Oocytes Polar Body Detection for Automatic Enucleation
Abstract
:1. Introduction

2. Learning and Prediction-Based Polar Body Detection Method
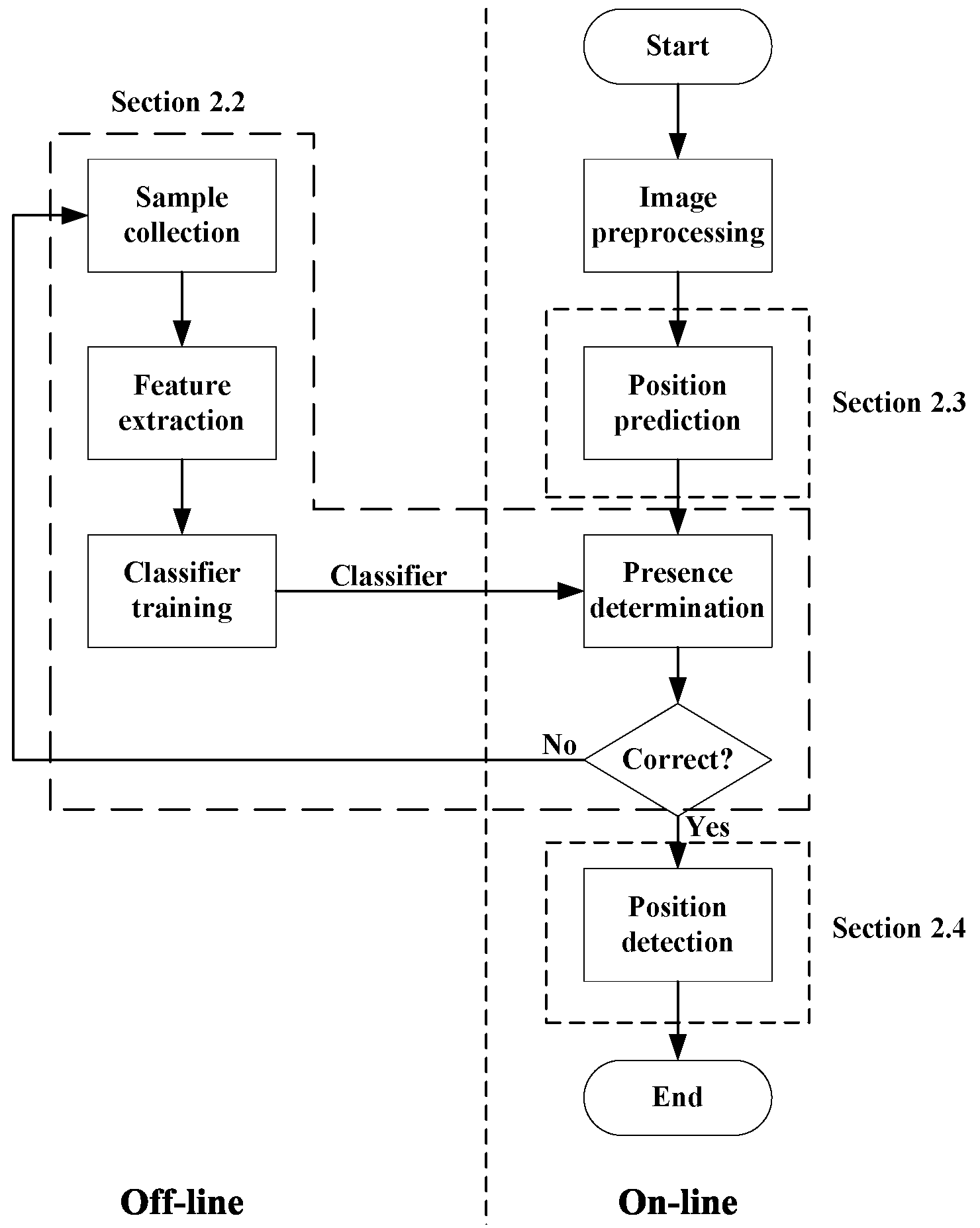
2.1. Polar Body Detection Process
- Obtain the image region of the oocyte as the region of interest (ROI). Divide the image region into small patches, and get the image patches that may contain the polar body by position prediction. We will analyze the method of position prediction in Section 2.3.
- Use an SVM classifier to determine whether a polar body is present in these image patches. The detected image patches will be added into the sample library and the classifier will be trained again.
- Detect the position of the polar body in the image patch, which is described in Section 2.4.
2.2. Learning-Based Polar Body Presence Determination
2.2.1. Improved HOG Algorithm

2.2.2. Polar Body Presence Determination Based on SVM Algorithm
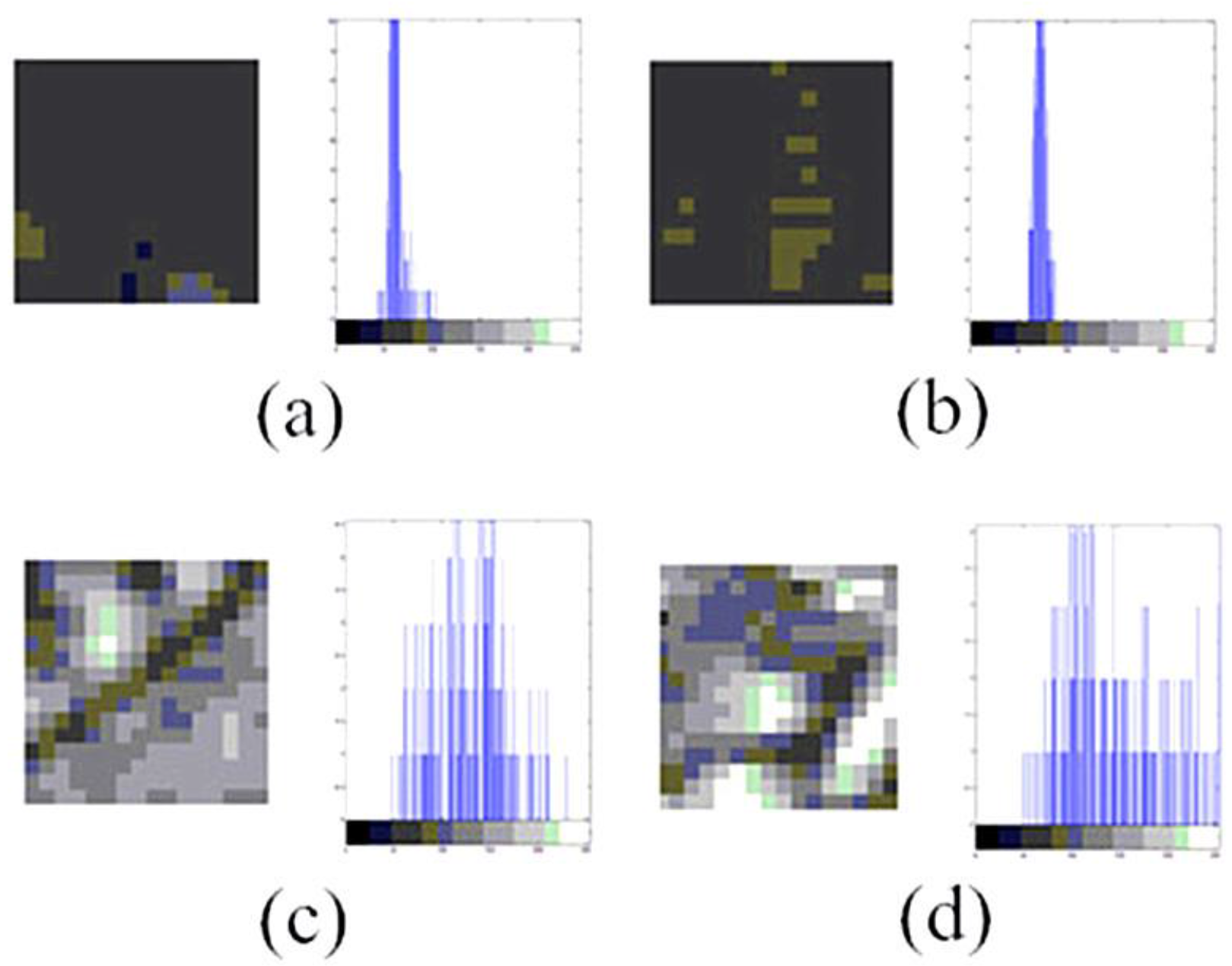
- Extract features of polar body and non-polar body image patches (positive and negative samples) using improved HOG algorithm.
- Reduce the dimension of features using PCA algorithm [19].
- Train the polar body classifier using SVM algorithm.
- Extract features of the detected image patch and determine whether a polar body is present in it using the polar body classifier.
2.3. Polar Body Position Prediction
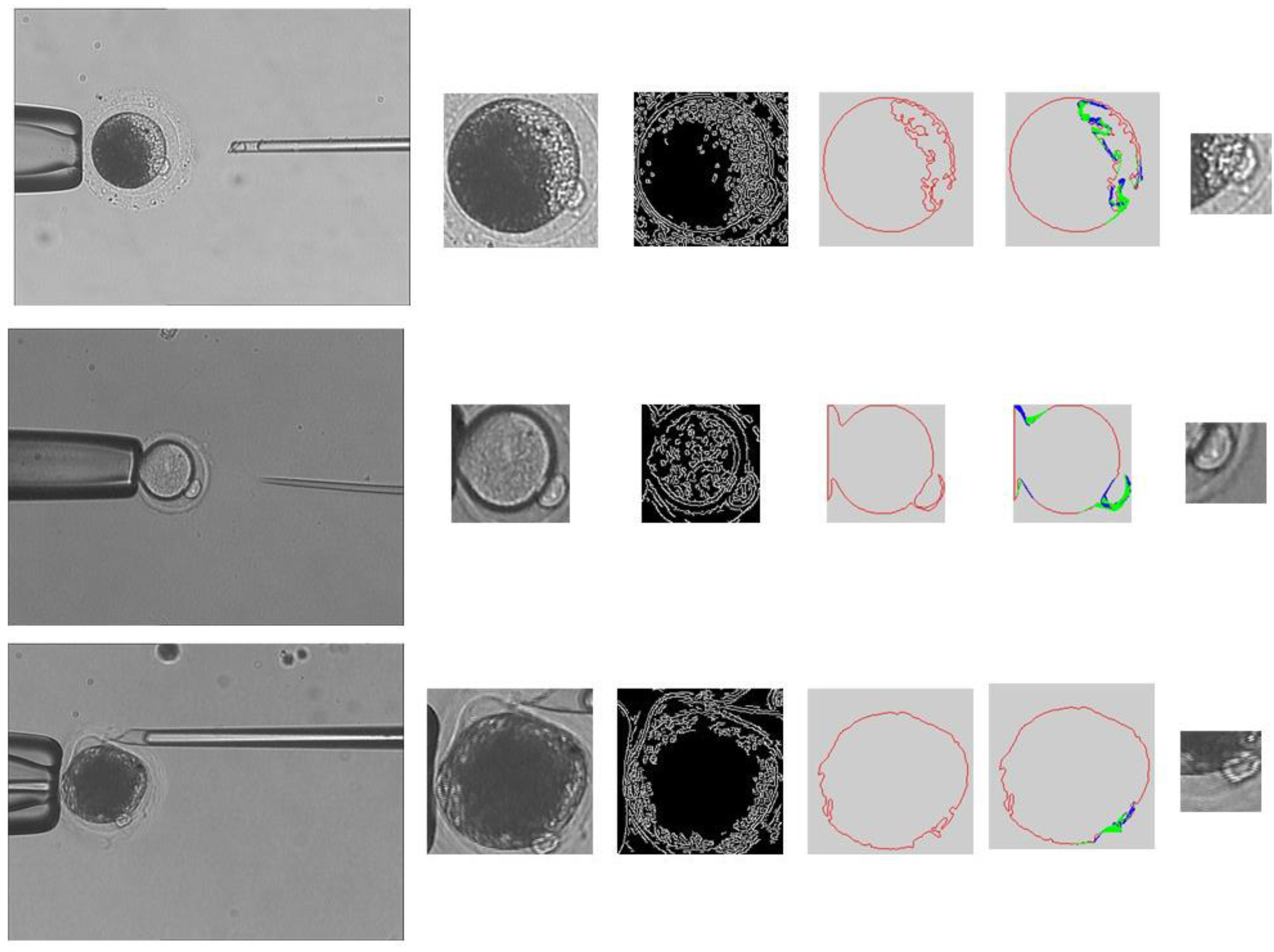
- Obtain cytoplasmic membrane contour. The polar body is located between the cytoplasm and the zona pellucida. We use the cytoplasmic membrane contour to determine the candidate positions of the polar body. The main steps in obtaining the cytoplasmic membrane contour are listed as follows:
- (1)
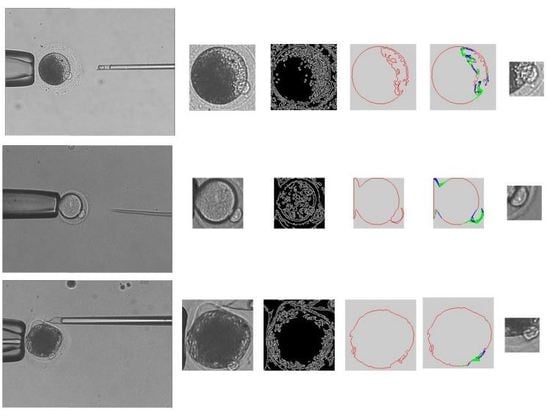
- The de-noised image is processed by using intensity transformation, so that the zona pellucida is eliminated in the image. The region of cytoplasmic membrane is detected by using binarization.
- (2)
- The region of cytoplasm is further detected by using Hoff algorithm [22].
- (3)
- The edge information is acquired by using Canny algorithm [23].
- (4)
- The cytoplasmic membrane contour is detected by using the active contour method.
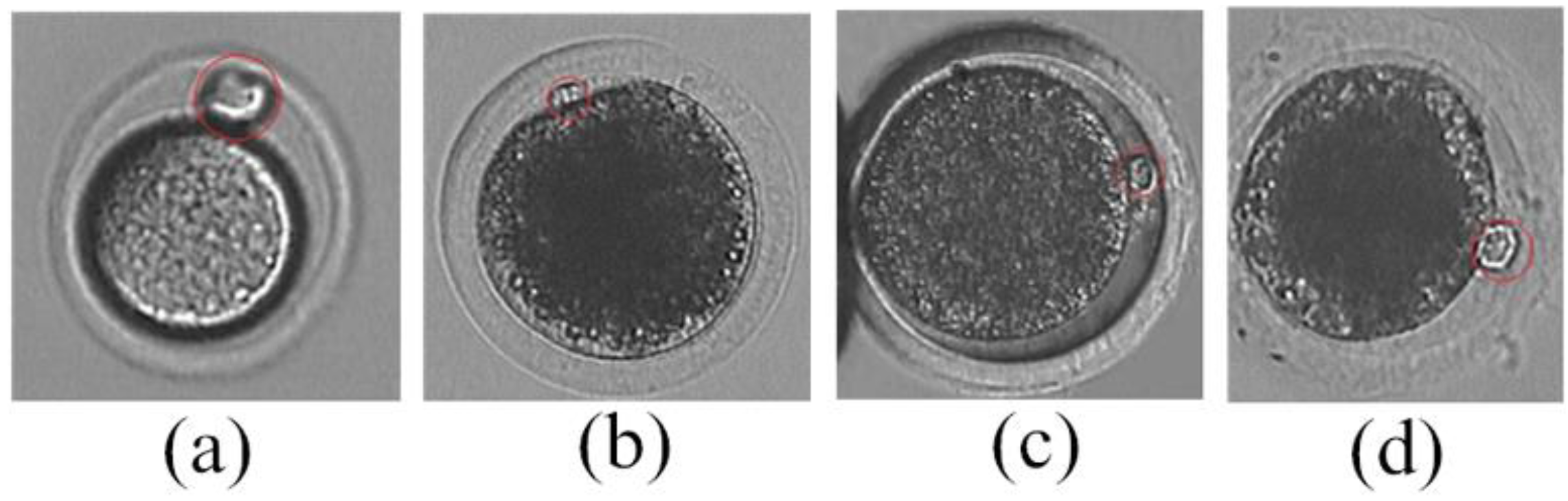
- Analyze the curvature of cytoplasm membrane contour to narrow the detection range of polar body. As shown in Figure 1, the curvature of cytoplasm membrane contour in polar body position is obviously different with the non-polar body position. The main steps are listed as follows:
- (1)
- For each polar body candidate position, we select two neighbor points in the contour with same interval. These three contour points compose a triangle. Figure 5 shows the schematic diagram, where Point i represents the candidate position, Point pre and Point next are the neighbor points.
- (2)
- We calculate the cosine value of angle A instead of the curvature.
- (3)
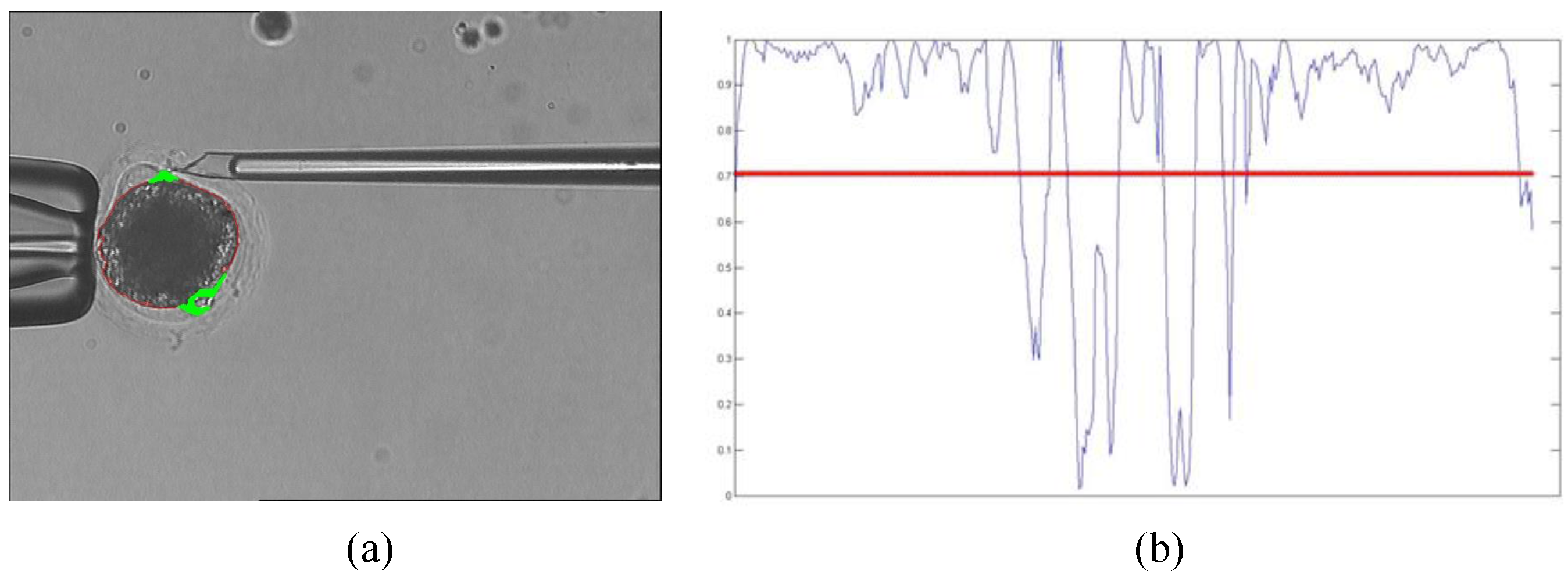
- The cosine value is smaller if Point i is in the position of the polar body; We set a threshold to distinguish the position of polar body and non-polar body. The threshold is obtained experimentally.

2.4. Polar Body Position Detection
- The polar body is detected by using SVM classifier in the image patch.
- The contour of the polar body is detected by using image binarization.
- The polar body position is obtained by calculating the center of the polar body contour in the binarization result.
3. Experiments
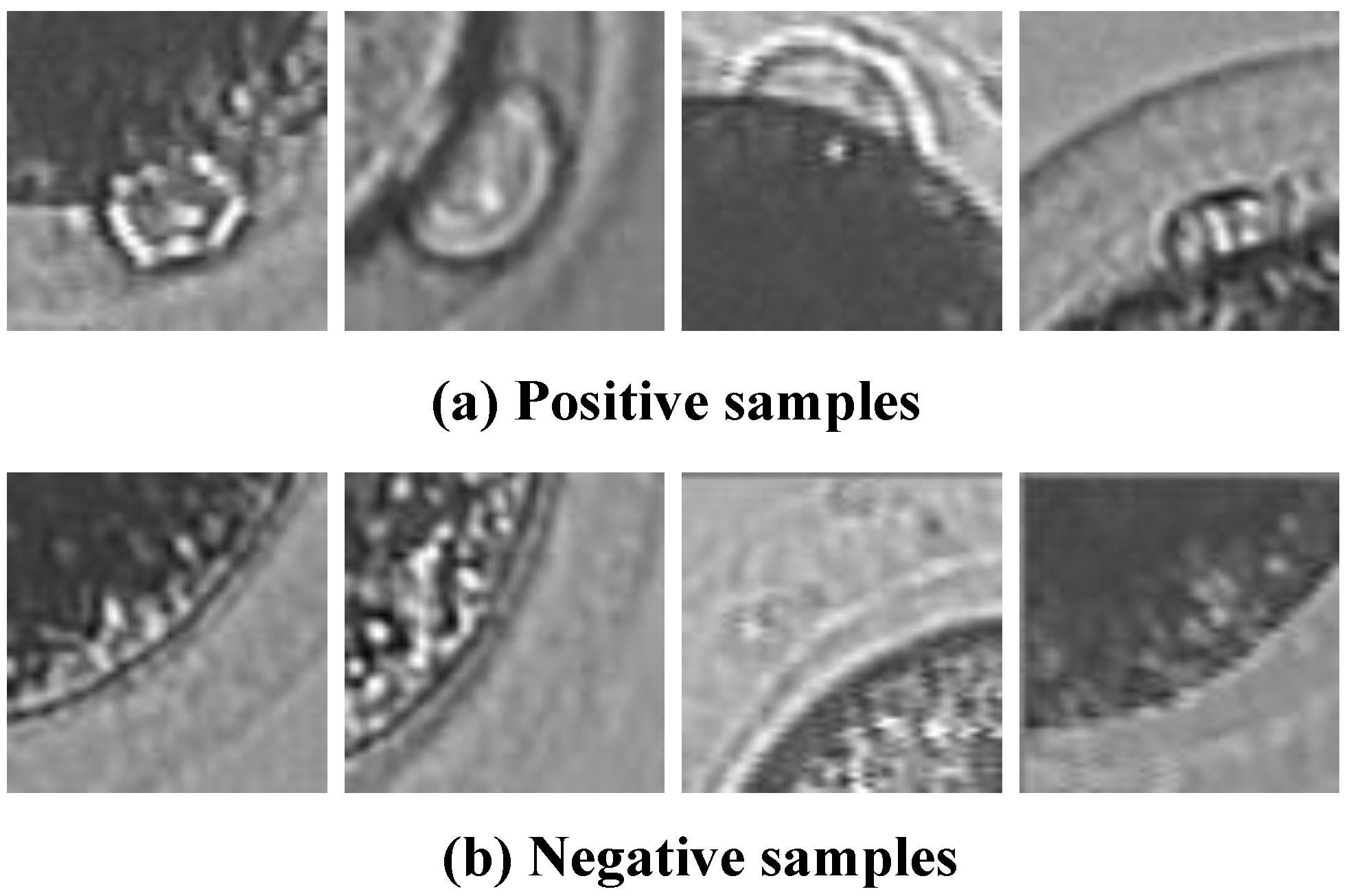
3.1. Sample Library Establishment
- We should consider detection efficiency when choosing samples.
- There should be obvious differences between positive samples and negative samples.
- There should be various types of polar bodies in positive samples.

3.2. HOG Feature Parameters Selection
- Block size.We change block size with the same feature dimension. The test results are shown in Table 1, which indicate that the block size has little effect on detection results.
| Block Size | Correct Number | Wrong Number | Success Rate |
|---|---|---|---|
| 32 × 32 | 67 | 5 | 93% |
| 24 × 24 | 67 | 5 | 93% |
| 16 × 16 | 67 | 5 | 93% |
- 2.
- Cell size in a block.We change cell size in a block with the same feature dimension. The test results are shown in Table 2, which indicate that the success rate increases with large cell size in a block.
| Cell Number | Correct Number | Wrong Number | Success Rate |
|---|---|---|---|
| 4 | 67 | 5 | 93% |
| 9 | 69 | 3 | 95.8% |
- 3.
- Step length and the size of overlapping.
| Block Step Length | Correct Number | Wrong Number | Success Rate |
|---|---|---|---|
| 20 | 58 | 14 | 81% |
| 10 | 61 | 11 | 85% |
| 8 | 62 | 10 | 86% |
3.3. PCA Feature Parameters Selection
| Dimension | Correct Number | Wrong Number | Success Rate | Time (ms/Frame) |
|---|---|---|---|---|
| 10 | 47 | 25 | 34.7% | 115 |
| 20 | 61 | 11 | 84.7% | 123 |
| 30 | 69 | 3 | 95.8% | 130 |
| 40 | 69 | 3 | 95.8% | 147 |
| 2025 | 70 | 2 | 97.2% | 605 |
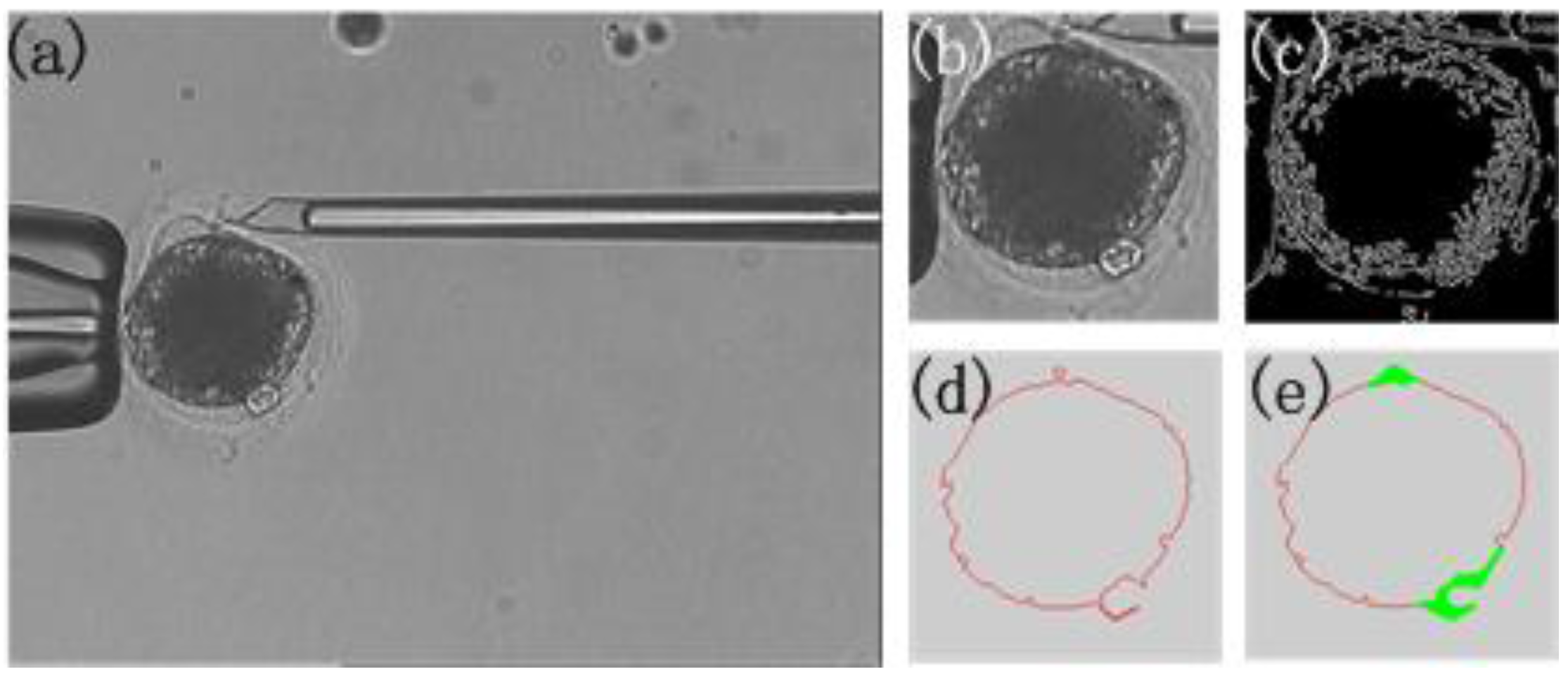
3.4. Polar Body Position Prediction Results

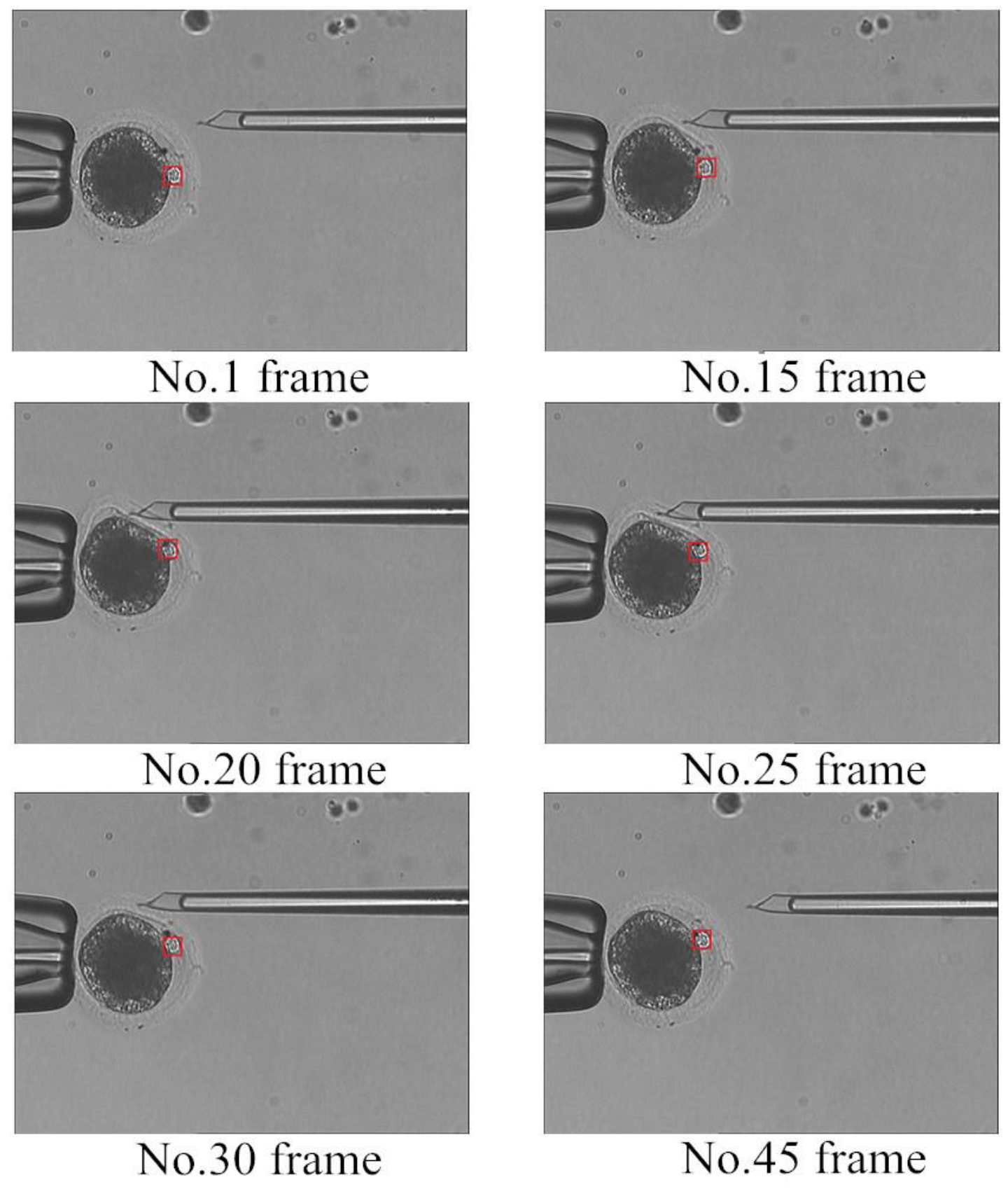
3.5. Polar Body Detection Results
| TP | TN | FP | FN | Success Rate | |
|---|---|---|---|---|---|
| Number | 2277 | 6131 | 107 | 107 | 96.4% |
4. Application of Polar Body Detection Method in Automated Enucleation
- (1)
- Hold the oocyte using a holder micropipette.
- (2)
- Rotate the oocyte using an injection micropipette until the polar body is in the proper position.
- (3)
- Localize the polar body and remove the nucleus and polar body using an injection micropipette.
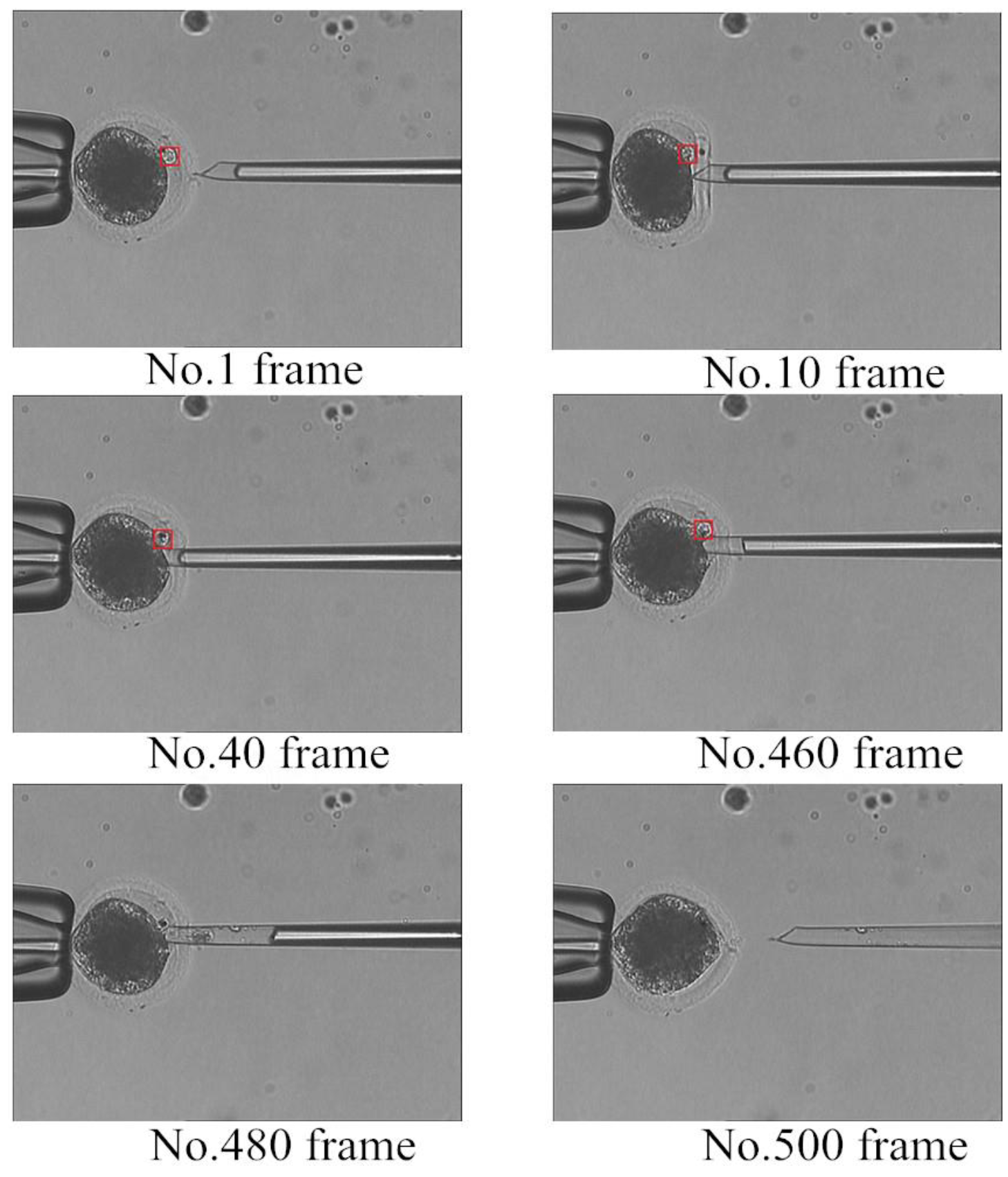


5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Onishi, A.; Iwamoto, M.; Akita, T.; Takashi, K.; Awata, T.; Hanada, H.; Perry, A.C. Pig Cloning by Microinjection of Fetal Fibroblast Nuclei. Science 2000, 289, 1188–1190. [Google Scholar] [CrossRef] [PubMed]
- Wakayama, T.; Perry, A.C.; Zuccotti, M.; Johnson, K.R.; Yannagimachi, R. Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 1998, 394, 369–374. [Google Scholar] [PubMed]
- Sun, Y.; Nelson, B.J. Biological cell injection using an autonomous microrobotic system. Int. J. Robot. Res. 2002, 21, 861–868. [Google Scholar] [CrossRef]
- Mattos, L.; Grant, E.; Thresher, R.; Kluckman, K. From teleoperated to automatic blastocyst microinjections: Designing a new system from expert-controlled operations. In Proceedings of IEEE RSJ International Conference on Intelligent Robots and Systems, Piscataway, NJ, USA, 18–20 July 2008.
- Gitlin, S.A.; Gibbons, W.E.; Gosden, R.G. Oocyte biology and genetics revelations from polar bodies. Reprod. BioMed. Online 2003, 6, 403–409. [Google Scholar] [CrossRef]
- Schmerler, S.; Wessel, G.M. Polar bodies-more a lack of understanding than a lack of respect. Mol. Reprod. Dev. 2011, 78, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Gianaroli, L. Preimplantation genetic diagnosis: Polar body and embryo biopsy. Hum. Reprod. 2002, 15, 941–955. [Google Scholar] [CrossRef]
- Wilmut, I.; Schnieke, A.E.; McWhir, J.; Kind, A.J.; Campbell, K.H. Viable offspring derived from fetal and adult mammalian cells. Nature 2007, 385, 810–813. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.H.; Liu, X.Y.; Sun, Y. High-throughput automated injection of individual biological cells. IEEE Trans. Autom. Sci. Eng. 2009, 6, 209–219. [Google Scholar] [CrossRef]
- Lu, Z.; Zhang, X.; Leung, C.; Esfandiari, N.; Casper, R.F.; Sun, Y. Robotic ICSI (intracytoplasmic sperm injection). IEEE Trans. Biomed. Eng. 2011, 58, 209–219. [Google Scholar]
- Leung, C.; Lu, Z.; Esfandiari, N.; Casper, R.F.; Sun, Y. Automated sperm immobilization for intracytoplasmic sperm injection. IEEE Trans. Biomed. Eng.. 2011, 58, 935–942. [Google Scholar]
- Vajta, G.; Lewsis, I.M.; Hyttel, P.; Thouas, G.A.; Trounson, A. Somatic cell cloning without micromanipulators. Cloning 2001, 3, 89–95. [Google Scholar]
- Ichikawa, A.; Honda, A.; Ejima, M.; Tanikawa, T.; Arai, F.; Fukuda, T. In situ formation of a gel microbead for laser micromanipulation of microorganisms. In Proceedings of International Symposium on Micro-NanoMechatronics and Human Science, Nagoya, Japan, 6–8 November 2006.
- Ichikawa, A.; Takahashi, S.; Matsukawa, K.; Tanikawa, T.; Ohba, K. Injection and cutting methods of animal cells using a microfuidic chip. In Proceedings of IEEE International Conference on Robotics and Automation, Pasadena, CA, USA, 19–23 May 2008.
- Tatham, B.G.; Dowsing, A.T.; Trounson, A.O. Enucleation by centrifugation of in vitro-matured bovine oocytes for use in nuclear transfer. Biol. Reprod. 1995, 53, 1088–1094. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.L.; Sun, M.Z.; Cui, M.S.; Yu, J.; Qin, Y.D.; Zhao, X. Robotic Cell Rotation Based on the Minimum Rotation Force. IEEE Trans. Autom. Sci. Eng. 2015, 12, 1504–1515. [Google Scholar] [CrossRef]
- Leung, C.; Lu, Z.; Zhang, X.; Sun, Y. Three-dimensional rotation of mouse embryos. IEEE Trans. Biomed. Eng. 2012, 59, 1049–1056. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.L.; Zhao, X.; Zhao, Q.L.; Lu, G.Z. Illumination intensity evaluation of microscopic image based on texture information and application on locating polar body in oocytes. In Proceedings of China Automation Conference, Beijing, China, 7–10 August 2011.
- Wold, S.; Esbensen, K.; Geladi, P. Principal Component Analysis. Chemometr. Intell. Lab. Syst. 1987, 2, 37–52. [Google Scholar] [CrossRef]
- Chen, Y.; Zhou, X.S.; Huang, T.S. One-class SVM for learning in image retrieval. In Proceedings of 2001 International Conference on Image Processing, Thessaloniki, Greece, 7–10 October 2001; pp. 34–37.
- Dalal, N.; Triggs, B. Histograms of oriented gradients for human detection. In Proceedings of IEEE Computer Society Conference on Computer Vision and Pattern Recognition, San Diego, CA, USA, 20–26 June 2005.
- Yuen, H.K.; Princen, J.; lllingworth, J.; Kittler, J. Comparative study of Hough Transform methods for circle finding. Image Vis. Comput. 1990, 8, 71–77. [Google Scholar] [CrossRef]
- Canny, J. A computational approach to edge detection. IEEE Trans. Pattern Anal. Mach. Intell. 1986, 8, 679–698. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, D.; Sun, M.; Zhao, X. Oocytes Polar Body Detection for Automatic Enucleation. Micromachines 2016, 7, 27. https://doi.org/10.3390/mi7020027
Chen D, Sun M, Zhao X. Oocytes Polar Body Detection for Automatic Enucleation. Micromachines. 2016; 7(2):27. https://doi.org/10.3390/mi7020027
Chicago/Turabian StyleChen, Di, Mingzhu Sun, and Xin Zhao. 2016. "Oocytes Polar Body Detection for Automatic Enucleation" Micromachines 7, no. 2: 27. https://doi.org/10.3390/mi7020027
APA StyleChen, D., Sun, M., & Zhao, X. (2016). Oocytes Polar Body Detection for Automatic Enucleation. Micromachines, 7(2), 27. https://doi.org/10.3390/mi7020027