Circular RNAs: Characteristics, Function and Clinical Significance in Hepatocellular Carcinoma
Abstract
1. Introduction
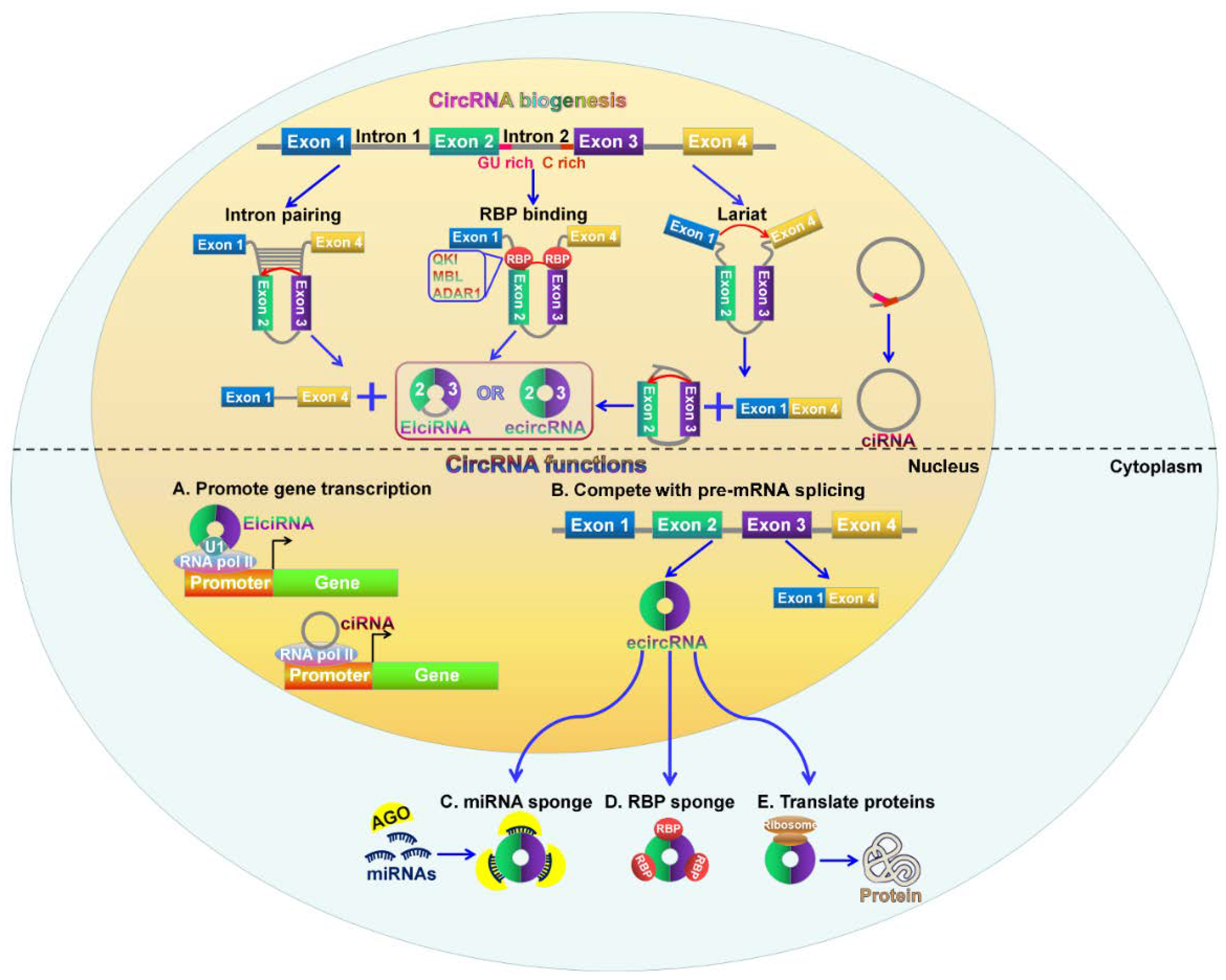
2. Biogenesis of CircRNAs
3. Biological Function of CircRNAs
3.1. CircRNAs can Act as miRNA Sponges
3.2. CircRNAs Function as Transcriptional and Translational Regulators
3.3. CircRNAs Compete with Linear Splicing of Pre-mRNAs
3.4. CircRNAs can Interact with RBPs to Regulate Gene Expression
3.5. CircRNAs Possess Translational Ability
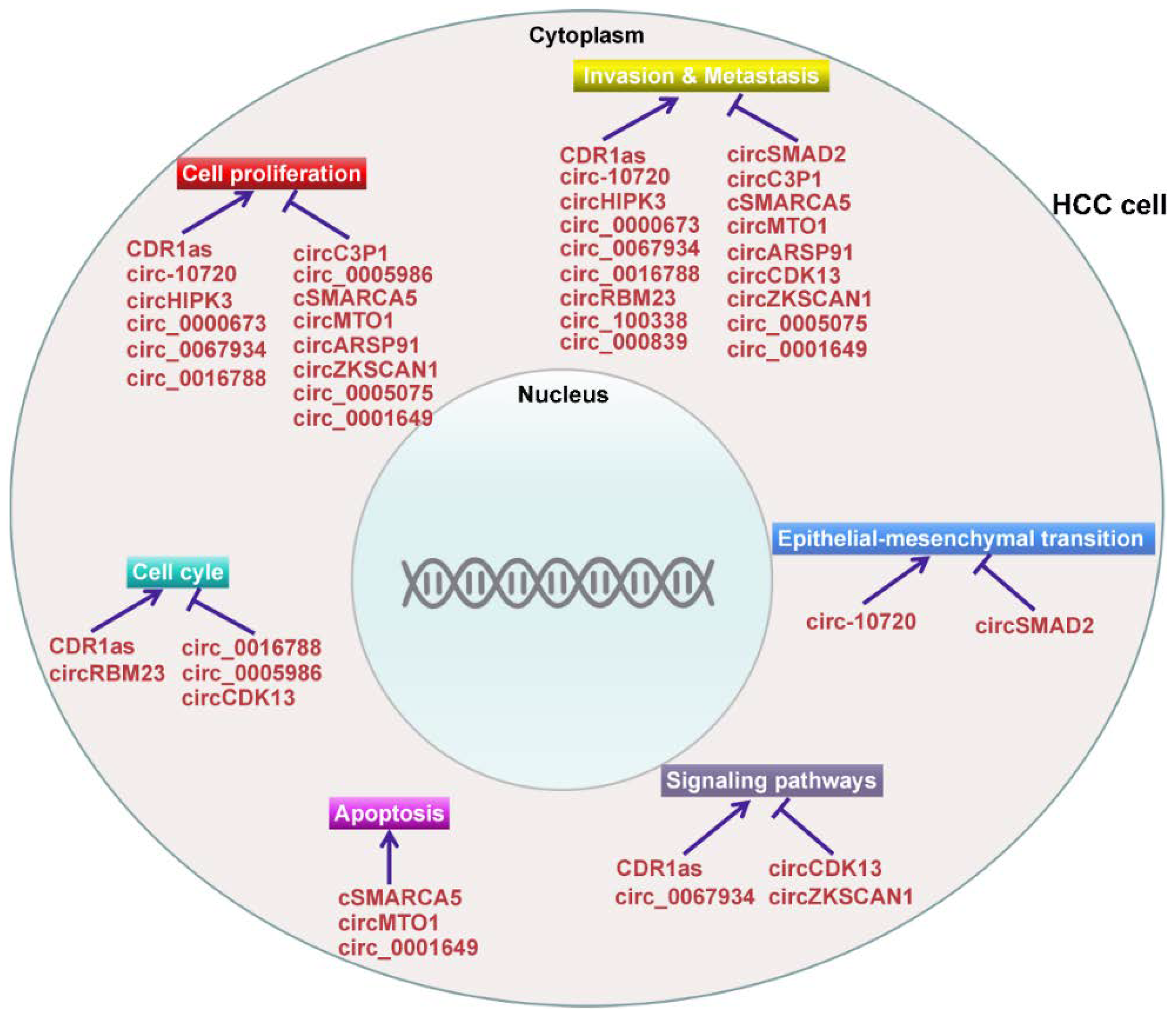
4. The Functional Roles of CircRNAs in HCC
4.1. CircRNAs Act as Oncogenes in HCC
4.2. CircRNAs Act as Tumor Suppressors in HCC
4.3. CircRNAs Interfere with Hepatitis Virus Infection
4.4. CircRNAs May Serve as Potential Diagnostic Biomarkers for HCC
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin. 2011, 61, 69–90. [Google Scholar] [CrossRef] [PubMed]
- Llovet, J.M.; Zucman-Rossi, J.; Pikarsky, E.; Sangro, B.; Schwartz, M.; Sherman, M.; Gores, G. Hepatocellular carcinoma. Nat. Rev. Dis. Primers 2016, 2, 16018. [Google Scholar] [CrossRef] [PubMed]
- Singal, A.G.; El-Serag, H.B. Hepatocellular carcinoma from epidemiology to prevention: Translating knowledge into practice. Clin. Gastroenterol. Hepatol. 2015, 13, 2140–2151. [Google Scholar] [CrossRef] [PubMed]
- Levrero, M. Viral hepatitis and liver cancer: The case of hepatitis c. Oncogene 2006, 25, 3834–3847. [Google Scholar] [CrossRef] [PubMed]
- Kuo, Y.H.; Wu, I.P.; Wang, J.H.; Hung, C.H.; Rau, K.M.; Chen, C.H.; Kee, K.M.; Hu, T.H.; Lu, S.N. The outcome of sorafenib monotherapy on hepatocellular carcinoma with portal vein tumor thrombosis. Investig. New Drugs 2018, 36, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.X.; Qiu, X.Y.; Hu, D.X.; Chen, X.Q. Advances in hypoxia-mediated mechanisms in hepatocellular carcinoma. Mol. Pharmacol. 2017, 92, 246–255. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Mewalal, R.; Hu, R.; Tuskan, G.A.; Yang, X. New technologies accelerate the exploration of non-coding RNAs in horticultural plants. Hortic. Res. 2017, 4, 17031. [Google Scholar] [CrossRef] [PubMed]
- Peschansky, V.J.; Wahlestedt, C. Non-coding RNAs as direct and indirect modulators of epigenetic regulation. Epigenetics 2014, 9, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.Q.; Nolasco, S.; Soares, H. Non-coding RNAs: Multi-tasking molecules in the cell. Int. J. Mol. Sci. 2013, 14, 16010–16039. [Google Scholar] [CrossRef] [PubMed]
- Taft, R.J.; Pang, K.C.; Mercer, T.R.; Dinger, M.; Mattick, J.S. Non-coding RNAs: Regulators of disease. J. Pathol. 2010, 220, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Sun, B.K.; Erwin, J.A.; Song, J.J.; Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse x chromosome. Science 2008, 322, 750–756. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.R.; Mondal, T.; Mohammad, F.; Enroth, S.; Redrup, L.; Komorowski, J.; Nagano, T.; Mancini-Dinardo, D.; Kanduri, C. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 2008, 32, 232–246. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Arai, S.; Song, X.; Reichart, D.; Du, K.; Pascual, G.; Tempst, P.; Rosenfeld, M.G.; Glass, C.K.; Kurokawa, R. Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature 2008, 454, 126–130. [Google Scholar] [CrossRef] [PubMed]
- Khandelwal, A.; Bacolla, A.; Vasquez, K.M.; Jain, A. Long non-coding RNA: A new paradigm for lung cancer. Mol. Carcinog. 2015, 54, 1235–1251. [Google Scholar] [CrossRef] [PubMed]
- Iyer, M.K.; Niknafs, Y.S.; Malik, R.; Singhal, U.; Sahu, A.; Hosono, Y.; Barrette, T.R.; Prensner, J.R.; Evans, J.R.; Zhao, S.; et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 2015, 47, 199–208. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; He, X.X.; Tian, A. Emerging role of microRNA in hepatocellular carcinoma (review). Oncol. Lett. 2015, 9, 1027–1033. [Google Scholar] [CrossRef] [PubMed]
- Conn, S.J.; Pillman, K.A.; Toubia, J.; Conn, V.M.; Salmanidis, M.; Phillips, C.A.; Roslan, S.; Schreiber, A.W.; Gregory, P.A.; Goodall, G.J. The RNA binding protein quaking regulates formation of circRNAs. Cell 2015, 160, 1125–1134. [Google Scholar] [CrossRef] [PubMed]
- Greene, J.; Baird, A.M.; Brady, L.; Lim, M.; Gray, S.G.; McDermott, R.; Finn, S.P. Circular RNAs: Biogenesis, function and role in human diseases. Front. Mol. Biosci. 2017, 4, 38. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.; Tsukahara, T. A view of pre-mRNA splicing from RNase R resistant RNAs. Int. J. Mol. Sci. 2014, 15, 9331–9342. [Google Scholar] [CrossRef] [PubMed]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. CircRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Kos, A.; Dijkema, R.; Arnberg, A.C.; van der Meide, P.H.; Schellekens, H. The hepatitis delta (delta) virus possesses a circular RNA. Nature 1986, 323, 558–560. [Google Scholar] [CrossRef] [PubMed]
- Sanger, H.L.; Klotz, G.; Riesner, D.; Gross, H.J.; Kleinschmidt, A.K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl. Acad. Sci. USA 1976, 73, 3852–3856. [Google Scholar] [CrossRef] [PubMed]
- Huo, L.; Zhang, P.; Li, C.; Rahim, K.; Hao, X.; Xiang, B.; Zhu, X. Genome-wide identification of circRNAs in Pathogenic Basidiomycetous Yeast Cryptococcus Neoformans suggests conserved circRNA host genes over kingdoms. Genes 2018, 9, 118. [Google Scholar] [CrossRef] [PubMed]
- Ye, C.Y.; Chen, L.; Liu, C.; Zhu, Q.H.; Fan, L. Widespread noncoding circular RNAs in plants. New Phytol. 2015, 208, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Thomas, L.F.; Saetrom, P. Circular RNAs are depleted of polymorphisms at microRNA binding sites. Bioinformatics 2014, 30, 2243–2246. [Google Scholar] [CrossRef] [PubMed]
- Abdelmohsen, K.; Panda, A.C.; Munk, R.; Grammatikakis, I.; Dudekula, D.B.; De, S.; Kim, J.; Noh, J.H.; Kim, K.M.; Martindale, J.L.; et al. Identification of hur target circular RNAs uncovers suppression of pabpn1 translation by circpabpn1. RNA Biol. 2017, 14, 361–369. [Google Scholar] [CrossRef] [PubMed]
- Holdt, L.M.; Stahringer, A.; Sass, K.; Pichler, G.; Kulak, N.A.; Wilfert, W.; Kohlmaier, A.; Herbst, A.; Northoff, B.H.; Nicolaou, A.; et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat. Commun. 2016, 7, 12429. [Google Scholar] [CrossRef] [PubMed]
- Dudekula, D.B.; Panda, A.C.; Grammatikakis, I.; De, S.; Abdelmohsen, K.; Gorospe, M. Circinteractome: A web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol. 2016, 13, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with Alu repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Salzman, J.; Chen, R.E.; Olsen, M.N.; Wang, P.L.; Brown, P.O. Cell-type specific features of circular RNA expression. PLoS Genet. 2013, 9, e1003777. [Google Scholar] [CrossRef]
- Yao, Z.; Luo, J.; Hu, K.; Lin, J.; Huang, H.; Wang, Q.; Zhang, P.; Xiong, Z.; He, C.; Huang, Z.; et al. Zkscan1 gene and its related circular RNA (circzkscan1) both inhibit hepatocellular carcinoma cell growth, migration, and invasion but through different signaling pathways. Mol. Oncol. 2017, 11, 422–437. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Chen, Q.; Yao, T.; Li, T.; Ying, S.; Hu, Y.; Guo, J. Hsa_circ_0005986 inhibits carcinogenesis by acting as a mir-129-5p sponge and is used as a novel biomarker for hepatocellular carcinoma. Oncotarget 2017, 8, 43878–43888. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Shi, Y.; Liu, M.; Sun, J. Circhipk3 regulates cell proliferation and migration by sponging mir-124 and regulating aqp3 expression in hepatocellular carcinoma. Cell Death Dis. 2018, 9, 175. [Google Scholar] [CrossRef] [PubMed]
- Granados-Riveron, J.T.; Aquino-Jarquin, G. The complexity of the translation ability of circRNAs. Biochim. Biophys. Acta 2016, 1859, 1245–1251. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, X.O.; Chen, T.; Xiang, J.F.; Yin, Q.F.; Xing, Y.H.; Zhu, S.; Yang, L.; Chen, L.L. Circular intronic long noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.L.; Yang, L. Regulation of circRNA biogenesis. RNA Biol. 2015, 12, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Li, X.; Zhang, P.; Wang, J.; Zhou, Y.; Chen, M. Circular RNA: An emerging key player in RNA world. Brief. Bioinform. 2017, 18, 547–557. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.; Chen, C.Y.; Chuang, T.J. Biogenesis, identification, and function of exonic circular RNAs. Wiley Interdiscip. Rev. RNA 2015, 6, 563–579. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, A.; Memczak, S.; Wyler, E.; Torti, F.; Porath, H.T.; Orejuela, M.R.; Piechotta, M.; Levanon, E.Y.; Landthaler, M.; Dieterich, C.; et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 2015, 10, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Trelles, F.; Tarrio, R.; Ayala, F.J. Origins and evolution of spliceosomal introns. Annu. Rev. Genet. 2006, 40, 47–76. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Valencia-Sanchez, M.A.; Liu, J.; Hannon, G.J.; Parker, R. Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev. 2006, 20, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Bezzi, M.; Guarnerio, J.; Pandolfi, P.P. A circular twist on microRNA regulation. Cell Res. 2017, 27, 1401–1402. [Google Scholar] [CrossRef] [PubMed]
- Capel, B.; Swain, A.; Nicolis, S.; Hacker, A.; Walter, M.; Koopman, P.; Goodfellow, P.; Lovell-Badge, R. Circular transcripts of the testis-determining gene sry in adult mouse testis. Cell 1993, 73, 1019–1030. [Google Scholar] [CrossRef]
- Dubin, R.A.; Kazmi, M.A.; Ostrer, H. Inverted repeats are necessary for circularization of the mouse testis sry transcript. Gene 1995, 167, 245–248. [Google Scholar] [CrossRef]
- Zheng, Q.; Bao, C.; Guo, W.; Li, S.; Chen, J.; Chen, B.; Luo, Y.; Lyu, D.; Li, Y.; Shi, G.; et al. Circular RNA profiling reveals an abundant circhipk3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 2016, 7, 11215. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, L.; Wu, Z.; Wang, M.; Jin, F.; Wang, N.; Hu, X.; Liu, Z.; Zhang, C.Y.; Zen, K.; et al. Mir-124-3p functions as a tumor suppressor in breast cancer by targeting cbl. BMC Cancer 2016, 16, 826. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Zhang, L.; Li, W.; Deng, J.; Zheng, J.; An, M.; Lu, J.; Zhou, Y. Circular RNA itch has inhibitory effect on escc by suppressing the wnt/beta-catenin pathway. Oncotarget 2015, 6, 6001–6013. [Google Scholar] [PubMed]
- Wan, L.; Zhang, L.; Fan, K.; Cheng, Z.X.; Sun, Q.C.; Wang, J.J. Circular RNA-itch suppresses lung cancer proliferation via inhibiting the wnt/beta-catenin pathway. Biomed. Res. Int. 2016, 2016, 1579490. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wei, X.; Yang, J.; Dong, D.; Hao, D.; Huang, Y.; Lan, X.; Plath, M.; Lei, C.; Ma, Y.; et al. Circfgfr4 promotes differentiation of myoblasts via binding mir-107 to relieve its inhibition of wnt3a. Mol. Ther. Nucleic Acids 2018, 11, 272–283. [Google Scholar] [CrossRef] [PubMed]
- Kleaveland, B.; Shi, C.Y.; Stefano, J.; Bartel, D.P. A network of noncoding regulatory RNAs acts in the mammalian brain. Cell 2018. [Google Scholar] [CrossRef] [PubMed]
- Piwecka, M.; Glazar, P.; Hernandez-Miranda, L.R.; Memczak, S.; Wolf, S.A.; Rybak-Wolf, A.; Filipchyk, A.; Klironomos, F.; Cerda Jara, C.A.; Fenske, P.; et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 2017, 357, eaam8526. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Wiklund, E.D.; Bramsen, J.B.; Villadsen, S.B.; Statham, A.L.; Clark, S.J.; Kjems, J. MiRNA-dependent gene silencing involving ago2-mediated cleavage of a circular antisense RNA. EMBO J. 2011, 30, 4414–4422. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef] [PubMed]
- Chao, C.W.; Chan, D.C.; Kuo, A.; Leder, P. The mouse formin (fmn) gene: Abundant circular RNA transcripts and gene-targeted deletion analysis. Mol. Med. 1998, 4, 614–628. [Google Scholar] [PubMed]
- Zhang, X.O.; Wang, H.B.; Zhang, Y.; Lu, X.; Chen, L.L.; Yang, L. Complementary sequence-mediated exon circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef] [PubMed]
- Kelly, S.; Greenman, C.; Cook, P.R.; Papantonis, A. Exon skipping is correlated with exon circularization. J. Mol. Biol. 2015, 427, 2414–2417. [Google Scholar] [CrossRef] [PubMed]
- Starke, S.; Jost, I.; Rossbach, O.; Schneider, T.; Schreiner, S.; Hung, L.H.; Bindereif, A. Exon circularization requires canonical splice signals. Cell Rep. 2015, 10, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Gruner, H.; Cortes-Lopez, M.; Cooper, D.A.; Bauer, M.; Miura, P. CircRNA accumulation in the aging mouse brain. Sci. Rep. 2016, 6, 38907. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Liu, T.; Wang, X.; He, A. Circles reshaping the RNA world: From waste to treasure. Mol. Cancer 2017, 16, 58. [Google Scholar] [CrossRef] [PubMed]
- Du, W.W.; Zhang, C.; Yang, W.; Yong, T.; Awan, F.M.; Yang, B.B. Identifying and characterizing circRNA-protein interaction. Theranostics 2017, 7, 4183–4191. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.G.; Awan, F.M.; Du, W.W.; Zeng, Y.; Lyu, J.; Wu, D.; Gupta, S.; Yang, W.; Yang, B.B. The circular RNA interacts with STAT3, increasing its nuclear translocation and wound repair by modulating DNMT3A and mir-17 function. Mol. Ther. 2017, 25, 2062–2074. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Du, W.W.; Wu, N.; Yang, W.; Awan, F.M.; Fang, L.; Ma, J.; Li, X.; Zeng, Y.; Yang, Z.; et al. A circular RNA promotes tumorigenesis by inducing c-MYC nuclear translocation. Cell Death Differ. 2017, 24, 1609–1620. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Liu, X.; Xue, Y.; Gong, W.; Ma, J.; Xi, Z.; Que, Z.; Liu, Y. Ttbk2 circular RNA promotes glioma malignancy by regulating mir-217/hnf1beta/derlin-1 pathway. J. Hematol. Oncol. 2017, 10, 52. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, Z. Efficient backsplicing produces translatable circular mRNAs. RNA 2015, 21, 172–179. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.Y.; Sarnow, P. Initiation of protein synthesis by the eukaryotic translational apparatus on circular RNAs. Science 1995, 268, 415–417. [Google Scholar] [CrossRef] [PubMed]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-znf609 is a circular RNA that can be translated and functions in myogenesis. Mol. Cell 2017, 66, 22–37. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Gao, X.; Zhang, M.; Yan, S.; Sun, C.; Xiao, F.; Huang, N.; Yang, X.; Zhao, K.; Zhou, H.; et al. Novel role of fbxw7 circular RNA in repressing glioma tumorigenesis. J. Natl. Cancer Inst. 2018, 110, 304–315. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Fan, X.; Mao, M.; Song, X.; Wu, P.; Zhang, Y.; Jin, Y.; Chen, L.L.; Wang, Y.; Wong, C.C.; et al. Extensive translation of circular RNAs driven by n(6)-methyladenosine. Cell Res. 2017, 27, 626–641. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Zhang, M.; Zheng, X.; Yi, P.; Lan, C.; Xu, M. The circular RNA cirs-7 (cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J. Cancer Res. Clin. Oncol. 2017, 143, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Gong, X.; Sun, L.; Zhou, Q.; Lu, B.; Zhu, L. The circular RNA cdr1as act as an oncogene in hepatocellular carcinoma through targeting mir-7 expression. PLoS ONE 2016, 11, e0158347. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Xiong, Q.; Wu, Y.; Li, S.; Ge, F. Quantitative proteomics reveals the regulatory networks of circular RNA cdr1as in hepatocellular carcinoma cells. J. Proteome Res. 2017, 16, 3891–3902. [Google Scholar] [CrossRef] [PubMed]
- Meng, J.; Chen, S.; Han, J.; Qian, B.; Wang, X.; Zhong, W.; Qin, Y.; Zhang, H.; Gao, W.; Lei, Y.; et al. Twist1 regulates vimentin through cul2 circular RNA to promote EMT in hepatocellular carcinoma. Cancer Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Wen, D.; Gong, L.; Wang, Y.; Liu, Z.; Yin, F. Circular RNA hsa_circ_0000673 promotes hepatocellular carcinoma malignance by decreasing mir-767-3p targeting set. Biochem. Biophys. Res. Commun. 2018, 500, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Lu, G.; Luo, Z.; Gui, F.; Wu, J.; Zhang, D.; Ni, Y. CircRNA circ_0067934 promotes tumor growth and metastasis in hepatocellular carcinoma through regulation of mir-1324/fzd5/wnt/beta-catenin axis. Biochem. Biophys. Res. Commun. 2018, 497, 626–632. [Google Scholar] [CrossRef] [PubMed]
- Guan, Z.; Tan, J.; Gao, W.; Li, X.; Yang, Y.; Li, Y.; Wang, Q. Circular RNA hsa_circ_0016788 regulates hepatocellular carcinoma tumorigenesis through mir-486/cdk4 pathway. J. Cell. Physiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Chen, H.; Zhang, C.; Yang, T.; Zhao, Q.; Yan, Y.; Zhang, Y.; Xu, F. Effects of hsa_circrbm23 on hepatocellular carcinoma cell viability and migration as produced by regulating mir-138 expression. Cancer Biother. Radiopharm. 2018, 33, 194–202. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.Y.; Huang, Z.L.; Xu, Y.H.; Zheng, Q.; Chen, Z.; Song, W.; Zhou, J.; Tang, Z.Y. Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-100338/mir-141-3p pathway in hepatitis b-related hepatocellular carcinoma. Sci. Rep. 2017, 7, 5428. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Luo, P.; Jing, W.; Zhou, H.; Liang, C.; Tu, J. Circsmad2 inhibits the epithelial-mesenchymal transition by targeting mir-629 in hepatocellular carcinoma. Onco Targets Ther. 2018, 11, 2853–2863. [Google Scholar] [CrossRef] [PubMed]
- Zhong, L.; Wang, Y.; Cheng, Y.; Wang, W.; Lu, B.; Zhu, L.; Ma, Y. Circular RNA circc3p1 suppresses hepatocellular carcinoma growth and metastasis through mir-4641/pck1 pathway. Biochem. Biophys. Res. Commun. 2018, 499, 1044–1049. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Xu, Q.G.; Wang, Z.G.; Yang, Y.; Zhang, L.; Ma, J.Z.; Sun, S.H.; Yang, F.; Zhou, W.P. Circular RNA csmarca5 inhibits growth and metastasis in hepatocellular carcinoma. J. Hepatol. 2018, 68, 1214–1227. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhou, H.; Jing, W.; Luo, P.; Qiu, S.; Liu, X.; Zhu, M.; Liang, C.; Yu, M.; Tu, J. The circular RNA hsa_circ_0001445 regulates the proliferation and migration of hepatocellular carcinoma and may serve as a diagnostic biomarker. Dis. Markers 2018, 2018, 3073467. [Google Scholar] [CrossRef] [PubMed]
- Han, D.; Li, J.; Wang, H.; Su, X.; Hou, J.; Gu, Y.; Qian, C.; Lin, Y.; Liu, X.; Huang, M.; et al. Circular RNA circmto1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology 2017, 66, 1151–1164. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Yan, P.; Liang, Y.; Sun, Y.; Shen, J.; Zhou, S.; Lin, H.; Liang, X.; Cai, X. Circular RNA expression is suppressed by androgen receptor (ar)-regulated adenosine deaminase that acts on RNA (adar1) in human hepatocellular carcinoma. Cell Death Dis. 2017, 8, e3171. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Ling, Y.B.; Chen, J.W.; Zhou, C.R.; Chen, J.; Li, X.; Huang, M.S. Circular RNA circcdk13 suppresses cell proliferation, migration and invasion by modulating the jak/stat and pi3k/akt pathways in liver cancer. Int. J. Oncol. 2018, 53, 246–256. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.G.; Li, J.S.; Liu, Y.F.; Xu, Q. MicroRNA-200b suppresses the invasion and migration of hepatocellular carcinoma by downregulating RHOA and circRNA_000839. Tumour Biol. 2017, 39, 1010428317719577. [Google Scholar] [CrossRef] [PubMed]
- Shang, X.; Li, G.; Liu, H.; Li, T.; Liu, J.; Zhao, Q.; Wang, C. Comprehensive circular RNA profiling reveals that hsa_circ_0005075, a new circular RNA biomarker, is involved in hepatocellular crcinoma development. Medicine 2016, 95, e3811. [Google Scholar] [CrossRef] [PubMed]
- Li, M.F.; Li, Y.H.; He, Y.H.; Wang, Q.; Zhang, Y.; Li, X.F.; Meng, X.M.; Huang, C.; Li, J. Emerging roles of hsa_circ_0005075 targeting mir-431 in the progress of HCC. Biomed. Pharmacother. 2018, 99, 848–858. [Google Scholar] [CrossRef] [PubMed]
- Qin, M.; Liu, G.; Huo, X.; Tao, X.; Sun, X.; Ge, Z.; Yang, J.; Fan, J.; Liu, L.; Qin, W. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016, 16, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Qiu, S.; Luo, P.; Zhou, H.; Jing, W.; Liang, C.; Tu, J. Down-regulation of hsa_circ_0001649 in hepatocellular carcinoma predicts a poor prognosis. Cancer Biomark. 2018, 22, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Kulcheski, F.R.; Christoff, A.P.; Margis, R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J. Biotechnol. 2016, 238, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Abu, N.; Jamal, R. Circular RNAs as promising biomarkers: A mini-review. Front. Physiol. 2016, 7, 355. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Nazarali, A.J.; Ji, S. Circular RNAs as potential biomarkers for cancer diagnosis and therapy. Am. J. Cancer Res. 2016, 6, 1167–1176. [Google Scholar] [PubMed]
- Peng, L.; Yuan, X.Q.; Li, G.C. The emerging landscape of circular RNA cirs-7 in cancer (review). Oncol. Rep. 2015, 33, 2669–2674. [Google Scholar] [CrossRef] [PubMed]
- Eastham, A.M.; Spencer, H.; Soncin, F.; Ritson, S.; Merry, C.L.; Stern, P.L.; Ward, C.M. Epithelial-mesenchymal transition events during human embryonic stem cell differentiation. Cancer Res. 2007, 67, 11254–11262. [Google Scholar] [CrossRef] [PubMed]
- Sidhu, K.; Kapoor, N.R.; Pandey, V.; Kumar, V. The “macro” world of microRNAs in hepatocellular carcinoma. Front. Oncol. 2015, 5, 68. [Google Scholar] [CrossRef] [PubMed]
- Marlar, S.; Jensen, H.H.; Login, F.H.; Nejsum, L.N. Aquaporin-3 in cancer. Int. J. Mol. Sci. 2017, 18, 2106. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Sun, T.; Yang, M.; Li, Z.; Gao, Y. Prognostic value of combined aquaporin 3 and aquaporin 5 overexpression in hepatocellular carcinoma. Biomed. Res. Int. 2013, 2013, 206525. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.F.; Li, C.F.; Lu, L.; Mei, Z.C. Expression and clinical significance of aquaglyceroporins in human hepatocellular carcinoma. Mol. Med. Rep. 2016, 13, 5283–5289. [Google Scholar] [CrossRef] [PubMed]
- Hung, M.H.; Wang, C.Y.; Chen, Y.L.; Chu, P.Y.; Hsiao, Y.J.; Tai, W.T.; Chao, T.T.; Yu, H.C.; Shiau, C.W.; Chen, K.F. Set antagonist enhances the chemosensitivity of non-small cell lung cancer cells by reactivating protein phosphatase 2a. Oncotarget 2016, 7, 638–655. [Google Scholar] [CrossRef] [PubMed]
- Rincon, R.; Cristobal, I.; Zazo, S.; Arpi, O.; Menendez, S.; Manso, R.; Lluch, A.; Eroles, P.; Rovira, A.; Albanell, J.; et al. Pp2a inhibition determines poor outcome and doxorubicin resistance in early breast cancer and its activation shows promising therapeutic effects. Oncotarget 2015, 6, 4299–4314. [Google Scholar] [CrossRef] [PubMed]
- Hung, M.H.; Chen, Y.L.; Chu, P.Y.; Shih, C.T.; Yu, H.C.; Tai, W.T.; Shiau, C.W.; Chen, K.F. Upregulation of the oncoprotein set determines poor clinical outcomes in hepatocellular carcinoma and shows therapeutic potential. Oncogene 2016, 35, 4891–4902. [Google Scholar] [CrossRef] [PubMed]
- Zipeto, M.A.; Jiang, Q.; Melese, E.; Jamieson, C.H. RNA rewriting, recoding, and rewiring in human disease. Trends Mol. Med. 2015, 21, 549–559. [Google Scholar] [CrossRef] [PubMed]
- Chan, T.H.; Lin, C.H.; Qi, L.; Fei, J.; Li, Y.; Yong, K.J.; Liu, M.; Song, Y.; Chow, R.K.; Ng, V.H.; et al. A disrupted RNA editing balance mediated by ADARS (adenosine deaminases that act on RNA) in human hepatocellular carcinoma. Gut 2014, 63, 832–843. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Chan, T.H.; Tenen, D.G.; Chen, L. RNA editome imbalance in hepatocellular carcinoma. Cancer Res. 2014, 74, 1301–1306. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.M.; Tsang, F.H.; Ng, I.O. Non-coding RNAs in hepatocellular carcinoma: Molecular functions and pathological implications. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 137–151. [Google Scholar] [CrossRef] [PubMed]
- Gramantieri, L.; Fornari, F.; Callegari, E.; Sabbioni, S.; Lanza, G.; Croce, C.M.; Bolondi, L.; Negrini, M. MicroRNA involvement in hepatocellular carcinoma. J. Cell. Mol. Med. 2008, 12, 2189–2204. [Google Scholar] [CrossRef] [PubMed]
- Klingenberg, M.; Matsuda, A.; Diederichs, S.; Patel, T. Non-coding RNA in hepatocellular carcinoma: Mechanisms, biomarkers and therapeutic targets. J. Hepatol. 2017, 67, 603–618. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Y.; Du, Y.; Yang, X.; Mo, Y.; Fan, C.; Xiong, F.; Ren, D.; Ye, X.; Li, C.; Wang, Y.; et al. Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol. Cancer 2018, 17, 79. [Google Scholar] [CrossRef] [PubMed]
- Qiu, L.P.; Wu, Y.H.; Yu, X.F.; Tang, Q.; Chen, L.; Chen, K.P. The emerging role of circular RNAs in hepatocellular carcinoma. J. Cancer 2018, 9, 1548–1559. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Li, P.; Song, Y.; Ge, Y.X.; Meng, X.M.; Huang, C.; Li, J.; Xu, T. Progress and prospects of circular RNAs in hepatocellular carcinoma: Novel insights into their function. J. Cell. Physiol. 2018, 233, 4408–4422. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Jiang, Z.; Li, T.; Hu, Y.; Guo, J. Circular RNAs in hepatocellular carcinoma: Functions and implications. Cancer Med. 2018. [Google Scholar] [CrossRef] [PubMed]
- Petruzziello, A. Epidemiology of hepatitis b virus (HBV) and hepatitis c virus (HCV) related hepatocellular carcinoma. Open Virol. J. 2018, 12, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Neuveut, C.; Wei, Y.; Buendia, M.A. Mechanisms of HBV-related hepatocarcinogenesis. J. Hepatol. 2010, 52, 594–604. [Google Scholar] [CrossRef] [PubMed]
- Webster, D.P.; Klenerman, P.; Dusheiko, G.M. Hepatitis c. Lancet 2015, 385, 1124–1135. [Google Scholar] [CrossRef]
- Zhou, T.C.; Li, X.; Chen, L.J.; Fan, J.H.; Lai, X.; Tang, Y.; Zhang, L.; Wei, J. Differential expression profile of hepatic circular RNAs in chronic hepatitis b. J. Viral Hepat. 2018. [Google Scholar] [CrossRef] [PubMed]
- Bandiera, S.; Pfeffer, S.; Baumert, T.F.; Zeisel, M.B. Mir-122—A key factor and therapeutic target in liver disease. J. Hepatol. 2015, 62, 448–457. [Google Scholar] [CrossRef] [PubMed]
- Tsai, W.C.; Hsu, S.D.; Hsu, C.S.; Lai, T.C.; Chen, S.J.; Shen, R.; Huang, Y.; Chen, H.C.; Lee, C.H.; Tsai, T.F.; et al. MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J. Clin. Investig. 2012, 122, 2884–2897. [Google Scholar] [CrossRef] [PubMed]
- Nakao, K.; Miyaaki, H.; Ichikawa, T. Antitumor function of microRNA-122 against hepatocellular carcinoma. J. Gastroenterol. 2014, 49, 589–593. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Nasser, M.W.; Wang, B.; Hsu, S.H.; Datta, J.; Kutay, H.; Yadav, A.; Nuovo, G.; Kumar, P.; Ghoshal, K. MicroRNA-122 inhibits tumorigenic properties of hepatocellular carcinoma cells and sensitizes these cells to sorafenib. J. Biol. Chem. 2009, 284, 32015–32027. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Xia, F.; Ma, L.; Shan, J.; Shen, J.; Yang, Z.; Liu, J.; Cui, Y.; Bian, X.; Bie, P.; et al. MicroRNA-122 sensitizes HCC cancer cells to adriamycin and vincristine through modulating expression of MDR and inducing cell cycle arrest. Cancer Lett. 2011, 310, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.H.; Wang, B.; Kota, J.; Yu, J.; Costinean, S.; Kutay, H.; Yu, L.; Bai, S.; La Perle, K.; Chivukula, R.R.; et al. Essential metabolic, anti-inflammatory, and anti-tumorigenic functions of MIR-122 in liver. J. Clin. Investig. 2012, 122, 2871–2883. [Google Scholar] [CrossRef] [PubMed]
- Masaki, T.; Arend, K.C.; Li, Y.; Yamane, D.; McGivern, D.R.; Kato, T.; Wakita, T.; Moorman, N.J.; Lemon, S.M. Mir-122 stimulates hepatitis c virus RNA synthesis by altering the balance of viral RNAs engaged in replication versus translation. Cell Host Microbe 2015, 17, 217–228. [Google Scholar] [CrossRef] [PubMed]
- Janssen, H.L.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y.; et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Van der Ree, M.H.; de Vree, J.M.; Stelma, F.; Willemse, S.; van der Valk, M.; Rietdijk, S.; Molenkamp, R.; Schinkel, J.; van Nuenen, A.C.; Beuers, U.; et al. Safety, tolerability, and antiviral effect of rg-101 in patients with chronic hepatitis c: A phase 1b, double-blind, randomised controlled trial. Lancet 2017, 389, 709–717. [Google Scholar] [CrossRef]
- Jost, I.; Shalamova, L.A.; Gerresheim, G.K.; Niepmann, M.; Bindereif, A.; Rossbach, O. Functional sequestration of microRNA-122 from hepatitis c virus by circular RNA sponges. RNA Biol. 2018, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.X.; Tang, H. Complex interactions between microRNAs and hepatitis b/c viruses. World J. Gastroenterol. 2014, 20, 13477–13492. [Google Scholar] [CrossRef] [PubMed]
- Xie, K.L.; Zhang, Y.G.; Liu, J.; Zeng, Y.; Wu, H. MicroRNAs associated with HBV infection and HBV-related HCC. Theranostics 2014, 4, 1176–1192. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.H.; Kim, J.H.; Lee, S.W. The role of microRNAs in hepatitis c virus replication and related liver diseases. J. Microbiol. 2014, 52, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Liu, C.X.; Xue, W.; Zhang, Y.; Jiang, S.; Yin, Q.F.; Wei, J.; Yao, R.W.; Yang, L.; Chen, L.L. Coordinated circRNA biogenesis and function with nf90/nf110 in viral infection. Mol. Cell 2017, 67, 214–227. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.G.; Kim, M.V.; Chen, X.; Batista, P.J.; Aoyama, S.; Wilusz, J.E.; Iwasaki, A.; Chang, H.Y. Sensing self and foreign circular RNAs by intron identity. Mol. Cell 2017, 67, 228–238. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Zhang, C.; Lin, J.; Song, X.; Wang, H. Screening differential circular RNA expression profiles reveal that hsa_circ_0128298 is a biomarker in the diagnosis and prognosis of hepatocellular carcinoma. Cancer Manag. Res. 2018, 10, 1275–1283. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Yao, T.; Chen, Q.; Mo, X.; Hu, Y.; Guo, J. Screening differential circular RNA expression profiles reveals hsa_circ_0004018 is associated with hepatocellular carcinoma. Oncotarget 2017, 8, 58405–58416. [Google Scholar] [CrossRef] [PubMed]
- Yao, T.; Chen, Q.; Shao, Z.; Song, Z.; Fu, L.; Xiao, B. Circular RNA 0068669 as a new biomarker for hepatocellular carcinoma metastasis. J. Clin. Lab. Anal. 2018, e22572. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Wu, S.; Yao, T.; Chen, Q.; Xie, Y.; Ying, S.; Chen, Z.; Xiao, B.; Hu, Y. Decreased expression of hsa_circ_0003570 in hepatocellular carcinoma and its clinical significance. J. Clin. Lab. Anal. 2018, 32, e22239. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Papavasileiou, P.; Peters, O.; Rajewsky, N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS ONE 2015, 10, e0141214. [Google Scholar] [CrossRef] [PubMed]
- Bahn, J.H.; Zhang, Q.; Li, F.; Chan, T.M.; Lin, X.; Kim, Y.; Wong, D.T.; Xiao, X. The landscape of microRNA, piwi-interacting Rna, and circular Rna in human saliva. Clin. Chem. 2015, 61, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Li, J.; Lu, R.; Li, T.; Yang, Y.; Xiao, B.; Guo, J. Global circular RNA expression profile of human gastric cancer and its clinical significance. Cancer Med. 2017, 6, 1173–1180. [Google Scholar] [CrossRef] [PubMed]
- Koh, W.; Pan, W.; Gawad, C.; Fan, H.C.; Kerchner, G.A.; Wyss-Coray, T.; Blumenfeld, Y.J.; El-Sayed, Y.Y.; Quake, S.R. Noninvasive in vivo monitoring of tissue-specific global gene expression in humans. Proc. Natl. Acad. Sci. USA 2014, 111, 7361–7366. [Google Scholar] [CrossRef] [PubMed]
- Caby, M.P.; Lankar, D.; Vincendeau-Scherrer, C.; Raposo, G.; Bonnerot, C. Exosomal-like vesicles are present in human blood plasma. Int. Immunol. 2005, 17, 879–887. [Google Scholar] [CrossRef] [PubMed]
- Pisitkun, T.; Shen, R.F.; Knepper, M.A. Identification and proteomic profiling of exosomes in human urine. Proc. Natl. Acad. Sci. USA 2004, 101, 13368–13373. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, Y.; Miura, Y.; Harazono, A.; Kanai-Azuma, M.; Akimoto, Y.; Kawakami, H.; Yamaguchi, T.; Toda, T.; Endo, T.; Tsubuki, M.; et al. Proteomic analysis of two types of exosomes in human whole saliva. Biol. Pharm. Bull. 2011, 34, 13–23. [Google Scholar] [CrossRef] [PubMed]
- Vella, L.J.; Sharples, R.A.; Lawson, V.A.; Masters, C.L.; Cappai, R.; Hill, A.F. Packaging of prions into exosomes is associated with a novel pathway of PRP processing. J. Pathol. 2007, 211, 582–590. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Chen, K.K.; Zhang, J.; Xiao, B.; Huang, Z.; Ju, C.; Sun, J.; Zhang, F.; Lv, X.B.; Huang, G. The decade of exosomal long RNA species: An emerging cancer antagonist. Mol. Cancer 2018, 17, 75. [Google Scholar] [CrossRef] [PubMed]
- Dou, Y.; Cha, D.J.; Franklin, J.L.; Higginbotham, J.N.; Jeppesen, D.K.; Weaver, A.M.; Prasad, N.; Levy, S.; Coffey, R.J.; Patton, J.G.; et al. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci. Rep. 2016, 6, 37982. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.P.; He, Y.J.; Hou, J.C.; Chen, X.; Zhou, S.Y.; Yang, S.J.; Li, J.; Zhang, H.D.; Hu, J.H.; Zhong, S.L.; et al. The role of circRNAs in cancers. Biosci. Rep. 2017, 37, BSR20170750. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zheng, Q.; Bao, C.; Li, S.; Guo, W.; Zhao, J.; Chen, D.; Gu, J.; He, X.; Huang, S. Circular RNA is enriched and stable in exosomes: A promising biomarker for cancer diagnosis. Cell Res. 2015, 25, 981–984. [Google Scholar] [CrossRef] [PubMed]
- Bortoluzzi, S.; Lovisa, F.; Gaffo, E.; Mussolin, L. Small RNAs in circulating exosomes of cancer patients: A minireview. High Throughput 2017, 6, 13. [Google Scholar] [CrossRef] [PubMed]
| CircRNA | Alias | Gene Symbol | Chromosome | Expression Change | Function | Possible Mechanism | Reference |
|---|---|---|---|---|---|---|---|
| CDR1as (ciRS-7) | hsa_circ_0001946 | CDR1 | chrX | Up | proliferation (+); invasion (+) | miRNA sponge | [73,74,75] |
| Cul2 circRNA | circ-10720 | CUL2 | chr10 | Up | proliferation (+); migration (+); invasion (+); EMT (+) | miRNA sponge | [76] |
| circHIPK3 | hsa_circ_0000284 | HIPK3 | chr11 | Up | proliferation (+); migration (+) | miRNA sponge | [36] |
| hsa_circ_0000673 | - | RSL1D1 | chr16 | Up | proliferation (+); invasion (+) | miRNA sponge | [77] |
| hsa_circ_0067934 | - | PRKC1 | chr3 | Up | proliferation (+); migration (+); invasion (+) | miRNA sponge | [78] |
| hsa_circ_0016788 | - | TRIM11 | chr1 | Up | proliferation (+); invasion (+); apoptosis (−) | miRNA sponge | [79] |
| circRBM23 | - | RBM23 | chr14 | Up | viability (+); migration (+) | miRNA sponge | [80] |
| hsa_circ_100338 | - | SNX27 | chr1 | Up | migration (+); invasion (+) | miRNA sponge | [81] |
| circSMAD2 | hsa_circ_0000847 | SMAD2 | chr18 | down | migration (−); invasion (−); EMT (−) | miRNA sponge | [82] |
| circC3P1 | - | C3P1 | chr19 | down | proliferation (−); migration (−); invasion (−) | miRNA sponge | [83] |
| hsa_circ_0005986 | - | PRDM2 | chr1 | down | proliferation (−) | miRNA sponge | [35] |
| cSMARCA5 | hsa_circ_0001445 | SMARCA5 | chr4 | down | proliferation (−); migration (−); invasion (−); apoptosis (+) | miRNA sponge | [84,85] |
| circMTO1 | hsa_circ_0007874/hsa_circ_104135 | MTO1 | chr6 | down | proliferation (−); invasion (−); apoptosis (+) | miRNA sponge | [86] |
| circARSP91 | hsa_circ_0085154 | PABPC1 | chr8 | down | proliferation (−); invasion (−) | regulated by ADAR1 | [87] |
| circCDK13 | hsa_circ_0001699 | CDK13 | chr7 | down | migration (−); invasion (−) | regulating JAK/STAT and PI3K/ATK signaling pathways | [88] |
| circZKSCAN1 | hsa_circ_0001727 | ZKSCAN1 | chr7 | down | proliferation (−); migration (−); invasion (−) | regulating cancer-related pathways | [34] |
| hsa_circ_000839 | hsa_circ_0000497 | SLAIN1 | chr13 | up | migration (+); invasion (+) | regulated by miR-200b | [89] |
| hsa_circ_0005075 | - | EIF4G3 | chr1 | up | proliferation (−); migration (−); invasion (−) | miRNA sponge | [90,91] |
| hsa_circ_0001649 | - | SHPRH | chr6 | down | proliferation (−); migration (−); invasion (−); apoptosis (+) | Promoting MMPs expression | [92,93] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, M.; Yu, F.; Li, P. Circular RNAs: Characteristics, Function and Clinical Significance in Hepatocellular Carcinoma. Cancers 2018, 10, 258. https://doi.org/10.3390/cancers10080258
Wang M, Yu F, Li P. Circular RNAs: Characteristics, Function and Clinical Significance in Hepatocellular Carcinoma. Cancers. 2018; 10(8):258. https://doi.org/10.3390/cancers10080258
Chicago/Turabian StyleWang, Man, Fei Yu, and Peifeng Li. 2018. "Circular RNAs: Characteristics, Function and Clinical Significance in Hepatocellular Carcinoma" Cancers 10, no. 8: 258. https://doi.org/10.3390/cancers10080258
APA StyleWang, M., Yu, F., & Li, P. (2018). Circular RNAs: Characteristics, Function and Clinical Significance in Hepatocellular Carcinoma. Cancers, 10(8), 258. https://doi.org/10.3390/cancers10080258






