Poly(ADP-Ribose) Polymerase Inhibitors in Pancreatic Cancer: A New Treatment Paradigms and Future Implications
Abstract
1. Introduction
2. Mechanism of Action of PARP Inhibitors
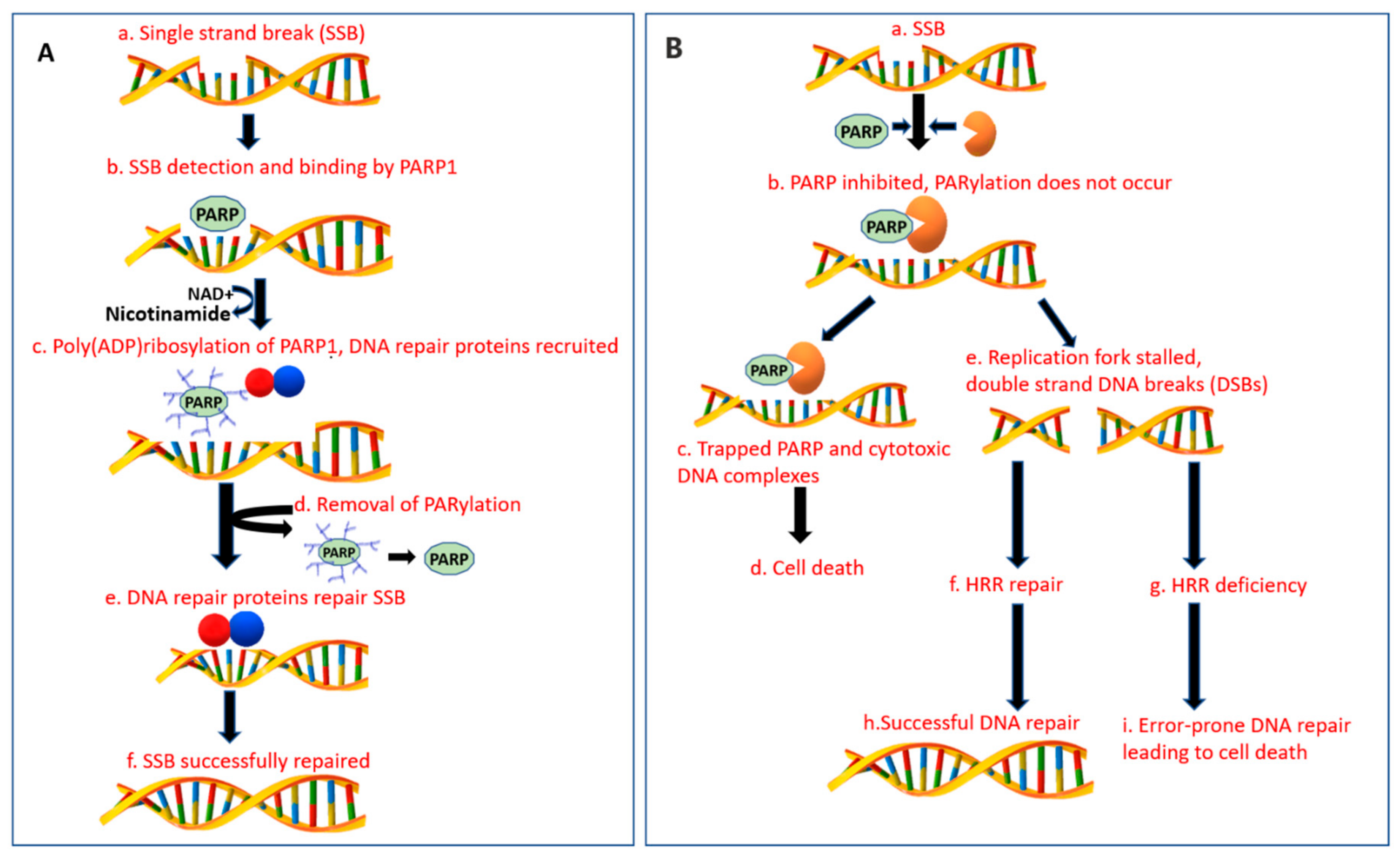
2.1. Repair Mechanisms in Healthy Cells
2.2. BRCA Deficiency
2.3. DNA Repair with BRCA-Deficient Cells in the Presence of PARPi
2.4. PARPi Pharmacology
2.5. Clinical Development of PARPi in PDAC
2.6. PARPi as Monotherapy in Advanced Disease
2.7. PARPi as Maintenance Therapy in Platinum-Sensitive Disease
3. PARPi in Combination with Other Therapies
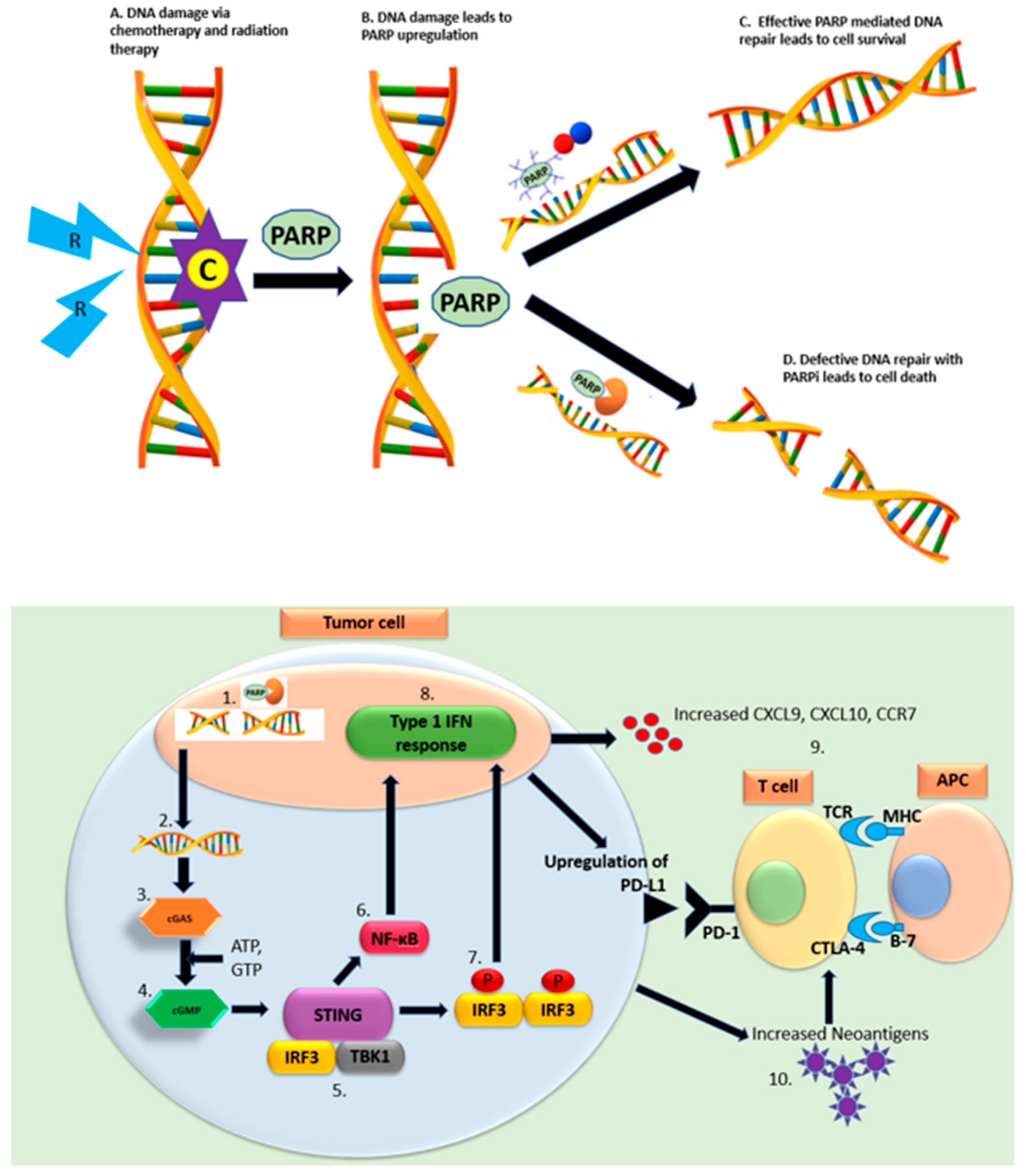
3.1. PARP in Combination with Chemotherapy
3.2. PARP in Combination with Radiotherapy
3.3. PARPi in Combination with Immunotherapy
3.4. Molecular Targeted Therapy Combinations
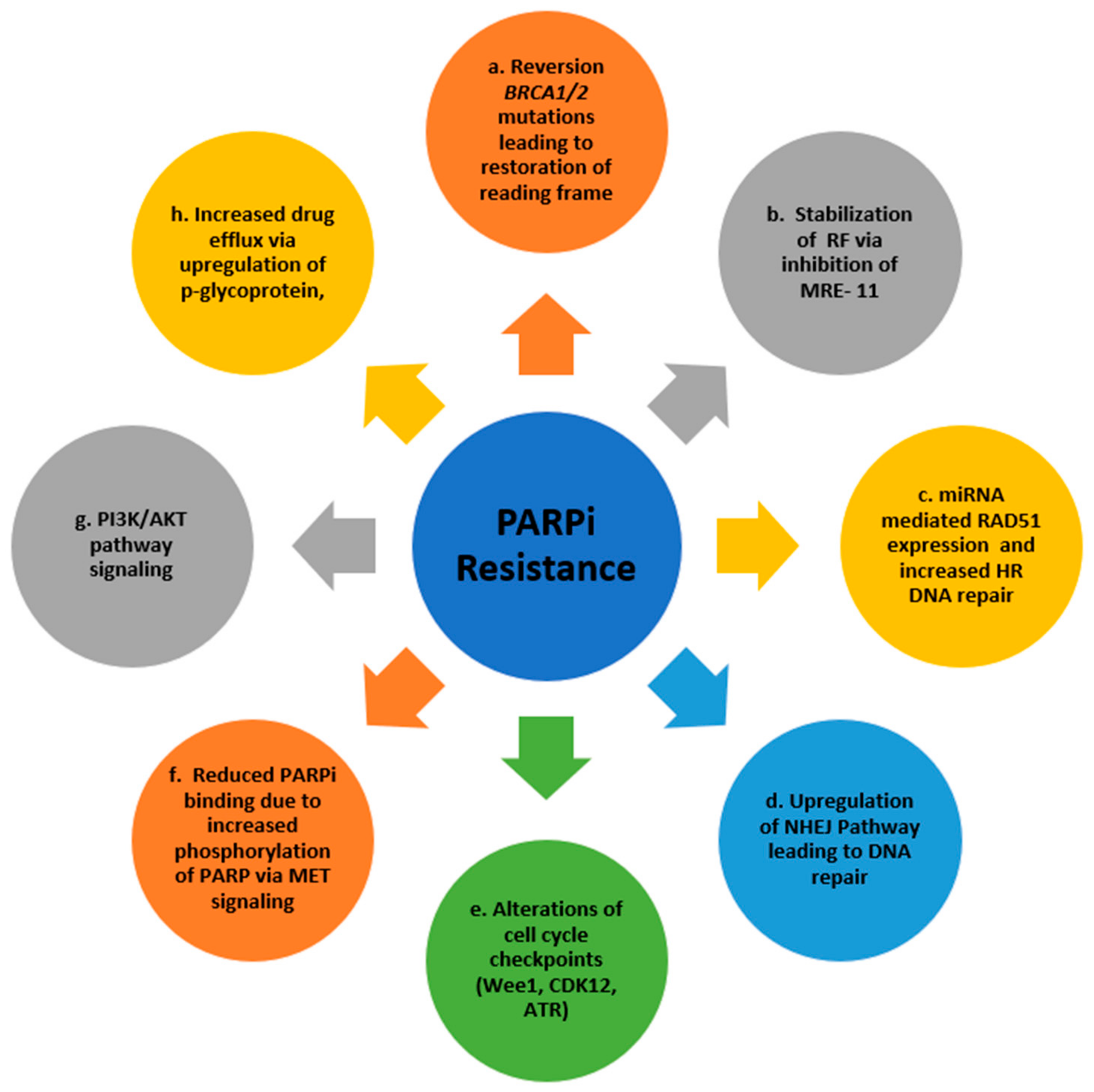
4. Resistance Mechanisms
5. Discussion
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2019. CA Cancer J. Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef] [PubMed]
- Rawla, P.; Sunkara, T.; Gaduputi, V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J. Oncol. 2019, 10, 10–27. [Google Scholar] [CrossRef] [PubMed]
- Conroy, T.; Desseigne, F.; Ychou, M.; Bouche, O.; Guimbaud, R.; Becouarn, Y.; Adenis, A.; Raoul, J.L.; Gourgou-Bourgade, S.; de la Fouchardiere, C.; et al. FOLFIRINOX versus gemcitabine for metastatic pancreatic cancer. N. Engl. J. Med. 2011, 364, 1817–1825. [Google Scholar] [CrossRef] [PubMed]
- Neuzillet, C.; Hentic, O.; Rousseau, B.; Rebours, V.; Bengrine-Lefevre, L.; Bonnetain, F.; Levy, P.; Raymond, E.; Ruszniewski, P.; Louvet, C.; et al. FOLFIRI regimen in metastatic pancreatic adenocarcinoma resistant to gemcitabine and platinum-salts. World J. Gastroenterol. 2012, 18, 4533–4541. [Google Scholar] [CrossRef] [PubMed]
- Von Hoff, D.D.; Ervin, T.; Arena, F.P.; Chiorean, E.G.; Infante, J.; Moore, M.; Seay, T.; Tjulandin, S.A.; Ma, W.W.; Saleh, M.N.; et al. Increased survival in pancreatic cancer with nab-paclitaxel plus gemcitabine. N. Engl. J. Med. 2013, 369, 1691–1703. [Google Scholar] [CrossRef]
- Wang-Gillam, A.; Li, C.P.; Bodoky, G.; Dean, A.; Shan, Y.S.; Jameson, G.; Macarulla, T.; Lee, K.H.; Cunningham, D.; Blanc, J.F.; et al. Nanoliposomal irinotecan with fluorouracil and folinic acid in metastatic pancreatic cancer after previous gemcitabine-based therapy (NAPOLI-1): A global, randomised, open-label, phase 3 trial. Lancet 2016, 387, 545–557. [Google Scholar] [CrossRef]
- Yoo, C.; Hwang, J.Y.; Kim, J.E.; Kim, T.W.; Lee, J.S.; Park, D.H.; Lee, S.S.; Seo, D.W.; Lee, S.K.; Kim, M.H.; et al. A randomised phase II study of modified FOLFIRI.3 vs. modified FOLFOX as second-line therapy in patients with gemcitabine-refractory advanced pancreatic cancer. Br. J. Cancer 2009, 101, 1658–1663. [Google Scholar] [CrossRef]
- Research C for DE and FDA Grants Accelerated Approval to Pembrolizumab for First Tissue/Site Agnostic Indication. FDA. 9 February 2019. Available online: http://www.fda.gov/drugs/resources-information-approved-drugs/fda-grants-accelerated-approval-pembrolizumab-first-tissuesite-agnostic-indication (accessed on 17 August 2019).
- Research C for DE and FDA Approves Larotrectinib for Solid Tumors with NTRK Gene Fusions. FDA. 9 February 2019. Available online: http://www.fda.gov/drugs/fda-approves-larotrectinib-solid-tumors-ntrk-gene-fusions-0 (accessed on 17 August 2019).
- Jones, S.; Hruban, R.H.; Kamiyama, M.; Borges, M.; Zhang, X.; Parsons, D.W.; Lin, J.C.; Palmisano, E.; Brune, K.; Jaffee, E.M.; et al. Exomic sequencing identifies PALB2 as a pancreatic cancer susceptibility gene. Science 2009, 324, e217. [Google Scholar] [CrossRef]
- McWilliams, R.R.; Wieben, E.D.; Rabe, K.G.; Pedersen, K.S.; Wu, Y.; Sicotte, H.; Petersen, G.M. Prevalence of CDKN2A mutations in pancreatic cancer patients: Implications for genetic counseling. Eur. J. Hum. Genet. 2011, 19, 472–478. [Google Scholar] [CrossRef]
- Su, G.H.; Hruban, R.H.; Bansal, R.K.; Bova, G.S.; Tang, D.J.; Shekher, M.C.; Westerman, A.M.; Entius, M.M.; Goggins, M.; Yeo, C.J.; et al. Germline and somatic mutations of the STK11/LKB1 Peutz-Jeghers gene in pancreatic and biliary cancers. Am. J. Pathol. 1999, 154, 1835–1840. [Google Scholar] [CrossRef]
- Tersmette, A.C.; Petersen, G.M.; Offerhaus, G.J.; Falatko, F.C.; Brune, K.A.; Goggins, M.; Rozenblum, E.; Wilentz, R.E.; Yeo, C.J.; Cameron, J.L.; et al. Increased risk of incident pancreatic cancer among first-degree relatives of patients with familial pancreatic cancer. Clin. Cancer Res. 2001, 7, 738–744. [Google Scholar] [PubMed]
- Holter, S.; Borgida, A.; Dodd, A.; Grant, R.; Semotiuk, K.; Hedley, D.; Dhani, N.; Narod, S.; Akbari, M.; Moore, M.; et al. Germline BRCA Mutations in a Large Clinic-Based Cohort of Patients With Pancreatic Adenocarcinoma. J. Clin. Oncol. 2015, 33, 3124–3129. [Google Scholar] [CrossRef] [PubMed]
- Salo-Mullen, E.E.; O’Reilly, E.M.; Kelsen, D.P.; Ashraf, A.M.; Lowery, M.A.; Yu, K.H.; Reidy, D.L.; Epstein, A.S.; Lincoln, A.; Saldia, A.; et al. Identification of germline genetic mutations in patients with pancreatic cancer. Cancer 2015, 121, 4382–4388. [Google Scholar] [CrossRef] [PubMed]
- Waddell, N.; Pajic, M.; Patch, A.M.; Chang, D.K.; Kassahn, K.S.; Bailey, P.; Johns, A.L.; Miller, D.; Nones, K.; Quek, K.; et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 2015, 518, 495–501. [Google Scholar] [CrossRef]
- Dedes, K.J.; Wilkerson, P.M.; Wetterskog, D.; Weigelt, B.; Ashworth, A.; Reis-Filho, J.S. Synthetic lethality of PARP inhibition in cancers lacking BRCA1 and BRCA2 mutations. Cell Cycle 2011, 10, 1192–1199. [Google Scholar] [CrossRef]
- Schreiber, V.; Dantzer, F.; Ame, J.C.; de Murcia, G. Poly(ADP-ribose): Novel functions for an old molecule. Nat. Rev. Mol. Cell Biol. 2006, 7, 517–528. [Google Scholar] [CrossRef]
- Moynahan, M.E.; Jasin, M. Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nat. Rev. Mol. Cell Biol. 2010, 11, 196–207. [Google Scholar] [CrossRef]
- Radhakrishnan, S.K.; Jette, N.; Lees-Miller, S.P. Non-homologous end joining: Emerging themes and unanswered questions. DNA Repair (Amst.) 2014, 17, 2–8. [Google Scholar] [CrossRef]
- Durkacz, B.W.; Omidiji, O.; Gray, D.A.; Shall, S. (ADP-ribose)n participates in DNA excision repair. Nature 1980, 283, 593–596. [Google Scholar] [CrossRef]
- Krishnakumar, R.; Kraus, W.L. The PARP side of the nucleus: Molecular actions, physiological outcomes, and clinical targets. Mol. Cell 2010, 39, 8–24. [Google Scholar] [CrossRef]
- Bai, P. Biology of Poly(ADP-Ribose) Polymerases: The Factotums of Cell Maintenance. Mol. Cell 2015, 58, 947–958. [Google Scholar] [CrossRef] [PubMed]
- El-Khamisy, S.F.; Masutani, M.; Suzuki, H.; Caldecott, K.W. A requirement for PARP-1 for the assembly or stability of XRCC1 nuclear foci at sites of oxidative DNA damage. Nucleic Acids Res. 2003, 31, 5526–5533. [Google Scholar] [CrossRef] [PubMed]
- Masson, M.; Niedergang, C.; Schreiber, V.; Muller, S.; Menissier-de Murcia, J.; de Murcia, G. XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage. Mol. Cell Biol. 1998, 18, 3563–3571. [Google Scholar] [CrossRef] [PubMed]
- Ahel, D.; Horejsi, Z.; Wiechens, N.; Polo, S.E.; Garcia-Wilson, E.; Ahel, I.; Flynn, H.; Skehel, M.; West, S.C.; Jackson, S.P.; et al. Poly(ADP-ribose)-dependent regulation of DNA repair by the chromatin remodeling enzyme ALC1. Science 2009, 325, 1240–1243. [Google Scholar] [CrossRef]
- Gagne, J.P.; Isabelle, M.; Lo, K.S.; Bourassa, S.; Hendzel, M.J.; Dawson, V.L.; Dawson, T.M.; Poirier, G.G. Proteome-wide identification of poly(ADP-ribose) binding proteins and poly(ADP-ribose)-associated protein complexes. Nucleic Acids Res. 2008, 36, 6959–6976. [Google Scholar] [CrossRef]
- Oka, S.; Kato, J.; Moss, J. Identification and characterization of a mammalian 39-kDa poly(ADP-ribose) glycohydrolase. J. Biol. Chem. 2006, 281, 705–713. [Google Scholar] [CrossRef]
- Satoh, M.S.; Lindahl, T. Role of poly(ADP-ribose) formation in DNA repair. Nature 1992, 356, 356–358. [Google Scholar] [CrossRef]
- Gudmundsdottir, K.; Ashworth, A. The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene 2006, 25, 5864–5874. [Google Scholar] [CrossRef]
- Lord, C.J.; Ashworth, A. RAD51, BRCA2 and DNA repair: A partial resolution. Nat. Struct. Mol. Biol. 2007, 14, 461–462. [Google Scholar] [CrossRef]
- Venkitaraman, A.R. Cancer susceptibility and the functions of BRCA1 and BRCA2. Cell 2002, 108, 171–182. [Google Scholar] [CrossRef]
- Lieber, M.R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 2010, 79, 181–211. [Google Scholar] [CrossRef] [PubMed]
- Haince, J.F.; Kozlov, S.; Dawson, V.L.; Dawson, T.M.; Hendzel, M.J.; Lavin, M.F.; Poirier, G.G. Ataxia telangiectasia mutated (ATM) signaling network is modulated by a novel poly(ADP-ribose)-dependent pathway in the early response to DNA-damaging agents. J. Biol. Chem. 2007, 282, 16441–16453. [Google Scholar] [CrossRef] [PubMed]
- Haince, J.F.; McDonald, D.; Rodrigue, A.; Dery, U.; Masson, J.Y.; Hendzel, M.J.; Poirier, G.G. PARP1-dependent kinetics of recruitment of MRE11 and NBS1 proteins to multiple DNA damage sites. J. Biol. Chem. 2008, 283, 1197–1208. [Google Scholar] [CrossRef] [PubMed]
- Ray Chaudhuri, A.; Callen, E.; Ding, X.; Gogola, E.; Duarte, A.A.; Lee, J.E.; Wong, N.; Lafarga, V.; Calvo, J.A.; Panzarino, N.J.; et al. Replication fork stability confers chemoresistance in BRCA-deficient cells. Nature 2016, 535, 382–387. [Google Scholar] [CrossRef] [PubMed]
- Ferrone, C.R.; Levine, D.A.; Tang, L.H.; Allen, P.J.; Jarnagin, W.; Brennan, M.F.; Offit, K.; Robson, M.E. BRCA germline mutations in Jewish patients with pancreatic adenocarcinoma. J. Clin. Oncol. 2009, 27, 433–438. [Google Scholar] [CrossRef] [PubMed]
- Friedenson, B. BRCA1 and BRCA2 pathways and the risk of cancers other than breast or ovarian. Medscape Gen. Med. 2005, 7, e60. [Google Scholar]
- Golan, T.; Kanji, Z.S.; Epelbaum, R.; Devaud, N.; Dagan, E.; Holter, S.; Aderka, D.; Paluch-Shimon, S.; Kaufman, B.; Gershoni-Baruch, R.; et al. Overall survival and clinical characteristics of pancreatic cancer in BRCA mutation carriers. Br. J. Cancer 2014, 111, 1132–1138. [Google Scholar] [CrossRef]
- Dobzhansky, T. Genetics of natural populations; recombination and variability in populations of Drosophila pseudoobscura. Genetics 1946, 31, 269–290. [Google Scholar]
- Ashworth, A. A synthetic lethal therapeutic approach: Poly(ADP) ribose polymerase inhibitors for the treatment of cancers deficient in DNA double-strand break repair. J. Clin. Oncol. 2008, 26, 3785–3790. [Google Scholar] [CrossRef]
- Murai, J.; Huang, S.Y.; Das, B.B.; Renaud, A.; Zhang, Y.; Doroshow, J.H.; Ji, J.; Takeda, S.; Pommier, Y. Trapping of PARP1 and PARP2 by Clinical PARP Inhibitors. Cancer Res. 2012, 72, 5588–5599. [Google Scholar] [CrossRef]
- D’Andrea, A.D. Susceptibility pathways in Fanconi’s anemia and breast cancer. N. Engl. J. Med. 2010, 362, 1909–1919. [Google Scholar] [CrossRef] [PubMed]
- McCabe, N.; Turner, N.C.; Lord, C.J.; Kluzek, K.; Bialkowska, A.; Swift, S.; Giavara, S.; O’Connor, M.J.; Tutt, A.N.; Zdzienicka, M.Z.; et al. Deficiency in the repair of DNA damage by homologous recombination and sensitivity to poly(ADP-ribose) polymerase inhibition. Cancer Res. 2006, 66, 8109–8115. [Google Scholar] [CrossRef] [PubMed]
- Turner, N.; Tutt, A.; Ashworth, A. Hallmarks of ‘BRCAness’ in sporadic cancers. Nat. Rev. Cancer 2004, 4, 814–819. [Google Scholar] [CrossRef] [PubMed]
- Moschetta, M.; George, A.; Kaye, S.B.; Banerjee, S. BRCA somatic mutations and epigenetic BRCA modifications in serous ovarian cancer. Ann. Oncol. 2016, 27, 1449–1455. [Google Scholar] [CrossRef]
- Carnevale, J.; Ashworth, A. Assessing the Significance of BRCA1 and BRCA2 Mutations in Pancreatic Cancer. J. Clin. Oncol. 2015, 33, 3080–3081. [Google Scholar] [CrossRef]
- Telli, M.L.; Timms, K.M.; Reid, J.; Hennessy, B.; Mills, G.B.; Jensen, K.C.; Szallasi, Z.; Barry, W.T.; Winer, E.P.; Tung, N.M.; et al. Homologous Recombination Deficiency (HRD) Score Predicts Response to Platinum-Containing Neoadjuvant Chemotherapy in Patients with Triple-Negative Breast Cancer. Clin. Cancer Res. 2016, 22, 3764–3773. [Google Scholar] [CrossRef]
- Shahda, S.; Timms, K.M.; Ibrahim, A.A.; Reid, J.E.; Cramer, H.M.; Radovich, M.; Ibrahim, S.; Allen, B.; O’Neil, B.H. Homologous Recombination Deficiency in Patients With Pancreatic Ductal Adenocarcinoma and Response to Chemotherapy. JCO Precis. Oncol. 2018, 2, 1–11. [Google Scholar] [CrossRef]
- Steffen, J.; Brody, J.; Armen, R.; Pascal, J. Structural Implications for Selective Targeting of PARPs. Front. Oncol. 2013, 3, e301. [Google Scholar] [CrossRef]
- Perkins, E.; Sun, D.; Nguyen, A.; Tulac, S.; Francesco, M.; Tavana, H.; Nguyen, H.; Tugendreich, S.; Barthmaier, P.; Couto, J.; et al. Novel inhibitors of poly(ADP-ribose) polymerase/PARP1 and PARP2 identified using a cell-based screen in yeast. Cancer Res. 2001, 61, 4175–4183. [Google Scholar]
- Murai, J.; Huang, S.Y.; Renaud, A.; Zhang, Y.; Ji, J.; Takeda, S.; Morris, J.; Teicher, B.; Doroshow, J.H.; Pommier, Y. Stereospecific PARP trapping by BMN 673 and comparison with olaparib and rucaparib. Mol. Cancer Ther. 2014, 13, e433. [Google Scholar] [CrossRef]
- Pommier, Y.; O’Connor, M.J.; de Bono, J. Laying a trap to kill cancer cells: PARP inhibitors and their mechanisms of action. Sci. Transl. Med. 2016, 8, 362ps17. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Rehman, F.L.; Feng, Y.; Boshuizen, J.; Bajrami, I.; Elliott, R.; Wang, B.; Lord, C.J.; Post, L.E.; Ashworth, A. BMN 673, a novel and highly potent PARP1/2 inhibitor for the treatment of human cancers with DNA repair deficiency. Clin. Cancer Res. 2013, 19, 5003–5015. [Google Scholar] [CrossRef] [PubMed]
- Lickliter, J.D.; Gan, H.K.; Meniawy, T.; Yang, J.; Wang, L.; Luo, L.; Lu, N.; Millward, M. A phase I dose-escalation study of BGB-290, a novel PARP1/2 selective inhibitor in patients with advanced solid tumors. J. Clin. Oncol. 2016, 34, e17049. [Google Scholar] [CrossRef]
- Kaufman, B.; Shapira-Frommer, R.; Schmutzler, R.K.; Audeh, M.W.; Friedlander, M.; Balmana, J.; Mitchell, G.; Fried, G.; Stemmer, S.M.; Hubert, A.; et al. Olaparib monotherapy in patients with advanced cancer and a germline BRCA1/2 mutation. J. Clin. Oncol. 2015, 33, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Ledermann, J.; Harter, P.; Gourley, C.; Friedlander, M.; Vergote, I.; Rustin, G.; Scott, C.; Meier, W.; Shapira-Frommer, R.; Safra, T.; et al. Olaparib maintenance therapy in platinum-sensitive relapsed ovarian cancer. N. Engl. J. Med. 2012, 366, 1382–1392. [Google Scholar] [CrossRef]
- Pujade-Lauraine, E.; Ledermann, J.A.; Selle, F.; Gebski, V.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Poveda, A.; Pignata, S.; et al. Olaparib tablets as maintenance therapy in patients with platinum-sensitive, relapsed ovarian cancer and a BRCA1/2 mutation (SOLO2/ENGOT-Ov21): A double-blind, randomised, placebo-controlled, phase 3 trial. Lancet Oncol. 2017, 18, 1274–1284. [Google Scholar] [CrossRef]
- Moore, K.; Colombo, N.; Scambia, G.; Kim, B.G.; Oaknin, A.; Friedlander, M.; Lisyanskaya, A.; Floquet, A.; Leary, A.; Sonke, G.S.; et al. Maintenance Olaparib in Patients with Newly Diagnosed Advanced Ovarian Cancer. N. Engl. J. Med. 2018, 379, 2495–2505. [Google Scholar] [CrossRef]
- Robson, M.; Im, S.A.; Senkus, E.; Xu, B.; Domchek, S.M.; Masuda, N.; Delaloge, S.; Li, W.; Tung, N.; Armstrong, A.; et al. Olaparib for Metastatic Breast Cancer in Patients with a Germline BRCA Mutation. N. Engl. J. Med. 2017, 377, 523–533. [Google Scholar] [CrossRef]
- Kristeleit, R.; Shapiro, G.I.; Burris, H.A.; Oza, A.M.; LoRusso, P.; Patel, M.R.; Domchek, S.M.; Balmana, J.; Drew, Y.; Chen, L.M.; et al. A Phase I-II Study of the Oral PARP Inhibitor Rucaparib in Patients with Germline BRCA1/2-Mutated Ovarian Carcinoma or Other Solid Tumors. Clin. Cancer Res. 2017, 23, 4095–4106. [Google Scholar] [CrossRef]
- Swisher, E.M.; Lin, K.K.; Oza, A.M.; Scott, C.L.; Giordano, H.; Sun, J.; Konecny, G.E.; Coleman, R.L.; Tinker, A.V.; O’Malley, D.M.; et al. Rucaparib in relapsed, platinum-sensitive high-grade ovarian carcinoma (ARIEL2 Part 1): An international, multicentre, open-label, phase 2 trial. Lancet Oncol. 2017, 18, 75–87. [Google Scholar] [CrossRef]
- Coleman, R.L.; Oza, A.M.; Lorusso, D.; Aghajanian, C.; Oaknin, A.; Dean, A.; Colombo, N.; Weberpals, J.I.; Clamp, A.; Scambia, G.; et al. Rucaparib maintenance treatment for recurrent ovarian carcinoma after response to platinum therapy (ARIEL3): A randomised, double-blind, placebo-controlled, phase 3 trial. Lancet 2017, 390, 1949–1961. [Google Scholar] [CrossRef]
- Mirza, M.R.; Monk, B.J.; Herrstedt, J.; Oza, A.M.; Mahner, S.; Redondo, A.; Fabbro, M.; Ledermann, J.A.; Lorusso, D.; Vergote, I.; et al. Niraparib Maintenance Therapy in Platinum-Sensitive, Recurrent Ovarian Cancer. N. Engl. J. Med. 2016, 375, 2154–2164. [Google Scholar] [CrossRef] [PubMed]
- Litton, J.K.; Rugo, H.S.; Ettl, J.; Hurvitz, S.A.; Goncalves, A.; Lee, K.H.; Fehrenbacher, L.; Yerushalmi, R.; Mina, L.A.; Martin, M.; et al. Talazoparib in Patients with Advanced Breast Cancer and a Germline BRCA Mutation. N. Engl. J. Med. 2018, 379, 753–763. [Google Scholar] [CrossRef] [PubMed]
- Golan, T.; Hammel, P.; Reni, M.; Van Cutsem, E.; Macarulla, T.; Hall, M.J.; Park, J.O.; Hochhauser, D.; Arnold, D.; Oh, D.Y.; et al. Maintenance Olaparib for Germline BRCA-Mutated Metastatic Pancreatic Cancer. N. Engl. J. Med. 2019, 381, 317–327. [Google Scholar] [CrossRef] [PubMed]
- Coleman, R.L.; Sill, M.W.; Bell-McGuinn, K.; Aghajanian, C.; Gray, H.J.; Tewari, K.S.; Rubin, S.C.; Rutherford, T.J.; Chan, J.K.; Chen, A.; et al. A phase II evaluation of the potent, highly selective PARP inhibitor veliparib in the treatment of persistent or recurrent epithelial ovarian, fallopian tube, or primary peritoneal cancer in patients who carry a germline BRCA1 or BRCA2 mutation—An NRG Oncology/Gynecologic Oncology Group study. Gynecol. Oncol. 2015, 137, 386–391. [Google Scholar] [CrossRef] [PubMed]
- LaFargue, C.J.; Dal Molin, G.Z.; Sood, A.K.; Coleman, R.L. Exploring and comparing adverse events between PARP inhibitors. Lancet Oncol. 2019, 20, 15–28. [Google Scholar] [CrossRef]
- LoRusso, P.M.; Li, J.; Burger, A.; Heilbrun, L.K.; Sausville, E.A.; Boerner, S.A.; Smith, D.; Pilat, M.J.; Zhang, J.; Tolaney, S.M.; et al. Phase I Safety, Pharmacokinetic, and Pharmacodynamic Study of the Poly(ADP-ribose) Polymerase (PARP) Inhibitor Veliparib (ABT-888) in Combination with Irinotecan in Patients with Advanced Solid Tumors. Clin. Cancer Res. 2016, 22, 3227–3237. [Google Scholar] [CrossRef]
- Shroff, R.T.; Hendifar, A.; McWilliams, R.R.; Geva, R.; Epelbaum, R.; Rolfe, L.; Goble, S.; Lin, K.K.; Biankin, A.V.; Giordano, H.; et al. Rucaparib Monotherapy in Patients With Pancreatic Cancer and a Known Deleterious BRCA Mutation. JCO Precis. Oncol. 2018, 2, 1–15. [Google Scholar] [CrossRef]
- Lowery, M.A.; Kelsen, D.P.; Capanu, M.; Smith, S.C.; Lee, J.W.; Stadler, Z.K.; Moore, M.J.; Kindler, H.L.; Golan, T.; Segal, A.; et al. Phase II trial of veliparib in patients with previously treated BRCA-mutated pancreas ductal adenocarcinoma. Eur. J. Cancer 2018, 89, 19–26. [Google Scholar] [CrossRef]
- Golan, T.; Varadhachary, G.R.; Sela, T.; Fogelman, D.R.; Halperin, N.; Shroff, R.T.; Halparin, S.; Xiao, L.; Aderka, D.; Maitra, A.; et al. Phase II study of olaparib for BRCAness phenotype in pancreatic cancer. J. Clin. Oncol. 2018, 36, e297. [Google Scholar] [CrossRef]
- Binder, K.A.R.; Mick, R.; O’Hara, M.; Teitelbaum, U.; Karasic, T.; Schneider, C.; O’Dwyer, P.J.; Carpenter, E.; Pantel, A.; Makvandi, M.; et al. Abstract CT234: A Phase II, single arm study of maintenance rucaparib in patients with platinum-sensitive advanced pancreatic cancer and a pathogenic germline or somatic mutation in BRCA1, BRCA2 or PALB2. Cancer Res. 2019, 79, CT234. [Google Scholar] [CrossRef]
- Chiorean, E.G.; Guthrie, K.A.; Philip, P.A.; Swisher, E.M.; Jalikis, F.; Pishvaian, M.J.; Berlin, J.; Noel, M.S.; Suga, J.M.; Garrido-Laguna, I.; et al. Randomized phase II study of second-line modified FOLFIRI with PARP inhibitor ABT-888 (Veliparib) (NSC-737664) versus FOLFIRI in metastatic pancreatic cancer (mPC): SWOG S1513. J. Clin. Oncol. 2019, 37, e4014. [Google Scholar] [CrossRef]
- Pishvaian, M.J.; Wang, H.; Parenti, S.; He, A.R.; Hwang, J.J.; Ley, L.; Difebo, H.; Smaglo, B.G.; Kim, S.S.; Weinberg, B.A.; et al. Final report of a phase I/II study of veliparib (Vel) in combination with 5-FU and oxaliplatin (FOLFOX) in patients (pts) with metastatic pancreatic cancer (mPDAC). J. Clin. Oncol. 2019, 37, e4015. [Google Scholar] [CrossRef]
- De Bono, J.; Ramanathan, R.K.; Mina, L.; Chugh, R.; Glaspy, J.; Rafii, S.; Kaye, S.; Sachdev, J.; Heymach, J.; Smith, D.C.; et al. Phase I, Dose-Escalation, Two-Part Trial of the PARP Inhibitor Talazoparib in Patients with Advanced Germline BRCA1/2 Mutations and Selected Sporadic Cancers. Cancer Discov. 2017, 7, 620–629. [Google Scholar] [CrossRef] [PubMed]
- Kasi, A.; Chalise, P.; Williamson, S.K.; Baranda, J.C.; Sun, W.; Al-Rajabi, R.M.D.T.; Saeed, A.; Kumer, S.; Schmitt, T.; Foster, C.; et al. Niraparib in metastatic pancreatic cancer after previous chemotherapy (NIRA-PANC): A phase 2 trial. J. Clin. Oncol. 2019, 37, TPS4168. [Google Scholar] [CrossRef]
- Melisi, D.; Ossovskaya, V.; Zhu, C.; Rosa, R.; Ling, J.; Dougherty, P.M.; Sherman, B.M.; Abbruzzese, J.L.; Chiao, P.J. Oral Poly(ADP-Ribose) Polymerase-1 Inhibitor BSI-401 Has Antitumor Activity and Synergizes with Oxaliplatin against Pancreatic Cancer, Preventing Acute Neurotoxicity. Clin. Cancer Res. 2009, 15, 6367–6377. [Google Scholar] [CrossRef]
- Turk, A.A.; Deming, D.A.; Lubner, S.J.; Mulkerin, D.; LoConte, N.K.; Tevaarwerk, A.; Wisinski, K.B.; Eickhoff, J.C.; Liu, G. A phase I study of veliparib (Vel) in combination with oxaliplatin (Ox) and capecitabine (Cap) in advanced solid tumors. J. Clin. Oncol. 2018, 36, e314. [Google Scholar] [CrossRef]
- O’Reilly, E.M.; Lowery, M.A.; Yu, K.H.; Capanu, M.; Stadler, Z.K.; Epstein, A.S.; Golan, T.; Segal, A.; Segal, M.; Salo-Mullen, E.E.; et al. Randomized phase II study of gemcitabine (G), cisplatin (C) with or without veliparib (V) (arms A, B) and a phase II single-arm study of single-agent veliparib (arm C) in patients with BRCA or PALB2-mutated pancreas adenocarcinoma (PC). J. Clin. Oncol. 2013, 31, TPS4144. [Google Scholar] [CrossRef]
- Pfister, T.D.; Reinhold, W.C.; Agama, K.; Gupta, S.; Khin, S.A.; Kinders, R.J.; Parchment, R.E.; Tomaszewski, J.E.; Doroshow, J.H.; Pommier, Y. Topoisomerase I levels in the NCI-60 cancer cell line panel determined by validated ELISA and microarray analysis and correlation with indenoisoquinoline sensitivity. Mol. Cancer Ther. 2009, 8, 1878–1884. [Google Scholar] [CrossRef]
- Patel, A.G.; Flatten, K.S.; Schneider, P.A.; Dai, N.T.; McDonald, J.S.; Poirier, G.G.; Kaufmann, S.H. Enhanced killing of cancer cells by poly(ADP-ribose) polymerase inhibitors and topoisomerase I inhibitors reflects poisoning of both enzymes. J. Biol. Chem. 2012, 287, 4198–4210. [Google Scholar] [CrossRef]
- Jacob, D.A.; Bahra, M.; Langrehr, J.M.; Boas-Knoop, S.; Stefaniak, R.; Davis, J.; Schumacher, G.; Lippert, S.; Neumann, U.P. Combination therapy of poly (ADP-ribose) polymerase inhibitor 3-aminobenzamide and gemcitabine shows strong antitumor activity in pancreatic cancer cells. J. Gastroenterol. Hepatol. 2007, 22, 738–748. [Google Scholar] [CrossRef] [PubMed]
- Bendell, J.; O’Reilly, E.M.; Middleton, M.R.; Chau, I.; Hochster, H.; Fielding, A.; Burke, W.; Burris, H., 3rd. Phase I study of olaparib plus gemcitabine in patients with advanced solid tumours and comparison with gemcitabine alone in patients with locally advanced/metastatic pancreatic cancer. Ann. Oncol. 2015, 26, 804–811. [Google Scholar] [CrossRef] [PubMed]
- Porcelli, L.; Quatrale, A.E.; Mantuano, P.; Leo, M.G.; Silvestris, N.; Rolland, J.F.; Carioggia, E.; Lioce, M.; Paradiso, A.; Azzariti, A. Optimize radiochemotherapy in pancreatic cancer: PARP inhibitors a new therapeutic opportunity. Mol. Oncol. 2013, 7, 308–322. [Google Scholar] [CrossRef] [PubMed]
- Tuli, R.; Surmak, A.J.; Reyes, J.; Armour, M.; Hacker-Prietz, A.; Wong, J.; DeWeese, T.L.; Herman, J.M. Radiosensitization of Pancreatic Cancer Cells In Vitro and In Vivo through Poly (ADP-ribose) Polymerase Inhibition with ABT-888. Transl. Oncol. 2014, 7, 439–445. [Google Scholar] [CrossRef]
- Strickland, K.C.; Howitt, B.E.; Shukla, S.A.; Rodig, S.; Ritterhouse, L.L.; Liu, J.F.; Garber, J.E.; Chowdhury, D.; Wu, C.J.; D’Andrea, A.D.; et al. Association and prognostic significance of BRCA1/2-mutation status with neoantigen load, number of tumor-infiltrating lymphocytes and expression of PD-1/PD-L1 in high grade serous ovarian cancer. Oncotarget 2016, 7, 13587–13598. [Google Scholar] [CrossRef]
- Parkes, E.E.; Walker, S.M.; Taggart, L.E.; McCabe, N.; Knight, L.A.; Wilkinson, R.; McCloskey, K.D.; Buckley, N.E.; Savage, K.I.; Salto-Tellez, M.; et al. Activation of STING-Dependent Innate Immune Signaling By S-Phase-Specific DNA Damage in Breast Cancer. J. Natl. Cancer Inst. 2017, 109, djw199. [Google Scholar] [CrossRef]
- Ishikawa, H.; Ma, Z.; Barber, G.N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 2009, 461, 788–792. [Google Scholar] [CrossRef]
- Simmons, D.P.; Wearsch, P.A.; Canaday, D.H.; Meyerson, H.J.; Liu, Y.C.; Wang, Y.; Boom, W.H.; Harding, C.V. Type I IFN drives a distinctive dendritic cell maturation phenotype that allows continued class II MHC synthesis and antigen processing. J. Immunol. (Baltim. Md. 1950) 2012, 188, 3116–3126. [Google Scholar] [CrossRef]
- Ding, L.; Kim, H.J.; Wang, Q.; Kearns, M.; Jiang, T.; Ohlson, C.E.; Li, B.B.; Xie, S.; Liu, J.F.; Stover, E.H.; et al. PARP Inhibition Elicits STING-Dependent Antitumor Immunity in Brca1-Deficient Ovarian Cancer. Cell Rep. 2018, 25, 2972–2980. [Google Scholar] [CrossRef]
- Bacher, N.; Raker, V.; Hofmann, C.; Graulich, E.; Schwenk, M.; Baumgrass, R.; Bopp, T.; Zechner, U.; Merten, L.; Becker, C.; et al. Interferon-alpha suppresses cAMP to disarm human regulatory T cells. Cancer Res. 2013, 73, 5647–5656. [Google Scholar] [CrossRef]
- Jiao, S.; Xia, W.; Yamaguchi, H.; Wei, Y.; Chen, M.K.; Hsu, J.M.; Hsu, J.L.; Yu, W.H.; Du, Y.; Lee, H.H.; et al. PARP Inhibitor Upregulates PD-L1 Expression and Enhances Cancer-Associated Immunosuppression. Clin. Cancer Res. 2017, 23, 3711–3720. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Sun, K.; Xiao, Y.; Feng, B.; Mikule, K.; Ma, X.; Feng, N.; Vellano, C.P.; Federico, L.; Marszalek, J.R.; et al. Niraparib activates interferon signaling and potentiates anti-PD-1 antibody efficacy in tumor models. Sci. Rep. 2019, 9, e1853. [Google Scholar] [CrossRef] [PubMed]
- Sato, H.; Niimi, A.; Yasuhara, T.; Permata, T.B.M.; Hagiwara, Y.; Isono, M.; Nuryadi, E.; Sekine, R.; Oike, T.; Kakoti, S.; et al. DNA double-strand break repair pathway regulates PD-L1 expression in cancer cells. Nat. Commun. 2017, 8, e1751. [Google Scholar] [CrossRef] [PubMed]
- Sen, T.; Rodriguez, B.L.; Chen, L.; Corte, C.M.D.; Morikawa, N.; Fujimoto, J.; Cristea, S.; Nguyen, T.; Diao, L.; Li, L.; et al. Targeting DNA Damage Response Promotes Antitumor Immunity through STING-Mediated T-cell Activation in Small Cell Lung Cancer. Cancer Discov. 2019, 9, 646–661. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, T.; Flies, D.B.; Marjon, N.A.; Mantia-Smaldone, G.; Ronner, L.; Gimotty, P.A.; Adams, S.F. CTLA-4 Blockade Synergizes Therapeutically with PARP Inhibition in BRCA1-Deficient Ovarian Cancer. Cancer Immunol. Res. 2015, 3, 1257–1268. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-M.; Zimmer, A.D.S.; Lipkowitz, S.; Annunziata, C.M.; Ho, T.W.; Chiou, V.L.; Minasian, L.M.; Houston, N.D.; Ekwede, I.; Kohn, E.C. Phase I study of the PD-L1 inhibitor, durvalumab (MEDI4736; D) in combination with a PARP inhibitor, olaparib (O) or a VEGFR inhibitor, cediranib (C) in women’s cancers (NCT02484404). J. Clin. Oncol. 2016, 34, e3015. [Google Scholar] [CrossRef]
- Bang, Y.-J.; Kaufman, B.; Geva, R.; Stemmer, S.M.; Hong, S.-H.; Lee, J.-S.; Domchek, S.M.; Lanasa, M.C.; Tang, M.; Gresty, C.; et al. An open-label, phase II basket study of olaparib and durvalumab (MEDIOLA): Results in patients with relapsed gastric cancer. J. Clin. Oncol. 2019, 37, e140. [Google Scholar] [CrossRef]
- Vinayak, S.; Tolaney, S.M.; Schwartzberg, L.S.; Mita, M.M.; McCann, G.A.-L.; Tan, A.R.; Hendrickson, A.E.W.; Forero-Torres, A.; Anders, C.K.; Wulf, G.M.; et al. TOPACIO/Keynote-162: Niraparib + pembrolizumab in patients (pts) with metastatic triple-negative breast cancer (TNBC), a phase 2 trial. J. Clin. Oncol. 2018, 36, e1011. [Google Scholar] [CrossRef]
- Konstantinopoulos, P.A.; Waggoner, S.; Vidal, G.A.; Mita, M.; Moroney, J.W.; Holloway, R.; Van Le, L.; Sachdev, J.C.; Chapman-Davis, E.; Colon-Otero, G.; et al. Single-Arm Phases 1 and 2 Trial of Niraparib in Combination With Pembrolizumab in Patients With Recurrent Platinum-Resistant Ovarian Carcinoma. JAMA Oncol. 2019, 5, 1141–1149. [Google Scholar] [CrossRef]
- Zimmer, A.S.; Nichols, E.; Cimino-Mathews, A.; Peer, C.; Cao, L.; Lee, M.J.; Kohn, E.C.; Annunziata, C.M.; Lipkowitz, S.; Trepel, J.B.; et al. A phase I study of the PD-L1 inhibitor, durvalumab, in combination with a PARP inhibitor, olaparib, and a VEGFR1-3 inhibitor, cediranib, in recurrent women’s cancers with biomarker analyses. J. Immunother. Cancer 2019, 7, e197. [Google Scholar] [CrossRef]
- Reiss, K.A.; Mick, R.; O’Hara, M.H.; Teitelbaum, U.R.; Karasic, T.B.; Schneider, C.J.; O’Dwyer, P.J.; Karlson, D.; Cowden, S.; Fuhrer, M.J.; et al. A randomized phase II trial of niraparib plus either nivolumab or ipilimumab in patients with advanced pancreatic cancer whose cancer has not progressed on platinum-based therapy. J. Clin. Oncol. 2019, 37, TPS4161. [Google Scholar] [CrossRef]
- Sun, C.; Fang, Y.; Yin, J.; Chen, J.; Ju, Z.; Zhang, D.; Chen, X.; Vellano, C.P.; Jeong, K.J.; Ng, P.K.; et al. Rational combination therapy with PARP and MEK inhibitors capitalizes on therapeutic liabilities in RAS mutant cancers. Sci. Transl. Med. 2017, 9, eaal5148. [Google Scholar] [CrossRef] [PubMed]
- Tentori, L.; Lacal, P.M.; Muzi, A.; Dorio, A.S.; Leonetti, C.; Scarsella, M.; Ruffini, F.; Xu, W.; Min, W.; Stoppacciaro, A.; et al. Poly(ADP-ribose) polymerase (PARP) inhibition or PARP-1 gene deletion reduces angiogenesis. Eur. J. Cancer 2007, 43, 2124–2133. [Google Scholar] [CrossRef] [PubMed]
- Sakai, W.; Swisher, E.M.; Karlan, B.Y.; Agarwal, M.K.; Higgins, J.; Friedman, C.; Villegas, E.; Jacquemont, C.; Farrugia, D.J.; Couch, F.J.; et al. Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers. Nature 2008, 451, 1116–1120. [Google Scholar] [CrossRef]
- Norquist, B.; Wurz, K.A.; Pennil, C.C.; Garcia, R.; Gross, J.; Sakai, W.; Karlan, B.Y.; Taniguchi, T.; Swisher, E.M. Secondary somatic mutations restoring BRCA1/2 predict chemotherapy resistance in hereditary ovarian carcinomas. J. Clin. Oncol. 2011, 29, 3008–3015. [Google Scholar] [CrossRef]
- Barber, L.J.; Sandhu, S.; Chen, L.; Campbell, J.; Kozarewa, I.; Fenwick, K.; Assiotis, I.; Rodrigues, D.N.; Reis Filho, J.S.; Moreno, V.; et al. Secondary mutations in BRCA2 associated with clinical resistance to a PARP inhibitor. J. Pathol. 2013, 229, 422–429. [Google Scholar] [CrossRef]
- Bunting, S.F.; Callen, E.; Wong, N.; Chen, H.T.; Polato, F.; Gunn, A.; Bothmer, A.; Feldhahn, N.; Fernandez-Capetillo, O.; Cao, L.; et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 2010, 141, 243–254. [Google Scholar] [CrossRef]
- Cao, L.; Xu, X.; Bunting, S.F.; Liu, J.; Wang, R.H.; Cao, L.L.; Wu, J.J.; Peng, T.N.; Chen, J.; Nussenzweig, A.; et al. A selective requirement for 53BP1 in the biological response to genomic instability induced by Brca1 deficiency. Mol. Cell 2009, 35, 534–541. [Google Scholar] [CrossRef]
- Oplustilova, L.; Wolanin, K.; Mistrik, M.; Korinkova, G.; Simkova, D.; Bouchal, J.; Lenobel, R.; Bartkova, J.; Lau, A.; O’Connor, M.J.; et al. Evaluation of candidate biomarkers to predict cancer cell sensitivity or resistance to PARP-1 inhibitor treatment. Cell Cycle 2012, 11, 3837–3850. [Google Scholar] [CrossRef]
- Neijenhuis, S.; Bajrami, I.; Miller, R.; Lord, C.J.; Ashworth, A. Identification of miRNA modulators to PARP inhibitor response. DNA Repair (Amst.) 2013, 12, 394–402. [Google Scholar] [CrossRef]
- Choi, Y.E.; Meghani, K.; Brault, M.E.; Leclerc, L.; He, Y.J.; Day, T.A.; Elias, K.M.; Drapkin, R.; Weinstock, D.M.; Dao, F.; et al. Platinum and PARP Inhibitor Resistance Due to Overexpression of MicroRNA-622 in BRCA1-Mutant Ovarian Cancer. Cell Rep. 2016, 14, 429–439. [Google Scholar] [CrossRef] [PubMed]
- Blazek, D.; Kohoutek, J.; Bartholomeeusen, K.; Johansen, E.; Hulinkova, P.; Luo, Z.; Cimermancic, P.; Ule, J.; Peterlin, B.M. The Cyclin K/Cdk12 complex maintains genomic stability via regulation of expression of DNA damage response genes. Genes Dev. 2011, 25, 2158–2172. [Google Scholar] [CrossRef] [PubMed]
- Johnson, S.F.; Cruz, C.; Greifenberg, A.K.; Dust, S.; Stover, D.G.; Chi, D.; Primack, B.; Cao, S.; Bernhardy, A.J.; Coulson, R.; et al. CDK12 Inhibition Reverses De Novo and Acquired PARP Inhibitor Resistance in BRCA Wild-Type and Mutated Models of Triple-Negative Breast Cancer. Cell Rep. 2016, 17, 2367–2381. [Google Scholar] [CrossRef]
- Alagpulinsa, D.A.; Ayyadevara, S.; Yaccoby, S.; Shmookler Reis, R.J. A Cyclin-Dependent Kinase Inhibitor, Dinaciclib, Impairs Homologous Recombination and Sensitizes Multiple Myeloma Cells to PARP Inhibition. Mol. Cancer Ther. 2016, 15, 241–250. [Google Scholar] [CrossRef]
- Lloyd, R.; Falenta, K.; Wijnhoven, P.W.; Chabbert, C.; Stott, J.; Yates, J.; Lau, A.Y.; Young, L.A.; Hollingsworth, S.J. Abstract 337: The PARP inhibitor olaparib is synergistic with the ATR inhibitor AZD6738 in ATM deficient cancer cells. Cancer Res. 2018, 78, e337. [Google Scholar] [CrossRef]
- Hamilton, E.; Falchook, G.S.; Wang, J.S.; Fu, S.; Oza, A.; Karen, S.; Imedio, E.R.; Kumar, S.; Ottesen, L.; Mugundu, G.M.; et al. Abstract CT025: Phase Ib study of adavosertib in combination with olaparib in patients with refractory solid tumors: Dose escalation. Cancer Res. 2019, 79, CT025. [Google Scholar] [CrossRef]
- Du, Y.; Yamaguchi, H.; Wei, Y.; Hsu, J.L.; Wang, H.L.; Hsu, Y.H.; Lin, W.C.; Yu, W.H.; Leonard, P.G.; Lee, G.R.t.; et al. Blocking c-Met-mediated PARP1 phosphorylation enhances anti-tumor effects of PARP inhibitors. Nat. Med. 2016, 22, 194–201. [Google Scholar] [CrossRef]
- Han, Y.; Chen, M.K.; Wang, H.L.; Hsu, J.L.; Li, C.W.; Chu, Y.Y.; Liu, C.X.; Nie, L.; Chan, L.C.; Yam, C.; et al. Synergism of PARP inhibitor fluzoparib (HS10160) and MET inhibitor HS10241 in breast and ovarian cancer cells. Am. J. Cancer Res. 2019, 9, 608–618. [Google Scholar]
- Liu, Q.; Turner, K.M.; Alfred Yung, W.K.; Chen, K.; Zhang, W. Role of AKT signaling in DNA repair and clinical response to cancer therapy. Neuro-Oncology 2014, 16, 1313–1323. [Google Scholar] [CrossRef]
- Ibrahim, Y.H.; Garcia-Garcia, C.; Serra, V.; He, L.; Torres-Lockhart, K.; Prat, A.; Anton, P.; Cozar, P.; Guzman, M.; Grueso, J.; et al. PI3K inhibition impairs BRCA1/2 expression and sensitizes BRCA-proficient triple-negative breast cancer to PARP inhibition. Cancer Discov. 2012, 2, 1036–1047. [Google Scholar] [CrossRef]
- Rottenberg, S.; Jaspers, J.E.; Kersbergen, A.; van der Burg, E.; Nygren, A.O.; Zander, S.A.; Derksen, P.W.; de Bruin, M.; Zevenhoven, J.; Lau, A.; et al. High sensitivity of BRCA1-deficient mammary tumors to the PARP inhibitor AZD2281 alone and in combination with platinum drugs. Proc. Natl. Acad. Sci. USA 2008, 105, 17079–17084. [Google Scholar] [CrossRef] [PubMed]
- Oplustil O’Connor, L.; Rulten, S.L.; Cranston, A.N.; Odedra, R.; Brown, H.; Jaspers, J.E.; Jones, L.; Knights, C.; Evers, B.; Ting, A.; et al. The PARP Inhibitor AZD2461 Provides Insights into the Role of PARP3 Inhibition for Both Synthetic Lethality and Tolerability with Chemotherapy in Preclinical Models. Cancer Res. 2016, 76, 6084–6094. [Google Scholar] [CrossRef] [PubMed]
- Tempero, M.A.; Malafa, M.P.; Chiorean, E.G.; Czito, B.; Scaife, C.; Narang, A.K.; Fountzilas, C.; Wolpin, B.M.; Al-Hawary, M.; Asbun, H.; et al. Pancreatic Adenocarcinoma, Version 1.2019. J. Natl. Compr. Cancer Netw. 2019, 17, 202–210. [Google Scholar] [CrossRef] [PubMed]
- Dougherty, B.A.; Lai, Z.; Hodgson, D.R.; Orr, M.C.M.; Hawryluk, M.; Sun, J.; Yelensky, R.; Spencer, S.K.; Robertson, J.D.; Ho, T.W.; et al. Biological and clinical evidence for somatic mutations in BRCA1 and BRCA2 as predictive markers for olaparib response in high-grade serous ovarian cancers in the maintenance setting. Oncotarget 2017, 8, 43653–43661. [Google Scholar] [CrossRef]
- Dahan, L.; Phelip, J.M.; Malicot, K.L.; Williet, N.; Desrame, J.; Volet, J.; Petorin, C.; Malka, D.; Rebischung, C.; Aparicio, T.; et al. FOLFIRINOX until progression, FOLFIRINOX with maintenance treatment, or sequential treatment with gemcitabine and FOLFIRI.3 for first-line treatment of metastatic pancreatic cancer: A randomized phase II trial (PRODIGE 35-PANOPTIMOX). J. Clin. Oncol. 2018, 36, e4000. [Google Scholar] [CrossRef]
- Johnson, N.; Johnson, S.F.; Yao, W.; Li, Y.C.; Choi, Y.E.; Bernhardy, A.J.; Wang, Y.; Capelletti, M.; Sarosiek, K.A.; Moreau, L.A.; et al. Stabilization of mutant BRCA1 protein confers PARP inhibitor and platinum resistance. Proc. Natl. Acad. Sci. USA 2013, 110, 17041–17046. [Google Scholar] [CrossRef]
- Chalasani, P.; Kurtin, S.; Dragovich, T. Response to a third-line mitomycin C (MMC)-based chemotherapy in a patient with metastatic pancreatic adenocarcinoma carrying germline BRCA2 mutation. Jop 2008, 9, 305–308. [Google Scholar]
| Agent | # Potency for PARP Trapping | Mono Therapy Dose | Toxicities * | FDA Approvals ∫ | |
|---|---|---|---|---|---|
| Any (%) | Grade >/3 (%) | ||||
| Olaparib | ++ | 300 mg PO BID | Any 96% Fatigue 60% Nausea 45% Anemia 27% Abd pain 29% Diarrhea 29% Anorexia 25% Constipation 23% | Any 40% Fatigue 5% Anemia 11% Abd pain 2% Anorexia 3Vomiting 1% Arthralgia 1% | -2014: gBRCA mutation positive ovarian cancer in 4th line of therapy. ORR of 34%, PFS of 6.7 mo [56] -2017: Maintenance post-recurrence in ovarian, primary peritoneal or fallopian tube carcinoma with CR or PR after platinum-based chemotherapy irrespective of BRCA status. PFS: olaparib (19 mo) vs. placebo (5.5 mo), HR 0.3, p < 0.0001) [57,58] -2018: Front-line maintenance in gBRCA or sBRCA mutation positive ovarian or primary peritoneal or fallopian tube carcinoma with CR or PR after platinum-based chemotherapy. 3-yr OS: olaparib (84%) vs placebo (80%), HR 0.95 (95% CI 0.6–1.53) [59] -2018: gBRCA mutation positive, Her-2 negative metastatic breast cancer </2 lines of therapy. PFS: olaparib (7 mo) vs std Rx (4 mo), HR 0.58 (p < 0.001); OS: olaparib (19.3 mo) vs. standard therapy (19.6mo), HR 0.9 (p = 0.57) [60] |
| Rucaparib | ++ | 600 mg PO BID | Any 100% Nausea 75% Fatigue 69% Dysgeusia 39% Anemia 37% Constipation, Vomiting 37% Transaminitis 34% Diarrhea 32% Abd pain 30% Thrombocytopenia 28% | Any 48% Anemia 18% Asthenia 7% Neutropenia 5% Thrombocytopenia 3% Transaminitis 10% Nausea, Vomiting 4% Abd pain, Diarrhea, Anorexia, Arthralgia 1% | -2016: gBRCA or sBRCA mutation positive ovarian cancer after >/2 lines of therapy. PFS: BRCA mutation positive (13 mo), LOH high (6 mo), LOH low (5.2mo) [61,62] -2018: Maintenance post-recurrence in ovarian or primary peritoneal or fallopian tube carcinoma with CR or PR after platinum-based chemotherapy PFS: rucaparib (17mo) vs placebo (5 mo), HR 0.23 (p < 0.0001) [63] |
| Veliparib | + | 400 mg PO BID | Nausea 43% Vomiting 29% Anemia 24% Leukopenia 20% Thrombocytopenia 9% | ||
| Niraparib | +++ | 300 mg PO QD | Nausea 74% Thrombocytopenia 61% Fatigue 59% Anemia 50% Constipation 40% Vomiting 34% Neutropenia 30% Headache 26% Anorexia 25% Insomnia 24% Abd pain 23% | Thrombocytopenia 34% Anemia 25% Neutropenia 20% Fatigue 8% Nausea 3% Hypertension 8% Vomiting 2% Abd pain, Dyspnea 1% | -2017: Maintenance post-recurrence in ovarian or primary peritoneal or fallopian tube carcinoma with CR or PR after platinum-based chemotherapy. PFS: niraparib (21mo) vs placebo (5.5 mo), HR 0.27 (95% CI 0.17–0.41) [64] |
| Talazoparib | ++++ | 1 mg PO QD | Anemia 53% Fatigue 50% Nausea 49% Headache 32% Neutropenia 35% Thrombocytopenia 27% Vomiting 25% | Anemia 39% Neutropenia 18% Thrombocytopenia 11% Leukopenia 6% Lymphopenia 3% Fatigue, Headache, Vomiting, back pain, dyspnea 2% | -2018: gBRCA mutation, Her-2 negative metastatic breast cancer </3 lines of therapy. PFS: talazoparib (8.6mo) vs placebo (5.6 mo), HR 0.54; p < 0.001. Interim OS HR 0.76 (95% CI 0.55–1.06, p = 0.11) [65] |
| Pamiparib | 60 mg PO BID | Nausea 50% Fatigue 33% Anemia 20% Vomiting 15% Neutropenia 13% | Anemia 13% Neutropenia 8% Fatigue 5% | No FDA approved indications yet | |
| Clinical Study | Phase | Patient Population | Intervention | Outcome | ADEs |
|---|---|---|---|---|---|
| PARPi as Monotherapy | |||||
| NCT01078662 Kauffman et al. [56] | II | gBRCA1/2 mutation positive advanced recurrent cancers, PDAC cohort with progression on gemcitabine (65% prior platinum-based regimen) | Single arm olaparib 400mg PO BID | PEP: ORR (PDAC cohort) 22% SEP: Stable disease at > 8 weeks 35%, DOR 134 days, PFS 4.6 mo, OS 9.8 mo | Any grade: Fatigue (74%), Nausea (48%), Vomiting, Anemia (40%) Grade >/3: Anemia (17%) Fatigue (13%) |
| NCT02042378 Shroff et al. [70] | II | sBRCA1/2 or gBRCA1/2 mutation positive advanced PDAC, 1–2 prior lines of therapy, prior platinum exposure in 79% pts, platinum refractory disease in 42% patients | Single arm rucaparib 600 mg BID | PEP: ORR 16% (3/19, 1CR and 2PR), SEP: DCR 32%, 44% with 1 prior line of therapy | Any grade: Nausea (63%) Anemia (47%), Abdominal pain, fatigue (37%) Grade >/3: Anemia (32%) Fatigue, Ascites (16%) Nausea, abdominal pain, increased AST, ALT (10%) |
| Lowery et al. [71] | II | gBRCA1/2 or PALB2 mutation positive advanced PDAC patients, prior 1–2 lines of therapies (88% prior platinum-based regimen, 64% of these pts had PD on platinum-based regimen) | Single arm veliparib 300mg BID PO (n = 3), 400 mg BID (n = 15) | PEP: ORR-No CR or PR, Stable disease 25% pts at 8 weeks SEP: PFS 1.7 mo, OS 3.1 mo | Fatigue (25%) Elevated bilirubin (19%) Thrombocytopenia, dehydration, increased alkaline phosphatase (13%) |
| NCT02677038, NCT02511223 Golan et al. [72] | II | Advanced PDAC, >/1 lines of therapy with BRCAness phenotype | Single arm olaparib PO BID | PEP: ORR Israel-5SD, 12 PD; U.S.-2PR, 6 SD, 3 PD SEP: PFS-14 wks (Israel) and 25 wks (U.S.) | Grade 1–2 anemia, fatigue, nausea |
| PARPi as Maintenance Therapy | |||||
| NCT02184195, Golan et al. [66] | III | gBRCA1/2 mutation positive, mPDAC, non-progressive disease during first line platinum-based therapy for at least 16 weeks | 3:2 randomization to olaparib versus placebo | PEP: PFS-7.4 mo vs 3.8 mo (HR 0.53, p = 0.004) SEP: OS (46% data maturity)-19 mo vs. 18 mo (p = 0.7), no difference in HrQOL scores | Any grade: Olaparib vs. placebo (96% vs. 93%), Fatigue (60% vs. 35%), nausea (45% vs. 23%), abdominal pain (29% vs. 25%), anemia (27% vs. 17%) Grade >/3: Olaparib vs placebo (40% vs. 23%) Anemia (11% vs. 3%) Fatigue (5% vs. 2%) Abdominal pain (2% each) |
| NCT 03140670, Binder et al. [73] | II | gBRCA1/2, gPALB2, sBRCA1/2, or sPALB2 mutation positive advanced PDAC, non-progressive disease during first line platinum-based therapy for at least 16 weeks | Single arm rucaparib 600mg PO BID | PEP: PFS Prelim data (19/24 pts enrolled, 42 planned) -mPFS of 9 mo SEP: ORR 37% (1CR, 6 PRs), DCR-90% for at least 8 weeks | Most common (grade 1,2): Nausea (46%) Dysgeusia (33%) Fatigue (25%) No grade 3 ADEs |
| PARP in combination with chemotherapy | |||||
| NCT02890355, Chiorean et al. [74] | II | mPDAC, second line therapy with (1st line Rx with non-irinotecan-based therapy), 9% (11/123) pts with HRD (4 germline BRCA1/2, ATM; 7 somatic BCRA2, PALB2, ATM, CDK12), 20% (24/123) pts with DDR, non HRD mutations | 1:1 randomization to either veliparib + FOLFIRI vs FOLFIRI alone | Planned interim futility analysis at 35% PFS events PEP: OS 5.1 vs. 5.9 mo (HR 1.3, p = 0.2) SEP: PFS 2 mo vs. 3 mo (HR 1.5, p = 0.05) | Most common Grade >/3 ADEs: Veliparib + mFOLFIRI vs FOLFIRI Neutropenia (33% vs. 20%) Fatigue (19% vs. 4%) Nausea (11% vs. 4%) |
| NCT01489865, Pishvaian et al. [75] | I/II | mPDAC pts, phase I (31 pts), phase II (33pts). Phase II pts preselected for germline or somatic BRCA1/2, PALB2 mutations (27%) or FH of breast /ovarian syndrome (69%); both previously treated (18/33) and untreated (15/33) | Veliparib + mFOLFOX6 | 57/64 pts evaluable PEP: ORR-26% all pts, 58% in pts with +FH, +DDR, platinum naïve (12 pts) SEP: OS 8.5 mo, PFS 3.7 mo | Grade >/3: Myelosuppression (16%) Nausea, vomiting (6%) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gupta, M.; Iyer, R.; Fountzilas, C. Poly(ADP-Ribose) Polymerase Inhibitors in Pancreatic Cancer: A New Treatment Paradigms and Future Implications. Cancers 2019, 11, 1980. https://doi.org/10.3390/cancers11121980
Gupta M, Iyer R, Fountzilas C. Poly(ADP-Ribose) Polymerase Inhibitors in Pancreatic Cancer: A New Treatment Paradigms and Future Implications. Cancers. 2019; 11(12):1980. https://doi.org/10.3390/cancers11121980
Chicago/Turabian StyleGupta, Medhavi, Renuka Iyer, and Christos Fountzilas. 2019. "Poly(ADP-Ribose) Polymerase Inhibitors in Pancreatic Cancer: A New Treatment Paradigms and Future Implications" Cancers 11, no. 12: 1980. https://doi.org/10.3390/cancers11121980
APA StyleGupta, M., Iyer, R., & Fountzilas, C. (2019). Poly(ADP-Ribose) Polymerase Inhibitors in Pancreatic Cancer: A New Treatment Paradigms and Future Implications. Cancers, 11(12), 1980. https://doi.org/10.3390/cancers11121980





