Activity and Transcriptional Responses of Hepatopancreatic Biotransformation and Antioxidant Enzymes in the Oriental River Prawn Macrobrachium nipponense Exposed to Microcystin-LR
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animals and Chemicals
2.2. MC-LR Exposure
2.3. Sampling, RNA Isolation, and Reverse Transcription (RT)
2.4. Cloning of Full-Length Cat and Gst cDNAs from M. nipponense
2.5. Protein Alignment and Phylogenetic Analyses
2.6. qRT-PCR
2.7. Tissue Distribution of the Cat and Gst Genes
2.8. Enzyme Activity Assays Following MC-LR Exposure
2.9. mRNA Expression of the Enzymes Following MC-LR Exposure
3. Results
3.1. Cloning and Molecular Characterization of the Cat and Gst Gene cDNAs
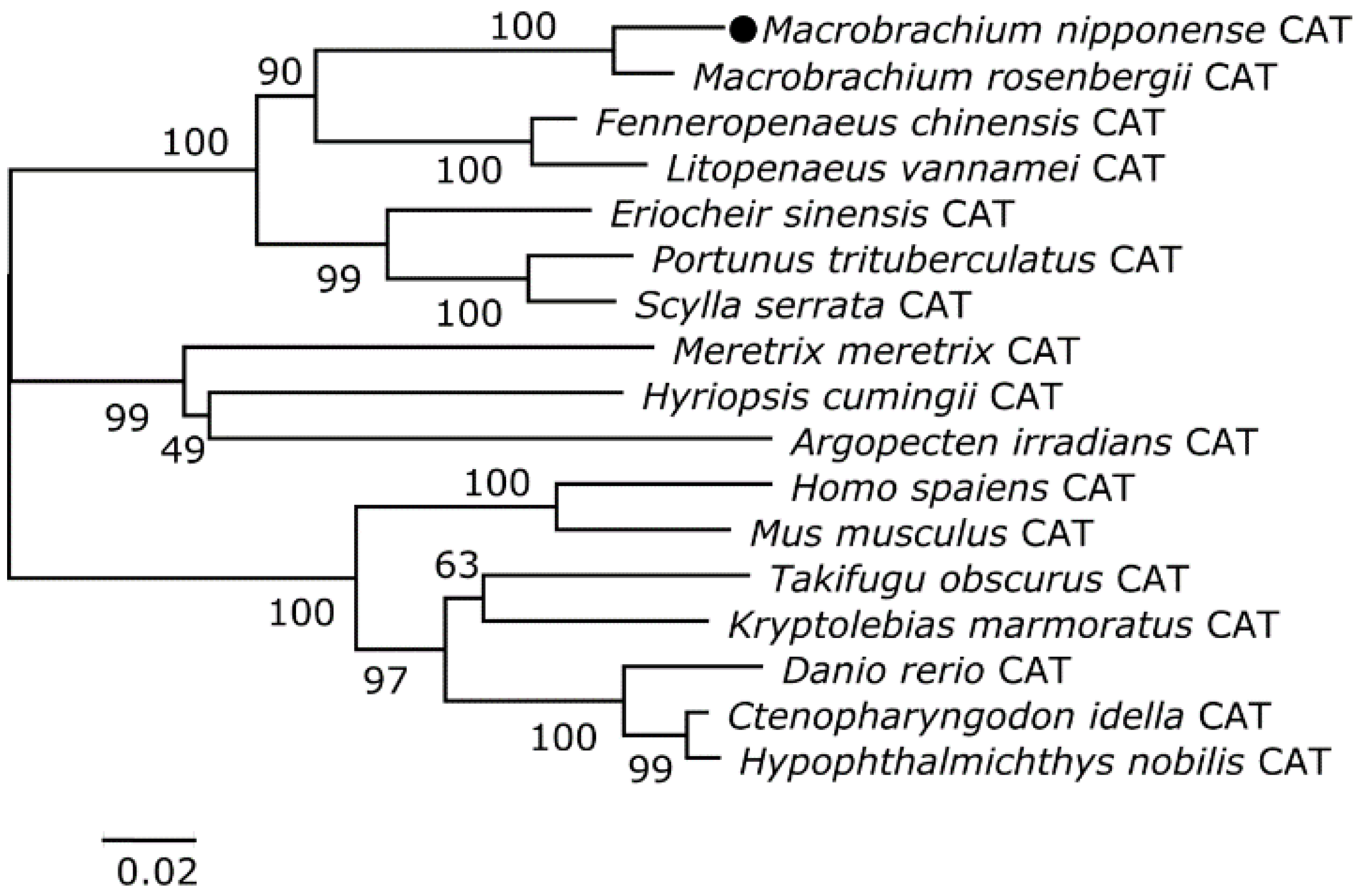
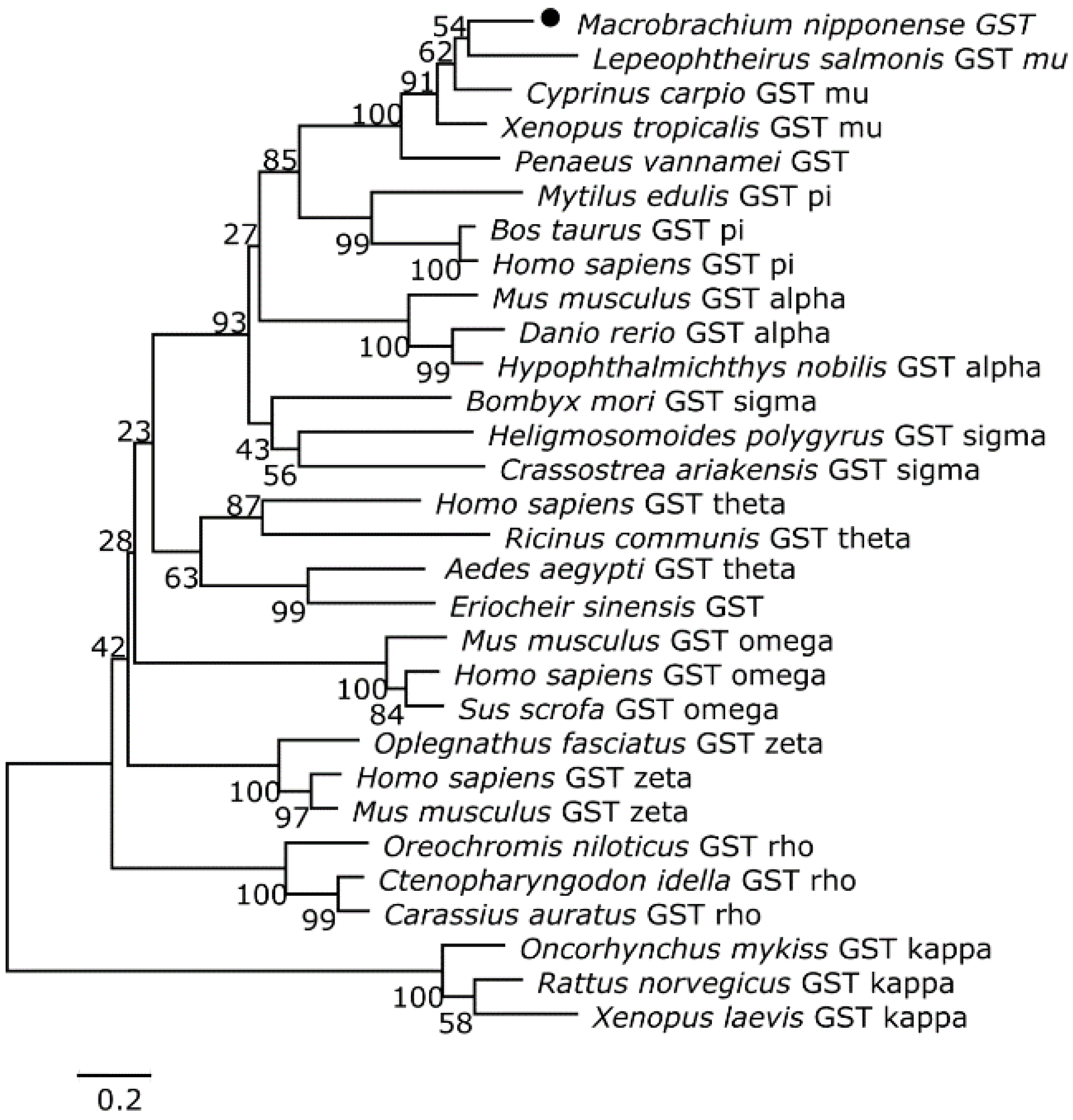
3.2. Homology and Phylogenetic Analyses of the Putative Cat and Gst Amino Acid Sequences
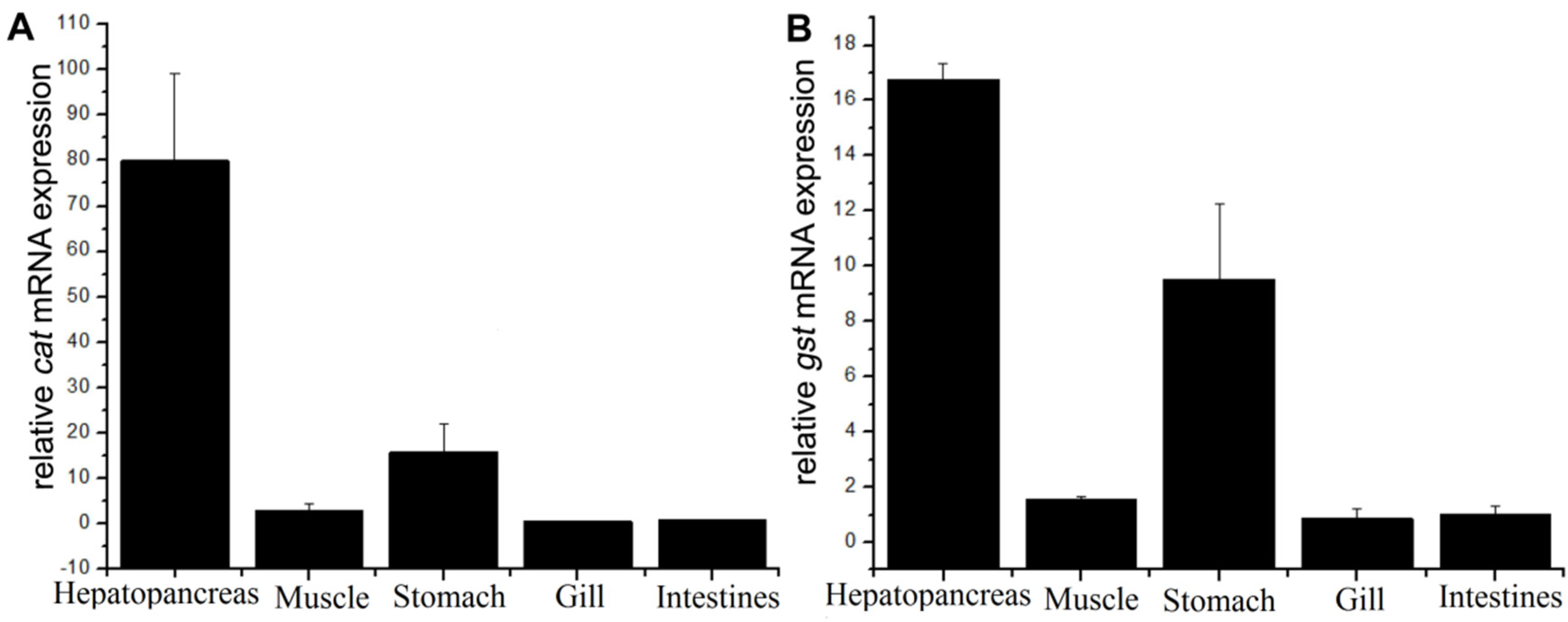
3.3. Tissue Distributions for Cat and Gst mRNAs
3.4. Effect of MC-LR on Biotransformation Enzyme and Antioxidant Enzymes
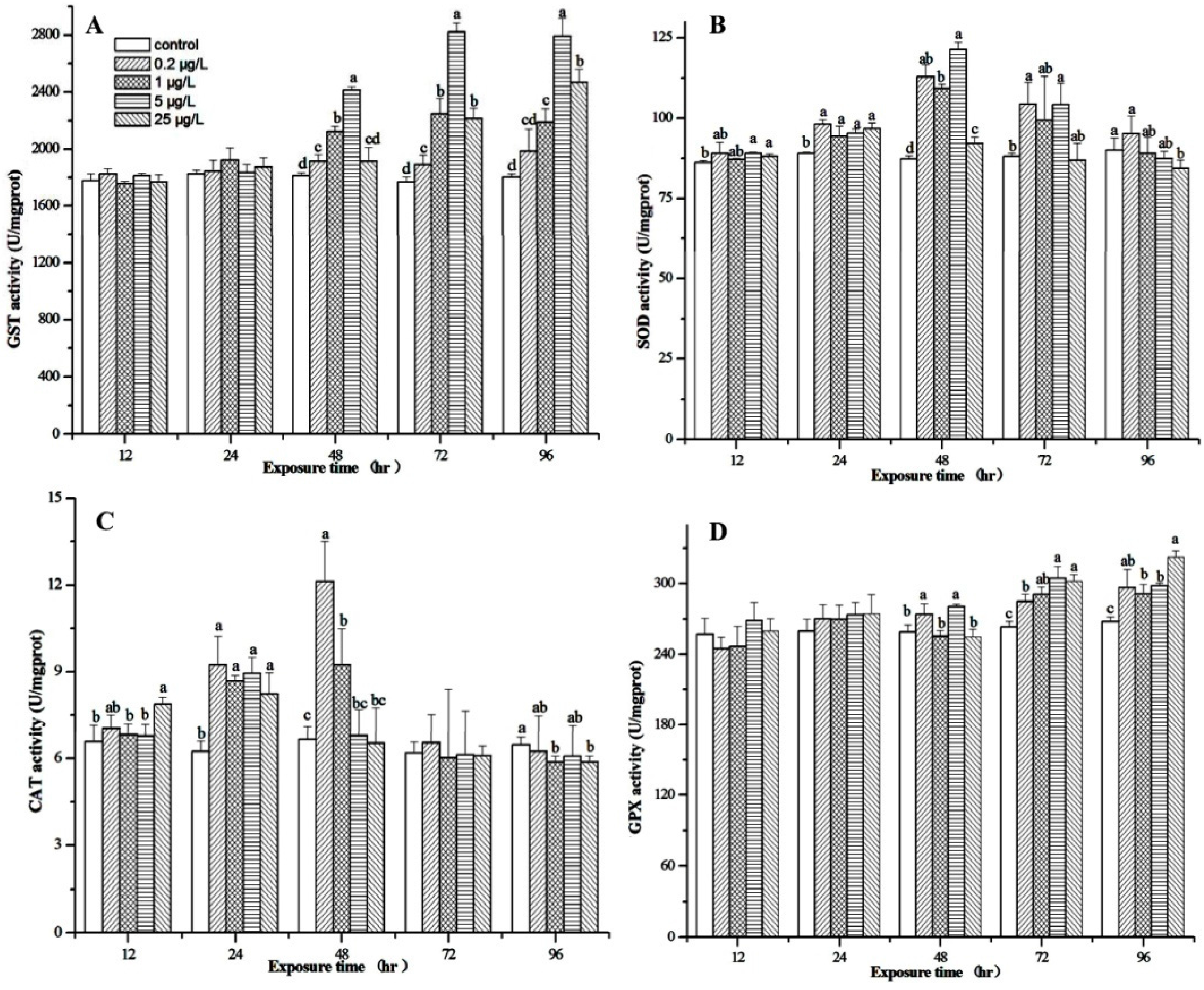
3.5. Expression Profiles of M. nipponense Gst, Cu/Zn-sod, Cat, and Gpx in the Hepatopancreas After MC-LR Exposure
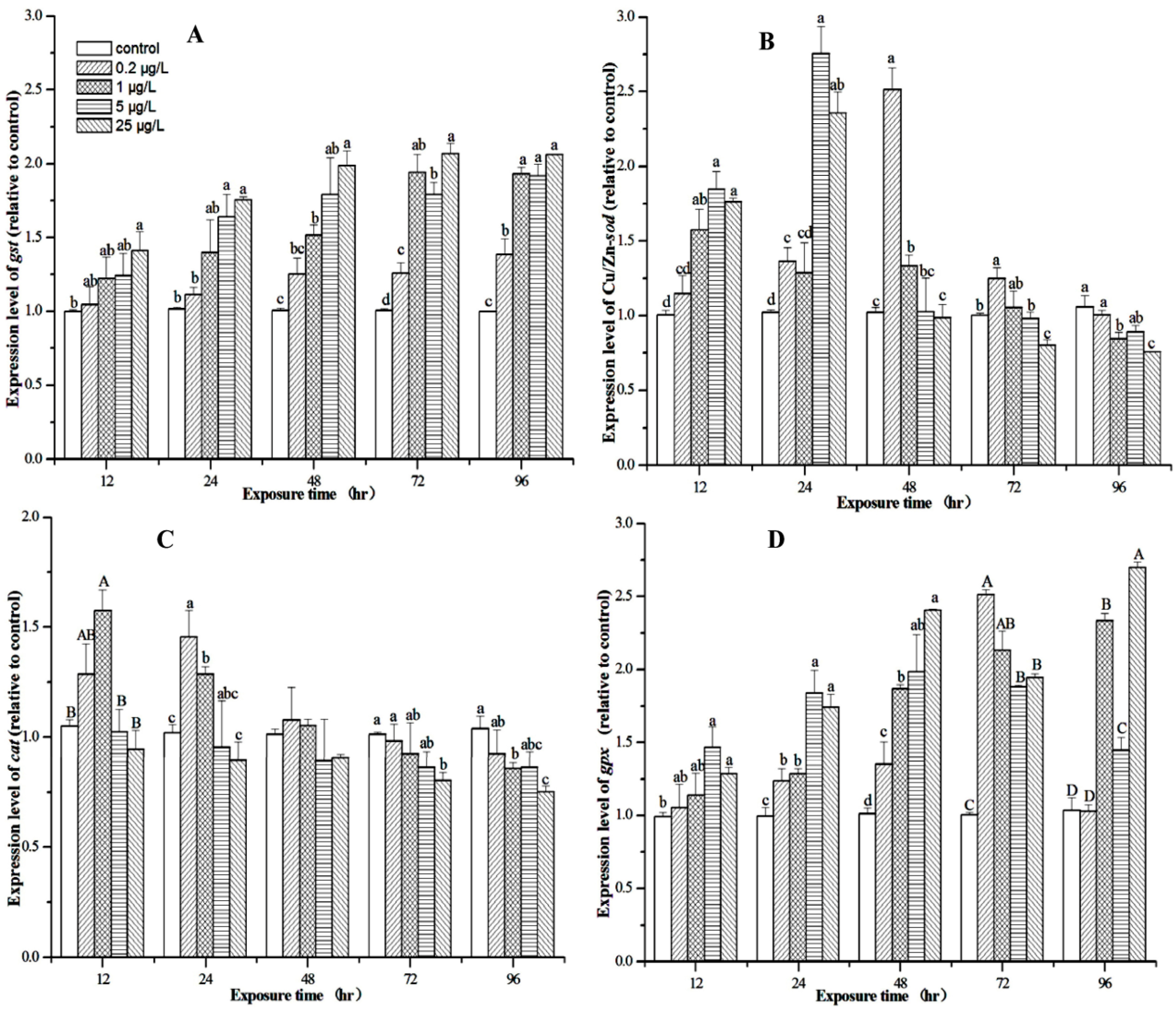
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Pearson, L.; Mihali, T.; Moffitt, M.; Kellmann, R.; Neilan, B. On the chemistry toxicology and genetics of the cyanobacterial toxins, microcystin, nodularin saxitoxin and cylindrospermopsin. Mar. Drugs 2010, 8, 1650–1680. [Google Scholar] [PubMed]
- Bieczynski, F.; Anna, J.D.; Pirez, M.; Brena, B.M.; Villanueva, S.S.M.; Luquet, C.M. Cellular transport of microcystin-LR in rainbow trout (Oncorhynchus mykiss) across the intestinal wall: Possible involvement of multidrug resistance-associated proteins. Aquat. Toxicol. 2014, 154, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Honkanen, R.E.; Codispoti, B.A.; Tse, K.; Boynton, A.L. Characterization of natural toxins with inhibitory activity against serine/threonine protein phosphatases. Toxicon 1994, 32, 339–350. [Google Scholar] [CrossRef]
- Wang, X.T.; Chen, Y.; Zuo, X.T.; Ding, N.Q.; Zeng, H.J.; Zou, X.; Han, X.D. Microcystin (-LR) induced cell apoptosis via up-regulating apoptosis-related genes in vivo. Food Chem. Toxicol. 2013, 60, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Van der Merwe, D.; Sebbag, L.; Nietfeld, J.C.; Aubel, M.T.; Foss, A.; Carney, E. Investigation of a Microcystis aeruginosa cyanobacterial freshwater harmful algal bloom associated with acute microcystin toxicosis in a dog. J. Vet. Diagn. Investig. 2012, 24, 679–687. [Google Scholar] [CrossRef] [PubMed]
- Kamjunke, N.; Mendonca, R.; Hardewig, I.; Mehner, T. Assimilation of different cyanobacteria as food and the consequences for internal energy stores of juvenile roach. J. Fish Biol. 2002, 60, 731–738. [Google Scholar] [CrossRef]
- Fischer, W.J.; Dietrich, P.J. Pathological and biochemical characterization of microcystin-induced hepatopancreas and kidney damage in carp (Cyprinus carpio). Toxicol. Appl. Pharm. 2000, 164, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Best, J.H.; Eddy, F.B.; Codd, G.A. Effect of purified microcystin-LR and cell extracts of Microcystis strains PCC7813 and CYA43 on cardiac function in brown trout (Salmo trutta) alevins. Fish Physiol. Biochem. 2001, 24, 171–178. [Google Scholar] [CrossRef]
- Qiu, T.; Xie, P.; Guo, L.; Zhang, D. Plasma biochemical response of the planktivorous filter-feeding silver carp (Hypophthalmichthys molitrix) and bighead carp (Aristichthys nobilis) to prolonged toxic cyanobacterial blooms in natural water. Environ. Toxicol. Pharmacol. 2009, 27, 350–356. [Google Scholar] [CrossRef] [PubMed]
- Cazenave, J.; Wunderlin, D.A.; Bistoni, M.; Amé, M.V.; Krause, E.P.; Flugmacher, S. Uptake, tissue, distribution and accumulation of microcystin-RR in Corydoras paleatus, Jenynsia multidentata and Odontesthes bonariensis: A field and laboratory study. Aquat. Toxicol. 2005, 75, 178–190. [Google Scholar] [CrossRef] [PubMed]
- Pires, L.M.D.; Kusserow, R.; Van, D.E. Influence of toxic and non-toxic phytoplankton on feeding and survival of Dreissena polymorpha (Pallas) larvae. Hydrobiologia 2003, 491, 193–200. [Google Scholar] [CrossRef]
- Juhel, G.; Davenport, J.; Ohalloran, J.; Culloty, S.; Ramsay, R.; James, K. Pseudodiarrhoea in zebra mussels Dreissena polymorpha (Pallas) exposed to microcystins. J. Exp. Biol. 2006, 209, 810–816. [Google Scholar] [CrossRef] [PubMed]
- DeMott, W.; Dhawale, S. Inhibition of protein phosphatase activity in three zooplankton species by microcystin-LR, a toxin from cyanobacteria. Arch. Hydrobiol. 1995, 34, 417–424. [Google Scholar]
- Winston, G.W.; Giulio, R.T. Prooxidant and antioxidant mechanisms in aquatic organisms. Aquat. Toxicol. 1991, 19, 137–161. [Google Scholar] [CrossRef]
- Cadenas, E. Biochemistry of oxygen toxicity. Annu. Rev. Biochem. 1989, 58, 79–110. [Google Scholar] [CrossRef] [PubMed]
- Prieto, A.I.; Pichardo, S.; Jos, Á.; Moreno, I.; Cameán, A.M. Time-dependent oxidative stress responses after acute exposure to toxic cyanobacterial cells containing microcystins in tilapia fish (Oreochromis niloticus) under laboratory condition. Aquat. Toxicol. 2007, 84, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Lü, K.; Minterc, E.J.A.; Chen, Y.; Yang, Z.; Montagnes, D.J.S. Combined effects of ammonia and microcystin on survival, growth, antioxidant responses, and lipid peroxidation of bighead carp Hypophthalmythys nobilis larvae. J. Hazard. Mater. 2012, 221–222, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Atencio, L.; Moreno, I.; Jos, A.; Pichardo, S.; Moyano, R.; Blanco, A.; Cameán, A.M. Dose-dependent antioxidant responses and pathological changes in tenca (Tinca tinca) after acute oral exposure to Microcystis under laboratory condition. Toxicon 2008, 52, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Chen, J.; Xie, P.; He, J.; Guo, X.C.; Tuo, X.; Zhang, W.; Wu, L.Y. Rapid conversion and reversible conjugation of glutathione detoxification of microcystins in bighead carp (Aridtichthys nobilis). Aquat. Toxicol. 2014, 147, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Pflugmacher, S.; Wiegand, C.; Oberemm, A.; Beattie, K.A.; Krause, E.; Codd, G.A. Identification of an enzymatically formed glutathione conjugate of the cyanobacterial hepatoxin microcystin-LR: The first step of detoxication. Biochim. Biophys. Acta 1998, 1425, 527–533. [Google Scholar] [CrossRef]
- Wang, H.; Wang, J.; Wu, T.; Qin, F.; Hu, X.; Wang, L.; Wang, Z. Molecular characterization of estrogen receptor genes in Gobiocypris rarus and their expression upon endocrine disrupting chemicals exposure in juveniles. Aquat. Toxicol. 2011, 101, 276–287. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The ClustalX windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method-a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, R. Quantification on the Light Cycler. In Rapid Cycle Real-Time PCR, Methods and Applications; Meuer, S., Wittwer, C., Nakagawara, K., Eds.; Springer Press: Heidelberg, Germany, 2001; pp. 21–34. [Google Scholar]
- Sun, S.; Ge, X.; Fu, H.; Zhu, J.; Zhang, S.; Qiao, H. Molecular cloning and expression analysis of the hemocyanin gene from oriental river prawn (Macrobrachium nipponense). J. Fish. Sci. China 2014, 21, 229–238. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Li, J.; Liu, P.; Gao, B.; Wang, Q.; Li, J. cDNA cloning, characterization and expression analysis of catalase in swimming crab Portunus trituberculatus. Mol. Biol. Rep. 2012, 39, 9979–9987. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Li, F.; Zhang, X.; Dong, B.; Zhang, J.; Xie, Y.; Xiang, J. cDNA cloning, characterization and expression analysis of the antioxidant enzyme gene, catalase of Chinese shrimp Fenneropenaeus chinensis. Fish Shellfish Immunol. 2008, 24, 584–591. [Google Scholar] [CrossRef] [PubMed]
- Tavares-Sánchez, O.L.; Gómez-Anduro, G.A.; Felipe-Orteg, X.; Islas-Osuna, M.A.; Sotelo-Mundo, R.R.; Barillas-Mury, C.; Yepiz-Plascencia, G. Catalase from the white shrimp Penaeus (Litopenaeus) vannamei: Molecular cloning and protein detection. Comp. Biochem. Physiol. B 2004, 138, 331–337. [Google Scholar] [CrossRef] [PubMed]
- Arockiaraj, J.; Easwvaran, S.; Vanaraja, P.; Singh, A.; Othman, R.Y.; Bhassu, S. Molecular cloning, characterization and gene expression of an antioxidant enzyme catalase (MrCat) from Macrobrachium rosenbergii. Fish Shellfish Immunol. 2012, 32, 670–682. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Li, G.; Wen, C.; Hu, B.; Deng, L.; Pei, P.; Xie, Y. A catalase from the freshwater mussel Cristaria plicata with cloning, identification and protein characterization. Fish Shellfish Immunol. 2011, 31, 389–399. [Google Scholar] [PubMed]
- Hu, B.; Deng, L.; Wen, C.; Yang, X.; Pei, P.; Xie, Y.; Luo, S. Cloning, identification and functional characterization of a pi-class glutathione-S-transferase from the freshwater mussel Cristaria plicata. Fish Shellfish Immunol. 2012, 32, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Glisic, B.; Mihaljevic, I.; Popovic, M.; Zaja, R.; Loncar, J.; Fent, K.; Kovacevic, R.; Smital, T. Characterization of glutathione-S-transferases in zebrafish (Danio rerio). Aquat. Toxicol. 2015, 158, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Won, E.J.; Kim, R.O.; Rhee, J.S.; Park, G.S.; Lee, J.; Shin, K.H.; Lee, Y.M.; Lee, J.S. Response of glutathione S-transferase (GST) gene to cadmium exposure in the marine pollution indicator worm, Perinereis nuntia. Comp. Biochem. Physiol. C 2011, 154, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Carmen, A.; Contreras, V.; Citlalli, H.V.; Rogerio, R.; Sotelo, M.; Gloria, Y.P. A mu-class glutathione S-transferase from the marine shrimp Litopenaeus vannamei: Molecular cloning and active-site structure modeling. J. Biochem. Mol. Toxicol. 2004, 218, 245–252. [Google Scholar]
- Yu, Y.; Liang, X.-F.; Li, L.; He, S.; Wen, Z.-Y.; Shen, D. Two homologs of rho-class and polymorphism in alpha-class glutathione S-transferase genes in the liver of three tilapias. Ecotoxicol. Environ. Saf. 2014, 101, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, D.R. Contaminant-stimulated reactive oxygen species production and oxidative damage in aquatic organisms. Mar. Pollut. Bull. 2001, 42, 656–666. [Google Scholar] [CrossRef]
- Li, X.; Liu, Y.; Song, L.; Liu, L. Response of antioxidant systems in the hepatocytes of common carp (Cyprinus carpio L.) to the toxicity of mcrocystin-LR. Toxicon 2003, 42, 85–89. [Google Scholar] [CrossRef]
- Amado, L.L.; Monserrat, J.M. Oxidative stress generation by microcystins in aquatic animals: Why and how. Environ. Int. 2010, 36, 226–235. [Google Scholar] [CrossRef] [PubMed]
- Pandey, S.; Parvez, S.; Sayeed, I.; Haque, R.; Bin-Hanfeez, B.; Raisuddin, S. Biomarker of oxidative stress: A comparative study of river Yamuna fish Wallago attu (Bl. & Schn.). Sci. Total Environ. 2003, 309, 105–115. [Google Scholar] [PubMed]
- Velma, V.; Tchounwou, P.B. Hexavalent chromium-induced multiple biomarker responses in liver and kidney of goldfish, Carassius auratus. Environ. Toxicol. 2011, 26, 649–656. [Google Scholar] [CrossRef] [PubMed]
- Burmester, V.; Nimptsch, J.; Wiegand, C. Adaptation of freshwater mussels to cyanobacterial toxins: Response of the biotransformation and antioxidant enzymes. Ecotoxicol. Environ. Saf. 2012, 78, 296–309. [Google Scholar] [CrossRef] [PubMed]
- Pigeolet, E.; Corbisier, P.; Houbion, A.; Lambert, D.; Michiels, C.; Raes, M.; Zachary, M.D.; Remacle, J. Glutathione peroxidase, superoxide dismutase, and catalase inactivation by peroxides and oxygen derived free radicals. Mech. Ageing Dev. 1990, 51, 283–297. [Google Scholar] [CrossRef]
- Chance, B.; Sies, H.; Boveris, A. Hydroperoxide metabolism in mammalian organs. Physiol. Rev. 1979, 59, 527–605. [Google Scholar] [PubMed]
- Pinho, G.L.L.; da Rosa, C.M.; Maciel, F.E.; Bianchini, A.; Yunes, J.S.; Proenca, L.A.O.; Monserrat, J.M. Antioxidant responses and oxidative stress after microcystin exposure in the hepatopancreas of an eatuarine crab species. Ecotoxicol. Environ. Saf. 2005, 61, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Escobar, J.A.; Rubio, M.A.; Lissi, E.A. SOD and catalase inactivation by singlet oxygen and peroxyl radicals. Free Radic. Biol. Med. 1996, 20, 285–290. [Google Scholar] [CrossRef]
- Takenaka, S. Covalent glutathione to cyanobacterial hepatotoxin microcystin LR by F344 rat cytosolic and microsomal glutathione S-transferases. Environ. Toxicol. Pharm. 2001, 9, 135–139. [Google Scholar] [CrossRef]
- Ito, E.; Takai, A.; Kondo, F.; Masui, H.; Imanishi, S.; Harada, K. Comparison of protein phosphatase inhibitory activity and apparent toxicity of microcystins and related compounds. Toxicon 2002, 40, 1017–1025. [Google Scholar] [CrossRef]
- Hou, J.; Li, L.; Xue, T.; Long, M.; Su, Y.J.; Wu, N. Hepatic positive and negative antioxidant responses in zebrafish after intraperitoneal administration of toxic microcystin-LR. Chemosphere 2015, 120, 729–736. [Google Scholar] [CrossRef] [PubMed]
- Galanti, L.N.; Amé, M.V.; Wunderlin, D.A. Accumulation and detoxification dynamics of cyanotoxins in the freshwater shrimp Palaemonetes argentinus. Harmful Algae 2013, 27, 89–97. [Google Scholar] [CrossRef]
- Sabatni, S.E.; Brena, B.M.; Pirez, M.; Molina, M.C.R.; Luquet, C.M. Oxidative effects and toxin bioaccumulation after dietary microcystin intoxication in the hepayopancreas of the crab Neohelice (Chasmagnathus) granulata. Ecotoxicol. Environ. Saf. 2015, 120, 136–141. [Google Scholar] [CrossRef] [PubMed]
- Bieczynski, F.; Bianchi, V.A.; Luquet, C.M. Accumulation and biochemical effects of microcystin-LR on the Patagonianpe jerrey (Odontesthes hatcheri) fed with the toxic cyanobacteria Microcystis aeruginosa. Fish Physiol. Biochem. 2013, 39, 1309–1321. [Google Scholar] [CrossRef] [PubMed]
- Ríos, V.; Guzmán-Guilén, R.; Moreno, I.M.; Prieto, A.I.; Puerto, M.; Jos, A.; Cameán, A.M. Influence of two depuration periods on the activity and transcription of antioxidant enzymes in tilapia exposed to repeated doses of cylindrospermopsin under laboratory conditions. Toxins 2014, 6, 1062–1079. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Liang, X.; Sun, J.; Shen, D. Induction of liver GST transcriptions by tert-butylhydroquinone reduced microcystin-LR accumulation in Nile tilapia (Oreochromis niloticus). Ecotoxicol. Environ. Saf. 2013, 90, 128–135. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Xie, P.; Fu, J.; Hao, L.; Xiong, Q.; Li, H. Microcystin-induced variations in transcription of GSTs in an omnivorous freshwater fish, goldfish. Aquat. Toxicol. 2008, 88, 75–80. [Google Scholar] [CrossRef]
- Amado, L.L.; Garcia, M.L.; Pereira, T.C.B.; Yunes, J.S.; Bogo, M.R.; Monserrat, J.M. Chemoprotection of lipoic acid against microcystin-induced toxicosis in common carp (Cyprinus carpio, Cyprinidae). Comp. Biochem. Physiol. C 2011, 154, 146–153. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yuan, J.; Wang, X.; Gu, Z.; Zhang, Y.; Wang, Z. Activity and Transcriptional Responses of Hepatopancreatic Biotransformation and Antioxidant Enzymes in the Oriental River Prawn Macrobrachium nipponense Exposed to Microcystin-LR. Toxins 2015, 7, 4006-4022. https://doi.org/10.3390/toxins7104006
Yuan J, Wang X, Gu Z, Zhang Y, Wang Z. Activity and Transcriptional Responses of Hepatopancreatic Biotransformation and Antioxidant Enzymes in the Oriental River Prawn Macrobrachium nipponense Exposed to Microcystin-LR. Toxins. 2015; 7(10):4006-4022. https://doi.org/10.3390/toxins7104006
Chicago/Turabian StyleYuan, Julin, Xueqin Wang, Zhiming Gu, Yingying Zhang, and Zaizhao Wang. 2015. "Activity and Transcriptional Responses of Hepatopancreatic Biotransformation and Antioxidant Enzymes in the Oriental River Prawn Macrobrachium nipponense Exposed to Microcystin-LR" Toxins 7, no. 10: 4006-4022. https://doi.org/10.3390/toxins7104006
APA StyleYuan, J., Wang, X., Gu, Z., Zhang, Y., & Wang, Z. (2015). Activity and Transcriptional Responses of Hepatopancreatic Biotransformation and Antioxidant Enzymes in the Oriental River Prawn Macrobrachium nipponense Exposed to Microcystin-LR. Toxins, 7(10), 4006-4022. https://doi.org/10.3390/toxins7104006




