Mass Spectrometric Detection of Bacterial Protein Toxins and Their Enzymatic Activity
Abstract
:1. Introduction
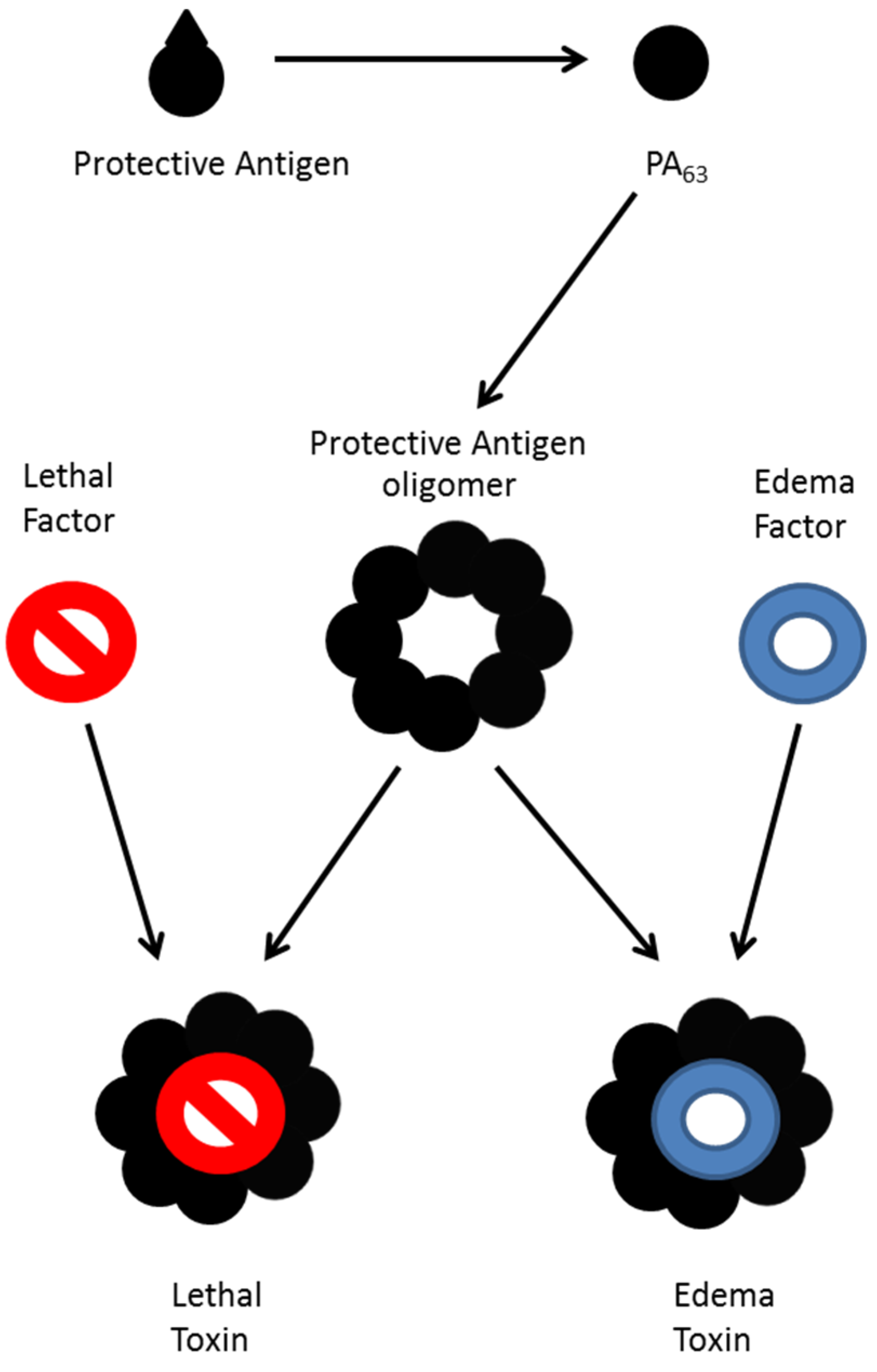
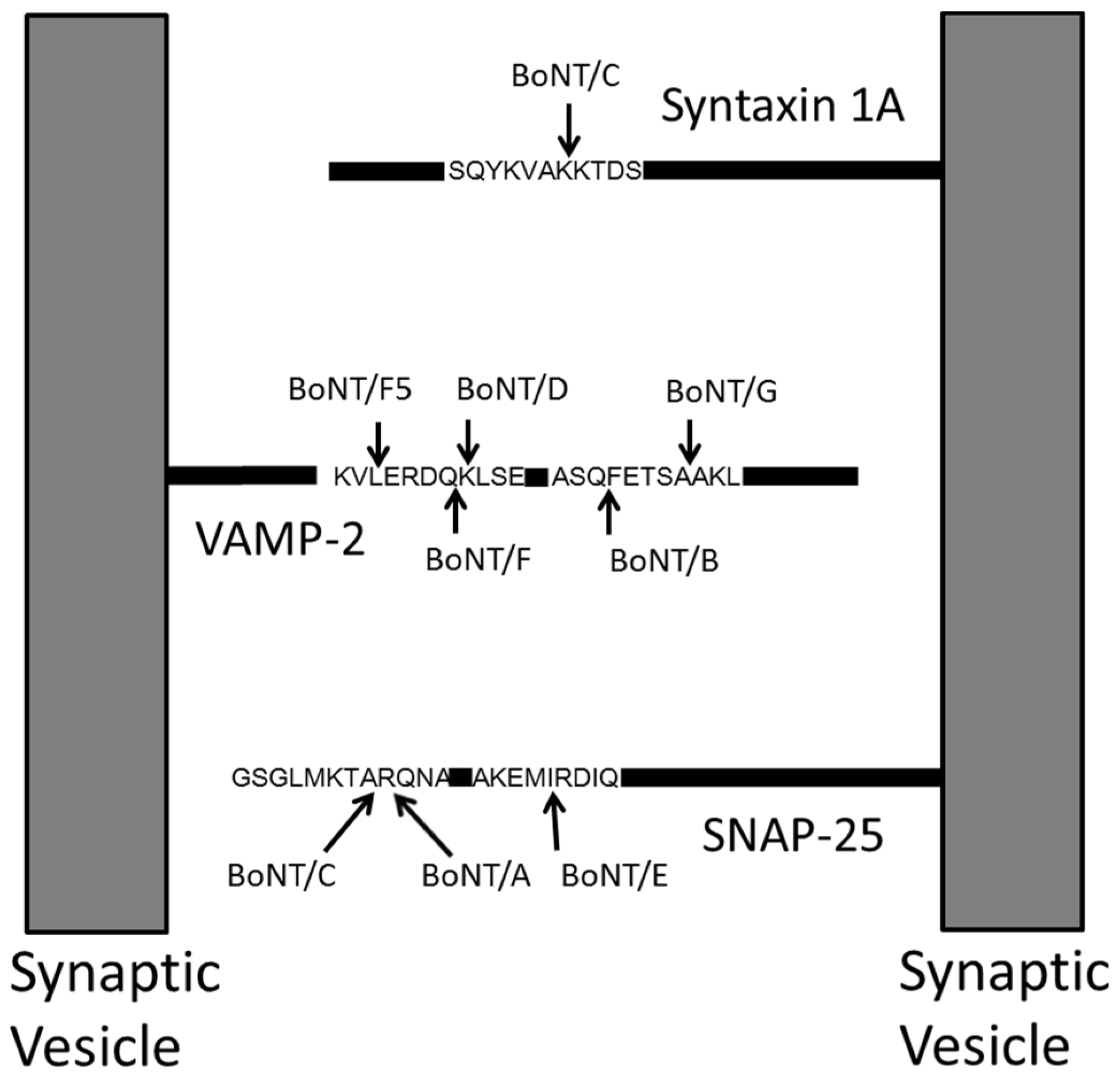
2. Mass Spectrometric Detection of Amino Acid Sequence of Botulinum Neurotoxin
3. Qualitative Mass Spectrometric Detection of BoNT Activity
| Peptide | Sequence | m/z Observed |
|---|---|---|
| BoNT/A substrate | Acetyl-RGSNKPKIDAGNQRATRXLGGR-NH2 | 2406 |
| BoNT/B substrate | LSELDDRADALQAGASQFESSAAKLKRKYWWKNLK | 4025 |
| BoNT/E substrate | WWWAKLGQEIDTRNRQKDhRIMAKADSNKR-NH2 | 3614 |
| BoNT/F substrate | TSNRRLQQTQAQVDEVVDIMRVNVDKVLERDQKLSELDDRADAL | 5111 |
4. Quantitative Mass Spectrometric Detection of Anthrax Lethal Factor Activity
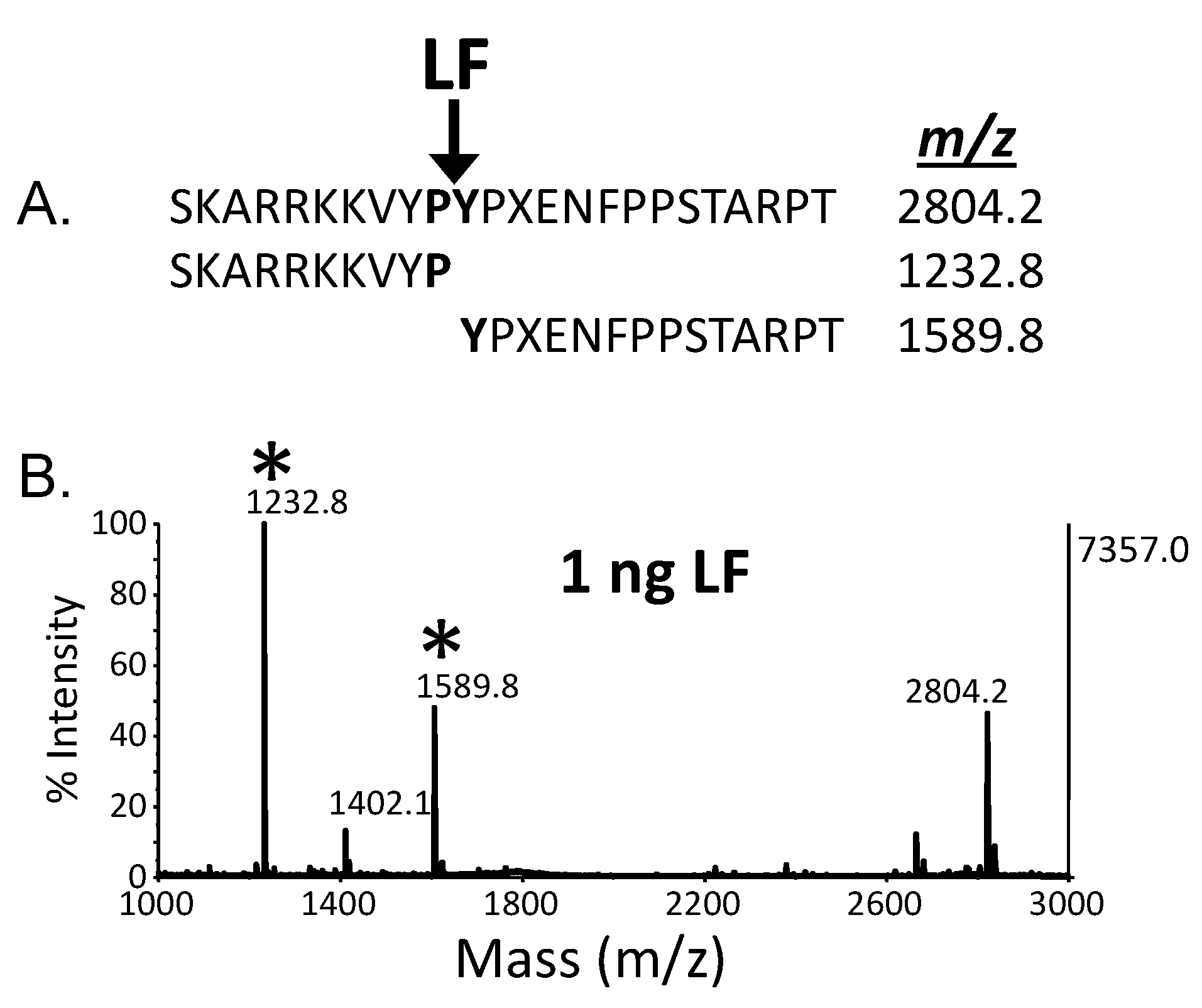
5. Discussion and Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Anhalt, J.P.; Fenselau, C. Identification of Bacteria Using Mass Spectrometry. Anal. Chem. 1975, 47, 219–225. [Google Scholar] [CrossRef]
- Collier, R.J.; Young, J.A.T. Anthrax toxin. Annu. Rev. Cell Dev. Biol. 2003, 19, 45–70. [Google Scholar] [CrossRef] [PubMed]
- Kintzer, A.F.; Thoren, K.L.; Sterling, H.J.; Dong, K.C.; Feld, G.K.; Tang, I.I.; Zhang, T.T.; Williams, E.R.; Berger, J.M.; Krantz, B.A. The Protective Antigen Component of Anthrax Toxin Forms Functional Octameric Complexes. J. Mol. Biol. 2009, 392, 614–629. [Google Scholar] [CrossRef] [PubMed]
- Lacy, D.B.; Wigelsworth, D.J.; Melnyk, R.A.; Harrison, S.C.; Collier, R.J. Structure of heptameric protective antigen bound to an anthrax toxin receptor: A role for receptor in pH-dependent pore formation. Proc. Natl. Acad. Sci. USA 2004, 101, 13147–13151. [Google Scholar] [CrossRef] [PubMed]
- Petosa, C.; Collier, R.J.; Klimpel, K.R.; Leppla, S.H.; Liddington, R.C. Crystal structure of the anthrax toxin protective antigen. Nature 1997, 385, 833–838. [Google Scholar] [CrossRef] [PubMed]
- Vitale, G.; Bernardi, L.; Napolitani, G.; Mock, M.; Montecucco, C. Susceptibility of mitogen-activated protein kinase kinase family members to proteolysis by anthrax lethal factor. Biochem. J. 2000, 352, 739–745. [Google Scholar] [CrossRef] [PubMed]
- Leppla, S.H. Anthrax Toxin Edema Factor—A Bacterial Adenylate-Cyclase that Increases Cyclic-Amp Concentrations in Eukaryotic Cells. Proc. Natl. Acad. Sci. USA 1982, 79, 3162–3166. [Google Scholar] [CrossRef] [PubMed]
- Kuo, S.R.; Willingham, M.C.; Bour, S.H.; Andreas, E.A.; Park, S.K.; Jackson, C.; Duesbery, N.S.; Leppla, S.H.; Tang, W.J.; Frankel, A.E. Anthrax toxin-induced shock in rats is associated with pulmonary edema and hemorrhage. Microb. Pathog. 2008, 44, 467–472. [Google Scholar] [CrossRef] [PubMed]
- Mock, M.; Mignot, T. Anthrax toxins and the host: A story of intimacy. Cell. Microbiol. 2003, 5, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Tournier, J.N.; Quesnel-Hellmann, A.; Cleret, A.; Vidal, D.R. Contribution of toxins to the pathogenesis of inhalational anthrax. Cell. Microbiol. 2007, 9, 555–565. [Google Scholar] [CrossRef] [PubMed]
- Binz, T.; Blasi, J.; Yamasaki, S.; Baumeister, A.; Link, E.; Sudhof, T.C.; Jahn, R.; Niemann, H. Proteolysis of SNAP-25 by types E and A botulinal neurotoxins. J. Biol. Chem. 1994, 269, 1617–1620. [Google Scholar] [PubMed]
- Blasi, J.; Chapman, E.R.; Link, E.; Binz, T.; Yamasaki, S.; de Camilli, P.; Sudhof, T.C.; Niemann, H.; Jahn, R. Botulinum neurotoxin A selectively cleaves the synaptic protein SNAP-25. Nature 1993, 365, 160–163. [Google Scholar] [CrossRef] [PubMed]
- Foran, P.; Lawrence, G.W.; Shone, C.C.; Foster, K.A.; Dolly, J.O. Botulinum neurotoxin C1 cleaves both syntaxin and SNAP-25 in intact and permeabilized chromaffin cells: Correlation with its blockade of catecholamine release. Biochemistry 1996, 35, 2630–2636. [Google Scholar] [CrossRef] [PubMed]
- Schiavo, G.; Rossetto, O.; Catsicas, S.; Polverino de Laureto, P.; DasGupta, B.R.; Benfenati, F.; Montecucco, C. Identification of the nerve terminal targets of botulinum neurotoxin serotypes A, D, and E. J. Biol. Chem. 1993, 268, 23784–23787. [Google Scholar] [PubMed]
- Schiavo, G.; Santucci, A.; Dasgupta, B.R.; Mehta, P.P.; Jontes, J.; Benfenati, F.; Wilson, M.C.; Montecucco, C. Botulinum neurotoxins serotypes A and E cleave SNAP-25 at distinct COOH-terminal peptide bonds. FEBS Lett. 1993, 335, 99–103. [Google Scholar] [CrossRef]
- Williamson, L.C.; Halpern, J.L.; Montecucco, C.; Brown, J.E.; Neale, E.A. Clostridial neurotoxins and substrate proteolysis in intact neurons: Botulinum neurotoxin C acts on synaptosomal-associated protein of 25 kDa. J. Biol. Chem. 1996, 271, 7694–7699. [Google Scholar] [PubMed]
- Kalb, S.R.; Baudys, J.; Webb, R.P.; Wright, P.; Smith, T.J.; Smith, L.A.; Fernandez, R.; Raphael, B.H.; Maslanka, S.E.; Pirkle, J.L.; et al. Discovery of a novel enzymatic cleavage site for botulinum neurotoxin F5. FEBS Lett. 2012, 586, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Schiavo, G.; Benfenati, F.; Poulain, B.; Rossetto, O.; Polverino de Laureto, P.; DasGupta, B.R.; Montecucco, C. Tetanus and botulinum-B neurotoxins block neurotransmitter release by proteolytic cleavage of synaptobrevin. Nature 1992, 359, 832–835. [Google Scholar] [CrossRef] [PubMed]
- Schiavo, G.; Malizio, C.; Trimble, W.S.; Polverino de Laureto, P.; Milan, G.; Sugiyama, H.; Johnson, E.A.; Montecucco, C. Botulinum G neurotoxin cleaves VAMP/synaptobrevin at a single Ala-Ala peptide bond. J. Biol. Chem. 1994, 269, 20213–20216. [Google Scholar] [PubMed]
- Schiavo, G.; Shone, C.C.; Rossetto, O.; Alexander, F.C.; Montecucco, C. Botulinum neurotoxin serotype F is a zinc endopeptidase specific for VAMP/synaptobrevin. J. Biol. Chem. 1993, 268, 11516–11519. [Google Scholar] [PubMed]
- Yamasaki, S.; Baumeister, A.; Binz, T.; Blasi, J.; Link, E.; Cornille, F.; Roques, B.; Fykse, E.M.; Sudhof, T.C.; Jahn, R.; et al. Cleavage of members of the synaptobrevin/VAMP family by types D and F botulinal neurotoxins and tetanus toxin. J. Biol. Chem. 1994, 269, 12764–12772. [Google Scholar] [PubMed]
- Yamasaki, S.; Binz, T.; Hayashi, T.; Szabo, E.; Yamasaki, N.; Eklund, M.; Jahn, R.; Niemann, H. Botulinum neurotoxin type G proteolyses the Ala81-Ala82 bond of rat synaptobrevin 2. Biochem. Biophys. Res. Commun. 1994, 200, 829–835. [Google Scholar] [CrossRef] [PubMed]
- Van Baar, B.L.; Hulst, A.G.; de Jong, A.L.; Wils, E.R. Characterisation of botulinum toxins type A and B, by matrix-assisted laser desorption ionisation and electrospray mass spectrometry. J. Chromatogr. A 2002, 970, 95–115. [Google Scholar] [CrossRef]
- Van Baar, B.L.; Hulst, A.G.; de Jong, A.L.; Wils, E.R. Characterisation of botulinum toxins type C, D, E, and F by matrix-assisted laser desorption ionisation and electrospray mass spectrometry. J. Chromatogr. A 2004, 1035, 97–114. [Google Scholar] [CrossRef] [PubMed]
- Klaubert, B.; Vujtovic-Ockenga, N.; Wermter, R.; Schad, K.; von Meyer, L. Determination of botulinum toxins after peptic sample pre-treatment by multidimensional nanoscale liquid chromatography and nano-electrospray ion-trap mass spectrometry. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2009, 877, 1084–1092. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Goodnough, M.C.; Malizio, C.J.; Pirkle, J.L.; Barr, J.R. Detection of botulinum neurotoxin A in a spiked milk sample with subtype identification through toxin proteomics. Anal. Chem. 2005, 77, 6140–6146. [Google Scholar] [CrossRef] [PubMed]
- Kull, S.; Pauly, D.; Stormann, B.; Kirchner, S.; Stammler, M.; Dorner, M.B.; Lasch, P.; Naumann, D.; Dorner, B.G. Multiplex detection of microbial and plant toxins by immunoaffinity enrichment and matrix-assisted laser desorption/ionization mass spectrometry. Anal. Chem. 2010, 82, 2916–2924. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Baudys, J.; Rees, J.; Marshall, K.M.; Kalb, S.R.; Parks, B.A.; Nowaczyk, L., II; Pirkle, J.L.; Barr, J.R. Subtyping botulinum neurotoxins by sequential multiple endoproteases in-gel digestion coupled with mass spectrometry. Anal. Chem. 2012, 84, 4652–4658. [Google Scholar] [CrossRef] [PubMed]
- Morineaux, V.; Mazuet, C.; Hilaire, D.; Enche, J.; Popoff, M.R. Characterization of botulinum neurotoxin type A subtypes by immunocapture enrichment and liquid chromatography-tandem mass spectrometry. Anal. Bioanal. Chem. 2015, 407, 5559–5570. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Baudys, J.; Rees, J.C.; Smith, T.J.; Smith, L.A.; Helma, C.H.; Hill, K.; Kull, S.; Kirchner, S.; Dorner, M.B.; et al. De novo subtype and strain identification of botulinum neurotoxin type B through toxin proteomics. Anal. Bioanal. Chem. 2012, 403, 215–226. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Garcia-Rodriguez, C.; Lou, J.; Baudys, J.; Smith, T.J.; Marks, J.D.; Smith, L.A.; Pirkle, J.L.; Barr, J.R. Extraction of BoNT/A, /B, /E, and /F with a single, high affinity monoclonal antibody for detection of botulinum neurotoxin by Endopep-MS. PLoS ONE 2010, 5, e12237. [Google Scholar] [CrossRef] [PubMed]
- Barr, J.R.; Moura, H.; Boyer, A.E.; Woolfitt, A.R.; Kalb, S.R.; Pavlopoulos, A.; McWilliams, L.G.; Schmidt, J.G.; Martinez, R.A.; Ashley, D.L. Botulinum neurotoxin detection and differentiation by mass spectrometry. Emerg. Infect. Dis. 2005, 11, 1578–1583. [Google Scholar] [CrossRef] [PubMed]
- Boyer, A.E.; Moura, H.; Woolfitt, A.R.; Kalb, S.R.; McWilliams, L.G.; Pavlopoulos, A.; Schmidt, J.G.; Ashley, D.L.; Barr, J.R. From the mouse to the mass spectrometer: Detection and differentiation of the endoproteinase activities of botulinum neurotoxins A-G by mass spectrometry. Anal. Chem. 2005, 77, 3916–3924. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Moura, H.; Boyer, A.E.; McWilliams, L.G.; Pirkle, J.L.; Barr, J.R. The use of Endopep-MS for the detection of botulinum toxins A, B, E, and F in serum and stool samples. Anal. Biochem. 2006, 351, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Baudys, J.; Kalb, S.R.; Barr, J.R. Improved detection of botulinum neurotoxin type A in stool by mass spectrometry. Anal. Biochem. 2011, 412, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Baudys, J.; Ye, Y.M.; Rees, J.C.; Barr, J.R.; Pirkle, J.L.; Kalb, S.R. Improved detection of botulinum neurotoxin serotype A by Endopep-MS through peptide substrate modification. Anal. Biochem. 2013, 432, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Rosen, O.; Feldberg, L.; Gura, S.; Zichel, R. A new peptide substrate for enhanced BoNT/B detection by Endopep-LC-MS-MS/MRM assay. Anal. Biochem. 2015, 473, 7–10. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Krilich, J.; Baudys, J.; Barr, J.R.; Kalb, S.R. Enhanced detection of type C botulinum neurotoxin by the Endopep-MS assay through optimization of peptide substrates. Bioorg. Med. Chem. 2015, 23, 3667–3673. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Krilich, J.; Baudys, J.; Barr, J.R.; Kalb, S.R. Optimization of peptide substrates for botulinum neurotoxin E improves detection sensitivity in the Endopep-MS assay. Anal. Biochem. 2015, 468, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Baudys, J.; Egan, C.; Smith, T.J.; Smith, L.A.; Pirkle, J.L.; Barr, J.R. Different substrate recognition requirements for cleavage of synaptobrevin-2 by Clostridium baratii and Clostridium botulinum type F neurotoxins. Appl. Environ. Microbiol. 2011, 77, 1301–1318. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Lou, J.; Garcia-Rodriguez, C.; Geren, I.N.; Smith, T.J.; Moura, H.; Marks, J.D.; Smith, L.A.; Pirkle, J.L.; Barr, J.R. Extraction and inhibition of enzymatic activity of botulinum neurotoxins/A1, /A2, and /A3 by a panel of monoclonal anti-BoNT/A antibodies. PLoS ONE 2009, 4, e5355. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Santana, W.I.; Geren, I.N.; Garcia-Rodriguez, C.; Lou, J.L.; Smith, T.J.; Marks, J.D.; Smith, L.A.; Pirkle, J.L.; Barr, J.R. Extraction and inhibition of enzymatic activity of botulinum neurotoxins/B1, /B2, /B3, /B4, and /B5 by a panel of monoclonal anti-BoNT/B antibodies. BMC Biochem. 2011. [Google Scholar] [CrossRef] [PubMed]
- Kalb, S.R.; Smith, T.J.; Moura, H.; Hill, K.; Lou, J.; Geren, I.N.; Garcia-Rodriguez, C.; Marks, J.D.; Smith, L.A.; Pirkle, J.L.; et al. The use of Endopep-MS to detect multiple subtypes of botulinum neurotoxins A, B, E, and F. Int. J. Mass Spectrom. 2008, 278, 101–108. [Google Scholar] [CrossRef]
- Raphael, B.H.; Lautenschlager, M.; Kalb, S.R.; de Jong, L.I.T.; Frace, M.; Luquez, C.; Barr, J.R.; Fernandez, R.A.; Maslanka, S.E. Analysis of a unique Clostridium botulinum strain from the Southern hemisphere producing a novel type E botulinum neurotoxin subtype. BMC Microbiol. 2012. [Google Scholar] [CrossRef] [PubMed]
- Boyer, A.E.; Quinn, C.P.; Woolfitt, A.R.; Pirkle, J.L.; McWilliams, L.G.; Stamey, K.L.; Bagarozzi, D.A.; Hart, J.C.; Barr, J.R. Detection and quantification of anthrax lethal factor in serum by mass spectrometry. Anal. Chem. 2007, 79, 8463–8470. [Google Scholar] [CrossRef] [PubMed]
- Kuklenyik, Z.; Boyer, A.E.; Lins, R.; Quinn, C.P.; Gallegos-Candela, M.; Woolfitt, A.; Pirkle, J.L.; Barr, J.R. Comparison of MALDI-TOF-MS and HPLC-ESI-MS/MS for Endopeptidase Activity-Based Quantification of Anthrax Lethal Factor in Serum. Anal. Chem. 2011, 83, 1760–1765. [Google Scholar] [CrossRef] [PubMed]
- Boyer, A.E.; Quinn, C.P.; Hoffmaster, A.R.; Kozel, T.R.; Saile, E.; Marston, C.K.; Percival, A.; Plikaytis, B.D.; Woolfitt, A.R.; Gallegos, M.; et al. Kinetics of Lethal Factor and Poly-d-Glutamic Acid Antigenemia during Inhalation Anthrax in Rhesus Macaques. Infect. Immun. 2009, 77, 3432–3441. [Google Scholar] [CrossRef] [PubMed]
- Boyer, A.E.; Quinn, C.P.; Beesley, C.A.; Gallegos-Candela, M.; Marston, C.K.; Cronin, L.X.; Lins, R.C.; Stoddard, R.A.; Li, H.; Schiffer, J.; et al. Lethal Factor Toxemia and Anti-Protective Antigen Antibody Activity in Naturally Acquired Cutaneous Anthrax. J. Infect. Dis. 2011, 204, 1321–1327. [Google Scholar] [CrossRef] [PubMed]
- Walsh, J.J.; Pesik, N.; Quinn, C.P.; Urdaneta, V.; Dykewicz, C.A.; Boyer, A.E.; Guarner, J.; Wilkins, P.; Norville, K.J.; Barr, J.R.; et al. A case of naturally acquired inhalation anthrax: Clinical care and analyses of anti-protective antigen immunoglobulin G and lethal factor. Clin. Infect. Dis. 2007, 44, 968–971. [Google Scholar] [CrossRef] [PubMed]
- Sprenkle, M.D.; Griffith, J.; Marinelli, W.; Boyer, A.E.; Quinn, C.P.; Pesik, N.T.; Hoffmaster, A.; Keenan, J.; Juni, B.A.; Blaney, D.D. Lethal Factor and Anti-Protective Antigen IgG Levels Associated with Inhalation Anthrax, Minnesota, USA. Emerg. Infect. Dis. 2014, 20, 310–314. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kalb, S.R.; Boyer, A.E.; Barr, J.R. Mass Spectrometric Detection of Bacterial Protein Toxins and Their Enzymatic Activity. Toxins 2015, 7, 3497-3511. https://doi.org/10.3390/toxins7093497
Kalb SR, Boyer AE, Barr JR. Mass Spectrometric Detection of Bacterial Protein Toxins and Their Enzymatic Activity. Toxins. 2015; 7(9):3497-3511. https://doi.org/10.3390/toxins7093497
Chicago/Turabian StyleKalb, Suzanne R., Anne E. Boyer, and John R. Barr. 2015. "Mass Spectrometric Detection of Bacterial Protein Toxins and Their Enzymatic Activity" Toxins 7, no. 9: 3497-3511. https://doi.org/10.3390/toxins7093497
APA StyleKalb, S. R., Boyer, A. E., & Barr, J. R. (2015). Mass Spectrometric Detection of Bacterial Protein Toxins and Their Enzymatic Activity. Toxins, 7(9), 3497-3511. https://doi.org/10.3390/toxins7093497






