Electrochemical Biosensing for Antibiotic-Resistant Bacteria: Advances, Challenges, and Future Directions
Abstract
1. Introduction
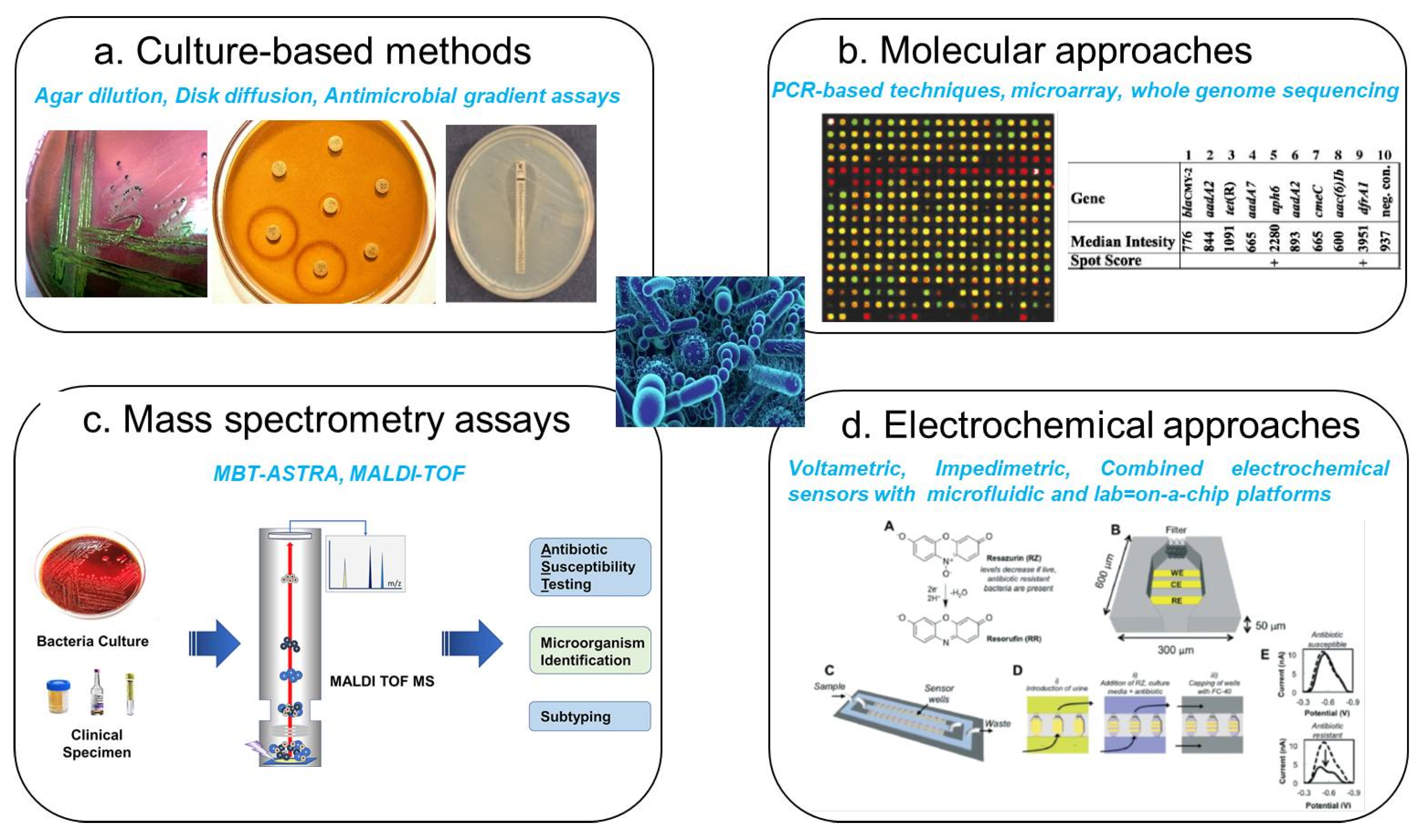
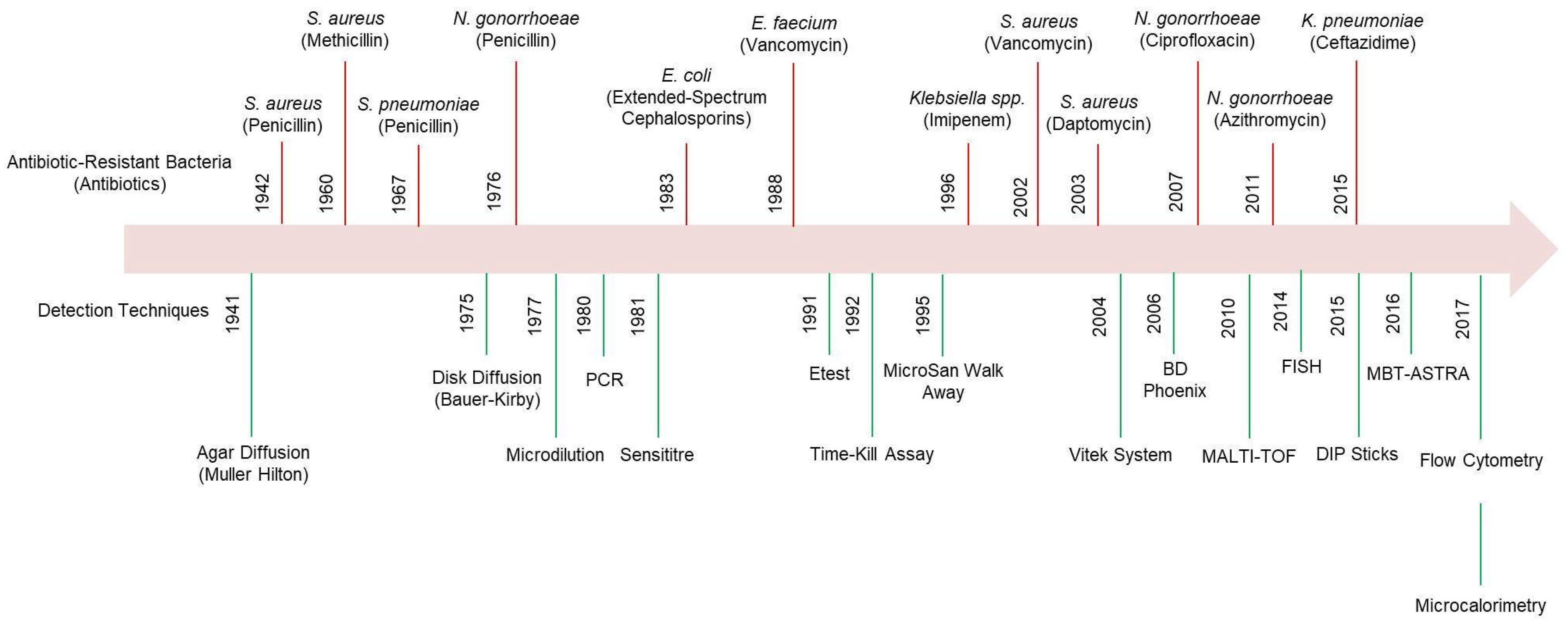
2. Clinical and Economic Significance of Detecting Antibiotic-Resistant Bacteria
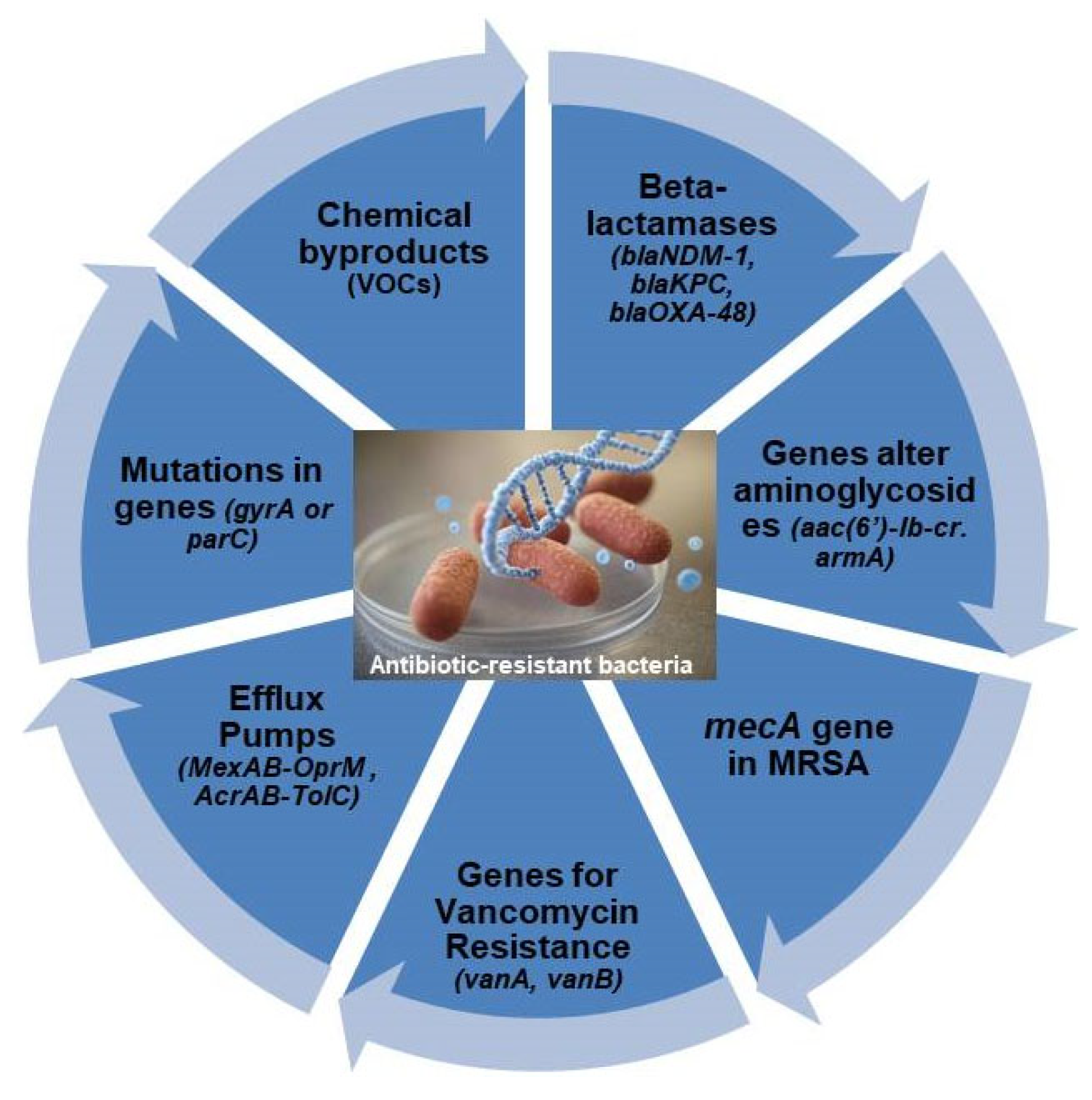
3. Common Biomarkers for Antibiotic-Resistant Bacteria Detection
4. Electrochemical Detection Techniques of Antibiotic-Resistant Bacteria
4.1. Electrochemical Detection of Antibiotic-Resistant Bacteria Based on Bio-Receptors
4.1.1. DNA-Based Sensors
4.1.2. mRNA-Based Sensors
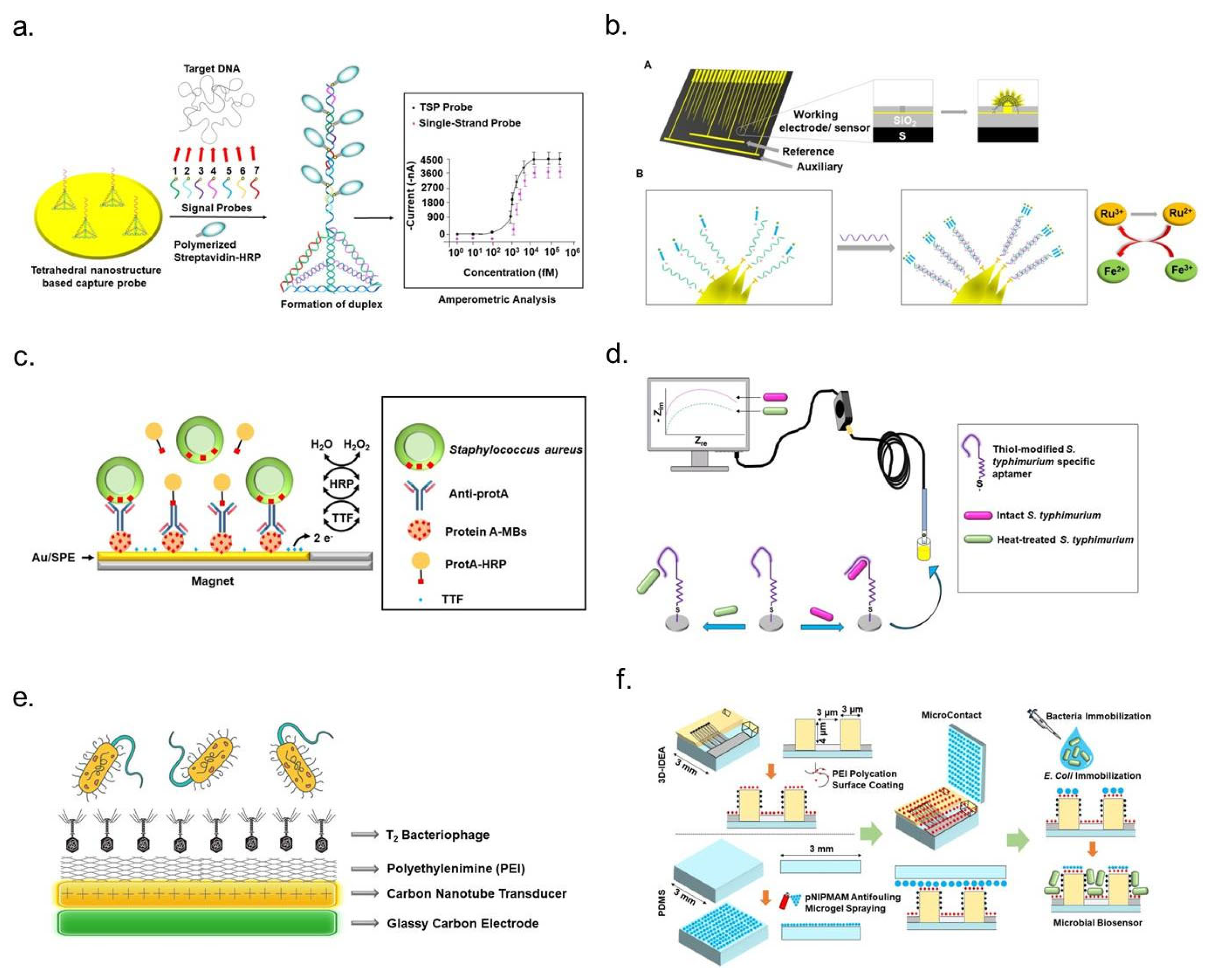
4.1.3. Antibody- and Antibody Fragment-Based Sensors
4.1.4. Aptamer-Based Sensors
4.1.5. Bacteriophage-Based Sensors
4.1.6. Cell- and Molecularly Imprinted Polymer-Based Sensors
4.1.7. Carbohydrate-Based Sensors
4.2. Electrochemical Detection of Antibiotic-Resistant Bacteria Based on Electrode Materials and Modified Electrode Surfaces
4.2.1. Carbon-Based Sensors (Carbon Nanotubes and Graphene)
4.2.2. Paper-Based Sensors
4.2.3. Nanoparticle-Based Sensors
4.3. Electrochemical Detection of Antibiotic-Resistant Bacteria Based on Redox-Active Cellular Metabolites
4.4. Electrochemical Detection of Antibiotic-Resistant Bacteria Based on Microbial Enzymes
4.5. Electrochemical Detection of Antibiotic-Resistant Bacteria Based on Combined Electrochemical Sensing Techniques
4.5.1. Microfluidic Based
4.5.2. Lab-on-a-Chip Based
5. Biological Challenges in Electrochemical Detection of Antibiotic-Resistant Bacteria
5.1. Influence of Sample Matrix on Assay Reliability
5.2. Challenges Posed by Polymicrobial Infections
5.3. Impact of Inoculum Size on Antibiotic Susceptibility Testing
6. Technical Challenges in Electrochemical Detection of Antibiotic-Resistant Bacteria
6.1. Growth Media Compatibility and Nutritional Stability
6.2. Limitations in Molecular Biology Approaches
6.3. Challenges of Non-Specific Biomolecule Interactions
6.4. Sensitivity Constraints in Miniaturized Platforms
7. Potential Strategies and Solutions for Addressing Challenges in Electrochemical Detection of Antibiotic-Resistant Bacteria
7.1. Improving Sample Matrix Compatibility
7.2. Addressing Polymicrobial Infections
7.3. Mitigating the Inoculum Effect
7.4. Enhancing Growth Media Performance
7.5. Use of Functional Materials
7.6. Leveraging Molecular Biology Tools
7.7. Minimising Non-Specific Responses
7.8. Maximizing Sensitivity in Miniaturized Systems
7.9. Overcoming Logistical and Cost Constraints
8. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Serwecińska, L. Antimicrobials and Antibiotic-Resistant Bacteria: A Risk to the Environment and to Public Health. Water 2020, 12, 3313. [Google Scholar] [CrossRef]
- Blair, J.M.A.; Webber, M.A.; Baylay, A.J.; Ogbolu, D.O.; Piddock, L.J.V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Larsson, D.G.J.; Andremont, A.; Bengtsson-Palme, J.; Brandt, K.K.; de Roda Husman, A.M.; Fagerstedt, P.; Fick, J.; Flach, C.F.; Gaze, W.H.; Kuroda, M.; et al. Critical knowledge gaps and research needs related to the environmental dimensions of antibiotic resistance. Environ. Int. 2018, 117, 132–138. [Google Scholar] [CrossRef]
- Martinez, J.L. Ecology and Evolution of Chromosomal Gene Transfer between Environmental Microorganisms and Pathogens. Microbiol. Spectr. 2018, 6. [Google Scholar] [CrossRef]
- Gulen, T.A.; Guner, R.; Celikbilek, N.; Keske, S.; Tasyaran, M. Clinical importance and cost of bacteremia caused by nosocomial multi drug resistant acinetobacter baumannii. Int. J. Infect. Dis. 2015, 38, 32–35. [Google Scholar] [CrossRef]
- Founou, R.C.; Founou, L.L.; Essack, S.Y. Clinical and economic impact of antibiotic resistance in developing countries: A systematic review and meta-analysis. PLoS ONE 2017, 12, e0189621. [Google Scholar] [CrossRef]
- Ehrlich, P. Chemotherapeutische Trypanosomen-Studien. Berliner Munch. Tierarztl. Wochenschr. 1907, 11, 233–238. [Google Scholar]
- Barber, M. Staphylococcal infection due to penicillin-resistant strains. Br. Med. J. 1947, 2, 863–865. [Google Scholar] [CrossRef]
- Syal, K.; Mo, M.; Yu, H.; Iriya, R.; Jing, W.; Guodong, S.; Wang, S.; Grys, T.E.; Haydel, S.E.; Tao, N. Current and emerging techniques for antibiotic susceptibility tests. Theranostics 2017, 7, 1795–1805. [Google Scholar] [CrossRef] [PubMed]
- Anjum, M.F. Screening methods for the detection of antimicrobial resistance genes present in bacterial isolates and the microbiota. Future Microbiol. 2015, 10, 317–320. [Google Scholar] [CrossRef]
- Váradi, L.; Luo, J.L.; Hibbs, D.E.; Perry, J.D.; Anderson, R.J.; Orenga, S.; Groundwater, P.W. Methods for the detection and identification of pathogenic bacteria: Past, present, and future. Chem. Soc. Rev. 2017, 46, 4818–4832. [Google Scholar] [CrossRef]
- Goluch, E.D. Microbial Identification Using Electrochemical Detection of Metabolites. Trends Biotechnol. 2017, 35, 1125–1128. [Google Scholar] [CrossRef]
- Behzadi, P. DNA Microarrays and Multidrug Resistant Bacteria. Eur. Pharm. Rev. 2018, 1, 30–32. [Google Scholar]
- Frye, J.G.; Jesse, T.; Long, F.; Rondeau, G.; Porwollik, S.; McClelland, M.; Jackson, C.R.; Englen, M.; Fedorka-Cray, P.J. DNA microarray detection of antimicrobial resistance genes in diverse bacteria. Int. J. Antimicrob. Agents 2006, 27, 138–151. [Google Scholar] [CrossRef]
- Pulido, M.R.; García-Quintanilla, M.; Martín-Peña, R.; Cisneros, J.M.; McConnell, M.J. Progress on the development of rapid methods for antimicrobial susceptibility testing. J. Antimicrob. Chemother. 2013, 68, 2710–2717. [Google Scholar] [CrossRef]
- Hou, T.Y.; Chiang-Ni, C.; Teng, S.H. Current status of MALDI-TOF mass spectrometry in clinical microbiology. J. Food Drug Anal. 2019, 27, 404–414. [Google Scholar] [CrossRef] [PubMed]
- Doern, C.D.; Butler-Wu, S.M. Emerging and Future Applications of Matrix-Assisted Laser Desorption Ionization Time-of-Flight (MALDI-TOF) Mass Spectrometry in the Clinical Microbiology Laboratory: A Report of the Association for Molecular Pathology. J. Mol. Diagn. 2016, 18, 789–802. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Zhu, J. Differentiating Antibiotic-Resistant Staphylococcus aureus Using Secondary Electrospray Ionization Tandem Mass Spectrometry. Anal. Chem. 2018, 90, 12108–12115. [Google Scholar] [CrossRef] [PubMed]
- Amiri, M.; Bezaatpour, A.; Jafari, H.; Boukherroub, R.; Szunerits, S. Electrochemical Methodologies for the Detection of Pathogens. ACS Sens. 2018, 3, 1069–1086. [Google Scholar] [CrossRef]
- Simoska, O.; Stevenson, K.J. Electrochemical sensors for rapid diagnosis of pathogens in real time. Analyst 2019, 144, 6461–6478. [Google Scholar] [CrossRef]
- Besant, J.D.; Sargent, E.H.; Kelley, S.O. Rapid electrochemical phenotypic profiling of antibiotic-resistant bacteria. Lab Chip 2015, 15, 2799–2807. [Google Scholar] [CrossRef]
- Kelley, S.O. New Technologies for Rapid Bacterial Identification and Antibiotic Resistance Profiling. SLAS Technol. 2017, 22, 113–121. [Google Scholar] [CrossRef]
- Kuss, S.; Amin, H.M.A.; Compton, R.G. Electrochemical Detection of Pathogenic Bacteria-Recent Strategies, Advances and Challenges. Chem. Asian J. 2018, 13, 2758–2769. [Google Scholar] [CrossRef] [PubMed]
- Cesewski, E.; Johnson, B.N. Electrochemical biosensors for pathogen detection. Biosens. Bioelectron. 2020, 159, 112214. [Google Scholar] [CrossRef]
- Munteanu, F.D.; Titoiu, A.M.; Marty, J.L.; Vasilescu, A. Detection of Antibiotics and Evaluation of Antibacterial Activity with Screen-Printed Electrodes. Sensors 2018, 18, 901. [Google Scholar] [CrossRef]
- Saucedo, N.M.; Srinives, S.; Mulchandani, A. Electrochemical Biosensor for Rapid Detection of Viable Bacteria and Antibiotic Screening. J. Anal. Test. 2019, 3, 117–122. [Google Scholar] [CrossRef]
- Monzó, J.; Insua, I.; Fernandez-Trillo, F.; Rodriguez, P. Fundamentals, achievements and challenges in the electrochemical sensing of pathogens. Analyst 2015, 140, 7116–7128. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Yoo, S. Electrochemical Sensors for Antibiotic Susceptibility Testing: Strategies and Applications. Chemosensors 2022, 10, 53. [Google Scholar] [CrossRef]
- Madhu, S.; Ramasamy, S.; Choi, J. Recent Developments in Electrochemical Sensors for the Detection of Antibiotic-Resistant Bacteria. Pharmaceuticals 2022, 15, 1488. [Google Scholar] [CrossRef] [PubMed]
- Manasa, G.; Rout, C.S. Versatile MXenes as electrochemical sensors for heavy metal ions and phenolic moiety-containing industrial chemicals: Recent development and prospects. Mater. Adv. 2024, 5, 83–122. [Google Scholar] [CrossRef]
- Zheng, A.L.T.; Teo, E.Y.L.; Yiu, P.H.; Boonyuen, S.; Chung, E.L.T.; Andou, Y. Recent advances in ultrasensitive electrochemical sensors and biosensors for determination of antibiotics in environment samples. Clean Technol. Environ. Policy 2025, 27, 549–576. [Google Scholar] [CrossRef]
- Du, Z.; Yang, R.; Guo, Z.; Song, Y.; Wang, J. Identification of Staphylococcus aureus and determination of its methicillin resistance by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Anal. Chem. 2002, 74, 5487–5491. [Google Scholar] [CrossRef]
- World Health Organization. Global Priority List of Antibiotic-Resistance Bacteria to Guide Research, Discovery, and Development of New Antibiotics; World Health Organization: Geneva, Switzerland, 2017. [Google Scholar]
- CDC. 2019 Antibiotic Resistance Threats Report. 2024. Available online: https://www.cdc.gov/antimicrobial-resistance/data-research/threats/index.html (accessed on 25 August 2025).
- Ripa, M.; Galli, L.; Poli, A.; Oltolini, C.; Spagnuolo, V.; Mastrangelo, A.; Muccini, C.; Monti, G.; De Luca, G.; Landoni, G.; et al. Secondary infections in patients hospitalized with COVID-19: Incidence and predictive factors. Clin. Microbiol. Infect. 2021, 27, 451–457. [Google Scholar] [CrossRef]
- Blot, S.; Ruppé, E.; Harbarth, S.; Asehnoune, K.; Poulakou, G.; Luyt, C.E.; Rello, J.; Klompas, M.; Depuydt, P.; Eckmann, C.; et al. Healthcare-associated infections in adult intensive care unit patients: Changes in epidemiology, diagnosis, prevention and contributions of new technologies. Intensive Crit. Care Nurs. 2022, 70, 103227. [Google Scholar] [CrossRef] [PubMed]
- Desai, V.; Kumar, S.; Patel, B.; Patel, S.N.; Patadiya, H.H.; Asawa, D.; Pathan, M.S.H.; Haque, M. Navigating Antimicrobials and Combating Antimicrobial Resistance: Challenges, Impacts, and Strategies for Global Action. Cureus 2025, 17, e82064. [Google Scholar] [CrossRef]
- O’Neill, J. Tackling drug-resistant infections globally: Final report and recommendations. In Review on Antimicrobial Resistance; HM Government: London, UK; Wellcome Trust: London, UK, 2016; Available online: https://amr-review.org/sites/default/files/160518_Final%20paper_with%20cover.pdf (accessed on 12 January 2019).
- Sonpar, A.; Hundal, C.O.; Totté, J.E.E.; Wang, J.; Klein, S.D.; Twyman, A.; Allegranzi, B.; Zingg, W. Multimodal strategies for the implementation of infection prevention and control interventions—Update of a systematic review for the WHO guidelines on core components of infection prevention and control programmes at the facility level. Clin. Microbiol. Infect. 2025, 31, 948–957. [Google Scholar] [CrossRef] [PubMed]
- Commission, E. The New EU One Health Action Plan against Antimicrobial Resistance. 2022. Available online: https://health.ec.europa.eu/system/files/2020-01/amr_2017_summary-action-plan_0.pdf (accessed on 25 August 2025).
- CDC. Antimicrobial Resistance Facts and Stats. 2025. Available online: https://www.cdc.gov/antimicrobial-resistance/data-research/facts-stats/index.html (accessed on 25 August 2025).
- Nordmann, P.; Naas, T.; Poirel, L. Global spread of Carbapenemase-producing Enterobacteriaceae. Emerg. Infect. Dis. 2011, 17, 1791–1798. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, M.S.; Tolmasky, M.E. Aminoglycoside modifying enzymes. Drug Resist. Updates 2010, 13, 151–171. [Google Scholar] [CrossRef]
- Chambers, H.F.; Deleo, F.R. Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat. Rev. Microbiol. 2009, 7, 629–641. [Google Scholar] [CrossRef]
- Cetinkaya, Y.; Falk, P.; Mayhall, C.G. Vancomycin-resistant enterococci. Clin. Microbiol. Rev. 2000, 13, 686–707. [Google Scholar] [CrossRef]
- Li, X.Z.; Ma, D.; Livermore, D.M.; Nikaido, H. Role of efflux pump(s) in intrinsic resistance of Pseudomonas aeruginosa: Active efflux as a contributing factor to beta-lactam resistance. Antimicrob. Agents Chemother. 1994, 38, 1742–1752. [Google Scholar] [CrossRef] [PubMed]
- Hooper, D.C.; Jacoby, G.A. Mechanisms of drug resistance: Quinolone resistance. Ann. N. Y. Acad. Sci. 2015, 1354, 12–31. [Google Scholar] [CrossRef]
- Bos, L.D.; Sterk, P.J.; Schultz, M.J. Volatile metabolites of pathogens: A systematic review. PLoS Pathog. 2013, 9, e1003311. [Google Scholar] [CrossRef]
- Drummond, T.G.; Hill, M.G.; Barton, J.K. Electrochemical DNA sensors. Nat. Biotechnol. 2003, 21, 1192–1199. [Google Scholar] [CrossRef]
- Pellitero, M.A.; Shaver, A.; Arroyo-Currás, N. Critical Review—Approaches for the Electrochemical Interrogation of DNA-Based Sensors: A Critical Review. J. Electrochem. Soc. 2020, 167, 037529. [Google Scholar] [CrossRef]
- Xu, B.; Zheng, D.; Qiu, W.; Gao, F.; Jiang, S.; Wang, Q. An ultrasensitive DNA biosensor based on covalent immobilization of probe DNA on fern leaf-like α-Fe2O3 and chitosan Hybrid film using terephthalaldehyde as arm-linker. Biosens. Bioelectron. 2015, 72, 175–181. [Google Scholar] [CrossRef]
- Ulianas, A.; Heng, L.Y.; Abu Hanifah, S.; Ling, T.L. An electrochemical DNA microbiosensor based on succinimide-modified acrylic microspheres. Sensors 2012, 12, 5445–5460. [Google Scholar] [CrossRef]
- Pöhlmann, C.; Sprinzl, M. Electrochemical Detection of RNA. In RNA and DNA Diagnostics; Erdmann, V.A., Jurga, S., Barciszewski, J., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 21–45. [Google Scholar]
- Soleymani, L.; Fang, Z.; Lam, B.; Bin, X.; Vasilyeva, E.; Ross, A.J.; Sargent, E.H.; Kelley, S.O. Hierarchical nanotextured microelectrodes overcome the molecular transport barrier to achieve rapid, direct bacterial detection. ACS Nano 2011, 5, 3360–3366. [Google Scholar] [CrossRef]
- Das, J.; Kelley, S.O. Tuning the bacterial detection sensitivity of nanostructured microelectrodes. Anal. Chem. 2013, 85, 7333–7338. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Liang, W.; Wen, Y.; Wang, L.; Yang, X.; Ren, S.; Jia, N.; Zuo, X.; Liu, G. An ultrasensitive electrochemical biosensor for the detection of mecA gene in methicillin-resistant Staphylococcus aureus. Biosens. Bioelectron. 2018, 99, 424–430. [Google Scholar] [CrossRef] [PubMed]
- Lam, B.; Fang, Z.; Sargent, E.H.; Kelley, S.O. Polymerase chain reaction-free, sample-to-answer bacterial detection in 30 minutes with integrated cell lysis. Anal. Chem. 2012, 84, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Esteban-Fernández de Ávila, B.; Pedrero, M.; Campuzano, S.; Escamilla-Gómez, V.; Pingarrón, J.M. Sensitive and rapid amperometric magnetoimmunosensor for the determination of Staphylococcus aureus. Anal. Bioanal. Chem. 2012, 403, 917–925. [Google Scholar] [CrossRef]
- Labib, M.; Zamay, A.S.; Kolovskaya, O.S.; Reshetneva, I.T.; Zamay, G.S.; Kibbee, R.J.; Sattar, S.A.; Zamay, T.N.; Berezovski, M.V. Aptamer-based viability impedimetric sensor for bacteria. Anal. Chem. 2012, 84, 8966–8969. [Google Scholar] [CrossRef]
- Wang, L.; Lin, X.; Liu, T.; Zhang, Z.; Kong, J.; Yu, H.; Yan, J.; Luan, D.; Zhao, Y.; Bian, X. Reusable and universal impedimetric sensing platform for the rapid and sensitive detection of pathogenic bacteria based on bacteria-imprinted polythiophene film. Analyst 2022, 147, 4433–4441. [Google Scholar] [CrossRef]
- Brosel-Oliu, S.; Mergel, O.; Uria, N.; Abramova, N.; van Rijn, P.; Bratov, A. 3D impedimetric sensors as a tool for monitoring bacterial response to antibiotics. Lab Chip 2019, 19, 1436–1447. [Google Scholar] [CrossRef]
- Lim, S.A.; Ahmed, M.U. Electrochemical immunosensors and their recent nanomaterial-based signal amplification strategies: A review. RSC Adv. 2016, 6, 24995–25014. [Google Scholar] [CrossRef]
- Samuel, V.r.; Rao, K.J. A review on label free biosensors. Biosens. Bioelectron. X 2022, 11, 100216. [Google Scholar] [CrossRef]
- Ghindilis, A.L.; Atanasov, P.; Wilkins, E. Enzyme-catalyzed direct electron transfer: Fundamentals and analytical applications. Electroanalysis 1997, 9, 661–674. [Google Scholar] [CrossRef]
- Chikkaveeraiah, B.V.; Bhirde, A.A.; Morgan, N.Y.; Eden, H.S.; Chen, X. Electrochemical immunosensors for detection of cancer protein biomarkers. ACS Nano 2012, 6, 6546–6561. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.S. Electrochemical immunosensors for the simultaneous detection of two tumor markers. Anal. Chem. 2005, 77, 1496–1502. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Lin, Y. Nanomaterial labels in electrochemical immunosensors and immunoassays. Talanta 2007, 74, 308–317. [Google Scholar] [CrossRef]
- Mollarasouli, F.; Kurbanoglu, S.; Ozkan, S.A. The Role of Electrochemical Immunosensors in Clinical Analysis. Biosensors 2019, 9, 86. [Google Scholar] [CrossRef] [PubMed]
- Tombelli, S.; Minunni, M.; Mascini, M. Analytical applications of aptamers. Biosens. Bioelectron. 2005, 20, 2424–2434. [Google Scholar] [CrossRef]
- Hong, P.; Li, W.; Li, J. Applications of aptasensors in clinical diagnostics. Sensors 2012, 12, 1181–1193. [Google Scholar] [CrossRef] [PubMed]
- Jo, N.; Kim, B.; Lee, S.M.; Oh, J.; Park, I.H.; Jin Lim, K.; Shin, J.S.; Yoo, K.H. Aptamer-functionalized capacitance sensors for real-time monitoring of bacterial growth and antibiotic susceptibility. Biosens. Bioelectron. 2018, 102, 164–170. [Google Scholar] [CrossRef]
- Reynoso, E.C.; Laschi, S.; Palchetti, I.; Torres, E. Advances in Antimicrobial Resistance Monitoring Using Sensors and Biosensors: A Review. Chemosensors 2021, 9, 232. [Google Scholar] [CrossRef]
- Harada, L.K.; Júnior, W.B.; Silva, E.C.; Oliveira, T.J.; Moreli, F.C.; Júnior, J.M.O.; Tubino, M.; Vila, M.; Balcão, V.M. Bacteriophage-Based Biosensing of Pseudomonas aeruginosa: An Integrated Approach for the Putative Real-Time Detection of Multi-Drug-Resistant Strains. Biosensors 2021, 11, 124. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Guo, Y.; Qiu, T.; Gao, M.; Wang, X. Bacteriophages: Underestimated vehicles of antibiotic resistance genes in the soil. Front. Microbiol. 2022, 13, 936267. [Google Scholar] [CrossRef]
- Roy, S.; Arshad, F.; Eissa, S.; Safavieh, M.; Alattas, S.G.; Ahmed, M.U.; Zourob, M. Recent developments towards portable point-of-care diagnostic devices for pathogen detection. Sens. Diagn. 2022, 1, 87–105. [Google Scholar] [CrossRef]
- Singh, A.; Poshtiban, S.; Evoy, S. Recent advances in bacteriophage based biosensors for food-borne pathogen detection. Sensors 2013, 13, 1763–1786. [Google Scholar] [CrossRef]
- Zhang, H.; Chiao, M. Anti-fouling Coatings of Poly(dimethylsiloxane) Devices for Biological and Biomedical Applications. J. Med. Biol. Eng. 2015, 35, 143–155. [Google Scholar] [CrossRef]
- Yang, L.; Bashir, R. Electrical/electrochemical impedance for rapid detection of foodborne pathogenic bacteria. Biotechnol. Adv. 2008, 26, 135–150. [Google Scholar] [CrossRef]
- Hargol Zadeh, S.; Kashanian, S.; Nazari, M. A Label-Free Carbohydrate-Based Electrochemical Sensor to Detect Escherichia coli Pathogenic Bacteria Using D-mannose on a Glassy Carbon Electrode. Biosensors 2023, 13, 619. [Google Scholar] [CrossRef]
- Wang, Z.; Zhang, J.; Chen, P.; Zhou, X.; Yang, Y.; Wu, S.; Niu, L.; Han, Y.; Wang, L.; Chen, P.; et al. Label-free, electrochemical detection of methicillin-resistant Staphylococcus aureus DNA with reduced graphene oxide-modified electrodes. Biosens. Bioelectron. 2011, 26, 3881–3886. [Google Scholar] [CrossRef]
- Kumar, N.; Wang, W.; Ortiz-Marquez, J.C.; Catalano, M.; Gray, M.; Biglari, N.; Hikari, K.; Ling, X.; Gao, J.; van Opijnen, T.; et al. Dielectrophoresis assisted rapid, selective and single cell detection of antibiotic resistant bacteria with G-FETs. Biosens. Bioelectron. 2020, 156, 112123. [Google Scholar] [CrossRef]
- Altintas, Z.; Akgun, M.; Kokturk, G.; Uludag, Y. A fully automated microfluidic-based electrochemical sensor for real-time bacteria detection. Biosens. Bioelectron. 2018, 100, 541–548. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Wang, Y.; Zhang, D. A novel multifunctional electrochemical platform for simultaneous detection, elimination, and inactivation of pathogenic bacteria based on the Vancomycin-functionalised AgNPs/3D-ZnO nanorod arrays. Biosens. Bioelectron. 2017, 98, 248–253. [Google Scholar] [CrossRef] [PubMed]
- Crane, B.; Hughes, J.P.; Rowley Neale, S.J.; Rashid, M.; Linton, P.E.; Banks, C.E.; Shaw, K.J. Rapid antibiotic susceptibility testing using resazurin bulk modified screen-printed electrochemical sensing platforms. Analyst 2021, 146, 5574–5583. [Google Scholar] [CrossRef]
- Laliwala, A.; Pant, A.; Svechkarev, D.; Sadykov, M.R.; Mohs, A.M. Advancements of paper-based sensors for antibiotic-resistant bacterial species identification. NPJ Biosensing 2024, 1, 17. [Google Scholar] [CrossRef] [PubMed]
- Boehle, K.E.; Gilliand, J.; Wheeldon, C.R.; Holder, A.; Adkins, J.A.; Geiss, B.J.; Ryan, E.P.; Henry, C.S. Utilizing Paper-Based Devices for Antimicrobial-Resistant Bacteria Detection. Angew. Chem. Int. Ed. Engl. 2017, 56, 6886–6890. [Google Scholar] [CrossRef]
- Mazur, F.; Tjandra, A.D.; Zhou, Y.; Gao, Y.; Chandrawati, R. Paper-based sensors for bacteria detection. Nat. Rev. Bioeng. 2023, 1, 180–192. [Google Scholar] [CrossRef]
- Keskin, D.; Mergel, O.; van der Mei, H.C.; Busscher, H.J.; van Rijn, P. Inhibiting Bacterial Adhesion by Mechanically Modulated Microgel Coatings. Biomacromolecules 2019, 20, 243–253. [Google Scholar] [CrossRef]
- Aurchey, N.S.; Hasan, S.; Reshad, R.A.I.; Bakul, M.M.H.; Zabeen, N.; Zahan, T.; Shiddiky, M.J.A.; Biswas, G.C. Paper strip embedded masking tape device for low-cost, fast, length-based colorimetric and multiple antibiotic susceptibility testing. Sens. Actuators B Chem. 2025, 436, 137689. [Google Scholar] [CrossRef]
- Choopara, I.; Suea-Ngam, A.; Teethaisong, Y.; Howes, P.D.; Schmelcher, M.; Leelahavanichkul, A.; Thunyaharn, S.; Wongsawaeng, D.; deMello, A.J.; Dean, D.; et al. Fluorometric Paper-Based, Loop-Mediated Isothermal Amplification Devices for Quantitative Point-of-Care Detection of Methicillin-Resistant Staphylococcus aureus (MRSA). ACS Sens. 2021, 6, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhou, X.; Liu, H.; Deng, H.; Huang, R.; Xing, D. Simultaneous Detection of Antibiotic Resistance Genes on Paper-Based Chip Using [Ru(phen)(2)dppz](2+) Turn-on Fluorescence Probe. ACS Appl. Mater. Interfaces 2018, 10, 4494–4501. [Google Scholar] [CrossRef]
- Njoku, D.I.; Guo, Q.; Dai, W.; Chen, J.L.; Mao, G.; Sun, Q.; Sun, H.; Peng, Y.-K. The multipurpose application of resazurin in micro-analytical techniques: Trends from the microbial, catalysis and single molecule detection assays. TrAC Trends Anal. Chem. 2023, 167, 117288. [Google Scholar] [CrossRef]
- Alatraktchi, F.A.a.; Dimaki, M.; Støvring, N.; Johansen, H.K.; Molin, S.; Svendsen, W.E. Nanograss sensor for selective detection of Pseudomonas aeruginosa by pyocyanin identification in airway samples. Anal. Biochem. 2020, 593, 113586. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Numan, A.; Cinti, S. Point-of-Care for Evaluating Antimicrobial Resistance through the Adoption of Functional Materials. Anal. Chem. 2022, 94, 26–40. [Google Scholar] [CrossRef]
- Ortiz, M.; Jauset-Rubio, M.; Skouridou, V.; Machado, D.; Viveiros, M.; Clark, T.G.; Simonova, A.; Kodr, D.; Hocek, M.; O’Sullivan, C.K. Electrochemical Detection of Single-Nucleotide Polymorphism Associated with Rifampicin Resistance in Mycobacterium tuberculosis Using Solid-Phase Primer Elongation with Ferrocene-Linked Redox-Labeled Nucleotides. ACS Sens. 2021, 6, 4398–4407. [Google Scholar] [CrossRef]
- Butterworth, A.; Pratibha, P.; Marx, A.; Corrigan, D.K. Electrochemical Detection of Oxacillin Resistance using Direct-Labeling Solid-Phase Isothermal Amplification. ACS Sens. 2021, 6, 3773–3780. [Google Scholar] [CrossRef]
- Rao, R.P.; Sharma, S.; Mehrotra, T.; Das, R.; Kumar, R.; Singh, R.; Roy, I.; Basu, T. Rapid Electrochemical Monitoring of Bacterial Respiration for Gram-Positive and Gram-Negative Microbes: Potential Application in Antimicrobial Susceptibility Testing. Anal. Chem. 2020, 92, 4266–4274. [Google Scholar] [CrossRef]
- Li, X.; Lu, J.; Feng, L.; Zhang, L.; Gong, J. Smart pH-Regulated Switchable Nanoprobes for Photoelectrochemical Multiplex Detection of Antibiotic Resistance Genes. Anal. Chem. 2020, 92, 11476–11483. [Google Scholar] [CrossRef] [PubMed]
- Wan, C.; Qu, A.; Li, M.; Tang, R.; Fu, L.; Liu, X.; Wang, P.; Wu, C. Electrochemical Sensor for Directional Recognition and Measurement of Antibiotic Resistance Genes in Water. Anal. Chem. 2022, 94, 732–739. [Google Scholar] [CrossRef] [PubMed]
- Safavieh, M.; Pandya, H.J.; Venkataraman, M.; Thirumalaraju, P.; Kanakasabapathy, M.K.; Singh, A.; Prabhakar, D.; Chug, M.K.; Shafiee, H. Rapid Real-Time Antimicrobial Susceptibility Testing with Electrical Sensing on Plastic Microchips with Printed Electrodes. ACS Appl. Mater. Interfaces 2017, 9, 12832–12840. [Google Scholar] [CrossRef] [PubMed]
- Norouz Dizaji, A.; Ali, Z.; Ghorbanpoor, H.; Ozturk, Y.; Akcakoca, I.; Avci, H.; Dogan Guzel, F. Electrochemical-based ‘’antibiotsensor’’ for the whole-cell detection of the vancomycin-susceptible bacteria. Talanta 2021, 234, 122695. [Google Scholar] [CrossRef]
- Hannah, S.; Addington, E.; Alcorn, D.; Shu, W.; Hoskisson, P.A.; Corrigan, D.K. Rapid antibiotic susceptibility testing using low-cost, commercially available screen-printed electrodes. Biosens. Bioelectron. 2019, 145, 111696. [Google Scholar] [CrossRef]
- Li, C.; Sun, F. Graphene-Assisted Sensor for Rapid Detection of Antibiotic Resistance in Escherichia coli. Front. Chem. 2021, 9, 696906. [Google Scholar] [CrossRef]
- Ren, Y.; Ji, J.; Sun, J.; Pi, F.; Zhang, Y.; Sun, X. Rapid detection of antibiotic resistance in Salmonella with screen printed carbon electrodes. J. Solid State Electrochem. 2020, 24, 1539–1549. [Google Scholar] [CrossRef]
- Bolotsky, A.; Muralidharan, R.; Butler, D.; Root, K.; Murray, W.; Liu, Z.; Ebrahimi, A. Organic redox-active crystalline layers for reagent-free electrochemical antibiotic susceptibility testing (ORACLE-AST). Biosens. Bioelectron. 2021, 172, 112615. [Google Scholar] [CrossRef]
- Mishra, P.; Singh, D.; Mishra, K.P.; Kaur, G.; Dhull, N.; Tomar, M.; Gupta, V.; Kumar, B.; Ganju, L. Rapid antibiotic susceptibility testing by resazurin using thin film platinum as a bio-electrode. J. Microbiol. Methods. 2019, 162, 69–76. [Google Scholar] [CrossRef]
- Wang, T.; Zhang, Z.; Li, Y.; Xie, G. Amplified electrochemical detection of mecA gene in methicillin-resistant Staphylococcus aureus based on target recycling amplification and isothermal strand-displacement polymerization reaction. Sens. Actuators B Chem. 2015, 221, 148–154. [Google Scholar] [CrossRef]
- Suea-Ngam, A.; Howes, P.D.; deMello, A.J. An amplification-free ultra-sensitive electrochemical CRISPR/Cas biosensor for drug-resistant bacteria detection. Chem. Sci. 2021, 12, 12733–12743. [Google Scholar] [CrossRef]
- Cihalova, K.; Hegerova, D.; Dostalova, S.; Jelinkova, P.; Krejcova, L.; Milosavljevic, V.; Krizkova, S.; Kopel, P.; Adam, V. Particle-based immunochemical separation of methicillin resistant Staphylococcus aureus with indirect electrochemical detection of labeling oligonucleotides. Anal. Methods 2016, 8, 5123–5128. [Google Scholar] [CrossRef]
- Watanabe, K.; Kuwata, N.; Sakamoto, H.; Amano, Y.; Satomura, T.; Suye, S. A smart DNA sensing system for detecting methicillin-resistant Staphylococcus aureus using modified nanoparticle probes. Biosens. Bioelectron. 2015, 67, 419–423. [Google Scholar] [CrossRef] [PubMed]
- Mandal, S.S.; Navratna, V.; Sharma, P.; Gopal, B.; Bhattacharyya, A.J. Titania nanotube-modified screen printed carbon electrodes enhance the sensitivity in the electrochemical detection of proteins. Bioelectrochemistry 2014, 98, 46–52. [Google Scholar] [CrossRef]
- Khue, V.Q.; Huy, T.Q.; Phan, V.N.; Tuan-Le, A.; Thanh Le, D.T.; Tonezzer, M.; Hong Hanh, N.T. Electrochemical stability of screen-printed electrodes modified with Au nanoparticles for detection of methicillin-resistant Staphylococcus aureus. Mater. Chem. Phys. 2020, 255, 123562. [Google Scholar] [CrossRef]
- Patel, D.; Zhou, Y.; Ramasamy, R.P. A Bacteriophage-Based Electrochemical Biosensor for Detection of Methicillin-Resistant Staphylococcus aureus. J. Electrochem. Soc. 2021, 168, 057523. [Google Scholar] [CrossRef]
- Liu, M.; Hua, X.; Erhui, H.; Li, W.; Xiaoying, J.; Xianqing, C.; Shangchun, S.; and Xie, G. Ultrasensitive Electrochemical Biosensor for the Detection of the mecA Gene Sequence in Methicillin Resistant Strains of Staphylococcus aureus Employing Gold Nanoparticles. Anal. Lett. 2014, 47, 579–591. [Google Scholar] [CrossRef]
- Dai, G.; Li, Z.; Luo, F.; Lu, Y.; Chu, Z.; Zhang, J.; Zhang, F.; Wang, Q.; He, P. Simultaneous electrochemical determination of nuc and mecA genes for identification of methicillin-resistant Staphylococcus aureus using N-doped porous carbon and DNA-modified MOF. Microchim. Acta 2021, 188, 39. [Google Scholar] [CrossRef]
- Nemr, C.R.; Smith, S.J.; Liu, W.; Mepham, A.H.; Mohamadi, R.M.; Labib, M.; Kelley, S.O. Nanoparticle-Mediated Capture and Electrochemical Detection of Methicillin-Resistant Staphylococcus aureus. Anal. Chem. 2019, 91, 2847–2853. [Google Scholar] [CrossRef]
- Lee, K.S.; Lee, S.M.; Oh, J.; Park, I.H.; Song, J.H.; Han, M.; Yong, D.; Lim, K.J.; Shin, J.S.; Yoo, K.H. Electrical antimicrobial susceptibility testing based on aptamer-functionalized capacitance sensor array for clinical isolates. Sci. Rep. 2020, 10, 13709. [Google Scholar] [CrossRef]
- Khoshroo, A.; Mavaei, M.; Rostami, M.; Valinezhad-Saghezi, B.; Fattahi, A. Recent advances in electrochemical strategies for bacteria detection. Bioimpacts 2022, 12, 567–588. [Google Scholar] [CrossRef]
- Kelly, A.M.; Mathema, B.; Larson, E.L. Carbapenem-resistant Enterobacteriaceae in the community: A scoping review. Int. J. Antimicrob. Agents 2017, 50, 127–134. [Google Scholar] [CrossRef]
- Huang, J.M.; Henihan, G.; Macdonald, D.; Michalowski, A.; Templeton, K.; Gibb, A.P.; Schulze, H.; Bachmann, T.T. Rapid Electrochemical Detection of New Delhi Metallo-beta-lactamase Genes To Enable Point-of-Care Testing of Carbapenem-Resistant Enterobacteriaceae. Anal. Chem. 2015, 87, 7738–7745. [Google Scholar] [CrossRef]
- Pan, H.Z.; Yu, H.W.; Wang, N.; Zhang, Z.; Wan, G.C.; Liu, H.; Guan, X.; Chang, D. Electrochemical DNA biosensor based on a glassy carbon electrode modified with gold nanoparticles and graphene for sensitive determination of Klebsiella pneumoniae carbapenemase. J. Biotechnol. 2015, 214, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Subak, H.; Yilmaz, F.F.; Ozkan-Ariksoysal, D. Ready-to-use diagnostic kit based on electrochemical nanobiosensor for antibiotic resistance gene determination. Microchem. J. 2024, 204, 110976. [Google Scholar] [CrossRef]
- Yang, Y.; Gupta, K.; Ekinci, K.L. All-electrical monitoring of bacterial antibiotic susceptibility in a microfluidic device. Proc. Natl. Acad. Sci. USA 2020, 117, 10639–10644. [Google Scholar] [CrossRef]
- Mohan, R.; Mukherjee, A.; Sevgen, S.E.; Sanpitakseree, C.; Lee, J.; Schroeder, C.M.; Kenis, P.J. A multiplexed microfluidic platform for rapid antibiotic susceptibility testing. Biosens. Bioelectron. 2013, 49, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Sabhachandani, P.; Sarkar, S.; Zucchi, P.C.; Whitfield, B.A.; Kirby, J.E.; Hirsch, E.B.; Konry, T. Integrated microfluidic platform for rapid antimicrobial susceptibility testing and bacterial growth analysis using bead-based biosensor via fluorescence imaging. Microchim. Acta 2017, 184, 4619–4628. [Google Scholar] [CrossRef]
- Zhu, M.; Xu, T.; Cheng, Y.; Ma, B.; Xu, J.; Diao, Z.; Wu, F.; Dai, J.; Han, X.; Zhu, P.; et al. Integrated Microfluidic Chip for Rapid Antimicrobial Susceptibility Testing Directly from Positive Blood Cultures. Anal. Chem. 2023, 95, 14375–14383. [Google Scholar] [CrossRef]
- Arshavsky-Graham, S.; Segal, E. Lab-on-a-Chip Devices for Point-of-Care Medical Diagnostics. In Microfluidics in Biotechnology; Bahnemann, J., Grünberger, A., Eds.; Springer International Publishing: Cham, Switzerland, 2022; pp. 247–265. [Google Scholar]
- Mambatta, A.K.; Jayarajan, J.; Rashme, V.L.; Harini, S.; Menon, S.; Kuppusamy, J. Reliability of dipstick assay in predicting urinary tract infection. J. Family Med. Prim. Care 2015, 4, 265–268. [Google Scholar] [CrossRef] [PubMed]
- Ombelet, S.; Ronat, J.-B.; Walsh, T.; Yansouni, C.P.; Cox, J.; Vlieghe, E.; Martiny, D.; Semret, M.; Vandenberg, O.; Jacobs, J.; et al. Clinical bacteriology in low-resource settings: Today’s solutions. Lancet Infect. Dis. 2018, 18, e248–e258. [Google Scholar] [CrossRef]
- Weinstein, M.P.; Lewis, J.S., 2nd. The Clinical and Laboratory Standards Institute Subcommittee on Antimicrobial Susceptibility Testing: Background, Organization, Functions, and Processes. J. Clin. Microbiol. 2020, 58. [Google Scholar] [CrossRef]
- Chu, H.; Liu, C.; Liu, J.; Yang, J.; Li, Y.; Zhang, X. Recent advances and challenges of biosensing in point-of-care molecular diagnosis. Sens. Actuators B Chem. 2021, 348, 130708. [Google Scholar] [CrossRef]
- Smith Kenneth, P.; Kirby James, E. The Inoculum Effect in the Era of Multidrug Resistance: Minor Differences in Inoculum Have Dramatic Effect on MIC Determination. Antimicrob. Agents Chemother. 2018, 62. [Google Scholar] [CrossRef]
- Baker, C.N.; Thornsberry, C.; Hawkinson, R.W. Inoculum standardization in antimicrobial susceptibility testing: Evaluation of overnight agar cultures and the Rapid Inoculum Standardization System. J. Clin. Microbiol. 1983, 17, 450–457. [Google Scholar] [CrossRef] [PubMed]
- Huys, G.; D’Haene, K.; Swings, J. Influence of the culture medium on antibiotic susceptibility testing of food-associated lactic acid bacteria with the agar overlay disc diffusion method. Lett. Appl. Microbiol. 2002, 34, 402–406. [Google Scholar] [CrossRef]
- Fang, X.; Zheng, Y.; Duan, Y.; Liu, Y.; Zhong, W. Recent Advances in Design of Fluorescence-Based Assays for High-Throughput Screening. Anal. Chem. 2019, 91, 482–504. [Google Scholar] [CrossRef]
- Alves, I.P.; Reis, N.M. Immunocapture of Escherichia coli in a fluoropolymer microcapillary array. J. Chromatogr. A 2019, 1585, 46–55. [Google Scholar] [CrossRef] [PubMed]
- Vollstedt, A.; Baunoch, D.; Wolfe, A.; Luke, N.; Wojno, K.J.; Cline, K.; Belkoff, L.; Milbank, A.; Sherman, N.; Haverkorn, R.; et al. Bacterial Interactions as Detected by Pooled Antibiotic Susceptibility Testing (P-AST) in Polymicrobial Urine Specimens. J. Surg. Urol. 2020, 1, 101. [Google Scholar]
- Zhao, Y.; Xin, H.; Wang, C. Biomarker Multiplexing with Rational Design of Nucleic Acid Probe Complex. Anal. Sens. 2024, 4, e202400009. [Google Scholar] [CrossRef]
- Welling, M.M.; Hensbergen, A.W.; Bunschoten, A.; Velders, A.H.; Scheper, H.; Smits, W.K.; Roestenberg, M.; van Leeuwen, F.W.B. Fluorescent imaging of bacterial infections and recent advances made with multimodal radiopharmaceuticals. Clin. Transl. Imaging 2019, 7, 125–138. [Google Scholar] [CrossRef]
- Mistretta, M.; Gangneux, N.; Manina, G. Microfluidic dose–response platform to track the dynamics of drug response in single mycobacterial cells. Sci. Rep. 2022, 12, 19578. [Google Scholar] [CrossRef]
- Wang, Z.; Cai, R.; Gao, Z.; Yuan, Y.; Yue, T. Immunomagnetic separation: An effective pretreatment technology for isolation and enrichment in food microorganisms detection. Compr. Rev. Food Sci. Food Saf. 2020, 19, 3802–3824. [Google Scholar] [CrossRef]
- Bonnet, M.; Lagier, J.C.; Raoult, D.; Khelaifia, S. Bacterial culture through selective and non-selective conditions: The evolution of culture media in clinical microbiology. New Microbes New Infect. 2020, 34, 100622. [Google Scholar] [CrossRef]
- Choi, Y.H.; Zhang, C.; Liu, Z.; Tu, M.J.; Yu, A.X.; Yu, A.M. A Novel Integrated Pharmacokinetic-Pharmacodynamic Model to Evaluate Combination Therapy and Determine In Vivo Synergism. J. Pharmacol. Exp. Ther. 2021, 377, 305–315. [Google Scholar] [CrossRef]
- Dou, M.; Dominguez, D.C.; Li, X.; Sanchez, J.; Scott, G. A versatile PDMS/paper hybrid microfluidic platform for sensitive infectious disease diagnosis. Anal. Chem. 2014, 86, 7978–7986. [Google Scholar] [CrossRef] [PubMed]
- Bernholc, J.; Brenner, D.; Buongiorno Nardelli, M.; Meunier, V.; Roland, C. Mechanical and Electrical Properties of Nanotubes. Annu. Rev. Mater. Res. 2002, 32, 347–375. [Google Scholar] [CrossRef]
- Dreyer, D.R.; Park, S.; Bielawski, C.W.; Ruoff, R.S. The chemistry of graphene oxide. Chem. Soc. Rev. 2010, 39, 228–240. [Google Scholar] [CrossRef]
- Cheng, W.; Chen, H.; Liu, C.; Ji, C.; Ma, G.; Yin, M. Functional organic dyes for health-related applications. VIEW 2020, 1, 20200055. [Google Scholar] [CrossRef]
- Banerjee, I.; Pangule, R.C.; Kane, R.S. Antifouling Coatings: Recent Developments in the Design of Surfaces That Prevent Fouling by Proteins, Bacteria, and Marine Organisms. Adv. Mater. 2011, 23, 690–718. [Google Scholar] [CrossRef] [PubMed]
- Labib, M.; Sargent, E.H.; Kelley, S.O. Electrochemical Methods for the Analysis of Clinically Relevant Biomolecules. Chem. Rev. 2016, 116, 9001–9090. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, B.; Gangadhar, S.; Conrad, J.M. A survey of multisensor fusion techniques, architectures and methodologies. In Proceedings of the SoutheastCon 2017, Charlotte, NC, USA, 30 March–2 April 2017; pp. 1–8. [Google Scholar]
- Natalia, A.; Zhang, L.; Sundah, N.R.; Zhang, Y.; Shao, H. Analytical device miniaturization for the detection of circulating biomarkers. Nat. Rev. Bioeng. 2023, 1, 481–498. [Google Scholar] [CrossRef]
- Fang, W.; Liu, X.; Maiga, M.; Cao, W.; Mu, Y.; Yan, Q.; Zhu, Q. Digital PCR for Single-Cell Analysis. Biosensors 2024, 14, 64. [Google Scholar] [CrossRef]
- Mumtaz, H.; Riaz, M.H.; Wajid, H.; Saqib, M.; Zeeshan, M.H.; Khan, S.E.; Chauhan, Y.R.; Sohail, H.; Vohra, L.I. Current challenges and potential solutions to the use of digital health technologies in evidence generation: A narrative review. Front. Digit. Health 2023, 5, 1203945. [Google Scholar] [CrossRef] [PubMed]
| Method | Working Electrode | Probe | Target Pathogens | Tested Antibiotics | Assay Time | Linear Range | Detection Limit | Advantages and Limitations | Ref. |
|---|---|---|---|---|---|---|---|---|---|
| Square wave voltammetry (SWV) | Gold electrode | Solid-phase isothermal primer | Mycobacterium tuberculosis | Rifampicin | 25 min | 6 µM–140 µM | 3 pM | Rapid approach and detecting multiple SNPs; additional development and validation may require | [95] |
| Chronoamperometry | SPGE | HRP-labeled thymine nucleotides | E. coli | Oxacillin | 60 min | 208 × 103 to 2.08 × 103 cfu/mL | 319 CFU/mL | Rapid detection; direct coupling of the modified RPA reaction to electrochemical measurement simplifies the detection process | [96] |
| Cyclic voltammetry (CV) | L-lysine coated CeO/ITO | K3Fe [CN]6 | E. coli | Ciprofloxacin, cefixime, amoxicillin | 15 min | 1.07 × 105 to 1.3 × 108 CFU/mL | 4.5×104 CFU/mL | Faster than traditional AST method | [97] |
| Photoclectrochemistry | SDNA1-SbT@SiO2NSs complex and Sb2S3/ZnS/ITO | DNA | E. coli | Penicillin | Not reported | 1 nM to 10 µM | 1 nM | It can be successfully used for measuring bla-CTX-M-1 and bla-TEM in real E. coli plasmids | [98] |
| Impedance | Gold electrode with surface graphene ink | ssDNA-GE | b-lactam gene | Ampicillin | 1 h | 6.3–900 ng/mL | 6.3 ng/mL | Specific recognition ability for single-base, double-base, and three-base mismatch DNA | [99] |
| Impedance | Tantalum silicide electrode | PEI/p(NIPMAM/PDMS microgel | E. coli | Ampicillin | 60–120 min | 2–8 mg/L | 2 mg/L | High sensitivity, real-time monitoring; requires complex sample preparation | [59] |
| Impedance | Silver interdigitated carbon working electrode, counter electrode, and reference electrode | Label free | E. coli, Methicillin-resistant Staphylococcus aureus (MRSA) | Ampicillin, erythromycin, ciprofloxacin, methicillin, daptomycin, gentamicin | <90 min | 0.1 µM–100 µM | 0.1 µM | Rapid, label-free detection, capable of isolating bacteria from whole blood | [100] |
| Impedance | SPGE immobilized with thiolated vancomycin | HS-Van | S. aureus | Vancomycin | 10 min | 101–108 CFU/mL | <39 CFU/mL | Exhibits high performance for the detection of vancomycin susceptible bacteria | [101] |
| Impedance, DPV | Gold WE, gold CE, silver RE | Agrose-based hydrogel | S. aureus, MRSA | Amoxicillin, oxacillin | <45 min | 8 µg/mL and 50 µg/mL | 104 CFU/mL | Rapid detection within 45 min, low-cost, commercially available | [102] |
| DPV | Miniaturized incubation chamber containing WE, CE, and RE | Resazurin | E. coli, Klebsiella pneumoniae | Ampicillin, ciprofloxacin | 30 min | 1–1000 CFU/mL | 100 CFU/μL | Faster result, effective with clinically relevant levels of bacteria | [21] |
| DPV | Glassy carbon electrode | Graphene-modified electrode | E. coli | Ofloxacin, penicillin, cefepime | 30 min | 1 × 105 CFU–5 × 107 CFU/mL | 10 CFU/mL | Rapid, label-free detection | [103] |
| DPV | Resazurin modified Graphite SPE | Resazurin | E. coli | Gentamycin sulphate | 90 min | 0–1000 µM | 15.6 µM | Rapid, cost-effective | [84] |
| DPV | SPCE modified with multiwelled carbon tube and gold nanoparticle | Salmonella gallinarum | Ofloxacin, penicillin | 60 min | 102 to 107 CFU/mL | 100 CFU/mL | Rapid, highly sensitive | [104] | |
| DPV | Nafion-coated organic redox-active crystal layers on planar pyrolytic graphite sheets | E. coli | Ampicillin, kanamycin | 60 min | 0.001–10 µM, 0 µg/mL, 16 µg/mL | 16 µg/mL | Sensors are stable after 60 days of storage in ambient conditions and enable analysis of microbial viability in complex solutions | [105] | |
| DPV, resazurin detection | Platinum WE and CE, Ag/AgCl RE | E. coli, K. pneumoniae | Ampicillin, kanamycin, tetracycline | <4 h | 103 to 108 CFU/mL | 1 × 104 cells/mL | Easy, rapid, reliable, and inexpensive | [106] | |
| Capacitance | Gold electrode on a glass substrate | Aptamer | E. coli, S. aureus | Gentamycin | 2 h | 0–50 µg/mL | 10 CFU/mL | Enhanced sensitivity due to AuNP modification | [71] |
| SWV | mecAgene/MCH/hairpin probe/Au electrode | E-DNA | mecA DNA from MRSA | Methicillin | 1–2 h | 0–400 pM | 63 fM | Hairpin probe design ensures specific binding to the mecA gene | [107] |
| SWV | E-Si-CRISPR | Aptamer gRNA | mecA DNA from MRSA | Methicillin | 60 min | 10 fM–0.1 nM | 3.5 fM and 10 fM | Amplification-free, ultrasensitive; requires precise gRNA design for different targets | [108] |
| SWV | Au nanoparticles modified by anti-Pls | MRSA-specific antibody | Antigen | Methicillin | 2–3 h | 0.2–10 µM 4 × 107–2 × 104 CFU/mL | 2 × 104 CFU/mL | Early identification of inflammatory diseases in resistant bacteria | [109] |
| Amperometry | Screen printed gold electrode (SPGE) | Solid-phase RPA primers | DNA AMR gene of E. coli | Oxacillin | 60 min | 319–20,830 CFU/mL | 319 CFU/mL | Rapid detection; direct coupling of the modified RPA reaction to electrochemical measurement simplifies the detection process | [96] |
| CV | MNP/DNA1- Au/DNA-2 | Ferrocene-labeled probes | mecA DNA from MRSA | Methicillin | 2 h | 10–166 pM | 10 pM | Combination of MNPs and AuNPs enhances detection sensitivity Risk of non-specific binding | [110] |
| CV | TiO2-NTs | PBP2a Protein | S. aureus | Methicillin | Few minutes | 1–100 ng/mL | 1 ng/µL | rapid detection of target antigenic proteins | [111] |
| CV, DPV | Au/SPCE | Monoclonal anti-MRSA antibody and Aptamer gRNA | Antigen | Methicillin | 1–2 h | 10–106 CFU/mL | 13 CFU/mL | Rapid and label-free detection of highly pathogenic bacteria | [112] |
| EIS | GCE-APTES-rGOdsDNA | ssDNA | Methicillin-resistant Staphylococcus aureus MRSA) | Methicillin | 2 h | 0.1 pM–1 µM | 0.1 pM | Presence of rGO is favored to anchor both ssDNA and dsDNA, which provides the stable response of impedance | [80] |
| EIS | MCH-sDNA-GE | ssDNA-GE | b-lactam gene | Ampicillin | 1 h | 3.1–480 pM | 3.1 pM | Specific recognition ability for single-base, double-base, and three-base mismatch DNA | [99] |
| EIS | MSP-TSP/Au electrode | Multisignal Probes | 130 nt synthetic ssDNA and gDNA | Methicillin | 2.5 h | 100 nM–10 fM | 10 fM and 57 fM | Specifically, detection of mecA gene, portable on-site detection of MRSA | [56] |
| EIS | PEI-f-CNT | SATA-8505, bacteriophage | MRSA USA300 strain | Methicillin | 30 min | 102–107 CFU/mL | 1.29 × 102 CFU/mL in blood plasma | Potentially be integrated into a lab-on-a-chip platform for point of care use | [113] |
| DPV | mecA gene/Au/GCE | mecA gene | mecA DNA from MRSA | Methicillin | <2 h | 50–250 pM | 23 pM | Successful determination of mecA gene from MRSA | [114] |
| DPV | UiO-66/BMZIFderived NPCs | ssDNA | mecA and nuc gene DNA from MRSA | Methicillin | 2 h | 5–1×105 fM | 1.6 fM and 3.6 fM | MRSA and SA can be distinguished; great potential in practical applications | [115] |
| DPV | Au electrode | Monoclonal anti-MRSA antibody | PBP2a antibody | Methicillin | <4.5 h | 3–105 CFU/mL | 3 CFU/mL | Can easily be modified to capture various bacteria of interest by selecting appropriate capture and detection antibody pairs | [116] |
| Capacitance | e-AST system on Au | 60 aptamers | E. coli U433 | 11 antibiotic drugs | 6 h | 0.5–128 mg/mL | 103 CFU/mL | Rapid AST can increase survival rate of sepsis patients; diagnosis of sepsis by e-AST costly than gold standard method | [117] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khan, M.U.; Rahman, M.M.; Zahan, N.; Masud, M.K.; Sarker, S.; Haque, M.H. Electrochemical Biosensing for Antibiotic-Resistant Bacteria: Advances, Challenges, and Future Directions. Micromachines 2025, 16, 986. https://doi.org/10.3390/mi16090986
Khan MU, Rahman MM, Zahan N, Masud MK, Sarker S, Haque MH. Electrochemical Biosensing for Antibiotic-Resistant Bacteria: Advances, Challenges, and Future Directions. Micromachines. 2025; 16(9):986. https://doi.org/10.3390/mi16090986
Chicago/Turabian StyleKhan, Muhib Ullah, Md. Munibur Rahman, Nusrat Zahan, Mostafa Kamal Masud, Subir Sarker, and Md. Hakimul Haque. 2025. "Electrochemical Biosensing for Antibiotic-Resistant Bacteria: Advances, Challenges, and Future Directions" Micromachines 16, no. 9: 986. https://doi.org/10.3390/mi16090986
APA StyleKhan, M. U., Rahman, M. M., Zahan, N., Masud, M. K., Sarker, S., & Haque, M. H. (2025). Electrochemical Biosensing for Antibiotic-Resistant Bacteria: Advances, Challenges, and Future Directions. Micromachines, 16(9), 986. https://doi.org/10.3390/mi16090986








