Polyketides, Toxins and Pigments in Penicillium marneffei
Abstract
:1. Introduction
2. Diversity and Phylogeny of pks Genes in P. marneffei

3. Pigments and Toxins in P. marneffei
3.1. Melanin
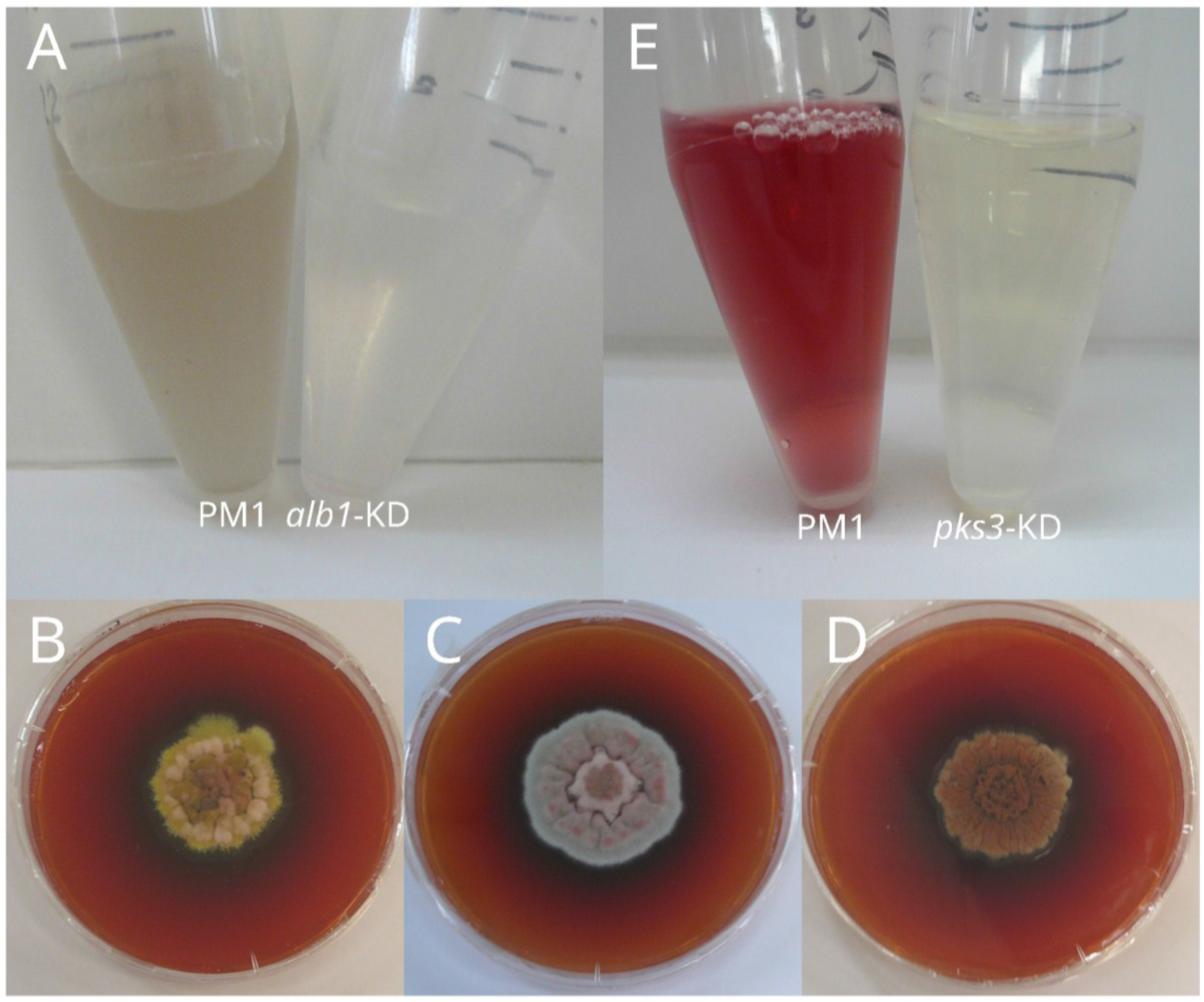
3.2. Yellow Pigment: Mitorubrinol and Mitorubrinic Acid
3.3. Toxin and Red Pigment Biosynthetic Pathway: Monascorubrin, Rubropunctatin, Citrinin and Ankaflavin
4. Concluding Remarks
Acknowledgments
Author contributions
Conflicts of Interest
References
- Hsueh, P.R.; Teng, L.J.; Hung, C.C.; Hsu, J.H.; Yang, P.C.; Ho, S.W.; Luh, K.T. Molecular evidence for strain dissemination of Penicillium marneffei: An emerging pathogen in Taiwan. J. Infect. Dis. 2000, 181, 1706–1712. [Google Scholar] [CrossRef] [PubMed]
- Supparatpinyo, K.; Khamwan, C.; Baosoung, V.; Nelson, K.E.; Sirisanthana, T. Disseminated Penicillium marneffei infection in southeast Asia. Lancet 1994, 344, 110–113. [Google Scholar] [CrossRef]
- Wong, S.S.; Siau, H.; Yuen, K.Y. Penicilliosis marneffei—West meets East. J. Med. Microbiol. 1999, 48, 973–975. [Google Scholar] [CrossRef] [PubMed]
- Yuen, K.Y.; Wong, S.S.; Tsang, D.N.; Chau, P.Y. Serodiagnosis of Penicillium marneffei infection. Lancet 1994, 344, 444–445. [Google Scholar] [CrossRef]
- Deng, Z.L.; Connor, D.H. Progressive disseminated penicilliosis caused by Penicillium marneffei. Report of eight cases and differentiation of the causative organism from Histoplasma capsulatum. Am. J. Clin. Pathol. 1985, 84, 323–327. [Google Scholar] [PubMed]
- Low, K.; Lee, S.S. The pattern of aids reporting and the implications on HIV surveillance. Public Health Epidemiol. Bull. 2002, 11, 41–49. [Google Scholar]
- Lo, C.Y.; Chan, D.T.; Yuen, K.Y.; Li, F.K.; Cheng, K.P. Penicillium marneffei infection in a patient with sle. Lupus 1995, 4, 229–231. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.L.; Hung, C.C.; Chang, S.C.; Chueh, S.C.; La, M.K. Disseminated Penicillium marneffei infection in a renal-transplant recipient successfully treated with liposomal amphotericin B. Transplantation 2003, 76, 1136–1137. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.S.; Woo, P.C.; Yuen, K.Y. Candida tropicalis and Penicillium marneffei mixed fungaemia in a patient with Waldenström’s macroglobulinaemia. Eur. J. Clin. Microbiol. Infect. Dis. 2001, 20, 132–135. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lau, S.K.; Lau, C.C.; Chong, K.T.; Hui, W.T.; Wong, S.S.; Yuen, K.Y. Penicillium marneffei fungaemia in an allogeneic bone marrow transplant recipient. Bone Marrow Transpl. 2005, 35, 831–833. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.P.; Mao, H.; Yang, W.; Chan, K.W.; Ho, M.H.; Lee, T.L.; Chan, J.F.; Woo, P.C.; Tu, W.; Lau, Y.L. Penicillium marneffei infection and impaired ifn-gamma immunity in humans with autosomal-dominant gain-of-phosphorylation stat1 mutations. J. Allergy Clin. Immunol. 2014, 133, 894–896. [Google Scholar] [CrossRef] [PubMed]
- Wongkulab, P.; Wipasa, J.; Chaiwarith, R.; Supparatpinyo, K. Autoantibody to interferon-gamma associated with adult-onset immunodeficiency in non-HIV individuals in northern Thailand. PLoS One 2013, 8, e76371. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.; Trendell-Smith, N.J.; Chan, J.C.; Hung, I.F.; Tang, B.S.; Cheng, V.C.; Yeung, C.K.; Yuen, K.Y. Reactive and infective dermatoses associated with adult-onset immunodeficiency due to anti-interferon-gamma autoantibody: Sweet’s syndrome and beyond. Dermatology 2013, 226, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.; Chan, T.S.; Gill, H.; Lam, F.Y.; Trendell-Smith, N.J.; Sridhar, S.; Tse, H.; Lau, S.K.; Hung, I.F.; Yuen, K.Y.; et al. Disseminated infections with Talaromyces marneffei in non-AIDS patients given monoclonal antibodies against CD20 and kinase inhibitors. Emerg. Infect. Dis. 2015, 21, 1101–1106. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Zhen, H.; Cai, J.J.; Yu, J.; Lau, S.K.; Wang, J.; Teng, J.L.; Wong, S.S.; Tse, R.H.; Chen, R.; et al. The mitochondrial genome of the thermal dimorphic fungus Penicillium marneffei is more closely related to those of molds than yeasts. FEBS Lett. 2003, 555, 469–477. [Google Scholar] [CrossRef]
- Woo, P.C.; Chong, K.T.; Tse, H.; Cai, J.J.; Lau, C.C.; Zhou, A.C.; Lau, S.K.; Yuen, K.Y. Genomic and experimental evidence for a potential sexual cycle in the pathogenic thermal dimorphic fungus Penicillium marneffei. FEBS Lett. 2006, 580, 3409–3416. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lau, C.C.; Chong, K.T.; Tse, H.; Tsang, D.N.; Lee, R.A.; Tse, C.W.; Que, T.L.; Chung, L.M.; Ngan, A.H.; et al. Mp1 homologue-based multilocus sequence system for typing the pathogenic fungus Penicillium marneffei: A novel approach using lineage-specific genes. J. Clin. Microbiol. 2007, 45, 3647–3654. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Tam, E.W.; Chong, K.T.; Cai, J.J.; Tung, E.T.; Ngan, A.H.; Lau, S.K.; Yuen, K.Y. High diversity of polyketide synthase genes and the melanin biosynthesis gene cluster in Penicillium marneffei. FEBS J. 2010, 277, 3750–3758. [Google Scholar] [CrossRef] [PubMed]
- Collemare, J.; Billard, A.; Bohnert, H.U.; Lebrun, M.H. Biosynthesis of secondary metabolites in the rice blast fungus Magnaporthe grisea: The role of hybrid PKS-NRPS in pathogenicity. Mycol. Res. 2008, 112, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lam, C.W.; Tam, E.W.; Lee, K.C.; Yung, K.K.; Leung, C.K.; Sze, K.H.; Lau, S.K.; Yuen, K.Y. The biosynthetic pathway for a thousand-year-old natural food colorant and citrinin in Penicillium marneffei. Sci. Rep. 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lam, C.W.; Tam, E.W.; Leung, C.K.; Wong, S.S.; Lau, S.K.; Yuen, K.Y. First discovery of two polyketide synthase genes for mitorubrinic acid and mitorubrinol yellow pigment biosynthesis and implications in virulence of Penicillium marneffei. PLoS Negl. Trop. Dis. 2012, 6, e1871. [Google Scholar] [CrossRef] [PubMed]
- Kroken, S.; Glass, N.L.; Taylor, J.W.; Yoder, O.C.; Turgeon, B.G. Phylogenomic analysis of type I polyketide synthase genes in pathogenic and saprobic ascomycetes. Proc. Natl. Acad. Sci. USA 2003, 100, 15670–15675. [Google Scholar] [CrossRef] [PubMed]
- Noverr, M.C.; Williamson, P.R.; Fajardo, R.S.; Huffnagle, G.B. CNLAC1 is required for extrapulmonary dissemination of Cryptococcus neoformans but not pulmonary persistence. Infect. Immun. 2004, 72, 1693–1699. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, M.B.; Marques, A.F.; Nosanchuk, J.D.; Casadevall, A.; Travassos, L.R.; Taborda, C.P. Melanin in the dimorphic fungal pathogen Paracoccidioides brasiliensis: Effects on phagocytosis, intracellular resistance and drug susceptibility. Microbes Infect. 2006, 8, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.B.; Thomaz, L.; Marques, A.F.; Svidzinski, A.E.; Nosanchuk, J.D.; Casadevall, A.; Travassos, L.R.; Taborda, C.P. Resistance of melanized yeast cells of Paracoccidioides brasiliensis to antimicrobial oxidants and inhibition of phagocytosis using carbohydrates and monoclonal antibody to cd18. Mem. Inst. Oswaldo. Cruz. 2009, 104, 644–648. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Shao, Y.; Chen, F. Monascus pigments. Appl. Microbiol. Biotechnol. 2012, 96, 1421–1440. [Google Scholar] [CrossRef] [PubMed]
- Carels, M.; Shepherd, D. The effect of different nitrogen sources on pigment production and sporulation of Monascus species in submerged, shaken culture. Can. J. Microbiol. 1977, 23, 1360–1372. [Google Scholar] [CrossRef] [PubMed]
- Fabre, C.E.; Santerre, A.L.; Loret, M.O.; Baberian, R.; Pareilleux, A.; Goma, G.; Blanc, P.J. Production and food applications of the red pigments of Monascus ruber. J. Food Sci. 1993, 58, 1099–1102. [Google Scholar] [CrossRef]
- Pihet, M.; Vandeputte, P.; Tronchin, G.; Renier, G.; Saulnier, P.; Georgeault, S.; Mallet, R.; Chabasse, D.; Symoens, F.; Bouchara, J.P. Melanin is an essential component for the integrity of the cell wall of Aspergillus fumigatus conidia. BMC Microbiol. 2009, 9. [Google Scholar] [CrossRef] [PubMed]
- San-Blas, G.; Guanipa, O.; Moreno, B.; Pekerar, S.; San-Blas, F. Cladosporium carrionii and Hormoconis resinae (c. Resinae): Cell wall and melanin studies. Curr. Microbiol. 1996, 32, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Romero-Martinez, R.; Wheeler, M.; Guerrero-Plata, A.; Rico, G.; Torres-Guerrero, H. Biosynthesis and functions of melanin in Sporothrix schenckii. Infect. Immun. 2000, 68, 3696–3703. [Google Scholar] [PubMed]
- Van Duin, D.; Casadevall, A.; Nosanchuk, J.D. Melanization of Cryptococcus neoformans and Histoplasma capsulatum reduces their susceptibilities to amphotericin B and caspofungin. Antimicrob. Agents Chemother. 2002, 46, 3394–3400. [Google Scholar] [CrossRef] [PubMed]
- Nosanchuk, J.D.; van Duin, D.; Mandal, P.; Aisen, P.; Legendre, A.M.; Casadevall, A. Blastomyces dermatitidis produces melanin in vitro and during infection. FEMS Microbiol. Lett. 2004, 239, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Nosanchuk, J.D.; Yu, J.J.; Hung, C.Y.; Casadevall, A.; Cole, G.T. Coccidioides posadasii produces melanin in vitro and during infection. Fungal Genet. Biol. 2007, 44, 517–520. [Google Scholar] [CrossRef] [PubMed]
- Youngchim, S.; Hay, R.J.; Hamilton, A.J. Melanization of Penicillium marneffei in vitro and in vivo. Microbiology 2005, 151, 291–299. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.F.; Chang, Y.C.; Washburn, R.G.; Wheeler, M.H.; Kwon-Chung, K.J. The developmentally regulated alb1 gene of Aspergillus fumigatus: Its role in modulation of conidial morphology and virulence. J. Bacteriol. 1998, 180, 3031–3038. [Google Scholar] [PubMed]
- Tsai, H.F.; Wheeler, M.H.; Chang, Y.C.; Kwon-Chung, K.J. A developmentally regulated gene cluster involved in conidial pigment biosynthesis in Aspergillus fumigatus. J. Bacteriol. 1999, 181, 6469–6477. [Google Scholar] [PubMed]
- Alviano, C.S.; Farbiarz, S.R.; De Souza, W.; Angluster, J.; Travassos, L.R. Characterization of Fonsecaea pedrosoi melanin. J. Gen. Microbiol. 1991, 137, 837–844. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, M.H. Comparisons of fungal melanin biosynthesis in ascomycetous, imperfect and basidiomycetous fungi. Trans. Br. Mycol. Soc. 1983, 81, 29–36. [Google Scholar] [CrossRef]
- Youngchim, S.; Morris-Jones, R.; Hay, R.J.; Hamilton, A.J. Production of melanin by Aspergillus fumigatus. J. Med. Microbiol. 2004, 53, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Adachi, K.; Hamer, J.E. Divergent camp signaling pathways regulate growth and pathogenesis in the rice blast fungus Magnaporthe grisea. Plant Cell 1998, 10, 1361–1374. [Google Scholar] [CrossRef] [PubMed]
- Fujii, I.; Mori, Y.; Watanabe, A.; Kubo, Y.; Tsuji, G.; Ebizuka, Y. Enzymatic synthesis of 1,3,6,8-tetrahydroxynaphthalene solely from malonyl coenzyme a by a fungal iterative type I polyketide synthase PKS1. Biochemistry 2000, 39, 8853–8858. [Google Scholar] [CrossRef] [PubMed]
- Langfelder, K.; Streibel, M.; Jahn, B.; Haase, G.; Brakhage, A.A. Biosynthesis of fungal melanins and their importance for human pathogenic fungi. Fungal Genet. Biol. 2003, 38, 143–158. [Google Scholar] [CrossRef]
- Chai, L.Y.; Netea, M.G.; Sugui, J.; Vonk, A.G.; van de Sande, W.W.; Warris, A.; Kwon-Chung, K.J.; Kullberg, B.J. Aspergillus fumigatus conidial melanin modulates host cytokine response. Immunobiology 2010, 215, 915–920. [Google Scholar] [CrossRef] [PubMed]
- Salas, S.D.; Bennett, J.E.; Kwon-Chung, K.J.; Perfect, J.R.; Williamson, P.R. Effect of the laccase gene CNLAC1, on virulence of Cryptococcus neoformans. J. Exp. Med. 1996, 184, 377–386. [Google Scholar] [CrossRef] [PubMed]
- Bailao, A.M.; Schrank, A.; Borges, C.L.; Dutra, V.; Walquiria Ines Molinari-Madlum, E.E.; Soares Felipe, M.S.; Soares Mendes-Giannini, M.J.; Martins, W.S.; Pereira, M.; Maria de Almeida Soares, C. Differential gene expression by Paracoccidioides brasiliensis in host interaction conditions: Representational difference analysis identifies candidate genes associated with fungal pathogenesis. Microbes Infect. 2006, 8, 2686–2697. [Google Scholar] [CrossRef] [PubMed]
- Madrid, I.M.; Xavier, M.O.; Mattei, A.S.; Fernandes, C.G.; Guim, T.N.; Santin, R.; Schuch, L.F.; Nobre Mde, O.; Araujo Meireles, M.C. Role of melanin in the pathogenesis of cutaneous sporotrichosis. Microbes Infect. 2010, 12, 162–165. [Google Scholar] [CrossRef] [PubMed]
- Nosanchuk, J.D.; Casadevall, A. Cellular charge of Cryptococcus neoformans: contributions from the capsular polysaccharide, melanin, and monoclonal antibody binding. Infect. Immun. 1997, 65, 1836–1841. [Google Scholar] [PubMed]
- Cunha, M.M.; Franzen, A.J.; Alviano, D.S.; Zanardi, E.; Alviano, C.S.; De Souza, W.; Rozental, S. Inhibition of melanin synthesis pathway by tricyclazole increases susceptibility of Fonsecaea pedrosoi against mouse macrophages. Microsc. Res. Tech. 2005, 68, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Peltroche-Llacsahuanga, H.; Schnitzler, N.; Jentsch, S.; Platz, A.; De Hoog, S.; Schweizer, K.G.; Haase, G. Analyses of phagocytosis, evoked oxidative burst, and killing of black yeasts by human neutrophils: A tool for estimating their pathogenicity? Med. Mycol. 2003, 41, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Aisen, P.; Casadevall, A. Melanin, melanin “ghosts,” and melanin composition in Cryptococcus neoformans. Infect. Immun. 1996, 64, 2420–2424. [Google Scholar] [PubMed]
- Taborda, C.P.; da Silva, M.B.; Nosanchuk, J.D.; Travassos, L.R. Melanin as a virulence factor of Paracoccidioides brasiliensis and other dimorphic pathogenic fungi: A minireview. Mycopathologia 2008, 165, 331–339. [Google Scholar] [CrossRef] [PubMed]
- Nosanchuk, J.D.; Casadevall, A. Impact of melanin on microbial virulence and clinical resistance to antimicrobial compounds. Antimicrob. Agents Chemother. 2006, 50, 3519–3528. [Google Scholar] [CrossRef] [PubMed]
- Gan, E.V.; Lam, K.M.; Haberman, H.F.; Menon, I.A. Oxidizing and reducing properties of melanins. Br. J. Dermatol. 1977, 96, 25–28. [Google Scholar] [CrossRef] [PubMed]
- Volling, K.; Thywissen, A.; Brakhage, A.A.; Saluz, H.P. Phagocytosis of melanized Aspergillus conidia by macrophages exerts cytoprotective effects by sustained PI3K/Akt signalling. Cell. Microbiol. 2011, 13, 1130–1148. [Google Scholar] [CrossRef] [PubMed]
- Mednick, A.J.; Nosanchuk, J.D.; Casadevall, A. Melanization of Cryptococcus neoformans affects lung inflammatory responses during cryptococcal infection. Infect. Immun. 2005, 73, 2012–2019. [Google Scholar] [CrossRef] [PubMed]
- Rosas, A.L.; MacGill, R.S.; Nosanchuk, J.D.; Kozel, T.R.; Casadevall, A. Activation of the alternative complement pathway by fungal melanins. Clin. Diagn. Lab. Immunol. 2002, 9, 144–148. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Porco, J.A., Jr. Asymmetric syntheses of (−)-mitorubrin and related azaphilone natural products. Org. Lett. 2006, 8, 5169–5171. [Google Scholar] [CrossRef] [PubMed]
- Buechi, G.; White, J.D.; Wogan, G.N. The structures of mitorubrin and mitorubrinol. J. Am. Chem. Soc. 1965, 87, 3484–3489. [Google Scholar] [CrossRef] [PubMed]
- Moss, M.O.; Robinson, F.V.; Wood, A.B. Rubratoxin B, a toxic metabolite of Penicillium rubrum. Chem. Ind. 1968, 18, 587–588. [Google Scholar] [PubMed]
- Natori, S.; Sakaki, S.; Kurata, H.; Udagawa, S.I.; Ichinoe, M. Production of rubratoxin B by Penicillium purpurogenum stoll. Appl. Microbiol. 1970, 19, 613–617. [Google Scholar] [PubMed]
- Suzuki, S.; Hosoe, T.; Nozawa, K.; Yaguchi, T.; Udagawa, S.; Kawai, K. Mitorubrin derivatives on ascomata of some Talaromyces species of ascomycetous fungi. J. Nat. Prod. 1999, 62, 1328–1329. [Google Scholar] [CrossRef] [PubMed]
- Natsume, M.; Takahashi, Y.; Marumo, S. (−)-mitorubrinic acid, a morphogenic substance inducing chlamydospore-like cells, and its related new metabolite, (+)-mitorubrinic acid b, isolated from Penicillium funiculosum. Agric. Biol. Chem. 1985, 49, 2517–2519. [Google Scholar] [CrossRef]
- Cox, R.J. Polyketides, proteins and genes in fungi: Programmed nano-machines begin to reveal their secrets. Org. Biomol. Chem. 2007, 5, 2010–2026. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.T.; Lee, Y.R.; Jin, J.; Han, K.H.; Kim, H.; Kim, J.C.; Lee, T.; Yun, S.H.; Lee, Y.W. Two different polyketide synthase genes are required for synthesis of zearalenone in Gibberella zeae. Mol. Microbiol. 2005, 58, 1102–1113. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, S.; Shukla, A.; Mukherjee, S.; Sharma, S.; Guptasarma, P.; Chakraborti, A.K.; Chakrabarti, A. Putative structure and characteristics of a red water-soluble pigment secreted by Penicillium marneffei. Med. Mycol. 2007, 45, 419–427. [Google Scholar] [CrossRef] [PubMed]
- Mapari, S.A.; Meyer, A.S.; Thrane, U.; Frisvad, J.C. Identification of potentially safe promising fungal cell factories for the production of polyketide natural food colorants using chemotaxonomic rationale. Microb. Cell Factor 2009, 8, 24. [Google Scholar] [CrossRef] [PubMed]
- Jůzlová, P.; Martínková, L.; Křen, V. Secondary metabolites of the fungus Monascus: A review. J. Ind. Microbiol. 1996, 16, 163–170. [Google Scholar] [CrossRef]
- Fu, G.; Xu, Y.; Li, Y.; Tan, W. Construction of a replacement vector to disrupt pksCT gene for the mycotoxin citrinin biosynthesis in Monascus aurantiacus and maintain food red pigment production. Asia Pac. J. Clin. Nutr. 2007, 16, 137–142. [Google Scholar] [PubMed]
- Flajs, D.; Peraica, M. Toxicological properties of citrinin. Arch. Hig. Rada. Toksikol. 2009, 60, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, T.; Kinoshita, H.; Ishihara, S.; Sakai, K.; Nagai, S.; Nihira, T. Polyketide synthase gene responsible for citrinin biosynthesis in Monascus purpureus. Appl. Environ. Microbiol. 2005, 71, 3453–3457. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, T.; Kinoshita, H.; Nihira, T. Identification and in vivo functional analysis by gene disruption of ctnA, an activator gene involved in citrinin biosynthesis in Monascus purpureus. Appl. Environ. Microbiol. 2007, 73, 5097–5103. [Google Scholar] [CrossRef] [PubMed]
- Balakrishnan, B.; Karki, S.; Chiu, S.H.; Kim, H.J.; Suh, J.W.; Nam, B.; Yoon, Y.M.; Chen, C.C.; Kwon, H.J. Genetic localization and in vivo characterization of a Monascus azaphilone pigment biosynthetic gene cluster. Appl. Microbiol. Biotechnol. 2013, 97, 6337–6345. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tam, E.W.T.; Tsang, C.-C.; Lau, S.K.P.; Woo, P.C.Y. Polyketides, Toxins and Pigments in Penicillium marneffei. Toxins 2015, 7, 4421-4436. https://doi.org/10.3390/toxins7114421
Tam EWT, Tsang C-C, Lau SKP, Woo PCY. Polyketides, Toxins and Pigments in Penicillium marneffei. Toxins. 2015; 7(11):4421-4436. https://doi.org/10.3390/toxins7114421
Chicago/Turabian StyleTam, Emily W. T., Chi-Ching Tsang, Susanna K. P. Lau, and Patrick C. Y. Woo. 2015. "Polyketides, Toxins and Pigments in Penicillium marneffei" Toxins 7, no. 11: 4421-4436. https://doi.org/10.3390/toxins7114421
APA StyleTam, E. W. T., Tsang, C.-C., Lau, S. K. P., & Woo, P. C. Y. (2015). Polyketides, Toxins and Pigments in Penicillium marneffei. Toxins, 7(11), 4421-4436. https://doi.org/10.3390/toxins7114421







