Assessment of the Antimicrobial Activity and the Entomocidal Potential of Bacillus thuringiensis Isolates from Algeria
Abstract
:1. Introduction
2. Results
2.1. Isolation and Distribution of B. thuringiensis Isolates
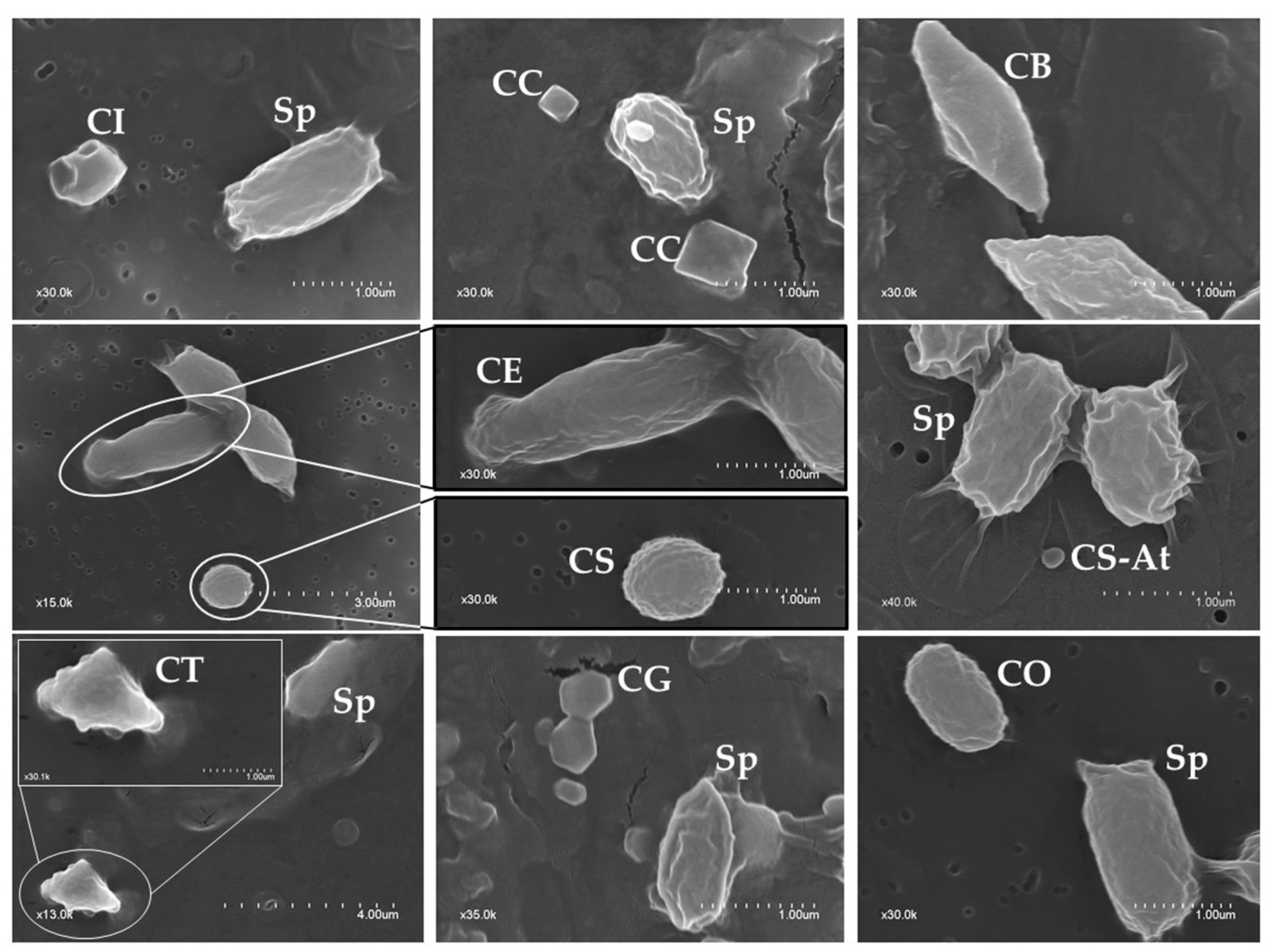
2.2. Phenotypic Characterization of Parasporal Crystals
2.3. Screening of the Biological Activity
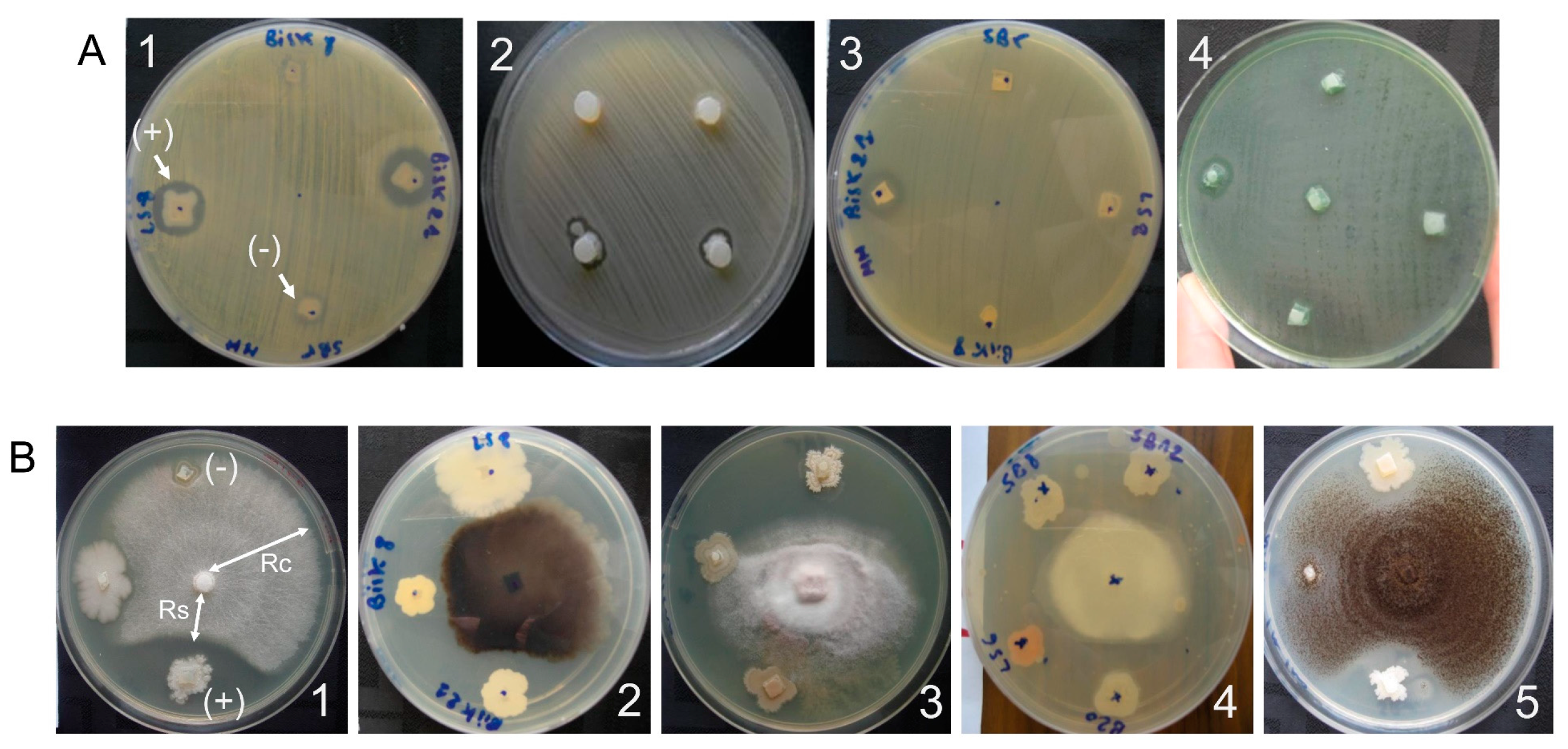
2.3.1. Antibacterial Activity
2.3.2. Antifungal Activity
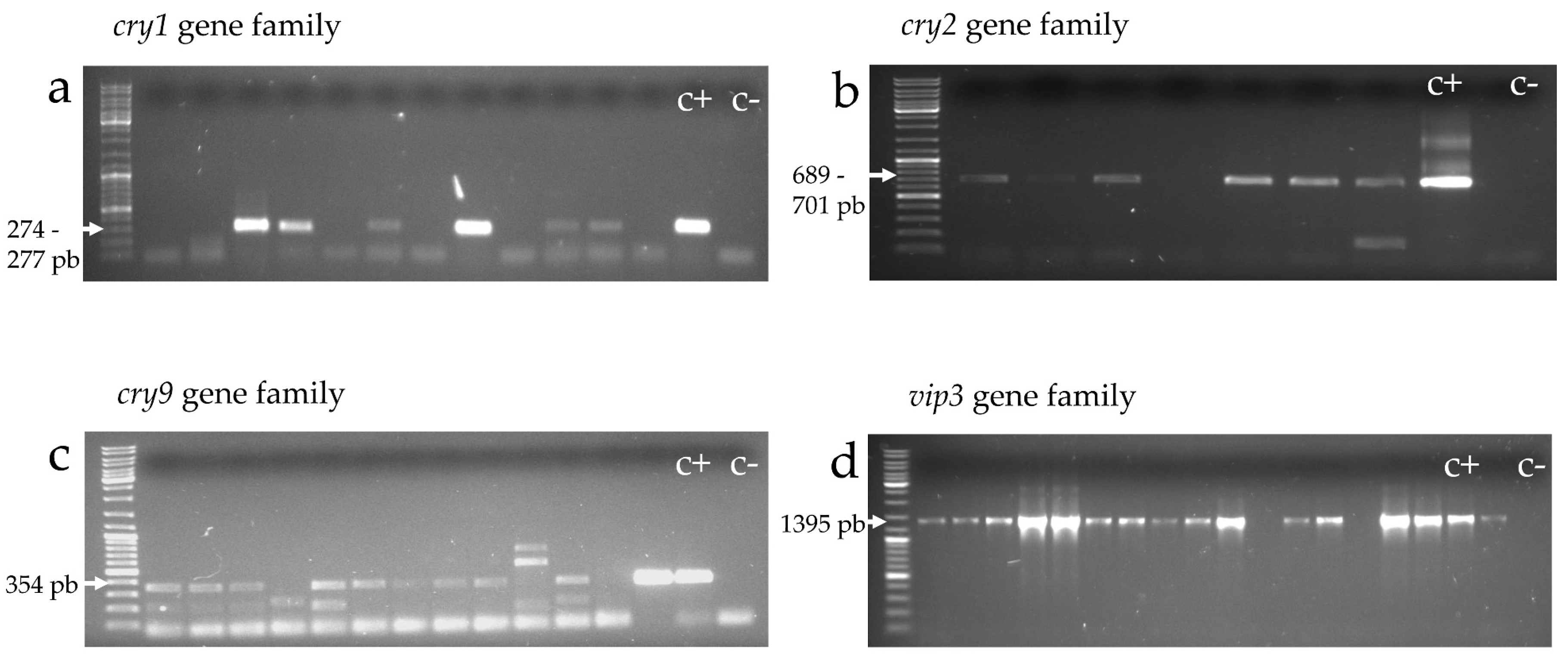
2.4. Molecular Screening
2.4.1. cry and vip Gene Families (cry1, cry2, cry9, and vip3)
2.4.2. Exochitinase (chi36) and Endochitinase (chit) Genes
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Sample Collection
5.2. Reference Strains
5.3. Bacillus Thuringiensis Culturing and Isolation
5.4. Screening for Antibacterial Activity with the Agar Plug Diffusion Method
5.5. Screening for the Antifungal Activity
5.6. DNA Extraction and PCR Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kfir, R.; Overholt, W.A.; Khan, Z.R.; Polaszek, A. Biology and management of economically important lepidopteran cereal stem borers in Africa. Annu. Rev. Entomol. 2002, 47, 701–731. [Google Scholar] [CrossRef] [PubMed]
- Midega, C.A.O.; Bruce, T.J.A.; Pickett, J.A.; Khan, Z.R. Ecological management of cereal stemborers in African smallholder agriculture through behavioural manipulation. Ecol. Entomol. 2015, 40, 70–81. [Google Scholar] [CrossRef] [PubMed]
- Gitau, C.W.; Gurr, G.M.; Dewhurst, C.F.; Fletcher, M.J.; Mitchell, A. Insect pests and insect-vectored diseases of palms. Aust. J. Entomol. 2009, 48, 328–342. [Google Scholar] [CrossRef]
- El-Shafie, H. Review: List of arthropod pests and their natural enemies identified worldwide on date palm, Phoenix dactylifera L. Agric. Biol. J. N. Am. 2012, 3, 516–524. [Google Scholar] [CrossRef]
- Meadows, M.P.; Ellis, D.J.; Butt, J.; Jarrett, P.; Burges, H.D. Distribution, frequency, and diversity of Bacillus thuringiensis in an animal feed mill. Appl. Environ. Microbiol. 1992, 58, 1344–1350. [Google Scholar] [PubMed]
- Ohba, M.; Aratake, Y. Comparative study of the frequency and flagellar serotype flora of Bacillus thuringiensis in soils and silkworm-breeding environments. J. Appl. Bacteriol. 1994, 76, 203–209. [Google Scholar] [CrossRef]
- Iriarte, J.; Porcar, M.; Lecadet, M.M.; Caballero, P. Isolation and characterization of Bacillus thuringiensis strains from aquatic environments in Spain. Curr. Microbiol. 2000, 40, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.H.; Machii, J.; Ohba, M. High frequency of Bacillus thuringiensis in feces of herbivorous animals maintained in a zoological garden in Japan. Appl. Entomol. Zool. 2002, 37, 509–516. [Google Scholar] [CrossRef]
- Hernández-Rodríguez, C.S.; Ferré, J. Ecological distribution and characterization of four collections of Bacillus thuringiensis strains. J. Basic Microbiol. 2008, 49, 152–157. [Google Scholar] [CrossRef] [PubMed]
- Seifinejad, A.; Jouzani, G.R.S.; Hosseinzadeh, A.; Abdmishani, C. Characterization of Lepidoptera-active cry and vip genes in Iranian Bacillus thuringiensis strain collection. Biol. Control. 2008, 44, 216–226. [Google Scholar] [CrossRef]
- Baig, D.N.; Mehnaz, S. Determination and distribution of cry-type genes in halophilic Bacillus thuringiensis isolates of Arabian Sea sedimentary rocks. Microbiol. Res. 2010, 165, 376–383. [Google Scholar] [CrossRef] [PubMed]
- Arora, N.; Ahmad, T.; Rajagopal, R.; Bhatnagar, R.K. A constitutively expressed 36 kDa exochitinase from Bacillus thuringiensis HD-1. Biochem. Biophys. Res. Commun. 2003, 307, 620–625. [Google Scholar] [CrossRef]
- Liu, D.; Cai, J.; Xie, C.C.; Liu, C.; Chen, Y.H. Purification and partial characterization of a 36-kDa chitinase from Bacillus thuringiensis subsp. colmeri, and its biocontrol potential. Enzym. Microb. Technol. 2010, 46, 252–256. [Google Scholar] [CrossRef]
- Lee, S.J.; Park, S.Y.; Lee, J.J.; Yum, D.Y.; Koo, B.T.; Lee, J.K. Genes encoding the N-acyl homoserine lactone-degrading enzyme are widespread in many subspecies of Bacillus thuringiensis. Appl. Environ. Microbiol. 2002, 68, 3919–3924. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Choi, Y.L.; Sun, M.; Yu, Z. Novel roles of Bacillus thuringiensis to control plant diseases. Appl. Microbiol. Biotechnol. 2008, 80, 563–572. [Google Scholar] [CrossRef] [PubMed]
- Ongena, M.; Jacques, P. Bacillus lipopeptides: Versatile weapons for plant disease biocontrol. Trends Microbiol. 2008, 16, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Abderrahmani, A.; Tapi, A.; Nateche, F.; Chollet, M.; Leclère, V.; Wathelet, B.; Hacene, H.; Jacques, P. Bioinformatics and molecular approaches to detect NRPS genes involved in the biosynthesis of kurstakin from Bacillus thuringiensis. Appl. Microbiol. Biotechnol. 2011, 92, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Ben Khedher, S.; Boukedi, H.; Dammak, M.; Kilani-Feki, O.; Sellami-Boudawara, T.; Abdelkefi-Mesrati, L.; Tounsi, S. Combinatorial effect of Bacillus amyloliquefaciens AG1 biosurfactant and Bacillus thuringiensis Vip3Aa16 toxin on Spodoptera littoralis larvae. J. Invertebr. Pathol. 2017, 144, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Broderick, N.A.; Goodman, R.M.; Raffa, K.F.; Handelsman, J. Synergy between Zwittermicin A and Bacillus thuringiensis subsp. kurstaki against gypsy moth (Lepidoptera: Lymantriidae). Environ. Entomol. 2000, 29, 101–107. [Google Scholar]
- Zhao, C.; Luo, Y.; Song, C.; Liu, Z.; Chen, S.; Yu, Z.; Sun, M. Identification of three Zwittermicin A biosynthesis-related genes from Bacillus thuringiensis subsp. kurstaki strain YBT-1520. Arch. Microbiol. 2007, 187, 313–319. [Google Scholar] [PubMed]
- Crickmore, N.; Zeigler, D.R.; Feitelson, J.; Schnepf, E.; Van Rie, J.; Lereclus, D.; Baum, J.; Dean, D.H. Revision of the nomenclature for the Bacillus thuringiensis pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 1998, 62, 807–813. [Google Scholar] [PubMed]
- Schnepf, E.; Crickmore, N.; Van Rie, J.; Lereclus, D.; Baum, J.; Feitelson, J.; Zeigler, D.R.; Dean, D.H. Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 1998, 62, 775–806. [Google Scholar] [PubMed]
- Deng, C.; Peng, Q.; Song, F.; Lereclus, D. Regulation of cry gene expression in Bacillus thuringiensis. Toxins (Basel) 2014, 6, 2194–2209. [Google Scholar] [CrossRef] [PubMed]
- Höfte, H.; Whiteley, H.R. Insecticidal crystal proteins of Bacillus thuringiensis. Microbiol. Rev. 1989, 53, 242–255. [Google Scholar] [PubMed]
- Van Frankenhuyzen, K. Insecticidal activity of Bacillus thuringiensis crystal proteins. J. Invertebr. Pathol. 2009, 101, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Bravo, A.; Likitvivatanavong, S.; Gill, S.S.; Soberón, M. Bacillus thuringiensis: A story of a successful bioinsecticide. Insect Biochem. Mol. Biol. 2011, 41, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Wang, B.C.; Yu, Z.; Sun, M. Structural insights into Bacillus thuringiensis Cry, Cyt and parasporin toxins. Toxins 2014, 6, 2732–2770. [Google Scholar] [CrossRef] [PubMed]
- Jurat-Fuentes, J.L.; Crickmore, N. Specificity determinants for Cry insecticidal proteins: Insights from their mode of action. J. Invertebr. Pathol. 2016, 142, 5–10. [Google Scholar] [CrossRef] [PubMed]
- Mendelsohn, M.; Kough, J.; Vaituzis, Z.; Matthews, K. Are Bt crops safe? Nat. Biotechnol. 2003, 21, 1003–1009. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.L.; Chen, P.H.; Pang, J.C.; Lin, C.W.; Hwang, C.F.; Tsen, H.Y. The correlation of the presence and expression levels of cry genes with the insecticidal activities against Plutella xylostella for Bacillus thuringiensis Strains. Toxins (Basel) 2014, 6, 2453–2470. [Google Scholar] [CrossRef] [PubMed]
- Monnerat, R.; Pereira, E.; Teles, B.; Martins, E.; Praça, L.; Queiroz, P.; Soberón, M.; Bravo, A.; Ramos, F.; Soares, C.M. Synergistic activity of Bacillus thuringiensis toxins against Simulium spp. larvae. J. Invertebr. Pathol. 2014, 121, 70–73. [Google Scholar] [CrossRef] [PubMed]
- Chakroun, M.; Banyuls, N.; Bel, Y.; Escriche, B.; Ferré, J. Bacterial vegetative insecticidal proteins (Vip) from entomopathogenic bacteria. Microbiol. Mol. Biol. Rev. 2016, 80, 329–350. [Google Scholar] [CrossRef] [PubMed]
- Sena, J.A.D.; Hernández-Rodríguez, C.S.; Ferré, J. Interaction of Bacillus thuringiensis Cry1 and Vip3A proteins with Spodoptera frugiperda midgut binding sites. Appl. Environ. Microbiol. 2009, 75, 2236–2237. [Google Scholar] [CrossRef] [PubMed]
- Chakroun, M.; Ferré, J. In vivo and in vitro binding of Vip3Aa to Spodoptera frugiperda midgut and characterization of binding sites by 125I radiolabeling. Appl. Environ. Microbiol. 2014, 80, 6258–6265. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Martínez, P.; Hernández-Rodríguez, C.S.; Van Rie, J.; Escriche, B.; Ferré, J. Insecticidal activity of Vip3Aa, Vip3Ad, Vip3Ae, and Vip3Af from Bacillus thuringiensis against lepidopteran corn pests. J. Invertebr. Pathol. 2013, 113, 78–81. [Google Scholar] [CrossRef] [PubMed]
- Palma, L.; de Escudero, I.R.; Maeztu, M.; Caballero, P.; Muñoz, D. Screening of vip genes from a Spanish Bacillus thuringiensis collection and characterization of two Vip3 proteins highly toxic to five lepidopteran crop pests. Biol. Control. 2013, 66, 141–149. [Google Scholar] [CrossRef]
- Lemes, A.R.N.; Davolos, C.C.; Legori, P.C.B.C.; Fernandes, O.A.; Ferré, J.; Lemos, M.V.F.; Desiderio, J.A. Synergism and antagonism between Bacillus thuringiensis Vip3A and Cry1 proteins in Heliothis virescens, Diatraea saccharalis and Spodoptera frugiperda. PLoS ONE 2014, 9, e107196. [Google Scholar] [CrossRef] [PubMed]
- Gomis-Cebolla, J.; Ruiz de Escudero, I.; Vera-Velasco, N.M.; Hernández-Martínez, P.; Hernández-Rodríguez, C.S.; Ceballos, T.; Palma, L.; Escriche, B.; Caballero, P.; Ferré, J. Insecticidal spectrum and mode of action of the Bacillus thuringiensis Vip3Ca insecticidal protein. J. Invertebr. Pathol. 2017, 142, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Ferre, J.; Van Rie, J. Biochemistry and genetics of insect resistance to Bacillus thuringiensis. Annu. Rev. Entomol. 2002, 47, 501–533. [Google Scholar] [CrossRef] [PubMed]
- Bravo, A.; Soberón, M. How to cope with insect resistance to Bt toxins? Trends Biotechnol. 2008, 26, 573–579. [Google Scholar] [CrossRef] [PubMed]
- Pardo-López, L.; Muñoz-Garay, C.; Porta, H.; Rodríguez-Almazán, C.; Soberón, M.; Bravo, A. Strategies to improve the insecticidal activity of Cry toxins from Bacillus thuringiensis. Peptides 2009, 30, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Tabashnik, B.E.; Van Rensburg, J.B.J.; Carrière, Y. Field-evolved insect resistance to Bt crops: Definition, theory, and data. J. Econ. Entomol. 2009, 102, 2011–2025. [Google Scholar] [CrossRef] [PubMed]
- Ghasemi, S.; Ahmadian, G.; Sadeghi, M.; Zeigler, D.R.; Rahimian, H.; Ghandili, S.; Naghibzadeh, N.; Dehestani, A. First report of a bifunctional chitinase/lysozyme produced by Bacillus pumilus SG2. Enzym. Microb. Technol. 2011, 48, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Hjort, K.; Presti, I.; Elväng, A.; Marinelli, F.; Sjöling, S. Bacterial chitinase with phytopathogen control capacity from suppressive soil revealed by functional metagenomics. Appl. Microbiol. Biotechnol. 2014, 98, 2819–2828. [Google Scholar] [CrossRef] [PubMed]
- El Guilli, M.; Hamza, A.; Clément, C.; Ibriz, M.; Ait Barka, E. Effectiveness of postharvest treatment with chitosan to control citrus green mold. Agriculture 2016, 6, 12. [Google Scholar] [CrossRef]
- Regev, A.; Keller, M.; Strizhov, N.; Sneh, B.; Prudovsky, E.; Chet, I.; Ginzberg, I.; Koncz-Kalman, Z.; Koncz, C.; Schell, J.; Zilberstein, A. Synergistic activity of a Bacillus thuringiensis delta-endotoxin and a bacterial endochitinase against Spodoptera littoralis larvae. Appl. Environ. Microbiol. 1996, 62, 3581–3586. [Google Scholar] [PubMed]
- Sampson, M.N.; Gooday, G.W. Involvement of chitinases of Bacillus thuringiensis during pathogenesis in insects. Microbiology 1998, 144, 2189–2194. [Google Scholar] [CrossRef] [PubMed]
- Wiwat, C.; Thaithanun, S.; Pantuwatana, S.; Bhumiratana, A. Toxicity of chitinase-producing Bacillus thuringiensis ssp. kurstaki HD-1 (G) toward Plutella xylostella. J. Invertebr. Pathol. 2000, 76, 270–277. [Google Scholar]
- Sirichotpakorn, N.; Rongnoparut, P.; Choosang, K.; Panbangred, W. Coexpression of chitinase and the cry11Aa1 toxin genes in Bacillus thuringiensis serovar israelensis. J. Invertebr. Pathol. 2001, 78, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Barboza-Corona, J.E.; Ortiz-Rodríguez, T.; de la Fuente-Salcido, N.; Bideshi, D.K.; Ibarra, J.E.; Salcedo-Hernández, R. Hyperproduction of chitinase influences crystal toxin synthesis and sporulation of Bacillus thuringiensis. Antonie Van Leeuwenhoek 2009, 96, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Lertcanawanichakul, M.; Wiwat, C.; Bhumiratana, A.; Dean, D.H. Expression of chitinase-encoding genes in Bacillus thuringiensis and toxicity of engineered B. thuringiensis subsp. aizawai toward Lymantria dispar larvae. Curr. Microbiol. 2004, 48, 175–181. [Google Scholar] [PubMed]
- Driss, F.; Rouis, S.; Azzouz, H.; Tounsi, S.; Zouari, N.; Jaoua, S. Integration of a recombinant chitinase into Bacillus thuringiensis parasporal insecticidal crystal. Curr. Microbiol. 2011, 62, 281–288. [Google Scholar] [CrossRef] [PubMed]
- Broglie, R.; Broglie, K. Chitinase gene expression in transgenic plants: A molecular approach to understanding plant defence responses. Philos. Trans. R. Soc. B Biol. Sci. 1993, 342, 265–270. [Google Scholar] [CrossRef]
- Cletus, J.; Balasubramanian, V.; Vashisht, D.; Sakthivel, N. Transgenic expression of plant chitinases to enhance disease resistance. Biotechnol. Lett. 2013, 35, 1719–1732. [Google Scholar] [CrossRef] [PubMed]
- Ben-Dov, E.; Zaritsky, A.; Dahan, E.; Barak, Z.; Sinai, R.; Manasherob, R.; Khamraev, A.; Troitskaya, E.; Dubitsky, A.; Berezina, N.; et al. Extended screening by PCR for seven cry-group genes from field-collected strains of Bacillus thuringiensis. Appl. Environ. Microbiol. 1997, 63, 4883–4890. [Google Scholar] [PubMed]
- Raddadi, N.; Belaouis, A.; Tamagnini, I.; Hansen, B.M.; Hendriksen, N.B.; Boudabous, A.; Cherif, A.; Daffonchio, D. Characterization of polyvalent and safe Bacillus thuringiensis strains with potential use for biocontrol. J. Basic Microbiol. 2009, 49, 293–303. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Rodríguez, C.S.; Boets, A.; Van Rie, J.; Ferré, J. Screening and identification of vip genes in Bacillus thuringiensis strains. J. Appl. Microbiol. 2009, 107, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Bel, Y.; Granero, F.; Alberola, T.M.; Martínez-Sebastián, M.J.; Ferré, J. Distribution, frequency and diversity of Bacillus thuringiensis in olive tree environments in Spain. Syst. Appl. Microbiol. 1997, 20, 652–658. [Google Scholar] [CrossRef]
- Vidal-Quist, J.C.; Castañera, P.; González-Cabrera, J. Diversity of Bacillus thuringiensis strains isolated from citrus orchards in Spain and evaluation of their insecticidal activity against Ceratitis capitata. J. Microbiol. Biotechnol. 2009, 19, 749–759. [Google Scholar] [PubMed]
- Alper, M.; Güneş, H.; Tatlipinar, A.; Çöl, B.; Civelek, H.S.; Özkan, C.; Poyraz, B. Distribution, occurrence of cry genes, and lepidopteran toxicity of native Bacillus thuringiensis isolated from fig tree environments in Aydän Province. Turk. J. Agric. For. 2014, 38, 898–907. [Google Scholar] [CrossRef]
- Ejiofor, A.O.; Johnson, T. Physiological and molecular detection of crystalliferous Bacillus thuringiensis strains from habitats in the South Central United States. J. Ind. Microbiol. Biotechnol. 2002, 28, 284–290. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Boets, A.; Van Rie, J.; Ren, G. Characterization of cryl, cry2, and cry9 genes in Bacillus thuringiensis isolates from China. J. Invertebr. Pathol. 2003, 82, 63–71. [Google Scholar] [CrossRef]
- DeLucca, A.J.; Simonson, J.G.; Larson, A.D. Bacillus thuringiensis distribution in soils of the United States. Can. J. Microbiol. 1981, 27, 865–870. [Google Scholar] [CrossRef] [PubMed]
- Ohba, M.; Aizawa, K. Distribution of Bacillus thuringiensis in soils of Japan. J. Invertebr. Pathol. 1986, 47, 277–282. [Google Scholar] [CrossRef]
- Ramalakshmi, A.; Udayasuriyan, V. Diversity of Bacillus thuringiensis isolated from Western Ghats of Tamil Nadu State, India. Curr. Microbiol. 2010, 61, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Asokan, R.; Mahadeva Swamy, H.M.; Thimmegowda, G.G.; Mahmood, R. Diversity analysis and characterization of Coleoptera, Hemiptera and Nematode active cry genes in native isolates of Bacillus thuringiensis. Ann. Microbiol. 2013, 64, 85–98. [Google Scholar] [CrossRef]
- Smith, R.A.; Couche, G.A. The phylloplane as a source of Bacillus thuringiensis variants. Appl. Environ. Microbiol. 1991, 57, 311–315. [Google Scholar] [PubMed]
- Mizuki, E.; Ichimatsu, T.; Hwang, S.H.; Park, Y.S.; Saitoh, H.; Higuchi, K.; Ohba, M. Ubiquity of Bacillus thuringiensis on phylloplanes of arboreous and herbaceous plants in Japan. J. Appl. Microbiol. 1999, 86, 979–984. [Google Scholar] [CrossRef]
- Maduell, P.; Callejas, R.; Cabrera, K.R.; Armengol, G.; Orduz, S. Distribution and Characterization of Bacillus thuringiensis on the phylloplane of species of piper (Piperaceae) in three altitudinal levels. Microb. Ecol. 2002, 44, 144–153. [Google Scholar] [CrossRef] [PubMed]
- Jara, S.; Maduell, P.; Orduz, S. Diversity of Bacillus thuringiensis strains in the maize and bean phylloplane and their respective soils in Colombia. J. Appl. Microbiol. 2006, 101, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Rosas-García, N.M.; Mireles-Martínez, M.; Hernández-Mendoza, J.L.; Ibarra, J.E. Screening of cry gene contents of Bacillus thuringiensis strains isolated from avocado orchards in Mexico, and their insecticidal activity towards Argyrotaenia sp. (Lepidoptera: Tortricidae) larvae. J. Appl. Microbiol. 2008, 104, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Mahadeva Swamy, H.M.; Asokan, R.; Mahmood, R.; Nagesha, S.N. Molecular characterization and genetic diversity of insecticidal crystal protein genes in native Bacillus thuringiensis isolates. Curr. Microbiol. 2013, 66, 323–330. [Google Scholar] [CrossRef] [PubMed]
- Saitoh, H.; Higuchi, K.; Mizuki, E.; Hwang, S.H.; Ohba, M. Characterization of mosquito larvicidal parasporal inclusions of a Bacillus thuringiensis serovar higo strain. J. Appl. Microbiol. 1998, 84, 883–888. [Google Scholar] [CrossRef] [PubMed]
- Aboussaid, H.; Vidal-Quist, J.C.; Oufdou, K.; El Messoussi, S.; Castañera, P.; González-Cabrera, J. Occurrence, characterization and insecticidal activity of Bacillus thuringiensis strains isolated from argan fields in Morocco. Environ. Technol. 2011, 32, 1383–1391. [Google Scholar] [CrossRef] [PubMed]
- Mahalakshmi, A.; Sujatha, K.; Kani, P.; Shenbagarathai, R. Distribution of cry and cyt genes among indigenous Bacillus thuringiensis isolates with mosquitocidal activity. Adv. Microbiol. 2012, 2, 216–226. [Google Scholar] [CrossRef]
- El-Kersh, T.A.; Ahmed, A.M.; Al-Sheikh, Y.A.; Tripet, F.; Ibrahim, M.S.; Metwalli, A.A.M. Isolation and characterization of native Bacillus thuringiensis strains from Saudi Arabia with enhanced larvicidal toxicity against the mosquito vector Anopheles gambiae (s.l.). Parasit Vectors 2016, 9, 647. [Google Scholar] [CrossRef] [PubMed]
- Ibarra, J.E.; Federici, B.A. Parasporal bodies of Bacillus thuringiensis subsp. morrisoni (PG-14) and Bacillus thuringiensis subsp. israelensis are similar in protein composition and toxicity. FEMS Microbiol. Lett. 1986, 34, 79–84. [Google Scholar] [CrossRef]
- Samasanti, W.; Tojo, A.; Aizawa, K. Insecticidal activity of bipyramidal and cuboidal inclusions of delta-endotoxin and distribution of their antigens among various strains of Bacillus thuringiensis. Agric. Biol. Chem. 1986, 50, 1731–1735. [Google Scholar] [CrossRef]
- López-Meza, J.E.; Ibarra, J.E. Characterization of a novel strain of Bacillus thuringiensis. Appl. Environ. Microbiol. 1996, 62, 1306–1310. [Google Scholar] [PubMed]
- Serfontein, S.; Logan, C.; Swanepoel, A.E.; Boelema, B.H.; Theron, D.J. A potato wilt disease in South Africa caused by Erwinia carotovora subspecies carotovora and E. chrysanthemi. Plant Pathol. 1991, 40, 382–386. [Google Scholar] [CrossRef]
- Jock, S.; Völksch, B.; Mansvelt, L.; Geider, K. Characterization of Bacillus strains from apple and pear trees in South Africa antagonistic to Erwinia amylovora. FEMS Microbiol. Lett. 2002, 211, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Djenane, Z. Criblage de Souches Autochtones de Bacillus en vue de la Mise en Evidence de Molécules Actives Présentant un Intérêt En Biotechnologie Industrielle Et Santé. Master’s Thesis, University of Science and Technology Houari Boumediene (USTHB), Bab Ezzouar, Algeria, 2012; p. 92. [Google Scholar]
- Mora, I.; Cabrefiga, J.; Montesinos, E. Cyclic lipopeptide biosynthetic genes and products, and inhibitory activity of plant associated Bacillus against phytopathogenic bacteria. PLoS ONE 2015, 10, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Yudina, T.G.; Konukhova, A.V.; Revina, L.P.; Kostina, L.I.; Zalunin, I.A.; Chestukhina, G.G. Antibacterial activity of Cry and Cyt proteins from Bacillus thuringiensis ssp. israelensis. Can. J. Microbiol. 2003, 49, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Yudina, T.G.; Brioukhanov, A.L.; Zalunin, I.A.; Revina, L.P.; Shestakov, A.I.; Voyushina, N.E.; Chestukhina, G.G.; Netrusov, A.I. Antimicrobial activity of different proteins and their fragments from Bacillus thuringiensis parasporal crystals against clostridia and archaea. Anaerobe 2007, 13, 6–13. [Google Scholar] [CrossRef] [PubMed]
- Silo-Suh, L.A.; Stabb, E.V.; Raffel, S.J.; Handelsman, J. Target range of Zwittermicin A, an aminopolyol antibiotic from Bacillus cereus. Curr. Microbiol. 1998, 37, 6–11. [Google Scholar] [PubMed]
- Béchet, M.; Caradec, T.; Hussein, W.; Abderrahmani, A.; Chollet, M.; Leclère, V.; Dubois, T.; Lereclus, D.; Pupin, M.; Jacques, P. Structure, biosynthesis, and properties of kurstakins, nonribosomal lipopeptides from Bacillus spp. Appl. Microbiol. Biotechnol. 2012, 95, 593–600. [Google Scholar] [CrossRef] [PubMed]
- El Arbi, A.; Rochex, A.; Chataigné, G.; Béchet, M.; Lecouturier, D.; Arnauld, S.; Gharsallah, N.; Jacques, P. The Tunisian oasis ecosystem is a source of antagonistic Bacillus spp. producing diverse antifungal lipopeptides. Res. Microbiol. 2016, 167, 46–57. [Google Scholar] [CrossRef] [PubMed]
- Abderrahmani, A. Identification du Mécanisme de Biosynthèse Non-Ribosomique d’un Nouveau Lipopeptide, la Kurstakine et Etude de son Influence sur le Phénotype de Souches de Bacillus thuringiensis Isolées en Algérie. Ph.D. Thesis, University of Science and Technology Houari Boumediene (USTHB), Bab Ezzouar, Algeria, 2011; p. 162. [Google Scholar]
- Roh, J.Y.; Liu, Q.; Choi, J.Y.; Wang, Y.; Shim, H.; Xu, H.G.; Choi, G.J.; Kim, J.C.; Je, Y.H. Construction of a recombinant Bacillus velezensis strain as an integrated control agent against plant diseases and insect pests. J. Microbiol. Biotechnol. 2009, 19, 1223–1229. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Roh, J.Y.; Wang, Y.; Choi, J.Y.; Tao, X.Y.; Kim, J.S.; Je, Y.H. Construction and characterisation of an antifungal recombinant Bacillus thuringiensis with an expanded host spectrum. J. Microbiol. 2012, 50, 874–877. [Google Scholar] [CrossRef] [PubMed]
- Abdalla, M.Y.; Al-Rokibah, A.; Moretti, A.; Mulè, G. Pathogenicity of toxigenic Fusarium proliferatum from date palm in Saudi Arabia. Plant Dis. 2000, 84, 321–324. [Google Scholar] [CrossRef]
- Flood, J. A review of fusarium wilt of oil palm caused by Fusarium oxysporum f. sp. elaeidis. Phytopathology 2006, 96, 660–662. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, S.K.; Asensio, L.; Monfort, E.; Gomez-Vidal, S.; Salinas, J.; López Lorca, L.; Jansson, H. Incidence of the two date palm pathogens, Thielaviopsis paradoxa and T. punctulata in soil from date palm plantations in Elx, South-East Spain. J. Plant Prot. Res. 2009, 49, 276–279. [Google Scholar] [CrossRef]
- Saeed, E.E.; Sham, A.; El-Tarabily, K.; Abu-Elsamen, F.; Iratni, R.; AbuQamar, S.F. Chemical control of black scorch disease on date palm caused by the fungal pathogen Thielaviopsis punctulata in United Arab Emirates. Plant Dis. 2016, 100, 2370–2376. [Google Scholar] [CrossRef]
- Yang, F.; Jacobsen, S.; Jorgensen, H.J.; Collinge, D.B.; Svensson, B.; Finnie, C. Fusarium graminearum and its interactions with cereal heads: Studies in the proteomics era. Front. Plant Sci. 2013, 4, 37. [Google Scholar] [CrossRef] [PubMed]
- Lazreg, F.; Belabid, L.; Sanchez, J.; Gallego, E.; Garrido-Cardenas, J.A.; Elhaitoum, A. First report of Fusarium redolens as a causal agent of Aleppo pine damping-off in Algeria. Plant Dis. 2013, 97, 997. [Google Scholar] [CrossRef]
- Lazreg, F.; Belabid, L.; Sánchez, J.; Gallego, E. Root rot and damping-off of Aleppo pine seedlings caused by Pythium spp. in Algerian forest nurseries. J. For. Sci. 2016, 62, 322–328. [Google Scholar] [CrossRef]
- Mohammed, A.S.; Kadar, N.H.; Kihal, M.; Henni, J.E.; Sanchez, J.; Gallego, E.; Garrido-cardenas, J.A.; Ahmed, O.; Bella, B.; Naouer, E.M.; et al. Characterization of Fusarium oxysporum isolates from tomato plants in Algeria. Afr. J. Microb. Res. 2016, 10, 1156–1163. [Google Scholar]
- Zemouli-Benfreha, F.; Djamel-eddine, H. Fusarium wilt of chickpea (Cicer arietinum L.) in North-West Algeria. Afr. J. 2014, 9, 168–175. [Google Scholar] [CrossRef]
- Thammasittirong, A.; Attathom, T. PCR-based method for the detection of cry genes in local isolates of Bacillus thuringiensis from Thailand. J. Invertebr. Pathol. 2008, 98, 121–126. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Wang, J.; Yin, W.; Shao, X.; Zheng, H.; Li, M.; Zhao, Y.; Sun, M.; Wang, S.; Yu, Z. Complete genome sequence of Bacillus thuringiensis subsp. chinensis strain CT-43. J. Bacteriol. 2011, 193, 3407–3408. [Google Scholar] [PubMed]
- Zhu, Y.; Shang, H.; Zhu, Q.; Ji, F.; Wang, P.; Fu, J.; Deng, Y.; Xu, C.; Ye, W.; Zheng, J.; et al. Complete genome sequence of Bacillus thuringiensis serovar finitimus strain YBT-020. J. Bacteriol. 2011, 193, 2379–2380. [Google Scholar] [CrossRef] [PubMed]
- Guan, P.; Ai, P.; Dai, X.; Zhang, J.; Xu, L.; Zhu, J.; Li, Q.; Deng, Q.; Li, S.; Wang, S.; et al. Complete genome sequence of Bacillus thuringiensis serovar sichuansis strain MC28. J. Bacteriol. 2012, 194, 6975. [Google Scholar] [CrossRef] [PubMed]
- Murawska, E.; Fiedoruk, K.; Bideshi, D.K.; Swiecicka, I. Complete genome sequence of Bacillus thuringiensis subsp. thuringiensis strain IS5056, an isolate highly toxic to Trichoplusia ni. Genome Announc. 2013, 1, e00108–e10013. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Peng, D.; Wang, Y.; Ye, W.; Zheng, J.; Zhao, C.; Han, D.; Geng, C.; Ruan, L.; He, J.; et al. Genomic and transcriptomic insights into the efficient entomopathogenicity of Bacillus thuringiensis. Sci. Rep. 2015, 4, 4585936. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Zheng, A.; Zhu, J.; Wang, S.; Wang, L.; Deng, Q.; Li, S.; Liu, H.; Li, P. Characterization of vegetative insecticidal protein vip genes of Bacillus thuringiensis from Sichuan Basin in China. Curr. Microbiol. 2011, 62, 752–757. [Google Scholar] [CrossRef] [PubMed]
- Travers, R.S.; Martin, P.A.W.; Reichelderfer, C.F. Selective process for efficient isolation of soil Bacillus spp. Appl. Environ. Microbiol. 1987, 53, 1263–1266. [Google Scholar] [PubMed]
- Paik, D.H.; Bae, S.S.; Park, H.S.; Pan, G.J. Identification and partial characterization of tochicin, a bacteriocin produced by Bacillus thuringiensis subsp tochigiensis. J. Ind. Microbiol. Biotechnol. 1997, 19, 294–298. [Google Scholar] [CrossRef] [PubMed]
- Jiménez-Esquilín, A.E.; Roane, T.M. Antifungal activities of actinomycete strains associated with high-altitude sagebrush rhizosphere. J. Ind. Microbiol. Biotechnol. 2005, 32, 378–381. [Google Scholar] [CrossRef] [PubMed]
- Stephen, J.; Cavalieri, S.J.; Ronald, J.; Harbeck, R.J.; McCarter, Y.S.; Ortez, J.H.; Rankin, I.D.; Sautter, R.L.; Sharp, S.E.; Spiegel, C.A. Manual of Antimicrobial Susceptibility Testing; Marie, B., Coyle, M.B., Eds.; American Society for Microbiology: Washington, DC, USA, 2005; pp. 25–52. [Google Scholar]
- Knaak, N.; Rohr, A.A.; Fiuza, L.M. In vitro effect of Bacillus thuringiensis strains and Cry proteins in phytopathogenic fungi of paddy rice-field. Braz. J. Microbiol. 2007, 38, 526–530. [Google Scholar] [CrossRef]
- Ferrandis, M.D.; Juárez-Pérez, V.M.; Frutos, R.; Bel, Y.; Ferré, J. Distribution of cryl, cryll and cryV Genes within Bacillus thuringiensis isolates from Spain. Syst. Appl. Microbiol. 1999, 22, 179–185. [Google Scholar] [CrossRef]
- Crickmore, N.; Zeigler, D.R.; Schnepf, E.; Van Rie, J.; Lereclus, D.; Baum, J.; Bravo, A.; Dean, D.H. Bacillus thuringiensis Toxin Nomenclature. Available online: http://www.lifesci.sussex.ac.uk/home/Neil_Crickmore/Bt/vip.html (accessed on 1 November 2016).


| Source of Samples | Samples | Mediterranean Area | Semi-Arid Area | Desert | Global Bt Index | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. of Isolates | Bt Index d | No. of Isolates | Bt Index d | No. of Isolates | Bt Index d | |||||||
| Total Analyzed | Bt Positive a | Bacillus-Like b | Bt c | Bacillus-Like b | Bt c | Bacillus-Like b | Bt c | |||||
| Telluric (soil) | ||||||||||||
| Rhizospheric | 18 | 18 | 68 | 35 | 0.51 | 39 | 14 | 0.36 | 77 | 39 | 0.51 | 0.48 |
| Non rhizospheric | 10 | 8 | 11 | 4 | 0.36 | 12 | 2 | 0.17 | 43 | 12 | 0.28 | 0.27 |
| Non telluric | ||||||||||||
| Sediment | 3 | 2 | 13 | 5 | 0.38 | 0 | 0 | / | 1 | 0 | 0 | 0.36 |
| Dead insects | 4 | 4 | 28 | 10 | 0.36 | 0 | 0 | / | 0 | 0 | / | 0.36 |
| Grain storage | 19 | 19 | 62 | 26 | 0.42 | 31 | 10 | 0.32 | 0 | 0 | / | 0.39 |
| Total | 54 | 51 | 182 | 80 | 0.44 e | 82 | 26 | 0.32 e | 121 | 51 | 0.42 e | 0.41 f |
| Crystal Shape | No. of Isolates Containing Crystals with a Given Shape | ||
|---|---|---|---|
| Alone | Combined with Other Crystals | Total (%) | |
| Spherical | 30 | 58 | 88 (64.2%) |
| Bipyramidal | 4 | 42 | 46 (33.6%) |
| Irregular/Geometrical | 19 | 36 | 55 (40.1%) |
| Triangular | 2 | 16 | 18 (13.1%) |
| Cuboidal | 0 | 16 | 16 (11.7%) |
| Ovoid | 2 | 8 | 10 (7.3%) |
| Elongate | 0 | 5 | 5 (3.6%) |
| Spectrum of Activity | Gram Positive a | Gram Negative b | |||
|---|---|---|---|---|---|
| SaSM | SaRM | Ec | Pa | n c | |
| Against both Gram positive and Gram negative pathogenic bacteria (n = 20) | + | + | + | + | 3 |
| + | + | + | − | 9 | |
| + | + | − | + | 4 | |
| + | − | + | − | 2 | |
| − | + | + | − | 2 | |
| Against Gram positive pathogenic bacteria (n = 14) | + | + | − | − | 7 |
| + | − | − | − | 5 | |
| − | + | − | − | 2 | |
| Against Gram negative pathogenic bacteria (n = 7) | − | − | + | − | 4 |
| − | − | − | + | 3 | |
| Total Bt isolates positive for each bacterium type | 30 | 27 | 20 | 10 | |
| Spectrum of Activity | Fusarium sp. | Monilia sp. | Colletotricum sp. | Thielaviopsis sp. | Aspergilus flavus | n a |
|---|---|---|---|---|---|---|
| Against five fungi (n = 24) | + | + | + | + | + | 24 |
| Against four fungi (n = 32) | + | + | + | + | − | 4 |
| + | + | + | − | + | 5 | |
| + | + | − | + | + | 9 | |
| + | − | + | + | + | 4 | |
| − | + | + | + | + | 10 | |
| Against three fungi (n = 25) | + | + | − | − | + | 2 |
| − | − | + | + | + | 2 | |
| + | + | − | + | − | 3 | |
| − | + | + | + | − | 2 | |
| − | + | − | + | + | 2 | |
| + | − | − | + | + | 1 | |
| − | + | + | − | + | 12 | |
| + | − | + | − | + | 1 | |
| Against two fungi (n = 27) | + | + | − | − | − | 1 |
| − | + | − | − | + | 5 | |
| − | + | + | − | − | 1 | |
| − | − | + | + | − | 3 | |
| − | − | − | + | + | 1 | |
| − | − | + | − | + | 16 | |
| Against one fungus (n = 27) | − | + | − | − | − | 1 |
| − | − | + | − | − | 14 | |
| − | − | − | − | + | 12 | |
| Total Bt isolates positive for each fungus type | 54 | 81 | 98 | 65 | 106 |
| Target Gene Family | Product Size (pb) | Primers Set | Sequence (5′ → 3′) | Tm a (°C) | Reference |
|---|---|---|---|---|---|
| cry1 | 274–277 | Un1(f) | CATGATTCATGCGGCAGATAAAC | 67.2 | [55] |
| Un1(r) | TTGTGACACTTCTGCTTCCCATT | 66.7 | |||
| cry2 | 689–701 | Un2(f) | GTTATTCTTAATGCAGATGAATGGG | 63.3 | [55] |
| Un2(r) | CGGATAAAATAATCTGGGAAATAGT | 61.1 | |||
| cry9 | 354 | Un9(f) | CGGTGTTACTATTAGCGAGGGCGG | 71.5 | [55] |
| Un9(r) | GTTTGAGCCGCTTCACAGCAATCC | 73.3 | |||
| endochitinase | 1997 | Chit(f) | ATTCACACTGCTATTACTATC | 50 | [56] |
| Chit(r) | TGACGGCATTTAAAAGTTCGGC | 68.7 | |||
| exochitinase 36 | 1083 | Chi36(f) | GATGTTAAACAGGTTCAA | 50.2 | [12] |
| Chi36(r) | TTATTTTTGCAAGGAAAG | 52.9 | |||
| vip3 | 1395 | vip3-sc(f) | TGCCACTGGTATCAARGA | 54.2 | [57] |
| vip3-scII(r) | CCATTAATYGGAKTCAAAAATGTTTCACTGAT | 71.1 | The current work |
| Presence/Absence of cry Gene Families | No. of Bt for Each cry Gene Profile | No. of Bt with a vip3 Gene | No. of Bt with Both chi36 and chit | No. of Bt with chi36 Only | No. of Bt with chit Only | No. of Bt without chi36 and chit |
|---|---|---|---|---|---|---|
| I. One cry gene family | ||||||
| cry1 | 10 | 6 | 1 | 0 | 3 | 6 |
| cry2 | 12 | 10 | 1 | 1 | 2 | 8 |
| cry9 | 12 | 6 | 11 | 0 | 0 | 1 |
| II. Two cry gene families | ||||||
| cry1 + cry2 | 21 | 18 | 6 | 0 | 5 | 10 |
| cry1 ± cry9 | 8 | 5 | 4 | 0 | 3 | 1 |
| cry2 + cry9 | 14 | 8 | 11 | 1 | 2 | 0 |
| III. Three cry gene families | ||||||
| cry1 + cry2 + cry9 | 35 | 29 | 15 | 4 | 4 | 12 |
| IV. No cry gene | 25 | 13 | 11 | 0 | 3 | 11 |
| Total Bt isolates (%) | 137 | 95 (69.3%) | 60 (43.8%) | 6 (4.4%) | 22 (16.1%) | 49 (35.8%) |
| Spectrum of the Antifungal Activity | Profile of Chitinase Genes | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Both chi36 and chit a | Only chi36 | Only chit | None | ||||||
| N | n | x | n | x | n | x | n | x | |
| Activity against at least three fungi | 81 | 50 | 0.62 | 5 | 0.06 | 4 | 0.05 | 22 | 0.27 |
| Activity against one or two fungi | 54 | 10 | 0.19 | 1 | 0.02 | 16 | 0.30 | 27 | 0.5 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Djenane, Z.; Nateche, F.; Amziane, M.; Gomis-Cebolla, J.; El-Aichar, F.; Khorf, H.; Ferré, J. Assessment of the Antimicrobial Activity and the Entomocidal Potential of Bacillus thuringiensis Isolates from Algeria. Toxins 2017, 9, 139. https://doi.org/10.3390/toxins9040139
Djenane Z, Nateche F, Amziane M, Gomis-Cebolla J, El-Aichar F, Khorf H, Ferré J. Assessment of the Antimicrobial Activity and the Entomocidal Potential of Bacillus thuringiensis Isolates from Algeria. Toxins. 2017; 9(4):139. https://doi.org/10.3390/toxins9040139
Chicago/Turabian StyleDjenane, Zahia, Farida Nateche, Meriam Amziane, Joaquín Gomis-Cebolla, Fairouz El-Aichar, Hassiba Khorf, and Juan Ferré. 2017. "Assessment of the Antimicrobial Activity and the Entomocidal Potential of Bacillus thuringiensis Isolates from Algeria" Toxins 9, no. 4: 139. https://doi.org/10.3390/toxins9040139
APA StyleDjenane, Z., Nateche, F., Amziane, M., Gomis-Cebolla, J., El-Aichar, F., Khorf, H., & Ferré, J. (2017). Assessment of the Antimicrobial Activity and the Entomocidal Potential of Bacillus thuringiensis Isolates from Algeria. Toxins, 9(4), 139. https://doi.org/10.3390/toxins9040139







