of Kruppel-like factors in cancer stem cells. J. Physiol. Biochem. 2015, 71, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Wang, Y.; Liu, N.; Ren, C.; Jiang, C.; Zhang, K.; Yu, S.; Chen, Y.; Tang, H.; Deng, Q.; et al. FBW7 regulates endothelial functions by targeting KLF2 for ubiquitination and degradation. Cell Res. 2013, 23, 803–819. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Jin, J.; Schlisio, S.; Harper, J.W.; Kaelin, W.G., Jr. The v-Jun point mutation allows c-Jun to escape GSK3-dependent recognition and destruction by the Fbw7 ubiquitin ligase. Cancer Cell 2005, 8, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Gotoh, M.; Nakamura, Y.; Arakawa, H. hCDC4b, a regulator of cyclin E, as a direct transcriptional target of p53. Cancer Sci. 2003, 94, 431–436. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; Ge, M.H.; Ling, Z.Q. Fbxw7 Tumor Suppressor: A Vital Regulator Contributes to Human Tumorigenesis. Medicine 2016, 95, e2496. [Google Scholar] [CrossRef] [PubMed]
- Balamurugan, K.; Sterneck, E. The many faces of C/EBPdelta and their relevance for inflammation and cancer. Int. J. Biol. Sci. 2013, 9, 917–933. [Google Scholar] [CrossRef] [PubMed]
- Gery, S.; Tanosaki, S.; Hofmann, W.K.; Koppel, A.; Koeffler, H.P. C/EBPdelta expression in a BCR-ABL-positive cell line induces growth arrest and myeloid differentiation. Oncogene 2005, 24, 1589–1597. [Google Scholar] [CrossRef] [PubMed]
- Welcker, M.; Clurman, B.E. FBW7 ubiquitin ligase: A tumour suppressor at the crossroads of cell division, growth and differentiation. Nat. Rev. Cancer 2008, 8, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Balamurugan, K.; Wang, J.M.; Tsai, H.H.; Sharan, S.; Anver, M.; Leighty, R.; Sterneck, E. The tumour suppressor C/EBPdelta inhibits FBXW7 expression and promotes mammary tumour metastasis. EMBO J. 2010, 29, 4106–4117. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.P.; Zhou, X.Z. The prolyl isomerase PIN1: A pivotal new twist in phosphorylation signalling and disease. Nat. Rev. Mol. Cell Biol. 2007, 8, 904–916. [Google Scholar] [CrossRef] [PubMed]
- Min, S.H.; Lau, A.W.; Lee, T.H.; Inuzuka, H.; Wei, S.; Huang, P.; Shaik, S.; Lee, D.Y.; Finn, G.; Balastik, M.; et al. Negative regulation of the stability and tumor suppressor function of Fbw7 by the Pin1 prolyl isomerase. Mol. Cell 2012, 46, 771–783. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Xing, H.; Kim, T.M.; Jung, Y.; Huang, W.; Yang, H.W.; Song, S.; Park, P.J.; Carroll, R.S.; Johnson, M.D. Numb regulates glioma stem cell fate and growth by altering epidermal growth factor receptor and Skp1-Cullin-F-box ubiquitin ligase activity. Stem Cells 2012, 30, 1313–1326. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, D.C.; Li, Z.F.; Liu, C.X.; Xiao, Y.M.; Zhang, B.; Li, X.D.; Zhao, J.; Chen, L.P.; Xing, X.M.; et al. Upregulation of miR-27a contributes to the malignant transformation of human bronchial epithelial cells induced by SV40 small T antigen. Oncogene 2011, 30, 3875–3886. [Google Scholar] [CrossRef] [PubMed]
- Lerner, M.; Lundgren, J.; Akhoondi, S.; Jahn, A.; Ng, H.F.; Akbari Moqadam, F.; Oude Vrielink, J.A.; Agami, R.; Den Boer, M.L.; Grander, D.; et al. MiRNA-27a controls FBW7/hCDC4-dependent cyclin E degradation and cell cycle progression. Cell Cycle 2011, 10, 2172–2183. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Guo, Y.; Liang, X.; Sun, M.; Wang, G.; De, W.; Wu, W. MicroRNA-223 functions as an oncogene in human gastric cancer by targeting FBXW7/hCdc4. J. Cancer Res. Clin. Oncol. 2012, 138, 763–774. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Palermo, R.; Talora, C.; Campese, A.F.; Checquolo, S.; Bellavia, D.; Tottone, L.; Testa, G.; Miele, E.; Indraccolo, S.; et al. Notch and NF-kB signaling pathways regulate miR-223/FBXW7 axis in T-cell acute lymphoblastic leukemia. Leukemia 2014, 28, 2324–2335. [Google Scholar] [CrossRef] [PubMed]
- Xia, W.; Zhou, J.; Luo, H.; Liu, Y.; Peng, C.; Zheng, W.; Ma, W. MicroRNA-32 promotes cell proliferation, migration and suppresses apoptosis in breast cancer cells by targeting FBXW7. Cancer Cell Int. 2017, 17, 14. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Shen, L.; Mao, L.; Wang, B.; Li, Y.; Yu, H. miR-92a is upregulated in cervical cancer and promotes cell proliferation and invasion by targeting FBXW7. Biochem. Biophys. Res. Commun. 2015, 458, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.; Liu, Y.H.; Wang, L.L.; Wang, J.; Zhao, Z.H.; Qu, J.F.; Wang, S.F. MiR-182 promotes cell proliferation by suppressing FBXW7 and FBXW11 in non-small cell lung cancer. Am. J. Transl. Res. 2018, 10, 1131–1142. [Google Scholar] [PubMed]
- Akhoondi, S.; Sun, D.; von der Lehr, N.; Apostolidou, S.; Klotz, K.; Maljukova, A.; Cepeda, D.; Fiegl, H.; Dafou, D.; Marth, C.; et al. FBXW7/hCDC4 is a general tumor suppressor in human cancer. Cancer Res. 2007, 67, 9006–9012. [Google Scholar] [CrossRef] [PubMed]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed]
- Mao, J.H.; Kim, I.J.; Wu, D.; Climent, J.; Kang, H.C.; DelRosario, R.; Balmain, A. FBXW7 targets mTOR for degradation and cooperates with PTEN in tumor suppression. Science 2008, 321, 1499–1502. [Google Scholar] [CrossRef] [PubMed]
- Davis, H.; Lewis, A.; Behrens, A.; Tomlinson, I. Investigation of the atypical FBXW7 mutation spectrum in human tumours by conditional expression of a heterozygous propellor tip missense allele in the mouse intestines. Gut 2014, 63, 792–799. [Google Scholar] [CrossRef] [PubMed]
- Sancho, R.; Jandke, A.; Davis, H.; Diefenbacher, M.E.; Tomlinson, I.; Behrens, A. F-box and WD repeat domain-containing 7 regulates intestinal cell lineage commitment and is a haploinsufficient tumor suppressor. Gastroenterology 2010, 139, 929–941. [Google Scholar] [CrossRef] [PubMed]
- King, B.; Trimarchi, T.; Reavie, L.; Xu, L.; Mullenders, J.; Ntziachristos, P.; Aranda-Orgilles, B.; Perez-Garcia, A.; Shi, J.; Vakoc, C.; et al. The ubiquitin ligase FBXW7 modulates leukemia-initiating cell activity by regulating MYC stability. Cell 2013, 153, 1552–1566. [Google Scholar] [CrossRef] [PubMed]
- Davis, H.; Lewis, A.; Spencer-Dene, B.; Tateossian, H.; Stamp, G.; Behrens, A.; Tomlinson, I. FBXW7 mutations typically found in human cancers are distinct from null alleles and disrupt lung development. J. Pathol. 2011, 224, 180–189. [Google Scholar] [CrossRef] [PubMed]
- Kitade, S.; Onoyama, I.; Kobayashi, H.; Yagi, H.; Yoshida, S.; Kato, M.; Tsunematsu, R.; Asanoma, K.; Sonoda, K.; Wake, N.; et al. FBXW7 is involved in the acquisition of the malignant phenotype in epithelial ovarian tumors. Cancer Sci. 2016, 107, 1399–1405. [Google Scholar] [CrossRef] [PubMed]
- Cremona, C.A.; Sancho, R.; Diefenbacher, M.E.; Behrens, A. Fbw7 and its counteracting forces in stem cells and cancer: Oncoproteins in the balance. Semin. Cancer Biol. 2016, 36, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Kunnumakkara, A.B.; Banik, K.; Bordoloi, D.; Harsha, C.; Sailo, B.L.; Padmavathi, G.; Roy, N.K.; Gupta, S.C.; Aggarwal, B.B. Googling the Guggul (Commiphora and Boswellia) for Prevention of Chronic Diseases. Front. Pharmacol. 2018, 9, 686. [Google Scholar] [CrossRef] [PubMed]
- Kunnumakkara, A.B.; Sailo, B.L.; Banik, K.; Harsha, C.; Prasad, S.; Gupta, S.C.; Bharti, A.C.; Aggarwal, B.B. Chronic diseases, inflammation, and spices: How are they linked? J. Transl. Med. 2018, 16, 14. [Google Scholar] [CrossRef] [PubMed]
- Bordoloi, D.; Banik, K.; Shabnam, B.; Padmavathi, G.; Monisha, J.; Arfuso, F.; Dharmarajan, A.; Mao, X.; Lim, L.H.K.; Wang, L.; et al. TIPE Family of Proteins and Its Implications in Different Chronic Diseases. Int. J. Mol. Sci. 2018, 19, 2974. [Google Scholar] [CrossRef] [PubMed]
- Kunnumakkara, A.B.; Bordoloi, D.; Harsha, C.; Banik, K.; Gupta, S.C.; Aggarwal, B.B. Curcumin mediates anticancer effects by modulating multiple cell signaling pathways. Clin. Sci. 2017, 131, 1781–1799. [Google Scholar] [CrossRef] [PubMed]
- Khwairakpam, A.D.; Bordoloi, D.; Thakur, K.K.; Monisha, J.; Arfuso, F.; Sethi, G.; Mishra, S.; Kumar, A.P.; Kunnumakkara, A.B. Possible use of Punica granatum (Pomegranate) in cancer therapy. Pharmacol. Res. 2018, 133, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, Y.; Lu, J.; Zhang, P.; Wang, Y.; Xu, Y.; Wang, Z.; Mao, J.H.; Wei, G. Rapamycin inhibits FBXW7 loss-induced epithelial-mesenchymal transition and cancer stem cell-like characteristics in colorectal cancer cells. Biochem. Biophys. Res. Commun. 2013, 434, 352–356. [Google Scholar] [CrossRef] [PubMed]
- Yumimoto, K.; Akiyoshi, S.; Ueo, H.; Sagara, Y.; Onoyama, I.; Ueo, H.; Ohno, S.; Mori, M.; Mimori, K.; Nakayama, K.I. F-box protein FBXW7 inhibits cancer metastasis in a non-cell-autonomous manner. J. Clin. Investig. 2015, 125, 621–635. [Google Scholar] [CrossRef] [PubMed]
- Teplyuk, N.M.; Uhlmann, E.J.; Wong, A.H.; Karmali, P.; Basu, M.; Gabriely, G.; Jain, A.; Wang, Y.; Chiocca, E.A.; Stephens, R.; et al. MicroRNA-10b inhibition reduces E2F1-mediated transcription and miR-15/16 activity in glioblastoma. Oncotarget 2015, 6, 3770–3783. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Woolard, K.; Lai, C.; Bauer, P.O.; Maric, D.; Song, H.; Li, A.; Kotliarova, S.; Zhang, W.; Fine, H.A. Gliomagenesis arising from Pten- and Ink4a/Arf-deficient neural progenitor cells is mediated by the p53-Fbxw7/Cdc4 pathway, which controls c-Myc. Cancer Res. 2012, 72, 6065–6075. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Mitsui, H.; Inomata, K.; Honda, M.; Endo, C.; Sakurada, A.; Sato, M.; Okada, Y.; Kondo, T.; Horii, A. The methylation status of FBXW7 beta-form correlates with histological subtype in human thymoma. Biochem. Biophys. Res. Commun. 2008, 377, 685–688. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Inomata, K.; Ishizawa, K.; Horii, A. The FBXW7 beta-form is suppressed in human glioma cells. Biochem. Biophys. Res. Commun. 2007, 354, 992–998. [Google Scholar] [CrossRef] [PubMed]
- Suryo Rahmanto, A.; Savov, V.; Brunner, A.; Bolin, S.; Weishaupt, H.; Malyukova, A.; Rosen, G.; Cancer, M.; Hutter, S.; Sundstrom, A.; et al. FBW7 suppression leads to SOX9 stabilization and increased malignancy in medulloblastoma. EMBO J. 2016, 35, 2192–2212. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Ji, A.; Qiu, G.; Feng, H.; Li, J.; Li, S.; Zou, Y.; Cui, Y.; Song, C.; He, H.; et al. FBW7 is associated with prognosis, inhibits malignancies and enhances temozolomide sensitivity in glioblastoma cells. Cancer Sci. 2018, 109, 1001–1011. [Google Scholar] [CrossRef] [PubMed]
- Cao, S.; Wang, Y.; Li, J.; Lv, M.; Niu, H.; Tian, Y. Tumor-suppressive function of long noncoding RNA MALAT1 in glioma cells by suppressing miR-155 expression and activating FBXW7 function. Am. J. Cancer Res. 2016, 6, 2561–2574. [Google Scholar] [PubMed]
- Yang, Y.; Gao, X.; Zhang, M.; Yan, S.; Sun, C.; Xiao, F.; Huang, N.; Yang, X.; Zhao, K.; Zhou, H.; et al. Novel Role of FBXW7 Circular RNA in Repressing Glioma Tumorigenesis. J. Natl. Cancer Inst. 2018, 110. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Zheng, H.Q.; Zhou, Z.; Chen, C. The Fbw7 tumor suppressor targets KLF5 for ubiquitin-mediated degradation and suppresses breast cell proliferation. Cancer Res. 2010, 70, 4728–4738. [Google Scholar] [CrossRef] [PubMed]
- Ibusuki, M.; Yamamoto, Y.; Shinriki, S.; Ando, Y.; Iwase, H. Reduced expression of ubiquitin ligase FBXW7 mRNA is associated with poor prognosis in breast cancer patients. Cancer Sci. 2011, 102, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Liu, Y.; Zhang, P.; Zhang, W.; Wang, W.; Curr, K.; Wei, G.; Mao, J.H. FAM83D promotes cell proliferation and motility by downregulating tumor suppressor gene FBXW7. Oncotarget 2013, 4, 2476–2486. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Li, X. The canonical wnt signal restricts the glycogen synthase kinase 3/fbw7-dependent ubiquitination and degradation of eya1 phosphatase. Mol. Cell. Biol. 2014, 34, 2409–2417. [Google Scholar] [CrossRef] [PubMed]
- Gasca, J.; Flores, M.L.; Giraldez, S.; Ruiz-Borrego, M.; Tortolero, M.; Romero, F.; Japon, M.A.; Saez, C. Loss of FBXW7 and accumulation of MCL1 and PLK1 promote paclitaxel resistance in breast cancer. Oncotarget 2016, 7, 52751–52765. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.M.; Liang, Z.R.; Zhou, K.R.; Zhou, H.; Qu, L.H. 27-Hydroxycholesterol increases Myc protein stability via suppressing PP2A, SCP1 and FBW7 transcription in MCF-7 breast cancer cells. Biochem. Biophys. Res. Commun. 2016, 480, 328–333. [Google Scholar] [CrossRef] [PubMed]
- Takada, M.; Zhuang, M.; Inuzuka, H.; Zhang, J.; Zurlo, G.; Zhang, J.; Zhang, Q. EglN2 contributes to triple negative breast tumorigenesis by functioning as a substrate for the FBW7 tumor suppressor. Oncotarget 2017, 8, 6787–6795. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, X.Y.; Long, M.; Wang, X.; Gao, Z.W.; Cui, Y.; Ren, J.; Zhang, Z.; Liu, C.; Dong, K.; et al. The FBXW7 tumor suppressor inhibits breast cancer proliferation and promotes apoptosis by targeting MTDH for degradation. Neoplasma 2018, 65, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Lu, X.; Liu, Z.; Chen, L.; Xu, Y.; Wang, Y.; Wei, G.; Chen, Y. FBXW7 suppresses epithelial-mesenchymal transition, stemness and metastatic potential of cholangiocarcinoma cells. Oncotarget 2015, 6, 6310–6325. [Google Scholar] [CrossRef] [PubMed]
- Ishii, N.; Araki, K.; Yokobori, T.; Watanabe, A.; Tsukagoshi, M.; Kubo, N.; Suzuki, H.; Saito, F.; Altan, B.; Hosouchi, Y.; et al. Poor prognosis in cholangiocarcinoma patients with low FBXW7 expression is improved by chemotherapy. Oncol. Lett. 2017, 13, 3653–3661. [Google Scholar] [CrossRef] [PubMed]
- Iwatsuki, M.; Mimori, K.; Ishii, H.; Yokobori, T.; Takatsuno, Y.; Sato, T.; Toh, H.; Onoyama, I.; Nakayama, K.I.; Baba, H.; et al. Loss of FBXW7, a cell cycle regulating gene, in colorectal cancer: Clinical significance. Int. J. Cancer 2010, 126, 1828–1837. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Zhou, Y.; Evers, B.M.; Wang, Q. Rictor regulates FBXW7-dependent c-Myc and cyclin E degradation in colorectal cancer cells. Biochem. Biophys. Res. Commun. 2012, 418, 426–432. [Google Scholar] [CrossRef] [PubMed]
- Grim, J.E.; Knoblaugh, S.E.; Guthrie, K.A.; Hagar, A.; Swanger, J.; Hespelt, J.; Delrow, J.J.; Small, T.; Grady, W.M.; Nakayama, K.I.; et al. Fbw7 and p53 cooperatively suppress advanced and chromosomally unstable intestinal cancer. Mol. Cell. Biol. 2012, 32, 2160–2167. [Google Scholar] [CrossRef] [PubMed]
- Zhan, P.; Wang, Y.; Zhao, S.; Liu, C.; Wang, Y.; Wen, M.; Mao, J.H.; Wei, G.; Zhang, P. FBXW7 negatively regulates ENO1 expression and function in colorectal cancer. Lab. Investig. 2015, 95, 995–1004. [Google Scholar] [CrossRef] [PubMed]
- Ou, B.; Zhao, J.; Guan, S.; Wangpu, X.; Zhu, C.; Zong, Y.; Ma, J.; Sun, J.; Zheng, M.; Feng, H.; et al. Plk2 promotes tumor growth and inhibits apoptosis by targeting Fbxw7/Cyclin E in colorectal cancer. Cancer Lett. 2016, 380, 457–466. [Google Scholar] [CrossRef] [PubMed]
- Tong, J.; Tan, S.; Nikolovska-Coleska, Z.; Yu, J.; Zou, F.; Zhang, L. FBW7-Dependent Mcl-1 Degradation Mediates the Anticancer Effect of Hsp90 Inhibitors. Mol. Cancer Ther. 2017, 16, 1979–1988. [Google Scholar] [CrossRef] [PubMed]
- Mu, Y.; Zou, H.; Chen, B.; Fan, Y.; Luo, S. FAM83D knockdown regulates proliferation, migration and invasion of colorectal cancer through inhibiting FBXW7/Notch-1 signalling pathway. Biomed. Pharmacother. 2017, 90, 548–554. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Zhao, Z.; Song, J.; Luo, B.; Huang, L. MiR-223 promotes the doxorubicin resistance of colorectal cancer cells via regulating epithelial-mesenchymal transition by targeting FBXW7. Acta Biochim. Biophys. Sin. 2018, 50, 597–604. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.Z.; Wang, K.P.; Song, H.J.; Xia, J.H.; Jiang, Y.; Wang, Y.L. MiR-27a-3p promotes esophageal cancer cell proliferation via F-box and WD repeat domain-containing 7 (FBXW7) suppression. Int. J. Clin. Exp. Med. 2015, 8, 15556–15562. [Google Scholar] [PubMed]
- Calcagno, D.Q.; Freitas, V.M.; Leal, M.F.; de Souza, C.R.; Demachki, S.; Montenegro, R.; Assumpcao, P.P.; Khayat, A.S.; Smith Mde, A.; dos Santos, A.K.; et al. MYC, FBXW7 and TP53 copy number variation and expression in gastric cancer. BMC Gastroenterol. 2013, 13, 141. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Cui, Z.; Li, L.; Ma, Q.; Wang, Q.; Gao, Y.; Sun, H. MicroRNA-25 promotes gastric cancer proliferation, invasion, and migration by directly targeting F-box and WD-40 Domain Protein 7, FBXW7. Tumour Biol. 2015, 36, 7831–7840. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, Z.; Zhang, W.; Qian, K.; Xu, W.; Zhang, S. Fbxw7 regulates tumor apoptosis, growth arrest and the epithelial-to-mesenchymal transition in part through the RhoA signaling pathway in gastric cancer. Cancer Lett. 2016, 370, 39–55. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Qi, X.; Liu, X.; Zhang, J.; Ji, J.; Zhu, Z.; Ren, J.; Yu, Y. Fbxw7 haploinsufficiency loses its protection against DNA damage and accelerates MNU-induced gastric carcinogenesis. Oncotarget 2017, 8, 33444–33456. [Google Scholar] [CrossRef] [PubMed]
- Li, M.R.; Zhu, C.C.; Ling, T.L.; Zhang, Y.Q.; Xu, J.; Zhao, E.H.; Zhao, G. FBXW7 expression is associated with prognosis and chemotherapeutic outcome in Chinese patients with gastric adenocarcinoma. BMC Gastroenterol. 2017, 17, 60. [Google Scholar] [CrossRef] [PubMed]
- O’Neil, J.; Grim, J.; Strack, P.; Rao, S.; Tibbitts, D.; Winter, C.; Hardwick, J.; Welcker, M.; Meijerink, J.P.; Pieters, R.; et al. FBW7 mutations in leukemic cells mediate NOTCH pathway activation and resistance to gamma-secretase inhibitors. J. Exp. Med. 2007, 204, 1813–1824. [Google Scholar] [CrossRef] [PubMed]
- Asnafi, V.; Buzyn, A.; Le Noir, S.; Baleydier, F.; Simon, A.; Beldjord, K.; Reman, O.; Witz, F.; Fagot, T.; Tavernier, E.; et al. NOTCH1/FBXW7 mutation identifies a large subgroup with favorable outcome in adult T-cell acute lymphoblastic leukemia (T-ALL): A Group for Research on Adult Acute Lymphoblastic Leukemia (GRAALL) study. Blood 2009, 113, 3918–3924. [Google Scholar] [CrossRef] [PubMed]
- Inuzuka, H.; Shaik, S.; Onoyama, I.; Gao, D.; Tseng, A.; Maser, R.S.; Zhai, B.; Wan, L.; Gutierrez, A.; Lau, A.W.; et al. SCF(FBW7) regulates cellular apoptosis by targeting MCL1 for ubiquitylation and destruction. Nature 2011, 471, 104–109. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.L.; Weng, H.Y.; Wang, L.Q.; Yu, C.H.; Huang, Q.J.; Zhao, P.P.; Wen, J.Z.; Zhou, H.; Qu, L.H. Triggering Fbw7-mediated proteasomal degradation of c-Myc by oridonin induces cell growth inhibition and apoptosis. Mol. Cancer Ther. 2012, 11, 1155–1165. [Google Scholar] [CrossRef] [PubMed]
- Malyukova, A.; Brown, S.; Papa, R.; O’Brien, R.; Giles, J.; Trahair, T.N.; Dalla Pozza, L.; Sutton, R.; Liu, T.; Haber, M.; et al. FBXW7 regulates glucocorticoid response in T-cell acute lymphoblastic leukaemia by targeting the glucocorticoid receptor for degradation. Leukemia 2013, 27, 1053–1062. [Google Scholar] [CrossRef] [PubMed]
- Mansour, M.R.; Sanda, T.; Lawton, L.N.; Li, X.; Kreslavsky, T.; Novina, C.D.; Brand, M.; Gutierrez, A.; Kelliher, M.A.; Jamieson, C.H.; et al. The TAL1 complex targets the FBXW7 tumor suppressor by activating miR-223 in human T cell acute lymphoblastic leukemia. J. Exp. Med. 2013, 210, 1545–1557. [Google Scholar] [CrossRef] [PubMed]
- Yeh, C.H.; Bellon, M.; Pancewicz-Wojtkiewicz, J.; Nicot, C. Oncogenic mutations in the FBXW7 gene of adult T-cell leukemia patients. Proc. Natl. Acad. Sci. USA 2016, 113, 6731–6736. [Google Scholar] [CrossRef] [PubMed]
- Yao, S.; Xu, F.; Chen, Y.; Ge, Y.; Zhang, F.; Huang, H.; Li, L.; Lin, D.; Luo, X.; Xu, J.; et al. Fbw7 regulates apoptosis in activated B-cell like diffuse large B-cell lymphoma by targeting Stat3 for ubiquitylation and degradation. J. Exp. Clin. Cancer Res. 2017, 36, 10. [Google Scholar] [CrossRef] [PubMed]
- Valliyammai, N.; Nancy, N.K.; Sagar, T.G.; Rajkumar, T. Study of NOTCH1 and FBXW7 Mutations and Its Prognostic Significance in South Indian T-Cell Acute Lymphoblastic Leukemia. J. Pediatr. Hematol. Oncol. 2018, 40, e1–e8. [Google Scholar] [CrossRef] [PubMed]
- Tu, K.; Zheng, X.; Zhou, Z.; Li, C.; Zhang, J.; Gao, J.; Yao, Y.; Liu, Q. Recombinant human adenovirus-p53 injection induced apoptosis in hepatocellular carcinoma cell lines mediated by p53-Fbxw7 pathway, which controls c-Myc and cyclin E. PLoS ONE 2013, 8, e68574. [Google Scholar] [CrossRef] [PubMed]
- Tu, K.; Yang, W.; Li, C.; Zheng, X.; Lu, Z.; Guo, C.; Yao, Y.; Liu, Q. Fbxw7 is an independent prognostic marker and induces apoptosis and growth arrest by regulating YAP abundance in hepatocellular carcinoma. Mol. Cancer 2014, 13, 110. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Wang, H.; Wang, J.; Huang, S.; Zhang, W. STAT1 inhibits human hepatocellular carcinoma cell growth through induction of p53 and Fbxw7. Cancer Cell Int. 2015, 15, 111. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.; Koo, J.; Guan, B.; Yue, P.; Deng, X.; Chen, M.; Khuri, F.R.; Sun, S.Y. The E3 ubiquitin ligases beta-TrCP and FBXW7 cooperatively mediates GSK3-dependent Mcl-1 degradation induced by the Akt inhibitor API-1, resulting in apoptosis. Mol. Cancer 2013, 12, 146. [Google Scholar] [CrossRef] [PubMed]
- Yokobori, T.; Yokoyama, Y.; Mogi, A.; Endoh, H.; Altan, B.; Kosaka, T.; Yamaki, E.; Yajima, T.; Tomizawa, K.; Azuma, Y.; et al. FBXW7 mediates chemotherapeutic sensitivity and prognosis in NSCLCs. Mol. Cancer Res. 2014, 12, 32–37. [Google Scholar] [CrossRef] [PubMed]
- Liao, S.Y.; Chiang, C.W.; Hsu, C.H.; Chen, Y.T.; Jen, J.; Juan, H.F.; Lai, W.W.; Wang, Y.C. CK1delta/GSK3beta/FBXW7alpha axis promotes degradation of the ZNF322A oncoprotein to suppress lung cancer progression. Oncogene 2017, 36, 5722–5733. [Google Scholar] [CrossRef] [PubMed]
- Xiao, G.; Zhang, B.; Meng, J.; Wang, J.; Xu, C.; Tang, S.C.; Li, X.; Zhang, J.; Liang, R.; Ren, H.; et al. miR-367 stimulates Wnt cascade activation through degrading FBXW7 in NSCLC stem cells. Cell Cycle 2017, 16, 2374–2385. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.; Zhang, Y.; Zhang, X.; Zhang, J.; Jing, P.; Cao, L.; Li, N.; Li, X.; Yao, L.; Zhang, J.; et al. Targeting FBW7 as a Strategy to Overcome Resistance to Targeted Therapy in Non-Small Cell Lung Cancer. Cancer Res. 2017, 77, 3527–3539. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Ma, J.; Xu, F.; Li, L. TINCR suppresses proliferation and invasion through regulating miR-544a/FBXW7 axis in lung cancer. Biomed. Pharmacother. 2018, 99, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Yin, C.; Wang, Y.; Lv, H.; Wang, W.; Huang, Y.; Perez-Losada, J.; Snijders, A.M.; Mao, J.H.; Zhang, P. FBXW7 deletion contributes to lung tumor development and confers resistance to gefitinib therapy. Mol. Oncol. 2018, 12, 883–895. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Hu, C.; Chi, J.; Li, J.; Peng, C.; Yun, X.; Li, D.; Yu, Y.; Li, Y.; Gao, M.; et al. miR-24 promotes the proliferation, migration and invasion in human tongue squamous cell carcinoma by targeting FBXW7. Oncol. Rep. 2016, 36, 1143–1149. [Google Scholar] [CrossRef] [PubMed]
- Arita, H.; Nagata, M.; Yoshida, R.; Matsuoka, Y.; Hirosue, A.; Kawahara, K.; Sakata, J.; Nakashima, H.; Kojima, T.; Toya, R.; et al. FBXW7 expression affects the response to chemoradiotherapy and overall survival among patients with oral squamous cell carcinoma: A single-center retrospective study. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Azmi, A.S.; Aboukameel, A.; Kauffman, M.; Shacham, S.; Abou-Samra, A.B.; Mohammad, R.M. Nuclear retention of Fbw7 by specific inhibitors of nuclear export leads to Notch1 degradation in pancreatic cancer. Oncotarget 2014, 5, 3444–3454. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.X.; Sun, C.Y.; Tian, S.; Yu, C.; Chen, M.Y.; Zhang, H. Tumor suppressor Fbxw7 antagonizes WNT signaling by targeting beta-catenin for degradation in pancreatic cancer. Tumour Biol. 2016, 37, 13893–13902. [Google Scholar] [CrossRef] [PubMed]
- Ishii, N.; Araki, K.; Yokobori, T.; Gantumur, D.; Yamanaka, T.; Altan, B.; Tsukagoshi, M.; Igarashi, T.; Watanabe, A.; Kubo, N.; et al. Reduced FBXW7 expression in pancreatic cancer correlates with poor prognosis and chemotherapeutic resistance via accumulation of MCL1. Oncotarget 2017, 8, 112636–112646. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.; Qin, Y.; Zhang, B.; Liang, C.; Ji, S.; Shi, S.; Xu, W.; Xiang, J.; Liang, D.; Ni, Q.; et al. FBW7 increases the chemosensitivity of pancreatic cancer cells to gemcitabine through upregulation of ENT1. Oncol. Rep. 2017, 38, 2069–2077. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Lin, Y.; Yang, Z.; Yang, G.; Li, G.; Liu, Y.; Tan, X.; Huang, Y.; Wu, X.; Wang, Y.; et al. FBXW7 overexpression suppresses renal cancer cell proliferation and induces apoptosis. Med. Oncol. 2015, 32, 215. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Zhang, M.; Qiu, X.; Wang, B.; Fu, Y.; Zeng, J.; Bai, J.; Yang, G. Upregulation of FBXW7 Suppresses Renal Cancer Metastasis and Epithelial Mesenchymal Transition. Dis. Markers 2017, 2017, 8276939. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, Y.; Hosogane, M.; Okuyama, R.; Aoyama, S.; Onoyama, I.; Nakayama, K.I.; Nakayama, K. Opposing functions of Fbxw7 in keratinocyte growth, differentiation and skin tumorigenesis mediated through negative regulation of c-Myc and Notch. Oncogene 2013, 32, 1921–1932. [Google Scholar] [CrossRef] [PubMed]
- Aydin, I.T.; Melamed, R.D.; Adams, S.J.; Castillo-Martin, M.; Demir, A.; Bryk, D.; Brunner, G.; Cordon-Cardo, C.; Osman, I.; Rabadan, R.; et al. FBXW7 mutations in melanoma and a new therapeutic paradigm. J. Natl. Cancer Inst. 2014, 106, dju107. [Google Scholar] [CrossRef] [PubMed]
- Wei, Z.; Lihua, L.; Qingqing, W. Effects of myeloid specific deficiency of FBXW7 on lung metastasis of murine melanoma. Zhejiang Da Xue Xue Bao Yi Xue Ban 2017, 46, 111–117. [Google Scholar] [PubMed]
- Abbate, F.; Badal, B.; Mendelson, K.; Aydin, I.T.; Serasinghe, M.N.; Iqbal, R.; Mohammed, J.N.; Solovyov, A.; Greenbaum, B.D.; Chipuk, J.E.; et al. FBXW7 regulates a mitochondrial transcription program by modulating MITF. Pigment Cell Melanoma Res. 2018, 31, 636–640. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Wang, Y.; Zhang, P.; Lu, J.; Mao, J.H. Evaluating the prognostic significance of FBXW7 expression level in human breast cancer by a meta-analysis of transcriptional profiles. J. Cancer Sci. Ther. 2012, 4, 299–305. [Google Scholar] [CrossRef] [PubMed]
- Strohmaier, H.; Spruck, C.H.; Kaiser, P.; Won, K.A.; Sangfelt, O.; Reed, S.I. Human F-box protein hCdc4 targets cyclin E for proteolysis and is mutated in a breast cancer cell line. Nature 2001, 413, 316–322. [Google Scholar] [CrossRef] [PubMed]
- Ong, P.S.; Wang, L.Z.; Dai, X.; Tseng, S.H.; Loo, S.J.; Sethi, G. Judicious Toggling of mTOR Activity to Combat Insulin Resistance and Cancer: Current Evidence and Perspectives. Front. Pharmacol. 2016, 7, 395. [Google Scholar] [CrossRef] [PubMed]
- Baek, S.H.; Ko, J.H.; Lee, J.H.; Kim, C.; Lee, H.; Nam, D.; Lee, J.; Lee, S.G.; Yang, W.M.; Um, J.Y.; et al. Ginkgolic Acid Inhibits Invasion and Migration and TGF-beta-Induced EMT of Lung Cancer Cells Through PI3K/Akt/mTOR Inactivation. J. Cell. Physiol. 2017, 232, 346–354. [Google Scholar] [CrossRef] [PubMed]
- Mohan, C.D.; Srinivasa, V.; Rangappa, S.; Mervin, L.; Mohan, S.; Paricharak, S.; Baday, S.; Li, F.; Shanmugam, M.K.; Chinnathambi, A.; et al. Trisubstituted-Imidazoles Induce Apoptosis in Human Breast Cancer Cells by Targeting the Oncogenic PI3K/Akt/mTOR Signaling Pathway. PLoS ONE 2016, 11, e0153155. [Google Scholar] [CrossRef] [PubMed]
- Ko, J.H.; Nam, D.; Um, J.Y.; Jung, S.H.; Sethi, G.; Ahn, K.S. Bergamottin Suppresses Metastasis of Lung Cancer Cells through Abrogation of Diverse Oncogenic Signaling Cascades and Epithelial-to-Mesenchymal Transition. Molecules 2018, 23, 1601. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Baek, S.H.; Lee, J.H.; Kim, C.; Ko, J.H.; Lee, S.G.; Chinnathambi, A.; Alharbi, S.A.; Yang, W.M.; Um, J.Y.; et al. Isorhynchophylline, a Potent Plant Alkaloid, Induces Apoptotic and Anti-Metastatic Effects in Human Hepatocellular Carcinoma Cells through the Modulation of Diverse Cell Signaling Cascades. Int. J. Mol. Sci. 2017, 18, 1095. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.S.; Yap, W.N.; Arfuso, F.; Kar, S.; Wang, C.; Cai, W.; Dharmarajan, A.M.; Sethi, G.; Kumar, A.P. Targeting the PI3K/Akt signaling pathway in gastric carcinoma: A reality for personalized medicine? World J. Gastroenterol. 2015, 21, 12261–12273. [Google Scholar] [CrossRef] [PubMed]
- Siveen, K.S.; Ahn, K.S.; Ong, T.H.; Shanmugam, M.K.; Li, F.; Yap, W.N.; Kumar, A.P.; Fong, C.W.; Tergaonkar, V.; Hui, K.M.; et al. Y-tocotrienol inhibits angiogenesis-dependent growth of human hepatocellular carcinoma through abrogation of AKT/mTOR pathway in an orthotopic mouse model. Oncotarget 2014, 5, 1897–1911. [Google Scholar] [CrossRef] [PubMed]
- Kannaiyan, R.; Manu, K.A.; Chen, L.; Li, F.; Rajendran, P.; Subramaniam, A.; Lam, P.; Kumar, A.P.; Sethi, G. Celastrol inhibits tumor cell proliferation and promotes apoptosis through the activation of c-Jun N-terminal kinase and suppression of PI3 K/Akt signaling pathways. Apoptosis 2011, 16, 1028–1041. [Google Scholar] [CrossRef] [PubMed]
- Park, K.R.; Nam, D.; Yun, H.M.; Lee, S.G.; Jang, H.J.; Sethi, G.; Cho, S.K.; Ahn, K.S. beta-Caryophyllene oxide inhibits growth and induces apoptosis through the suppression of PI3K/AKT/mTOR/S6K1 pathways and ROS-mediated MAPKs activation. Cancer Lett. 2011, 312, 178–188. [Google Scholar] [CrossRef] [PubMed]
- Sethi, G.; Ahn, K.S.; Sung, B.; Kunnumakkara, A.B.; Chaturvedi, M.M.; Aggarwal, B.B. SH-5, an AKT inhibitor potentiates apoptosis and inhibits invasion through the suppression of anti-apoptotic, proliferative and metastatic gene products regulated by IkappaBalpha kinase activation. Biochem. Pharmacol. 2008, 76, 1404–1416. [Google Scholar] [CrossRef] [PubMed]
- Kogita, A.; Yoshioka, Y.; Sakai, K.; Togashi, Y.; Sogabe, S.; Nakai, T.; Okuno, K.; Nishio, K. Inter- and intra-tumor profiling of multi-regional colon cancer and metastasis. Biochem. Biophys. Res. Commun. 2015, 458, 52–56. [Google Scholar] [CrossRef] [PubMed]
- Korphaisarn, K.; Morris, V.K.; Overman, M.J.; Fogelman, D.R.; Kee, B.K.; Raghav, K.P.S.; Manuel, S.; Shureiqi, I.; Wolff, R.A.; Eng, C.; et al. FBXW7 missense mutation: A novel negative prognostic factor in metastatic colorectal adenocarcinoma. Oncotarget 2017, 8, 39268–39279. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Lorenzi, F.; Kalakouti, E.; Normatova, M.; Babaei-Jadidi, R.; Tomlinson, I.; Nateri, A.S. FBXW7-mutated colorectal cancer cells exhibit aberrant expression of phosphorylated-p53 at Serine-15. Oncotarget 2015, 6, 9240–9256. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Sarver, A.L.; Khatri, R.; Hajeri, P.B.; Kamenev, I.; French, A.J.; Thibodeau, S.N.; Steer, C.J.; Subramanian, S. Sequential expression of miR-182 and miR-503 cooperatively targets FBXW7, contributing to the malignant transformation of colon adenoma to adenocarcinoma. J. Pathol. 2014, 234, 488–501. [Google Scholar] [CrossRef] [PubMed]
- Gong, L.; Ren, M.; Lv, Z.; Yang, Y.; Wang, Z. miR-92b-3p Promotes Colorectal Carcinoma Cell Proliferation, Invasion, and Migration by Inhibiting FBXW7 In vitro and In vivo. DNA Cell Biol. 2018, 37, 501–511. [Google Scholar] [CrossRef] [PubMed]
- Yokobori, T.; Mimori, K.; Iwatsuki, M.; Ishii, H.; Tanaka, F.; Sato, T.; Toh, H.; Sudo, T.; Iwaya, T.; Tanaka, Y.; et al. Copy number loss of FBXW7 is related to gene expression and poor prognosis in esophageal squamous cell carcinoma. Int. J. Oncol. 2012, 41, 253–259. [Google Scholar] [CrossRef] [PubMed]
- Naganawa, Y.; Ishiguro, H.; Kuwabara, Y.; Kimura, M.; Mitsui, A.; Katada, T.; Tanaka, T.; Shiozaki, M.; Fujii, Y.; Takeyama, H. Decreased expression of FBXW7 is correlated with poor prognosis in patients with esophageal squamous cell carcinoma. Exp. Ther. Med. 2010, 1, 841–846. [Google Scholar] [CrossRef] [PubMed]
- Kurashige, J.; Watanabe, M.; Iwatsuki, M.; Kinoshita, K.; Saito, S.; Hiyoshi, Y.; Kamohara, H.; Baba, Y.; Mimori, K.; Baba, H. Overexpression of microRNA-223 regulates the ubiquitin ligase FBXW7 in oesophageal squamous cell carcinoma. Br. J. Cancer 2012, 106, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Soung, Y.H.; Kim, H.J.; Park, W.S.; Nam, S.W.; Kim, S.H.; Lee, J.Y.; Yoo, N.J.; Lee, S.H. Mutational analysis of the hCDC4 gene in gastric carcinomas. Eur. J. Cancer 2006, 42, 2369–2373. [Google Scholar] [CrossRef] [PubMed]
- Milne, A.N.; Leguit, R.; Corver, W.E.; Morsink, F.H.; Polak, M.; de Leng, W.W.; Carvalho, R.; Offerhaus, G.J. Loss of CDC4/FBXW7 in gastric carcinoma. Cell Oncol. 2010, 32, 347–359. [Google Scholar] [CrossRef] [PubMed]
- Eto, K.; Iwatsuki, M.; Watanabe, M.; Ishimoto, T.; Ida, S.; Imamura, Y.; Iwagami, S.; Baba, Y.; Sakamoto, Y.; Miyamoto, Y.; et al. The sensitivity of gastric cancer to trastuzumab is regulated by the miR-223/FBXW7 pathway. Int. J. Cancer 2015, 136, 1537–1545. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Shen, Y.; Wang, S.H.; Xiang, Z.W.; Xie, Y.S.; Jiang, X. Effects of UO-126 on proliferation and fbw7 expression of HeLa cells. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 2010, 26, 138–140. [Google Scholar] [PubMed]
- Zhou, Z.Y.; Tu, K.S.; Zhang, J.; Zheng, X.; Gao, J.; Yao, Y.M.; Liu, Q.G. Expression of Fbxw7 and its correlation with cell proliferation in human hepatocellular carcinoma. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi 2012, 28, 1303–1306. [Google Scholar] [PubMed]
- Song, J.H.; Schnittke, N.; Zaat, A.; Walsh, C.S.; Miller, C.W. FBXW7 mutation in adult T-cell and B-cell acute lymphocytic leukemias. Leuk. Res. 2008, 32, 1751–1755. [Google Scholar] [CrossRef] [PubMed]
- Kraszewska, M.D.; Dawidowska, M.; Kosmalska, M.; Sedek, L.; Grzeszczak, W.; Kowalczyk, J.R.; Szczepanski, T.; Witt, M. BCL11B, FLT3, NOTCH1 and FBXW7 mutation status in T-cell acute lymphoblastic leukemia patients. Blood Cells Mol. Dis. 2013, 50, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Thompson, B.J.; Buonamici, S.; Sulis, M.L.; Palomero, T.; Vilimas, T.; Basso, G.; Ferrando, A.; Aifantis, I. The SCFFBW7 ubiquitin ligase complex as a tumor suppressor in T cell leukemia. J. Exp. Med. 2007, 204, 1825–1835. [Google Scholar] [CrossRef] [PubMed]
- Baldus, C.D.; Thibaut, J.; Goekbuget, N.; Stroux, A.; Schlee, C.; Mossner, M.; Burmeister, T.; Schwartz, S.; Bloomfield, C.D.; Hoelzer, D.; et al. Prognostic implications of NOTCH1 and FBXW7 mutations in adult acute T-lymphoblastic leukemia. Haematologica 2009, 94, 1383–1390. [Google Scholar] [CrossRef] [PubMed]
- Mihashi, Y.; Mizoguchi, M.; Takamatsu, Y.; Ishitsuka, K.; Iwasaki, H.; Koga, M.; Urabe, K.; Momosaki, S.; Sakata, T.; Kiyomi, F.; et al. C-MYC and Its Main Ubiquitin Ligase, FBXW7, Influence Cell Proliferation and Prognosis in Adult T-cell Leukemia/Lymphoma. Am. J. Surg. Pathol. 2017, 41, 1139–1149. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef] [PubMed]
- Villaruz, L.C.; Socinski, M.A. Temsirolimus therapy in a patient with lung adenocarcinoma harboring an FBXW7 mutation. Lung Cancer 2014, 83, 300–301. [Google Scholar] [CrossRef] [PubMed]
- Calhoun, E.S.; Jones, J.B.; Ashfaq, R.; Adsay, V.; Baker, S.J.; Valentine, V.; Hempen, P.M.; Hilgers, W.; Yeo, C.J.; Hruban, R.H.; et al. BRAF and FBXW7 (CDC4, FBW7, AGO, SEL10) mutations in distinct subsets of pancreatic cancer: Potential therapeutic targets. Am. J. Pathol. 2003, 163, 1255–1260. [Google Scholar] [CrossRef]
- Koh, M.S.; Ittmann, M.; Kadmon, D.; Thompson, T.C.; Leach, F.S. CDC4 gene expression as potential biomarker for targeted therapy in prostate cancer. Cancer Biol. Ther. 2006, 5, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Sun, Y.; Chen, X.; Squires, J.; Nowroozizadeh, B.; Liang, C.; Huang, J. p53 Mutation Directs AURKA Overexpression via miR-25 and FBXW7 in Prostatic Small Cell Neuroendocrine Carcinoma. Mol. Cancer Res. 2015, 13, 584–591. [Google Scholar] [CrossRef] [PubMed]
- Williams, R.D.; Al-Saadi, R.; Chagtai, T.; Popov, S.; Messahel, B.; Sebire, N.; Gessler, M.; Wegert, J.; Graf, N.; Leuschner, I.; et al. Subtype-specific FBXW7 mutation and MYCN copy number gain in Wilms’ tumor. Clin. Cancer Res. 2010, 16, 2036–2045. [Google Scholar] [CrossRef] [PubMed]
- Enkhbold, C.; Utsunomiya, T.; Morine, Y.; Imura, S.; Ikemoto, T.; Arakawa, Y.; Kanamoto, M.; Iwahashi, S.; Saito, Y.; Ishikawa, D.; et al. Loss of FBXW7 expression is associated with poor prognosis in intrahepatic cholangiocarcinoma. Hepatol. Res. 2014, 44, E346–E352. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Yu, J.; Liu, T.; Meng, F.; Kong, D.; Lou, G. Loss of FBXW7 is related to the susceptibility and poor prognosis of cervical squamous carcinoma. Biomarkers 2016, 21, 379–385. [Google Scholar] [CrossRef] [PubMed]
- Takeishi, S.; Matsumoto, A.; Onoyama, I.; Naka, K.; Hirao, A.; Nakayama, K.I. Ablation of Fbxw7 eliminates leukemia-initiating cells by preventing quiescence. Cancer Cell 2013, 23, 347–361. [Google Scholar] [CrossRef] [PubMed]
- Izumi, D.; Ishimoto, T.; Miyake, K.; Eto, T.; Arima, K.; Kiyozumi, Y.; Uchihara, T.; Kurashige, J.; Iwatsuki, M.; Baba, Y.; et al. Colorectal Cancer Stem Cells Acquire Chemoresistance Through the Upregulation of F-Box/WD Repeat-Containing Protein 7 and the Consequent Degradation of c-Myc. Stem Cells 2017, 35, 2027–2036. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Zhang, W.; Gao, F.; Liu, Y.X.; Chen, Z.Y.; Cheng, L.Y.; Xie, S.F.; Zheng, S.S. FBW7 increases chemosensitivity in hepatocellular carcinoma cells through suppression of epithelial-mesenchymal transition. Hepatobiliary Pancreat. Dis. Int. 2014, 13, 184–191. [Google Scholar] [CrossRef]
- Yu, H.G.; Wei, W.; Xia, L.H.; Han, W.L.; Zhao, P.; Wu, S.J.; Li, W.D.; Chen, W. FBW7 upregulation enhances cisplatin cytotoxicity in non-small cell lung cancer cells. Asian Pac. J. Cancer Prev. 2013, 14, 6321–6326. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Zhou, X.; Bai, W.; Ma, X. FBW7 increases drug sensitivity to cisplatin in human nasopharyngeal carcinoma by downregulating the expression of multidrug resistance-associated protein. Tumour Biol. 2015, 36, 4197–4202. [Google Scholar] [CrossRef] [PubMed]
- Yeh, C.H.; Bellon, M.; Nicot, C. FBXW7: A critical tumor suppressor of human cancers. Mol. Cancer 2018, 17, 115. [Google Scholar] [CrossRef] [PubMed]

| Cancer | In Vitro/In Vivo/Clinical | Cell Lines | Expression of FBXW7 | Modulation of FBXW7 | Mechanism of Action | Ref. |
|---|---|---|---|---|---|---|
| Bone | In vitro | U2OS, MG-63 | ↑FBXW7 | Ectopic overexpression | ↓c-Myc; ↓cyclin E | [27] |
| In vivo | Mouse xenografts | ↑FBXW7 | Ectopic overexpression | ↓c-Myc; ↓cyclin E | [27] | |
| In vitro | BMSCs | ↓FBXW7 | - | ↑CCL2 | [34] | |
| Brain | In vitro | A172, U87MG, U251MG, U373MG | ↓FBXW7 | - | ↑CCNE1; ↑MYC; | [104] |
| ↑AURKA | ||||||
| In vivo | Mouse xenografts | ↓FBXW7 | Overexpressed p53 mutants (C132W and R270C) | ↑c-Myc | [102] | |
| In vitro | U87, SHG139 | ↑FBXW7 | MALAT1 induced overexpression | ↓Cell viability | [107] | |
| In vitro | U343MG, Daoy | ↓FBXW7 | siRNA-mediated silencing | ↑SOX9 | [105] | |
| In vitro | U251, U373 | ↑FBXW7 | Ectopic overexpression | ↓Aurora B; ↓MCL-1; | [106] | |
| ↓Notch1 | ||||||
| In vitro | U251, U373 | ↑FBXW7 | Ectopic overexpression | ↓c-Myc | [108] | |
| Breast | In vitro | DLD1 | ↑FBXW7 | Ectopic overexpression | ↓KLF5 protein | [109] |
| In vitro | T47D | ↓FBXW7 | siRNA-mediated silencing | ↑c-Myc; ↑cyclin E | [110] | |
| In vitro | MCF10A, BT549 | ↓FBXW7 | Overexpression of FAM83D | ↑mTOR | [111] | |
| In vivo | Wnt9b+/− & Eya1+/− | ↓FBXW7 | - | ↓Eya1 ubiquitination | [112] | |
| In vitro | MCF7, T47D, MDA-MB-231 | ↓FBXW7 | siRNA-mediated silencing | ↑MCL-l; ↑PLK-1 | [113] | |
| In vitro | MCF-7 | ↓FBXW7 | Suppression by 27-HC at transcriptional level | ↑Myc | [114] | |
| In vitro | MDA-MB-453, MCF-7 and MCF-10A | ↓FBXW7 | shRNA-mediated silencing | ↑EglN2 | [115] | |
| In vitro | MCF-7, MDA-MB-231 | ↓FBXW7 | Binding of miR-32 to the 3-UTR of FBXW7 | ↓Apoptosis | [82] | |
| In vitro | MDA-MB-23, SKBR | ↑FBXW7 | Ectopic overexpression | ↓MTDH | [116] | |
| Cholangioc-arcinoma | In vitro | HuCCT1, RBE | ↓FBXW7 | shRNA-mediated silencing | ↑EMT | [117] |
| In vivo | HuCCT1-shFBXW7 injected mice | ↓FBXW7 | shRNA-mediated silencing | ↑EMT | [117] | |
| In vitro | Tissue samples | ↓FBXW7 | - | ↑c-Myc; ↑Ki-67 | [118] | |
| Colorectal | In vitro | LoVo, Colo 201 | ↓FBXW7 | siRNA-mediated silencing | ↑c-Myc; ↑cyclin E | [119] |
| In vitro | SW620, HT29, HCT116 | ↓FBXW7 | Knockdown of Rictor | ↑c-Myc; ↑cyclin E | [120] | |
| In vivo | FBXW7flox/flox mice | ↓FBXW7 | Conditional deletion | ↑c-Myc; ↑cyclin E | [121] | |
| In vitro | HCT116, DLD-1 | ↓FBXW7 | Depletion | ↑EMT | [99] | |
| In vitro | HCT116 | ↑FBXW7 | Ectopic overexpression | ↓ENO1 | [122] | |
| In vitro | SW480, RKO | ↓FBXW7 | Degradation by PLK2 | ↑Cyclin E | [123] | |
| In vivo | Mouse xenografts | ↓FBXW7 | Degradation by PLK2 | ↑Cyclin E | [123] | |
| In vitro | HCT116, DLD1, RKO, LoVo | ↑FBXW7 mutant | - | ↑MCL-1 | [124] | |
| In vitro | SW480, HCT116 | ↑FBXW7 | FAM83D knockdown | ↓Notch1 | [125] | |
| In vitro | HT-29, SW480, SW620, LoVo | ↓FBXW7 | Binding of miR-223 to the 3-UTR of FBXW7 | ↑EMT | [126] | |
| Esophageal | In vitro | TE8, Eca109, EC9706, KYSE30 | ↓FBXW7 | Binding of miR-27a-3p to the 3-UTR of FBXW7 | ↓G1/S arrest | [127] |
| Gastric | In vitro | ACP02, ACP03 | ↓FBXW7 | Deletion of one copy of FBXW7 | ↑c-Myc | [128] |
| In vitro | - | ↓FBXW7 | Binding of miR-25 to the 3-UTR of FBXW7 | - | [129] | |
| In vitro | AZ-521, MGC-803, BGC-823, SGC-7901 | ↑FBXW7 | Ectopic overexpression | ↓RhoA | [130] | |
| In vivo | Mouse xenografts | ↑FBXW7 | Ectopic overexpression | ↓RhoA | [130] | |
| In vivo | FBXW7 knockout mice | ↓FBXW7 | Haploinsufficiency | ↑c-Myc | [131] | |
| In vitro | Tissue samples | ↓FBXW7 | - | ↓Survival; ↓response | [132] | |
| Leukemia | In vitro | DU528, CEM, Jurkat | ↑FBXW7 mutant | Missense mutations of arginine (R465 & R505) | ↑MYC; ↑DELTEX1 | [133] |
| In vitro | Tissue samples | ↑FBXW7 mutant | Arginine substitutions at R479, R465, R505, and R689 | ↑NOTCH1; favorable outcome | [134] | |
| In vitro | FBXW7−/− DLD1 | ↑FBXW7 | Ectopic overexpression | ↓MCL-1 | [135] | |
| In vitro | Jurkat, CCRF-CEM | ↑FBXW7 | Oridonin- mediated upregulation | ↓c-Myc | [136] | |
| In vivo | FBXW7 knock-in mice | ↑FBXW7 mutants | Missense mutation | ↑c-Myc stability | [90] | |
| In vitro | Molt4, K562 | ↓FBXW7 | shRNA-mediated silencing | ↑GRα | [137] | |
| In vivo | T-ALL xenografts | ↑FBXW7 mutant | R479Q mutation | ↑GR stability | [137] | |
| In vitro | Jurkat cells | ↑FBXW7 | Knockdown of TAL1 | ↓Myc; ↓Notch1; | [138] | |
| ↓Cyclin E | ||||||
| In vitro | MT1 | ↑FBXW7 mutant | Mutation at arginine residues R479Q, R505C, and R465H | ↑Notch 1 | [139] | |
| In vitro | SU-DHL-2, OCI-LY-3. | ↑FBXW7 | Ectopic overexpression | ↓STAT3 | [140] | |
| Clinical | 50 patients | ↑FBXW7 mutant | - | Better clinical outcome | [141] | |
| Liver | In vitro | SMMC-7721, HepG2, Hep3B, Huh7 | ↑FBXW7 | Adenoviral delivery of p53 | ↓c-Myc; ↓cyclin E | [142] |
| In vitro | HepG2, Hep3B | ↑FBXW7 | Flag-FBXW7 overexpression | ↓YAP | [143] | |
| In vivo | Mouse xenografts | ↑FBXW7 | Flag-FBXW7 overexpression | ↓YAP | [143] | |
| In vitro | SMMC7721, HepG2 | ↑FBXW7 | STAT1 overexpression | ↓Cyclin A, D1, E; ↓CDK2; | [144] | |
| ↓Hes-1; ↓NF-κB p65 | ||||||
| Lung | In vitro | A549, HCT116 | ↓FBXW7 | siRNA-mediated silencing | ↑MCL-1 | [145] |
| In vitro | H2009, H1975 | ↓FBXW7 | siRNA-mediated silencing | ↑MCL-1 | [146] | |
| In vitro | H1299, H460 | ↑FBXW7 | - | ↓ZNF322A | [147] | |
| In vivo | Mouse xenografts | ↑FBXW7 | - | ↓ZNF322A | [147] | |
| In vitro | A549, H460, H1299 | ↓FBXW7 | Binding of miR-367 to the 3-UTR of FBXW7 | ↑Wnt signaling | [148] | |
| In vitro | PC-9, HCC827, H3122, H3255, H1975, H1299 | ↓FBXW7 | shRNA-mediated silencing | ↑MCL-1 | [149] | |
| In vitro | A549, H322, H460, GLC-82, SPC-A1 | ↓FBXW7 | MiR-544a overexpression/ TINCR knockdown | ↑Proliferation; ↑invasion | [150] | |
| In vitro | PC9, H1299 | ↓FBXW7 | shRNA-mediated silencing | ↑EMT | [151] | |
| In vivo | FBXW7+/− mice | ↓FBXW7 | shRNA-mediated silencing | ↑Tumorigenesis | [151] | |
| Oral | In vitro | UM1, UM2, Cal27, SCC1, SCC15, SCC25 | ↓FBXW7 | Binding of miR-24 to the 3-UTR of FBXW7 | ↑Tumorigenesis | [152] |
| In vitro | Tissue samples | ↓FBXW7 | - | Poor prognosis | [153] | |
| Pancreas | In vitro | BxPC-3, Colo-357 | ↑FBXW7 | Nuclear retention by KPT-185 | ↓Notch1; ↓ c-Myc; ↓VEGF | [154] |
| In vivo | Colo-357 xenografts | ↑FBXW7 | Nuclear retention by KPT-185 | ↓Notch1 | [154] | |
| In vitro | MIAPaCa2, BxPC3, PANC1 | ↓FBXW7 | shRNA-mediated silencing | ↑β-catenin | [155] | |
| In vitro | SUIT-2 | ↓FBXW7 | siRNA-mediated silencing | ↑MCL-1 | [156] | |
| In vitro | PANC-1, Mia PaCa-2 | ↑FBXW7 | Ectopic overexpression | ↑ENT1 | [157] | |
| Renal | In vitro | AHCN, A704 | ↑FBXW7 | Ectopic overexpression | ↓c-Myc; ↓c-Jun | [158] |
| In vitro | 786-O, ACHN | ↑FBXW7 | Ectopic overexpression | ↓MMP-2, -9, -13 | [159] | |
| Skin | In vitro | MMRU, RPEP | ↓FBXW7 | siRNA-mediated silencing | ↑MAPK/ERK | [15] |
| In vivo | FBXW7F/F mice | ↓FBXW7 | - | ↑c-Myc | [160] | |
| In vitro | WC00125, WM39, WM3702, WM3862 | ↓FBXW7 | Nonsynonymous mutations; shRNA-mediated silencing | ↑Notch | [161] | |
| In vivo | Lysm−FBXW7f/f, Lysm+FBXW7 f/f mice | ↓FBXW7 | Myeloid cell-specific deletion | ↓MAM | [162] | |
| In vitro | 501mel, SKMEL28, SKMEL24, WM3862, WM39 | ↓FBXW7 | shRNA- mediated silencing | ↓Nuclear HSF1 | [65] | |
| In vitro | MM415, MM485, HT144, A2058, SH4 | ↓FBXW7 | siRNA-mediated silencing | ↑MITF/PGC-1 signaling | [163] |
| Cancer | Combination | In Vitro/In Vivo | Mechanism of Action | Ref. |
|---|---|---|---|---|
| Breast | ↑FBXW7+ paclitaxel | In vitro | ↓MCL-1; ↓PLK-1 | [113] |
| Colorectal | ↑FBXW7+ doxorubicin | In vitro | ↓EMT | [126] |
| ↓FBXW7+ irinotecan | In vitro | ↑c-Myc; ↓CSC | [203] | |
| CML | ↓FBXW7+ imatinib | In vivo | ↓LICs | [202] |
| Gastric | ↑FBXW7+ trastuzumab | In vitro | ↓MCL-1; ↓c-Myc; ↓c-Jun | [186] |
| Glioblastoma | ↑FBXW7+ temozolomide | In vitro | ↓Aurora B; ↓MCL-1; ↓ Notch-1 | [106] |
| Liver | ↑FBXW7+ doxorubicin | In vitro | ↓EMT | [204] |
| Lung | ↑FBXW7+ cisplatin | In vitro | ↓EMT | [205] |
| Lung | ↑FBXW7+ TKI | In vitro | ↓MCL-1 | [149] |
| Lung | ↑FBXW7+ TKI | In vivo | ↓MCL-1 | [149] |
| Nasopharynx | ↑FBXW7+ cisplatin | In vitro | ↓MRP | [206] |
| Pancreas | ↑FBXW7+ gemcitabine | In vitro | ↑ENT1 | [157] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sailo, B.L.; Banik, K.; Girisa, S.; Bordoloi, D.; Fan, L.; Halim, C.E.; Wang, H.; Kumar, A.P.; Zheng, D.; Mao, X.; et al. FBXW7 in Cancer: What Has Been Unraveled Thus Far? Cancers 2019, 11, 246. https://doi.org/10.3390/cancers11020246
Sailo BL, Banik K, Girisa S, Bordoloi D, Fan L, Halim CE, Wang H, Kumar AP, Zheng D, Mao X, et al. FBXW7 in Cancer: What Has Been Unraveled Thus Far? Cancers. 2019; 11(2):246. https://doi.org/10.3390/cancers11020246
Chicago/Turabian StyleSailo, Bethsebie Lalduhsaki, Kishore Banik, Sosmitha Girisa, Devivasha Bordoloi, Lu Fan, Clarissa Esmeralda Halim, Hong Wang, Alan Prem Kumar, Dali Zheng, Xinliang Mao, and et al. 2019. "FBXW7 in Cancer: What Has Been Unraveled Thus Far?" Cancers 11, no. 2: 246. https://doi.org/10.3390/cancers11020246
APA StyleSailo, B. L., Banik, K., Girisa, S., Bordoloi, D., Fan, L., Halim, C. E., Wang, H., Kumar, A. P., Zheng, D., Mao, X., Sethi, G., & Kunnumakkara, A. B. (2019). FBXW7 in Cancer: What Has Been Unraveled Thus Far? Cancers, 11(2), 246. https://doi.org/10.3390/cancers11020246







