Companion Animals as Models for Inhibition of STAT3 and STAT5
Abstract
1. Introduction
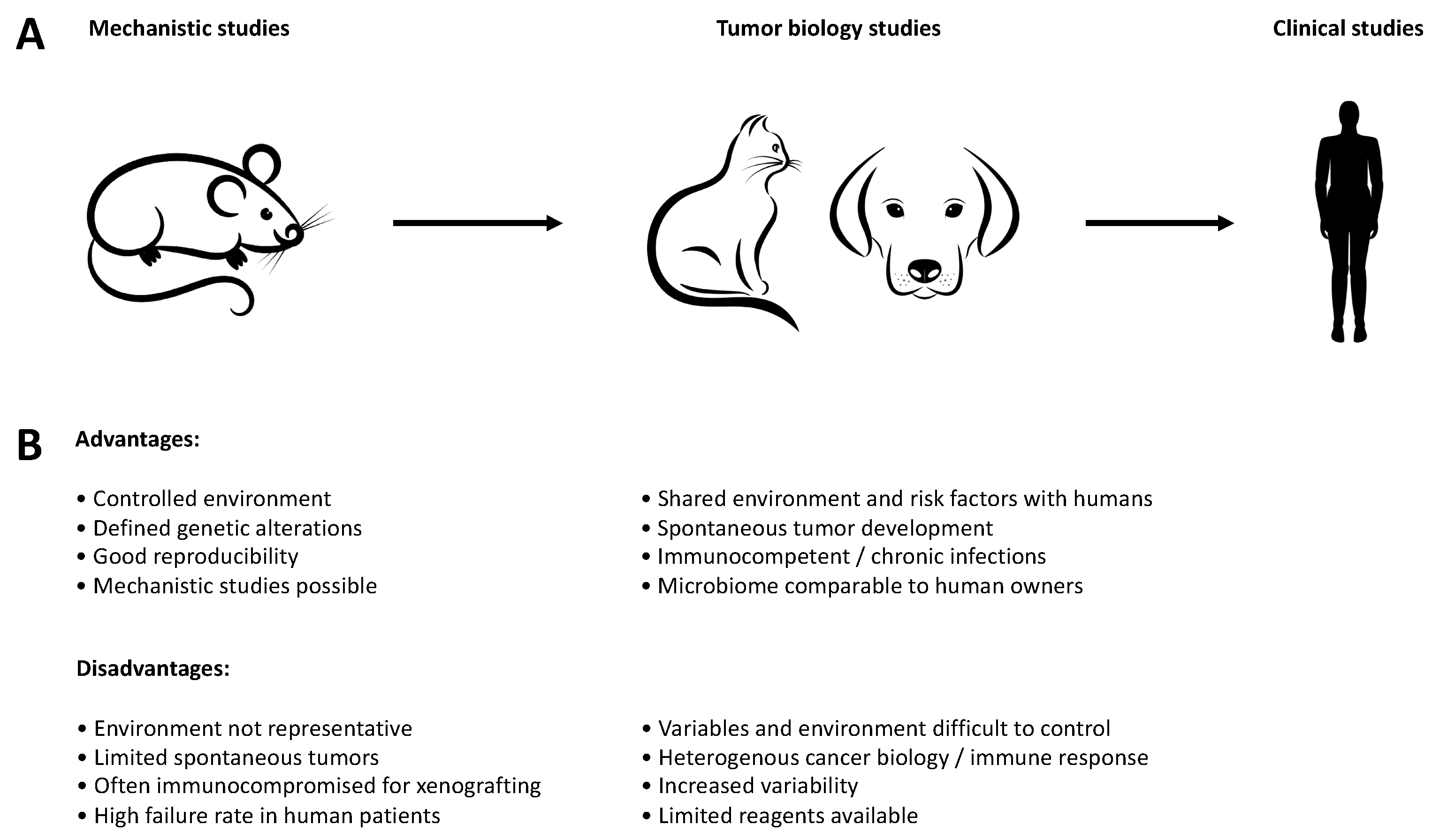
2. Preclinical Models
3. Advantages and Disadvantages of Canine Tumor Models
4. Relevance and Conservation of the JAK-STAT Signaling Pathway
5. Inhibition of STAT3 and STAT5 in Companion Animals: Current Status/Future Perspectives
6. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Vail, D.M.; MacEwen, E.G. Spontaneously occurring tumors of companion animals as models for human cancer. Cancer Invest. 2000, 18, 781–792. [Google Scholar] [CrossRef]
- Bonnett, B.N.; Egenvall, A.; Hedhammar, A.; Olson, P. Mortality in over 350,000 insured Swedish dogs from 1995-2000: I. Breed-, gender-, age- and cause-specific rates. Acta Vet. Scand. 2005, 46, 105–120. [Google Scholar] [CrossRef]
- Egenvall, A.; Bonnett, B.N.; Hedhammar, A.; Olson, P. Mortality in over 350,000 insured Swedish dogs from 1995-2000: II. Breed-specific age and survival patterns and relative risk for causes of death. Acta Vet. Scand. 2005, 46, 121–136. [Google Scholar] [CrossRef] [PubMed]
- Vascellari, M.; Baioni, E.; Ru, G.; Carminato, A.; Mutinelli, F. Animal tumour registry of two provinces in northern Italy: Incidence of spontaneous tumours in dogs and cats. BMC Vet. Res. 2009, 5, 39. [Google Scholar] [CrossRef] [PubMed]
- Withrow, S.J.; Vail, D.M. Withrow & MacEwen’s Small Animal Clinical Oncology, 5th ed.; Saunders Elsevier: Philadelphia, PA, USA, 2013. [Google Scholar]
- Paoloni, M.; Khanna, C. Translation of new cancer treatments from pet dogs to humans. Nat. Rev. Cancer 2008, 8, 147–156. [Google Scholar] [CrossRef] [PubMed]
- National Cancer Institute. Surveillance, Epidemiology and End Results (SEER) Program 2012–2016 Data. Available online: www.seer.cancer.gov/statfacts (accessed on 6 November 2019).
- Vascellari, M.; Capello, K.; Carminato, A.; Zanardello, C.; Baioni, E.; Mutinelli, F. Incidence of mammary tumors in the canine population living in the Veneto region (Northeastern Italy): Risk factors and similarities to human breast cancer. Prev. Vet. Med. 2016, 126, 183–189. [Google Scholar] [CrossRef]
- Hassan, B.B.; Elshafae, S.M.; Supsavhad, W.; Simmons, J.K.; Dirksen, W.P.; Sokkar, S.M.; Rosol, T.J. Feline Mammary Cancer. Vet. Pathol. 2017, 54, 32–43. [Google Scholar] [CrossRef]
- Graf, R.; Pospischil, A.; Guscetti, F.; Meier, D.; Welle, M.; Dettwiler, M. Cutaneous tumors in Swiss dogs: Retrospective data from the Swiss canine cancer registry, 2008–2013. Vet. Pathol. 2018, 55, 809–820. [Google Scholar] [CrossRef]
- Merlo, D.F.; Rossi, L.; Pellegrino, C.; Ceppi, M.; Cardellino, U.; Capurro, C.; Ratto, A.; Sambucco, P.L.; Sestito, V.; Tanara, G.; et al. Cancer incidence in pet dogs: Findings of the animal tumor registry of Genoa, Italy. J. Vet. Intern. Med. 2008, 22, 976–984. [Google Scholar] [CrossRef]
- Lomas, A.; Leonardi-Bee, J.; Bath-Hextall, F. A systematic review of worldwide incidence of nonmelanoma skin cancer. Br. J. Dermatol. 2012, 166, 1069–1080. [Google Scholar] [CrossRef]
- Pinello, K.C.; Niza-Ribeiro, J.; Fonseca, L.; de Matos, A.J. Incidence, characteristics and geographical distributions of canine and human non-Hodgkin’s lymphoma in the Porto region (North West Portugal). Vet. J. 2019, 245, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Grover, S. Gastronintestinal lymphoma in cats. Oncol. Compend. 2005, 27, 741–751. [Google Scholar]
- Egenvall, A.; Nodtvedt, A.; von Euler, H. Bone tumors in a population of 400 000 insured Swedish dogs up to 10 y of age: Incidence and survival. Can. J. Vet. Res. 2007, 71, 292–299. [Google Scholar] [PubMed]
- Dimopoulou, M.; Kirpensteijn, J.; Moens, H.; Kik, M. Histologic prognosticators in feline osteosarcoma: A comparison with phenotypically similar canine osteosarcoma. Vet. Surg. 2008, 37, 466–471. [Google Scholar] [CrossRef] [PubMed]
- Cheon, D.J.; Orsulic, S. Mouse models of cancer. Annu. Rev. Pathol 2011, 6, 95–119. [Google Scholar] [CrossRef] [PubMed]
- Ghebranious, N.; Donehower, L.A. Mouse models in tumor suppression. Oncogene 1998, 17, 3385–3400. [Google Scholar] [CrossRef] [PubMed]
- Fowlis, D.J.; Balmain, A. Oncogenes and tumour suppressor genes in transgenic mouse models of neoplasia. Eur. J. Cancer 1993, 29A, 638–645. [Google Scholar] [CrossRef]
- Murphy, W.J. Of mice and men. Biol. Blood Marrow Transplant. 2013, 19, 1140–1141. [Google Scholar] [CrossRef][Green Version]
- Cekanova, M.; Rathore, K. Animal models and therapeutic molecular targets of cancer: Utility and limitations. Drug Des. Devel. Ther. 2014, 8, 1911–1921. [Google Scholar] [CrossRef]
- Stark, G.R.; Darnell, J.E., Jr. The JAK-STAT pathway at twenty. Immunity 2012, 36, 503–514. [Google Scholar] [CrossRef]
- Igelmann, S.; Neubauer, H.A.; Ferbeyre, G. STAT3 and STAT5 Activation in Solid Cancers. Cancers 2019, 11, 1428. [Google Scholar] [CrossRef] [PubMed]
- Logotheti, S.; Putzer, B.M. STAT3 and STAT5 targeting for simultaneous management of melanoma and autoimmune diseases. Cancers 2019, 11, 1448. [Google Scholar] [CrossRef] [PubMed]
- Rebe, C.; Ghiringhelli, F. STAT3, a master regulator of anti-tumor immune response. Cancers 2019, 11, 1280. [Google Scholar] [CrossRef] [PubMed]
- Bromberg, J. Stat proteins and oncogenesis. J. Clin. Invest. 2002, 109, 1139–1142. [Google Scholar] [CrossRef]
- O’Shea, J.J.; Schwartz, D.M.; Villarino, A.V.; Gadina, M.; McInnes, I.B.; Laurence, A. The JAK-STAT pathway: Impact on human disease and therapeutic intervention. Annu. Rev. Med. 2015, 66, 311–328. [Google Scholar] [CrossRef]
- Fagard, R.; Metelev, V.; Souissi, I.; Baran-Marszak, F. STAT3 inhibitors for cancer therapy: Have all roads been explored? JAKSTAT 2013, 2, e22882. [Google Scholar] [CrossRef]
- Rangarajan, A.; Weinberg, R.A. Opinion: Comparative biology of mouse versus human cells: Modelling human cancer in mice. Nat. Rev. Cancer 2003, 3, 952–959. [Google Scholar] [CrossRef]
- Dobrowolski, P.; Fischer, M.; Naumann, R. Novel insights into the genetic background of genetically modified mice. Transgenic Res. 2018, 27, 265–275. [Google Scholar] [CrossRef]
- Kilkenny, C.; Browne, W.J.; Cuthill, I.C.; Emerson, M.; Altman, D.G. Improving bioscience research reporting: The ARRIVE guidelines for reporting animal research. PLoS Biol. 2010, 8, e1000412. [Google Scholar] [CrossRef]
- Bouchlaka, M.N.; Murphy, W.J. Impact of aging in cancer immunotherapy: The importance of using accurate preclinical models. Oncoimmunology 2013, 2, e27186. [Google Scholar] [CrossRef]
- Hooper, L.V.; Littman, D.R.; Macpherson, A.J. Interactions between the microbiota and the immune system. Science 2012, 336, 1268–1273. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, I.I.; Honda, K. Intestinal commensal microbes as immune modulators. Cell Host Microbe 2012, 12, 496–508. [Google Scholar] [CrossRef] [PubMed]
- Sivan, A.; Corrales, L.; Hubert, N.; Williams, J.B.; Aquino-Michaels, K.; Earley, Z.M.; Benyamin, F.W.; Lei, Y.M.; Jabri, B.; Alegre, M.L.; et al. Commensal bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 2015, 350, 1084–1089. [Google Scholar] [CrossRef] [PubMed]
- Antonovic, L.; Martinez, M. Role of the cytochrome P450 enzyme system in veterinary pharmacokinetics: Where are we now? Where are we going? Future Med. Chem. 2011, 3, 855–879. [Google Scholar] [CrossRef] [PubMed]
- Brekke, T.D.; Steele, K.A.; Mulley, J.F. Inbred or outbred? Genetic diversity in laboratory rodent colonies. G3 (Bethesda) 2018, 8, 679–686. [Google Scholar] [CrossRef] [PubMed]
- Cutler, G.; Kassner, P.D. Copy number variation in the mouse genome: Implications for the mouse as a model organism for human disease. Cytogenet. Genome Res. 2008, 123, 297–306. [Google Scholar] [CrossRef]
- Hartman, J.L.t.; Garvik, B.; Hartwell, L. Principles for the buffering of genetic variation. Science 2001, 291, 1001–1004. [Google Scholar] [CrossRef]
- Srivastava, A.; Morgan, A.P.; Najarian, M.L.; Sarsani, V.K.; Sigmon, J.S.; Shorter, J.R.; Kashfeen, A.; McMullan, R.C.; Williams, L.H.; Giusti-Rodriguez, P.; et al. Genomes of the mouse collaborative cross. Genetics 2017, 206, 537–556. [Google Scholar] [CrossRef]
- Dunn, G.P.; Bruce, A.T.; Ikeda, H.; Old, L.J.; Schreiber, R.D. Cancer immunoediting: From immunosurveillance to tumor escape. Nat. Immunol. 2002, 3, 991–998. [Google Scholar] [CrossRef]
- Cespedes, M.V.; Casanova, I.; Parreno, M.; Mangues, R. Mouse models in oncogenesis and cancer therapy. Clin. Transl. Oncol. 2006, 8, 318–329. [Google Scholar] [CrossRef]
- Holzapfel, B.M.; Wagner, F.; Thibaudeau, L.; Levesque, J.P.; Hutmacher, D.W. Concise review: Humanized models of tumor immunology in the 21st century: Convergence of cancer Research and tissue engineering. Stem Cells 2015, 33, 1696–1704. [Google Scholar] [CrossRef] [PubMed]
- Richmond, A.; Su, Y. Mouse xenograft models vs. GEM models for human cancer therapeutics. Dis Model. Mech. 2008, 1, 78–82. [Google Scholar] [CrossRef] [PubMed]
- Wald, A.; Corey, L. Persistence in the population: Epidemiology, transmission. In Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; Cambridge University Press: Cambridge, UK, 2007. [Google Scholar]
- Smatti, M.K.; Al-Sadeq, D.W.; Ali, N.H.; Pintus, G.; Abou-Saleh, H.; Nasrallah, G.K. Epstein-Barr virus epidemiology, serology, and genetic variability of LMP-1 oncogene among healthy population: An update. Front. Oncol. 2018, 8, 211. [Google Scholar] [CrossRef] [PubMed]
- Colugnati, F.A.; Staras, S.A.; Dollard, S.C.; Cannon, M.J. Incidence of cytomegalovirus infection among the general population and pregnant women in the United States. BMC Infect. Dis 2007, 7, 71. [Google Scholar] [CrossRef] [PubMed]
- Day, C.P.; Merlino, G.; Van Dyke, T. Preclinical mouse cancer models: A maze of opportunities and challenges. Cell 2015, 163, 39–53. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Willyard, C. The mice with human tumours: Growing pains for a popular cancer model. Nature 2018, 560, 156–157. [Google Scholar] [CrossRef]
- Ireson, C.R.; Alavijeh, M.S.; Palmer, A.M.; Fowler, E.R.; Jones, H.J. The role of mouse tumour models in the discovery and development of anticancer drugs. Br. J. Cancer 2019, 121, 101–108. [Google Scholar] [CrossRef]
- Ciociola, A.A.; Cohen, L.B.; Kulkarni, P.; Gastroenterology, F.D.-R.M.C.o.t.A.C.o. How drugs are developed and approved by the FDA: Current process and future directions. Am. J. Gastroenterol. 2014, 109, 620–623. [Google Scholar] [CrossRef]
- Schiffman, J.D.; Breen, M. Comparative oncology: What dogs and other species can teach us about humans with cancer. Philos. Trans. R Soc. Lond. B Biol. Sci. 2015, 370. [Google Scholar] [CrossRef]
- Hahn, K.A.; Bravo, L.; Adams, W.H.; Frazier, D.L. Naturally occurring tumors in dogs as comparative models for cancer therapy research. In Vivo 1994, 8, 133–143. [Google Scholar] [PubMed]
- Reif, J.S.; Bruns, C.; Lower, K.S. Cancer of the nasal cavity and paranasal sinuses and exposure to environmental tobacco smoke in pet dogs. Am. J. Epidemiol. 1998, 147, 488–492. [Google Scholar] [CrossRef] [PubMed]
- Reif, J.S.; Dunn, K.; Ogilvie, G.K.; Harris, C.K. Passive smoking and canine lung cancer risk. Am. J. Epidemiol. 1992, 135, 234–239. [Google Scholar] [CrossRef] [PubMed]
- Laflamme, D.; Izquierdo, O.; Eirmann, L.; Binder, S. Myths and misperceptions about ingredients used in commercial pet foods. Vet. Clin. North. Am. Small Anim. Pract. 2014, 44, 689–698. [Google Scholar] [CrossRef]
- Song, S.J.; Lauber, C.; Costello, E.K.; Lozupone, C.A.; Humphrey, G.; Berg-Lyons, D.; Caporaso, J.G.; Knights, D.; Clemente, J.C.; Nakielny, S.; et al. Cohabiting family members share microbiota with one another and with their dogs. Elife 2013, 2, e00458. [Google Scholar] [CrossRef]
- Lindblad-Toh, K.; Wade, C.M.; Mikkelsen, T.S.; Karlsson, E.K.; Jaffe, D.B.; Kamal, M.; Clamp, M.; Chang, J.L.; Kulbokas, E.J., 3rd; Zody, M.C.; et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 2005, 438, 803–819. [Google Scholar] [CrossRef]
- Vonholdt, B.M.; Pollinger, J.P.; Lohmueller, K.E.; Han, E.; Parker, H.G.; Quignon, P.; Degenhardt, J.D.; Boyko, A.R.; Earl, D.A.; Auton, A.; et al. Genome-wide SNP and haplotype analyses reveal a rich history underlying dog domestication. Nature 2010, 464, 898–902. [Google Scholar] [CrossRef]
- Hoffman, M.M.; Birney, E. Estimating the neutral rate of nucleotide substitution using introns. Mol. Biol Evol 2007, 24, 522–531. [Google Scholar] [CrossRef]
- Paoloni, M.C.; Khanna, C. Comparative oncology today. Vet. Clin. North. Am. Small Anim Pract 2007, 37, 1023–1032. [Google Scholar] [CrossRef]
- Khanna, C.; Lindblad-Toh, K.; Vail, D.; London, C.; Bergman, P.; Barber, L.; Breen, M.; Kitchell, B.; McNeil, E.; Modiano, J.F.; et al. The dog as a cancer model. Nat. Biotechnol. 2006, 24, 1065–1066. [Google Scholar] [CrossRef]
- Di Cerbo, A.; Palmieri, B.; De Vico, G.; Iannitti, T. Onco-epidemiology of domestic animals and targeted therapeutic attempts: Perspectives on human oncology. J. Cancer Res. Clin. Oncol 2014, 140, 1807–1814. [Google Scholar] [CrossRef] [PubMed]
- Modiano, J.F.; Breen, M.; Burnett, R.C.; Parker, H.G.; Inusah, S.; Thomas, R.; Avery, P.R.; Lindblad-Toh, K.; Ostrander, E.A.; Cutter, G.C.; et al. Distinct B-cell and T-cell lymphoproliferative disease prevalence among dog breeds indicates heritable risk. Cancer Res. 2005, 65, 5654–5661. [Google Scholar] [CrossRef] [PubMed]
- Nishiya, A.T.; Massoco, C.O.; Felizzola, C.R.; Perlmann, E.; Batschinski, K.; Tedardi, M.V.; Garcia, J.S.; Mendonca, P.P.; Teixeira, T.F.; Zaidan Dagli, M.L. Comparative aspects of canine melanoma. Vet. Sci. 2016, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- Dobson, J.M. Breed-predispositions to cancer in pedigree dogs. ISRN Vet. Sci 2013, 2013, 941275. [Google Scholar] [CrossRef] [PubMed]
- Klopfenstein, M.; Howard, J.; Rossetti, M.; Geissbuhler, U. Life expectancy and causes of death in Bernese mountain dogs in Switzerland. BMC Vet. Res. 2016, 12, 153. [Google Scholar] [CrossRef] [PubMed]
- Lewis, T.W.; Wiles, B.M.; Llewellyn-Zaidi, A.M.; Evans, K.M.; O’Neill, D.G. Longevity and mortality in Kennel Club registered dog breeds in the UK in 2014. Canine Genet. Epidemiol. 2018, 5, 10. [Google Scholar] [CrossRef] [PubMed]
- Dobson, J.; Hoather, T.; McKinley, T.J.; Wood, J.L. Mortality in a cohort of flat-coated retrievers in the UK. Vet. Comp. Oncol. 2009, 7, 115–121. [Google Scholar] [CrossRef]
- Zink, M.C.; Farhoody, P.; Elser, S.E.; Ruffini, L.D.; Gibbons, T.A.; Rieger, R.H. Evaluation of the risk and age of onset of cancer and behavioral disorders in gonadectomized Vizslas. J. Am. Vet. Med. Assoc. 2014, 244, 309–319. [Google Scholar] [CrossRef]
- Henry, C.J.; Higginbotham, H.M. Cancer Management in Small Animal Practice; Saunders Elsevier: Philadelphia, PA, USA, 2009. [Google Scholar]
- Fan, T.M.; Khanna, C. Comparative Aspects of Osteosarcoma Pathogenesis in Humans and Dogs. Vet. Sci 2015, 2, 210–230. [Google Scholar] [CrossRef]
- Urfer, S.R.; Gaillard, C.; Steiger, A. Lifespan and disease predispositions in the Irish Wolfhound: A review. Vet. Q. 2007, 29, 102–111. [Google Scholar] [CrossRef]
- Zandvliet, M. Canine lymphoma: A review. Vet. Q. 2016, 36, 76–104. [Google Scholar] [CrossRef] [PubMed]
- Gajewski, T.F.; Schreiber, H.; Fu, Y.X. Innate and adaptive immune cells in the tumor microenvironment. Nat. Immunol. 2013, 14, 1014–1022. [Google Scholar] [CrossRef] [PubMed]
- Ribas, A. Adaptive immune resistance: How cancer protects from immune attack. Cancer Discov. 2015, 5, 915–919. [Google Scholar] [CrossRef] [PubMed]
- Goulart, M.R.; Pluhar, G.E.; Ohlfest, J.R. Identification of myeloid derived suppressor cells in dogs with naturally occurring cancer. PLoS ONE 2012, 7, e33274. [Google Scholar] [CrossRef]
- Pinheiro, D.; Chang, Y.M.; Bryant, H.; Szladovits, B.; Dalessandri, T.; Davison, L.J.; Yallop, E.; Mills, E.; Leo, C.; Lara, A.; et al. Dissecting the regulatory microenvironment of a large animal model of non-Hodgkin lymphoma: Evidence of a negative prognostic impact of FOXP3+ T cells in canine B cell lymphoma. PLoS ONE 2014, 9, e105027. [Google Scholar] [CrossRef]
- Coy, J.; Caldwell, A.; Chow, L.; Guth, A.; Dow, S. PD-1 expression by canine T cells and functional effects of PD-1 blockade. Vet. Comp. Oncol. 2017, 15, 1487–1502. [Google Scholar] [CrossRef]
- Zerdes, I.; Wallerius, M.; Sifakis, E.G.; Wallmann, T.; Betts, S.; Bartish, M.; Tsesmetzis, N.; Tobin, N.P.; Coucoravas, C.; Bergh, J.; et al. STAT3 activity promotes programmed-death ligand 1 expression and suppresses immune responses in breast cancer. Cancers 2019, 11, 1479. [Google Scholar] [CrossRef]
- Reardon, S. CRISPR gene-editing creates wave of exotic model organisms. Nature 2019, 568, 441–442. [Google Scholar] [CrossRef]
- Simoff, I.; Karlgren, M.; Backlund, M.; Lindstrom, A.C.; Gaugaz, F.Z.; Matsson, P.; Artursson, P. Complete knockout of endogenous Mdr1 (Abcb1) in MDCK cells by CRISPR-Cas9. J. Pharm. Sci. 2016, 105, 1017–1021. [Google Scholar] [CrossRef]
- Amoasii, L.; Hildyard, J.C.W.; Li, H.; Sanchez-Ortiz, E.; Mireault, A.; Caballero, D.; Harron, R.; Stathopoulou, T.R.; Massey, C.; Shelton, J.M.; et al. Gene editing restores dystrophin expression in a canine model of Duchenne muscular dystrophy. Science 2018, 362, 86–91. [Google Scholar] [CrossRef]
- Liongue, C.; Ward, A.C. Evolution of the JAK-STAT pathway. JAKSTAT 2013, 2, e22756. [Google Scholar] [CrossRef] [PubMed]
- Morris, R.; Kershaw, N.J.; Babon, J.J. The molecular details of cytokine signaling via the JAK/STAT pathway. Protein Sci. 2018, 27, 1984–2009. [Google Scholar] [CrossRef] [PubMed]
- Polak, K.L.; Chernosky, N.M.; Smigiel, J.M.; Tamagno, I.; Jackson, M.W. Balancing STAT Activity as a Therapeutic Strategy. Cancers 2019, 11, 1716. [Google Scholar] [CrossRef] [PubMed]
- Yue-Ting, K.L.; Ramaiyer, M.; E. Johnson, D.; R. Grandis, J. Targeting STAT3 in cancer with nucleotide therapeutics. Cancers 2019, 11, 1681. [Google Scholar] [CrossRef]
- Gough, D.J.; Corlett, A.; Schlessinger, K.; Wegrzyn, J.; Larner, A.C.; Levy, D.E. Mitochondrial STAT3 supports Ras-dependent oncogenic transformation. Science 2009, 324, 1713–1716. [Google Scholar] [CrossRef]
- Meier, J.A.; Larner, A.C. Toward a new STATe: The role of STATs in mitochondrial function. Semin. Immunol. 2014, 26, 20–28. [Google Scholar] [CrossRef]
- Frank, D.A. STAT3 as a central mediator of neoplastic cellular transformation. Cancer Lett. 2007, 251, 199–210. [Google Scholar] [CrossRef]
- Bar-Natan, M.; Nelson, E.A.; Xiang, M.; Frank, D.A. STAT signaling in the pathogenesis and treatment of myeloid malignancies. JAKSTAT 2012, 1, 55–64. [Google Scholar] [CrossRef]
- Roeser, J.C.; Leach, S.D.; McAllister, F. Emerging strategies for cancer immunoprevention. Oncogene 2015, 34, 6029–6039. [Google Scholar] [CrossRef]
- Yu, H.; Jove, R. The STATs of cancer--new molecular targets come of age. Nat. Rev. Cancer 2004, 4, 97–105. [Google Scholar] [CrossRef]
- Sen, M.; Thomas, S.M.; Kim, S.; Yeh, J.I.; Ferris, R.L.; Johnson, J.T.; Duvvuri, U.; Lee, J.; Sahu, N.; Joyce, S.; et al. First-in-human trial of a STAT3 decoy oligonucleotide in head and neck tumors: Implications for cancer therapy. Cancer Discov. 2012, 2, 694–705. [Google Scholar] [CrossRef] [PubMed]
- Park, J.S.; Kwok, S.K.; Lim, M.A.; Kim, E.K.; Ryu, J.G.; Kim, S.M.; Oh, H.J.; Ju, J.H.; Park, S.H.; Kim, H.Y.; et al. STA-21, a promising STAT-3 inhibitor that reciprocally regulates Th17 and Treg cells, inhibits osteoclastogenesis in mice and humans and alleviates autoimmune inflammation in an experimental model of rheumatoid arthritis. Arthritis Rheumatol. 2014, 66, 918–929. [Google Scholar] [CrossRef] [PubMed]
- Kunigal, S.; Lakka, S.S.; Sodadasu, P.K.; Estes, N.; Rao, J.S. Stat3-siRNA induces Fas-mediated apoptosis in vitro and in vivo in breast cancer. Int. J. Oncol. 2009, 34, 1209–1220. [Google Scholar] [PubMed]
- Darnell, J.E., Jr. Transcription factors as targets for cancer therapy. Nat. Rev. Cancer 2002, 2, 740–749. [Google Scholar] [CrossRef] [PubMed]
- Ferbeyre, G.; Moriggl, R. The role of Stat5 transcription factors as tumor suppressors or oncogenes. Biochim. Biophys. Acta 2011, 1815, 104–114. [Google Scholar] [CrossRef]
- Zhang, H.F.; Lai, R. STAT3 in Cancer-Friend or Foe? Cancers 2014, 6, 1408–1440. [Google Scholar] [CrossRef]
- Starr, R.; Hilton, D.J. Negative regulation of the JAK/STAT pathway. Bioessays 1999, 21, 47–52. [Google Scholar] [CrossRef]
- Wake, M.S.; Watson, C.J. STAT3 the oncogene - still eluding therapy? FEBS J. 2015, 282, 2600–2611. [Google Scholar] [CrossRef]
- Carmi-Levy, I.; Homey, B.; Soumelis, V. A modular view of cytokine networks in atopic dermatitis. Clin. Rev. Allergy Immunol. 2011, 41, 245–253. [Google Scholar] [CrossRef]
- Brandt, E.B.; Sivaprasad, U. Th2 Cytokines and Atopic Dermatitis. J. Clin. Cell Immunol 2011, 2. [Google Scholar] [CrossRef]
- Ong, P.Y.; Leung, D.Y. Immune dysregulation in atopic dermatitis. Curr. Allergy Asthma Rep. 2006, 6, 384–389. [Google Scholar] [CrossRef] [PubMed]
- Hill, P.B.; Lo, A.; Eden, C.A.; Huntley, S.; Morey, V.; Ramsey, S.; Richardson, C.; Smith, D.J.; Sutton, C.; Taylor, M.D.; et al. Survey of the prevalence, diagnosis and treatment of dermatological conditions in small animals in general practice. Vet. Rec. 2006, 158, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Olivry, T.; Bizikova, P. A systematic review of randomized controlled trials for prevention or treatment of atopic dermatitis in dogs: 2008-2011 update. Vet. Dermatol. 2013, 24, 97–117, e25–e26. [Google Scholar] [CrossRef] [PubMed]
- Marsella, R.; Olivry, T.; Maeda, S. Cellular and cytokine kinetics after epicutaneous allergen challenge (atopy patch testing) with house dust mites in high-IgE beagles. Vet. Dermatol. 2006, 17, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Nuttall, T.J.; Knight, P.A.; McAleese, S.M.; Lamb, J.R.; Hill, P.B. Expression of Th1, Th2 and immunosuppressive cytokine gene transcripts in canine atopic dermatitis. Clin. Exp. Allergy 2002, 32, 789–795. [Google Scholar] [CrossRef] [PubMed]
- Schlotter, Y.M.; Rutten, V.P.; Riemers, F.M.; Knol, E.F.; Willemse, T. Lesional skin in atopic dogs shows a mixed Type-1 and Type-2 immune responsiveness. Vet. Immunol. Immunopathol. 2011, 143, 20–26. [Google Scholar] [CrossRef]
- Gonzales, A.J.; Bowman, J.W.; Fici, G.J.; Zhang, M.; Mann, D.W.; Mitton-Fry, M. Oclacitinib (APOQUEL((R))) is a novel Janus kinase inhibitor with activity against cytokines involved in allergy. J. Vet. Pharmacol. Ther. 2014, 37, 317–324. [Google Scholar] [CrossRef]
- Cosgrove, S.B.; Wren, J.A.; Cleaver, D.M.; Walsh, K.F.; Follis, S.I.; King, V.I.; Tena, J.K.; Stegemann, M.R. A blinded, randomized, placebo-controlled trial of the efficacy and safety of the Janus kinase inhibitor oclacitinib (Apoquel(R)) in client-owned dogs with atopic dermatitis. Vet. Dermatol. 2013, 24, 587–597, e141–e142. [Google Scholar] [CrossRef]
- Cosgrove, S.B.; Wren, J.A.; Cleaver, D.M.; Martin, D.D.; Walsh, K.F.; Harfst, J.A.; Follis, S.L.; King, V.L.; Boucher, J.F.; Stegemann, M.R. Efficacy and safety of oclacitinib for the control of pruritus and associated skin lesions in dogs with canine allergic dermatitis. Vet. Dermatol. 2013, 24, 479-e114. [Google Scholar] [CrossRef]
- Collard, W.T.; Hummel, B.D.; Fielder, A.F.; King, V.L.; Boucher, J.F.; Mullins, M.A.; Malpas, P.B.; Stegemann, M.R. The pharmacokinetics of oclacitinib maleate, a Janus kinase inhibitor, in the dog. J. Vet. Pharmacol Ther 2014, 37, 279–285. [Google Scholar] [CrossRef]
- Lu, Z.; Hong, C.C.; Jark, P.C.; Assumpcao, A.; Bollig, N.; Kong, G.; Pan, X. JAK1/2 Inhibitors AZD1480 and CYT387 inhibit canine B-Cell lymphoma growth by increasing apoptosis and disrupting cell proliferation. J. Vet. Intern. Med. 2017, 31, 1804–1815. [Google Scholar] [CrossRef] [PubMed]
- Assumpcao, A.; Jark, P.C.; Hong, C.C.; Lu, Z.; Ruetten, H.M.; Heaton, C.M.; Pinkerton, M.E.; Pan, X. STAT3 Expression and Activity are Up-Regulated in Diffuse Large B Cell Lymphoma of Dogs. J. Vet. Intern. Med. 2018, 32, 361–369. [Google Scholar] [CrossRef] [PubMed]
- Krol, M.; Pawlowski, K.M.; Dolka, I.; Musielak, O.; Majchrzak, K.; Mucha, J.; Motyl, T. Density of Gr1-positive myeloid precursor cells, p-STAT3 expression and gene expression pattern in canine mammary cancer metastasis. Vet. Res. Commun. 2011, 35, 409–423. [Google Scholar] [CrossRef] [PubMed]
- Van Garderen, E.; Swennenhuis, J.F.; Hellmen, E.; Schalken, J.A. Growth hormone induces tyrosyl phosphorylation of the transcription factors Stat5a and Stat5b in CMT-U335 canine mammary tumor cells. Domest. Anim. Endocrinol. 2001, 20, 123–135. [Google Scholar] [CrossRef]
- Brown, M.E.; Bear, M.D.; Rosol, T.J.; Premanandan, C.; Kisseberth, W.C.; London, C.A. Characterization of STAT3 expression, signaling and inhibition in feline oral squamous cell carcinoma. BMC Vet. Res. 2015, 11, 206. [Google Scholar] [CrossRef]
- Petterino, C.; Podesta, G.; Ratto, A.; Drigo, M.; Pellegrino, C. Immunohistochemical study of phospho-Stat3-ser727 expression in feline mammary gland tumours. Vet. Res. Commun. 2007, 31, 173–184. [Google Scholar] [CrossRef]
- Petterino, C.; Ratto, A.; Podesta, G.; Drigo, M.; Pellegrino, C. Immunohistochemical evaluation of STAT3-p-tyr705 expression in feline mammary gland tumours and correlation with histologic grade. Res. Vet. Sci. 2007, 82, 218–224. [Google Scholar] [CrossRef]
- Willmann, M.; Hadzijusufovic, E.; Hermine, O.; Dacasto, M.; Marconato, L.; Bauer, K.; Peter, B.; Gamperl, S.; Eisenwort, G.; Jensen-Jarolim, E.; et al. Comparative oncology: The paradigmatic example of canine and human mast cell neoplasms. Vet. Comp. Oncol. 2019, 17, 1–10. [Google Scholar] [CrossRef]
- Makielski, K.M.; Mills, L.J.; Sarver, A.L.; Henson, M.S.; Spector, L.G.; Naik, S.; Modiano, J.F. Risk factors for development of canine and human osteosarcoma: A comparative review. Vet. Sci. 2019, 6, 48. [Google Scholar] [CrossRef]
- Rowell, J.L.; McCarthy, D.O.; Alvarez, C.E. Dog models of naturally occurring cancer. Trends Mol. Med. 2011, 17, 380–388. [Google Scholar] [CrossRef]
- Mirabello, L.; Troisi, R.J.; Savage, S.A. Osteosarcoma incidence and survival rates from 1973 to 2004: Data from the surveillance, epidemiology, and end results program. Cancer 2009, 115, 1531–1543. [Google Scholar] [CrossRef] [PubMed]
- Fenger, J.M.; London, C.A.; Kisseberth, W.C. Canine osteosarcoma: A naturally occurring disease to inform pediatric oncology. ILAR J. 2014, 55, 69–85. [Google Scholar] [CrossRef] [PubMed]
- Fossey, S.L.; Liao, A.T.; McCleese, J.K.; Bear, M.D.; Lin, J.; Li, P.K.; Kisseberth, W.C.; London, C.A. Characterization of STAT3 activation and expression in canine and human osteosarcoma. BMC Cancer 2009, 9, 81. [Google Scholar] [CrossRef] [PubMed]
- Fossey, S.L.; Bear, M.D.; Lin, J.; Li, C.; Schwartz, E.B.; Li, P.K.; Fuchs, J.R.; Fenger, J.; Kisseberth, W.C.; London, C.A. The novel curcumin analog FLLL32 decreases STAT3 DNA binding activity and expression, and induces apoptosis in osteosarcoma cell lines. BMC Cancer 2011, 11, 112. [Google Scholar] [CrossRef] [PubMed]
- Couto, J.I.; Bear, M.D.; Lin, J.; Pennel, M.; Kulp, S.K.; Kisseberth, W.C.; London, C.A. Biologic activity of the novel small molecule STAT3 inhibitor LLL12 against canine osteosarcoma cell lines. BMC Vet. Res. 2012, 8, 244. [Google Scholar] [CrossRef] [PubMed]
- Onimoe, G.I.; Liu, A.; Lin, L.; Wei, C.C.; Schwartz, E.B.; Bhasin, D.; Li, C.; Fuchs, J.R.; Li, P.K.; Houghton, P.; et al. Small molecules, LLL12 and FLLL32, inhibit STAT3 and exhibit potent growth suppressive activity in osteosarcoma cells and tumor growth in mice. Invest. New Drugs 2012, 30, 916–926. [Google Scholar] [CrossRef]
- Paoloni, M.; Davis, S.; Lana, S.; Withrow, S.; Sangiorgi, L.; Picci, P.; Hewitt, S.; Triche, T.; Meltzer, P.; Khanna, C. Canine tumor cross-species genomics uncovers targets linked to osteosarcoma progression. BMC Genomics 2009, 10, 625. [Google Scholar] [CrossRef]
- Rodriguez, C.O., Jr. Using canine osteosarcoma as a model to assess efficacy of novel therapies: Can old dogs teach us new tricks? Adv. Exp. Med. Biol. 2014, 804, 237–256. [Google Scholar] [CrossRef]
- London, C.A.; Seguin, B. Mast cell tumors in the dog. Vet. Clin. North. Am. Small Anim. Pract. 2003, 33, 473–489. [Google Scholar] [CrossRef]
- Misdorp, W. Mast cells and canine mast cell tumours. A review. Vet. Q. 2004, 26, 156–169. [Google Scholar] [CrossRef]
- Shoop, S.J.; Marlow, S.; Church, D.B.; English, K.; McGreevy, P.D.; Stell, A.J.; Thomson, P.C.; O’Neill, D.G.; Brodbelt, D.C. Prevalence and risk factors for mast cell tumours in dogs in England. Canine Genet. Epidemiol. 2015, 2, 1. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.H.; Tefferi, A.; Lasho, T.L.; Finke, C.; Patnaik, M.; Butterfield, J.H.; McClure, R.F.; Li, C.Y.; Pardanani, A. Systemic mastocytosis in 342 consecutive adults: Survival studies and prognostic factors. Blood 2009, 113, 5727–5736. [Google Scholar] [CrossRef] [PubMed]
- Letard, S.; Yang, Y.; Hanssens, K.; Palmerini, F.; Leventhal, P.S.; Guery, S.; Moussy, A.; Kinet, J.P.; Hermine, O.; Dubreuil, P. Gain-of-function mutations in the extracellular domain of KIT are common in canine mast cell tumors. Mol. Cancer Res. 2008, 6, 1137–1145. [Google Scholar] [CrossRef] [PubMed]
- London, C.A.; Galli, S.J.; Yuuki, T.; Hu, Z.Q.; Helfand, S.C.; Geissler, E.N. Spontaneous canine mast cell tumors express tandem duplications in the proto-oncogene c-kit. Exp. Hematol. 1999, 27, 689–697. [Google Scholar] [CrossRef]
- Nagata, H.; Worobec, A.S.; Oh, C.K.; Chowdhury, B.A.; Tannenbaum, S.; Suzuki, Y.; Metcalfe, D.D. Identification of a point mutation in the catalytic domain of the protooncogene c-kit in peripheral blood mononuclear cells of patients who have mastocytosis with an associated hematologic disorder. Proc. Natl. Acad. Sci. USA 1995, 92, 10560–10564. [Google Scholar] [CrossRef]
- Metcalfe, D.D. Mast cells and mastocytosis. Blood 2008, 112, 946–956. [Google Scholar] [CrossRef]
- Iemura, A.; Tsai, M.; Ando, A.; Wershil, B.K.; Galli, S.J. The c-kit ligand, stem cell factor, promotes mast cell survival by suppressing apoptosis. Am. J. Pathol. 1994, 144, 321–328. [Google Scholar]
- Moller, C.; Alfredsson, J.; Engstrom, M.; Wootz, H.; Xiang, Z.; Lennartsson, J.; Jonsson, J.I.; Nilsson, G. Stem cell factor promotes mast cell survival via inactivation of FOXO3a-mediated transcriptional induction and MEK-regulated phosphorylation of the proapoptotic protein Bim. Blood 2005, 106, 1330–1336. [Google Scholar] [CrossRef]
- Hahn, K.A.; Ogilvie, G.; Rusk, T.; Devauchelle, P.; Leblanc, A.; Legendre, A.; Powers, B.; Leventhal, P.S.; Kinet, J.P.; Palmerini, F.; et al. Masitinib is safe and effective for the treatment of canine mast cell tumors. J. Vet. Intern. Med. 2008, 22, 1301–1309. [Google Scholar] [CrossRef]
- London, C.A.; Malpas, P.B.; Wood-Follis, S.L.; Boucher, J.F.; Rusk, A.W.; Rosenberg, M.P.; Henry, C.J.; Mitchener, K.L.; Klein, M.K.; Hintermeister, J.G.; et al. Multi-center, placebo-controlled, double-blind, randomized study of oral toceranib phosphate (SU11654), a receptor tyrosine kinase inhibitor, for the treatment of dogs with recurrent (either local or distant) mast cell tumor following surgical excision. Clin. Cancer Res. 2009, 15, 3856–3865. [Google Scholar] [CrossRef]
- Chaix, A.; Lopez, S.; Voisset, E.; Gros, L.; Dubreuil, P.; De Sepulveda, P. Mechanisms of STAT protein activation by oncogenic KIT mutants in neoplastic mast cells. J. Biol. Chem. 2011, 286, 5956–5966. [Google Scholar] [CrossRef] [PubMed]
- Baumgartner, C.; Cerny-Reiterer, S.; Sonneck, K.; Mayerhofer, M.; Gleixner, K.V.; Fritz, R.; Kerenyi, M.; Boudot, C.; Gouilleux, F.; Kornfeld, J.W.; et al. Expression of activated STAT5 in neoplastic mast cells in systemic mastocytosis: Subcellular distribution and role of the transforming oncoprotein KIT D816V. Am. J. Pathol. 2009, 175, 2416–2429. [Google Scholar] [CrossRef] [PubMed]
- Keller, A.; Wingelhofer, B.; Peter, B.; Bauer, K.; Berger, D.; Gamperl, S.; Reifinger, M.; Cerny-Reiterer, S.; Moriggl, R.; Willmann, M.; et al. The JAK2/STAT5 signaling pathway as a potential therapeutic target in canine mastocytoma. Vet. Comp. Oncol. 2018, 16, 55–68. [Google Scholar] [CrossRef] [PubMed]
- Peter, B.; Bibi, S.; Eisenwort, G.; Wingelhofer, B.; Berger, D.; Stefanzl, G.; Blatt, K.; Herrmann, H.; Hadzijusufovic, E.; Hoermann, G.; et al. Drug-induced inhibition of phosphorylation of STAT5 overrides drug resistance in neoplastic mast cells. Leukemia 2018, 32, 1016–1022. [Google Scholar] [CrossRef] [PubMed]
- Page, B.D.; Khoury, H.; Laister, R.C.; Fletcher, S.; Vellozo, M.; Manzoli, A.; Yue, P.; Turkson, J.; Minden, M.D.; Gunning, P.T. Small molecule STAT5-SH2 domain inhibitors exhibit potent antileukemia activity. J. Med. Chem. 2012, 55, 1047–1055. [Google Scholar] [CrossRef]
- Park, J.S.; Withers, S.S.; Modiano, J.F.; Kent, M.S.; Chen, M.; Luna, J.I.; Culp, W.T.N.; Sparger, E.E.; Rebhun, R.B.; Monjazeb, A.M.; et al. Canine cancer immunotherapy studies: Linking mouse and human. J. Immunother. Cancer 2016, 4, 97. [Google Scholar] [CrossRef]
- Alvarez, C.E. Naturally occurring cancers in dogs: Insights for translational genetics and medicine. ILAR J. 2014, 55, 16–45. [Google Scholar] [CrossRef]
- Furdos, I.; Fazekas, J.; Singer, J.; Jensen-Jarolim, E. Translating clinical trials from human to veterinary oncology and back. J. Transl. Med. 2015, 13, 265. [Google Scholar] [CrossRef]
- Riccardo, F.; Aurisicchio, L.; Impellizeri, J.A.; Cavallo, F. The importance of comparative oncology in translational medicine. Cancer Immunol. Immunother. 2015, 64, 137–148. [Google Scholar] [CrossRef]

| Tissue | Human | Dog | Cat |
|---|---|---|---|
| Mammary | 127.5 [7] | 250 [8] | 13–25 [9] |
| Melanoma | 22.2 [7] | 19.8 [10] | ND |
| Testes | 5.9 [7] | 16.7 [11] | ND |
| Connective Tissue | 3.5 [7] | 40.1 [10] | 17.0 [1] |
| Skin | 98.85 [12] | 103.3 [10] | 34.7 [1] |
| Oral | 11.3 [7] | 20.4 [1] | 11.6 [1] |
| NHL/Leukemia | 33.7 [7] | 76.3 [13] | 41 [14] |
| Bone | 1.0 [7] | 27.2 [15] | 3.1–4.9 [16] |
| Breeds | Most Frequent Tumor Types |
|---|---|
| Bernese Mountain Dog | Histiocytic sarcoma [68], Lymphoma [68,69], Osteosarcoma [68] |
| Boxer | Glioma [67,69], Mast cell tumor [10,67] |
| Flat-Coated Retriever | Soft tissue sarcoma, Histiocytic sarcoma, Hemangiosarcoma [70] |
| Golden Retriever | Mast cell tumor, Lymphoma, Oral Melanoma, Fibrosarcoma [67] |
| Magyar Viszla | Mast cell tumor, Hemangiosarcoma, Lymphoma [10,71] |
| Giant Schnauzer | Epidermal tumor, Hair follicle tumor, Melanocytic tumor [10] |
| Airedale Terrier | Melanoma [72], Lymphoma [67], Prostatic carcinoma [67] |
| Bullmastiff | Mast cell tumor, Lymphoma [67] |
| St. Bernard | Lymphoma [67], Osteosarcoma [73] |
| Irish Wolfhound | Osteosarcoma [67,74], Lymphoma [75] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kieslinger, M.; Swoboda, A.; Kramer, N.; Pratscher, B.; Wolfesberger, B.; Burgener, I.A. Companion Animals as Models for Inhibition of STAT3 and STAT5. Cancers 2019, 11, 2035. https://doi.org/10.3390/cancers11122035
Kieslinger M, Swoboda A, Kramer N, Pratscher B, Wolfesberger B, Burgener IA. Companion Animals as Models for Inhibition of STAT3 and STAT5. Cancers. 2019; 11(12):2035. https://doi.org/10.3390/cancers11122035
Chicago/Turabian StyleKieslinger, Matthias, Alexander Swoboda, Nina Kramer, Barbara Pratscher, Birgitt Wolfesberger, and Iwan A. Burgener. 2019. "Companion Animals as Models for Inhibition of STAT3 and STAT5" Cancers 11, no. 12: 2035. https://doi.org/10.3390/cancers11122035
APA StyleKieslinger, M., Swoboda, A., Kramer, N., Pratscher, B., Wolfesberger, B., & Burgener, I. A. (2019). Companion Animals as Models for Inhibition of STAT3 and STAT5. Cancers, 11(12), 2035. https://doi.org/10.3390/cancers11122035





