An Electromagnetically Actuated Double-Sided Cell-Stretching Device for Mechanobiology Research
Abstract
:1. Introduction
2. Materials and Methods
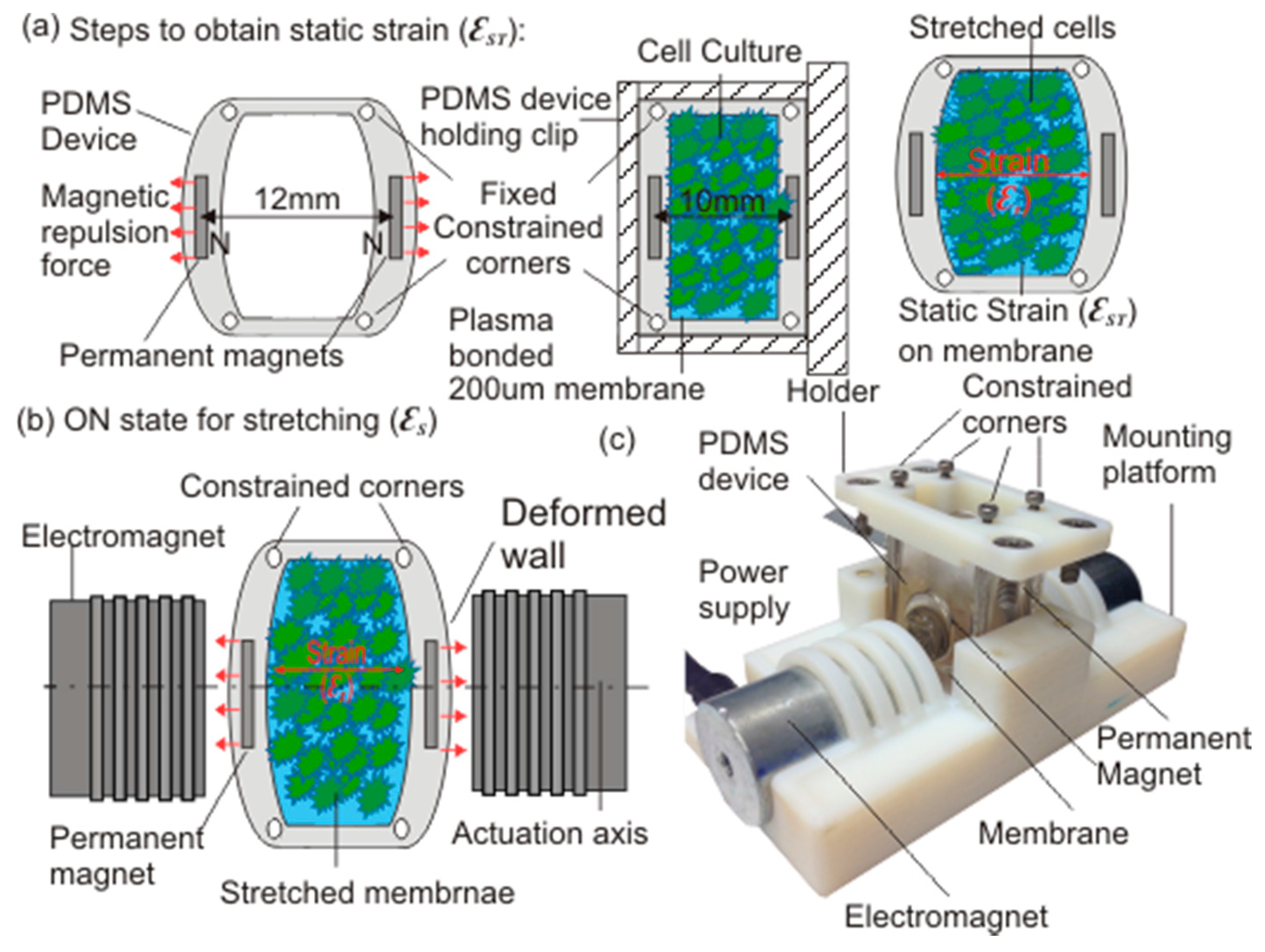
2.1. Device Design and Working Principle
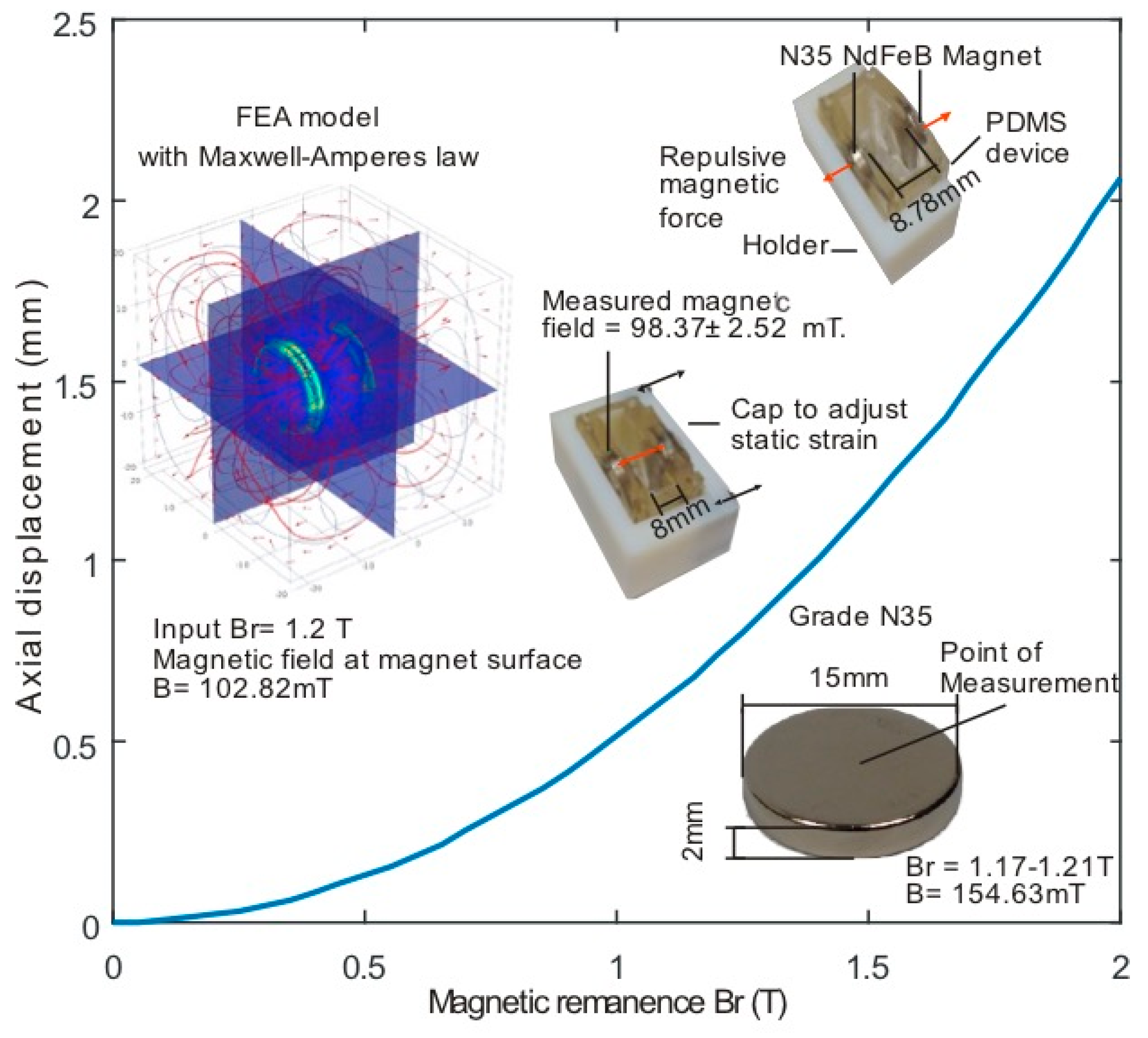
2.2. Modelling and Fabrication
2.3. Cell Culture
2.4. Application of Strain on Fibroblasts
2.5. Cell Fixing, Immunofluorescence Staining and Imaging
2.6. Image Analysis
3. Results and Discussion
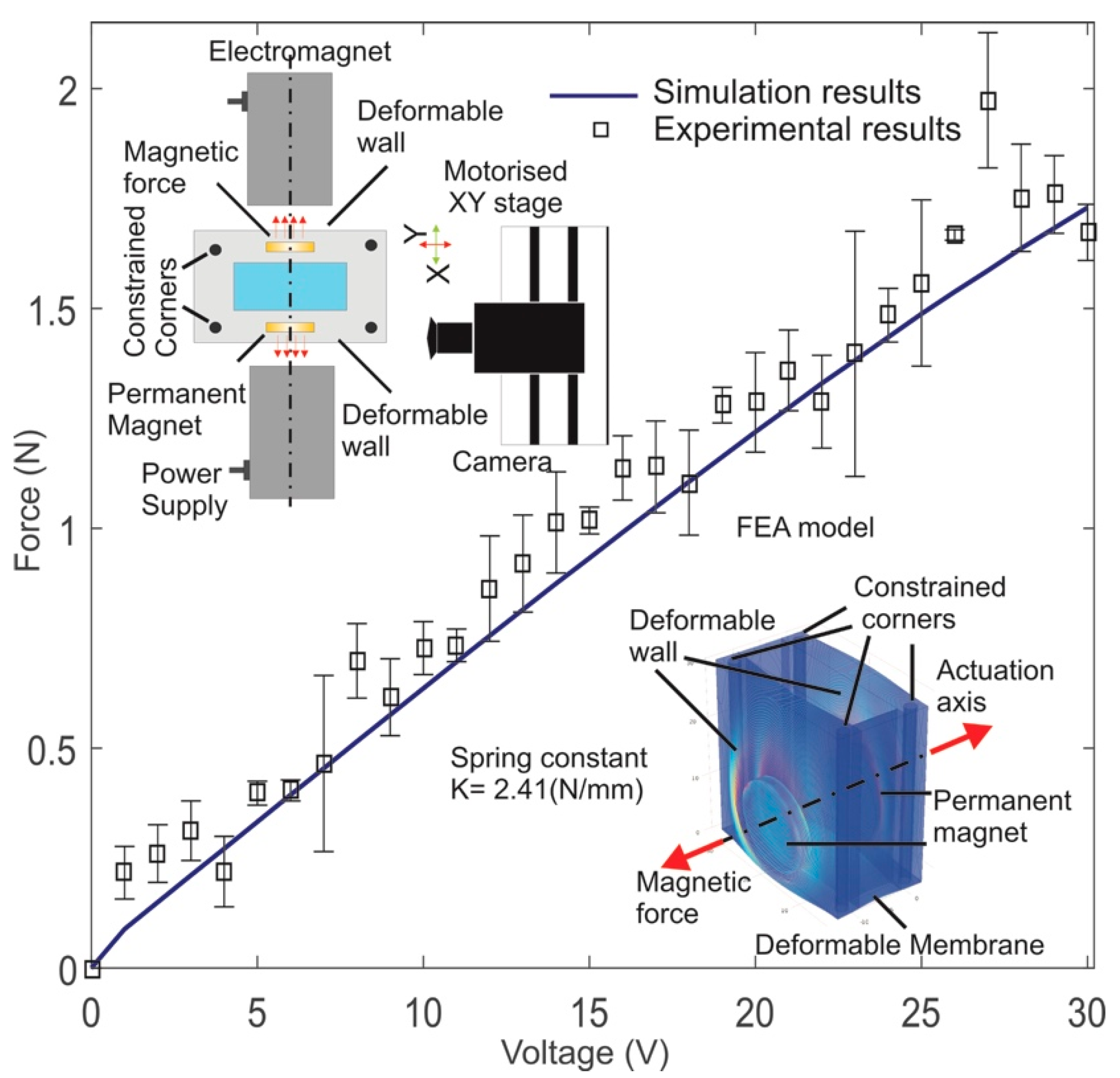
3.1. Force Calculation
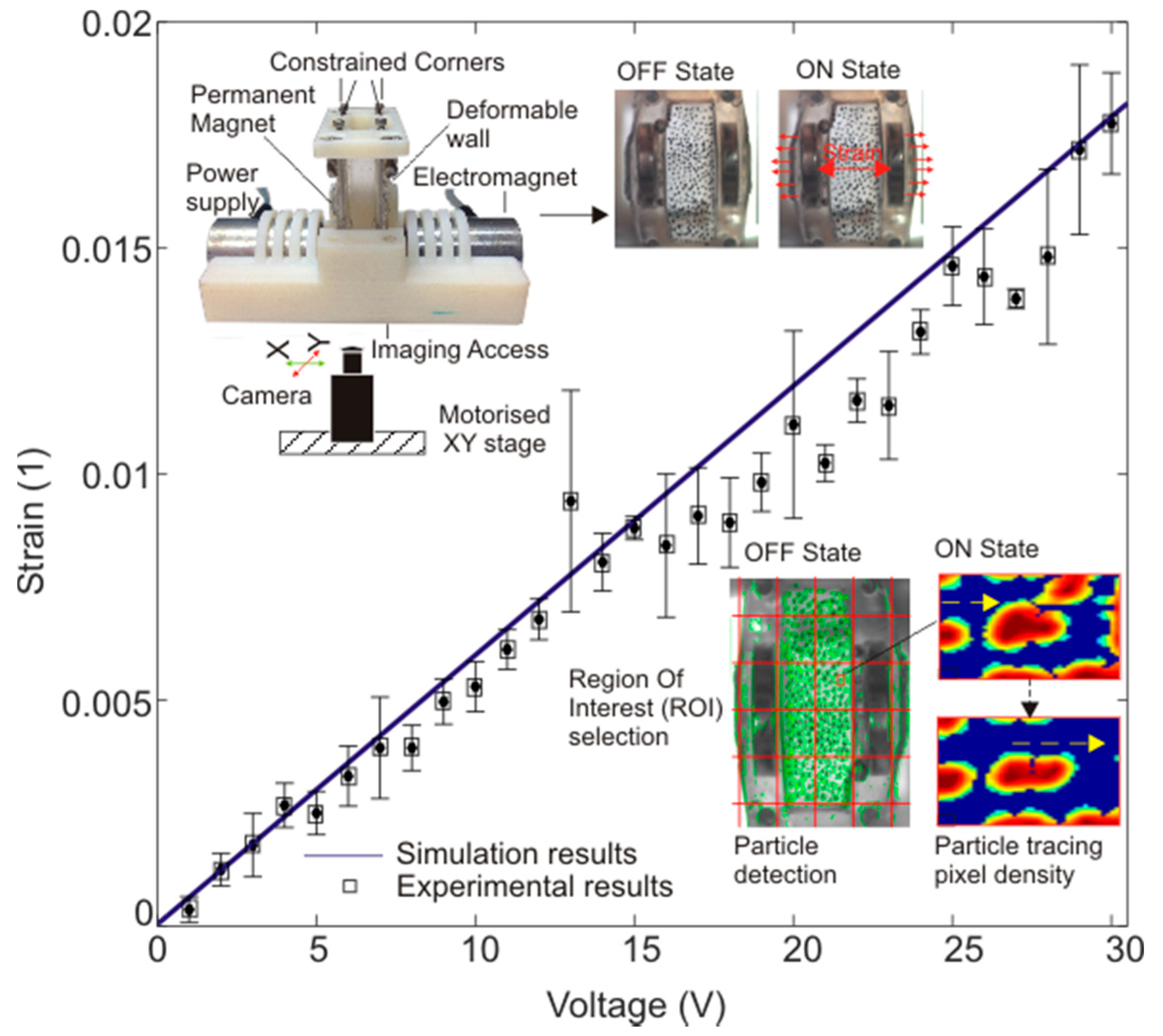
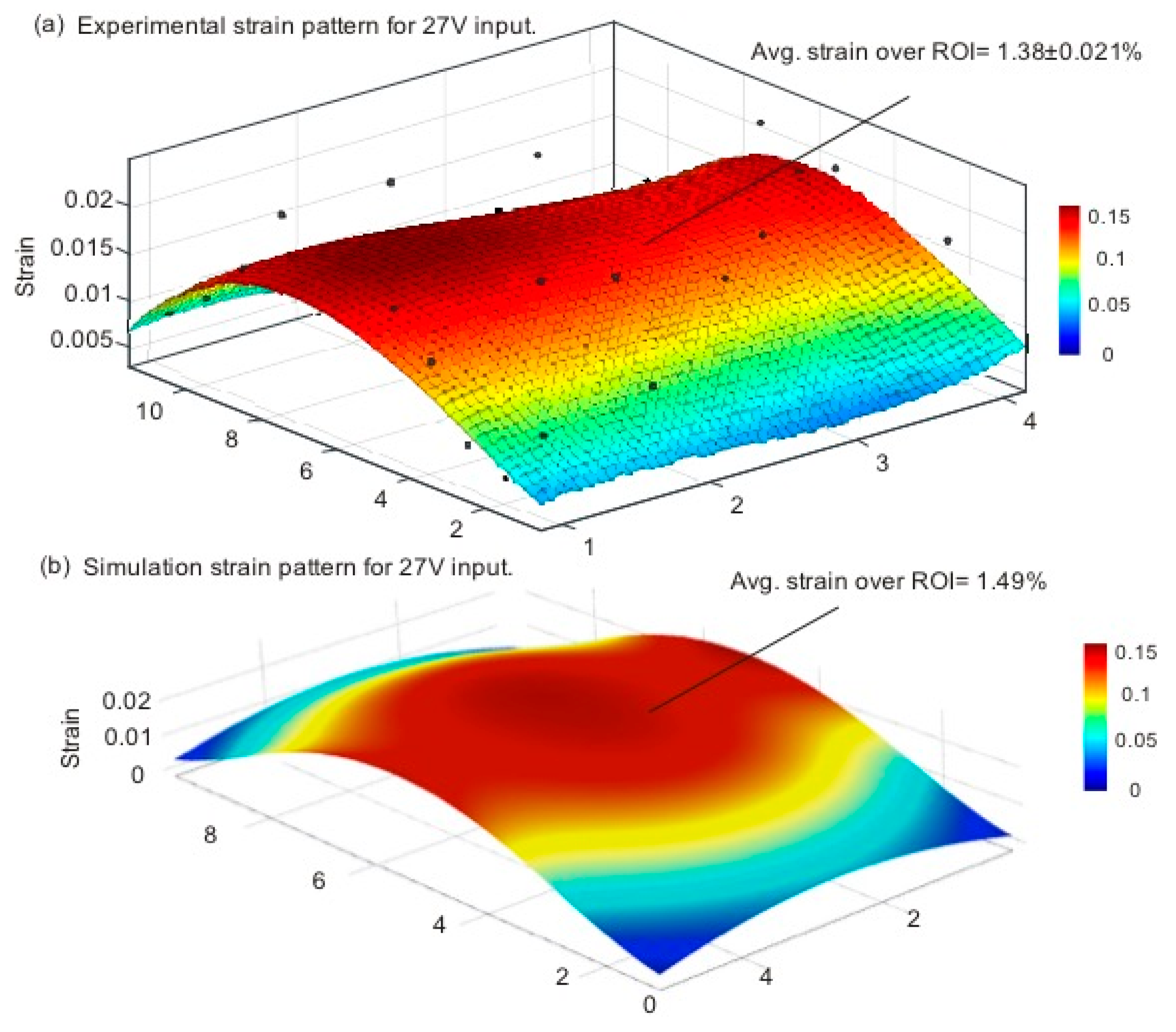
3.2. Strain Calculation
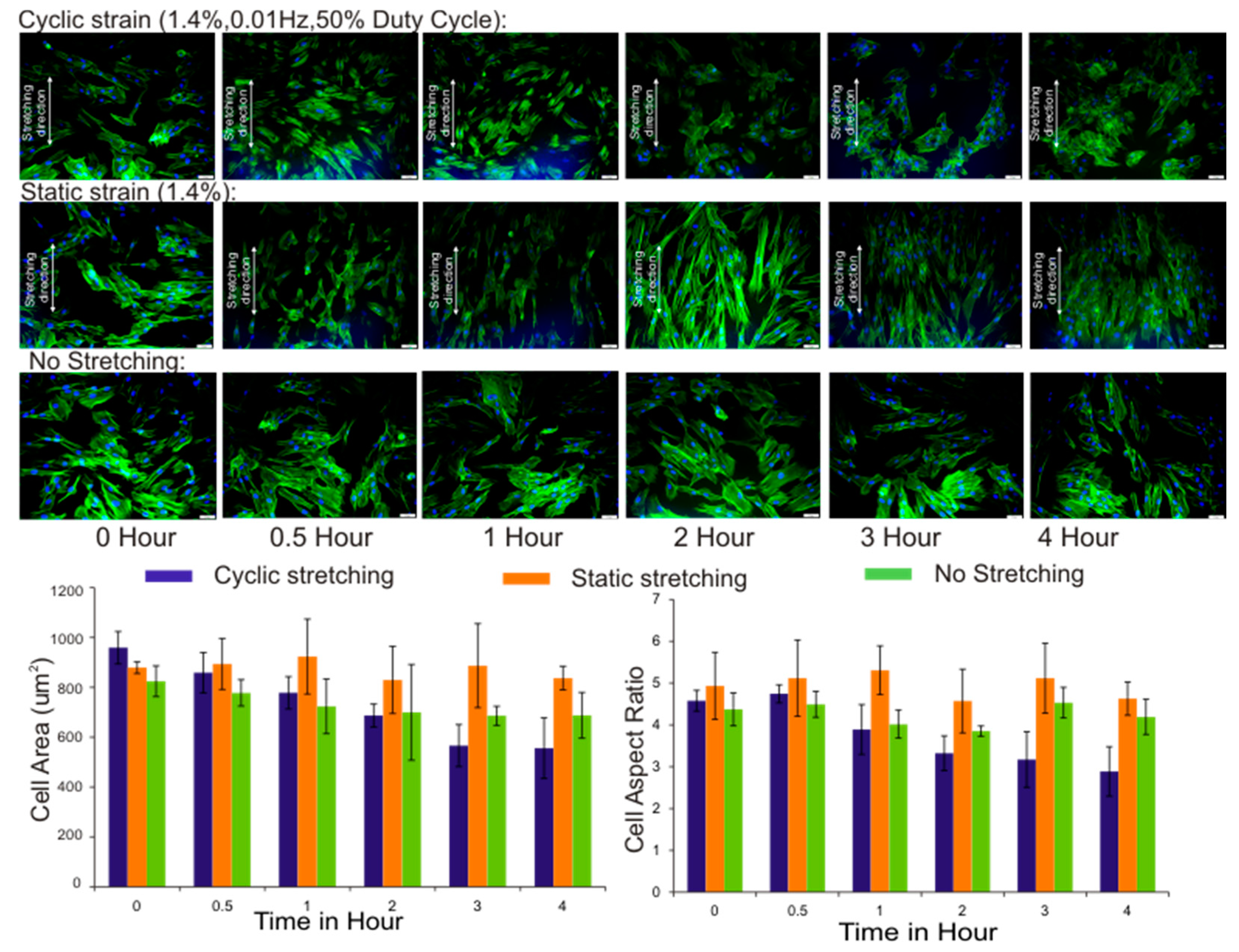
3.3. Cell Area and Aspect Ratio
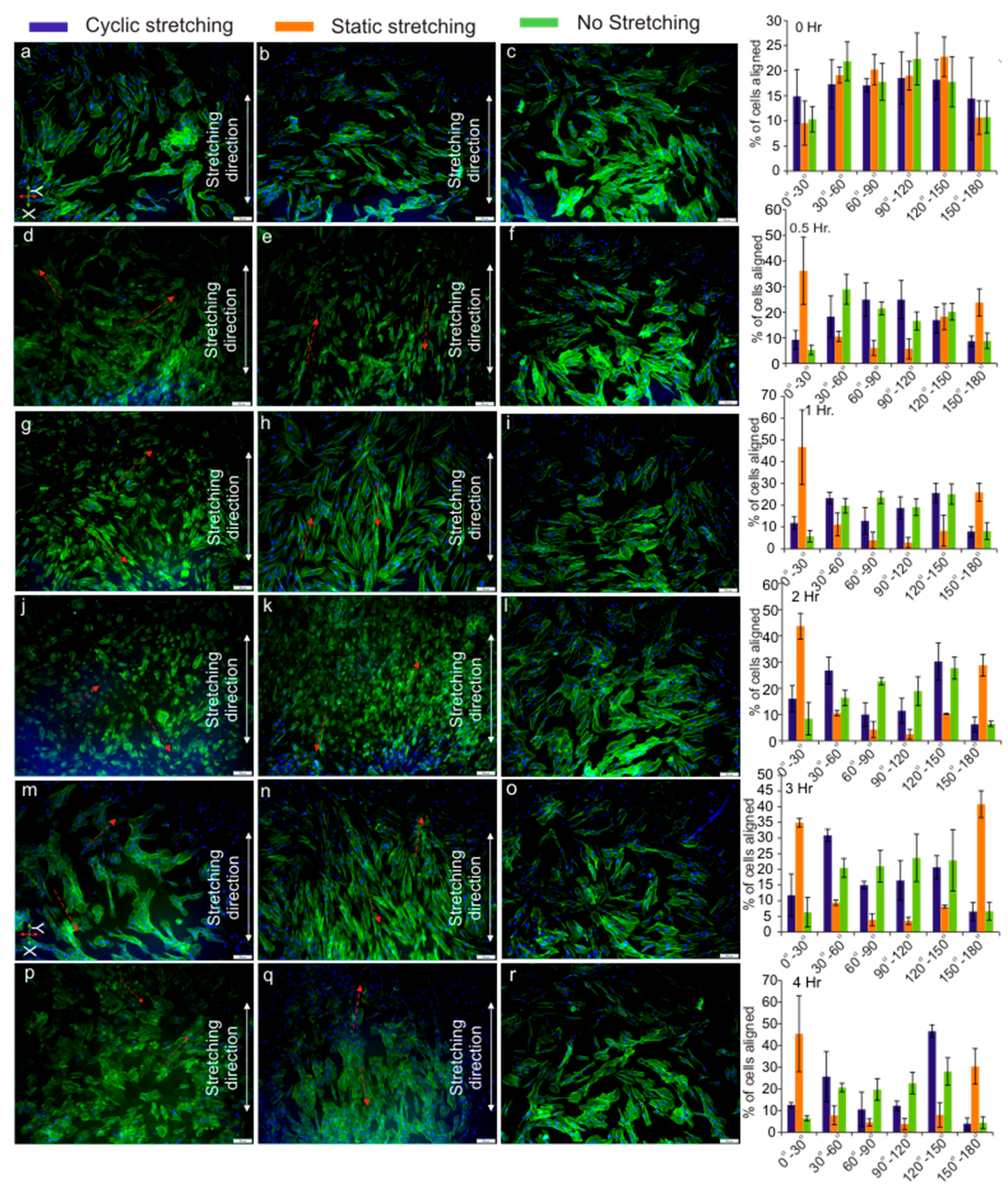
3.4. Cell Orientation
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Chen, C.S. Mechanotransduction—A field pulling together? J. Cell Sci. 2008, 121, 3285–3292. [Google Scholar] [CrossRef] [PubMed]
- Silver, F.H.; Siperko, L.M. Mechanosensing and mechanochemical transduction: How is mechanical energy sensed and converted into chemical energy in an extracellular matrix? Crit. Rev. Biomed. Eng. 2003, 31, 255–331. [Google Scholar] [CrossRef] [PubMed]
- Orr, A.W.; Helmke, B.P.; Blackman, B.R.; Schwartz, M.A. Mechanisms of mechanotransduction. Dev. Cell 2006, 10, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Ingber, D.E. Mechanobiology and diseases of mechanotransduction. Ann. Med. 2003, 35, 564–577. [Google Scholar] [CrossRef] [PubMed]
- DuFort, C.C.; Paszek, M.J.; Weaver, V.M. Balancing forces: Architectural control of mechanotransduction. Nat. Rev. Mol. Cell Biol. 2011, 12, 308–319. [Google Scholar] [CrossRef] [PubMed]
- Mammoto, T.; Ingber, D.E. Mechanical control of tissue and organ development. Development 2010, 137, 1407–1420. [Google Scholar] [CrossRef] [PubMed]
- Klingberg, F.; Hinz, B.; White, E.S. The myofibroblast matrix: Implications for tissue repair and fibrosis. J. Pathol. 2013, 229, 298–309. [Google Scholar] [CrossRef] [PubMed]
- Chiquet, M.; Renedo, A.S.; Huber, F.; Flück, M. How do fibroblasts translate mechanical signals into changes in extracellular matrix production? Matrix Biol. 2003, 22, 73–80. [Google Scholar] [CrossRef]
- Schwartz, M.A. Integrins and extracellular matrix in mechanotransduction. Cold Spring Harb. Perspect. Biol. 2010, 2, a005066. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, D.A.; Mullins, R.D. Cell mechanics and the cytoskeleton. Nature 2010, 463, 485–492. [Google Scholar] [CrossRef] [PubMed]
- Hoon, J.L.; Tan, M.H.; Koh, C.G. The regulation of cellular responses to mechanical cues by Rho GTPases. Cells 2016, 5, 17. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.H.-C.; Yang, G.; Li, Z.; Shen, W. Fibroblast responses to cyclic mechanical stretching depend on cell orientation to the stretching direction. J. Biomech. 2004, 37, 573–576. [Google Scholar] [CrossRef] [PubMed]
- De, R.; Zemel, A.; Safran, S.A. Dynamics of cell orientation. Nat. Phys. 2007, 3, 655–659. [Google Scholar] [CrossRef]
- Deguchi, S.; Kudo, S.; Matsui, T.S.; Huang, W.; Sato, M. Piezoelectric actuator-based cell microstretch device with real-time imaging capability. AIP Adv. 2015, 5, 067110. [Google Scholar] [CrossRef]
- Tremblay, D.; Chagnon-Lessard, S.; Mirzaei, M.; Pelling, A.E.; Godin, M. A microscale anisotropic biaxial cell stretching device for applications in mechanobiology. Biotechnol. Lett. 2014, 36, 657–665. [Google Scholar] [CrossRef] [PubMed]
- Kamble, H.; Barton, M.J.; Jun, M.; Park, S.; Nguyen, N.-T. Cell stretching devices as research tools: Engineering and biological considerations. Lab Chip 2016, 16, 3193–3203. [Google Scholar] [CrossRef] [PubMed]
- Esch, E.W.; Bahinski, A.; Huh, D. Organs-on-chips at the frontiers of drug discovery. Nat. Rev. Drug Discov. 2015, 14, 248–260. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, S.; Seo, K.; Nagasaki, M.; Hosoya, Y.; Yamashita, H.; Fujita, H.; Nagai, R.; Sugiura, S. Responses of single-ventricular myocytes to dynamic axial stretching. Prog. Biophys. Mol. Biol. 2008, 97, 282–297. [Google Scholar] [CrossRef] [PubMed]
- Chaudhuri, O.; Parekh, S.H.; Lam, W.A.; Fletcher, D.A. Combined atomic force microscopy and side-view optical imaging for mechanical studies of cells. Nat. Methods 2009, 6, 383–387. [Google Scholar] [CrossRef] [PubMed]
- Sniadecki, N.J.; Lamb, C.M.; Liu, Y.; Chen, C.S.; Reich, D.H. Magnetic microposts for mechanical stimulation of biological cells: Fabrication, characterization, and analysis. Rev. Sci. Instrum. 2008, 79, 044302. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.H.; Sun, D.; Wang, J.Z.; Huang, W.H. Mechanical characterization of human red blood cells under different osmotic conditions by robotic manipulation with optical tweezers. IEEE Trans. Biomed. Eng. 2010, 57, 1816–1825. [Google Scholar] [CrossRef] [PubMed]
- Dhein, S.; Schreiber, A.; Steinbach, S.; Apel, D.; Salameh, A.; Schlegel, F.; Kostelka, M.; Dohmen, P.M.; Mohr, F.W. Mechanical control of cell biology. Effects of cyclic mechanical stretch on cardiomyocyte cellular organization. Prog. Biophys. Mol. Biol. 2014, 115, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Higgins, S.; Lee, J.S.; Ha, L.; Lim, J.Y. Inducing neurite outgrowth by mechanical cell stretch. Biores. Open Access 2013, 2, 212–216. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Potter, R.; Li, X.; Flessner, M. Stretch of human mesothelial cells increases cytokine expression. Adv. Perit. Dial. 2012, 28, 2–9. [Google Scholar] [PubMed]
- Nava, G.; Bragheri, F.; Yang, T.; Minzioni, P.; Osellame, R.; Cristiani, I.; Berg-Sørensen, K. All-silica microfluidic optical stretcher with acoustophoretic prefocusing. Microfluid. Nanofluid. 2015, 19, 837–844. [Google Scholar] [CrossRef]
- Huang, Y.; Nguyen, N.T. A polymeric cell stretching device for real-time imaging with optical microscopy. Biomed. Microdevices 2013, 15, 1043–1054. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Tan, X.; Novitski, R.; Muqaddam, M.; List, P.; Williamson, L.; Fu, J.; Liu, A.P. Uniaxial cell stretching device for live-cell imaging of mechanosensitive cellular functions. Rev. Sci. Instrum. 2013, 84, 114304. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, K.; Shunori, A.; Morimoto, K.; Hashida, M.; Konishi, S. Development of a biochip with serially connected pneumatic balloons for cell-stretching culture. Sens. Actuators B Chem. 2011, 156, 486–493. [Google Scholar] [CrossRef]
- Kamotani, Y.; Bersano-Begey, T.; Kato, N.; Tung, Y.C.; Huh, D.; Song, J.W.; Takayama, S. Individually programmable cell stretching microwell arrays actuated by a braille display. Biomaterials 2008, 29, 2646–2655. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Mathieu, P.S.; Helmke, B.P. A stretching device for high-resolution live-cell imaging. Ann. Biomed. Eng. 2010, 38, 1728–1740. [Google Scholar] [CrossRef] [PubMed]
- Sraj, I.; Eggleton, C.D.; Jimenez, R.; Hoover, E.; Squier, J.; Chichester, J.; Marr, D.W.M. Cell deformation cytometry using diode-bar optical stretchers. J. Biomed. Opt. 2010, 15, 047010. [Google Scholar] [CrossRef] [PubMed]
- Scuor, N.; Gallina, P.; Panchawagh, H.V.; Mahajan, R.L.; Sbaizero, O.; Sergo, V. Design of a novel MEMS platform for the biaxial stimulation of living cells. Biomed. Microdevices 2006, 8, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Iwadate, Y.; Yumura, S. Cyclic stretch of the substratum using a shape-memory alloy induces directional migration in dictyostelium cells. Biotechniques 2009, 47, 757–767. [Google Scholar] [CrossRef] [PubMed]
- Guido, I.; Xiong, C.; Jaeger, M.S.; Duschl, C. Microfluidic system for cell mechanics analysis through dielectrophoresis. Microelectron. Eng. 2012, 97, 379–382. [Google Scholar] [CrossRef]
- Kamble, H.; Barton, M.J.; Nguyen, N.-T. Modelling of an uniaxial single-sided magnetically actuated cell-stretching device. Sens. Actuators A Phys. 2016, 252, 174–179. [Google Scholar] [CrossRef]
- Armani, D.; Liu, C.; Aluru, N. Re-configurable fluid circuits by PDMS elastomer micromachining. In Proceedings of the Twelfth IEEE International Conference on Micro Electro Mechanical Systems, Orlando, FL, USA, 17–21 January 1999; pp. 222–227. [Google Scholar]
- Choi, S.; Park, J.K. Tuneable hydrophoretic separation using elastic deformation of poly(dimethylsiloxane). Lab Chip 2009, 9, 1962–1965. [Google Scholar] [CrossRef] [PubMed]
- Roh, C.; Lee, J.; Kang, C. The deformation of polydimethylsiloxane (PDMS) microfluidic channels filled with embedded circular obstacles under certain circumstances. Molecules 2016, 21, 798. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.-L.; Huang, Y.-C.; Fang, W.; Hsieh, J. A novel electromagnetic elastomer membrane actuator with a semi-embedded coil. Sens. Actuators A Phys. 2007, 139, 194–202. [Google Scholar] [CrossRef]
- Koschwanez, J.H.; Carlson, R.H.; Meldrum, D.R. Thin PDMS films using long spin times or tert-butyl alcohol as a solvent. PLoS ONE 2009, 4, e4572. [Google Scholar] [CrossRef] [PubMed]
- Harshad, K.; Jun, M.; Park, S.; Barton, M.J.; Vadivelu, R.K.; St John, J.; Nguyen, N.T. An electromagnetic cell-stretching device for mechanotransduction studies of olfactory ensheathing cells. Biomed. Microdevices 2016, 18, 45. [Google Scholar] [CrossRef] [PubMed]
- Mann, J.M.; Lam, R.H.; Weng, S.; Sun, Y.; Fu, J. A silicone-based stretchable micropost array membrane for monitoring live-cell subcellular cytoskeletal response. Lab Chip 2012, 12, 731–740. [Google Scholar] [CrossRef] [PubMed]
- Mao, X.; Said, R.; Louis, H.; Max, J.P.; Bourhim, M.; Challande, P.; Wahl, D.; Li, Z.; Regnault, V.; Lacolley, P. Cyclic stretch-induced thrombin generation by rat vascular smooth muscle cells is mediated by the integrin αvβ3 pathway. Cardiovasc. Res. 2012, 96, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Sethi, K.; Cram, E.J.; Zaidel-Bar, R. Stretch-induced actomyosin contraction in epithelial tubes: Mechanotransduction pathways for tubular homeostasis. Semin. Cell Dev. Biol. 2017, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Geiger, B.; Yamada, K.M. Molecular architecture and function of matrix adhesions. Cold Spring Harb. Perspect. Biol. 2011, 3, a00503. [Google Scholar] [CrossRef] [PubMed]
- Boccafoschi, F.; Bosetti, M.; Sandra, P.M.; Leigheb, M.; Cannas, M. Effects of mechanical stress on cell adhesion: A possible mechanism for morphological changes. Cell Adhes. Migr. 2010, 4, 19–25. [Google Scholar] [CrossRef]
- Kassianidou, E.; Kumar, S. A biomechanical perspective on stress fiber structure and function. Biochim. Biophys. Acta 2015, 1853, 3065–3074. [Google Scholar] [CrossRef] [PubMed]
- Kaunas, R. Sarcomeric model of stretch-induced stress fiber reorganization. Cell Health Cytoskelet. 2010, 2011, 13–22. [Google Scholar] [CrossRef]
- Ngu, H.; Feng, Y.; Lu, L.; Oswald, S.J.; Longmore, G.D.; Yin, F.C. Effect of focal adhesion proteins on endothelial cell adhesion, motility and orientation response to cyclic strain. Ann. Biomed. Eng. 2010, 38, 208–222. [Google Scholar] [CrossRef] [PubMed]
- Bazellieres, E.; Conte, V.; Elosegui-Artola, A.; Serra-Picamal, X.; Bintanel-Morcillo, M.; Roca-Cusachs, P.; Munoz, J.J.; Sales-Pardo, M.; Guimera, R.; Trepat, X. Control of cell-cell forces and collective cell dynamics by the intercellular adhesome. Nat. Cell Biol. 2015, 17, 409–420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Parsons, J.T.; Horwitz, A.R.; Schwartz, M.A. Cell adhesion: Integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 2010, 11, 633–643. [Google Scholar] [CrossRef] [PubMed]
- Verkhovsky, A.B.; Borisy, G.G. Non-sarcomeric mode of myosin ii organization in the fibroblast lamellum. J. Cell Biol. 1993, 123, 637–652. [Google Scholar] [CrossRef] [PubMed]
- Tondon, A.; Kaunas, R. The direction of stretch-induced cell and stress fiber orientation depends on collagen matrix stress. PLoS ONE 2014, 9, e89592. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.; Grande-Allen, K.J. Effects of static and cyclic loading in regulating extracellular matrix synthesis by cardiovascular cells. Cardiovasc. Res. 2006, 72, 375–383. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Hameed, F.M.; Yang, B.; Lee, K.; Pan, C.Q.; Park, S.; Sheetz, M. Cyclic stretching of soft substrates induces spreading and growth. Nat. Commun. 2015, 6, 6333. [Google Scholar] [CrossRef] [PubMed]
- Ugolini, G.S.; Pavesi, A.; Rasponi, M.; Fiore, G.B.; Kamm, R.; Soncini, M. Human cardiac fibroblasts adaptive responses to controlled combined mechanical strain and oxygen changes in vitro. eLife 2017, 6, e22847. [Google Scholar] [CrossRef] [PubMed]
- Ugolini, G.S.; Rasponi, M.; Pavesi, A.; Santoro, R.; Kamm, R.; Fiore, G.B.; Pesce, M.; Soncini, M. On-chip assessment of human primary cardiac fibroblasts proliferative responses to uniaxial cyclic mechanical strain. Biotechnol. Bioeng. 2016, 113, 859–869. [Google Scholar] [CrossRef] [PubMed]
- Pavesi, A.; Adriani, G.; Rasponi, M.; Zervantonakis, I.K.; Fiore, G.B.; Kamm, R.D. Controlled electromechanical cell stimulation on-a-chip. Sci. Rep. 2015, 5, 11800. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Li, Y.; Chen, B.; Liu, S.; Li, M.; Zheng, L.; Wang, P.; Lu, T.J.; Xu, F. Patterning cellular alignment through stretching hydrogels with programmable strain gradients. ACS Appl. Mater. Interfaces 2015, 7, 15088–15097. [Google Scholar] [CrossRef] [PubMed]
- Chagnon-Lessard, S.; Jean-Ruel, H.; Godin, M.; Pelling, A.E. Cellular orientation is guided by strain gradients. Integr. Biol. 2017, 9, 607–618. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.H.; Yang, G.; Li, Z. Controlling cell responses to cyclic mechanical stretching. Ann. Biomed. Eng. 2005, 33, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Huang, H.; Wei, K.; Zhao, Y. Time-dependent combinatory effects of active mechanical loading and passive topographical cues on cell orientation. Biotechnol. Bioeng. 2016, 113, 2191–2201. [Google Scholar] [CrossRef] [PubMed]
- Weidenhamer, N.K.; Tranquillo, R.T. Influence of cyclic mechanical stretch and tissue constraints on cellular and collagen alignment in fibroblast-derived cell sheets. Tissue Eng. Part C Methods 2013, 19, 386–395. [Google Scholar] [CrossRef] [PubMed]
- Abiko, H.; Fujiwara, S.; Ohashi, K.; Hiatari, R.; Mashiko, T.; Sakamoto, N.; Sato, M.; Mizuno, K. Rho guanine nucleotide exchange factors involved in cyclic-stretch-induced reorientation of vascular endothelial cells. J. Cell Sci. 2015, 128, 1683–1695. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulou, A.; Iliadi, A.; Eliades, T.; Kletsas, D. Early responses of human periodontal ligament fibroblasts to cyclic and static mechanical stretching. Eur. J. Orthod. 2016, 39, 258–263. [Google Scholar] [CrossRef] [PubMed]
- Blanchoin, L.; Boujemaa-Paterski, R.; Sykes, C.; Plastino, J. Actin dynamics, architecture, and mechanics in cell motility. Physiol. Rev. 2014, 94, 235–263. [Google Scholar] [CrossRef] [PubMed]
- Burridge, K.; Guilluy, C. Focal adhesions, stress fibers and mechanical tension. Exp. Cell Res. 2016, 343, 14–20. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kamble, H.; Vadivelu, R.; Barton, M.; Boriachek, K.; Munaz, A.; Park, S.; Shiddiky, M.J.A.; Nguyen, N.-T. An Electromagnetically Actuated Double-Sided Cell-Stretching Device for Mechanobiology Research. Micromachines 2017, 8, 256. https://doi.org/10.3390/mi8080256
Kamble H, Vadivelu R, Barton M, Boriachek K, Munaz A, Park S, Shiddiky MJA, Nguyen N-T. An Electromagnetically Actuated Double-Sided Cell-Stretching Device for Mechanobiology Research. Micromachines. 2017; 8(8):256. https://doi.org/10.3390/mi8080256
Chicago/Turabian StyleKamble, Harshad, Raja Vadivelu, Mathew Barton, Kseniia Boriachek, Ahmed Munaz, Sungsu Park, Muhammad J. A. Shiddiky, and Nam-Trung Nguyen. 2017. "An Electromagnetically Actuated Double-Sided Cell-Stretching Device for Mechanobiology Research" Micromachines 8, no. 8: 256. https://doi.org/10.3390/mi8080256
APA StyleKamble, H., Vadivelu, R., Barton, M., Boriachek, K., Munaz, A., Park, S., Shiddiky, M. J. A., & Nguyen, N.-T. (2017). An Electromagnetically Actuated Double-Sided Cell-Stretching Device for Mechanobiology Research. Micromachines, 8(8), 256. https://doi.org/10.3390/mi8080256






