Influence of DPH1 and DPH5 Protein Variants on the Synthesis of Diphthamide, the Target of ADPRibosylating Toxins
Abstract
:1. Introduction
2. Results
2.1. Transient Expression of DPH1 and DPH5 cDNA in DPH Deficient MCF7 Cells
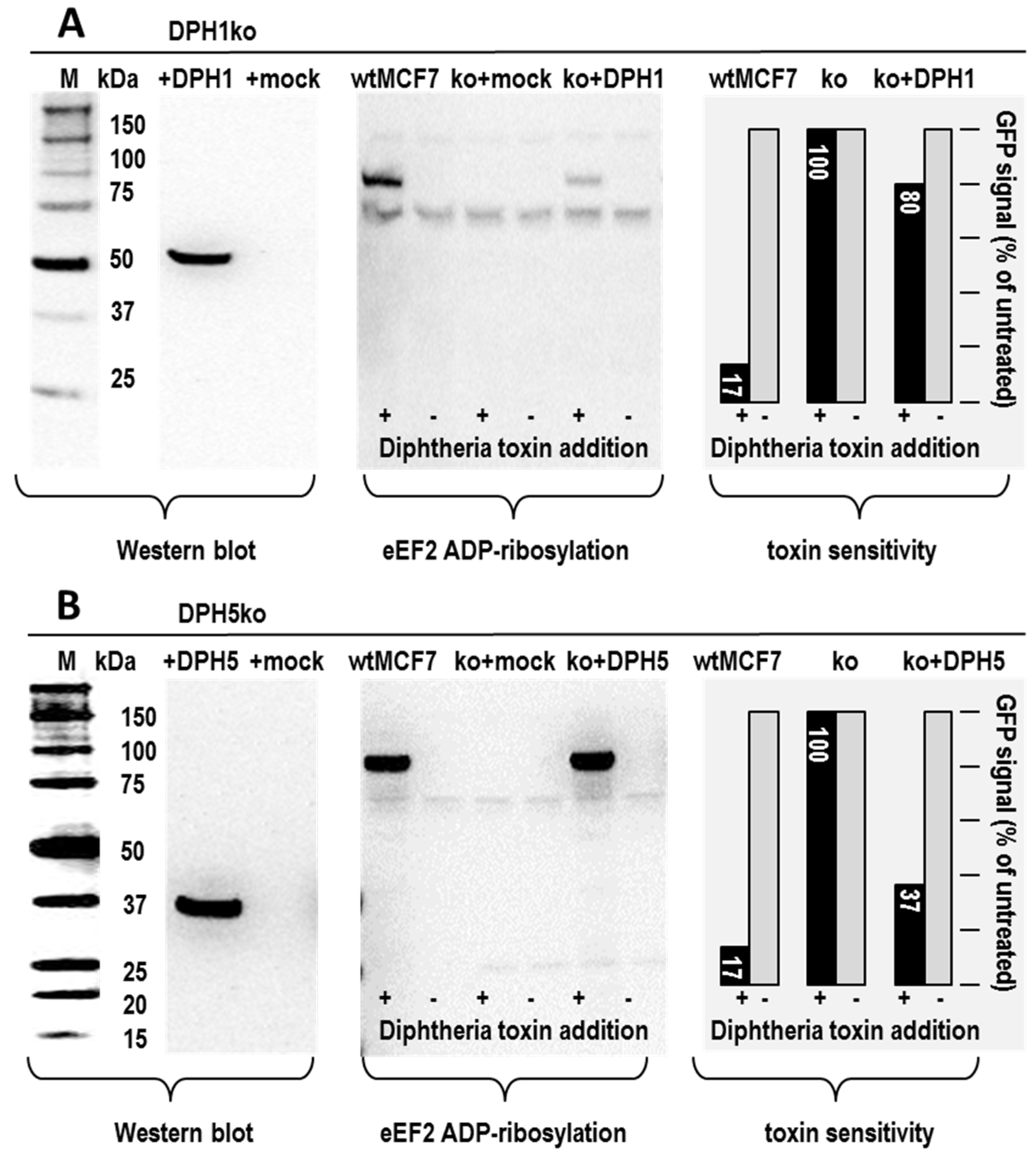
2.2. Plasmid Encoded DPH Proteins Restore Diphthamide Synthesis and Toxin Sensitivity in DPH1 or DPH5 Deficient Cells
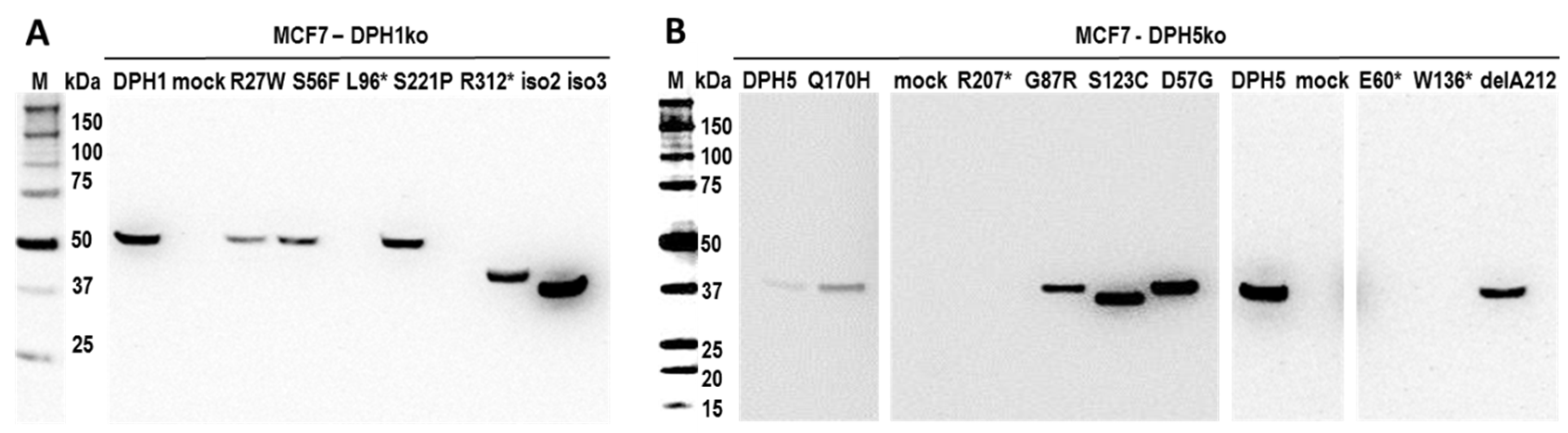
2.3. DPH1 and DPH5 Variants
2.4. Activity of DPH1 and DPH5 Protein Variants
3. Discussion
4. Materials and Methods
4.1. MCF7 Variants with Inactivated DPH Genes
4.2. Expression of DPH Proteins
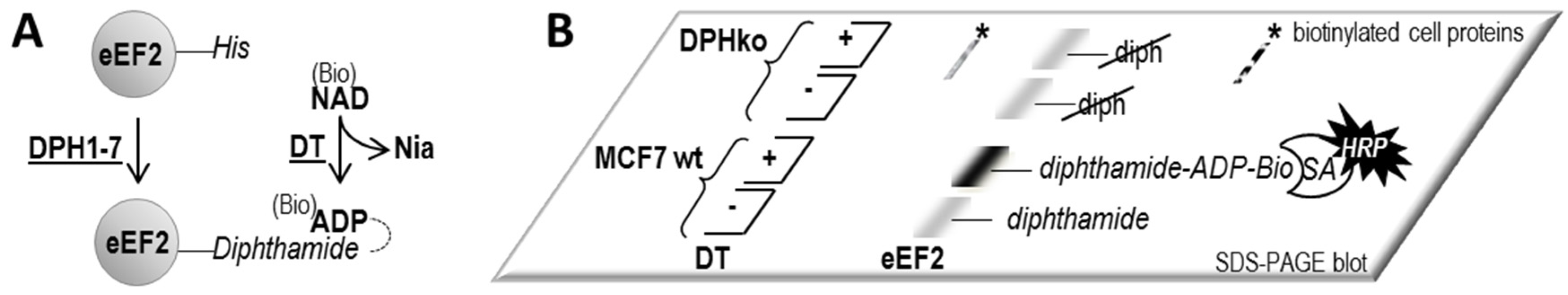
4.3. ADP-Ribosylation of eEF2
4.4. Cytotoxicity Assays
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Stahl, S.; Mueller, F.; Pastan, I.; Brinkmann, U. Factors that Determine Sensitivity and Resistances of Tumor Cells towards Antibody-Targeted Protein Toxins. In Resistance to Immunotoxins in Cancer Therapy; Verma, R.S., Bonavida, B., Eds.; Springer International Publishing: Cham, Switzerland, 2015. [Google Scholar]
- Abdel-Fattah, W.; Scheidt, V.; Uthman, S.; Stark, M.J.R.; Schaffrath, R. Insights into diphthamide, key diphtheria toxin effector. Toxins 2003, 5, 958–968. [Google Scholar] [CrossRef] [PubMed]
- Collier, R.J. Understanding the mode of action of diphtheria toxin: A perspective on progress during the 20th century. Toxicon Off. J. Int. Soc. Toxinol. 2001, 39, 1793–1803. [Google Scholar] [CrossRef]
- Liu, S.; Milne, G.T.; Kuremsky, J.G.; Fink, G.R.; Leppla, S.H. Identification of the proteins required for biosynthesis of diphthamide, the target of bacterial ADP-ribosylating toxins on translation elongation factor 2. Mol. Cell. Biol. 2004, 24, 9487–9497. [Google Scholar] [CrossRef] [PubMed]
- Shapira, A.; Benhar, I. Toxin-Based Therapeutic Approaches. Toxins 2010, 2, 2519–2583. [Google Scholar] [CrossRef] [PubMed]
- Pastan, I.; Hassan, R.; FitzGerald, D.J.; Kreitman, R.J. Immunotoxin therapy of cancer, Nature reviews. Cancer 2006, 6, 559–565. [Google Scholar] [PubMed]
- FitzGerald, D.J.; Wayne, A.S.; Kreitman, R.J.; Pastan, I. Treatment of hematologic malignancies with immunotoxins and antibody-drug conjugates. Cancer Res. 2011, 71, 6300–6309. [Google Scholar] [CrossRef] [PubMed]
- Hassan, R.; Alewine, C.; Pastan, I. New Life for Immunotoxin Cancer Therapy. Clin. Cancer Res. 2016, 22, 1055–1058. [Google Scholar] [CrossRef] [PubMed]
- Wayne, A.S.; FitzGerald, D.J.; Kreitman, R.J.; Pastan, I. Immunotoxins for leukemia. Blood 2014, 123, 2470–2477. [Google Scholar] [CrossRef] [PubMed]
- Weidle, U.H.; Tiefenthaler, G.; Schiller, C.; Weiss, E.H.; Georges, G.; Brinkmann, U. Prospects of bacterial and plant protein-based immunotoxins for treatment of cancer. Cancer Genom. Proteom. 2014, 11, 25–38. [Google Scholar]
- Antignani, A.; FitzGerald, D. Immunotoxins: The Role of the Toxin. Toxins 2013, 5, 1486–1502. [Google Scholar] [CrossRef] [PubMed]
- Bryan, D. Fleming and Mitchell Ho. Glypican-3 Targeting Immunotoxins for the Treatment of Liver Cancer. Toxins 2016, 8. [Google Scholar] [CrossRef]
- Wei, H.; Bera, T.K.; Wayne, A.S.; Xiang, L.; Colantonio, S.; Chertov, O.; Pastan, I. A modified form of diphthamide causes immunotoxin resistance in a lymphoma cell line with a deletion of the WDR85 gene. J. Boil. Chem. 2013, 288, 12305–12312. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.; Xiang, L.; Wayne, A.S.; Chertov, O.; FitzGerald, D.J.; Bera, T.K.; Pastan, I. Immunotoxin resistance via reversible methylation of the DPH4 promoter is a unique survival strategy. Proc. Natl. Acad. Sci. USA 2012, 109, 6898–6903. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Wei, H.; Xiang, L.; Chertov, O.; Wayne, A.S.; Bera, T.K.; Pastan, I. Methylation of the DPH1 promoter causes immunotoxin resistance in acute lymphoblastic leukemia cell line KOPN-8. Leuk. Res. 2013, 37, 1551–1556. [Google Scholar] [CrossRef] [PubMed]
- Schaffrath, R.; Abdel-Fattah, W.; Klassen, R.; Stark, M.J.R. The diphthamide modification pathway from Saccharomyces cerevisiae—Revisited. Mol. Microbiol. 2014, 94, 1213–1226. [Google Scholar] [CrossRef] [PubMed]
- Su, X.; Lin, Z.; Lin, H. The biosynthesis and biological function of diphthamide. Crit. Rev. Biochem. Mol. Biol. 2013, 48, 515–521. [Google Scholar] [CrossRef] [PubMed]
- Mattheakis, L.C.; Shen, W.H.; Collier, R.J. DPH5, a methyltransferase gene required for diphthamide biosynthesis in Saccharomyces cerevisiae. Mol. Cell. Biol. 1992, 12, 4026–4037. [Google Scholar] [CrossRef] [PubMed]
- Dong, M.; Su, X.; Dzikovski, B.; Dando, E.E.; Zhu, X.; Du, J.; Freed, J.H.; Lin, H. Dph3 is an electron donor for Dph1-Dph2 in the first step of eukaryotic diphthamide biosynthesis. J. Am. Chem. Soc. 2014, 136, 1754–1757. [Google Scholar] [CrossRef] [PubMed]
- Nobukuni, Y.; Kohno, K.; Miyagawa, K. Gene trap mutagenesis-based forward genetic approach reveals that the tumor suppressor OVCA1 is a component of the biosynthetic pathway of diphthamide on elongation factor 2. J. Biol. Chem. 2005, 280, 10572–10577. [Google Scholar] [CrossRef] [PubMed]
- Webb, T.R.; Cross, S.H.; McKie, L.; Edgar, R.; Vizor, L.; Harrison, J.; Peters, J.; Jackson, I.J. Diphthamide modification of eEF2 requires a J-domain protein and is essential for normal development. J. Cell Sci. 2008, 121, 3140–3145. [Google Scholar] [CrossRef] [PubMed]
- Stahl, S.; Seidl, D.M.; Rita, A.; Ducret, A.; Geijtenbeek, S.V.; Michel, S.; Racek, T.; Birzele, F.; Haas, A.K.; Rueger, R.; et al. Loss of diphthamide pre-activates NF-κB and death receptor pathways and renders MCF7 cells hypersensitive to tumor necrosis factor. Proc. Natl. Acad. Sci. USA 2015, 112, 10732–10737. [Google Scholar] [CrossRef] [PubMed]
- Bruening, W.; Prowse, A.H.; Schultz, D.C.; Holgado-Madruga, M.; Wong, A.; Godwin, A.K. Expression of OVCA1, a Candidate Tumor Suppressor, Is Reduced in Tumors and Inhibits Growth of Ovarian Cancer Cells. Cancer Res. 1999, 59, 4973–4983. [Google Scholar] [PubMed]
- Chen, C.-M.; Behringer, R.R. Ovca1 regulates cell proliferation, embryonic development, and tumorigenesis. Genes Dev. 2004, 18, 320–332. [Google Scholar] [CrossRef] [PubMed]
- Apweiler, R.; Bairoch, A.; Wu, C.; Barker, W.; Boeckmann, B.; Ferro, S.; Gasteiger, E.; HUnag, H.; Lopez, R.; Magrane, M.; et al. HThe UniProt Consortium. UniProt: The universal protein knowledgebase. Nucl. Acids Res. 2016. [Google Scholar] [CrossRef]
- Forbes, S.A.; Beare, D.; Boutselakis, H.; Bamford, S.; Bindal, N.; Tate, J.; Cole, C.G.; Ward, S.; Dawson, E.; Ponting, L.; et al. COSMIC: somatic cancer genetics at high-resolution. Nucl. Acids Res. 2016. [Google Scholar] [CrossRef] [PubMed]
- Sherry, S.T.; Ward, M.-H.; Kholodov, M.; Baker, J.; Phan, L.; Smigielski, E.M.; Sirotkin, K. dbSNP: The NCBI database of genetic variation. Nucl. Acids Res. 2001, 29, 308–311. [Google Scholar] [CrossRef] [PubMed]

| Protein | Variant | Type | Source | Expression * | Phenotype |
|---|---|---|---|---|---|
| DPH1 | - | reference sequence (wt) | Uniprot Q9BZG8-1 | +++ | functional |
| isotype 2 | N-term deletion (1-80) | Uniprot Q9BZG8-2 | ++ | loss of function | |
| isotype 3 | N-term deletion (1-140) | Uniprot Q9BZG8-3 | +++ | loss of function | |
| R27W | missense | COSM 178273 | + | functional | |
| S56F | missense | COSM 389195 | + | functional | |
| S221P | missense | COSM 1381407* | ++ | reduced function | |
| L96 * | frameshift-stop | COSM 190903 | ? | loss of function | |
| R312 * | nonsense-stop | COSM 137079 | ? | reduced function | |
| DPH5 | - | reference sequence (wt) | Uniprot Q9H2P9-1 | +-+++ | functional |
| del A212 | single aa deletion isoform | Uniprot Q9H2P9-6 | ++ | functional | |
| D57G | missense | COSM 893022 | +++ | functional | |
| G87R | missense | dbSNP rs376902046 | ++ | functional | |
| S123C | missense | COSM 4792156 | +++ | functional | |
| Q170H | missense | COSM 674720 | + | functional | |
| E60 * | nonsense-stop | COSM 260571 | ? | loss of function | |
| W136 * | frameshift-stop | COSM 5002373 | ? | loss of function | |
| R207 * | nonsense-stop | COSM 3417782 | ? | loss of function |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mayer, K.; Schröder, A.; Schnitger, J.; Stahl, S.; Brinkmann, U. Influence of DPH1 and DPH5 Protein Variants on the Synthesis of Diphthamide, the Target of ADPRibosylating Toxins. Toxins 2017, 9, 78. https://doi.org/10.3390/toxins9030078
Mayer K, Schröder A, Schnitger J, Stahl S, Brinkmann U. Influence of DPH1 and DPH5 Protein Variants on the Synthesis of Diphthamide, the Target of ADPRibosylating Toxins. Toxins. 2017; 9(3):78. https://doi.org/10.3390/toxins9030078
Chicago/Turabian StyleMayer, Klaus, Anna Schröder, Jerome Schnitger, Sebastian Stahl, and Ulrich Brinkmann. 2017. "Influence of DPH1 and DPH5 Protein Variants on the Synthesis of Diphthamide, the Target of ADPRibosylating Toxins" Toxins 9, no. 3: 78. https://doi.org/10.3390/toxins9030078
APA StyleMayer, K., Schröder, A., Schnitger, J., Stahl, S., & Brinkmann, U. (2017). Influence of DPH1 and DPH5 Protein Variants on the Synthesis of Diphthamide, the Target of ADPRibosylating Toxins. Toxins, 9(3), 78. https://doi.org/10.3390/toxins9030078






