Histopathological Effects of Bt and TcdA Insecticidal Proteins on the Midgut Epithelium of Western Corn Rootworm Larvae (Diabrotica virgifera virgifera)
Abstract
:1. Introduction
2. Results
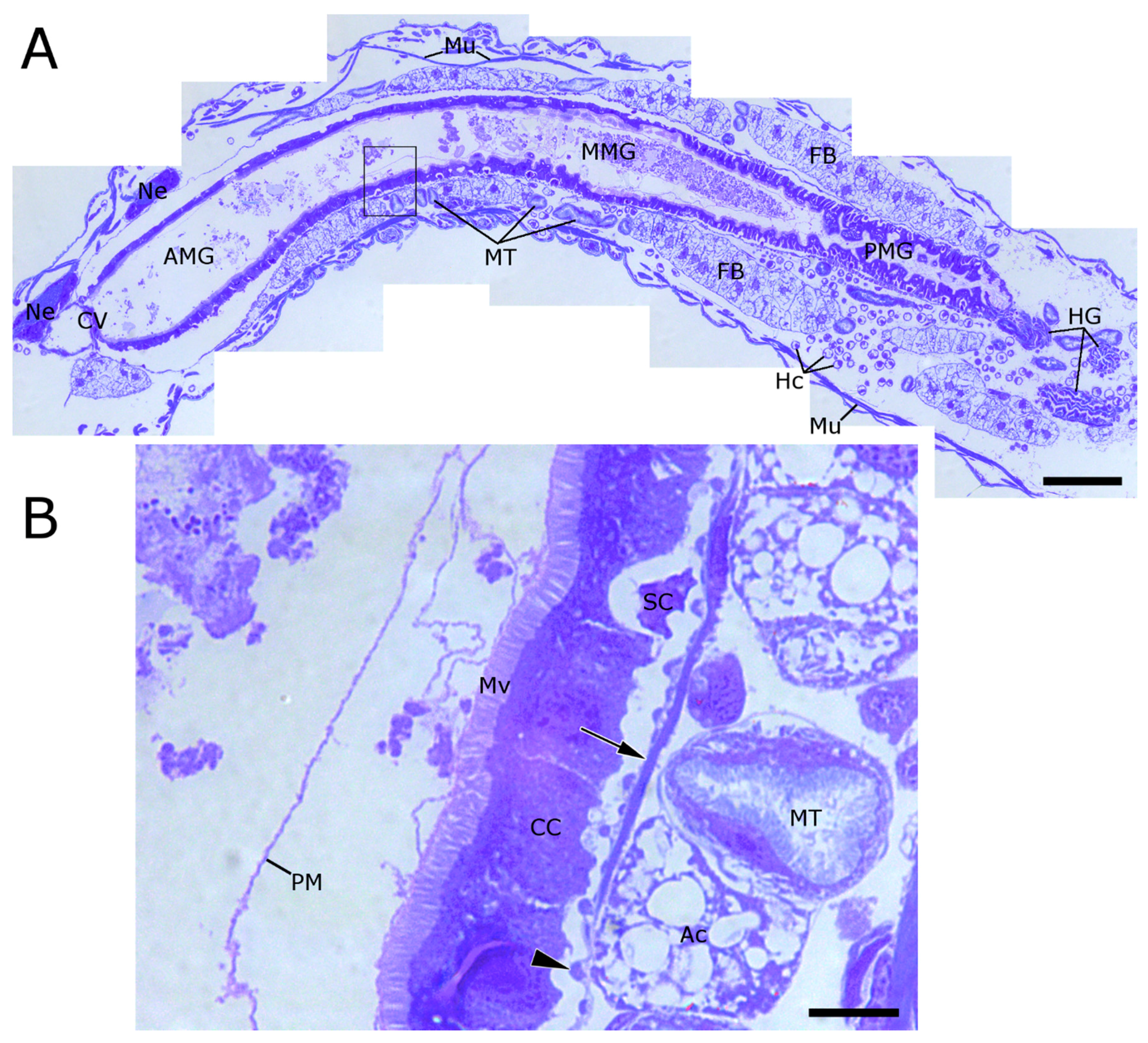
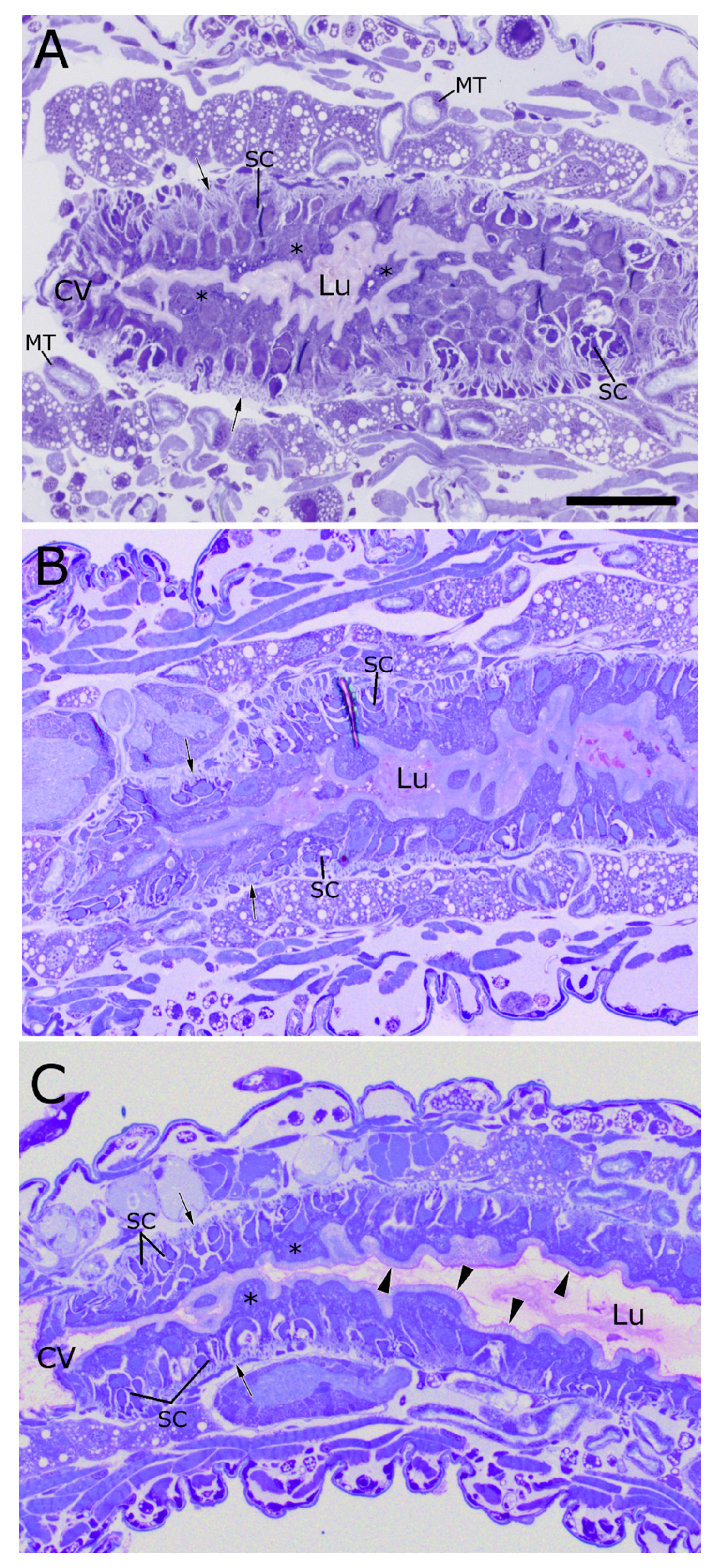
2.1. WCR Anatomy
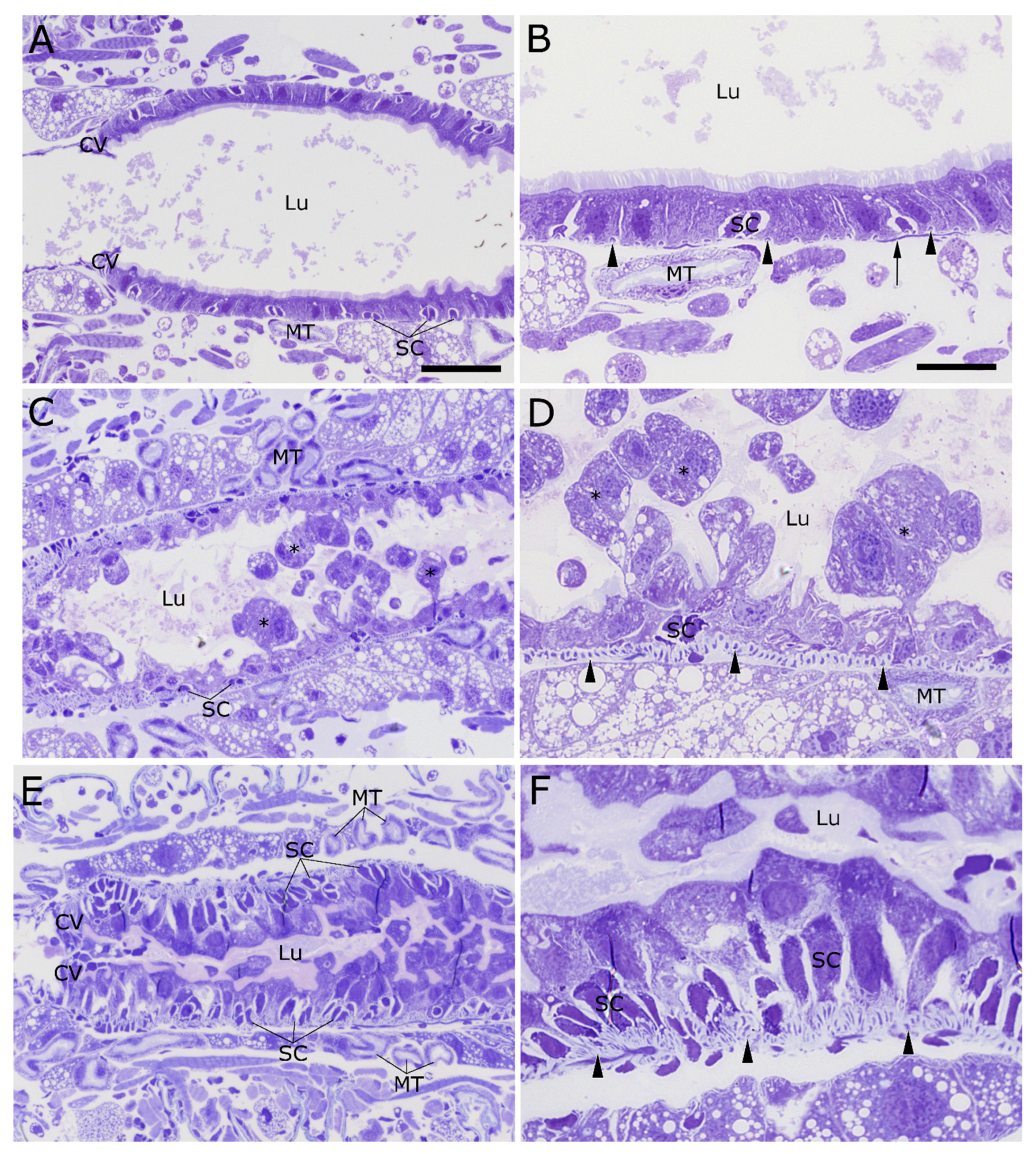
2.2. Cry34/35Ab1
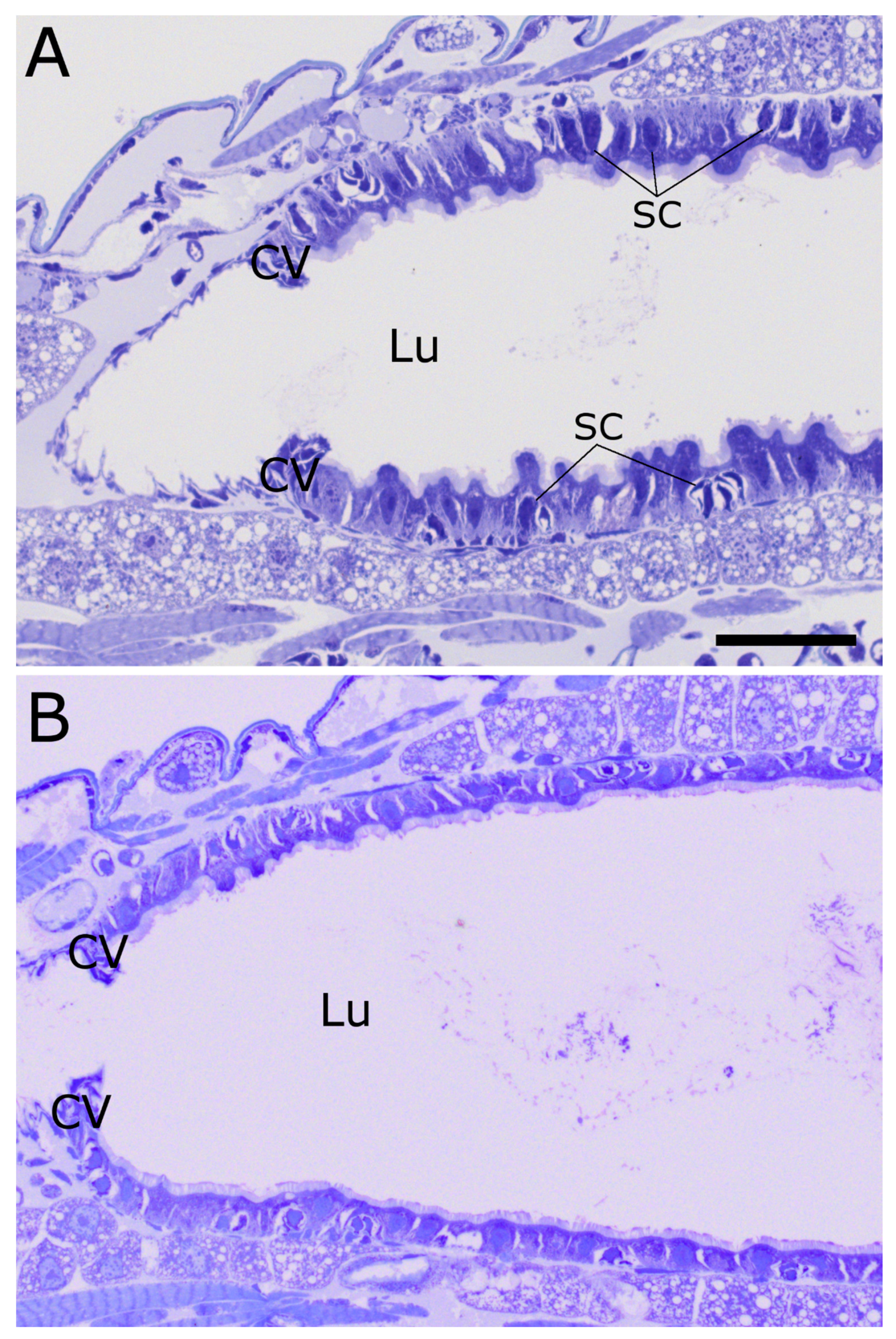
2.3. Cry34Ab1 or Cry35Ab1 Alone
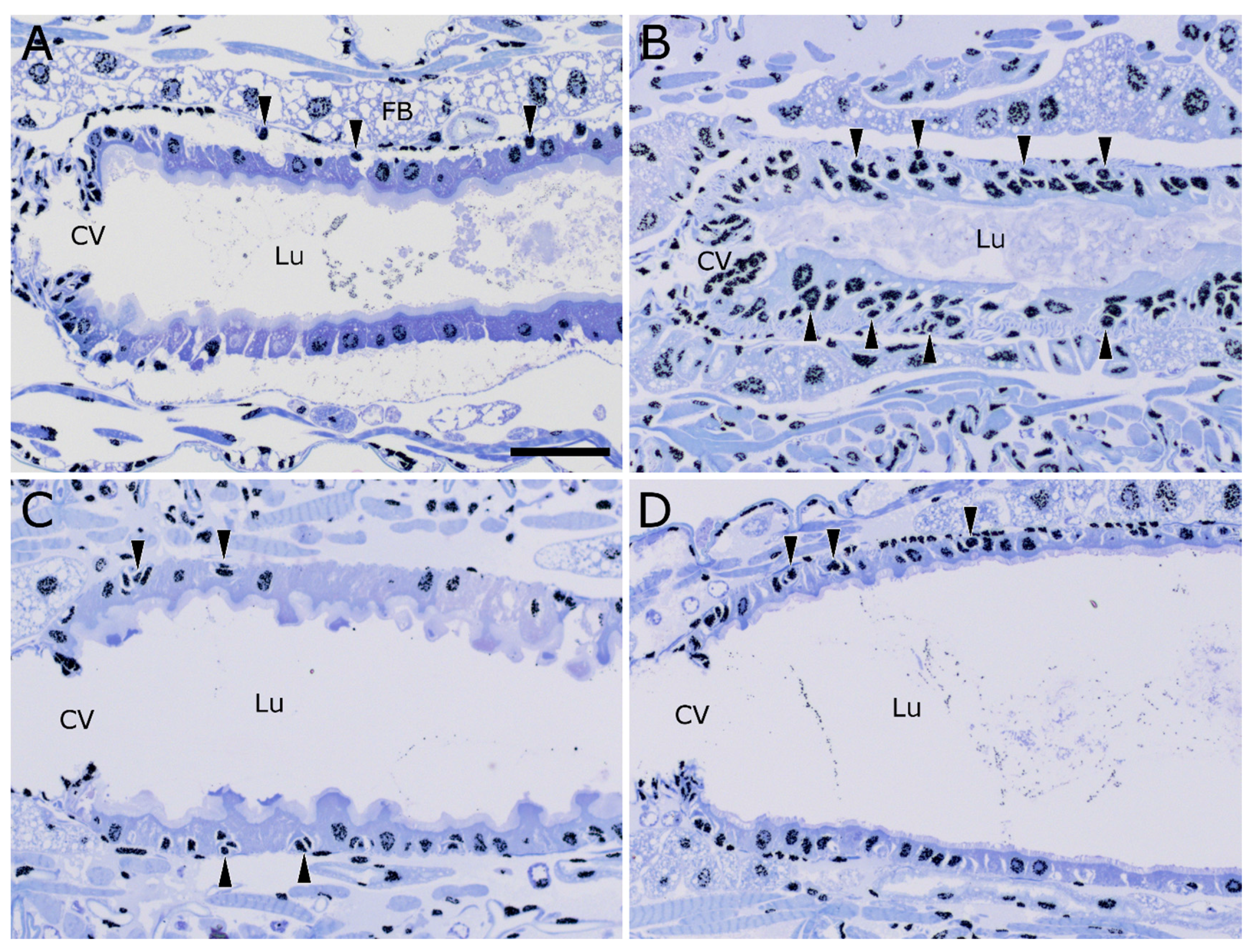
2.4. Stem Cell Activation upon Intoxication with Cry34/35Ab1
2.5. Other Insecticidal Proteins
3. Discussion
4. Materials and Methods
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gray, M.E.; Sappington, T.W.; Miller, N.J.; Moeser, J.; Bohn, M.O. Adaptation and invasiveness of western corn rootworm: Intensifying research on a worsening pest. Annu. Rev. Entomol. 2009, 54, 303–321. [Google Scholar] [CrossRef] [PubMed]
- Metcalf, R.L. Foreword. In Methods for the Study of Pest Diabrotica; Krysan, J.L., Miller, T.A., Eds.; Springer: New Youk, NY, USA, 1986; pp. vii–xv. [Google Scholar]
- Narva, K.E.; Siegfried, B.D.; Storer, N.P. Transgenic approaches to western corn rootworm control. Adv. Biochem. Eng. Biotechnol. 2013, 136, 135–162. [Google Scholar] [PubMed]
- Environmental Protection Agency (EPA). Biopesticides Registration Action Document. Bacillus thuringiensis Cry3Bb1 Protein and the Genetic Material Necessary for Its Production (Vector PV-ZMIR13L) in MON 863 Corn (OECD Unique Identifier: MON-ØØ863-5). Available online: https://www3.epa.gov/pesticides/chem_search/reg_actions/registration/decision_PC-006484_30-sep-10.pdf (accessed on 6 May 2017).
- Environmental Protection Agency (EPA). Biopesticides Registration Action Document. Modified Cry3A Protein and the Genetic Material Necessary for its Production (Via Elements of pZM26) in Event MIR604 Corn SYN-IR604-8. Available online: http://www3.epa.gov/pesticides/chem_search/reg_actions/pip/mcry3a-brad.pdf (accessed on 6 May 2017).
- Moellenbeck, D.J.; Peters, M.L.; Bing, J.W.; Rouse, J.R.; Higgins, L.S.; Sims, L.; Nevshemal, T.; Marshall, L.; Ellis, R.T.; Bystrak, P.G.; et al. Insecticidal proteins from bacillus thuringiensis protect corn from corn rootworms. Nat. Biotechnol. 2001, 19, 668–672. [Google Scholar] [CrossRef] [PubMed]
- Environmental Protection Agency (EPA). Biopesticides Registration Action Document. Bacillus thuringiensis Cry34Ab1 and Cry35Ab1 Proteins and the Genetic Material Necessary for Their Production (PHP17662 T-DNA) in Event DAS-59122-7 Corn (OECD Unique Identifier: DAS-59122-7). Available online: https://www3.epa.gov/pesticides/chem_search/reg_actions/pip/cry3435ab1-brad.pdf (accessed on 6 May 2017).
- Jakka, S.R.K.; Shrestha, R.B.; Gassmann, A.J. Broad-spectrum resistance to Bacillus thuringiensis toxins by western corn rootworm (Diabrotica virgifera virgifera). Sci. Rep. 2016. [Google Scholar] [CrossRef] [PubMed]
- Bravo, A.; Gill, S.S.; Soberon, M. Mode of action of bacillus thuringiensis cry and cyt toxins and their potential for insect control. Toxicon 2007, 49, 423–435. [Google Scholar] [CrossRef] [PubMed]
- Gahan, L.J.; Pauchet, Y.; Vogel, H.; Heckel, D.G. An abc transporter mutation is correlated with insect resistance to bacillus thuringiensis cry1ac toxin. PLoS Genet. 2010, 6, e1001248. [Google Scholar] [CrossRef] [PubMed]
- Vachon, V.; Laprade, R.; Schwartz, J.L. Current models of the mode of action of bacillus thuringiensis insecticidal crystal proteins: A critical review. J. Invertebr. Pathol. 2012, 111, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Heckel, D.G. A return to the pore - dissecting bacillus thuringiensis toxin mode of action via voltage clamp experiments. FEBS J. 2016, 283, 4458–4461. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.S.; Cowles, E.A.; Pietrantonio, P.V. The mode of action of bacillus thuringiensis endotoxins. Annu. Rev. Entomol. 1992, 37, 615–636. [Google Scholar] [CrossRef] [PubMed]
- Bravo, A.; Gomez, I.; Conde, J.; Munoz-Garay, C.; Sanchez, J.; Miranda, R.; Zhuang, M.; Gill, S.S.; Soberon, M. Oligomerization triggers binding of a bacillus thuringiensis cry1ab pore-forming toxin to aminopeptidase n receptor leading to insertion into membrane microdomains. Biochim. Biophys. Acta 2004, 1667, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Li, J.D.; Carroll, J.; Ellar, D.J. Crystal structure of insecticidal delta-endotoxin from bacillus thuringiensis at 2.5 a resolution. Nature 1991, 353, 815–821. [Google Scholar] [CrossRef] [PubMed]
- Kelker, M.S.; Berry, C.; Evans, S.L.; Pai, R.; McCaskill, D.G.; Wang, N.X.; Russell, J.C.; Baker, M.D.; Yang, C.; Pflugrath, J.W.; et al. Structural and biophysical characterization of bacillus thuringiensis insecticidal proteins cry34ab1 and cry35ab1. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Ellis, R.T.; Stockhoff, B.A.; Stamp, L.; Schnepf, H.E.; Schwab, G.E.; Knuth, M.; Russell, J.; Cardineau, G.A.; Narva, K.E. Novel bacillus thuringiensis binary insecticidal crystal proteins active on western corn rootworm, diabrotica virgifera virgifera leconte. Appl. Environ. Microbiol. 2002, 68, 1137–1145. [Google Scholar] [CrossRef] [PubMed]
- Dementiev, A.; Board, J.; Sitaram, A.; Hey, T.; Kelker, M.S.; Xu, X.; Hu, Y.; Vidal-Quist, C.; Chikwana, V.; Griffin, S.; et al. The pesticidal cry6aa toxin from bacillus thuringiensis is structurally similar to hlye-family alpha pore-forming toxins. BMC Biol. 2016, 14. [Google Scholar] [CrossRef] [PubMed]
- Sheets, J.J.; Hey, T.D.; Fencil, K.J.; Burton, S.L.; Ni, W.; Lang, A.E.; Benz, R.; Aktories, K. Insecticidal toxin complex proteins from xenorhabdus nematophilus: Structure and pore formation. J. Biol. Chem. 2011, 286, 22742–22749. [Google Scholar] [CrossRef] [PubMed]
- Blackburn, M.; Golubeva, E.; Bowen, D.; Ffrench-Constant, R.H. A novel insecticidal toxin from photorhabdus luminescens, toxin complex a (tca), and its histopathological effects on the midgut of manduca sexta. Appl. Environ. Microbiol. 1998, 64, 3036–3041. [Google Scholar] [PubMed]
- Bowen, D.; Rocheleau, T.A.; Blackburn, M.; Andreev, O.; Golubeva, E.; Bhartia, R.; ffrench-Constant, R.H. Insecticidal toxins from the bacterium photorhabdus luminescens. Science 1998, 280, 2129–2132. [Google Scholar] [CrossRef] [PubMed]
- Denolf, P.; Jansens, S.; Peferoen, M.; Degheele, D.; Van Rie, J. Two different bacillus thuringiensis delta-endotoxin receptors in the midgut brush border membrane of the european corn borer, ostrinia nubilalis (hubner) (lepidoptera: Pyralidae). Appl. Environ. Microbiol. 1993, 59, 1828–1837. [Google Scholar] [PubMed]
- BenFarhat-Touzri, D.; Saadaoui, M.; Abdelkefi-Mesrati, L.; Saadaoui, I.; Azzouz, H.; Tounsi, S. Histopathological effects and determination of the putative receptor of bacillus thuringiensis cry1da toxin in spodoptera littoralis midgut. J. Invertebr. Pathol. 2013, 112, 142–145. [Google Scholar] [CrossRef] [PubMed]
- Sass, J.E. Botanical Microtechnique, 3d ed.; Iowa State College Press: Ames, LA, USA, 1958; 228p. [Google Scholar]
- Berlyn, G.P.; Miksche, J.P.; Sass, J.E. Botanical Microtechnique and Cytochemistry, 1st ed.; Iowa State University Press: Ames, IA, USA, 1976; p. viii. 326p. [Google Scholar]
- Ruzin, S.E. Plant Microtechnique and Microscopy; Oxford University Press: New York, NY, USA, 1999; p. xi. 322p. [Google Scholar]
- Yiallouros, M.; Storch, V.; Becker, N. Impact of bacillus thuringiensis var. Israelensis on larvae of chironomus thummi thummi and psectrocladius psilopterus (diptera: Chironomidae). J. Invertebr. Pathol. 1999, 74, 39–47. [Google Scholar] [CrossRef] [PubMed]
- Koci, J.; Ramaseshadri, P.; Bolognesi, R.; Segers, G.; Flannagan, R.; Park, Y. Ultrastructural changes caused by snf7 rnai in larval enterocytes of western corn rootworm (diabrotica virgifera virgifera le conte). PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Richtman, N.M.; Zhao, J.Z.; Duncan, K.E.; Niu, X.; Procyk, L.A.; Oneal, M.A.; Kernodle, B.M.; Steimel, J.P.; Crane, V.C.; et al. Discovery of midgut genes for the rna interference control of corn rootworm. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Vaughn, K. Immunocytochemistry of Plant Cells; Springer: Berlin, Germany, 2013; pp. 1–41. [Google Scholar]
- Endo, Y.; Nishiitsutsujiuwo, J. Ultrastructural changes in the midgut epithelium of bombyx-mori l induced by bacillus-thuringiensis crystals. Jpn. J. Appl. Entomol. Zool. 1979, 23, 183–185. [Google Scholar] [CrossRef]
- Cavados, C.F.G.; Majerowicz, S.; Chaves, J.Q.; Araujo-Coutinho, C.J.P.C.; Rabinovitch, L. Histopathological and ultrastructural effects of delta-endotoxins of bacillus thuringiensis serovar israelensis in the midgut of simulium pertinax larvae (diptera, simuliidae). Mem. Inst. Oswaldo Cruz 2004, 99, 493–498. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Olson, M.; Lin, G.; Hey, T.; Tan, S.Y.; Narva, K.E. Bacillus thuringiensis cry34ab1/cry35ab1 interactions with western corn rootworm midgut membrane binding sites. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Herman, R.A.; Scherer, P.N.; Young, D.L.; Mihaliak, C.A.; Meade, T.; Woodsworth, A.T.; Stockhoff, B.A.; Narva, K.E. Binary insecticidal crystal protein from Bacillus thuringiensis, strain PS149B1: Effects of individual protein components and mixtures in laboratory bioassays. J. Econ. Entomol. 2002, 95, 635–639. [Google Scholar] [PubMed]
- Hakim, R.S.; Baldwin, K.; Smagghe, G. Regulation of midgut growth, development, and metamorphosis. Annu. Rev. Entomol. 2010, 55, 593–608. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Ramirez, A.C.; Gould, F.; Ferre, J. Histopathological effects and growth reduction in a susceptible and a resistant strain of heliothis virescens (lepidoptera: Noctuidae) caused by sublethal doses of pure cry1a crystal proteins from bacillus thuringiensis. Biocontrol Sci. Technol. 1999, 9, 239–246. [Google Scholar] [CrossRef]
- Castagnola, A.; Eda, S.; Jurat-Fuentes, J.L. Monitoring stem cell proliferation and differentiation in primary midgut cell cultures from heliothis virescens larvae using flow cytometry. Differentiation 2011, 81, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Gatsogiannis, C.; Lang, A.E.; Meusch, D.; Pfaumann, V.; Hofnagel, O.; Benz, R.; Aktories, K.; Raunser, S. A syringe-like injection mechanism in photorhabdus luminescens toxins. Nature 2013, 495, 520–523. [Google Scholar] [CrossRef] [PubMed]
- Meusch, D.; Gatsogiannis, C.; Efremov, R.G.; Lang, A.E.; Hofnagel, O.; Vetter, I.R.; Aktories, K.; Raunser, S. Mechanism of tc toxin action revealed in molecular detail. Nature 2014, 508, 61–65. [Google Scholar] [CrossRef] [PubMed]
- Blackburn, M.B.; Domek, J.M.; Gelman, D.B.; Hu, J.S. The broadly insecticidal photorhabdus luminescens toxin complex a (tca): Activity against the colorado potato beetle, leptinotarsa decemlineata, and sweet potato whitefly, bemisia tabaci. J. Insect Sci. 2005, 5. [Google Scholar] [CrossRef]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bowling, A.J.; Pence, H.E.; Li, H.; Tan, S.Y.; Evans, S.L.; Narva, K.E. Histopathological Effects of Bt and TcdA Insecticidal Proteins on the Midgut Epithelium of Western Corn Rootworm Larvae (Diabrotica virgifera virgifera). Toxins 2017, 9, 156. https://doi.org/10.3390/toxins9050156
Bowling AJ, Pence HE, Li H, Tan SY, Evans SL, Narva KE. Histopathological Effects of Bt and TcdA Insecticidal Proteins on the Midgut Epithelium of Western Corn Rootworm Larvae (Diabrotica virgifera virgifera). Toxins. 2017; 9(5):156. https://doi.org/10.3390/toxins9050156
Chicago/Turabian StyleBowling, Andrew J., Heather E. Pence, Huarong Li, Sek Yee Tan, Steven L. Evans, and Kenneth E. Narva. 2017. "Histopathological Effects of Bt and TcdA Insecticidal Proteins on the Midgut Epithelium of Western Corn Rootworm Larvae (Diabrotica virgifera virgifera)" Toxins 9, no. 5: 156. https://doi.org/10.3390/toxins9050156
APA StyleBowling, A. J., Pence, H. E., Li, H., Tan, S. Y., Evans, S. L., & Narva, K. E. (2017). Histopathological Effects of Bt and TcdA Insecticidal Proteins on the Midgut Epithelium of Western Corn Rootworm Larvae (Diabrotica virgifera virgifera). Toxins, 9(5), 156. https://doi.org/10.3390/toxins9050156




