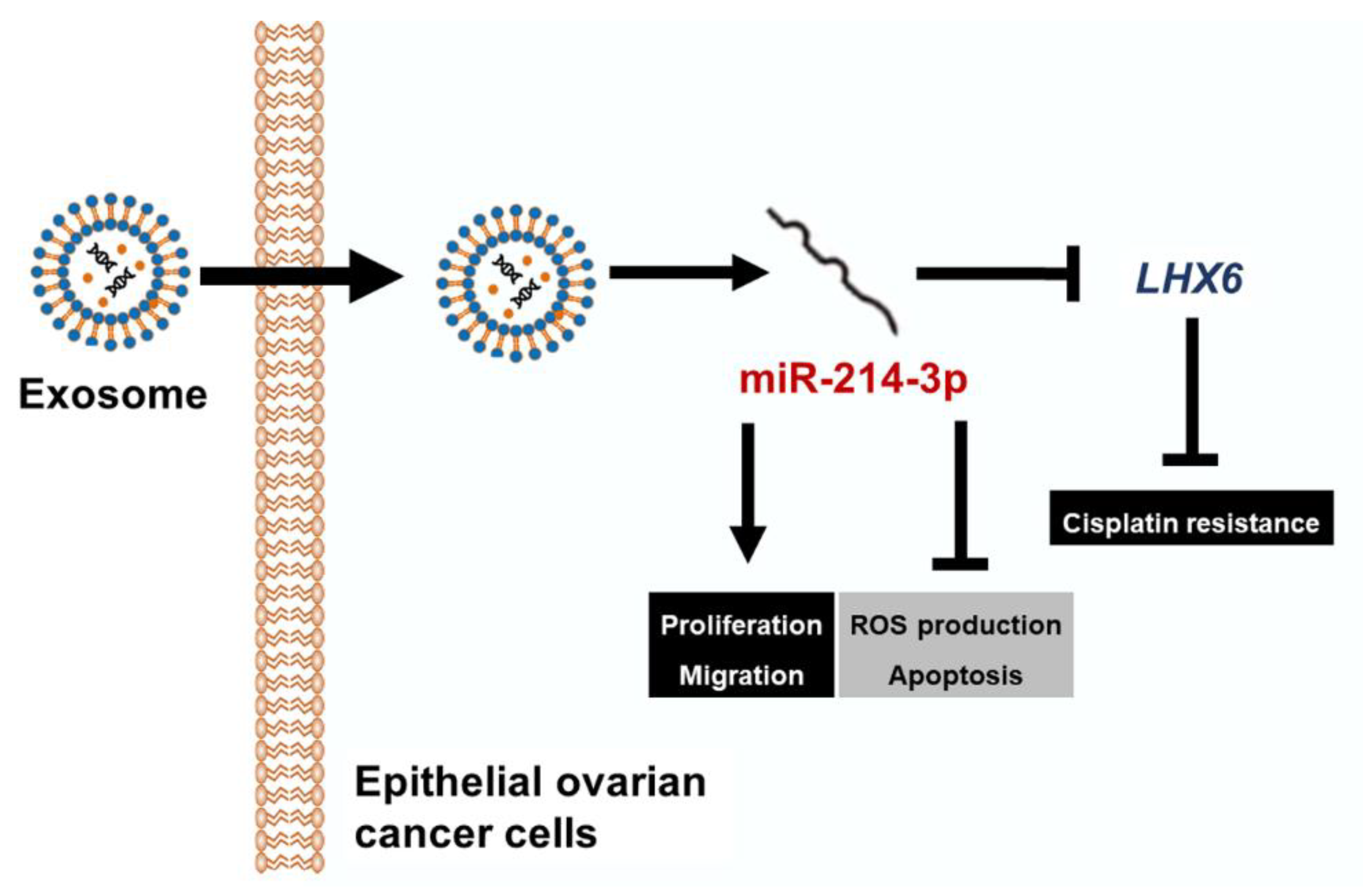
Inhibition of miR-214-3p Aids in Preventing Epithelial Ovarian Cancer Malignancy by Increasing the Expression of LHX6
Abstract
1. Introduction
2. Results
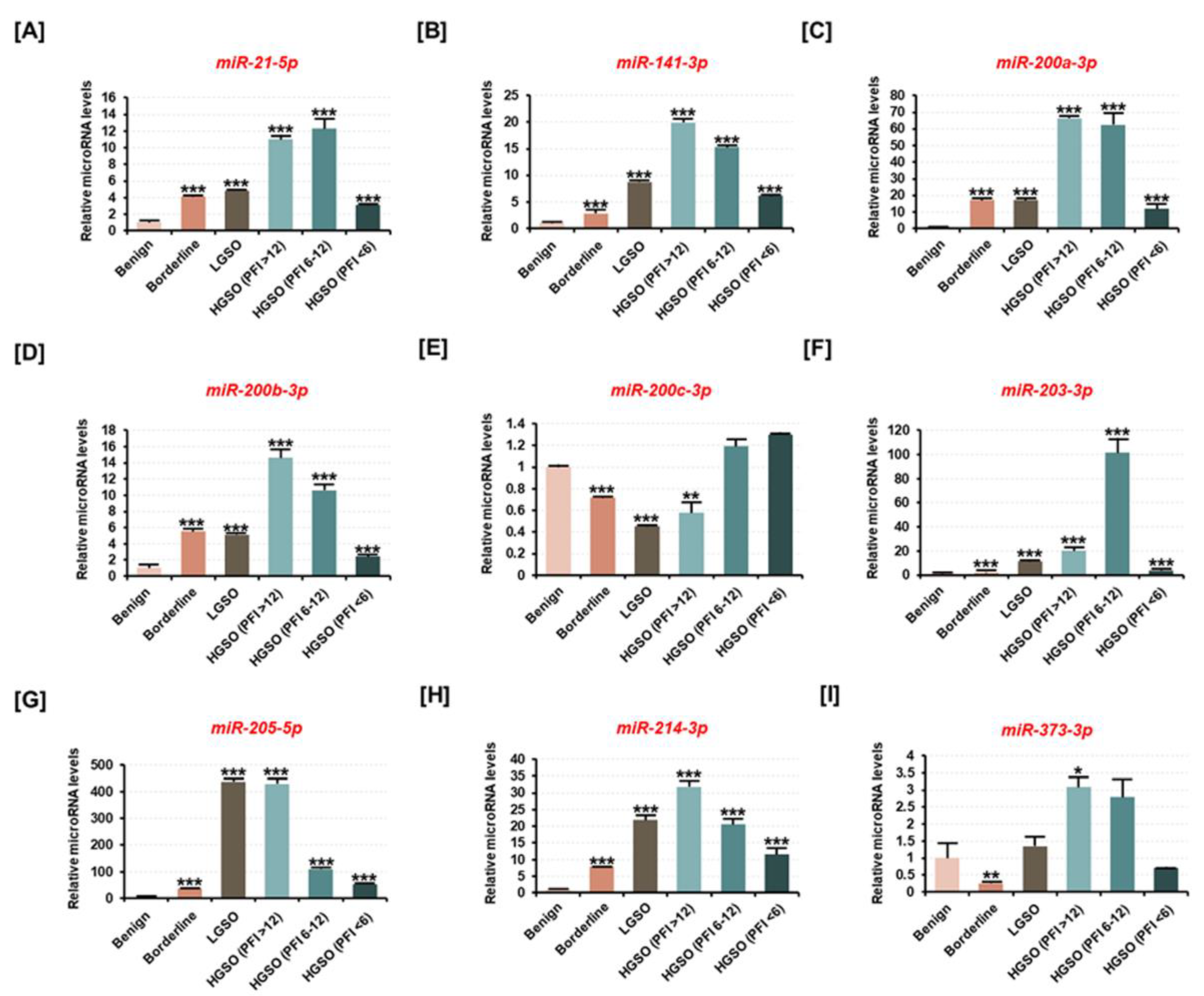
2.1. miR-214-3p Is Positively Correlated with the Malignancy of Ovarian Tumors in Human Tissues But Not with Drug Resistance
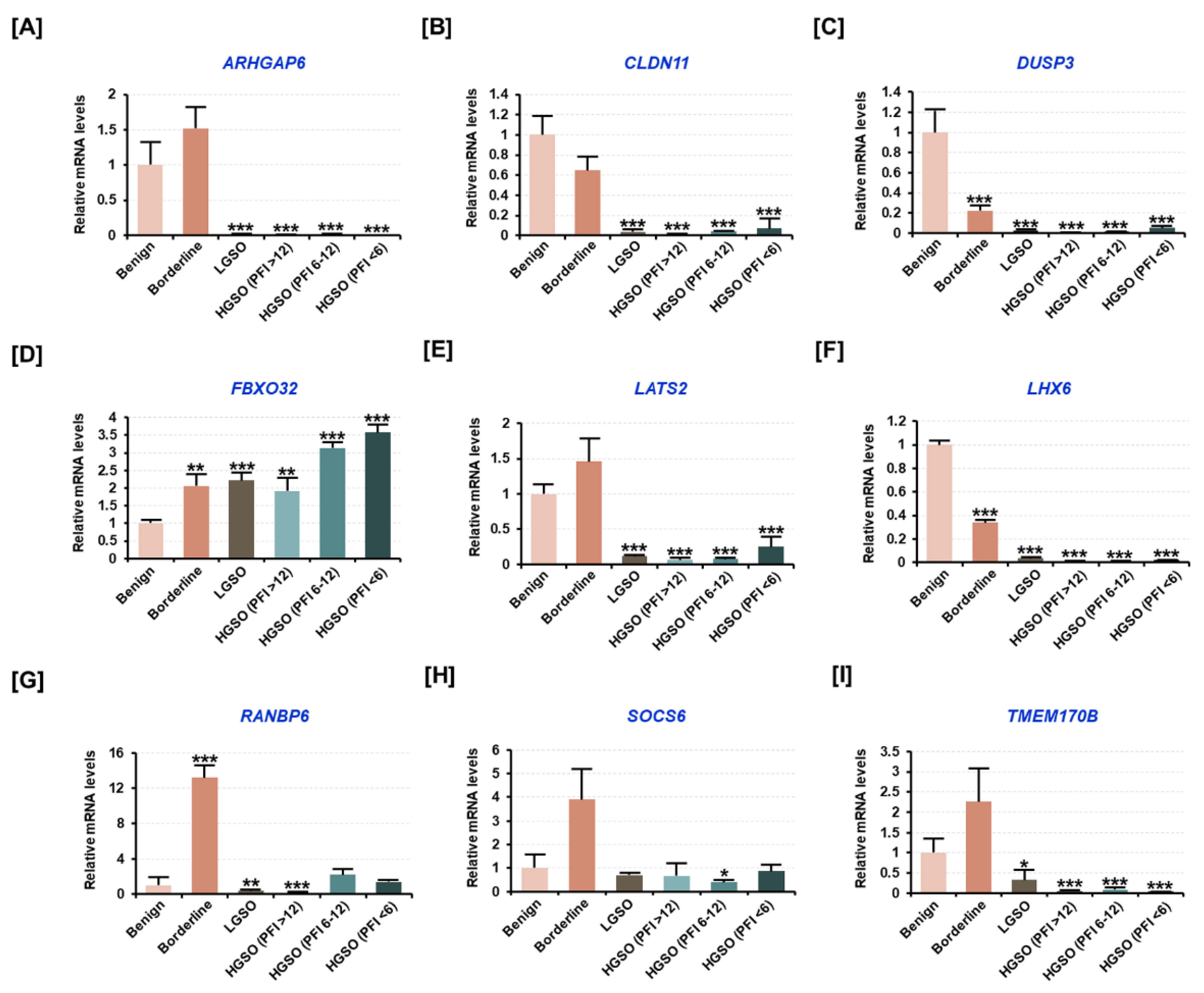
2.2. Target Genes Presumed to Be Regulated by Candidate miRNAs Were Altered by the Malignancy of Ovarian Tissue
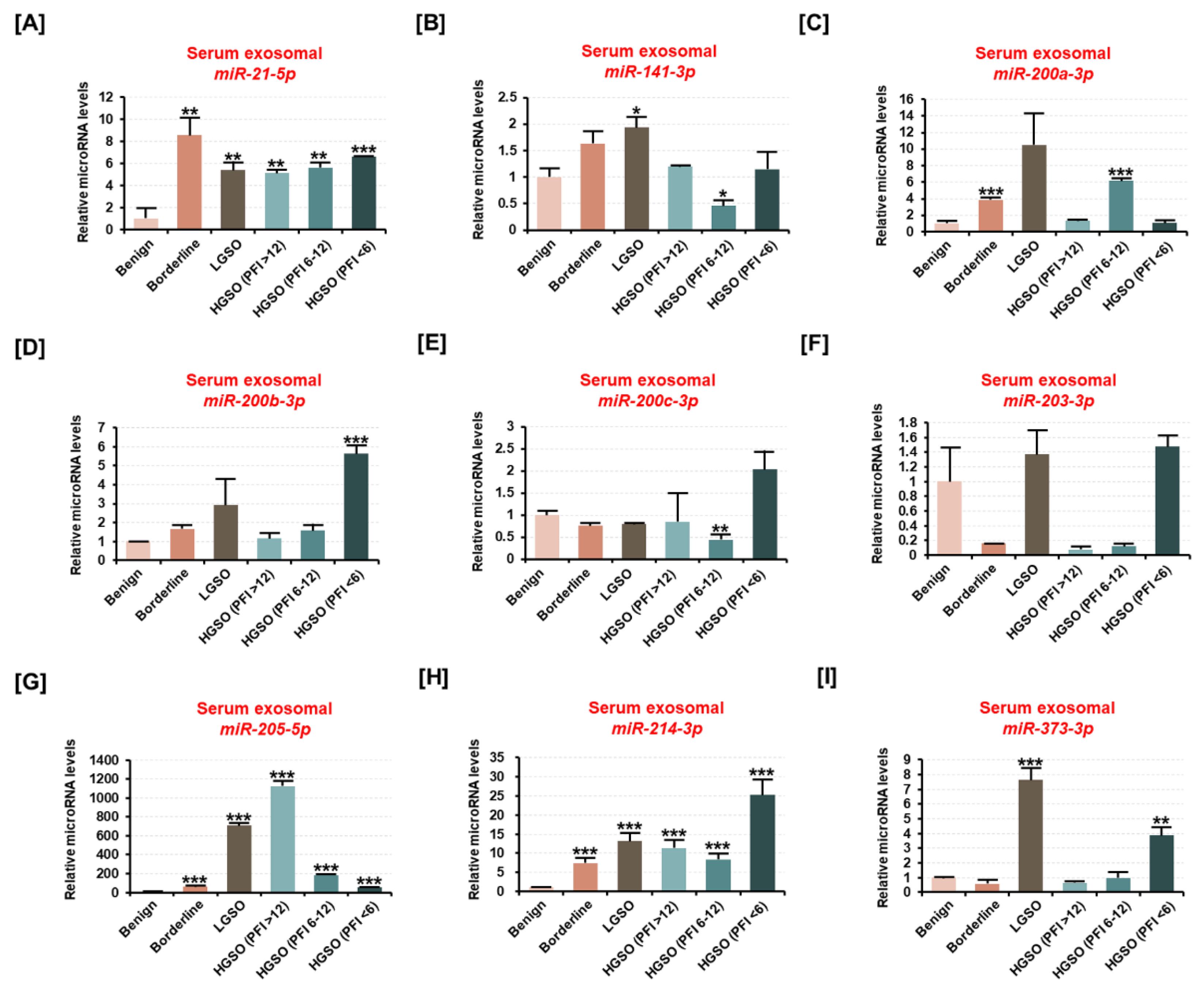
2.3. The Expression of Ovarian Cancer Patient-Derived Exosomal miR-214-3p Increases with Malignancy
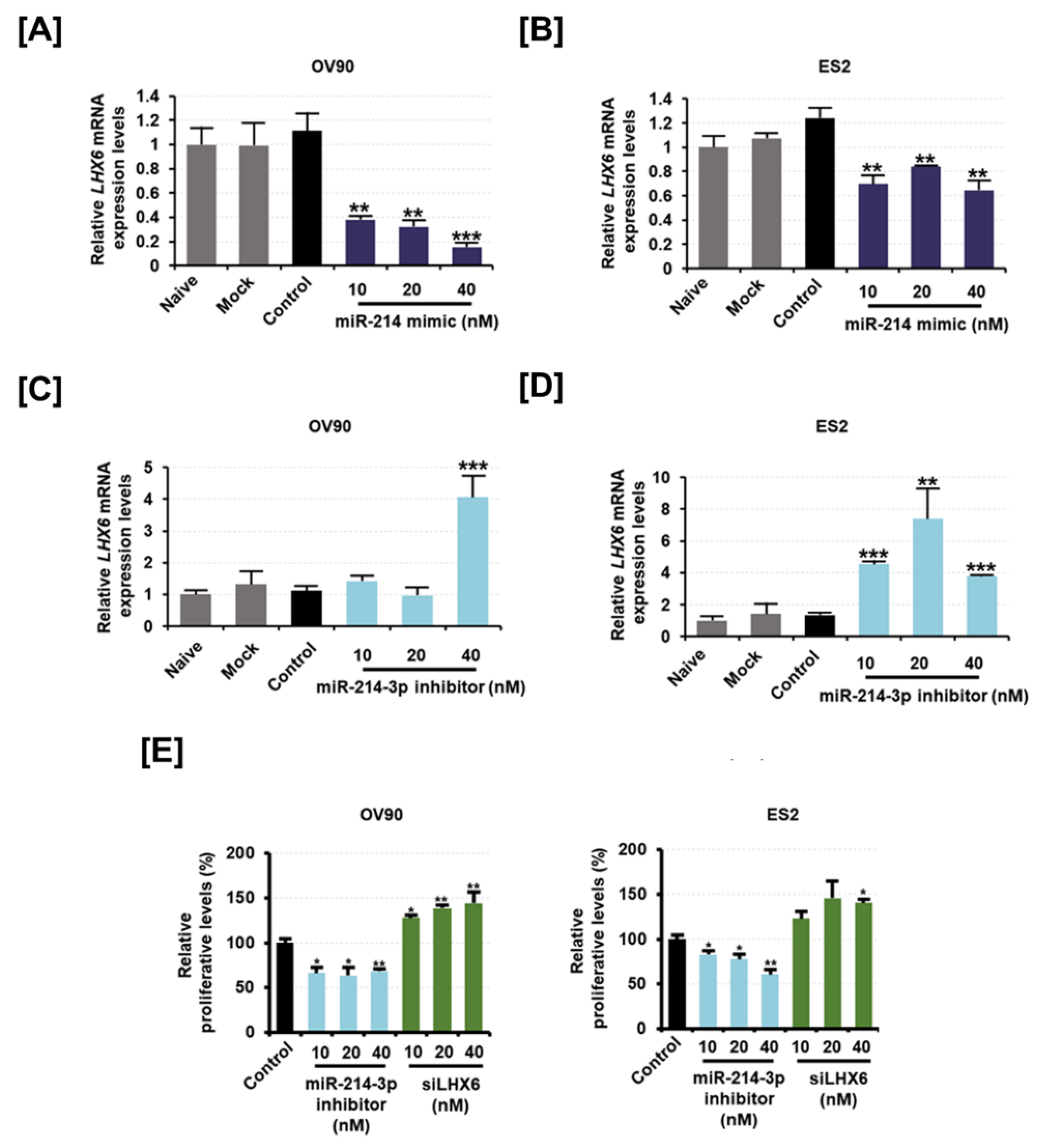
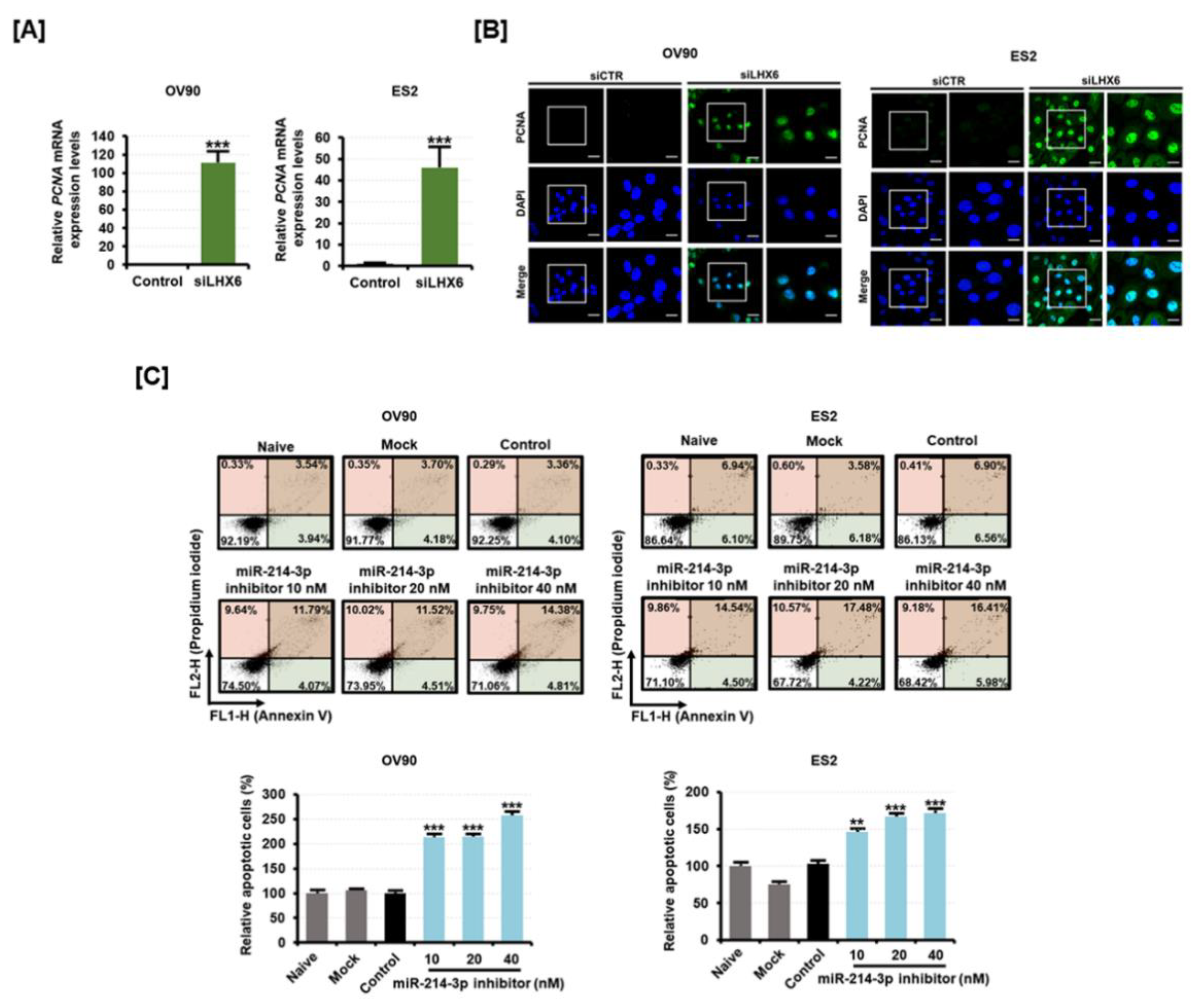
2.4. Inhibition of LXH6 Increases Proliferating Cell Nuclear Antigen (PCNA) Expression While Inhibition of miR-214-3p Induces Apoptosis on Epithelial Ovarian Cancer Cells
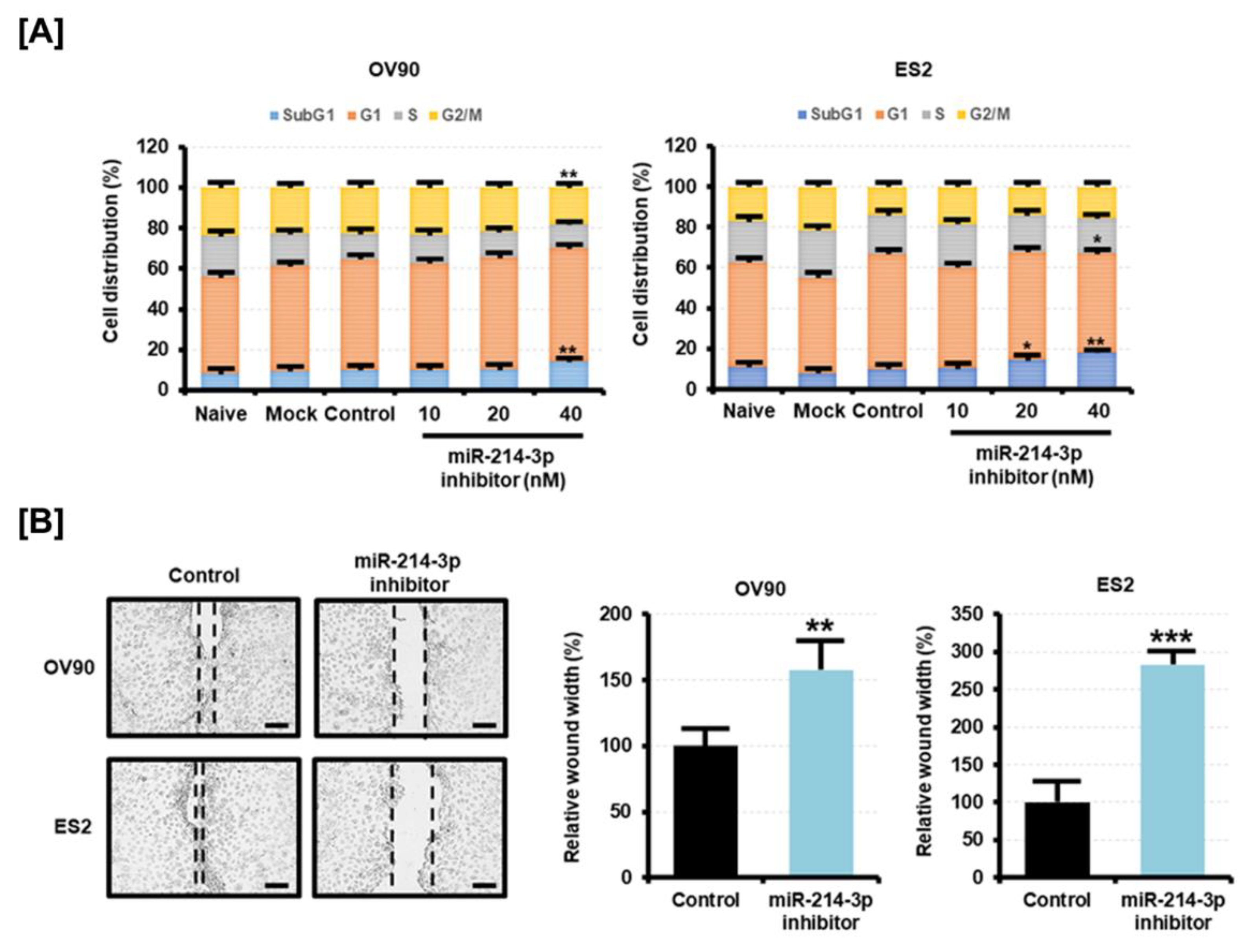
2.5. Inhibition of miR-214-3p Expression Is Involved with the Cell Cycle and Migration of Epithelial Ovarian Cancer Cells
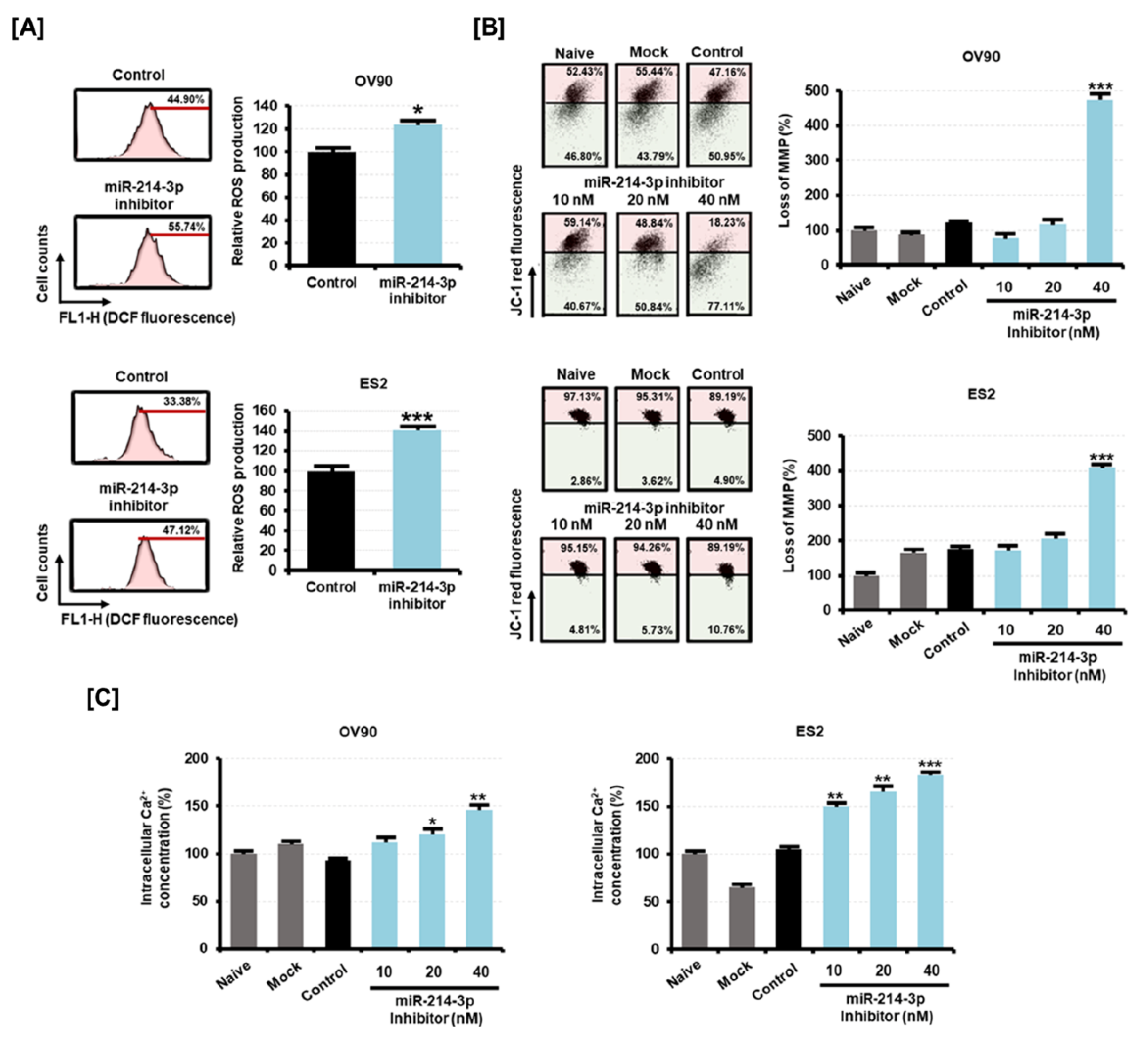
2.6. Mitochondrial Damage Due to Elevated Intracellular ROS Is Induced by the Inhibition of miR-214-3p Expression
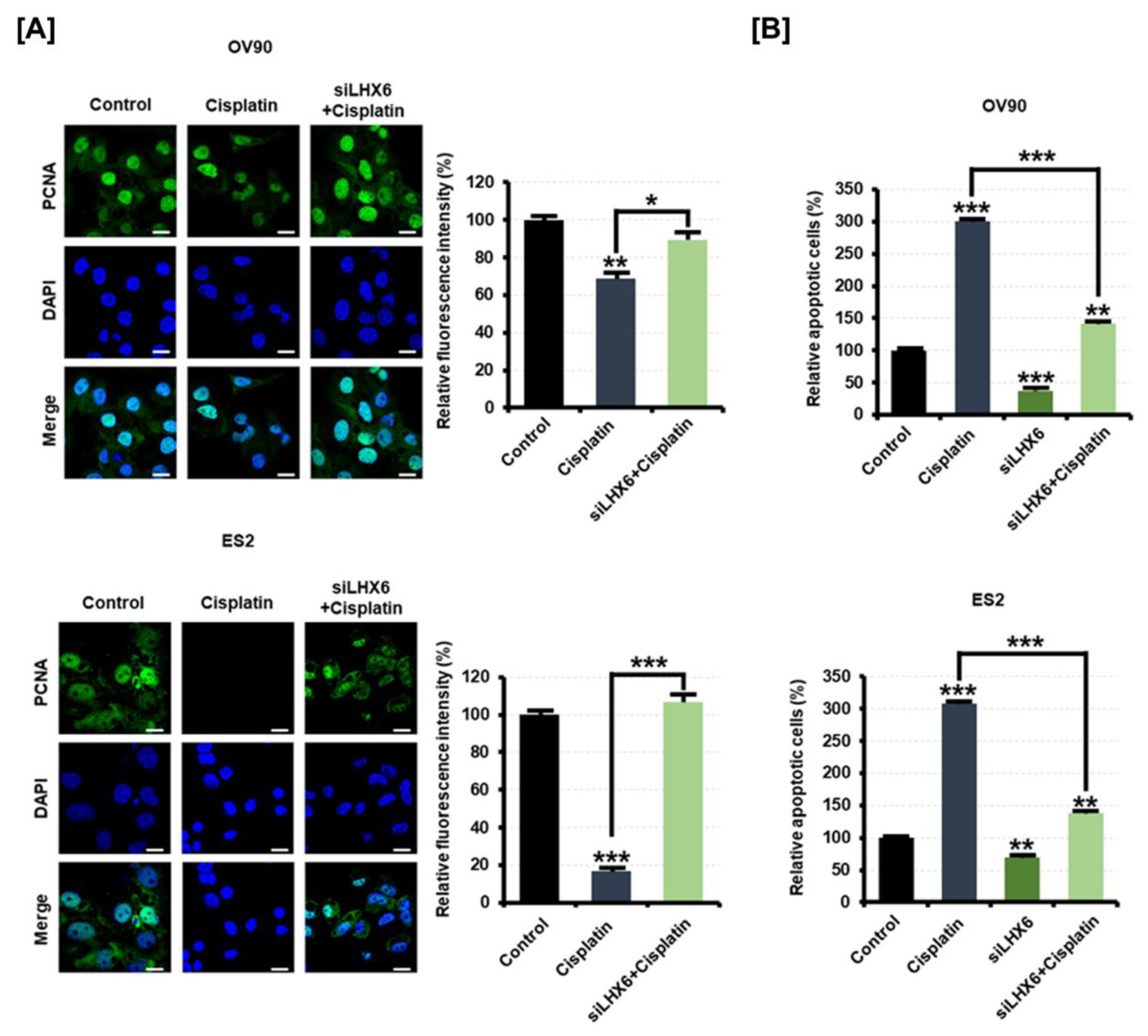
2.7. Inhibition of LHX6 Induces Resistance to Cisplatin in Epithelial Ovarian Cancer Cells
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Human Tissues and Serum
4.3. RNA Extraction and Quantitative RT-PCR
4.4. Quantitative RT-PCR Analysis for microRNA
4.5. Exosome Isolation
4.6. Transfection of microRNA Mimics, Inhibitors, and siRNA
4.7. Proliferation Assay
4.8. Immunofluorescence Microscopy
4.9. Determination of Apoptosis by Annexin V and Propidium Iodide (PI) Staining
4.10. Cell Cycle Analysis
4.11. Migration Assay
4.12. Cellular ROS Determination
4.13. JC-1 Mitochondrial Membrane Potential Assay
4.14. Measurement of the Intracellular Free Ca2+ Concentration
4.15. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jayson, G.C.; Kohn, E.C.; Kitchener, H.C.; Ledermann, J.A. Ovarian cancer. Lancet 2014, 384, 1376–1388. [Google Scholar] [CrossRef]
- Ambros, V. MicroRNA pathways in flies and worms: Growth, death, fat, stress, and timing. Cell 2003, 113, 673–676. [Google Scholar] [CrossRef]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef] [PubMed]
- Voorhoeve, P.M.; le Sage, C.; Schrier, M.; Gillis, A.J.; Stoop, H.; Nagel, R.; Liu, Y.P.; van Duijse, J.; Drost, J.; Griekspoor, A.; et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Adv. Exp. Med. Biol. 2007, 604, 17–46. [Google Scholar]
- Xiaohong, Z.; Lichun, F.; Na, X.; Kejian, Z.; Xiaolan, X.; Shaosheng, W. MiR-203 promotes the growth and migration of ovarian cancer cells by enhancing glycolytic pathway. Tumor Biol. 2016, 37, 14989–14997. [Google Scholar] [CrossRef]
- Li, J.; Hu, K.; Gong, G.; Zhu, D.; Wang, Y.; Liu, H.; Wu, X. Upregulation of MiR-205 transcriptionally suppresses SMAD4 and PTEN and contributes to human ovarian cancer progression. Sci. Rep. 2017, 7, 41330. [Google Scholar] [CrossRef]
- Ibrahim, F.F.; Jamal, R.; Syafruddin, S.E.; Ab Mutalib, N.S.; Saidin, S.; MdZin, R.R.; Hossain Mollah, M.M.; Mokhtar, N.M. MicroRNA-200c and microRNA-31 regulate proliferation, colony formation, migration and invasion in serous ovarian cancer. J. Ovarian Res. 2015, 8, 56. [Google Scholar] [CrossRef]
- Nakano, H.; Yamada, Y.; Miyazawa, T.; Yoshida, T. Gain-of-function microRNA screens identify miR-193a regulating proliferation and apoptosis in epithelial ovarian cancer cells. Int. J. Oncol. 2013, 42, 1875–1882. [Google Scholar] [CrossRef]
- Yu, X.; Zheng, H.; Chan, M.T.; Wu, W.K. Modulation of chemoresponsiveness to platinum-based agents by microRNAs in cancer. Am. J. Cancer Res. 2017, 7, 1769–1778. [Google Scholar]
- Teng, Y.; Su, X.; Zhang, X.; Zhang, Y.; Li, C.; Niu, W.; Liu, C.; Qu, K. Mirna-200a/c as potential biomarker in epithelial ovarian cancer (EOC): Evidence based on miRNA meta-signature and clinical investigations. Oncotarget 2016, 7, 81621–81633. [Google Scholar] [CrossRef]
- Baez-Vega, P.M.; Echevarria Vargas, I.M.; Valiyeva, F.; Encarnacion-Rosado, J.; Roman, A.; Flores, J.; Marcos-Martinez, M.J.; Vivas-Mejia, P.E. Targeting miR-21-3p inhibits proliferation and invasion of ovarian cancer cells. Oncotarget 2016, 7, 36321–36337. [Google Scholar] [CrossRef] [PubMed]
- Ying, H.C.; Xu, H.Y.; Lv, J.; Ying, T.S.; Yang, Q. MicroRNA signatures of platinum-resistance in ovarian cancer. Eur. J. Gynaecol. Oncol. 2015, 36, 16–20. [Google Scholar] [PubMed]
- Marchini, S.; Cavalieri, D.; Fruscio, R.; Calura, E.; Garavaglia, D.; Fuso Nerini, I.; Mangioni, C.; Cattoretti, G.; Clivio, L.; Beltrame, L.; et al. Association between miR-200c and the survival of patients with stage I epithelial ovarian cancer: A retrospective study of two independent tumour tissue collections. Lancet Oncol. 2011, 12, 273–285. [Google Scholar] [CrossRef]
- Feng, M.; Zhao, J.; Wang, L.; Liu, J. Upregulated Expression of Serum Exosomal microRNAs as Diagnostic Biomarkers of Lung Adenocarcinoma. Ann. Clin. Lab. Sci. 2018, 48, 712–718. [Google Scholar] [PubMed]
- Frassanito, M.A.; Desantis, V.; Di Marzo, L.; Craparotta, I.; Beltrame, L.; Marchini, S.; Annese, T.; Visino, F.; Arciuli, M.; Saltarella, I.; et al. Bone marrow fibroblasts overexpress miR-27b and miR-214 in step with multiple myeloma progression, dependent on tumour cell-derived exosomes. J. Pathol. 2019, 247, 241–253. [Google Scholar] [CrossRef]
- Meng, X.; Muller, V.; Milde-Langosch, K.; Trillsch, F.; Pantel, K.; Schwarzenbach, H. Circulating Cell-Free miR-373, miR-200a, miR-200b and miR-200c in Patients with Epithelial Ovarian Cancer. Adv. Exp. Med. Biol. 2016, 924, 3–8. [Google Scholar] [CrossRef]
- Liu, J.; Chen, W.; Zhang, H.; Liu, T.; Zhao, L. miR-214 targets the PTEN-mediated PI3K/Akt signaling pathway and regulates cell proliferation and apoptosis in ovarian cancer. Oncol. Lett. 2017, 14, 5711–5718. [Google Scholar] [CrossRef]
- Flynt, A.S.; Li, N.; Thatcher, E.J.; Solnica-Krezel, L.; Patton, J.G. Zebrafish miR-214 modulates Hedgehog signaling to specify muscle cell fate. Nat. Genet. 2007, 39, 259–263. [Google Scholar] [CrossRef]
- Yang, H.; Kong, W.; He, L.; Zhao, J.J.; O’Donnell, J.D.; Wang, J.; Wenham, R.M.; Coppola, D.; Kruk, P.A.; Nicosia, S.V.; et al. MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res. 2008, 68, 425–433. [Google Scholar] [CrossRef]
- Liu, Y.; Zhou, H.; Ma, L.; Hou, Y.; Pan, J.; Sun, C.; Yang, Y.; Zhang, J. MiR-214 suppressed ovarian cancer and negatively regulated semaphorin 4D. Tumor Biol. 2016, 37, 8239–8248. [Google Scholar] [CrossRef]
- Liu, Y.; Lin, J.; Zhai, S.; Sun, C.; Xu, C.; Zhou, H.; Liu, H. MicroRNA-214 Suppresses Ovarian Cancer by Targeting beta-Catenin. Cell Physiol. Biochem. 2018, 45, 1654–1662. [Google Scholar] [CrossRef] [PubMed]
- Sharma, T.; Hamilton, R.; Mandal, C.C. miR-214: A potential biomarker and therapeutic for different cancers. Future Oncol. 2015, 11, 349–363. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Lin, J.; Lin, D.; Zou, C.; Kurata, J.; Lin, R.; He, Z.; Su, Y. Down-regulation of miR-214 reverses erlotinib resistance in non-small-cell lung cancer through up-regulating LHX6 expression. Sci. Rep. 2017, 7, 781. [Google Scholar] [CrossRef] [PubMed]
- Yuan, X.; Zhang, Z.; Jiang, K.; Wang, X.; Li, Y. Preliminary Study of the Role F-Box Protein 32 (FBXO32) in Colorectal Neoplasms Through the Transforming Growth Factor beta (TGF-beta)/Smad4 Signalling Pathway. Med. Sci. Monit. 2018, 24, 1080–1088. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Zhang, M.; Shen, S.; Guo, Y.; Kuang, G.; Yang, Z.; Dong, Z. Aberrant methylation and decreased expression of the TGF-beta/Smad target gene FBXO32 in esophageal squamous cell carcinoma. Cancer 2014, 120, 2412–2423. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Zhang, M.; Guo, Y.; Shen, S.; Guo, X.; Dong, Z. FBXO32, a new TGF-beta/Smad signaling pathway target gene, is epigenetically inactivated in gastric cardia adenocarcinoma. Neoplasma 2015, 62, 646–657. [Google Scholar] [CrossRef][Green Version]
- Taylor, D.D.; Gercel-Taylor, C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol. Oncol. 2008, 110, 13–21. [Google Scholar] [CrossRef]
- Melo, S.A.; Sugimoto, H.; O’Connell, J.T.; Kato, N.; Villanueva, A.; Vidal, A.; Qiu, L.; Vitkin, E.; Perelman, L.T.; Melo, C.A.; et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell 2014, 26, 707–721. [Google Scholar] [CrossRef]
- Nakamura, K.; Sawada, K.; Kobayashi, M.; Miyamoto, M.; Shimizu, A.; Yamamoto, M.; Kinose, Y.; Kimura, T. Role of the Exosome in Ovarian Cancer Progression and Its Potential as a Therapeutic Target. Cancers 2019, 11, 1147. [Google Scholar] [CrossRef]
- Chen, X.; Zhou, J.; Li, X.; Wang, X.; Lin, Y.; Wang, X. Exosomes derived from hypoxic epithelial ovarian cancer cells deliver microRNAs to macrophages and elicit a tumor-promoted phenotype. Cancer Lett. 2018, 435, 80–91. [Google Scholar] [CrossRef]
- Gupta, V.; Yull, F.; Khabele, D. Bipolar Tumor-Associated Macrophages in Ovarian Cancer as Targets for Therapy. Cancers 2018, 10, 366. [Google Scholar] [CrossRef] [PubMed]
- Nathalia, E.; Theardy, M.S.; Elvira, S.; Rosellinny, G.; Liyanto, A.S.; Utama, M.P.; Sumarpo, A. Downregulation of tumor-suppressor gene LHX6 in cancer: A systematic review. Rom. J. Intern. Med. 2018, 56, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Xie, L. LHX6 inhibits breast cancer cell proliferation and invasion via repression of the Wnt/beta-catenin signaling pathway. Mol. Med. Rep. 2015, 12, 4634–4639. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.B.; Jiang, X.; Han, F.; Li, Y.H.; Chen, H.Q.; Liu, Y.; Cao, J.; Liu, J.Y. LHX6 acts as a novel potential tumour suppressor with epigenetic inactivation in lung cancer. Cell Death Dis. 2013, 4, e882. [Google Scholar] [CrossRef]
- Mehrabi, S.; Partridge, E.E.; Seffens, W.; Yao, X.; Aikhionbare, F.O. Oxidatively modified proteins in the serous subtype of ovarian carcinoma. Biomed. Res. Int. 2014, 2014, 585083. [Google Scholar] [CrossRef]
- Jasen, P. From the “silent killer” to the “whispering disease”: Ovarian cancer and the uses of metaphor. Med. Hist 2009, 53, 489–512. [Google Scholar] [CrossRef]
- Cohen, S.; Mehrabi, S.; Yao, X.; Millingen, S.; Aikhionbare, F.O. Reactive Oxygen Species and Serous Epithelial Ovarian Adenocarcinoma. Cancer Res. J. 2016, 4, 106–114. [Google Scholar] [CrossRef]
- Park, S.; Lim, W.; Jeong, W.; Bazer, F.W.; Lee, D.; Song, G. Sideroxylin (Callistemon lanceolatus) suppressed cell proliferation and increased apoptosis in ovarian cancer cells accompanied by mitochondrial dysfunction, the generation of reactive oxygen species, and an increase of lipid peroxidation. J. Cell Physiol. 2018, 233, 8597–8604. [Google Scholar] [CrossRef]
- Lim, W.; An, Y.; Yang, C.; Bazer, F.W.; Song, G. Chrysophanol induces cell death and inhibits invasiveness via mitochondrial calcium overload in ovarian cancer cells. J. Cell Biochem. 2018, 119, 10216–10227. [Google Scholar] [CrossRef]
- Wang, Y.; Ma, J.; Shen, H.; Wang, C.; Sun, Y.; Howell, S.B.; Lin, X. Reactive oxygen species promote ovarian cancer progression via the HIF-1alpha/LOX/E-cadherin pathway. Oncol. Rep. 2014, 32, 2150–2158. [Google Scholar] [CrossRef]
- Cui, C.; Merritt, R.; Fu, L.; Pan, Z. Targeting calcium signaling in cancer therapy. Acta Pharm. Sin. B 2017, 7, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Jin, Y.; Xu, T.; Zhou, S.; Cui, M. MicroRNA-215 targets NOB1 and inhibits growth and invasion of epithelial ovarian cancer. Am. J. Transl. Res. 2017, 9, 466–477. [Google Scholar] [PubMed]
- Liu, J.; Mao, Q.; Liu, Y.; Hao, X.; Zhang, S.; Zhang, J. Analysis of miR-205 and miR-155 expression in the blood of breast cancer patients. Chin. J. Cancer Res. 2013, 25, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Ng, E.K.; Chong, W.W.; Jin, H.; Lam, E.K.; Shin, V.Y.; Yu, J.; Poon, T.C.; Ng, S.S.; Sung, J.J. Differential expression of microRNAs in plasma of patients with colorectal cancer: A potential marker for colorectal cancer screening. Gut 2009, 58, 1375–1381. [Google Scholar] [CrossRef]
- Lim, W.; Yang, C.; Bazer, F.W.; Song, G. Chrysophanol Induces Apoptosis of Choriocarcinoma Through Regulation of ROS and the AKT and ERK1/2 Pathways. J. Cell Physiol. 2017, 232, 331–339. [Google Scholar] [CrossRef]
| Gene Symbol | Regulatory miRNA | Sense Primer (5′→3′) | Antisense Primer (5′→3′) | Accession Number |
|---|---|---|---|---|
| ARHGAP6 | miR-200b-3p/miR-200c-3p | TGTCGTCGTCAAAGTCAAGG | CCAGAGGAACCTGTCATTCC | NM_001287242.2 |
| CLDN11 | miR-205-5p | TGGTGTTTTGCTCATTCTGC | GCCTGCATACAGGGAGTAGC | NM_001185056.2 |
| DUSP3 | miR-200a-3p | ATCTCAACGACCTGCTCTCG | GGGTGATGCCTAGTTTCTGC | NM_004090.4 |
| FBXO32 | miR-214-3p | TTGTCCGATGTTACCCAAGG | GCAGCTCTCGGGTTATTGG | NM_001242463.2 |
| LATS2 | miR-373-3p | AGTGACACTTCCCTGGATGC | CCCGATTCATTAGCAAAAGG | NM_014572.3 |
| LHX6 | miR-214-3p | GGACCGATATCTGCTAAGG | CGAATCGGCTGAAGTAGTCC | NM_001242333.2 |
| RANBP6 | miR-141-3p | TTCAGGACTTGAAGCAAAAGC | CCACTCGAACATTGTCATGG | NM_001243202.2 |
| SOCS6 | miR-203-3p | ACCATTGCTACCTCCAATGC | TGACAGCGCATACTTTCAGC | NM_004232.4 |
| TMEM170B | miR-200a-3p | CGACCACTCCATGATCAACC | GAGAGCCCAGAGGAAGATCC | NM_001100829.3 |
| miRNA Name | miRNA Sequence (5′→3′) | NCBI Gene ID of Precursor miRNA |
|---|---|---|
| miR-21-5p | UAGCUUAUCAGACUGAUGUUGA | 406991 |
| miR-141-3p | UAACACUGUCUGGUAAAGAUGG | 406933 |
| miR-200a-3p | UAACACUGUCUGGUAACGAUGU | 406983 |
| miR-200b-3p | UAAUACUGCCUGGUAAUGAUGA | 406984 |
| miR-200c-3p | UAAUACUGCCGGGUAAUGAUGGA | 406985 |
| miR-203-3p | GUGAAAUGUUUAGGACCACUAG | 406986 |
| miR-205-5p | UCCUUCAUUCCACCGGAGUCUG | 406988 |
| miR-214-3p | ACAGCAGGCACAGACAGGCAGU | 406996 |
| miR-373-3p | GAAGUGCUUCGAUUUUGGGGUGU | 442918 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, C.; Kim, H.S.; Park, S.J.; Lee, E.J.; Kim, S.I.; Song, G.; Lim, W. Inhibition of miR-214-3p Aids in Preventing Epithelial Ovarian Cancer Malignancy by Increasing the Expression of LHX6. Cancers 2019, 11, 1917. https://doi.org/10.3390/cancers11121917
Yang C, Kim HS, Park SJ, Lee EJ, Kim SI, Song G, Lim W. Inhibition of miR-214-3p Aids in Preventing Epithelial Ovarian Cancer Malignancy by Increasing the Expression of LHX6. Cancers. 2019; 11(12):1917. https://doi.org/10.3390/cancers11121917
Chicago/Turabian StyleYang, Changwon, Hee Seung Kim, Soo Jin Park, Eun Ji Lee, Se Ik Kim, Gwonhwa Song, and Whasun Lim. 2019. "Inhibition of miR-214-3p Aids in Preventing Epithelial Ovarian Cancer Malignancy by Increasing the Expression of LHX6" Cancers 11, no. 12: 1917. https://doi.org/10.3390/cancers11121917
APA StyleYang, C., Kim, H. S., Park, S. J., Lee, E. J., Kim, S. I., Song, G., & Lim, W. (2019). Inhibition of miR-214-3p Aids in Preventing Epithelial Ovarian Cancer Malignancy by Increasing the Expression of LHX6. Cancers, 11(12), 1917. https://doi.org/10.3390/cancers11121917







