3D Cardiac Cell Culture on Nanofiber Bundle Substrates for the Investigation of Cell Morphology and Contraction
Abstract
:1. Introduction
2. Methodology
3. Experimental Section
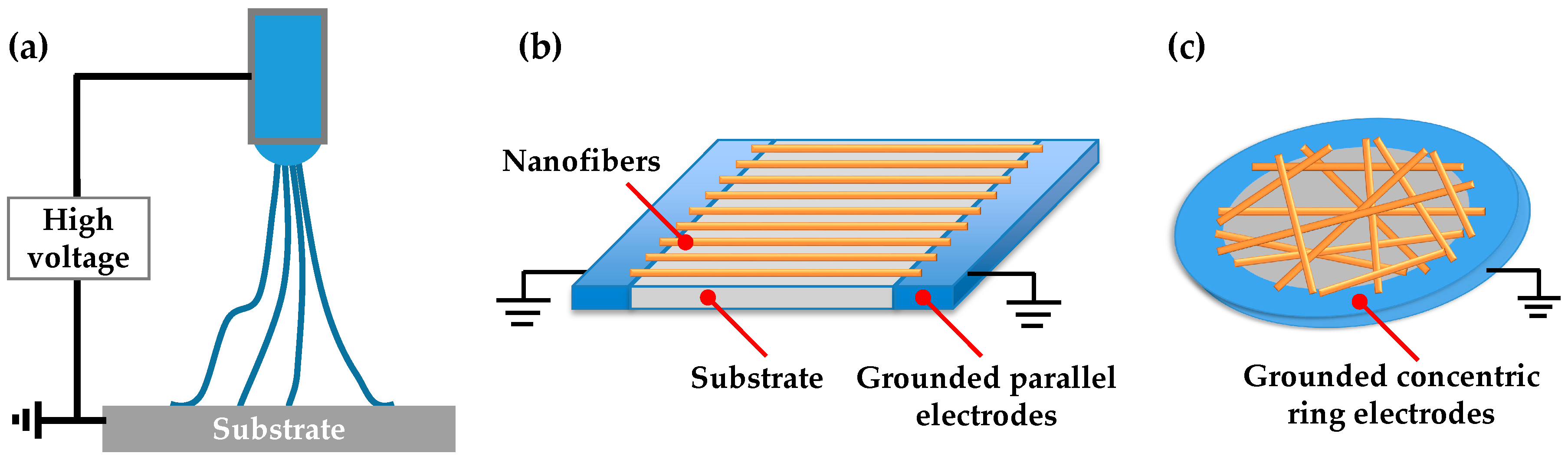
3.1. Nanofiber Preparation
3.2. Nanofiber Surface Functionalization
3.3. Cell Isolation and Cell Culture
3.4. Cellular Characterization
3.4.1. Environmental Scanning Electron Microscopy
3.4.2. Immunofluorescence Staining and Imaging
3.4.3. Atomic Force Microscopy Characterization
4. Results and Discussion
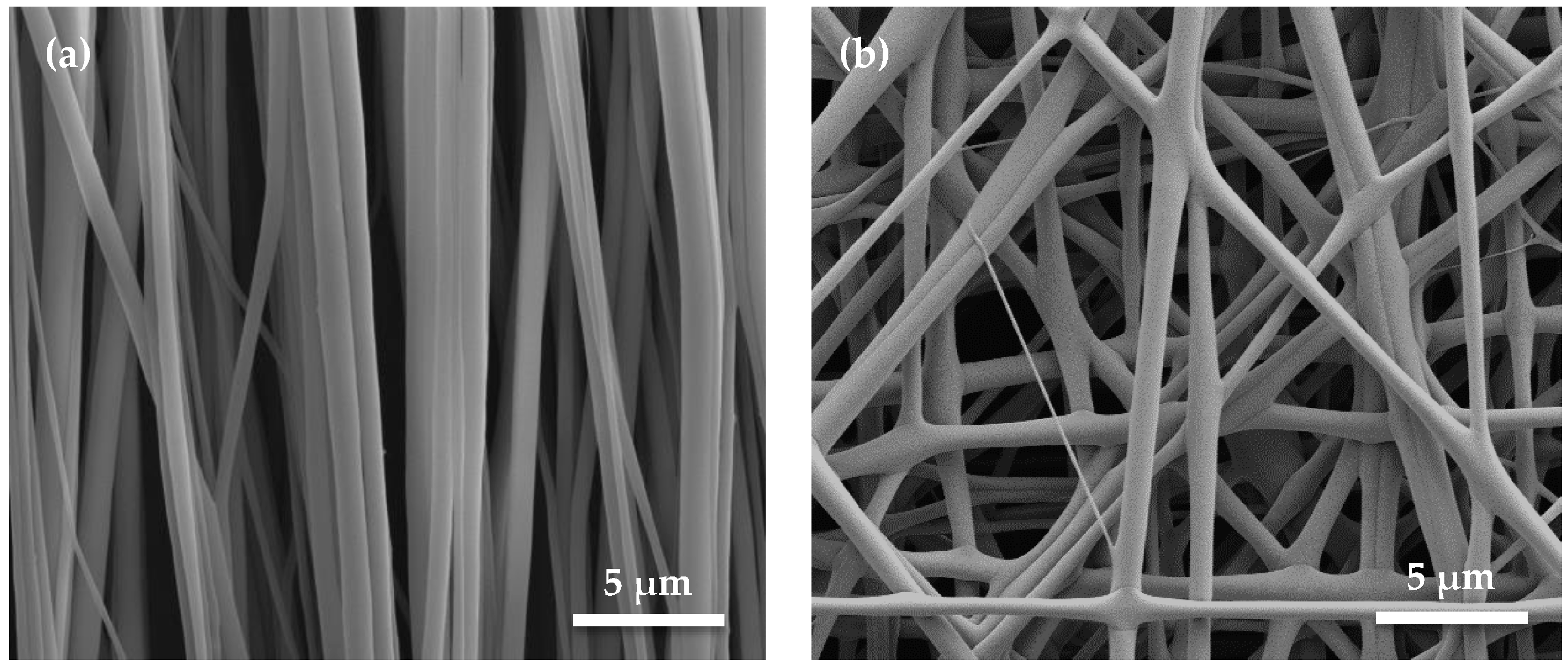
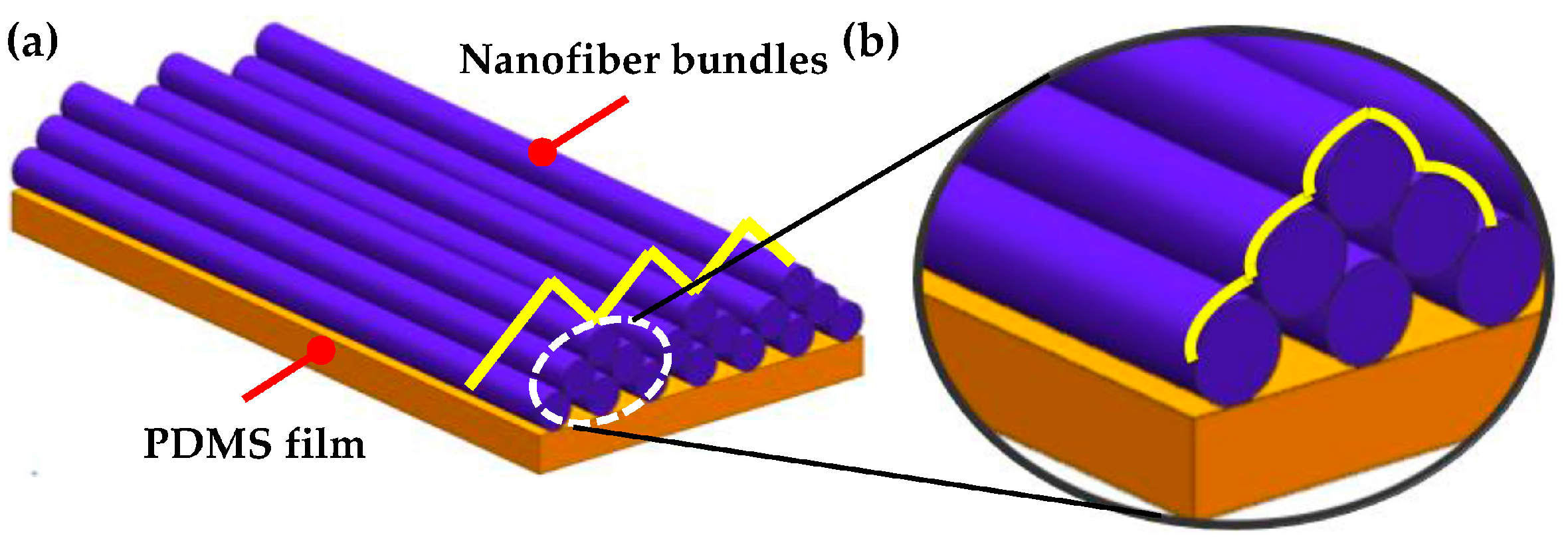
4.1. Pattern of Nanofiber Bundles
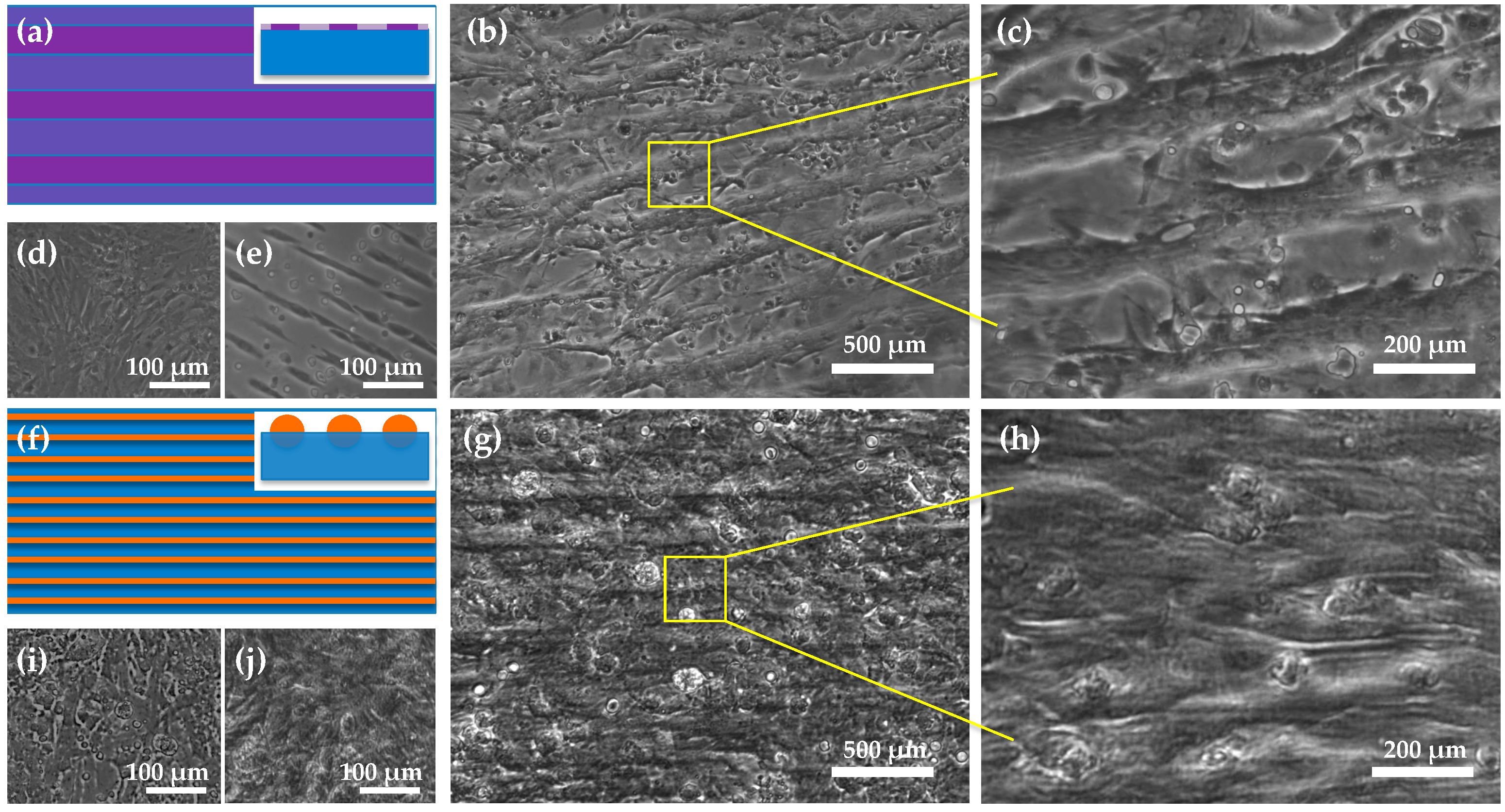
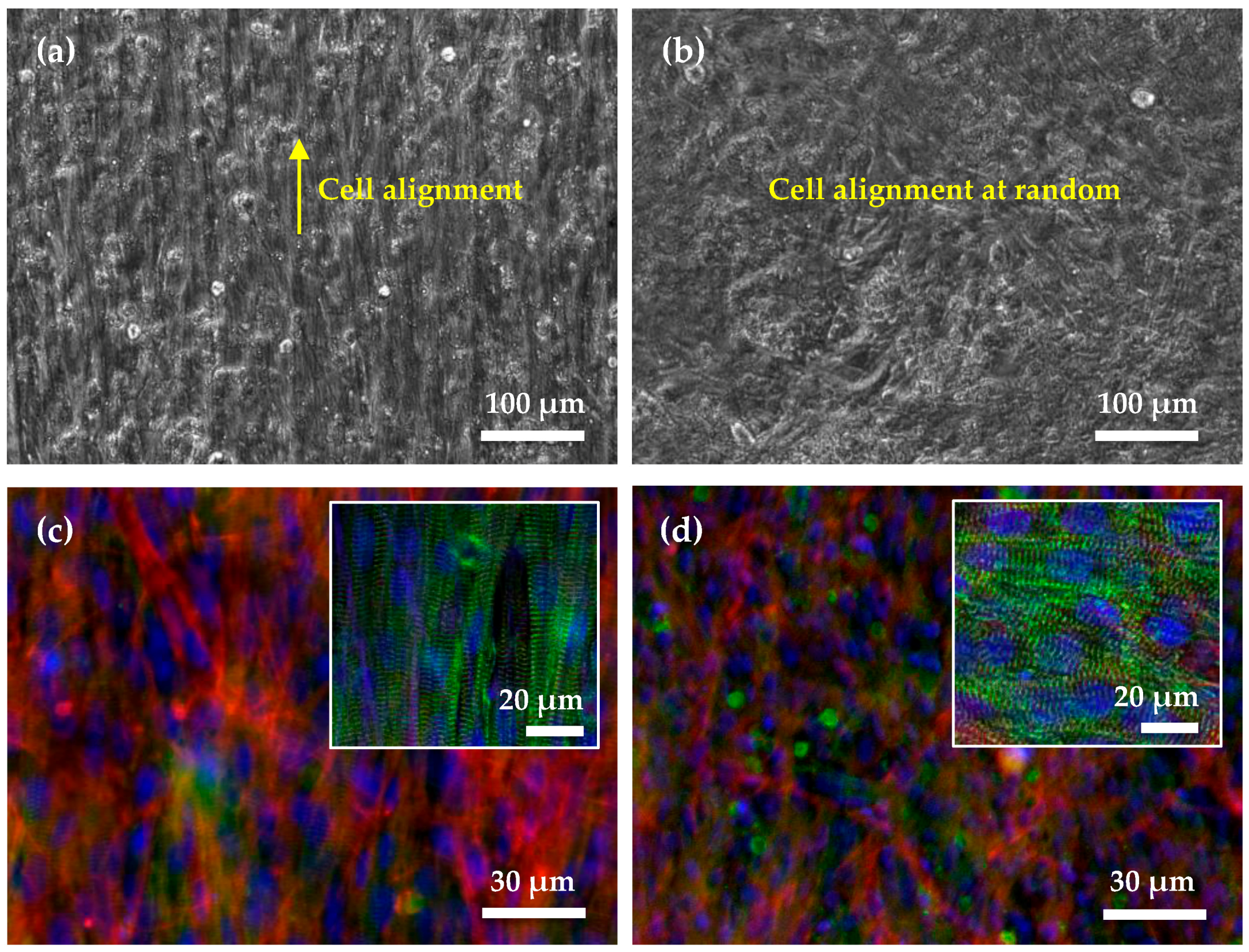
4.2. Microstructures of 2D Cardiac Cell Sheets
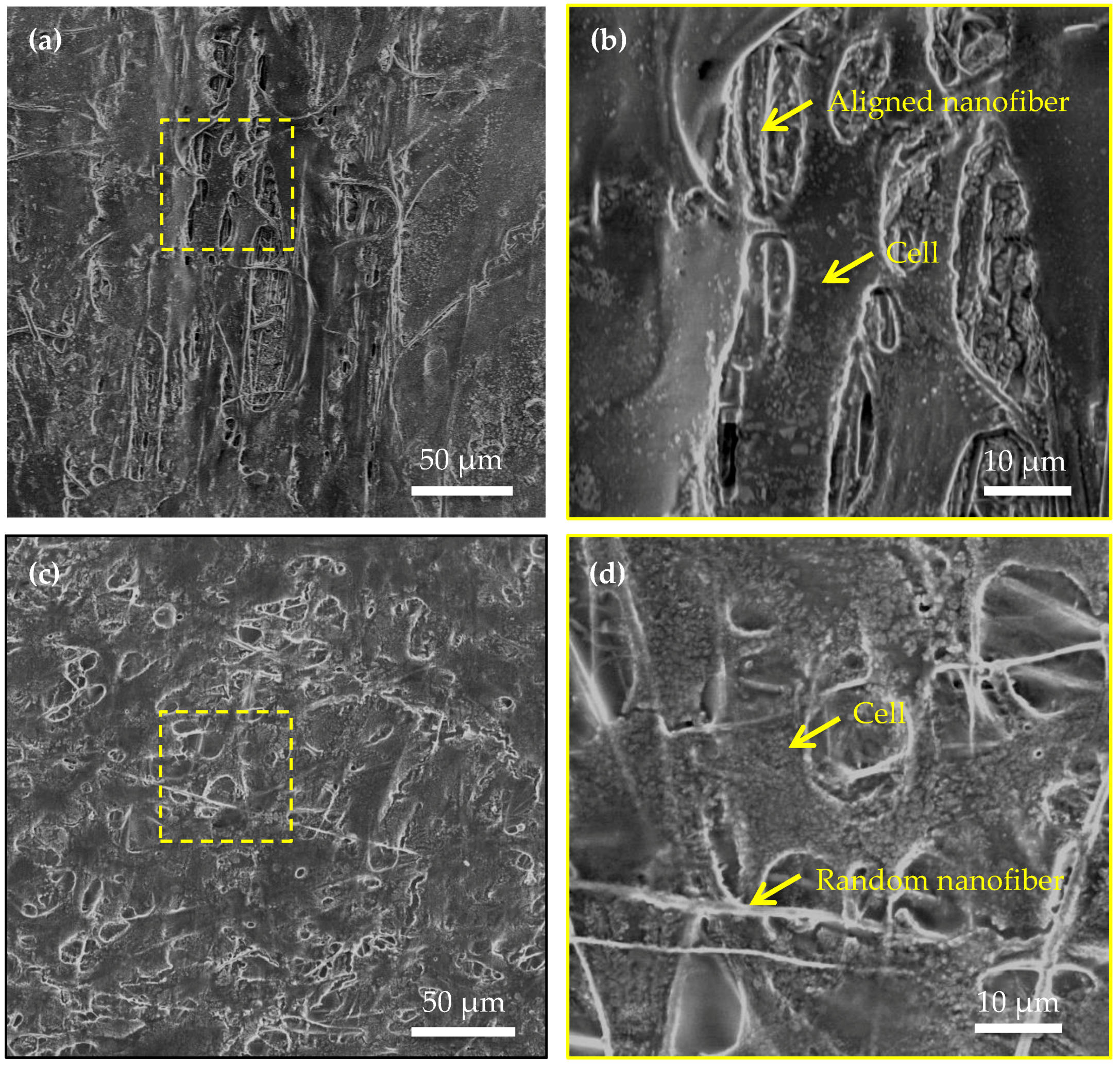
4.3. Shape of Single Cell
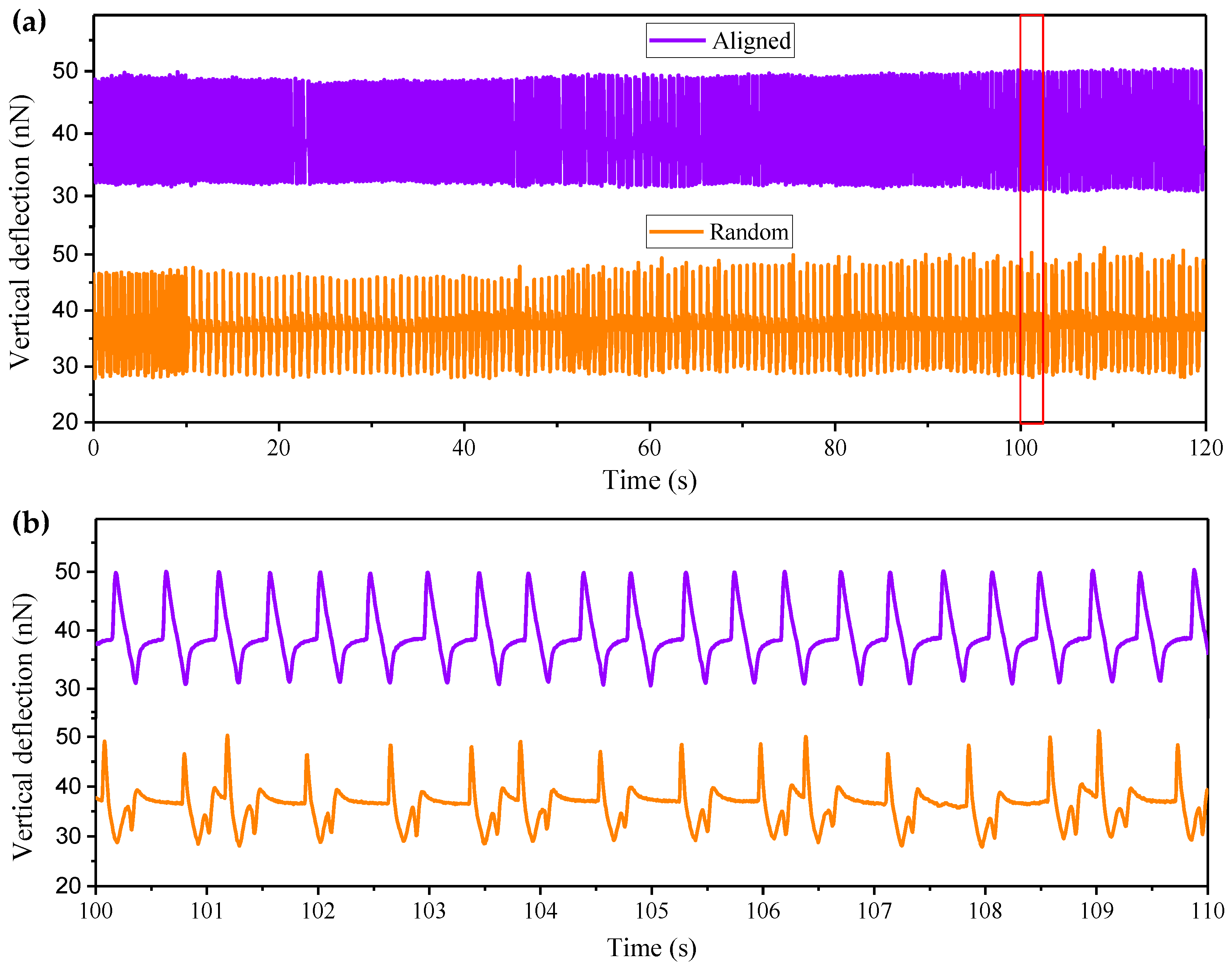
4.4. Contraction of Cell Sheet
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Zuppinger, C. 3D culture for cardiac cells. Biochim. Biophys. Acta 2015, 1863, 1873–1881. [Google Scholar] [CrossRef] [PubMed]
- Battista, S.; Guarnieri, D.; Borselli, C.; Zeppetelli, S.; Borzacchiello, A.; Mayol, L.; Gerbasio, D.; Keene, D.R.; Ambrosio, L.; Netti, P.A. The effect of matrix composition of 3D constructs on embryonic stem cell differentiation. Biomaterials 2005, 26, 6194–6207. [Google Scholar] [CrossRef] [PubMed]
- Ikonen, L.; Kerkelä, E.; Metselaar, G.; Stuart, M.C.; de Jong, M.R.; Aaltosetälä, K. 2D and 3D self-assembling nanofiber hydrogels for cardiomyocyte culture. BioMed Res. Int. 2013, 2013, 285678. [Google Scholar] [CrossRef] [PubMed]
- Emmert, M.Y.; Hitchcock, R.W.; Hoerstrup, S.P. Cell therapy, 3D culture systems and tissue engineering for cardiac regeneration. Adv. Drug Deliv. Rev. 2014, 69–70, 254–269. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Mhawechfauceglia, P.; Lee, N.; Parsanian, L.C.; Lin, Y.G.; Gayther, S.A.; Lawrenson, K. A three-dimensional microenvironment alters protein expression and chemosensitivity of epithelial ovarian cancer cells in vitro. Lab. Investig. 2013, 93, 528–542. [Google Scholar] [PubMed]
- Li, Z.; Guan, J. Hydrogels for cardiac tissue engineering. Polymers 2011, 3, 740–761. [Google Scholar] [CrossRef]
- Khaoustov, V.I.; Darlington, G.J.; Soriano, H.E.; Krishnan, B.; Risin, D.; Pellis, N.R.; Yoffe, B. Induction of threedimensional assembly of human liver cells by simulated microgravity. In Vitro Cell. Dev. Biol. Anim. 1999, 35, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Noto, T.; Tokuda, Y.; Nakamura, Y.; Suzuki, A.; Watanabe, K.; Yamamura, M.; Tajima, T.; Mitomi, T.; Nishijima, K. A new high-yield continuous cell-culture system for lymphokine-activated killer cells. Cancer Immunol. Immunother. 1989, 30, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Barcellos-Hoff, M.H.; Aggeler, J.; Ram, T.G.; Bissell, M.J. Functional differentiation and alveolar morphogenesis of primary mammary cultures on reconstituted basement membrane. Development 1989, 105, 223–235. [Google Scholar] [PubMed]
- Hutmacher, D.W. Scaffold design and fabrication technologies for engineering tissues—State of the art and future perspectives. J. Biomater. Sci. Polym. Ed. 2001, 12, 107–124. [Google Scholar] [CrossRef] [PubMed]
- Lawrenson, K.; Benjamin, E.; Turmaine, M.; Jacobs, I.; Gayther, S.; Dafou, D. In vitro three-dimensional modelling of human ovarian surface epithelial cells. Cell Prolif. 2009, 42, 385–393. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Spagnoli, G.C.; Martin, I.; Ploegert, S.; Demougin, P.; Heberer, M.; Reschner, A. Three-dimensional culture of melanoma cells profoundly affects gene expression profile: A high density oligonucleotide array study. J. Cell. Physiol. 2005, 204, 522–531. [Google Scholar] [CrossRef] [PubMed]
- Yeong, W.Y.; Sudarmadji, N.; Yu, H.Y.; Chua, C.K.; Leong, K.F.; Venkatraman, S.S.; Boey, Y.C.F.; Tan, L.P. Porous polycaprolactone scaffold for cardiac tissue engineering fabricated by selective laser sintering. Acta Biomater. 2010, 6, 2028–2034. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Leong, K.F.; Du, Z.; Chua, C.K. The design of scaffolds for use in tissue engineering. Part II. Rapid prototyping techniques. Tissue Eng. 2002, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Sing, S.L.; Tan, E.Y.S.; Yeong, W.Y. Bioprinting in cardiovascular tissue engineering: A review. Int. J. Bioprint. 2016, 2, 27–36. [Google Scholar] [CrossRef]
- Wang, H.; Vijayavenkataraman, S.; Wu, Y.; Shu, Z.; Sun, J.; Hsi, J.F.Y. Investigation of process parameters of electrohydro-dynamic jetting for 3D printed PCL fibrous scaffolds with complex geometries. Int. J. Bioprint. 2016, 2, 63–71. [Google Scholar] [CrossRef]
- Wang, M.; He, J.; Liu, Y.; Li, M.; Li, D.; Jin, Z. The trend towards in vivo bioprinting. Int. J. Bioprint. 2015, 1, 15–26. [Google Scholar] [CrossRef]
- Bhuthalingam, R.; Lim, P.Q.; Irvine, S.A.; Agrawal, A.; Mhaisalkar, P.S.; An, J.; Chua, C.K.; Venkatraman, S. A novel 3D printing method for cell alignment and differentiation. Int. J. Bioprint. 2015, 1, 57–65. [Google Scholar]
- Jahnke, H.G.; Steel, D.; Fleischer, S.; Seidel, D.; Kurz, R.; Vinz, S.; Dahlenborg, K.; Sartipy, P.; Robitzki, A.A. A novel 3D label-free monitoring system of hES-derived cardiomyocyte clusters: A step forward to in vitro cardiotoxicity testing. PLoS ONE 2013, 8, e68971. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; He, J.; Xu, F.; Liu, Y.; Li, D. Electrohydrodynamic printing: A potential tool for high-resolution hydrogel/cell patterning. Virtual Phys. Prototyp. 2016, 11, 57–63. [Google Scholar] [CrossRef]
- Tse, C.C.W.; Ng, S.S.; Stringer, J.; Macneil, S.; Haycock, J.W.; Smith, P.J. Utilising inkjet printed paraffin wax for cell patterning applications. Int. J. Bioprint. 2016, 2, 35–44. [Google Scholar] [CrossRef]
- Dan, K.; Prabhakaran, M.P.; Jin, G.; Ramakrishna, S. Guided orientation of cardiomyocytes on electrospun aligned nanofibers for cardiac tissue engineering. J. Biomed. Mater. Res. Part B Appl. Biomater. 2011, 98, 379–386. [Google Scholar]
- Chanthakulchan, A.; Koomsap, P.; Parkhi, A.A.; Supaphol, P. Environmental effects in fibre fabrication using electrospinning-based rapid prototyping. Virtual Phys. Prototyp. 2015, 10, 227–237. [Google Scholar] [CrossRef]
- Parajuli, D.; Koomsap, P.; Parkhi, A.A.; Supaphol, P. Experimental investigation on process parameters of near-field deposition of electrospinning-based rapid prototyping. Virtual Phys. Prototyp. 2016, 11, 1–15. [Google Scholar] [CrossRef]
- Liu, X.; Wang, X.; Li, S.; Lin, L. Energy harvesting using uniaxially aligned cardiomyocytes. In Proceedings of the 2014 IEEE 27th International Conference on Micro Electro Mechanical Systems (MEMS), San Francisco, CA, USA, 26–30 January 2014; pp. 159–162. [Google Scholar]
- Chen, Z.G.; Wang, P. Electrospun collagen-chitosan nanofiber: A biomimetic extracellular matrix for endothelial cell and smooth muscle cell. Acta Biomater. 2010, 6, 372–382. [Google Scholar] [CrossRef] [PubMed]
- Ghasemi-Mobarakeh, L.; Prabhakaran, M.P.; Morshed, M.; Nasr-Esfahani, M.H.; Ramakrishna, S. Electrical stimulation of nerve cells using conductive nanofibrous scaffolds for nerve tissue engineering. Tissue Eng. Part A 2009, 15, 3605–3619. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Cao, Y.; Pan, J.; Liu, Y. Macro-alignment of electrospun fibers for vascular tissue engineering. J. Biomed. Mater. Res. Part B Appl. Biomater. 2010, 92, 508–516. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zhao, H.; Lu, Y.; Li, S.; Lin, L.; Du, Y.; Wang, X. In vitro cardiomyocyte-driven biogenerator based on aligned piezoelectric nanofibers. Nanoscale 2016, 8, 7278–7286. [Google Scholar] [CrossRef] [PubMed]
- Grosberg, A.; Kuo, P.L.; Guo, C.L.; Geisse, N.A.; Bray, M.A.; Adams, W.J.; Sheehy, S.P.; Parker, K.K. Self-organization of muscle cell structure and function. PLoS Comput. Biol. 2011, 7, e1001088. [Google Scholar] [CrossRef] [PubMed]
- Alford, P.W.; Feinberg, A.W.; Sheehy, S.P.; Parker, K.K. Biohybrid thin films for measuring contractility in engineered cardiovascular muscle. Biomaterials 2010, 31, 3613–3621. [Google Scholar] [CrossRef] [PubMed]



© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Xu, S.; Kuang, X.; Wang, X. 3D Cardiac Cell Culture on Nanofiber Bundle Substrates for the Investigation of Cell Morphology and Contraction. Micromachines 2017, 8, 147. https://doi.org/10.3390/mi8050147
Liu X, Xu S, Kuang X, Wang X. 3D Cardiac Cell Culture on Nanofiber Bundle Substrates for the Investigation of Cell Morphology and Contraction. Micromachines. 2017; 8(5):147. https://doi.org/10.3390/mi8050147
Chicago/Turabian StyleLiu, Xia, Sixing Xu, Xuanlin Kuang, and Xiaohong Wang. 2017. "3D Cardiac Cell Culture on Nanofiber Bundle Substrates for the Investigation of Cell Morphology and Contraction" Micromachines 8, no. 5: 147. https://doi.org/10.3390/mi8050147
APA StyleLiu, X., Xu, S., Kuang, X., & Wang, X. (2017). 3D Cardiac Cell Culture on Nanofiber Bundle Substrates for the Investigation of Cell Morphology and Contraction. Micromachines, 8(5), 147. https://doi.org/10.3390/mi8050147




