KCNMA1 Expression Is Downregulated in Colorectal Cancer via Epigenetic Mechanisms
Abstract
:1. Introduction
2. Results
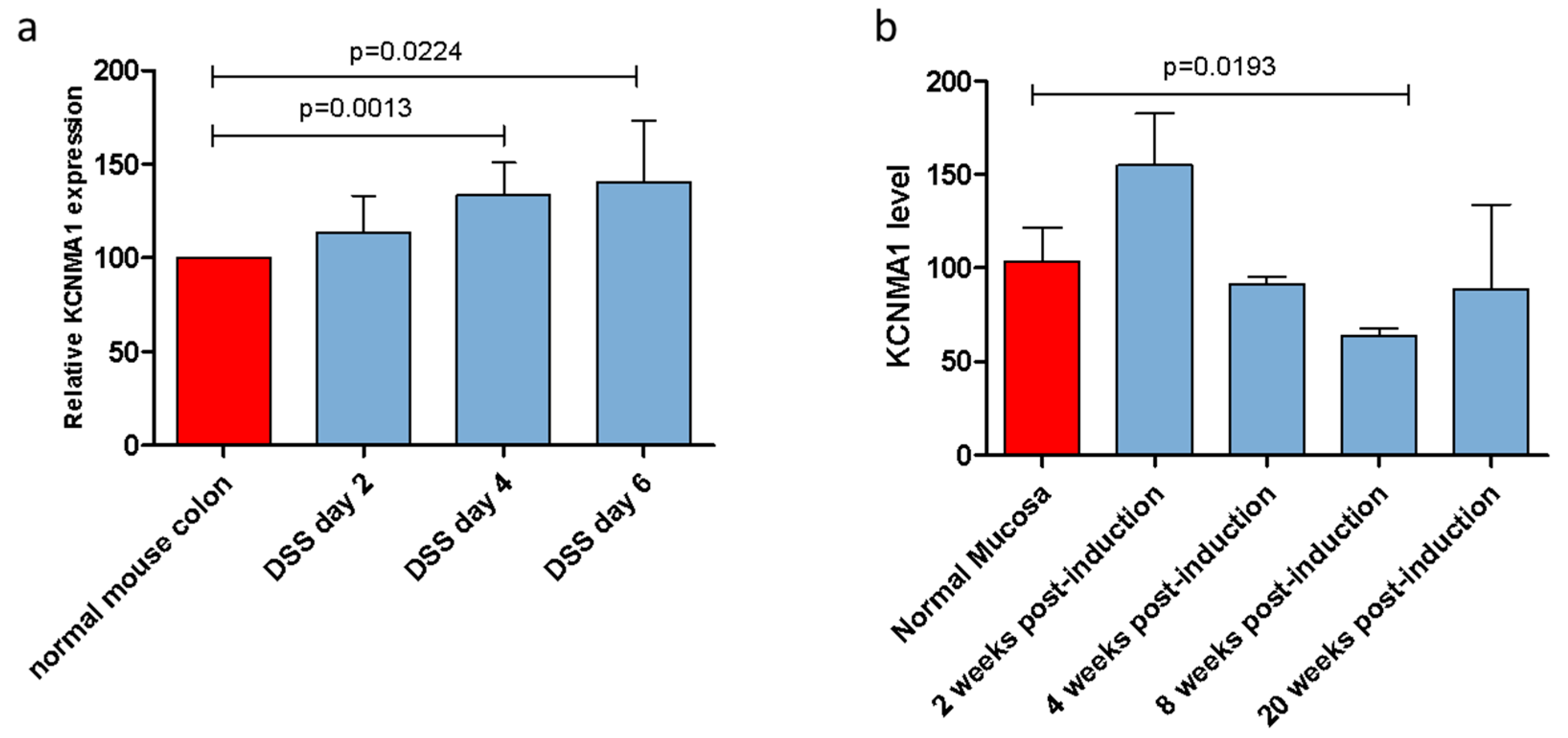
2.1. KCNMA1 Expression Is Modulated in Preclinical Models of Ulcerative Colitis (UC) and UC-Associated CRC
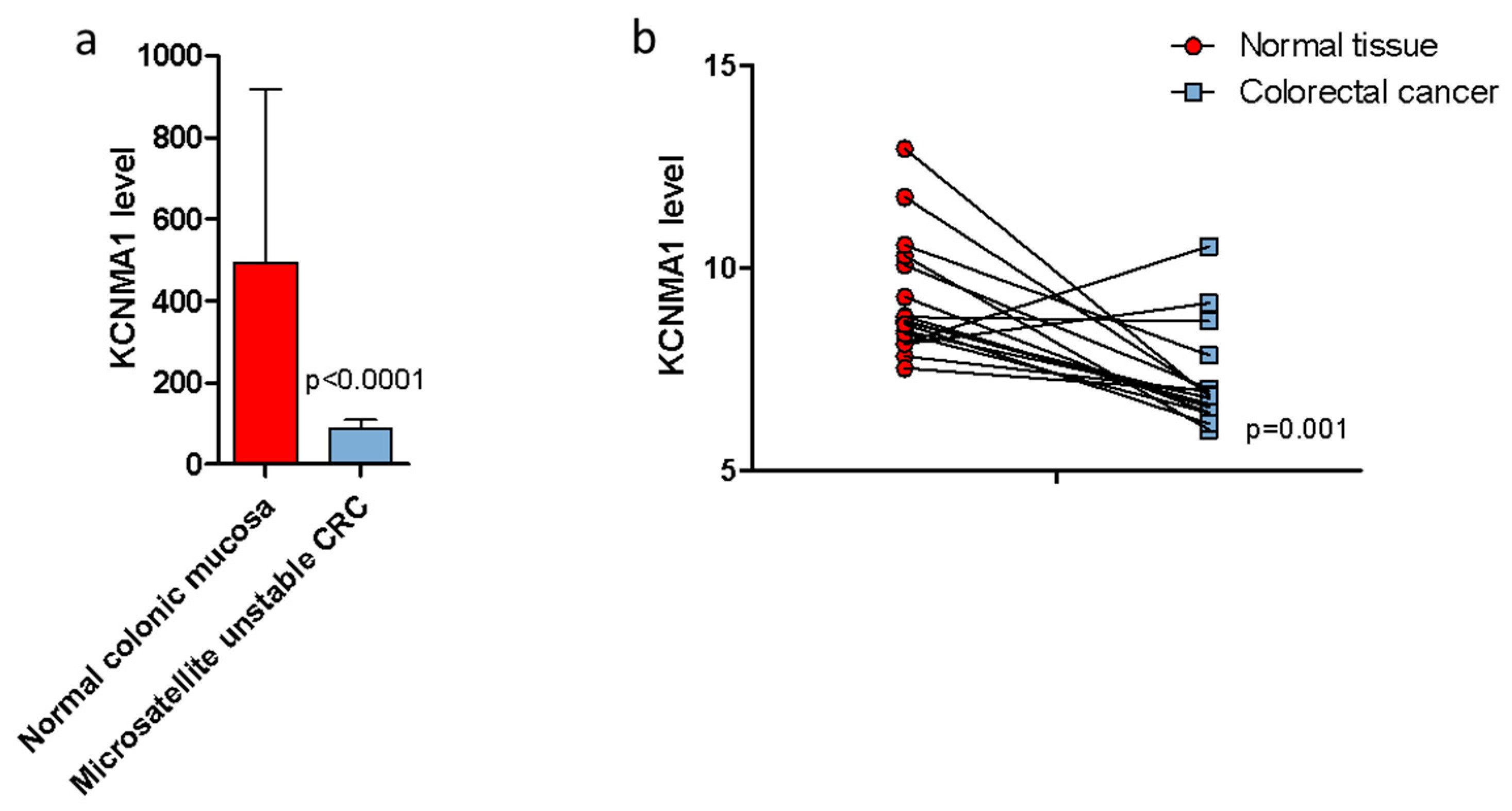
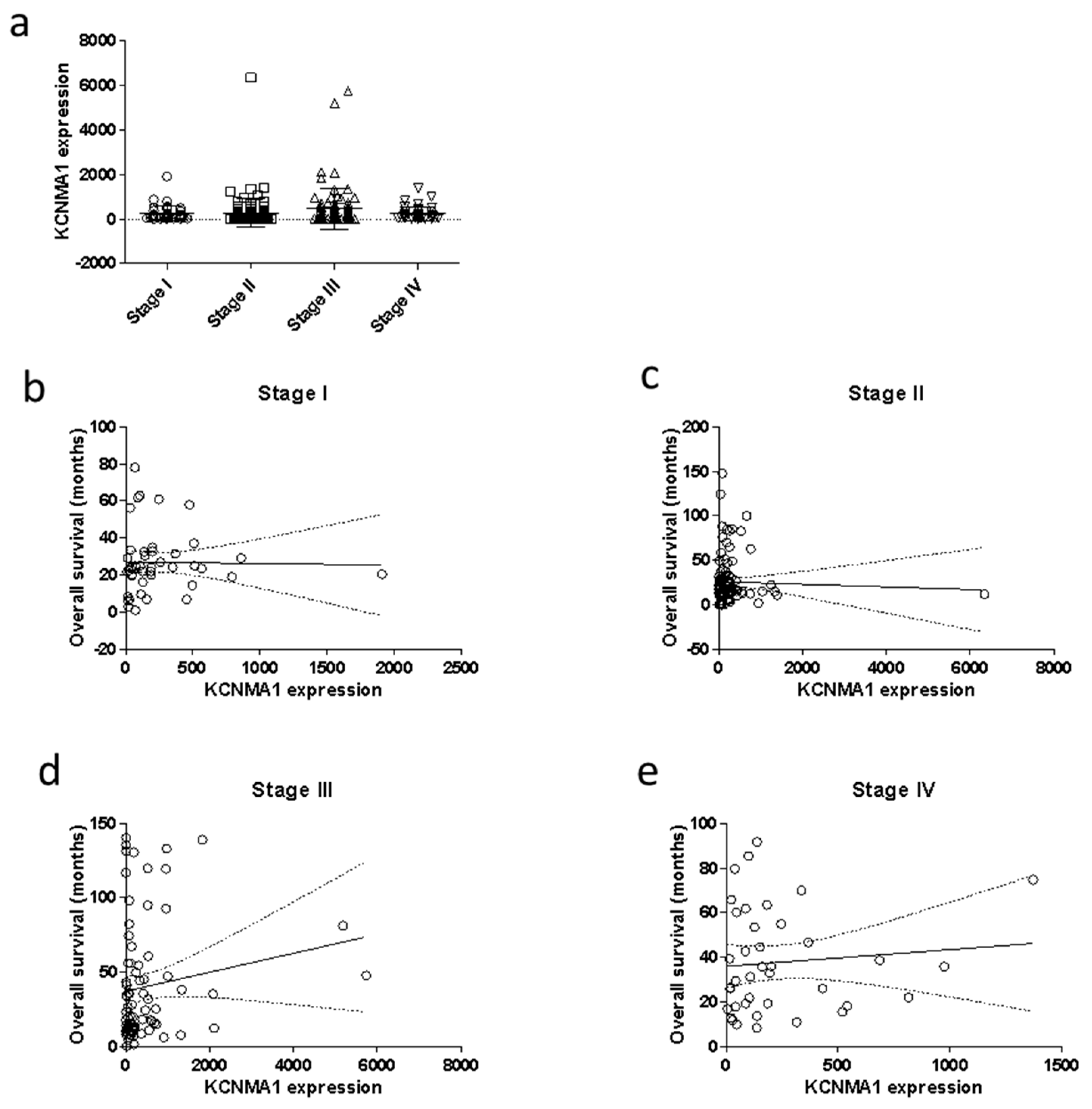
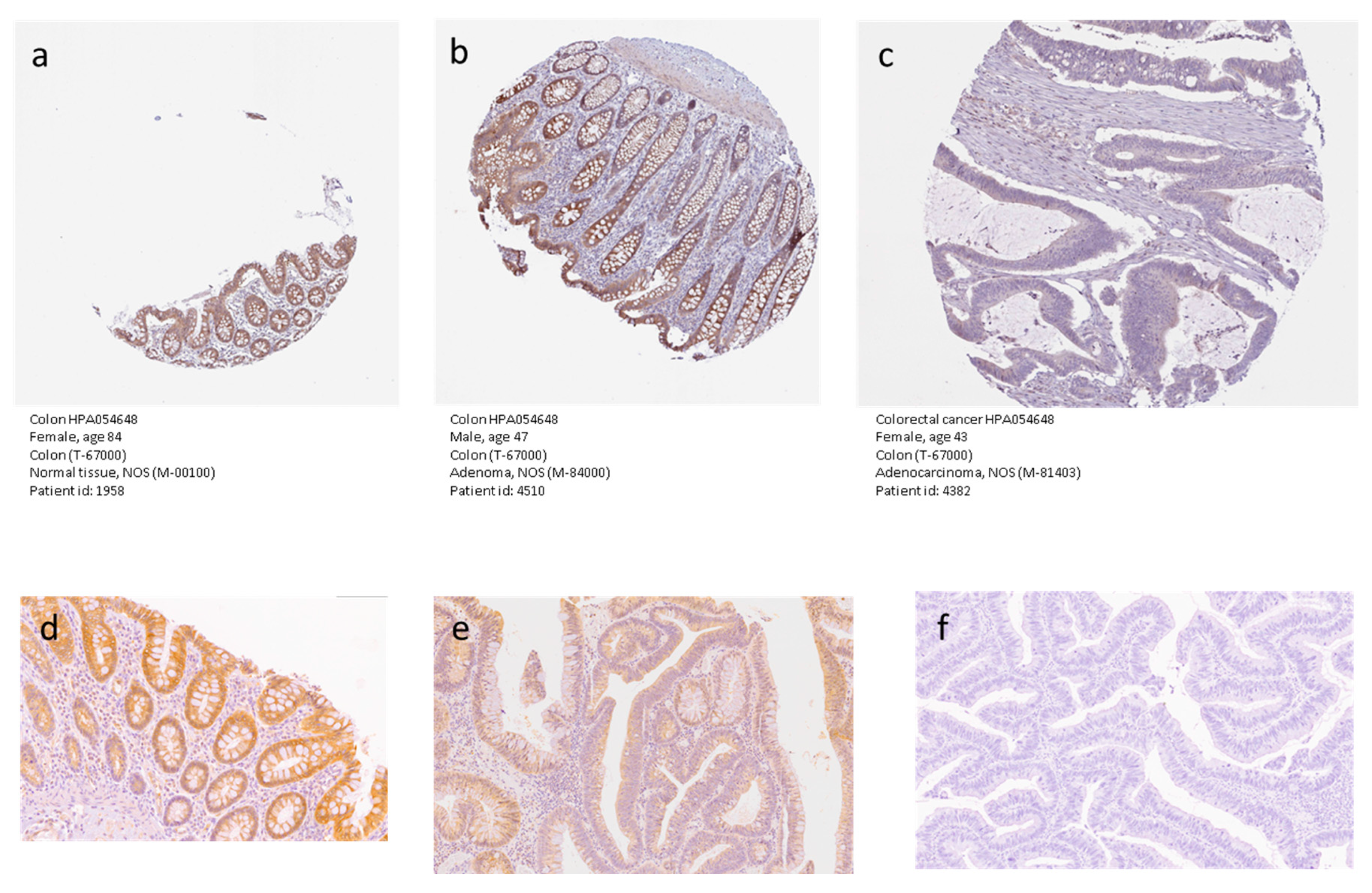
2.2. KCNMA1 Levels Are Reduced in Human CRC
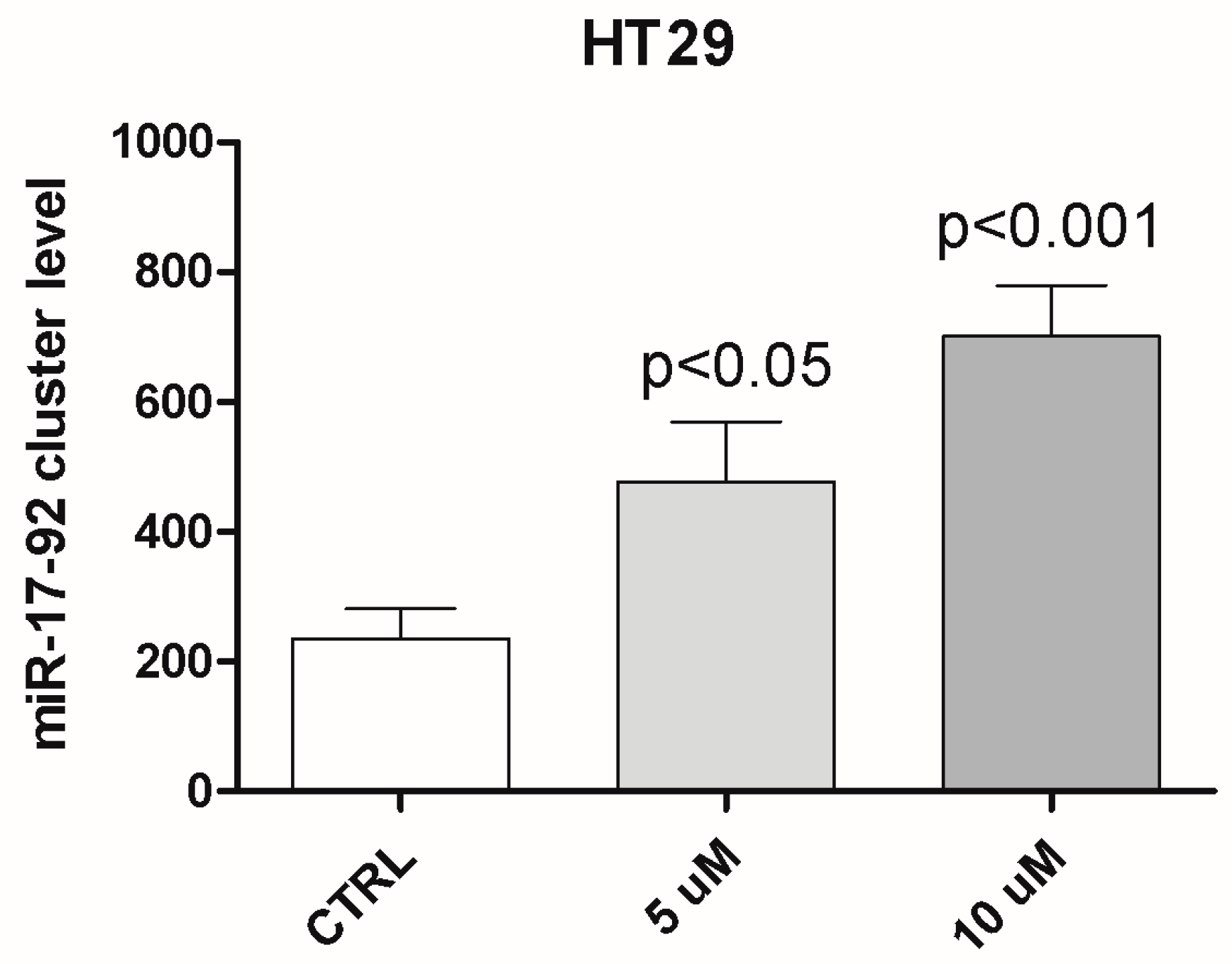
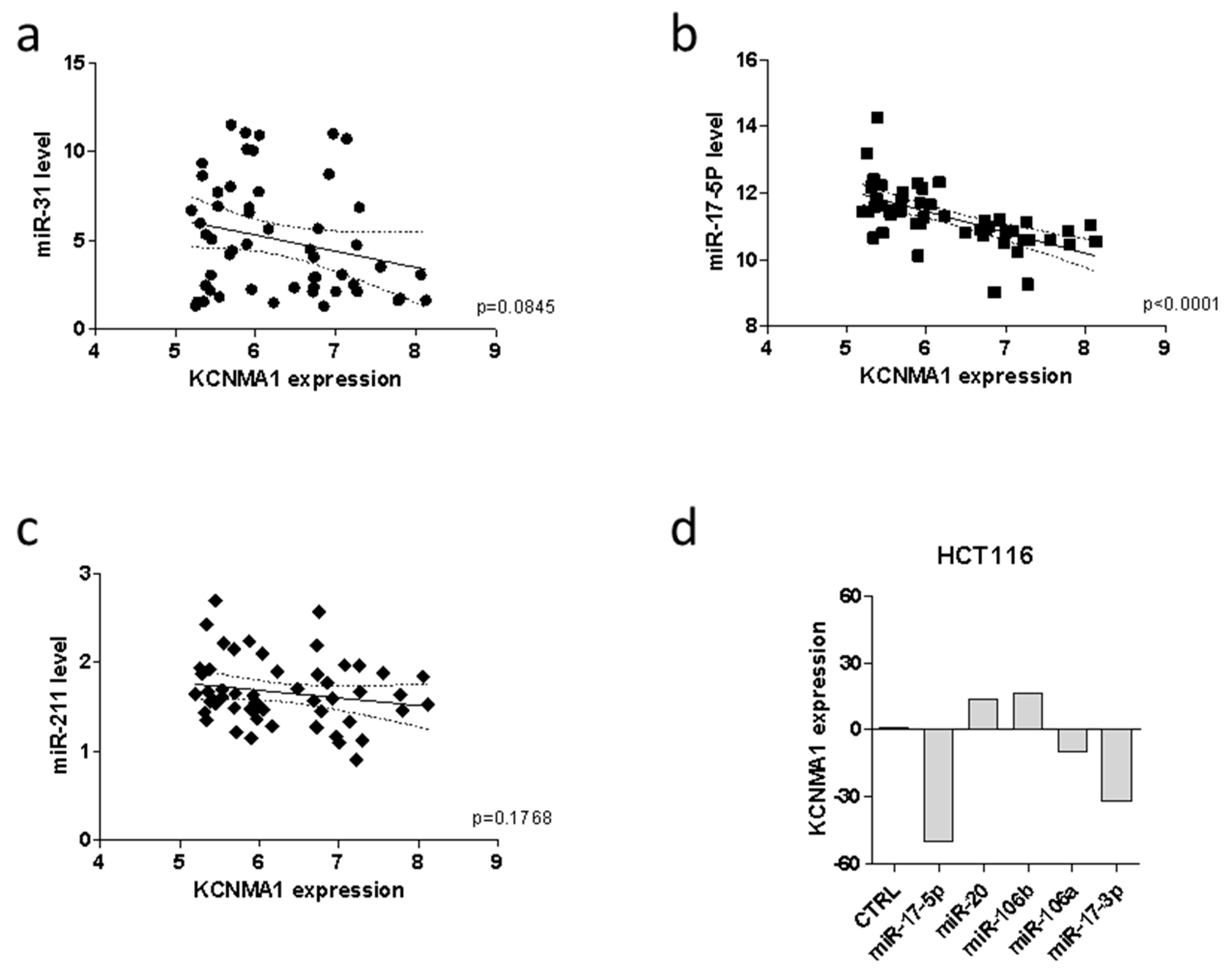
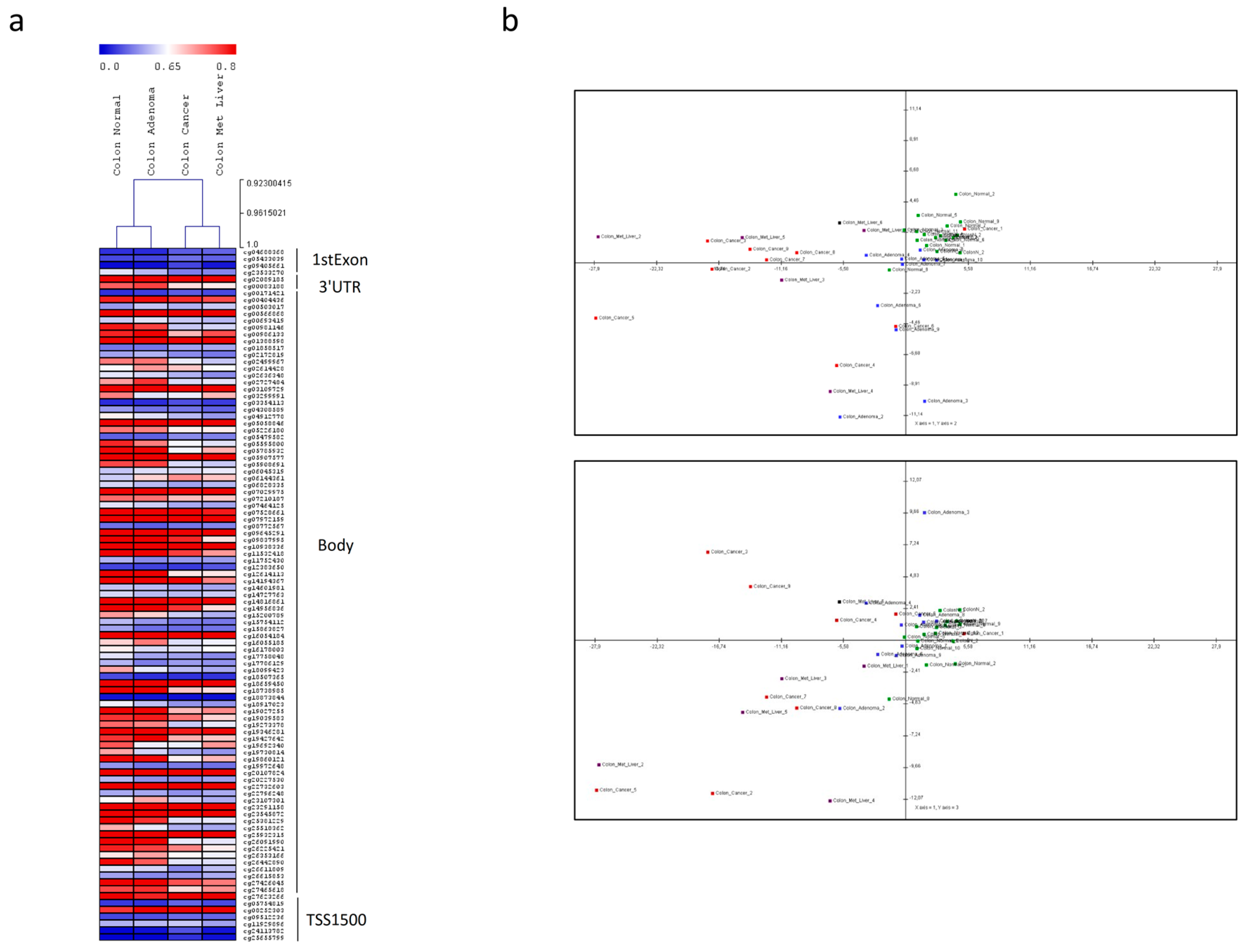
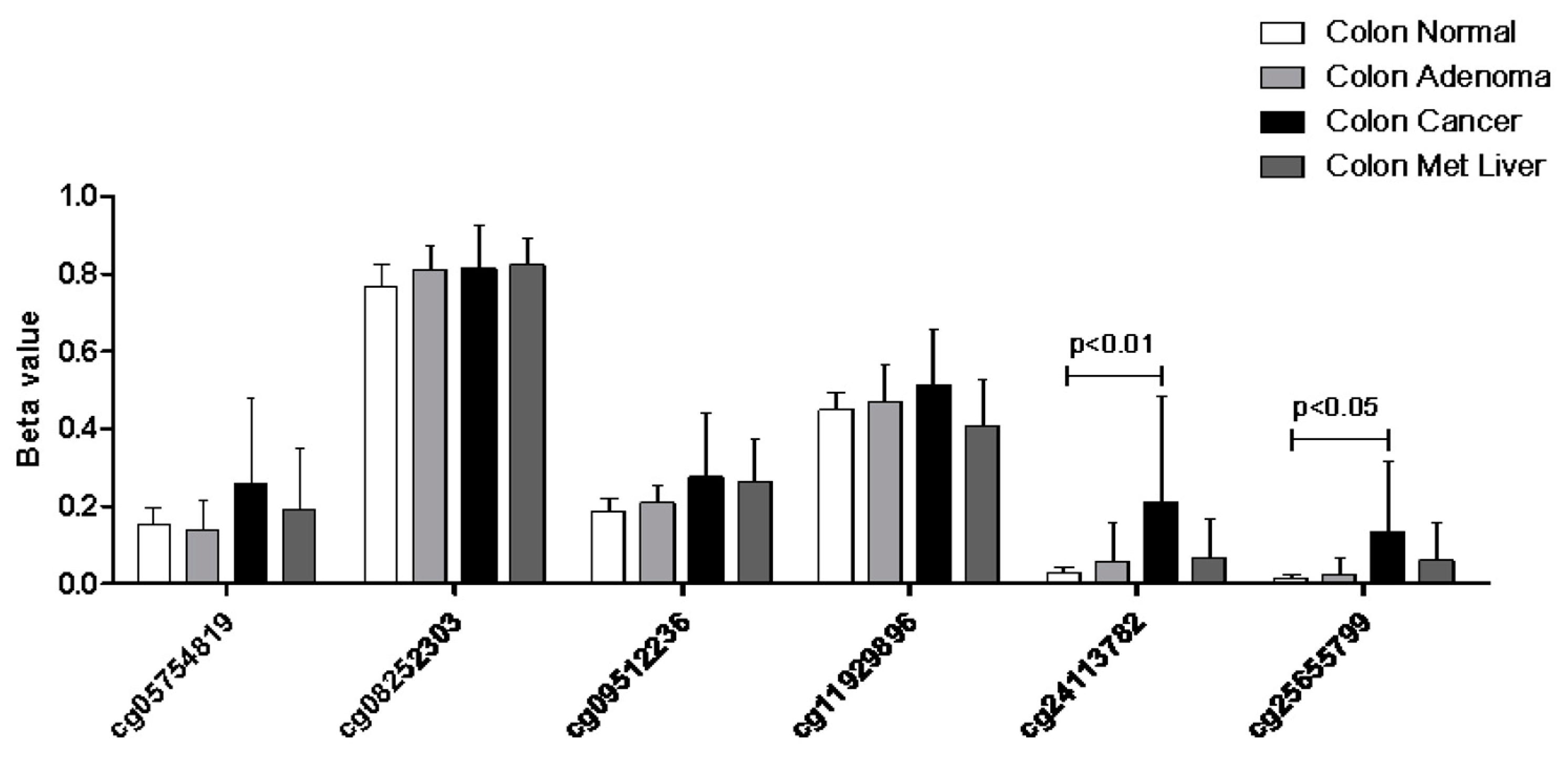
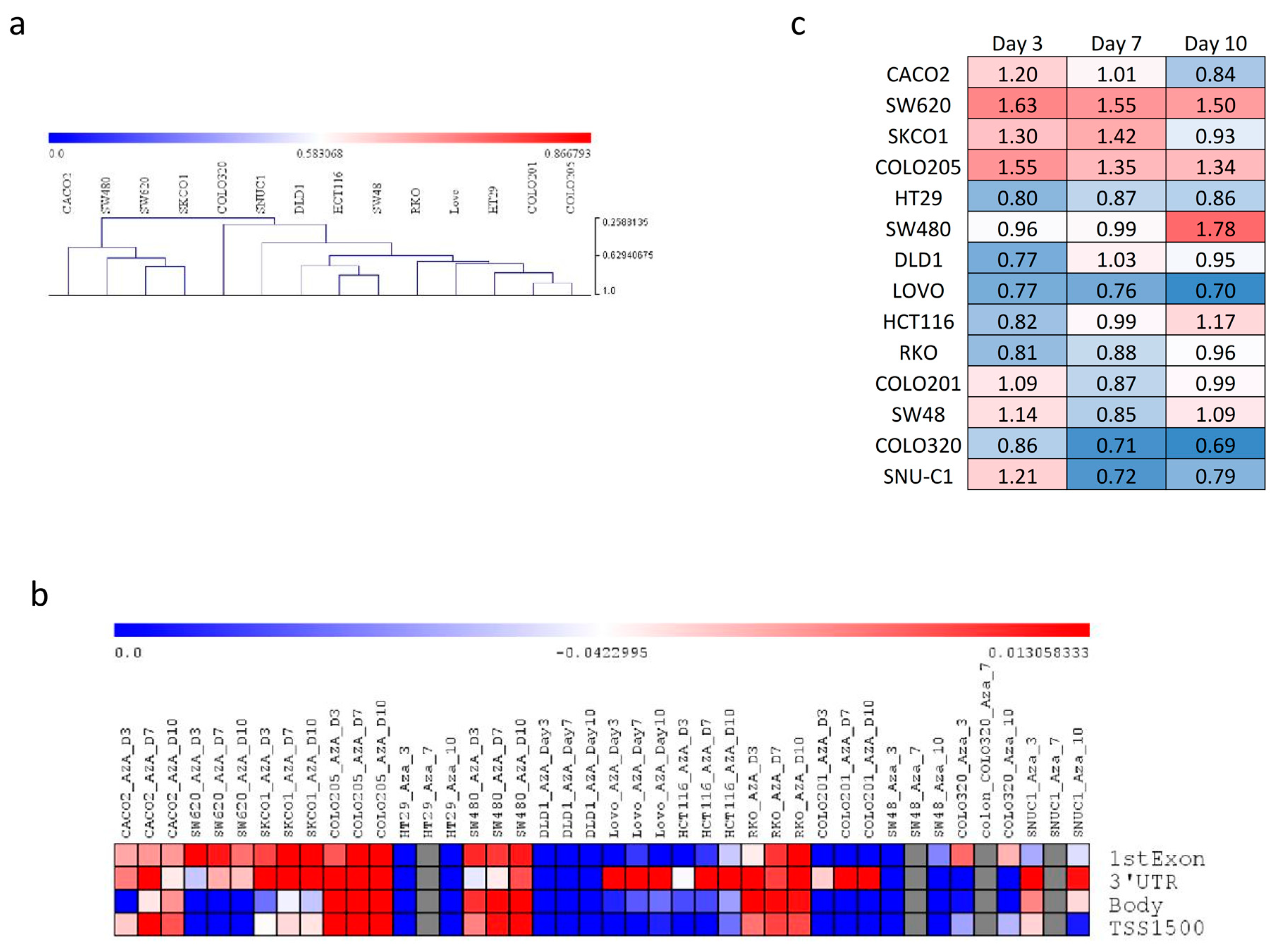
2.3. MiRNA Analysis and Methylation Profile of KCNMA1
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. Murine Models of UC and UC-Associated Cancer
4.3. RNA Isolation and Real-Time RT-PCR
4.4. Analysis of Transcription Profiles of KCNMA1 in Patients with CRC
4.5. Immunohistochemistry Analysis of KCNMA1 Expression in CRC Samples
4.6. Analysis of Epigenetic Regulation of KCNMA1 Expression in CRC
4.7. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A
| Sample | X | Y | Z |
|---|---|---|---|
| Colon_Normal_1 | 1.735467 | 1.358078 | −1.74759 |
| Colon_Normal_2 | 4.342013 | 5.111035 | −1.65102 |
| Colon_Normal_3 | −0.25661 | 2.49674 | 0.359764 |
| Colon_Normal_4 | 1.501905 | 2.161722 | 0.570626 |
| Colon_Normal_5 | 0.961287 | 3.563164 | 0.071522 |
| Colon_Normal_6 | 3.428178 | 1.750228 | 1.24613 |
| Colon_Normal_7 | 3.580552 | 2.792617 | 1.531011 |
| Colon_Normal_8 | −1.62841 | −0.42259 | −4.3392 |
| Colon_Normal_9 | 4.737718 | 3.108791 | 1.336153 |
| Colon_Normal_10 | 0.88916 | 1.758846 | −0.51744 |
| Colon_Normal_11 | 0.849407 | 2.382122 | 1.14948 |
| Colon_Normal_12 | 2.52812 | 1.950024 | 0.647312 |
| ColonN_2 | 2.949582 | 2.091265 | 2.387948 |
| ColonN_2 | 4.124992 | 1.94098 | 0.049728 |
| ColonN_2 | 4.724679 | 0.837779 | 2.424594 |
| ColonN_2 | 4.442956 | 2.109996 | 1.577544 |
| ColonN_2 | 2.655884 | 0.930409 | 1.088807 |
| Colon_Adenoma_1 | 1.481603 | 0.340708 | 1.481484 |
| Colon_Adenoma_2 | −6.04664 | −11.1386 | −5.05503 |
| Colon_Adenoma_3 | 1.561727 | −9.99486 | 9.808415 |
| Colon_Adenoma_4 | −3.68738 | 0.668282 | 2.941357 |
| Colon_Adenoma_5 | −0.52092 | 0.373219 | 1.291463 |
| Colon_Adenoma_6 | −2.653 | −3.03014 | −0.95898 |
| Colon_Adenoma_7 | −0.44732 | −0.00646 | −0.31317 |
| Colon_Adenoma_8 | 1.141704 | 1.037116 | 2.029349 |
| Colon_Adenoma_9 | −1.0016 | −4.7615 | −1.04647 |
| Colon_Adenoma_10 | 2.62769 | 0.301795 | 1.581635 |
| Colon_Cancer_1 | 5.112652 | 2.56774 | 0.636602 |
| Colon_Cancer_2 | −17.4902 | −0.3567 | −11.5103 |
| Colon_Cancer_3 | −17.9117 | 1.695159 | 6.819301 |
| Colon_Cancer_4 | −6.336 | −7.3712 | 1.640066 |
| Colon_Cancer_5 | −27.9047 | −3.92621 | −11.265 |
| Colon_Cancer_6 | −1.03183 | −4.53172 | 2.102261 |
| Colon_Cancer_7 | −12.6164 | 0.340978 | −4.18775 |
| Colon_Cancer_8 | −9.90718 | 0.857501 | −5.01775 |
| Colon_Cancer_9 | −14.0838 | 1.098967 | 4.202911 |
| Colon_Met_Liver_1 | −3.8611 | 2.457655 | −1.83603 |
| Colon_Met_Liver_2 | −27.6862 | 2.011681 | −9.32142 |
| Colon_Met_Liver_3 | −11.2724 | −1.14682 | −2.78756 |
| Colon_Met_Liver_4 | −6.92045 | −9.26846 | −12.0695 |
| Colon_Met_Liver_5 | −14.7814 | 1.954306 | −5.35754 |
| Colon_Met_Liver_6 | −6.07612 | 3.006146 | 3.031533 |
References
- Ge, L.; Hoa, N.T.; Wilson, Z.; Arismendi-Morillo, G.; Kong, X.-T.; Tajhya, R.B.; Beeton, C.; Jadus, M.R. Big Potassium (BK) ion channels in biology, disease and possible targets for cancer immunotherapy. Int. Immunopharmacol. 2014, 22, 427–443. [Google Scholar] [CrossRef] [PubMed]
- Bloch, M.; Ousingsawat, J.; Simon, R.; Schraml, P.; Gasser, T.C.; Mihatsch, M.J.; Kunzelmann, K.; Bubendorf, L. KCNMA1 gene amplification promotes tumor cell proliferation in human prostate cancer. Oncogene 2007, 26, 2525–2534. [Google Scholar] [CrossRef] [PubMed]
- Xie, M.; Zhou, L.; Chen, X.; Gainey, L.O.; Xiao, J.; Nanes, M.S.; Hou, A.; You, S.; Chen, Q. Progesterone and Src Family Inhibitor PP1 Synergistically Inhibit Cell Migration and Invasion of Human Basal Phenotype Breast Cancer Cells. Biomed. Res. Int. 2015, 2015, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Oeggerli, M.; Tian, Y.; Ruiz, C.; Wijker, B.; Sauter, G.; Obermann, E.; Güth, U.; Zlobec, I.; Sausbier, M.; Kunzelmann, K.; et al. Role of KCNMA1 in Breast Cancer. PLoS ONE 2012, 7, e41664. [Google Scholar] [CrossRef] [PubMed]
- Khaitan, D.; Sankpal, U.T.; Weksler, B.; Meister, E.A.; Romero, I.A.; Couraud, P.-O.; Ningaraj, N.S. Role of KCNMA1gene in breast cancer invasion and metastasis to brain. BMC Cancer 2009, 9, 258. [Google Scholar] [CrossRef] [PubMed]
- Du, C.; Zheng, Z.; Li, D.; Chen, L.; Li, N.; Yi, X.; Yang, Y.; Guo, F.; Liu, W.; Xie, X.; et al. BKCa promotes growth and metastasis of prostate cancer through facilitating the coupling between αvβ3 integrin and FAK. Oncotarget 2016, 7, 40174–40188. [Google Scholar] [CrossRef] [PubMed]
- Bury, M.; Girault, A.; Mégalizzi, V.; Spiegl-Kreinecker, S.; Mathieu, V.; Berger, W.; Evidente, A.; Kornienko, A.; Gailly, P.; Vandier, C.; et al. Ophiobolin A induces paraptosis-like cell death in human glioblastoma cells by decreasing BKCa channel activity. Cell Death Dis. 2013, 4, e561. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, A.; Vera, E.; Gamboa-Domínguez, A.; Lambert, P.; Gariglio, P.; Camacho, J. Calcium-activated potassium channels as potential early markers of human cervical cancer. Oncol. Lett. 2018, 15, 7249–7254. [Google Scholar] [CrossRef] [PubMed]
- Ma, G.; Liu, H.; Hua, Q.; Wang, M.; Du, M.; Lin, Y.; Ge, Y.; Gong, W.; Zhao, Q.; Qiang, F.; et al. KCNMA1 cooperating with PTK2 is a novel tumor suppressor in gastric cancer and is associated with disease outcome. Mol. Cancer 2017, 16, 46. [Google Scholar] [CrossRef] [PubMed]
- Samuel, P.; Pink, R.C.; Caley, D.P.; Currie, J.M.S.; Brooks, S.A.; Carter, D.R.F. Over-expression of miR-31 or loss of KCNMA1 leads to increased cisplatin resistance in ovarian cancer cells. Tumor Biol. 2016, 37, 2565–2573. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.Y.; Wright, C.M.; Kirschner, M.B.; Williams, M.; Sarun, K.H.; Sytnyk, V.; Leshchynska, I.; Edelman, J.J.; Vallely, M.P.; McCaughan, B.C.; et al. KCa1.1, a calcium-activated potassium channel subunit alpha 1, is targeted by miR-17-5p and modulates cell migration in malignant pleural mesothelioma. Mol. Cancer 2016, 15, 44. [Google Scholar] [CrossRef] [PubMed]
- Mazar, J.; DeYoung, K.; Khaitan, D.; Meister, E.; Almodovar, A.; Goydos, J.; Ray, A.; Perera, R.J. The Regulation of miRNA-211 Expression and Its Role in Melanoma Cell Invasiveness. PLoS ONE 2010, 5, e13779. [Google Scholar] [CrossRef] [PubMed]
- De Robertis, M.; Massi, E.; Poeta, M.L.; Carotti, S.; Morini, S.; Cecchetelli, L.; Signori, E.; Fazio, V.M. The AOM/DSS murine model for the study of colon carcinogenesis: From pathways to diagnosis and therapy studies. J. Carcinog. 2011, 10, 9. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T. Development of an Inflammation-Associated Colorectal Cancer Model and Its Application for Research on Carcinogenesis and Chemoprevention. Int. J. Inflam. 2012, 2012, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Alhopuro, P.; Sammalkorpi, H.; Niittymäki, I.; Biström, M.; Raitila, A.; Saharinen, J.; Nousiainen, K.; Lehtonen, H.J.; Heliövaara, E.; Puhakka, J.; et al. Candidate driver genes in microsatellite-unstable colorectal cancer. Int. J. Cancer 2012, 130, 1558–1566. [Google Scholar] [CrossRef] [PubMed]
- Khamas, A.; Ishikawa, T.; Shimokawa, K.; Mogushi, K.; Iida, S.; Ishiguro, M.; Mizushima, H.; Tanaka, H.; Uetake, H.; Sugihara, K. Screening for epigenetically masked genes in colorectal cancer Using 5-Aza-2′-deoxycytidine, microarray and gene expression profile. Cancer Genom. Proteom. 2012, 9, 67–75. [Google Scholar]
- Boland, C.R.; Goel, A. Microsatellite Instability in Colorectal Cancer. Gastroenterology 2010, 138, 2073–2087.e3. [Google Scholar] [CrossRef] [PubMed]
- Pizzini, S.; Bisognin, A.; Mandruzzato, S.; Biasiolo, M.; Facciolli, A.; Perilli, L.; Rossi, E.; Esposito, G.; Rugge, M.; Pilati, P.; et al. Impact of microRNAs on regulatory networks and pathways in human colorectal carcinogenesis and development of metastasis. BMC Genom. 2013, 14, 589. [Google Scholar] [CrossRef] [PubMed]
- Timp, W.; Bravo, H.C.; McDonald, O.G.; Goggins, M.; Umbricht, C.; Zeiger, M.; Feinberg, A.P.; Irizarry, R.A. Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors. Genome Med. 2014, 6, 61. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Chiappinelli, K.B.; Guzzetta, A.A.; Easwaran, H.; Yen, R.-W.C.; Vatapalli, R.; Topper, M.J.; Luo, J.; Connolly, R.M.; Azad, N.S.; et al. Immune regulation by low doses of the DNA methyltransferase inhibitor 5-azacitidine in common human epithelial cancers. Oncotarget 2014, 5, 587–598. [Google Scholar] [CrossRef] [PubMed]
| Sample ID | Cancer Type | Protein Change | Mutation Type | Copy # |
|---|---|---|---|---|
| TCGA-AA-3831-01 | Colon Adenocarcinoma | S60del | IF del | Diploid |
| TCGA-AA-3524-01 | Colon Adenocarcinoma | T905M | Missense | Diploid |
| TCGA-AA-A01V-01 | Colon Adenocarcinoma | R909W | Missense | Diploid |
| TCGA-AA-A00E-01 | Colon Adenocarcinoma | S580P | Missense | Diploid |
| TCGA-AA-A00L-01 | Colon Adenocarcinoma | A381T | Missense | Diploid |
| TCGA-AA-3984-01 | Colon Adenocarcinoma | S1190P | Missense | Diploid |
| TCGA-AA-A00N-01 | Mucinous Adenocarcinoma of the... | K1153T | Missense | Diploid |
| TCGA-AA-A00N-01 | Mucinous Adenocarcinoma of the... | N801S | Missense | Diploid |
| TCGA-AA-A010-01 | Colon Adenocarcinoma | L1236F | Missense | Diploid |
| TCGA-AA-A010-01 | Colon Adenocarcinoma | E824D | Missense | Diploid |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Basile, M.S.; Fagone, P.; Mangano, K.; Mammana, S.; Magro, G.; Salvatorelli, L.; Li Destri, G.; La Greca, G.; Nicoletti, F.; Puleo, S.; et al. KCNMA1 Expression Is Downregulated in Colorectal Cancer via Epigenetic Mechanisms. Cancers 2019, 11, 245. https://doi.org/10.3390/cancers11020245
Basile MS, Fagone P, Mangano K, Mammana S, Magro G, Salvatorelli L, Li Destri G, La Greca G, Nicoletti F, Puleo S, et al. KCNMA1 Expression Is Downregulated in Colorectal Cancer via Epigenetic Mechanisms. Cancers. 2019; 11(2):245. https://doi.org/10.3390/cancers11020245
Chicago/Turabian StyleBasile, Maria Sofia, Paolo Fagone, Katia Mangano, Santa Mammana, Gaetano Magro, Lucia Salvatorelli, Giovanni Li Destri, Gaetano La Greca, Ferdinando Nicoletti, Stefano Puleo, and et al. 2019. "KCNMA1 Expression Is Downregulated in Colorectal Cancer via Epigenetic Mechanisms" Cancers 11, no. 2: 245. https://doi.org/10.3390/cancers11020245
APA StyleBasile, M. S., Fagone, P., Mangano, K., Mammana, S., Magro, G., Salvatorelli, L., Li Destri, G., La Greca, G., Nicoletti, F., Puleo, S., & Pesce, A. (2019). KCNMA1 Expression Is Downregulated in Colorectal Cancer via Epigenetic Mechanisms. Cancers, 11(2), 245. https://doi.org/10.3390/cancers11020245







