Core-Shell Beads Made by Composite Liquid Marble Technology as A Versatile Microreactor for Polymerase Chain Reaction
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of Photopolymer Liquid
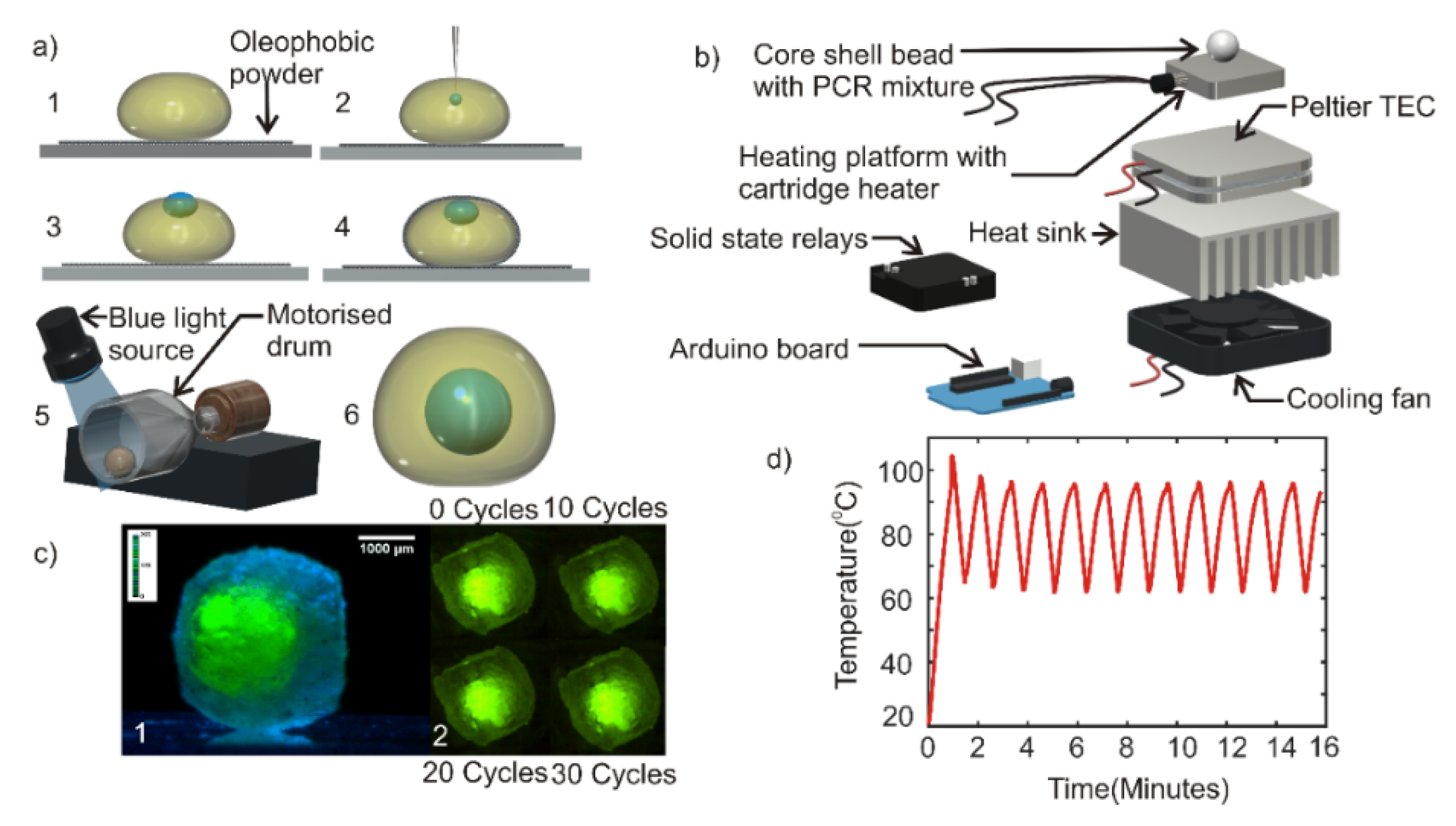
2.2. Synthesis of Composite Liquid Marbles and Core-Shell Beads
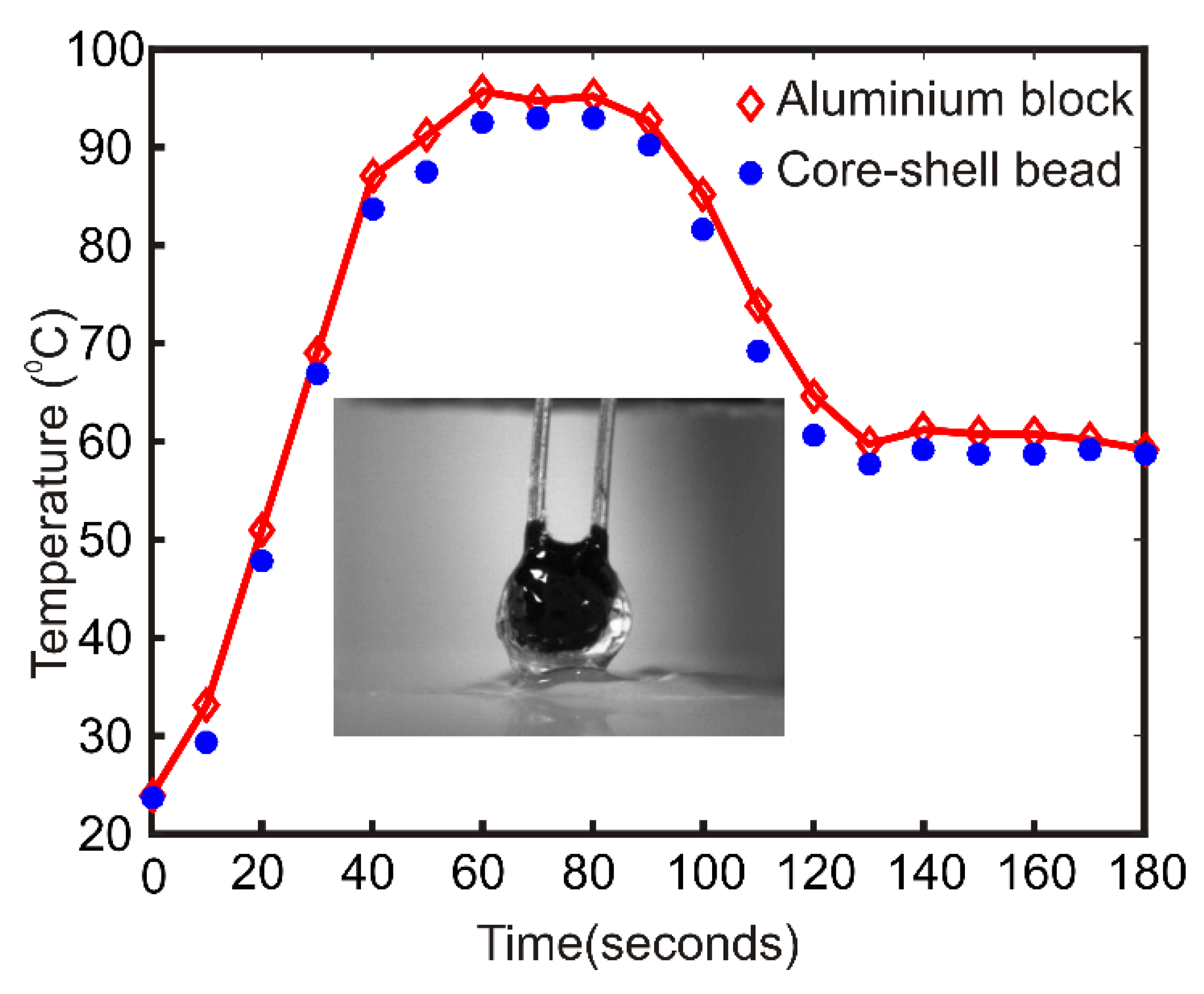
2.3. Design and Fabrication of Thermal Cycler
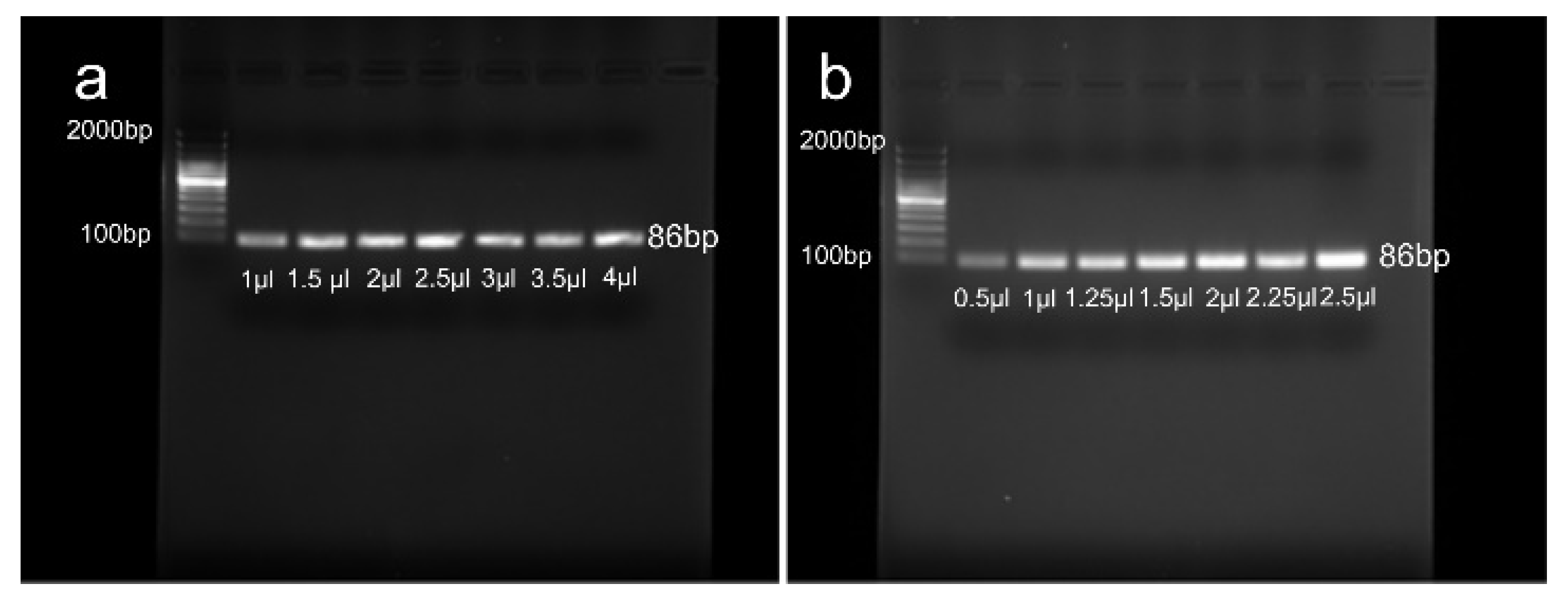
2.4. Preparation and Optimization of the PCR Mixture
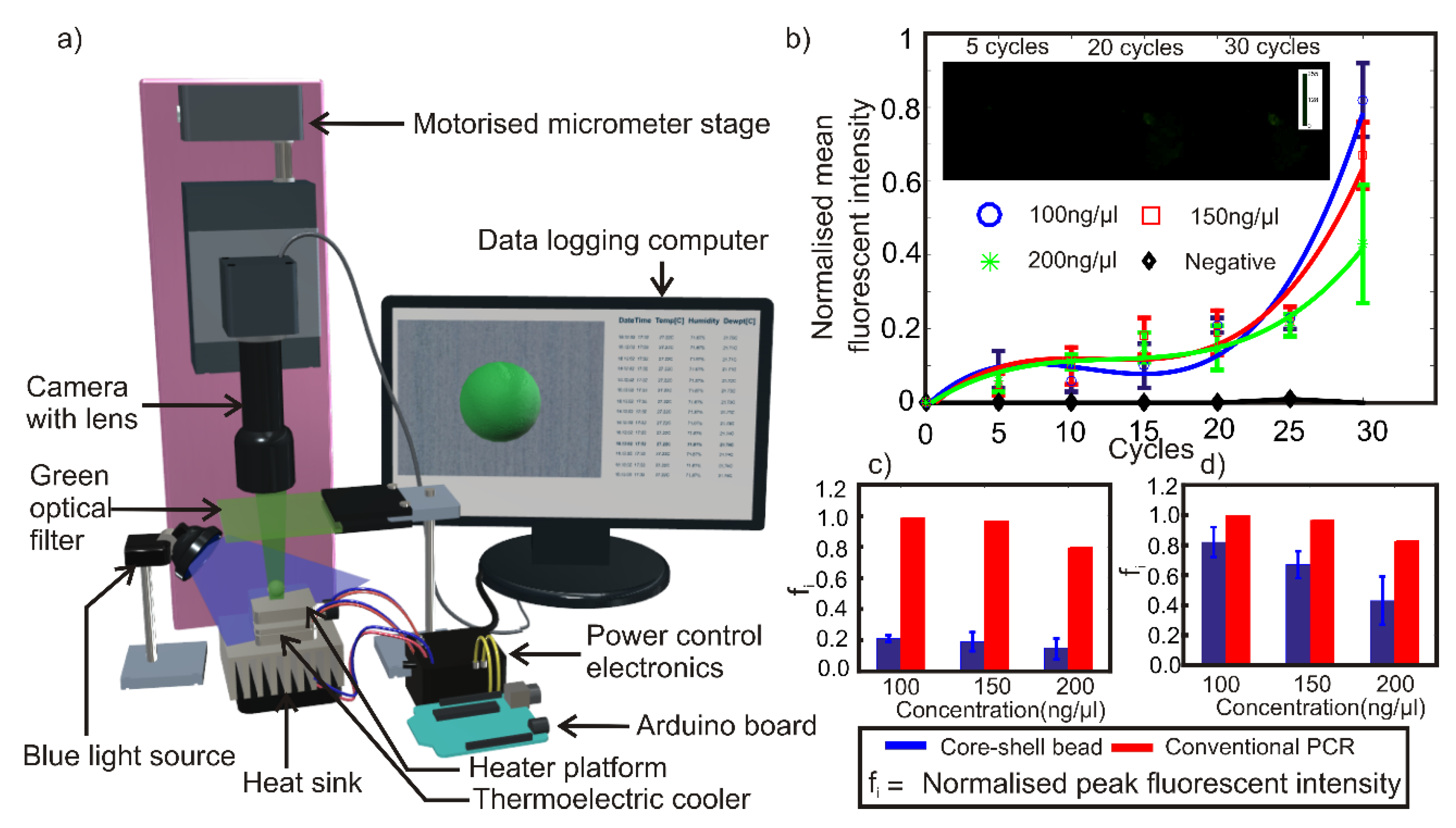
2.5. Design and Fabrication of Fluorescent Detection System
3. Experimental
4. Results and discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Ethical Approval
References
- Mullis, K.; Faloona, F.; Scharf, S.; Saiki, R.; Horn, G.; Erlich, H. Specific enzymatic amplification of DNA in vitro: The polymerase chain reaction. Cold Spring Harb. Symp. Quant. Biol. 1986, 51, 263–273. [Google Scholar] [CrossRef]
- Heyries, K.A.; Tropini, C.; Vaninsberghe, M.; Doolin, C.; Petriv, O.I.; Singhal, A.; Leung, K.; Hughesman, C.B.; Hansen, C.L. Megapixel digital PCR. Nat. Methods 2011, 8, 649–651. [Google Scholar] [CrossRef] [PubMed]
- Horsman, K.M.; Bienvenue, J.M.; Blasier, K.R.; Landers, J.P. Forensic DNA analysis on microfluidic devices: A review. J. Forensic. Sci. 2007, 52, 784–799. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Li, X.; Greenspoon, S.A.; Scherer, J.R.; Mathies, R.A. Integrated DNA purification, PCR, sample cleanup, and capillary electrophoresis microchip for forensic human identification. Lab Chip 2011, 11, 1041–1048. [Google Scholar] [CrossRef] [PubMed]
- Girones, R.; Ferrus, M.A.; Alonso, J.L.; Rodriguez-Manzano, J.; Calgua, B.; Correa Ade, A.; Hundesa, A.; Carratala, A.; Bofill-Mas, S. Molecular detection of pathogens in water--the pros and cons of molecular techniques. Water Res. 2010, 44, 4325–4339. [Google Scholar] [CrossRef]
- Baker, M. Clever PCR: More genotyping, smaller volumes. Nat. Methods 2010, 7, 351–356. [Google Scholar] [CrossRef]
- Zhang, H.; Jenkins, G.; Zou, Y.; Zhu, Z.; Yang, C.J. Massively parallel single-molecule and single-cell emulsion reverse transcription polymerase chain reaction using agarose droplet microfluidics. Anal. Chem. 2012, 84, 3599–3606. [Google Scholar] [CrossRef]
- Postollec, F.; Falentin, H.; Pavan, S.; Combrisson, J.; Sohier, D. Recent advances in quantitative PCR (qPCR) applications in food microbiology. Food Microbiol. 2011, 28, 848–861. [Google Scholar] [CrossRef]
- Urbina, M.A.; Watts, A.J.R.; Reardon, E.E. Labs should cut plastic waste too. Nature 2015, 528, 479. [Google Scholar] [CrossRef]
- Sreejith, K.R.; Ooi, C.H.; Jin, J.; Dao, D.; Nguyen, N.-T. Digital polymerase chain reaction technology – recent advances and future perspectives. Lab. A Chip 2018. [Google Scholar] [CrossRef]
- Aussillous, P.; Quéré, D. Liquid marbles. Nature 2001, 411, 924–927. [Google Scholar] [CrossRef] [PubMed]
- Polwaththe-Gallage, H.-N.; Ooi, C.H.; Jin, J.; Sauret, E.; Nguyen, N.-T.; Li, Z.; Gu, Y. The stress-strain relationship of liquid marbles under compression. Appl. Phys. Lett. 2019, 114, 043701. [Google Scholar] [CrossRef]
- Planchette, C.; Biance, A.-L.; Pitois, O.; Lorenceau, E. Coalescence of armored interface under impact. Phys. Fluids 2013, 25, 042104. [Google Scholar] [CrossRef]
- Draper, T.C.; Fullarton, C.; Mayne, R.; Phillips, N.; Canciani, G.E.; de Lacy Costello, B.P.J.; Adamatzky, A. Mapping outcomes of liquid marble collisions. Soft Matter 2019, 15, 3541–3551. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Fu, X.; Binks, B.P.; Shum, H.C. Mechanical Compression to Characterize the Robustness of Liquid Marbles. Langmuir 2015, 31, 11236–11242. [Google Scholar] [CrossRef]
- Sreejith, K.R.; Ooi, C.H.; Jin, J.; Dao, D.V.; Nguyen, N.-T. An automated on-demand liquid marble generator based on electrohydrodynamic pulling. Rev. Sci. Instrum. 2019, 90, 055102. [Google Scholar] [CrossRef]
- Ireland, P.M.; Kido, K.; Webber, G.B.; Fujii, S.; Wanless, E.J. pH-Responsive Particle-Liquid Aggregates-Electrostatic Formation Kinetics. Front. Chem. 2018, 6, 215. [Google Scholar] [CrossRef]
- Mahadevan, L.; Pomeau, Y. Rolling droplets. Phys. Fluids 1999, 11, 2449–2453. [Google Scholar] [CrossRef]
- Bormashenko, E.; Pogreb, R.; Balter, R.; Gendelman, O.; Aurbach, D. Composite non-stick droplets and their actuation with electric field. Appl. Phys. Lett. 2012, 100, 151601. [Google Scholar] [CrossRef]
- Khaw, M.K.; Ooi, C.H.; Mohd-Yasin, F.; Vadivelu, R.; John, J.S.; Nguyen, N.T. Digital microfluidics with a magnetically actuated floating liquid marble. Lab. Chip 2016, 16, 2211–2218. [Google Scholar] [CrossRef]
- Jin, J.; Ooi, C.H.; Sreejith, K.R.; Dao, D.V.; Nguyen, N.-T. Dielectrophoretic Trapping of a Floating Liquid Marble. Phys. Rev. Appl. 2019, 11, 044059. [Google Scholar] [CrossRef]
- Jin, J.; Ooi, C.H.; Sreejith, K.R.; Zhang, J.; Nguyen, A.V.; Evans, G.M.; Dao, D.V.; Nguyen, N.-T. Accurate dielectrophoretic positioning of a floating liquid marble with a two-electrode configuration. Microfluid. Nanofluidics 2019, 23, 85. [Google Scholar] [CrossRef]
- Ooi, C.H.; Jin, J.; Sreejith, K.R.; Nguyen, A.V.; Evans, G.M.; Nguyen, N.-T. Manipulation of a floating liquid marble using dielectrophoresis. Lab. A Chip 2018, 18, 3770–3779. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Sreejith, K.R.; Ooi, C.H.; Dao, D.V.; Nguyen, N.-T. Critical Trapping Conditions for Floating Liquid Marbles. Phys. Rev. Appl. 2020, 13, 014002. [Google Scholar] [CrossRef]
- Jin, J.; Nguyen, N.-T. Manipulation schemes and applications of liquid marbles for micro total analysis systems. Microelectron. Eng. 2018, 197, 87–95. [Google Scholar] [CrossRef]
- Ooi, C.H.; Jin, J.; Nguyen, A.V.; Evans, G.M.; Nguyen, N.-T. Picking up and placing a liquid marble using dielectrophoresis. Microfluid. Nanofluidics 2018, 22, 142. [Google Scholar] [CrossRef]
- Oliveira, N.M.; Correia, C.R.; Reis, R.L.; Mano, J.F. Liquid Marbles for High-Throughput Biological Screening of Anchorage-Dependent Cells. Adv. Healthc. Mater. 2015, 4, 264–270. [Google Scholar] [CrossRef]
- Vadivelu, R.; Kashaninejad, N.; Sreejith, K.R.; Bhattacharjee, R.; Cock, I.; Nguyen, N.-T. Cryoprotectant-Free Freezing of Cells Using Liquid Marbles Filled with Hydrogel. Acs Appl. Mater. Interfaces 2018, 10, 43439–43449. [Google Scholar] [CrossRef]
- Arbatan, T.; Li, L.; Tian, J.; Shen, W. Liquid marbles as micro-bioreactors for rapid blood typing. Adv. Healthc Mater. 2012, 1, 80–83. [Google Scholar] [CrossRef]
- Sarvi, F.; Arbatan, T.; Chan, P.P.Y.; Shen, W. A novel technique for the formation of embryoid bodies inside liquid marbles. Rsc Adv. 2013, 3, 14501. [Google Scholar] [CrossRef]
- Serrano, M.C.; Nardecchia, S.; Gutierrez, M.C.; Ferrer, M.L.; del Monte, F. Mammalian cell cryopreservation by using liquid marbles. Acs Appl. Mater. Interfaces 2015, 7, 3854–3860. [Google Scholar] [CrossRef] [PubMed]
- Sreejith, K.R.; Gorgannezhad, L.; Jin, J.; Ooi, C.H.; Stratton, H.; Dao, D.V.; Nguyen, N.-T. Liquid marbles as biochemical reactors for the polymerase chain reaction. Lab. A Chip 2019. [Google Scholar] [CrossRef] [PubMed]
- Sreejith, K.R.; Ooi, C.H.; Dao, D.V.; Nguyen, N.-T. Evaporation dynamics of liquid marbles at elevated temperatures. Rsc Adv. 2018, 8, 15436–15443. [Google Scholar] [CrossRef]
- CID=18689, N.C.f.B.I.P.D. Trimethylolpropane Trimethacrylate. Available online: https://pubchem.ncbi.nlm.nih.gov/compound/Trimethylolpropane-trimethacrylate#section=Information-Sources (accessed on 9 February 2020).
- Ono, S.; Kadoma, Y.; Morita, S.; Takakuda, K. Development of New Bone Cement utilizing Low Toxicity Monomers. J. Med. Dent. Sci. 2008, 55, 189–196. [Google Scholar] [CrossRef]
- Vining, K.H.; Scherba, J.C.; Bever, A.M.; Alexander, M.R.; Celiz, A.D.; Mooney, D.J. Synthetic Light-Curable Polymeric Materials Provide a Supportive Niche for Dental Pulp Stem Cells. Adv. Mater. 2018, 30, 1704486. [Google Scholar] [CrossRef]
- Hayase, G.; Kanamori, K.; Hasegawa, G.; Maeno, A.; Kaji, H.; Nakanishi, K. A Superamphiphobic Macroporous Silicone Monolith with Marshmallow-like Flexibility. Angew. Chem. Int. Ed. 2013, 52, 10788–10791. [Google Scholar] [CrossRef]
- Takei, T.; Yamasaki, Y.; Yuji, Y.; Sakoguchi, S.; Ohzuno, Y.; Hayase, G.; Yoshida, M. Millimeter-sized capsules prepared using liquid marbles: Encapsulation of ingredients with high efficiency and preparation of spherical core-shell capsules with highly uniform shell thickness using centrifugal force. J. Colloid Interface Sci. 2019, 536, 414–423. [Google Scholar] [CrossRef]
- Beer, N.R.; Hindson, B.J.; Wheeler, E.K.; Hall, S.B.; Rose, K.A.; Kennedy, I.M.; Colston, B.W. On-Chip, Real-Time, Single-Copy Polymerase Chain Reaction in Picoliter Droplets. Anal. Chem. 2007, 79, 8471–8475. [Google Scholar] [CrossRef]
- Li, L.; Shu, X.; Zhu, J. Low temperature depolymerization from a copper-based aqueous vinyl polymerization system. Polymer 2012, 53, 5010–5015. [Google Scholar] [CrossRef]
- Lloyd, D.J.; Nikolaou, V.; Collins, J.; Waldron, C.; Anastasaki, A.; Bassett, S.P.; Howdle, S.M.; Blanazs, A.; Wilson, P.; Kempe, K.; et al. Controlled aqueous polymerization of acrylamides and acrylates and “in situ” depolymerization in the presence of dissolved CO2. Chem. Commun. 2016, 52, 6533–6536. [Google Scholar] [CrossRef]
- Tang, H.; Luan, Y.; Yang, L.; Sun, H. A Perspective on Reversibility in Controlled Polymerization Systems: Recent Progress and New Opportunities. Molecules 2018, 23, 2870. [Google Scholar] [CrossRef] [PubMed]
- Vasantha, V.A.; Junhui, C.; Wenguang, Z.; van Herk, A.M.; Parthiban, A. Reversible Photo- and Thermoresponsive, Self-Assembling Azobenzene Containing Zwitterionic Polymers. Langmuir 2019, 35, 1465–1474. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sreejith, K.R.; Gorgannezhad, L.; Jin, J.; Ooi, C.H.; Takei, T.; Hayase, G.; Stratton, H.; Lamb, K.; Shiddiky, M.; Dao, D.V.; et al. Core-Shell Beads Made by Composite Liquid Marble Technology as A Versatile Microreactor for Polymerase Chain Reaction. Micromachines 2020, 11, 242. https://doi.org/10.3390/mi11030242
Sreejith KR, Gorgannezhad L, Jin J, Ooi CH, Takei T, Hayase G, Stratton H, Lamb K, Shiddiky M, Dao DV, et al. Core-Shell Beads Made by Composite Liquid Marble Technology as A Versatile Microreactor for Polymerase Chain Reaction. Micromachines. 2020; 11(3):242. https://doi.org/10.3390/mi11030242
Chicago/Turabian StyleSreejith, Kamalalayam Rajan, Lena Gorgannezhad, Jing Jin, Chin Hong Ooi, Takayuki Takei, Gen Hayase, Helen Stratton, Krystina Lamb, Muhammad Shiddiky, Dzung Viet Dao, and et al. 2020. "Core-Shell Beads Made by Composite Liquid Marble Technology as A Versatile Microreactor for Polymerase Chain Reaction" Micromachines 11, no. 3: 242. https://doi.org/10.3390/mi11030242
APA StyleSreejith, K. R., Gorgannezhad, L., Jin, J., Ooi, C. H., Takei, T., Hayase, G., Stratton, H., Lamb, K., Shiddiky, M., Dao, D. V., & Nguyen, N.-T. (2020). Core-Shell Beads Made by Composite Liquid Marble Technology as A Versatile Microreactor for Polymerase Chain Reaction. Micromachines, 11(3), 242. https://doi.org/10.3390/mi11030242










