Microenvironmental pH and Exosome Levels Interplay in Human Cancer Cell Lines of Different Histotypes
Abstract
1. Introduction
2. Results
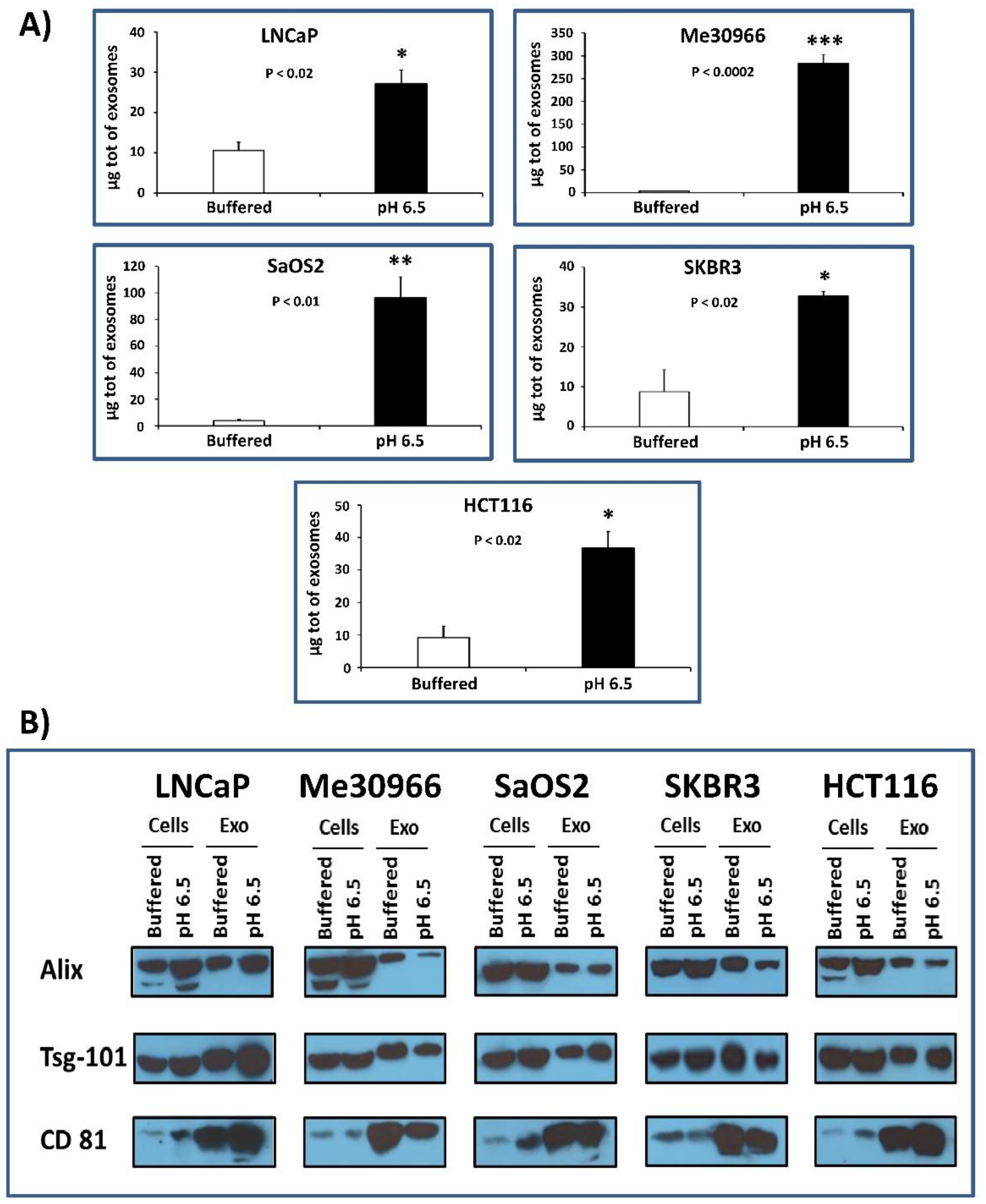
2.1. Protein Quantification and Characterization by Western Blot Analysis for Housekeeping Markers of Exosomes
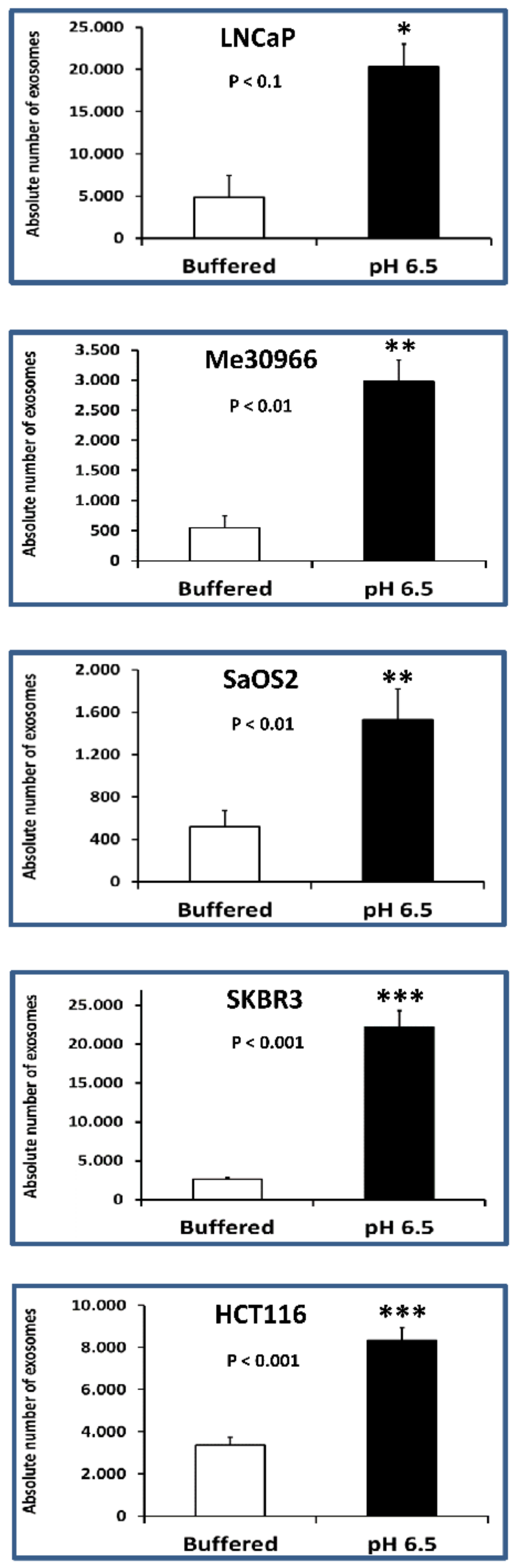
2.2. Nanoscale Flow Cytometry Quantification and Characterization of Exosomal Preparations Obtained from Supernatants of Tumor Cells Grown under Different pH Conditions
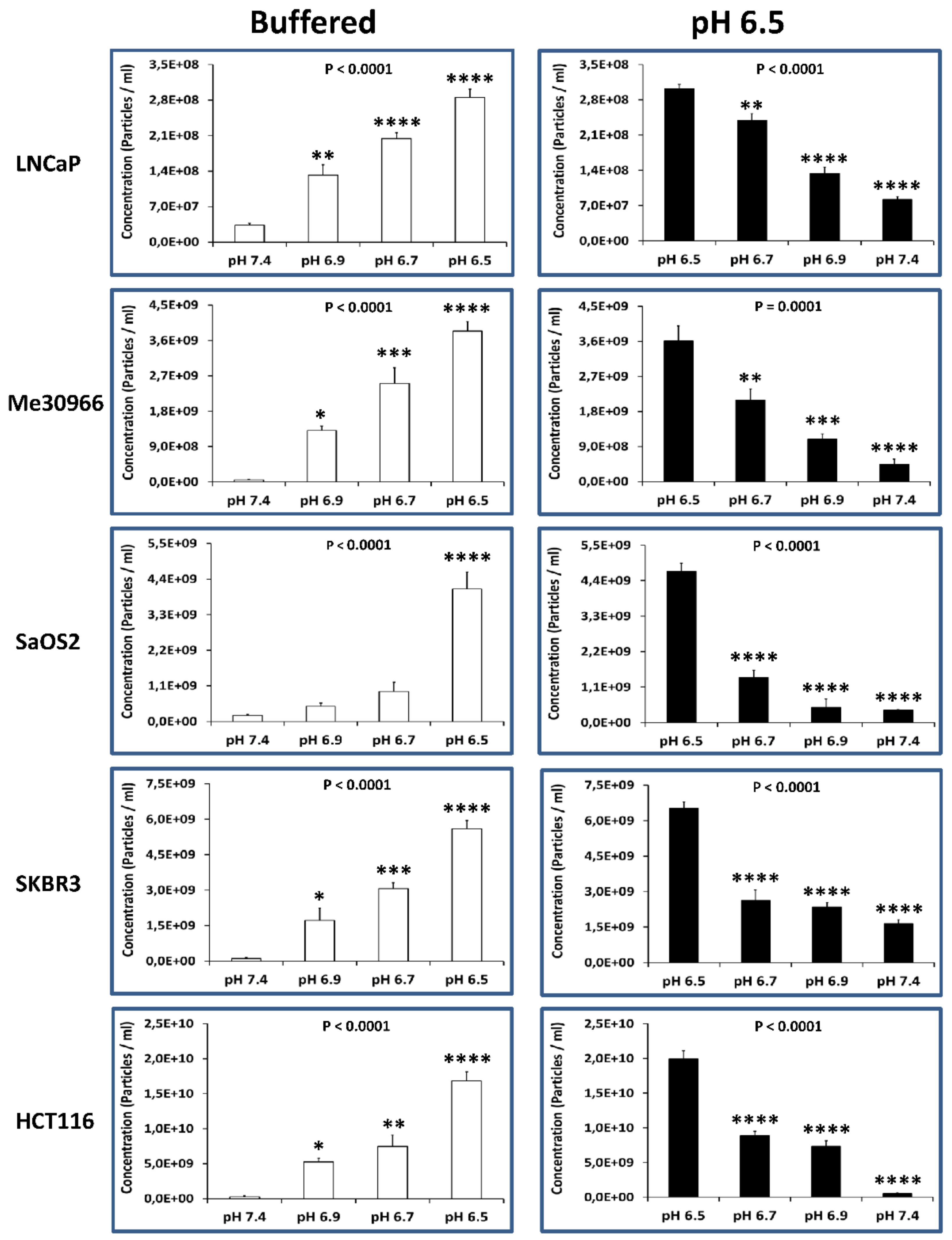
2.3. NTA Quantification of Exosome Preparations Obtained from Supernatants of Tumor Cells Grown under Different pH Conditions
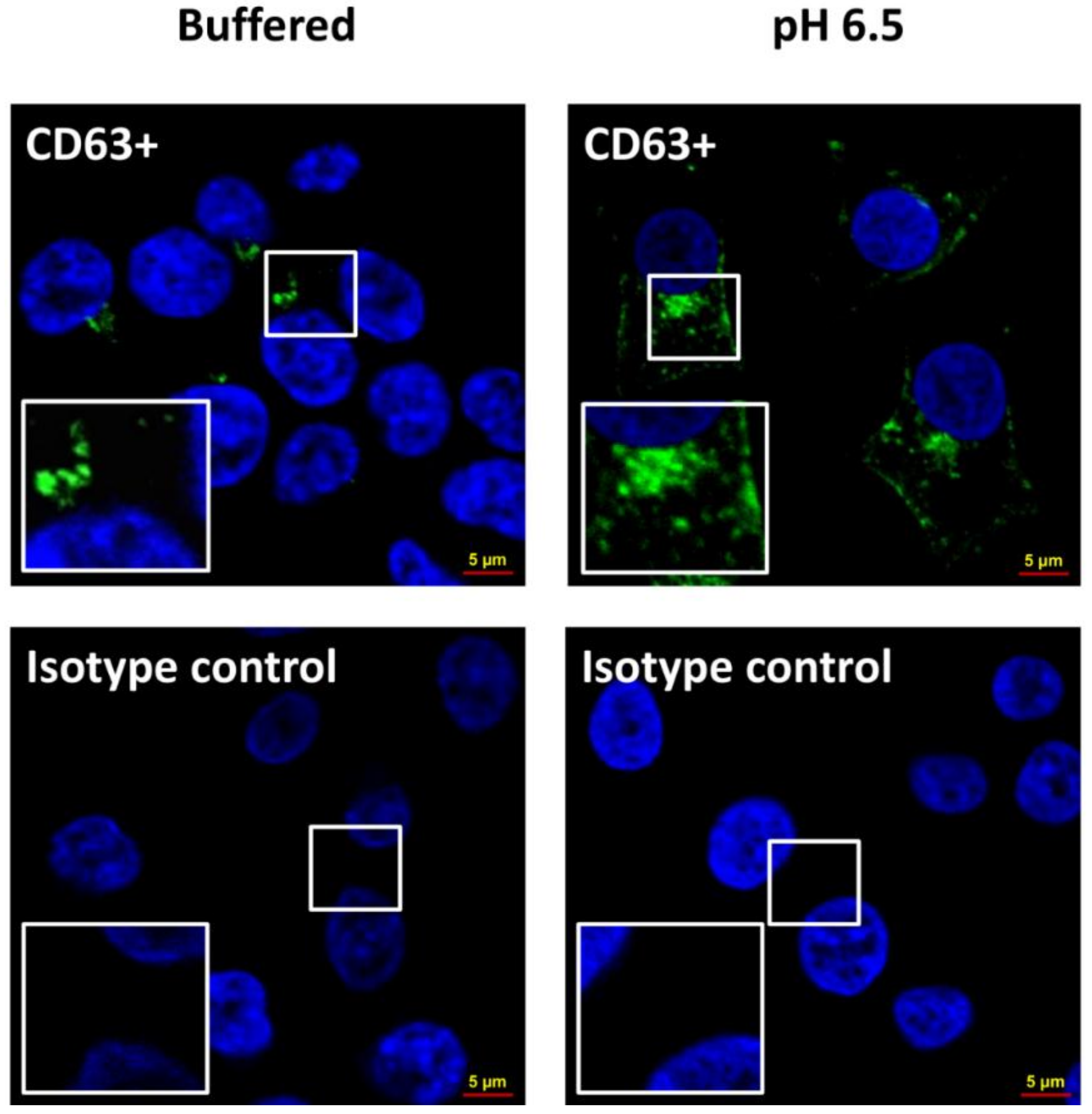
2.4. Alkalization of the Tumor Cell Microenvironment Induces a Reduction of Exosome Release
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. Exosome Purification from Cell-Culture Supernatants
4.3. Nanoparticle Tracking Analysis
4.4. Flow Cytometry Analysis of Exosomes
4.5. Western Blot
4.6. Confocal Analysis
4.7. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Federici:, C.; Petrucci, F.; Caimi, S.; Cesolini, A.; Logozzi, M.; Borghi, M.; D’Ilio, S.; Lugini, L.; Violante, N.; Azzarito, T.; et al. Exosome Release and Low pH Belong to a Framework of Resistance of Human Melanoma Cells to Cisplatin. PLoS ONE 2014, 9, e88193. [Google Scholar] [CrossRef] [PubMed]
- Iessi, E.; Logozzi, M.; Lugini, L.; Azzarito, T.; Federici, C.; Spugnini, E.P.; Mizzoni, D.; Di Raimo, R.; Angelini, D.F.; Battistini, L.; et al. Acridine Orange/exosomes increase the delivery and the effectiveness of Acridine Orange in human melanoma cells: A new prototype for theranostics of tumors. J. Enzyme Inhib. Med. Chem. 2017, 32, 648–657. [Google Scholar] [CrossRef] [PubMed]
- Fais, S.; Logozzi, M.; Lugini, L.; Federici, C.; Azzarito, T.; Zarovni, N.; Chiesi, A. Exosomes: the ideal nanovectors for biodelivery. Biol. Chem. 2013, 394, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Yáñez-Mó, M.; Siljander, P.R.-M.; Andreu, Z.; Bedina Zavec, A.; Borràs, F.E.; Buzas, E.I.; Buzas, K.; Casal, E.; Cappello, F.; Carvalho, J.; et al. Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 2015, 4, 27066. [Google Scholar] [CrossRef] [PubMed]
- Cocucci, E.; Meldolesi, J. Ectosomes and exosomes: Shedding the confusion between extracellular vesicles. Trends Cell Biol. 2015, 25, 364–372. [Google Scholar] [CrossRef] [PubMed]
- Parolini, I.; Federici, C.; Raggi, C.; Lugini, L.; Palleschi, S.; De Milito, A.; Coscia, C.; Iessi, E.; Logozzi, M.; Molinari, A.; et al. Microenvironmental pH Is a Key Factor for Exosome Traffic in Tumor Cells. J. Biol. Chem. 2009, 284, 34211–34222. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Iguchi, H.; Yoshioka, Y.; Takeshita, F.; Matsuki, Y.; Ochiya, T. Secretory Mechanisms and Intercellular Transfer of MicroRNAs in Living Cells. J. Biol. Chem. 2010, 285, 17442–17452. [Google Scholar] [CrossRef] [PubMed]
- Gallala, H.D.; Sandhoff, K. Biological Function of the Cellular Lipid BMP—BMP as a Key Activator for Cholesterol Sorting and Membrane Digestion. Neurochem. Res. 2011, 36, 1594–1600. [Google Scholar] [CrossRef] [PubMed]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Camussi, G.; Deregibus, M.-C.; Bruno, S.; Grange, C.; Fonsato, V.; Tetta, C. Exosome/microvesicle-mediated epigenetic reprogramming of cells. Am. J. Cancer Res. 2011, 1, 98–110. [Google Scholar] [PubMed]
- Nolte-’t Hoen, E.N.M.; Buermans, H.P.J.; Waasdorp, M.; Stoorvogel, W.; Wauben, M.H.M.; ’t Hoen, P.A.C. Deep sequencing of RNA from immune cell-derived vesicles uncovers the selective incorporation of small non-coding RNA biotypes with potential regulatory functions. Nucleic Acids Res. 2012, 40, 9272–9285. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Achreja, A.; Iessi, E.; Logozzi, M.; Mizzoni, D.; Di Raimo, R.; Nagrath, D.; Fais, S. The key role of extracellular vesicles in the metastatic process. Biochim. Biophys. Acta BBA Rev. Cancer 2018, 1869, 64–77. [Google Scholar] [CrossRef] [PubMed]
- Fais, S. A nonmainstream approach against cancer. J. Enzyme Inhib. Med. Chem. 2016, 31, 882–889. [Google Scholar] [CrossRef] [PubMed]
- Fais, S.; Venturi, G.; Gatenby, B. Microenvironmental acidosis in carcinogenesis and metastases: New strategies in prevention and therapy. Cancer Metastasis Rev. 2014, 33, 1095–1108. [Google Scholar] [CrossRef] [PubMed]
- Luciani, F.; Spada, M.; De Milito, A.; Molinari, A.; Rivoltini, L.; Montinaro, A.; Marra, M.; Lugini, L.; Logozzi, M.; Lozupone, F.; et al. Effect of proton pump inhibitor pretreatment on resistance of solid tumors to cytotoxic drugs. J. Natl. Cancer Inst. 2004, 96, 1702–1713. [Google Scholar] [CrossRef] [PubMed]
- De Milito, A.; Canese, R.; Marino, M.L.; Borghi, M.; Iero, M.; Villa, A.; Venturi, G.; Lozupone, F.; Iessi, E.; Logozzi, M.; et al. pH-dependent antitumor activity of proton pump inhibitors against human melanoma is mediated by inhibition of tumor acidity. Int. J. Cancer 2010, 127, 207–219. [Google Scholar] [CrossRef] [PubMed]
- Calcinotto, A.; Filipazzi, P.; Grioni, M.; Iero, M.; De Milito, A.; Ricupito, A.; Cova, A.; Canese, R.; Jachetti, E.; Rossetti, M.; et al. Modulation of Microenvironment Acidity Reverses Anergy in Human and Murine Tumor-Infiltrating T Lymphocytes. Cancer Res. 2012, 72, 2746–2756. [Google Scholar] [CrossRef] [PubMed]
- Azzarito, T.; Venturi, G.; Cesolini, A.; Fais, S. Lansoprazole induces sensitivity to suboptimal doses of paclitaxel in human melanoma. Cancer Lett. 2015, 356, 697–703. [Google Scholar] [CrossRef] [PubMed]
- Azzarito, T.; Lugini, L.; Spugnini, E.P.; Canese, R.; Gugliotta, A.; Fidanza, S.; Fais, S. Effect of Modified Alkaline Supplementation on Syngenic Melanoma Growth in CB57/BL Mice. PLOS ONE 2016, 11, e0159763. [Google Scholar] [CrossRef] [PubMed]
- Canitano, A.; Iessi, E.; Spugnini, E.P.; Federici, C.; Fais, S. Proton pump inhibitors induce a caspase-independent antitumor effect against human multiple myeloma. Cancer Lett. 2016, 376, 278–283. [Google Scholar] [CrossRef] [PubMed]
- Warburg, O. On the origin of cancer cells. Science 1956, 123, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Gatenby, R.A.; Gillies, R.J. Why do cancers have high aerobic glycolysis? Nat. Rev. Cancer 2004, 4, 891–899. [Google Scholar] [CrossRef] [PubMed]
- Vander Heiden, M.G.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg Effect: The Metabolic Requirements of Cell Proliferation. Science 2009, 324, 1029–1033. [Google Scholar] [CrossRef] [PubMed]
- Cairns, R.A.; Harris, I.S.; Mak, T.W. Regulation of cancer cell metabolism. Nat. Rev. Cancer 2011, 11, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Iessi, E.; Logozzi, M.; Mizzoni, D.; Di Raimo, R.; Supuran, C.; Fais, S. Rethinking the Combination of Proton Exchanger Inhibitors in Cancer Therapy. Metabolites 2017, 8, 2. [Google Scholar] [CrossRef] [PubMed]
- Logozzi, M.; De Milito, A.; Lugini, L.; Borghi, M.; Calabrò, L.; Spada, M.; Perdicchio, M.; Marino, M.L.; Federici, C.; Iessi, E.; et al. High Levels of Exosomes Expressing CD63 and Caveolin-1 in Plasma of Melanoma Patients. PLoS ONE 2009, 4, e5219. [Google Scholar] [CrossRef] [PubMed]
- Properzi, F.; Logozzi, M.; Fais, S. Exosomes: the future of biomarkers in medicine. Biomark. Med. 2013, 7, 769–778. [Google Scholar] [CrossRef] [PubMed]
- Logozzi, M.; Angelini, D.F.; Iessi, E.; Mizzoni, D.; Di Raimo, R.; Federici, C.; Lugini, L.; Borsellino, G.; Gentilucci, A.; Pierella, F.; et al. Increased PSA expression on prostate cancer exosomes in in vitro condition and in cancer patients. Cancer Lett. 2017, 403, 318–329. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Luecke, L.B.; Kahlert, C.; Fernandez, A.F.; Gammon, S.T.; Kaye, J.; LeBleu, V.S.; Mittendorf, E.A.; Weitz, J.; Rahbari, N.; et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 2015, 523, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Spugnini, E.P.; Sonveaux, P.; Stock, C.; Perez-Sayans, M.; De Milito, A.; Avnet, S.; Garcìa, A.G.; Harguindey, S.; Fais, S. Proton channels and exchangers in cancer. Biochim. Biophys. Acta BBA - Biomembr. 2015, 1848, 2715–2726. [Google Scholar] [CrossRef] [PubMed]
- Cappello, F.; Logozzi, M.; Campanella, C.; Bavisotto, C.C.; Marcilla, A.; Properzi, F.; Fais, S. Exosome levels in human body fluids: A tumor marker by themselves? Eur. J. Pharm. Sci. 2017, 96, 93–98. [Google Scholar] [CrossRef] [PubMed]
- Oosthuyzen, W.; Sime, N.E.L.; Ivy, J.R.; Turtle, E.J.; Street, J.M.; Pound, J.; Bath, L.E.; Webb, D.J.; Gregory, C.D.; Bailey, M.A.; et al. Quantification of human urinary exosomes by nanoparticle tracking analysis: Nanoparticle tracking analysis and exosomes. J. Physiol. 2013, 591, 5833–5842. [Google Scholar] [CrossRef] [PubMed]
- Dragovic, R.A.; Collett, G.P.; Hole, P.; Ferguson, D.J.P.; Redman, C.W.; Sargent, I.L.; Tannetta, D.S. Isolation of syncytiotrophoblast microvesicles and exosomes and their characterisation by multicolour flow cytometry and fluorescence Nanoparticle Tracking Analysis. Methods 2015, 87, 64–74. [Google Scholar] [CrossRef] [PubMed]
- Lugini, L.; Valtieri, M.; Federici, C.; Cecchetti, S.; Meschini, S.; Condello, M.; Signore, M.; Fais, S. Exosomes from human colorectal cancer induce a tumor-like behavior in colonic mesenchymal stromal cells. Oncotarget 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Fais, S.; Fauvarque, M.-O. TM9 and cannibalism: how to learn more about cancer by studying amoebae and invertebrates. Trends Mol. Med. 2012, 18, 4–5. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, F.; Perdicchio, M.; Brambilla, D.; Borghi, M.; Meschini, S.; Barca, S.; Marino, M.L.; Logozzi, M.; Federici, C.; Iessi, E.; et al. The human homologue of Dictyostelium discoideum phg1A is expressed by human metastatic melanoma cells. EMBO Rep. 2009, 10, 1348–1354. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, M.C.; Cunningham, J.J.; Bui, M.M.; Gillies, R.J.; Brown, J.S.; Gatenby, R.A. Darwinian Dynamics of Intratumoral Heterogeneity: Not Solely Random Mutations but Also Variable Environmental Selection Forces. Cancer Res. 2016, 76, 3136–3144. [Google Scholar] [CrossRef] [PubMed]
- Gillies, R.J.; Brown, J.S.; Anderson, A.R.A.; Gatenby, R.A. Eco-evolutionary causes and consequences of temporal changes in intratumoural blood flow. Nat. Rev. Cancer 2018, 18, 576–585. [Google Scholar] [CrossRef] [PubMed]
- Overholtzer, M.; Fais, S. Nat. Rev. Cancer 2018, in press.
- Fais, S.; O’Driscoll, L.; Borras, F.E.; Buzas, E.; Camussi, G.; Cappello, F.; Carvalho, J.; Cordeiro da Silva, A.; Del Portillo, H.; El Andaloussi, S.; et al. Evidence-Based Clinical Use of Nanoscale Extracellular Vesicles in Nanomedicine. ACS Nano 2016, 10, 3886–3899. [Google Scholar] [CrossRef] [PubMed]
- Huber, V.; Fais, S.; Iero, M.; Lugini, L.; Canese, P.; Squarcina, P.; Zaccheddu, A.; Colone, M.; Arancia, G.; Gentile, M.; et al. Human colorectal cancer cells induce T-cell death through release of proapoptotic microvesicles: role in immune escape. Gastroenterology 2005, 128, 1796–1804. [Google Scholar] [CrossRef] [PubMed]
- Andreola, G.; Rivoltini, L.; Castelli, C.; Huber, V.; Perego, P.; Deho, P.; Squarcina, P.; Accornero, P.; Lozupone, F.; Lugini, L.; et al. Induction of lymphocyte apoptosis by tumor cell secretion of FasL-bearing microvesicles. J. Exp. Med. 2002, 195, 1303–1316. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; Betts, C.; Lakhal, S.; Wood, M.J.A. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Gillies, R.J.; Raghunand, N.; Garcia-Martin, M.L.; Gatenby, R.A. pH imaging. A review of pH measurement methods and applications in cancers. IEEE Eng. Med. Biol. Mag. 2004, 23, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Zitvogel, L.; Amigorena, S. Exosomes: Composition, biogenesis and function. Nat. Rev. Immunol. 2002, 2, 569–579. [Google Scholar] [CrossRef] [PubMed]

| Nanoscale Flow Cytometry | ||
|---|---|---|
| Tumor Cell Lines | Absolute Number of Exosomes (Mean ± Standard Error (SE)) | |
| Buffered (pH 7.4) | pH 6.5 | |
| LNCaP | 4.9 × 103 ± 2.6 × 103 | 2.0 × 104 ± 2.6 × 103 |
| Me30966 | 5.5 × 102 ± 1.9 × 102 | 3.0 × 103 ± 3.5 × 102 |
| SaOS2 | 5.2 × 102 ± 3.7 × 10 | 1.5 × 103 ± 1.4 × 102 |
| SKBR3 | 2.7 × 103 ± 1.5 × 102 | 2.2 × 104 ± 2.1 × 103 |
| HCT116 | 3.4 × 103 ± 3.1 × 102 | 8.3 × 103 ± 3.5 × 102 |
| Nanoparticle Tracking Analysis (NTA) | ||
|---|---|---|
| Tumor Cell Lines | Concentration of Exosomes (Particles/mL) (Mean ± SE) | |
| Buffered (pH 7.4) | pH 6.5 | |
| LNCaP | 2.8 × 109 ± 3.9 × 107 | 2.4 × 1010 ± 8.6 × 108 |
| Me30966 | 9.2 × 108 ± 8.2 × 108 | 9.3 × 1010 ± 1.7 × 1010 |
| SaOS2 | 1.7 × 109 ± 5.4 × 108 | 2.3 × 1010 ± 3.2 × 109 |
| SKBR3 | 1.5 × 109 ± 5.1 × 108 | 8.6 × 109 ± 1.7 × 109 |
| HCT116 | 1.9 × 108 ± 1.1 × 108 | 1.0 × 1010 ± 1.1 × 109 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Logozzi, M.; Mizzoni, D.; Angelini, D.F.; Di Raimo, R.; Falchi, M.; Battistini, L.; Fais, S. Microenvironmental pH and Exosome Levels Interplay in Human Cancer Cell Lines of Different Histotypes. Cancers 2018, 10, 370. https://doi.org/10.3390/cancers10100370
Logozzi M, Mizzoni D, Angelini DF, Di Raimo R, Falchi M, Battistini L, Fais S. Microenvironmental pH and Exosome Levels Interplay in Human Cancer Cell Lines of Different Histotypes. Cancers. 2018; 10(10):370. https://doi.org/10.3390/cancers10100370
Chicago/Turabian StyleLogozzi, Mariantonia, Davide Mizzoni, Daniela F. Angelini, Rossella Di Raimo, Mario Falchi, Luca Battistini, and Stefano Fais. 2018. "Microenvironmental pH and Exosome Levels Interplay in Human Cancer Cell Lines of Different Histotypes" Cancers 10, no. 10: 370. https://doi.org/10.3390/cancers10100370
APA StyleLogozzi, M., Mizzoni, D., Angelini, D. F., Di Raimo, R., Falchi, M., Battistini, L., & Fais, S. (2018). Microenvironmental pH and Exosome Levels Interplay in Human Cancer Cell Lines of Different Histotypes. Cancers, 10(10), 370. https://doi.org/10.3390/cancers10100370






