MicroRNA-Mediated Metabolic Reprograming in Renal Cancer
Abstract
1. Introduction
2. Results
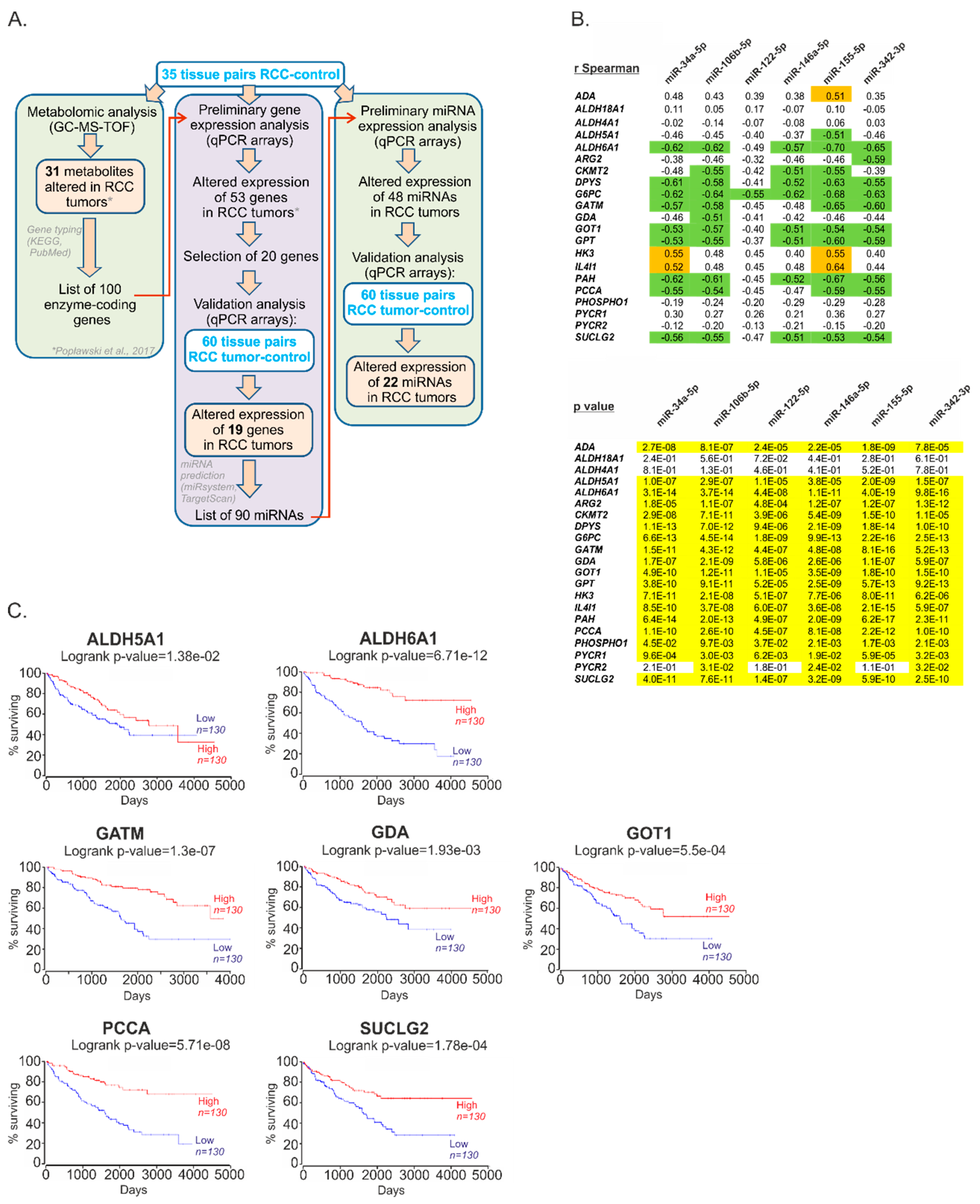
2.1. The Expression of miRs Predicted to Target Metabolic Genes Is Altered in Renal Tumors
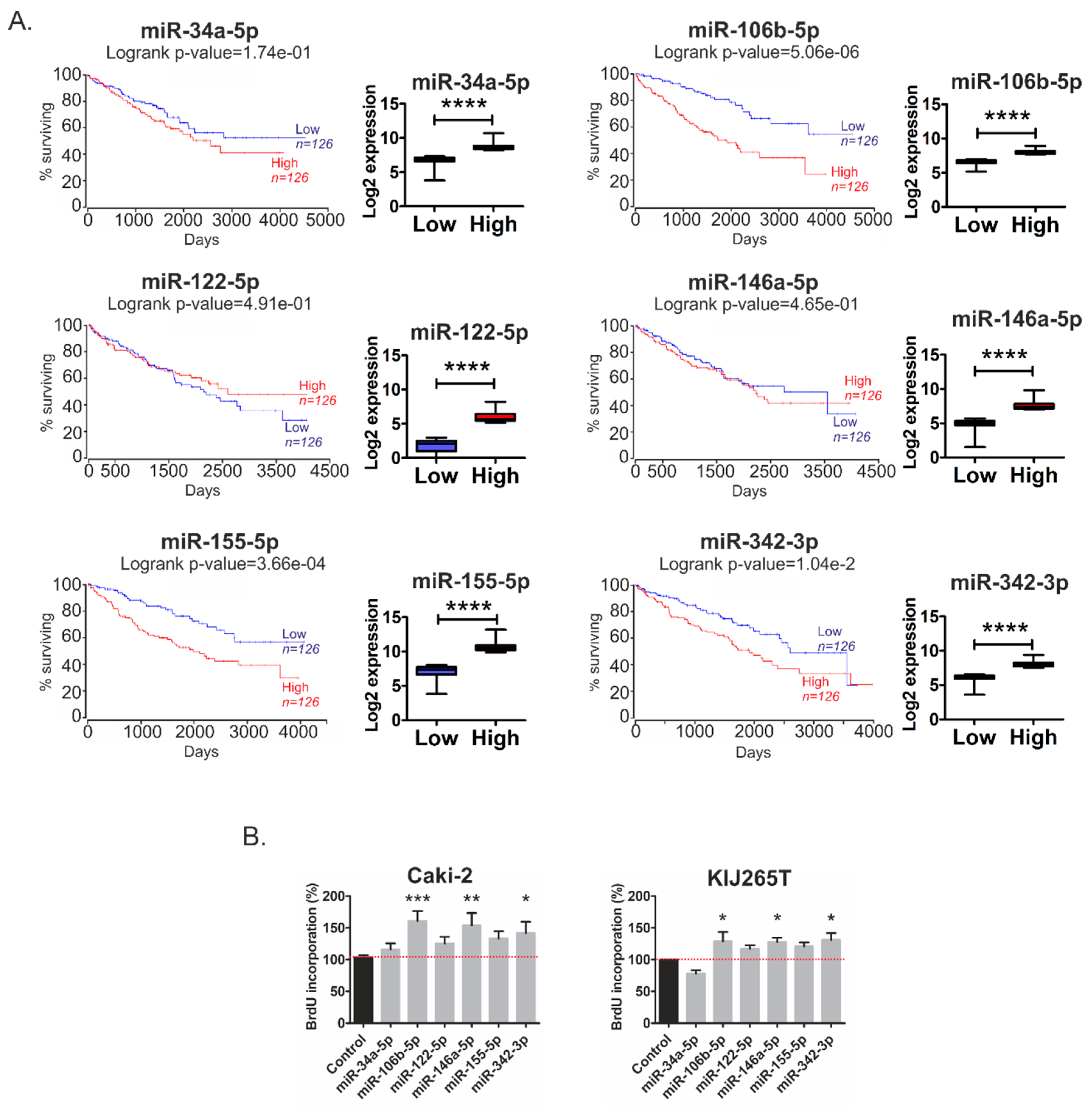
2.2. Metabolic miRNAs Affect Proliferation of RCC Cells and Correlate with Poor Survival of RCC Patients
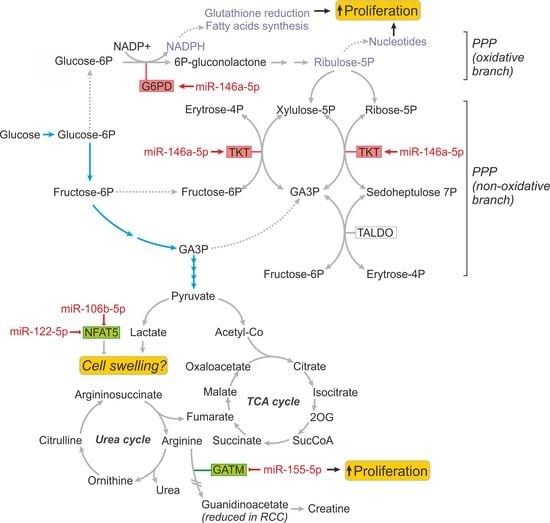
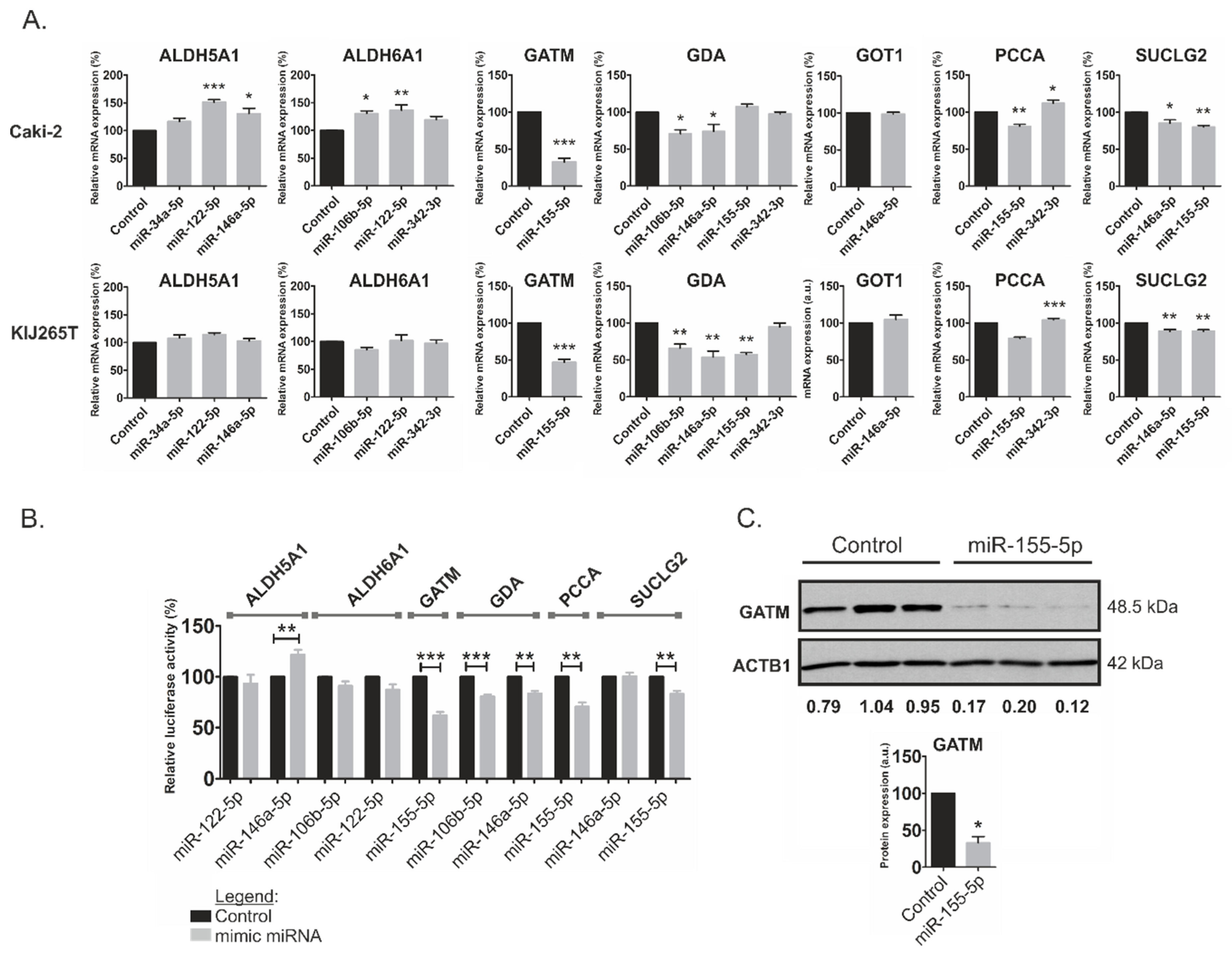
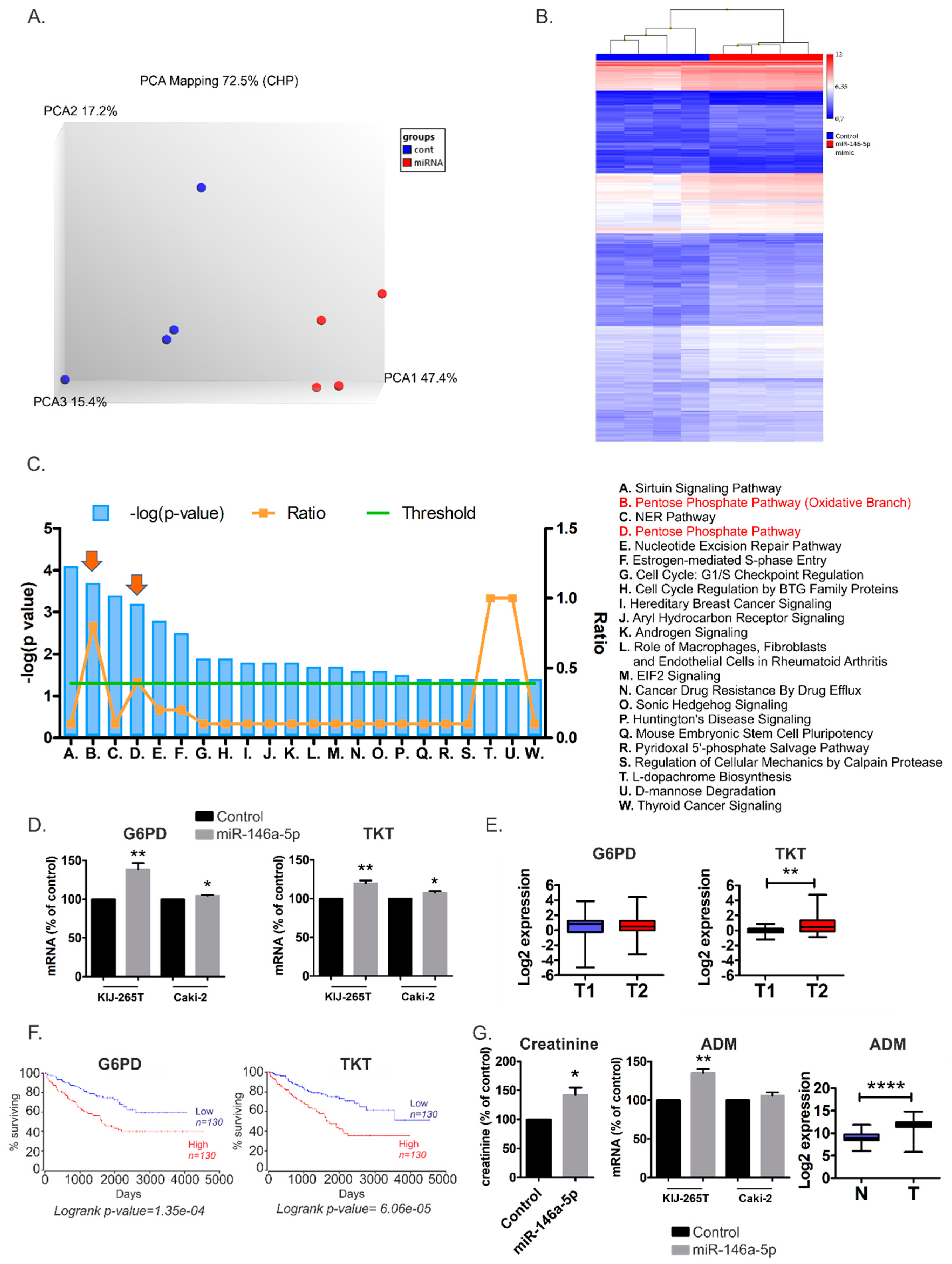
2.3. MiR-146a-5p is a Global Regulator of Key Metabolic Pathways in RCC
2.4. Metabolically-Relevant miRNAs Regulate the Expression of NFAT5
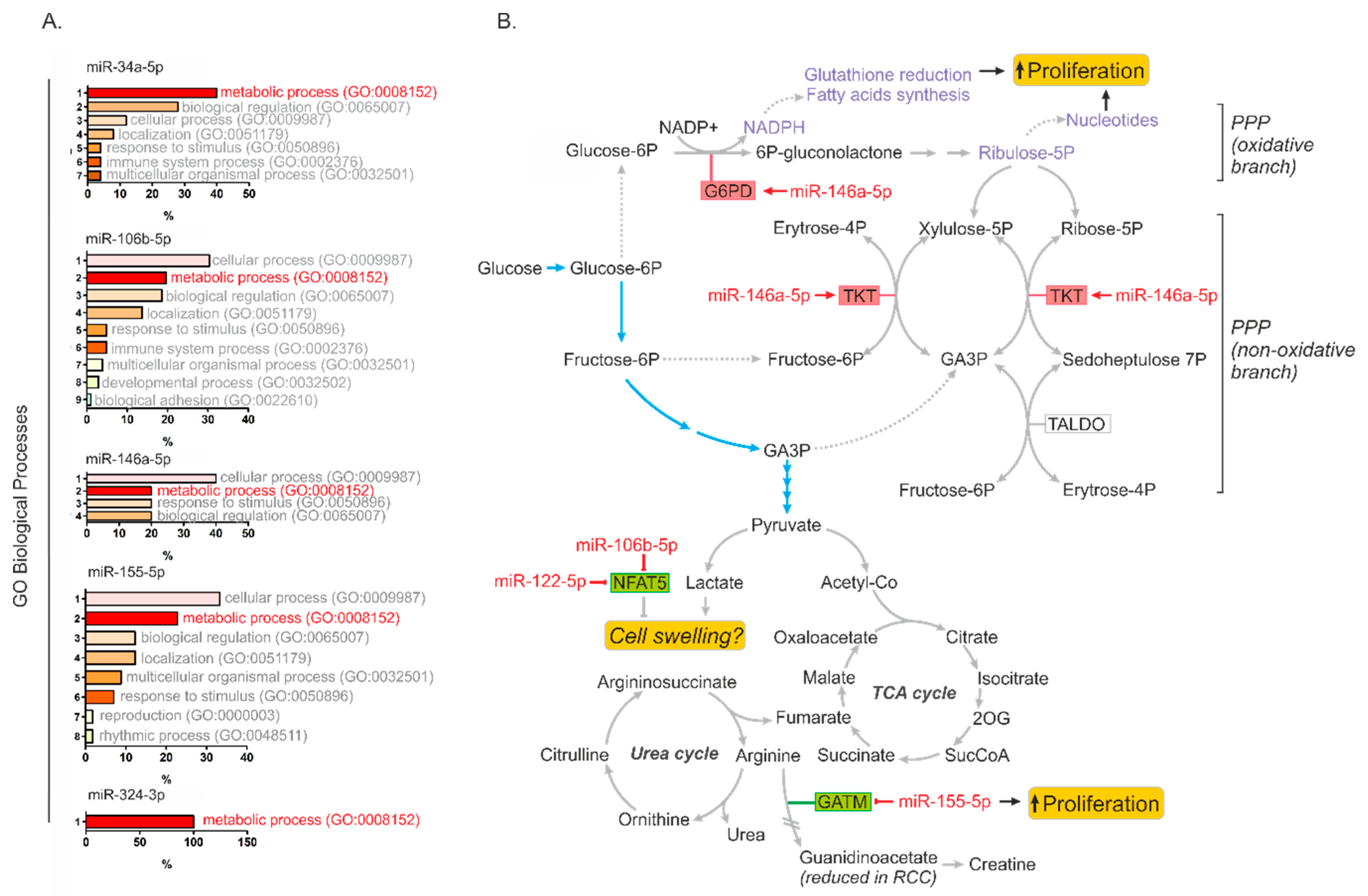
2.5. MiR-34a-5p, miR-106b-5p, miR-146a-5p and miR-155-5p Are PanCancer MetabomiRs
3. Discussion
4. Materials and Methods
4.1. Tissue Samples
4.2. Cell Lines
4.3. Transfections
4.4. Isolations of RNA and Proteins, Reverse Transcription
4.5. Cloning of miRNA Targets Sites and Luciferase Assays
4.6. Western Blots
4.7. Analysis of Proliferation
4.8. Transcriptomic Analysis
4.9. Metabolomic Analysis
4.10. Bioinformatics Analysis
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ljungberg, B.; Bensalah, K.; Canfield, S.; Dabestani, S.; Hofmann, F.; Hora, M.; Kuczyk, M.A.; Lam, T.; Marconi, L.; Merseburger, A.S.; et al. EAU guidelines on renal cell carcinoma: 2014 update. Eur. Urol. 2015, 67, 913–924. [Google Scholar] [CrossRef] [PubMed]
- Albiges, L.; Choueiri, T.; Escudier, B.; Galsky, M.; George, D.; Hofmann, F.; Lam, T.; Motzer, R.; Mulders, P.; Porta, C.; et al. A systematic review of sequencing and combinations of systemic therapy in metastatic renal cancer. Eur. Urol. 2015, 67, 100–110. [Google Scholar] [CrossRef] [PubMed]
- Wettersten, H.I.; Aboud, O.A.; Lara, P.N., Jr.; Weiss, R.H. Metabolic reprogramming in clear cell renal cell carcinoma. Nat. Rev. Nephrol. 2017, 13, 410–419. [Google Scholar] [CrossRef] [PubMed]
- Poplawski, P.; Tohge, T.; Boguslawska, J.; Rybicka, B.; Tanski, Z.; Trevino, V.; Fernie, A.R.; Piekielko-Witkowska, A. Integrated transcriptomic and metabolomic analysis shows that disturbances in metabolism of tumor cells contribute to poor survival of RCC patients. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 744–752. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, K.; Karolina, D.S.; Sepramaniam, S.; Armugam, A.; Wintour, E.M.; Bertram, J.F.; Jeyaseelan, K. Role of microRNAs in kidney homeostasis and disease. Kidney Int. 2012, 81, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Boguslawska, J.; Rodzik, K.; Poplawski, P.; Kedzierska, H.; Rybicka, B.; Sokol, E.; Tanski, Z.; Piekielko-Witkowska, A. TGF-beta1 targets a microRNA network that regulates cellular adhesion and migration in renal cancer. Cancer Lett. 2018, 412, 155–169. [Google Scholar] [CrossRef]
- Morris, M.R.; Latif, F. The epigenetic landscape of renal cancer. Nat. Rev. Nephrol. 2017, 13, 47–60. [Google Scholar] [CrossRef]
- Wang, Y.P.; Li, K.B. Correlation of expression profiles between microRNAs and mRNA targets using NCI-60 data. BMC Genomics 2009, 10, 218. [Google Scholar] [CrossRef]
- Heggermont, W.A.; Papageorgiou, A.P.; Quaegebeur, A.; Deckx, S.; Carai, P.; Verhesen, W.; Eelen, G.; Schoors, S.; van Leeuwen, R.; Alekseev, S.; et al. Inhibition of MicroRNA-146a and Overexpression of Its Target Dihydrolipoyl Succinyltransferase Protect Against Pressure Overload-Induced Cardiac Hypertrophy and Dysfunction. Circulation 2017, 136, 747–761. [Google Scholar] [CrossRef]
- Mori, Y.; Nishikimi, T.; Kobayashi, N.; Ono, H.; Kangawa, K.; Matsuoka, H. Long-term adrenomedullin infusion improves survival in malignant hypertensive rats. Hypertension 2002, 40, 107–113. [Google Scholar] [CrossRef]
- Nishikimi, T.; Mori, Y.; Kobayashi, N.; Tadokoro, K.; Wang, X.; Akimoto, K.; Yoshihara, F.; Kangawa, K.; Matsuoka, H. Renoprotective effect of chronic adrenomedullin infusion in Dahl salt-sensitive rats. Hypertension 2002, 39, 1077–1082. [Google Scholar] [CrossRef] [PubMed]
- Nishikimi, T.; Yoshihara, F.; Kanazawa, A.; Okano, I.; Horio, T.; Nagaya, N.; Yutani, C.; Matsuo, H.; Matsuoka, H.; Kangawa, K. Role of increased circulating and renal adrenomedullin in rats with malignant hypertension. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2001, 281, R2079–R2087. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.P.; Lee, C.Y.; Tsai, M.H.; Chiu, Y.C.; Hsiao, C.K.; Lai, L.C.; Chuang, E.Y. miRSystem: An integrated system for characterizing enriched functions and pathways of microRNA targets. PLoS ONE 2012, 7, e42390. [Google Scholar] [CrossRef] [PubMed]
- Ho, S.N. The role of NFAT5/TonEBP in establishing an optimal intracellular environment. Arch Biochem. Biophys. 2003, 413, 151–157. [Google Scholar] [CrossRef]
- The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 2013, 499, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Pinthus, J.H.; Whelan, K.F.; Gallino, D.; Lu, J.P.; Rothschild, N. Metabolic features of clear-cell renal cell carcinoma: Mechanisms and clinical implications. Can. Urol. Assoc. J. 2011, 5, 274–282. [Google Scholar] [CrossRef] [PubMed]
- Lucarelli, G.; Galleggiante, V.; Rutigliano, M.; Sanguedolce, F.; Cagiano, S.; Bufo, P.; Lastilla, G.; Maiorano, E.; Ribatti, D.; Giglio, A.; et al. Metabolomic profile of glycolysis and the pentose phosphate pathway identifies the central role of glucose-6-phosphate dehydrogenase in clear cell-renal cell carcinoma. Oncotarget 2015, 6, 13371–13386. [Google Scholar] [CrossRef]
- Markkanen, T.; Peltola, O.; Forsstrom, J.; Himanen, P. Pentose phosphate pathway of erythrocytes in uremia. Acta Haematol. 1972, 48, 269–277. [Google Scholar] [CrossRef]
- Liu, L.; Pang, X.L.; Shang, W.J.; Xie, H.C.; Wang, J.X.; Feng, G.W. Over-expressed microRNA-181a reduces glomerular sclerosis and renal tubular epithelial injury in rats with chronic kidney disease via down-regulation of the TLR/NF-kappaB pathway by binding to CRY1. Mol. Med. 2018, 24, 49. [Google Scholar] [CrossRef]
- Wei, Q.; Sun, H.; Song, S.; Liu, Y.; Liu, P.; Livingston, M.J.; Wang, J.; Liang, M.; Mi, Q.S.; Huo, Y.; et al. MicroRNA-668 represses MTP18 to preserve mitochondrial dynamics in ischemic acute kidney injury. J. Clin. Invest 2018, 128, 5448–5464. [Google Scholar] [CrossRef]
- Monteiro, M.S.; Barros, A.S.; Pinto, J.; Carvalho, M.; Pires-Luis, A.S.; Henrique, R.; Jeronimo, C.; Bastos, M.L.; Gil, A.M.; Guedes de Pinho, P. Nuclear Magnetic Resonance metabolomics reveals an excretory metabolic signature of renal cell carcinoma. Sci. Rep. 2016, 6, 37275. [Google Scholar] [CrossRef] [PubMed]
- Campos-Ferraz, P.L.; Gualano, B.; das Neves, W.; Andrade, I.T.; Hangai, I.; Pereira, R.T.; Bezerra, R.N.; Deminice, R.; Seelaender, M.; Lancha, A.H. Exploratory studies of the potential anti-cancer effects of creatine. Amino Acids 2016, 48, 1993–2001. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, C.A.; Askenasy, N.; Jain, R.K.; Koretsky, A.P. Creatine and cyclocreatine treatment of human colon adenocarcinoma xenografts: 31P and 1H magnetic resonance spectroscopic studies. Br. J. Cancer 1999, 79, 278–285. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Miller, E.E.; Evans, A.E.; Cohn, M. Inhibition of rate of tumor growth by creatine and cyclocreatine. Proc. Natl. Acad. Sci. USA 1993, 90, 3304–3308. [Google Scholar] [CrossRef] [PubMed]
- Patra, S.; Ghosh, A.; Roy, S.S.; Bera, S.; Das, M.; Talukdar, D.; Ray, S.; Wallimann, T.; Ray, M. A short review on creatine-creatine kinase system in relation to cancer and some experimental results on creatine as adjuvant in cancer therapy. Amino Acids 2012, 42, 2319–2330. [Google Scholar] [CrossRef]
- Hsin, J.P.; Lu, Y.; Loeb, G.B.; Leslie, C.S.; Rudensky, A.Y. The effect of cellular context on miR-155-mediated gene regulation in four major immune cell types. Nat. Immunol. 2018, 19, 1137–1145. [Google Scholar] [CrossRef]
- Nam, J.W.; Rissland, O.S.; Koppstein, D.; Abreu-Goodger, C.; Jan, C.H.; Agarwal, V.; Yildirim, M.A.; Rodriguez, A.; Bartel, D.P. Global analyses of the effect of different cellular contexts on microRNA targeting. Mol. Cell 2014, 53, 1031–1043. [Google Scholar] [CrossRef]
- Ho, S.N. Intracellular water homeostasis and the mammalian cellular osmotic stress response. J. Cell Physiol. 2006, 206, 9–15. [Google Scholar] [CrossRef]
- Halterman, J.A.; Kwon, H.M.; Wamhoff, B.R. Tonicity-independent regulation of the osmosensitive transcription factor TonEBP (NFAT5). Am. J. Physiol. Cell Physiol. 2012, 302, C1–C8. [Google Scholar] [CrossRef]
- Shorthouse, D.; Riedel, A.; Kerr, E.; Pedro, L.; Bihary, D.; Samarajiwa, S.; Martins, C.P.; Shields, J.; Hall, B.A. Exploring the role of stromal osmoregulation in cancer and disease using executable modelling. Nat. Commun. 2018, 9, 3011. [Google Scholar] [CrossRef]
- Levy, C.; Khaled, M.; Iliopoulos, D.; Janas, M.M.; Schubert, S.; Pinner, S.; Chen, P.H.; Li, S.; Fletcher, A.L.; Yokoyama, S.; et al. Intronic miR-211 assumes the tumor suppressive function of its host gene in melanoma. Mol. Cell 2010, 40, 841–849. [Google Scholar] [CrossRef] [PubMed]
- Xin, Y.; Cai, H.; Lu, T.; Zhang, Y.; Yang, Y.; Cui, Y. miR-20b Inhibits T Cell Proliferation and Activation via NFAT Signaling Pathway in Thymoma-Associated Myasthenia Gravis. Biomed. Res. Int. 2016, 2016, 9595718. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Li, C.; Guo, T.; Chen, J.; Wang, H.T.; Wang, Y.T.; Xiao, Y.S.; Li, J.; Liu, P.; Liu, Z.S.; et al. Upregulation of DARS2 by HBV promotes hepatocarcinogenesis through the miR-30e-5p/MAPK/NFAT5 pathway. J. Exp. Clin. Cancer Res. 2017, 36, 148. [Google Scholar] [CrossRef] [PubMed]
- Hinske, L.C.; Heyn, J.; Hubner, M.; Rink, J.; Hirschberger, S.; Kreth, S. Intronic miRNA-641 controls its host Gene’s pathway PI3K/AKT and this relationship is dysfunctional in glioblastoma multiforme. Biochem. Biophys. Res. Commun. 2017, 489, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Kastle, M.; Bartel, S.; Geillinger-Kastle, K.; Irmler, M.; Beckers, J.; Ryffel, B.; Eickelberg, O.; Krauss-Etschmann, S. microRNA cluster 106a~363 is involved in T helper 17 cell differentiation. Immunology 2017, 152, 402–413. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Diaz, S.; Valle, N.; Ferrer-Mayorga, G.; Lombardia, L.; Herrera, M.; Dominguez, O.; Segura, M.F.; Bonilla, F.; Hernando, E.; Munoz, A. MicroRNA-22 is induced by vitamin D and contributes to its antiproliferative, antimigratory and gene regulatory effects in colon cancer cells. Hum. Mol. Genet. 2012, 21, 2157–2165. [Google Scholar] [CrossRef]
- Li, W.; Kong, L.B.; Li, J.T.; Guo, Z.Y.; Xue, Q.; Yang, T.; Meng, Y.L.; Jin, B.Q.; Wen, W.H.; Yang, A.G. MiR-568 inhibits the activation and function of CD4(+) T cells and Treg cells by targeting NFAT5. Int. Immunol. 2014, 26, 269–281. [Google Scholar] [CrossRef]
- Luo, Y.; Liu, Y.; Liu, M.; Wei, J.; Zhang, Y.; Hou, J.; Huang, W.; Wang, T.; Li, X.; He, Y.; et al. Sfmbt2 10th intron-hosted miR-466(a/e)-3p are important epigenetic regulators of Nfat5 signaling, osmoregulation and urine concentration in mice. Biochim. Biophys. Acta 2014, 1839, 97–106. [Google Scholar] [CrossRef]
- Pedersen, S.F.; Hoffmann, E.K.; Novak, I. Cell volume regulation in epithelial physiology and cancer. Front Physiol. 2013, 4, 233. [Google Scholar] [CrossRef]
- Okada, Y.; Maeno, E.; Shimizu, T.; Manabe, K.; Mori, S.; Nabekura, T. Dual roles of plasmalemmal chloride channels in induction of cell death. Pflugers Arch 2004, 448, 287–295. [Google Scholar] [CrossRef]
- Sforna, L.; Cenciarini, M.; Belia, S.; Michelucci, A.; Pessia, M.; Franciolini, F.; Catacuzzeno, L. Hypoxia Modulates the Swelling-Activated Cl Current in Human Glioblastoma Cells: Role in Volume Regulation and Cell Survival. J. Cell Physiol. 2017, 232, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Usher-Smith, J.A.; Fraser, J.A.; Bailey, P.S.; Griffin, J.L.; Huang, C.L. The influence of intracellular lactate and H+ on cell volume in amphibian skeletal muscle. J. Physiol. 2006, 573, 799–818. [Google Scholar] [CrossRef] [PubMed]
- Heins, J.; Zwingmann, C. Organic osmolytes in hyponatremia and ammonia toxicity. Metab. Brain Dis. 2010, 25, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Lang, F. Effect of cell hydration on metabolism. Nestle Nutr. Inst. Workshop Ser. 2011, 69, 115–126; discussion 126–130. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.H.; Chao, H.N.; Walker, C.S.; Choong, S.Y.; Phillips, A.; Loomes, K.M. Renal depletion of myo-inositol is associated with its increased degradation in animal models of metabolic disease. Am. J. Physiol. Renal Physiol. 2015, 309, F755–F763. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Research Network; Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Shaw, K.R.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef]
- Hoadley, K.A.; Yau, C.; Hinoue, T.; Wolf, D.M.; Lazar, A.J.; Drill, E.; Shen, R.; Taylor, A.M.; Cherniack, A.D.; Thorsson, V.; et al. Cell-of-Origin Patterns Dominate the Molecular Classification of 10,000 Tumors from 33 Types of Cancer. Cell 2018, 173, 291–304.e6. [Google Scholar] [CrossRef]
- Liu, J.; Lichtenberg, T.; Hoadley, K.A.; Poisson, L.M.; Lazar, A.J.; Cherniack, A.D.; Kovatich, A.J.; Benz, C.C.; Levine, D.A.; Lee, A.V.; et al. An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell 2018, 173, 400–416.e11. [Google Scholar] [CrossRef]
- Kruger, R. Charting a Course to a Cure. Cell 2018, 173, 277. [Google Scholar] [CrossRef]
- Sanchez-Vega, F.; Mina, M.; Armenia, J.; Chatila, W.K.; Luna, A.; La, K.C.; Dimitriadoy, S.; Liu, D.L.; Kantheti, H.S.; Saghafinia, S.; et al. Oncogenic Signaling Pathways in The Cancer Genome Atlas. Cell 2018, 173, 321–337.e10. [Google Scholar] [CrossRef]
- Cooper, L.A.; Demicco, E.G.; Saltz, J.H.; Powell, R.T.; Rao, A.; Lazar, A.J. PanCancer insights from The Cancer Genome Atlas: The pathologist’s perspective. J. Pathol. 2018, 244, 512–524. [Google Scholar] [CrossRef] [PubMed]
- Peng, X.; Chen, Z.; Farshidfar, F.; Xu, X.; Lorenzi, P.L.; Wang, Y.; Cheng, F.; Tan, L.; Mojumdar, K.; Du, D.; et al. Molecular Characterization and Clinical Relevance of Metabolic Expression Subtypes in Human Cancers. Cell Rep. 2018, 23, 255–269.e4. [Google Scholar] [CrossRef] [PubMed]
- Boguslawska, J.; Kedzierska, H.; Poplawski, P.; Rybicka, B.; Tanski, Z.; Piekielko-Witkowska, A. Expression of Genes Involved in Cellular Adhesion and Extracellular Matrix Remodeling Correlates with Poor Survival of Patients with Renal Cancer. J. Urol. 2016, 195, 1892–1902. [Google Scholar] [CrossRef] [PubMed]
- Usarek, M.; Jagielski, A.K.; Krempa, P.; Dylewska, A.; Kiersztan, A.; Drozak, J.; Girstun, A.; Derlacz, R.A.; Bryla, J. Proinsulin C-peptide potentiates the inhibitory action of insulin on glucose synthesis in primary cultured rabbit kidney-cortex tubules: Metabolic studies. Biochem. Cell Biol. 2014, 92, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Anaya, J. OncoLnc: Linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. Peerj. Comput. Sci. 2016, ARTN e6710.7717/peerj-cs.67. [Google Scholar] [CrossRef]
- Aguirre-Gamboa, R.; Gomez-Rueda, H.; Martinez-Ledesma, E.; Martinez-Torteya, A.; Chacolla-Huaringa, R.; Rodriguez-Barrientos, A.; Tamez-Pena, J.G.; Trevino, V. SurvExpress: An online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS ONE 2013, 8, e74250. [Google Scholar] [CrossRef]
- Li, J.H.; Liu, S.; Zhou, H.; Qu, L.H.; Yang, J.H. starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, D92–D97. [Google Scholar] [CrossRef]

| A. Expression of Metabolic Genes in RCC | ||
| Gene | FC | p Value |
| Increased expression in tumors | ||
| 1. ADA | +5.77 | <0.0001 |
| 2. IL4I1 | +4.20 | <0.0001 |
| 3. HK3 | +3.96 | <0.0001 |
| 4. PYCR1 | +1.56 | <0.0001 |
| Decreased expression in tumors | ||
| 5. PAH | −70.47 | <0.0001 |
| 6. ALDH6A1 | −21.41 | <0.0001 |
| 7. CMKT2 | −18.36 | <0.0001 |
| 8. ALDH4A1 | −14.72 | <0.0001 |
| 9. GATM | −12.99 | <0.0001 |
| 10. DPYS | −10.83 | <0.0001 |
| 11. G6PC | −10.83 | <0.0001 |
| 12. PCCA | −6.87 | <0.0001 |
| 13. GPT | −6.62 | <0.0001 |
| 14. GDA | −6.37 | <0.0001 |
| 15. ALDH5A1 | −5.54 | <0.0001 |
| 16. SUCLG2 | −5.35 | <0.0001 |
| 17. ARG2 | −4.45 | <0.0001 |
| 18. GOT1 | −3.68 | <0.0001 |
| 19. PHOSPHO1 | −1.35 | =0.0215 |
| B. Expression of miRNAs Predicted to Regulate Metabolic Genes in RCC | ||
| MicroRNA | FC | p Value |
| Increased expression in tumors | ||
| 1. miR-122-5p | +107.7 | <0.0001 |
| 2. miR-210-3p | +10.2 | <0.0001 |
| 3. miR-155-5p | +8.3 | <0.0001 |
| 4. miR-34a-5p | +3.1 | <0.0001 |
| 5. miR-146a-5p | +2.1 | <0.0001 |
| 6. miR-106b-5p | +2.1 | <0.0001 |
| 7. miR-342-3p | +1.9 | <0.0001 |
| 8. miR-454-3p | +1.6 | <0.0001 |
| 9. miR-28-5p | +1.5 | <0.0001 |
| 10. miR-126-3p | +1.5 | <0.0001 |
| 11. miR-340-5p | +1.5 | <0.0001 |
| 12. miR-20-5p | +1.4 | <0.0001 |
| Decreased expression in tumors | ||
| 13. miR-129-1-3p | −17.0 | <0.0001 |
| 14. miR-129-2-3p | −6.6 | <0.0001 |
| 15. miR-200b-3p | −4.3 | <0.0001 |
| 16. miR-370-3p | −2.6 | <0.0001 |
| 17. miR-20b-5p | −2.4 | <0.0001 |
| 18. miR-133a-3p | −2.2 | 0.0262 |
| 19. miR-154-5p | −2.1 | <0.0001 |
| 20. miR-135b-5p | −2.0 | 0.0003 |
| 21. miR-27b-3p | −1.6 | <0.0001 |
| 22. miR-543 | −1.5 | 0.0337 |
| Symbol | Entrez Gene Description | Metabolic Pathway | Fold Change | p-Value |
|---|---|---|---|---|
| ACO2 | aconitase 2 | TCA cycle, Amino acid metabolism, Metabolic reprogramming in colon cancer | 1.53 | 3.40 × 10−3 |
| AHCY | Adenosylhomocysteinase | Trans-sulfuration pathway; Trans-sulfuration and one carbon metabolism | 1.76 | 6.00 × 10−4 |
| ALDH1A1 | aldehyde dehydrogenase 1 family member A1 | Tryptophan metabolism | 2.2 | 5.00 × 10−4 |
| CANT1 | calcium activated nucleotidase 1 | Pyrimidine metabolism | 1.53 | 1.14 × 10−2 |
| CBS/CBSL | cystathionine-beta-synthase | Amino acid metabolism; Trans-sulfuration pathway; Trans-sulfuration and one carbon metabolism; One carbon metabolism and related pathways | 1.57 | 2.00 × 10−4 |
| CEBPD | CCAAT enhancer binding protein delta | Adipogenesis | 1.58 | 4.50 × 10−3 |
| CHDH | choline dehydrogenase | One carbon metabolism and related pathways | 1.63 | 4.50 × 10−3 |
| CKB | creatine kinase B | Trans-sulfuration; Urea cycle and metabolism of amino groups | 1.58 | 5.13 × 10−2 |
| CPT2 | carnitine palmitoyltransferase 2 | Fatty Acids Beta Oxidation | 1.61 | 2.20 × 10−3 |
| DHODH | dihydroorotate dehydrogenase (quinone) | Pyrimidine metabolism | 1.88 | 1.00 × 10−4 |
| DNMT3B | DNA methyltransferase 3 beta | Trans-sulfuration; Trans-sulfuration and one carbon metabolism; One carbon metabolism and related pathways | 1.5 | 6.20 × 10−3 |
| E2F1 | E2F transcription factor 1 | Adipogenesis | 1.82 | 9.00 × 10−4 |
| E2F4 | E2F transcription factor 4 | Adipogenesis | 2.01 | 8.00 × 10−4 |
| ECHS1 | enoyl-CoA hydratase, short chain 1 | Fatty Acid Biosynthesis; Fatty Acid Beta oxidation; Tryptophan metabolism | 1.55 | 1.29 × 10−2 |
| ECSIT | ECSIT signalling integrator | Mitochondrial complex I assembly model OXPHOS system | 1.61 | 3.60 × 10−3 |
| ENTPD4 | ectonucleoside triphosphate diphosphohydrolase 4 | Pyrimidine metabolism | 1.58 | 3.42 × 10−1 |
| ESRRA | estrogen related receptor alpha | Energy metabolism | 1.69 | 1.00 × 10−4 |
| G6PD | glucose-6-phosphate dehydrogenase | Pentose Phosphate Pathway; Metabolic reprogramming in colon cancer; Glutathione metabolism | 1.64 | 6.00 × 10−4 |
| GK | glycerol kinase | Fatty Acids Beta Oxidation | -1.75 | 4.30 × 10−3 |
| GPX4 | glutathione peroxidase 4 | One carbon metabolism and related pathways; Glutathion metabolism | 1.82 | 3.40 × 10−3 |
| H6PD | hexose-6-phosphate dehydrogenase/glucose 1-dehydrogenase | Pentose Phosphate Pathway | 1.72 | 4.40 × 10−3 |
| IDH2 | isocitrate dehydrogenase (NADP (+)) 2, mitochondrial | TCA cycle; Metabolic reprogramming in colon cancer | 1.91 | 9.25 × 10−5 |
| LMNA | lamin A/C | Adipogenesis | 1.77 | 8.90 × 10−3 |
| LPIN3 | lipin 3 | Adipogenesis | 2.13 | 2.30 × 10−3 |
| MEF2D | myocyte enhancer factor 2D | Adipogenesis; Energy metabolism | 1.7 | 1.83 × 10−2 |
| MYBBP1A | MYB binding protein 1a | Energy metabolism | 1.77 | 6.00 × 10−4 |
| NDUFAF8 | NADH:ubiquinone oxidoreductase complex assembly factor 8 | Electron Transport Chain (OXPHOS system in mitochondria) | 1.55 | 8.00 × 10−4 |
| NDUFB7 | NADH:ubiquinone oxidoreductase subunit B7 | Electron Transport Chain (OXPHOS system in mitochondria); Mitochondrial complex I assembly model OXPHOS system | 1.64 | 3.67 × 10−2 |
| NDUFS3 | NADH:ubiquinone oxidoreductase core subunit S3 | Electron Transport Chain (OXPHOS system in mitochondria); Mitochondrial complex I assembly model OXPHOS system | 1.52 | 9.00 × 10−4 |
| PGAM5 | PGAM family member 5, mitochondrial serine/threonine protein phosphatase | Metabolic reprogramming in colon cancer | 1.52 | 1.20 × 10−2 |
| PGLS | 6-phosphogluconolactonase | Pentose Phosphate Pathway | 1.53 | 6.40 × 10−3 |
| PYCR2 | pyrroline-5-carboxylate reductase 2 | Metabolic reprogramming in colon cancer | 1.5 | 6.00 × 10−3 |
| RAPGEF3 | Rap guanine nucleotide exchange factor 3 | Integration of energy metabolism | 1.58 | 4.00 × 10−4 |
| SDHA | succinate dehydrogenase complex flavoprotein subunit A | Amino acid metablism; TCA cycle | 1.52 | 2.60 × 10−2 |
| SEMA6B | semaphorin 6B | TCA cycle | 1.5 | 1.60 × 10−3 |
| SHPK | sedoheptulokinase | Pentose Phosphate Pathway | 1.63 | 4.00 × 10−4 |
| SOCS3 | suppressor of cytokine signaling 3 | Adipogenesis | 1.53 | 1.28 × 10−2 |
| STK11 | serine/threonine kinase 11 | Integration of energy metabolism | 1.69 | 6.00 × 10−4 |
| TKT | Transketolase | Pentose Phosphate Pathway; Metabolic reprogramming in colon cancer | 1.56 | 2.00 × 10−3 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bogusławska, J.; Popławski, P.; Alseekh, S.; Koblowska, M.; Iwanicka-Nowicka, R.; Rybicka, B.; Kędzierska, H.; Głuchowska, K.; Hanusek, K.; Tański, Z.; et al. MicroRNA-Mediated Metabolic Reprograming in Renal Cancer. Cancers 2019, 11, 1825. https://doi.org/10.3390/cancers11121825
Bogusławska J, Popławski P, Alseekh S, Koblowska M, Iwanicka-Nowicka R, Rybicka B, Kędzierska H, Głuchowska K, Hanusek K, Tański Z, et al. MicroRNA-Mediated Metabolic Reprograming in Renal Cancer. Cancers. 2019; 11(12):1825. https://doi.org/10.3390/cancers11121825
Chicago/Turabian StyleBogusławska, Joanna, Piotr Popławski, Saleh Alseekh, Marta Koblowska, Roksana Iwanicka-Nowicka, Beata Rybicka, Hanna Kędzierska, Katarzyna Głuchowska, Karolina Hanusek, Zbigniew Tański, and et al. 2019. "MicroRNA-Mediated Metabolic Reprograming in Renal Cancer" Cancers 11, no. 12: 1825. https://doi.org/10.3390/cancers11121825
APA StyleBogusławska, J., Popławski, P., Alseekh, S., Koblowska, M., Iwanicka-Nowicka, R., Rybicka, B., Kędzierska, H., Głuchowska, K., Hanusek, K., Tański, Z., Fernie, A. R., & Piekiełko-Witkowska, A. (2019). MicroRNA-Mediated Metabolic Reprograming in Renal Cancer. Cancers, 11(12), 1825. https://doi.org/10.3390/cancers11121825