Perfusion System for Modification of Luminal Contents of Human Intestinal Organoids and Realtime Imaging Analysis of Microbial Populations
Abstract
:1. Introduction
2. Materials and Methods
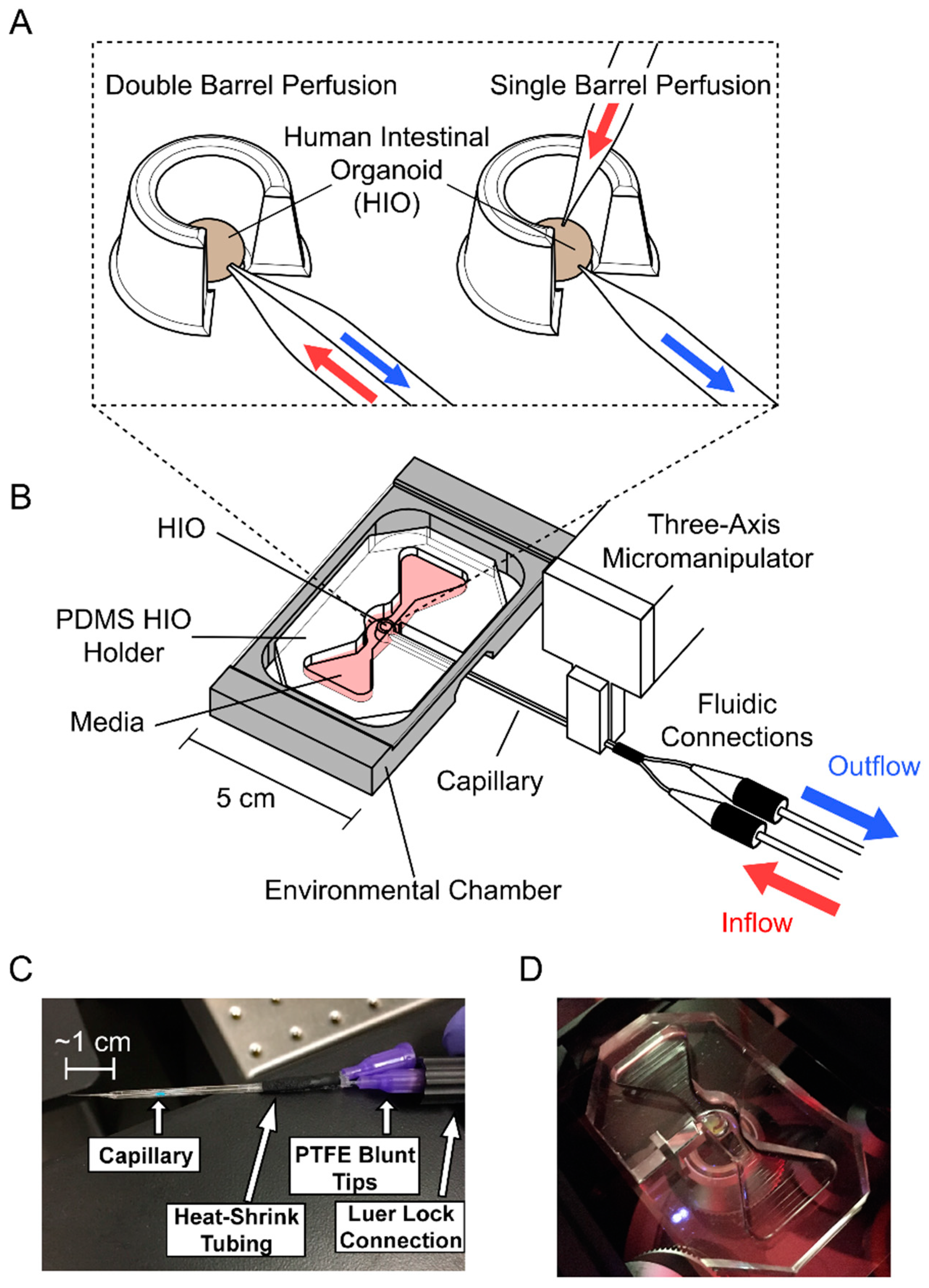
2.1. Perfusion System Fabrication
2.1.1. HIO Holder
2.1.2. Perfusion System Assembly
2.1.3. Capillary Tip Fabrication
2.2. HIO Production
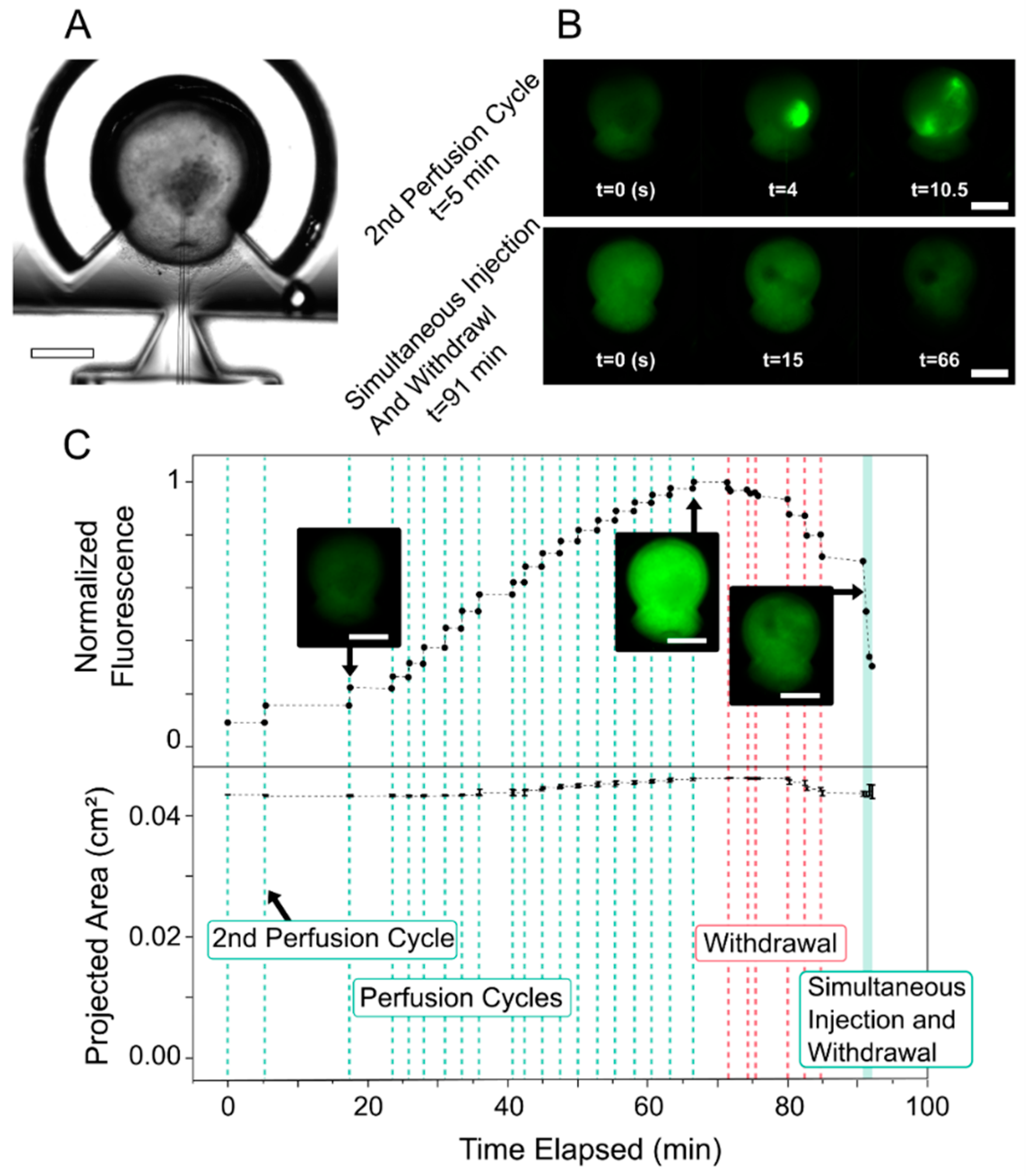
2.3. Perfusing HIOs with FITC Dextran
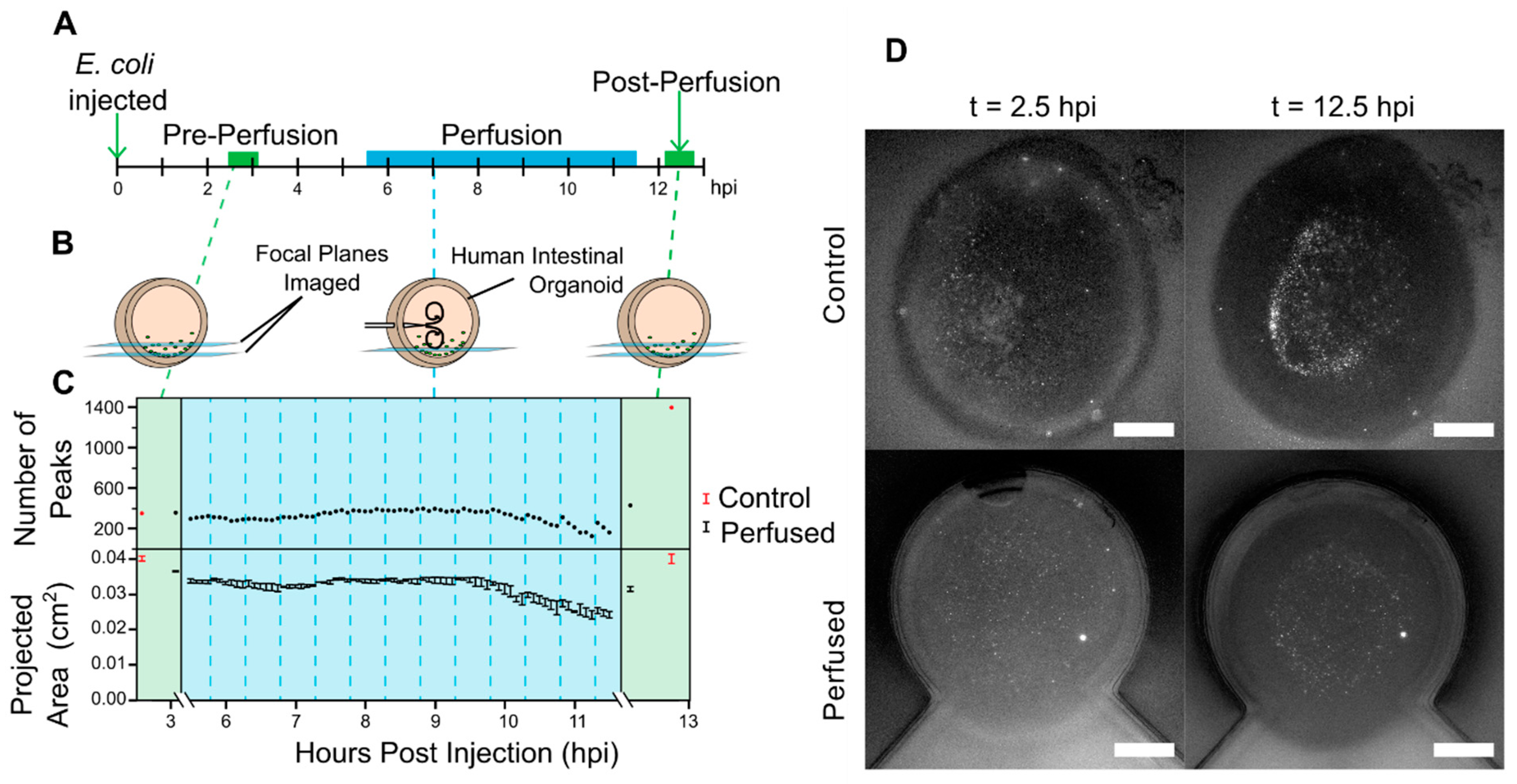
2.4. Colonizing HIOs with GFP-Labeled E. coli and Perfusion
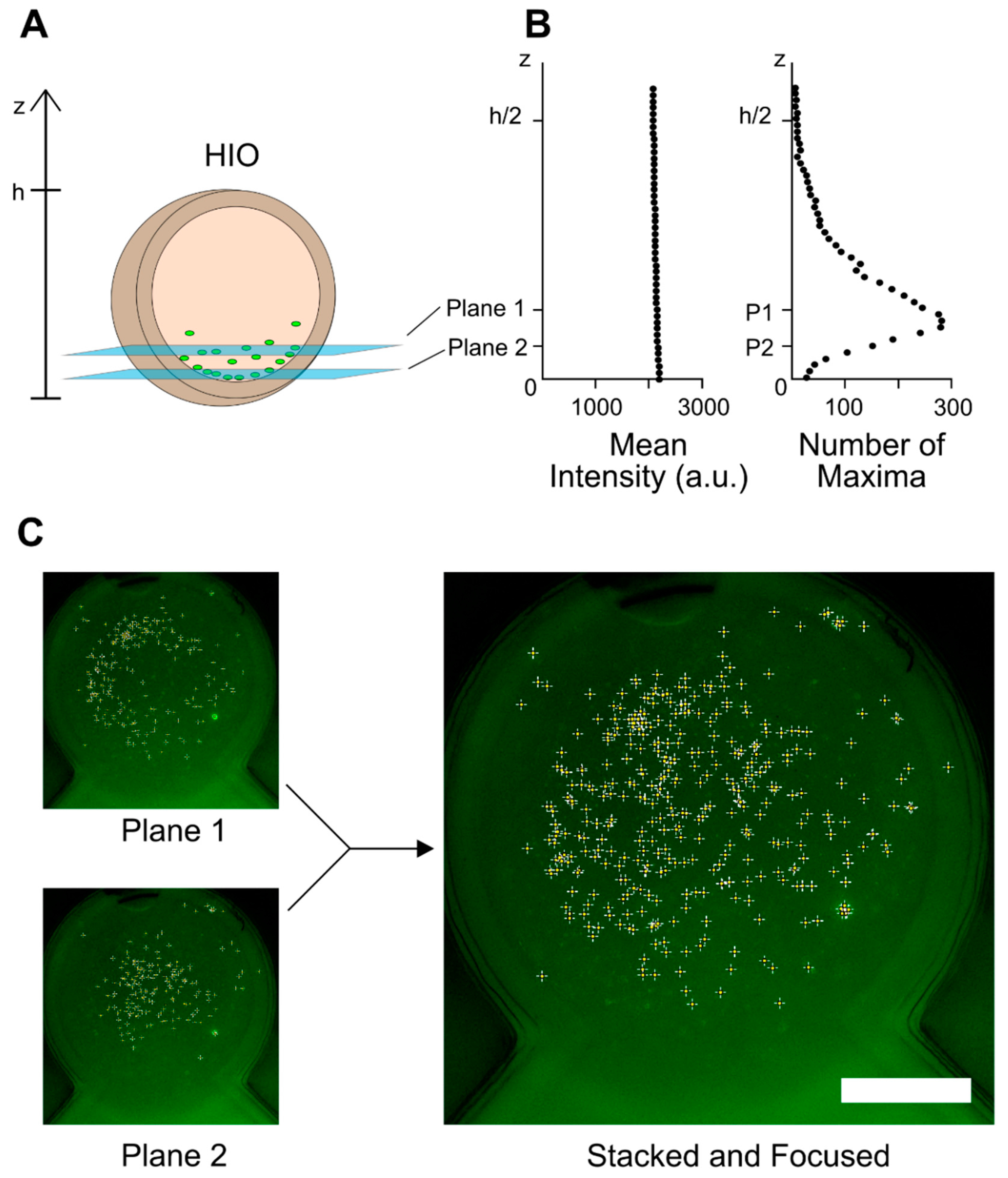
2.5. Image Analysis
2.6. Organoid Histology
3. Results and Discussion
3.1. Fluorescent Dye Perfusion Experiment
3.2. GFP-Labeled E. coli Imaging
3.3. GFP-Labeled E. coli-Perfused HIOs Experiment
3.4. Organoid Strain during Perfusion
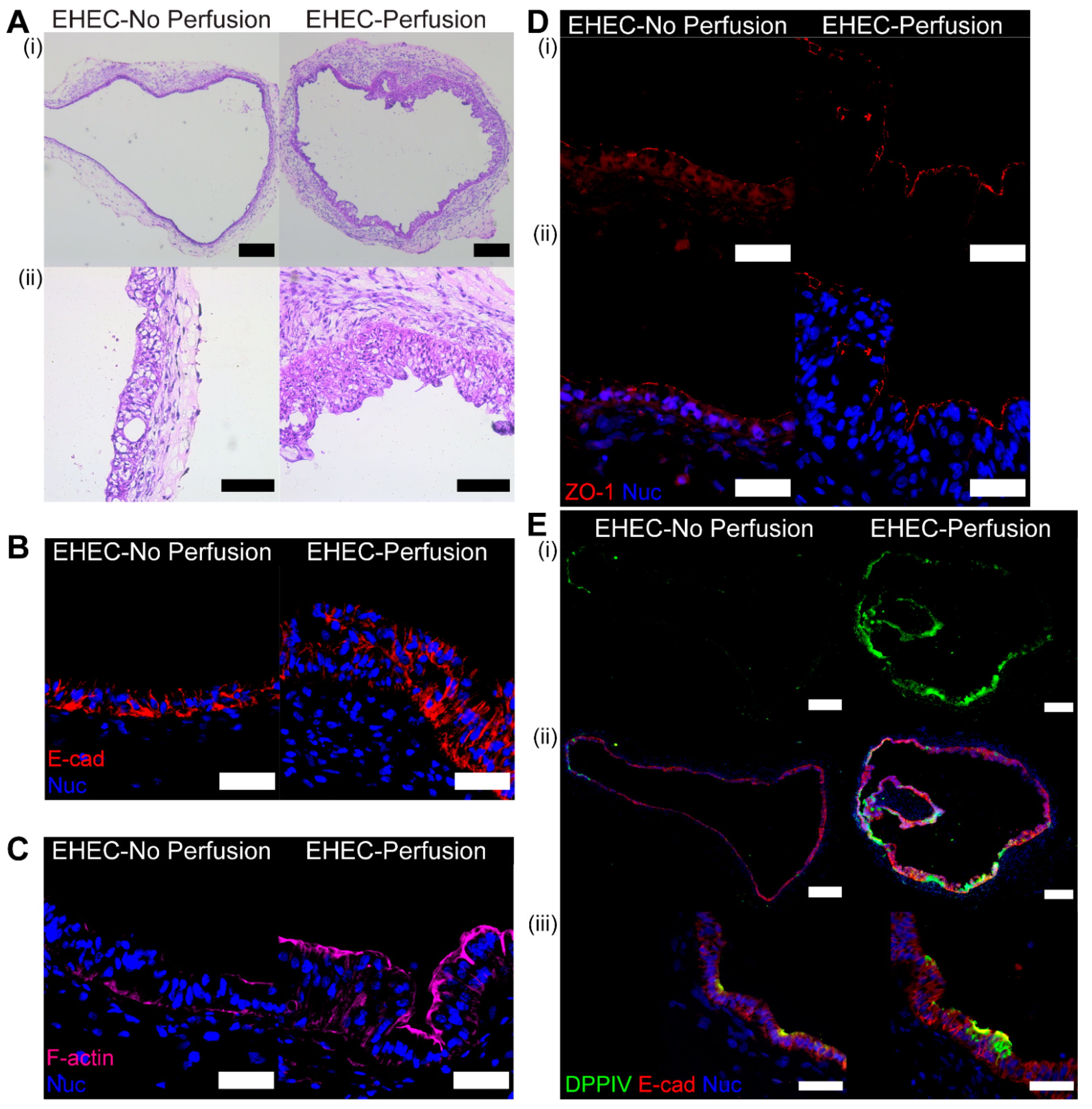
3.5. Histological Examination of Organoid Following Perfusion with E. coli
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Huch, M.; Koo, B.-K. Modeling mouse and human development using organoid cultures. Developement 2015, 142, 3113–3125. [Google Scholar] [CrossRef] [Green Version]
- Sato, T.; Vries, R.G.; Snippert, H.J.; van de Wetering, M.; Barker, N.; Stange, D.E.; van Es, J.H.; Abo, A.; Kujala, P.; Peters, P.J.; et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265. [Google Scholar] [CrossRef] [PubMed]
- McCracken, K.W.; Howel, J.C.; Wells, J.M.; Spence, J.R. Generating human intestinal tissue from pluripotent stem cells in vitro. Nat. Protoc. 2011, 6, 1920–1928. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Co, J.Y.; Margalef-Català, M.; Li, X.; Mah, A.T.; Kuo, C.J.; Monack, D.M.; Amieva, M.R. Controlling epithelial polarity: A human enteroid model for host-pathogen interactions. Cell Rep. 2019, 26, 2509–2520.e4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rossi, G.; Manfrin, A.; Lutolf, M.P. Progress and potential in organoid research. Nat. Rev. Genet. 2018, 19, 671–687. [Google Scholar] [CrossRef]
- Crawford, S.; Ramani, S.; Blutt, S.; Estes, M. Organoids to Dissect Gastrointestinal Virus–Host Interactions: What Have We Learned? Viruses 2021, 13, 999. [Google Scholar] [CrossRef]
- Noel, G.; Baetz, N.W.; Staab, J.F.; Donowitz, M.; Kovbasnjuk, O.; Pasetti, M.F.; Zachos, N.C. A primary human macrophage-enteroid co-culture model to investigate mucosal gut physiology and host-pathogen interactions. Sci. Rep. 2017, 7, 1–14. [Google Scholar]
- Kim, G.-A.; Ginga, N.J.; Takayama, S. Integration of sensors in gastrointestinal organoid culture for biological analysis. Cell. Mol. Gastroenterol. Hepatol. 2018, 6, 123–131.e1. [Google Scholar] [CrossRef] [Green Version]
- Sidar, B.; Jenkins, B.R.; Huang, S.; Spence, J.R.; Walk, S.T.; Wilking, J.N. Long-term flow through human intestinal organoids with the gut organoid flow chip (GOFlowChip). Lab Chip 2019, 19, 3552–3562. [Google Scholar] [CrossRef] [Green Version]
- Lee, K.K.; McCauley, H.A.; Broda, T.R.; Kofron, M.J.; Wells, J.M.; Hong, C.I. Human stomach-on-a-chip with luminal flow and peristaltic-like motility. Lab Chip 2018, 18, 3079–3085. [Google Scholar] [CrossRef] [PubMed]
- Williamson, I.A.; Arnold, J.W.; Samsa, L.A.; Gaynor, L.; DiSalvo, M.; Cocchiaro, J.L.; Carroll, I.; Azcarate-Peril, M.A.; Rawls, J.F.; Allbritton, N.L.; et al. A High-Throughput Organoid Microinjection Platform to Study Gastrointestinal Microbiota and Luminal Physiology. Cell. Mol. Gastroenterol. Hepatol. 2018, 6, 301–319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Puschhof, J.; Pleguezuelos-Manzano, C.; Martinez-Silgado, A.; Akkerman, N.; Saftien, A.; Boot, C.; de Waal, A.; Beumer, J.; Dutta, D.; Heo, I.; et al. Intestinal organoid cocultures with microbes. Nat. Protoc. 2021, 16, 4633–4649. [Google Scholar] [CrossRef] [PubMed]
- Duque-Correa, M.A.; Maizels, R.M.; Grencis, R.K.; Berriman, M. Organoids—New models for host–helminth interactions. Trends Parasitol. 2020, 36, 170–181. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Salahudeen, A.A.; Choi, S.S.; Rustagi, A.; Zhu, J.; van Unen, V.; Sean, M.; Flynn, R.A.; Mar, M.-C.; Santos, A.J.M. Progenitor identification and SARS-CoV-2 infection in human distal lung organoids. Nature 2020, 588, 670–675. [Google Scholar] [CrossRef]
- Heinen, N.; Klöhn, M.; Steinmann, E.; Pfaender, S. In vitro lung models and their application to study SARS-CoV-2 pathogenesis and disease. Viruses 2021, 13, 792. [Google Scholar] [CrossRef]
- Dutta, D.; Heo, I.; Clevers, H. Disease modeling in stem cell-derived 3D organoid systems. Trends Mol. Med. 2017, 23, 393–410. [Google Scholar] [CrossRef] [PubMed]
- Parigoris, E.; Lee, S.; Mertz, D.; Turner, M.; Liu, A.Y.; Sentosa, J.; Djomehri, S.; Chang, H.C.; Luker, K.; Luker, G.; et al. Cancer cell invasion of mammary organoids with basal-in phenotype. Adv. Healthc. Mater. 2021, 10, e2000810. [Google Scholar] [CrossRef]
- Hill, D.R.; Huang, S.; Nagy, M.S.; Yadagiri, V.K.; Fields, C.; Mukherjee, D.; Bons, B.; Dedhia, P.H.; Chin, A.M.; Tsai, Y.-H.; et al. Bacterial colonization stimulates a complex physiological response in the immature human intestinal epithelium. eLife 2017, 6, e29132. [Google Scholar] [CrossRef] [PubMed]
- Capelinga, M.; Huangb, S.; Mulero-Russec, A.; Ciezad, R.; Tsaib, Y.-H.; Garciac, A.; Hill, D.R. Chapter 7—Generation of Small Intestinal Organoids for Experimental Intestinal Physiology. In Human Pluripotent Stem Cell Derived Organoid Models; Spence, J.R., Ed.; Academic Press: Cambridge, MA, USA, 2020; Volume 159, pp. 143–174. [Google Scholar]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [Green Version]
- Selinummi, J.; Seppälä, J.; Yli-Harja, O.; Puhakka, J.A. Software for quantification of labeled bacteria from digital microscope images by automated image analysis. Future Sci. 2018, 39, 859–862. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Forster, B.; Van De Ville, D.; Berent, J.; Sage, D.; Unser, M. Extended depth-of-focus for multi-channel microscopy images: A complex wavelet approach. In Proceedings of the 2nd IEEE International Symposium on Biomedical Imaging Macro to Nano, Arlington, VA, USA, 15–18 April 2004; pp. 660–663. [Google Scholar]
- Gorbach, S.L. Chapter 95 Microbiology of the Gastrointestinal Tract Composition and Distribution of the Microflora Metabolic Activites of the Microflora The Intestinal Microflora and Infection. In Medocal Microbiology, 4th ed.; Baron, S., Ed.; Univeristy of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996; pp. 1–7. [Google Scholar]
- Jones, R.M. The Influence of the Gut Microbiota on Host Physiology: In Pursuit of Mechanisms. Yale J. Boil. Med. 2016, 89, 285–297. [Google Scholar]
- Kim, H.J.; Dongeun, H.; Hamilton, G.; Ingber, D.E. Human gut-on-a-chip inhabited by microbial flora that experiences intestinal peristalsis-like motions and flow. Lab Chip 2012, 12, 2165–2174. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.J.; Dye, B.R.; Ferrer-Torres, D.; Hill, D.R.; Overeem, A.W.; Shea, L.D.; Spenc, J.R. Generation of lung organoids from human pluripotent stem cells in vitro. Nat. Protoc. 2019, 14, 518–540. [Google Scholar] [CrossRef]
- Sharma, K.; Dhar, N.; Thacker, V.V.; Simonet, T.M.; Signorino-Gelor, F.; Knott, G.; McKinney, J.D. Dynamic persistence of intracellular bacterial communities of uropathogenic escherichia coli in a human bladder-chip model of urinary tract infections. eLife 2021, 10, 66481. [Google Scholar] [CrossRef]
- Karve, S.S.; Pradhan, S.; Ward, D.V.; Weiss, A.A. Intestinal organoids model human responses to infection by commensal and Shiga toxin producing Escherichia coli. PLoS ONE 2017, 12, e0178966. [Google Scholar] [CrossRef] [Green Version]
- Grassart, A.; Malarde, V.; Gobaa, S.; Sartori-Rup, A.; Kerns, J.; Karalis, K.; Marteyn, B.; Sansonetti, P.; Sauvonnet, N. Bioengineered human organ-on-chip reveals intestinal microenvironment and mechanical forces impacting shigella infection. Cell Host Microbe 2019, 26, 435–444.e4. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ginga, N.J.; Slyman, R.; Kim, G.-A.; Parigoris, E.; Huang, S.; Yadagiri, V.K.; Young, V.B.; Spence, J.R.; Takayama, S. Perfusion System for Modification of Luminal Contents of Human Intestinal Organoids and Realtime Imaging Analysis of Microbial Populations. Micromachines 2022, 13, 131. https://doi.org/10.3390/mi13010131
Ginga NJ, Slyman R, Kim G-A, Parigoris E, Huang S, Yadagiri VK, Young VB, Spence JR, Takayama S. Perfusion System for Modification of Luminal Contents of Human Intestinal Organoids and Realtime Imaging Analysis of Microbial Populations. Micromachines. 2022; 13(1):131. https://doi.org/10.3390/mi13010131
Chicago/Turabian StyleGinga, Nicholas J., Raleigh Slyman, Ge-Ah Kim, Eric Parigoris, Sha Huang, Veda K. Yadagiri, Vincent B. Young, Jason R. Spence, and Shuichi Takayama. 2022. "Perfusion System for Modification of Luminal Contents of Human Intestinal Organoids and Realtime Imaging Analysis of Microbial Populations" Micromachines 13, no. 1: 131. https://doi.org/10.3390/mi13010131
APA StyleGinga, N. J., Slyman, R., Kim, G.-A., Parigoris, E., Huang, S., Yadagiri, V. K., Young, V. B., Spence, J. R., & Takayama, S. (2022). Perfusion System for Modification of Luminal Contents of Human Intestinal Organoids and Realtime Imaging Analysis of Microbial Populations. Micromachines, 13(1), 131. https://doi.org/10.3390/mi13010131





