Omics Meets Biology: Application to the Design and Preclinical Assessment of Antivenoms
Abstract
:1. Introduction: Snakebite Envenoming and the Challenge of Generating Effective Antivenoms
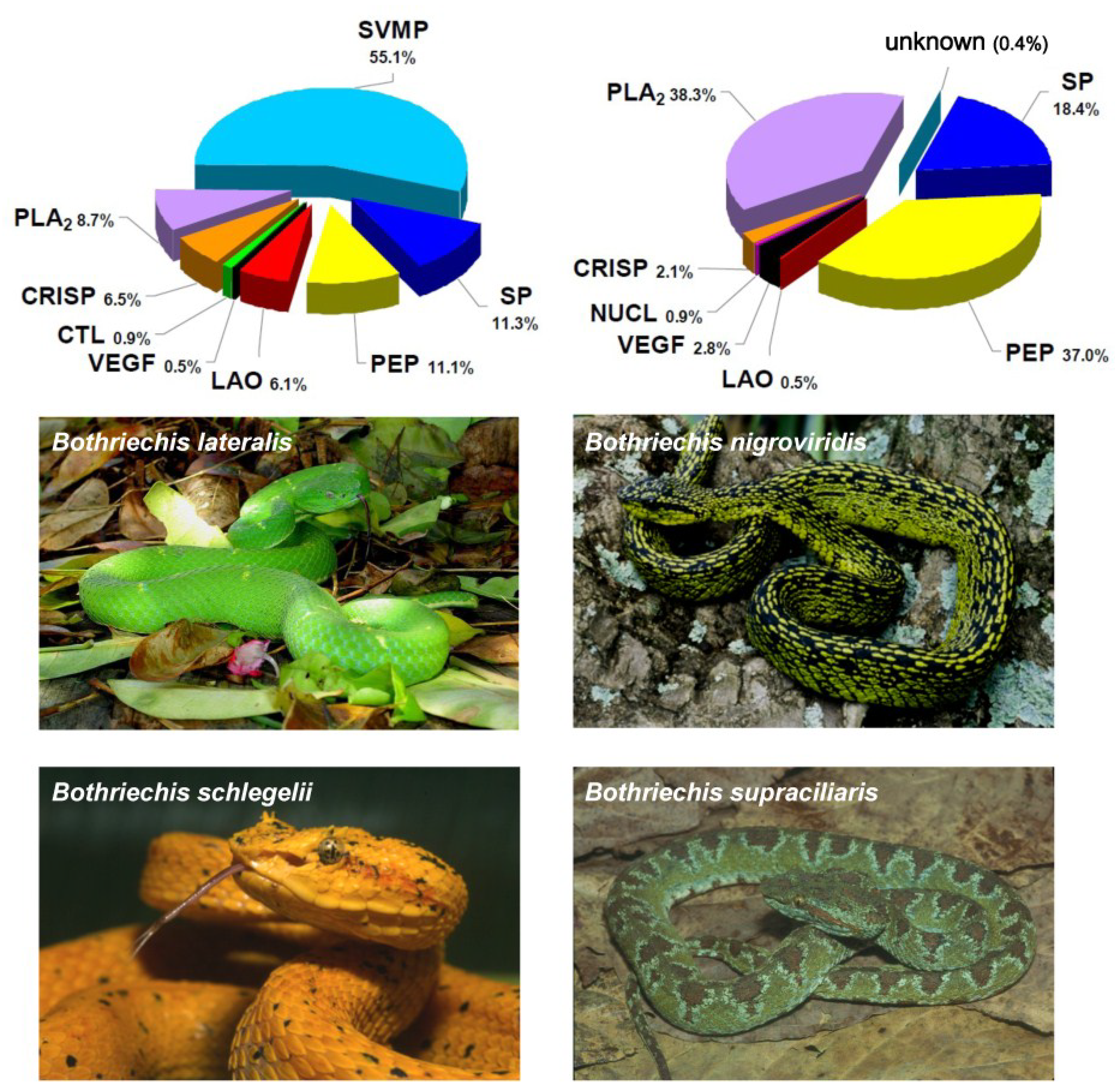
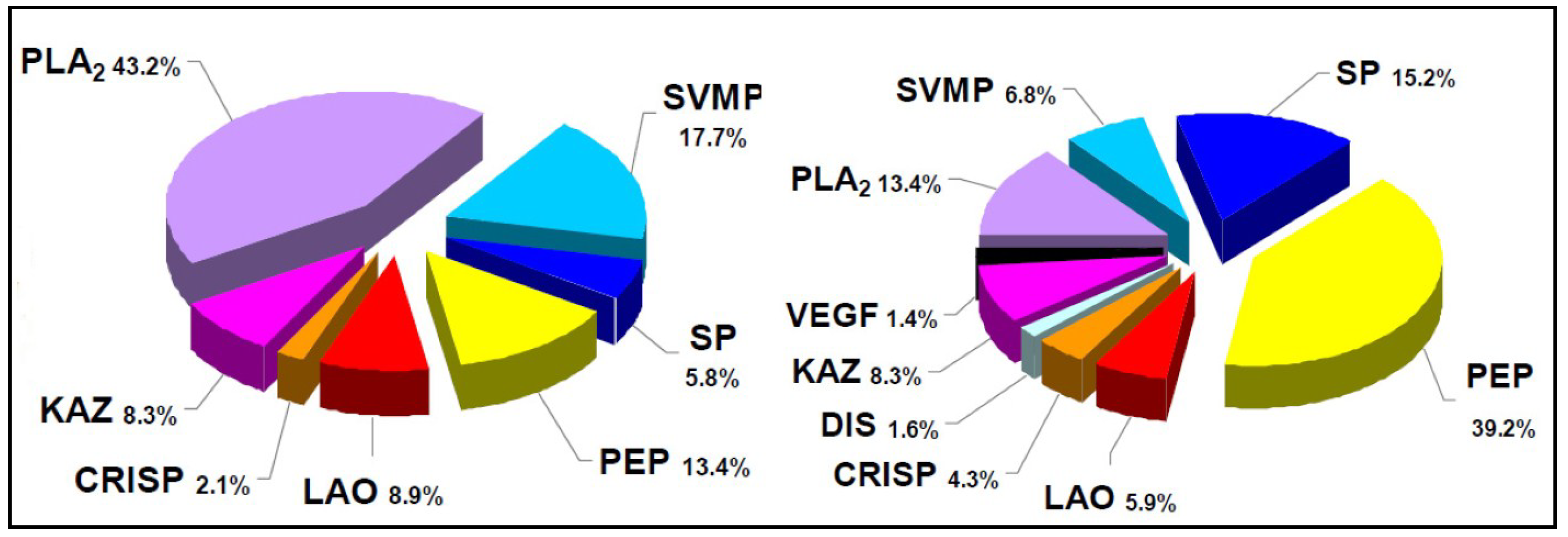
2. Biochemical and Toxinological Toolbox for the Preclinical Assessment of Antivenom Efficacy
- (1)
- Hemorrhagic activity: The most widely-used method is based on the intradermal injection of venom solutions, followed, several hours later, by the measurement of the area of the hemorrhagic spot in the inner side of the skin [36,44,45]. Venom activity is expressed as the minimum hemorrhagic dose (MHD), which corresponds to the dose of venom that induces a hemorrhagic halo of 10 mm in diameter. More recently, the analysis of systemic, i.e., pulmonary, hemorrhage has been introduced. Mice are injected i.v. with venom, and one hour later, the animals are sacrificed and the thoracic cavity exposed for observation of hemorrhagic spots on the surface of the lungs. The minimum pulmonary hemorrhagic dose (MPHD) corresponds to the lowest amount of venom that induces hemorrhagic spots in the lungs of all mice injected [46].
- (2)
- Myotoxic activity: Venom-induced skeletal muscle necrosis can be assessed by histological examination of muscle tissue injected with venom. Mice receiving an intramuscular injection of venom solution, for example in the gastrocnemius muscle, are sacrificed 24 h later, and the injected muscle is processed for histological analysis. The number of necrotic cells and the total number of muscle cells are quantified by microscopic assessment, and the myotoxic effect is expressed as the necrotic index, i.e., the ratio of necrotic muscle fibers to total muscle fibers [47]. Venom activity can be expressed as the dose inducing a necrotic index of 0.5. Since histological analysis is time consuming and not all laboratories have facilities for tissue processing for histology, a convenient alternative for the histological analysis is the quantification of the activity of the plasmatic enzyme, creatine kinase (CK), which is released from muscle fibers when the plasma membrane of muscle cells is disrupted. Mice are injected intramuscularly, as described, and a blood sample is collected usually 3 h after injection. After separation of plasma by centrifugation, the plasma CK activity is quantified by using commercial kits. The minimum myotoxic dose (MMD) is defined as the dose of venom that increases the plasma CK activity four times as compared to mice injected with saline solution [48].
- (3)
- Dermonecrotic activity: This effect is assessed in either rats or mice by carrying out intradermal injections of venom solutions, followed by the measurement of the necrotic area in the inner side of the skin 72 h after venom injection [36].
- (4)
- Coagulant activity: In vitro coagulant activity of venoms is assessed by the addition of various doses of venom to samples of citrated human plasma, obtained from healthy donors, followed by the determination of clotting time. Activity is expressed as the minimum coagulant dose (MCD), defined as the dose of venom that induces clotting in 60 seconds [36,39]. For assessing the thrombin-like activity of venoms, a similar test is performed on fibrinogen solutions instead of plasma [36].
- (5)
- Defibrinogenating activity: This is assessed in rats or mice by intravenous injection of venom solutions. After a defined period of time, a blood sample is collected, placed in a glass tube and incubated at room temperature for observation of clot formation. Activity is expressed as the minimum defibrinogenating dose (MDD), defined as the dose of venom that induces incoagulability, i.e., blood remains unclottable, in all animals injected [36,39].
- (6)
- Other tests: Assays for the determination of other toxic activities, such as edema-forming activity, ex vivo neurotoxic activity and thrombocytopenic effect, have been described [49,50,51]. The analysis of the neutralization of venom enzyme activities, including neutralization of proteinase, phospholipase A2 and hyaluronidase activities, has been also investigated [41,52,53]. Similarly to the neutralization of lethality, for the analysis of the neutralization of venom enzyme activities, various mixtures of venom:antivenom incubated for 30 min at 37 °C are tested, and neutralization of venom activity is expressed as ED50, i.e., the antivenom/venom ratio in which the effect of venom is neutralized by 50% [54]. In the case of coagulant and defibrinogenating activities, neutralization is expressed as the effective dose (ED), defined as the antivenom/venom ratio at which the clotting time of plasma is prolonged three times when compared to plasma incubated with venom alone (for coagulant activity) or as the antivenom/venom ratio in which blood clots form in all animals injected (for defibrinogenating effect) [39].
3. Omics Toolbox for the Preclinical Assessment of Antivenom Efficacy: The Venomics-Antivenomics Platform
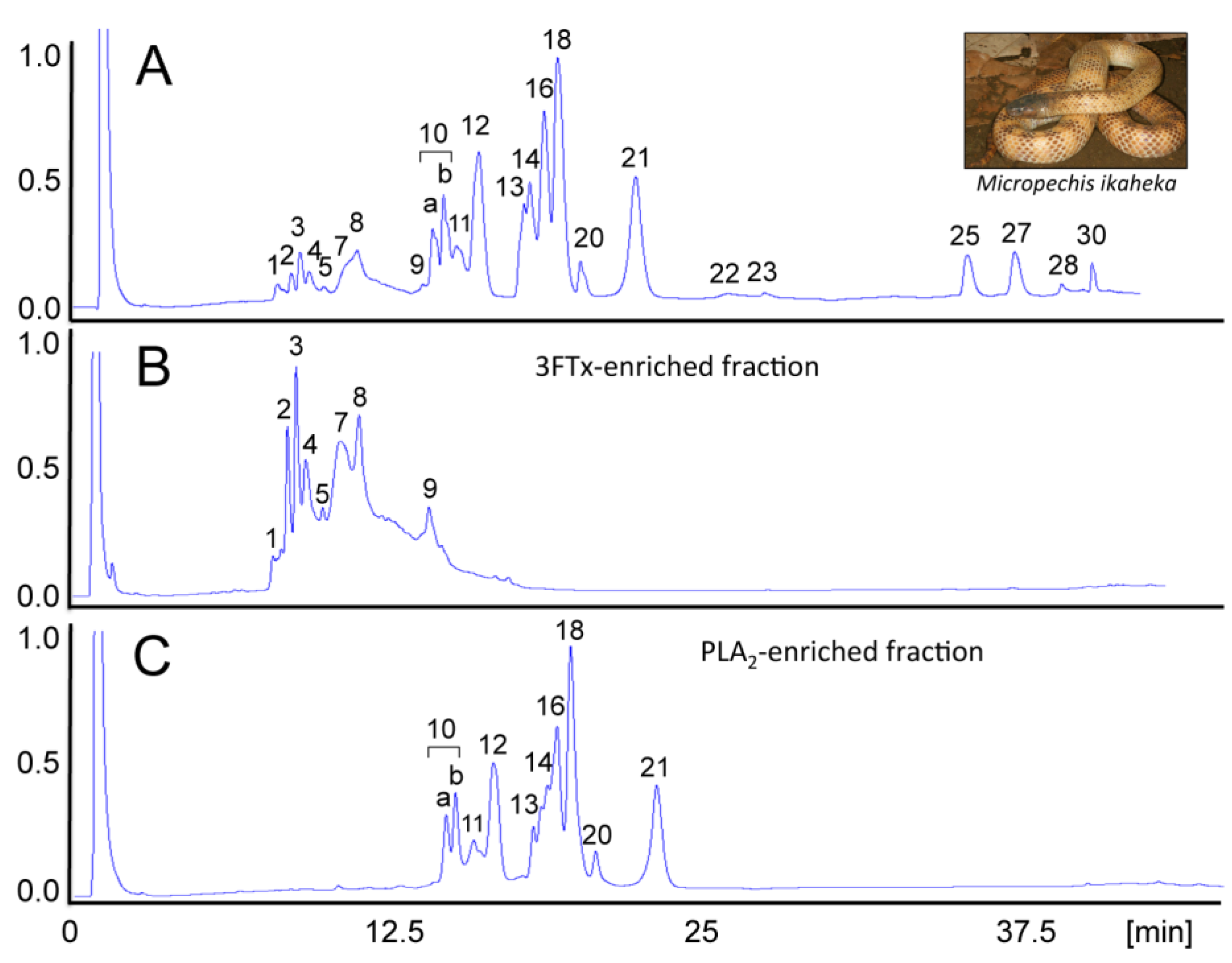
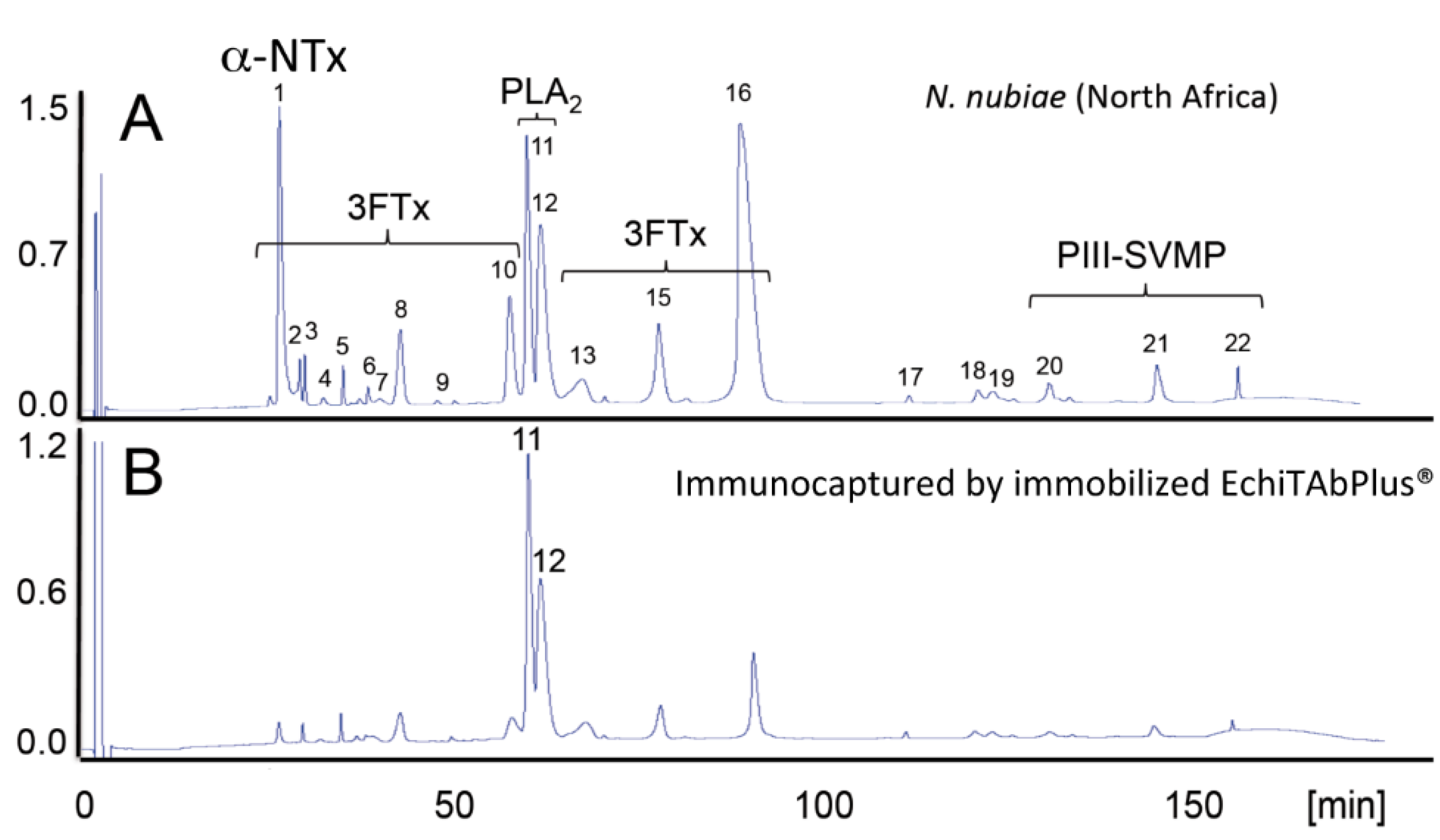
4. Concluding Remarks
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kasturiratne, A.; Wickremasinghe, A.R.; de Silva, N.; Gunawardena, N.K.; Pathmeswaran, A.; Premaratna, R.; Savioli, L.; Lalloo, D.G.; de Silva, H.J. The global burden of snakebite: A literature analysis and modeling based on regional estimates of envenoming and deaths. PLoS Med. 2008, 5, e218. [Google Scholar] [CrossRef] [PubMed]
- Harrison, R.A.; Hargreaves, A.; Wagstaff, S.C.; Faragher, B.; Lalloo, D.G. Snake envenoming: A disease of poverty. PLoS Negl. Trop. Dis. 2009, 3, e569. [Google Scholar] [CrossRef] [PubMed]
- Williams, D.J.; Gutiérrez, J.M.; Calvete, J.J.; Wüster, W.; Ratanabanangkoon, K.; Paiva, O.; Brown, N.I.; Casewell, N.R.; Harrison, R.A.; Rowley, P.D.; et al. Ending the drought: New strategies for improving the flow of affordable, effective antivenoms in Asia and Africa. J. Proteomics 2011, 74, 1735–1767. [Google Scholar] [CrossRef]
- Warrell, D.A.; Gutiérrez, J.M.; Calvete, J.J.; Williams, D. New approaches and technologies of venomics to meet the challenge of human envenoming by snakebites in India. Indian J. Med. Res. 2013, 138, 38–59. [Google Scholar] [PubMed]
- Gutiérrez, J.M.; Williams, D.J.; Fan, H.W.; Warrell, D.A. Snakebite envenoming from a global perspective: Towards an integrated approach. Toxicon 2010, 56, 1223–1235. [Google Scholar]
- Gutiérrez, J.M.; León, G.; Burnouf, T. Antivenoms for the treatment of snakebite envenomings: The road ahead. Biologicals 2011, 39, 129–142. [Google Scholar] [CrossRef] [PubMed]
- Harrison, R.A.; Cook, D.A.; Renjifo, C.; Casewell, N.R.; Currier, R.B.; Wagstaff, S.C. Research strategies to improve snakebite treatment: Challenges and progress. J. Proteomics 2011, 74, 1768–1780. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M. Improving antivenom availability and accessibility: Science, technology, and beyond. Toxicon 2012, 60, 676–687. [Google Scholar] [CrossRef] [PubMed]
- Lalloo, D.G.; Theakston, R.D.G. Snake antivenoms. J. Toxicol. Clin. Toxicol. 2003, 41, 277–290. [Google Scholar] [CrossRef] [PubMed]
- Calmette, A. L'immunisation artificielle des animaux contre le venin des serpents, et la thérapeutic expérimentale des morsures venimeuses. Compte. Rendus Soc. Biol. 1894, 46, 120–124. [Google Scholar]
- Calmette, A. Contribution à l'étude du venin des serpents. Immunisation des animaux et traitement de l'envenimation. Ann. l'Institut Pasteur 1894, 8, 275–291. (In French) [Google Scholar]
- Hawgood, B.J. Doctor Albert Calmette 1863–1933: Founder of antivenomous serotherapy and of antituberculous BCG vaccination. Toxicon 1999, 37, 1241–1258. [Google Scholar] [CrossRef] [PubMed]
- Phisalix, C.; Bertrand, G. Sur la propriété antitoxique du sang des animaux vaccinée contre le venin de vipére. Compte. Rendus Soc. Biol. 1894, 46, 111–113. [Google Scholar]
- Phisalix, C.; Bertrand, G. Propriétés antitoxique du sang des animaux vaccineé contre le venin de vipère. Contribution à l'étude du mécanisme de la vaccination contre ce venin. Arch. Physiol. 1894, 6, 611–619. (In French) [Google Scholar]
- World Health Organization: Venomous snakes and antivenoms search interface. Available online: http://apps.who.int/bloodproducts/snakeantivenoms/database (accessed on 11 December 2014).
- Global Snakebite Initiative. Available online: http://www.snakebiteinitiative.org (accessed on 11 December 2014).
- Williams, D.J.; Gutiérrez, J.M.; Harrison, R.; Warrell, D.A.; White, J.; Winkel, K.D.; Gopalakrishnakone, P.; Global Snakebite Initiative Working Group; International Society on Toxinology. The Global Snakebite Initiative: an antidote for snakebite. Lancet 2010, 375, 89–91. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Warrell, D.A.; Williams, D.J.; Jensen, S.; Brown, N.; Calvete, J.J.; Harrison, R.A.; Global Snakebite Initiative. The need for full integration of snakebite envenoming within a global strategy to combat the neglected tropical diseases: the way forward. PLoS Negl. Trop. Dis. 2013, 7, e2162. [Google Scholar] [CrossRef]
- Gutiérrez, J.M.; Burnouf, T.; Harrison, R.A.; Calvete, J.J.; Kuch, U.; Warrell, D.A.; Williams, D.J.; Global Snakebite Initiative. A multicomponent strategy to improve the availability of antivenom for treating snakebite envenoming. Bull. World Health Organ. 2014, 92, 526–532. [Google Scholar] [CrossRef] [PubMed]
- Georgieva, D.; Arni, R.K.; Betzel, C. Proteome analysis of snake venom toxins: pharmacological insights. Expert Rev. Proteomics 2008, 5, 787–797. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J. Proteomic tools against the neglected pathology of snakebite envenoming. Expert Rev. Proteomics 2011, 8, 739–758. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Solano, G.; Pla, D.; Herrera, M.; Segura, A.; Villalta, M.; Vargas, M.; Sanz, L.; Lomonte, B.; Calvete, J.J.; León, G. Assessing the preclinical efficacy of antivenoms: From the lethality neutralization assay to antivenomics. Toxicon 2013, 69, 168–179. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J. Snake venomics: from the inventory of toxins to biology. Toxicon 2013, 75, 44–62. [Google Scholar] [CrossRef] [PubMed]
- Chippaux, J.-P.; Williams, V.; White, J. Snake venom variability: Methods of study, results and interpretation. Toxicon 1991, 29, 1279–1303. [Google Scholar] [CrossRef] [PubMed]
- Massey, D.J.; Calvete, J.J.; Sánchez, E.E.; Sanz, L.; Richards, K.; Curtis, R.; Boesen, K. Venom variability and envenoming severity outcomes of the Crotalus scutulatus scutulatus (Mojave rattlesnake) from Southern Arizona. J. Proteomics 2012, 75, 2576–2587. [Google Scholar] [CrossRef] [PubMed]
- Durban, J.; Pérez, A.; Sanz, L.; Gómez, A.; Bonilla, F.; Rodríguez, S.; Chacón, D.; Sasa, M.; Angulo, Y.; Gutiérrez, J.M.; Calvete, J.J. Integrated “omics” profiling indicates that miRNAs are modulators of the ontogenetic venom composition shift in the Central American rattlesnake, Crotalus simus simus. BMC Genomics 2013, 14, 234. [Google Scholar] [CrossRef] [PubMed]
- Casewell, N.R.; Wagstaff, S.C.; Wüster, W.; Cook, D.A.; Bolton, F.M.; King, S.I.; Pla, D.; Sanz, L.; Calvete, J.J.; Harrison, R.A. Medically important differences in snake venom composition are dictated by distinct postgenomic mechanisms. Proc. Natl. Acad. Sci. USA 2014, 111, 9205–9210. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, H.L.; Sanz, L.; Sovic, M.G.; Calvete, J.J. Phylogeny-based comparative analysis of venom proteome variation in a clade of rattlesnakes (Sistrurus sp.). PLoS One 2013, 8, e67220. [Google Scholar] [CrossRef] [PubMed]
- Lomonte, B.; Tsai, W.C.; Ureña-Diaz, J.M.; Sanz, L.; Mora-Obando, D.; Sánchez, E.E.; Fry, B.G.; Gutiérrez, J.M.; Gibbs, H.L.; Sovic, M.G.; Calvete, J.J. Venomics of New World pit vipers: Genus-wide comparisons of venom proteomes across Agkistrodon. J. Proteomics 2014, 96, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Mackessy, S.P. Venom composition in rattlesnakes: Trends and biological significance. In The Biology of Rattlesnakes; Hayes, H.K., Beaman, K.R., Cardwell, M.D., Bush, S.P., Eds.; Loma Linda University Press: Loma Linda, CA, USA, 2008; pp. 495–510. [Google Scholar]
- Fernández, J.; Lomonte, B.; Sanz, L.; Angulo, Y.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics of Bothriechis nigroviridis reveals extreme variability among palm pitviper venoms: different evolutionary solutions for the same trophic purpose. J. Proteome Res. 2009, 9, 4234–4241. [Google Scholar] [CrossRef]
- Lomonte, B.; Fernández, J.; Sanz, L.; Angulo, Y.; Sasa, M.; Gutiérrez, J.M.; Calvete, J.J. Venomous snakes of Costa Rica: biological and medical implications of their venom proteomic profiles analyzed through the strategy of snake venomics. J. Proteomics 2014, 105, 323–339. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Guidelines for the Production, Control and Regulation of Snake Antivenom Immunoglobulins. Available online: www.who.int/bloodproducts/snakeantivenoms (accessed on 11 December 2014).
- Theakston, R.D.G. Characterization of venoms and standardization of antivenoms. In Natural Toxins. Animal, Plant and Microbial; Harris, J.B., Ed.; Clarendon Press: Oxford, UK, 1986; pp. 287–303. [Google Scholar]
- Gutiérrez, J.M.; Rojas, G.; Bogarín, G.; Lomonte, B. Evaluation of the neutralizing ability of antivenoms for the treatment of snakebite envenoming in Central America. In Envenomings and Their Treatments; Bon, C., Goyffon, M., Eds.; Fondation Marcel Mérieux: Lyon, France, 1996; pp. 223–231. [Google Scholar]
- Theakston, R.D.G.; Reid, H.A. Development of simple standard assay procedures for the characterization of snake venom. Bull. World Health Org. 1983, 61, 949–956. [Google Scholar] [PubMed]
- Theakston, R.D.G.; Warrell, D.A.; Griffiths, E. Report of a WHO workshop on the standardization and control of antivenoms. Toxicon 2003, 41, 541–557. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Arroyo, O.; Bolaños, R. Mionecrosis, hemorragia y edema inducidos por el veneno de Bothrops asper en ratón blanco. Toxicon 1980, 18, 603–610. [Google Scholar] [CrossRef] [PubMed]
- Gené, J.A.; Roy, A.; Rojas, G.; Gutiérrez, J.M.; Cerdas, L. Comparative study on coagulant, defibrinating, fibrinolytic and fibrinogenolytic activities of Costa Rican crotaline snake venoms and their neutralization by a polyvalent antivenom. Toxicon 1989, 27, 841–848. [Google Scholar] [CrossRef] [PubMed]
- Warrell, D.A. Snakebite. Lancet 2010, 375, 77–88. [Google Scholar] [CrossRef] [PubMed]
- Warrell, D.A. Clinical toxicology of snakebite in Africa and the Middle East/Arabian Peninsula. In Handbook of Clinical Toxicology of Animal Venoms and Poisons; Meier, J., White, J., Eds.; CRC Press: Boca Raton, FL, USA, 1995; pp. 433–492. [Google Scholar]
- Petras, D.; Sanz, L.; Segura, A.; Herrera, M.; Villalta, M.; Solano, D.; Vargas, M.; León, G.; Warrell, D.A.; Theakston, R.D.G.; Harrison, R.A.; et al. Snake venomics of African spitting cobras: Toxin composition and assessment of congeneric cross-reactivity of the pan-African EchiTAb-Plus-ICP® antivenom by antivenomics and neutralization approaches. J. Proteome Res. 2011, 10, 1266–1280. [Google Scholar] [CrossRef] [PubMed]
- Vargas, M.; Segura, A.; Herrera, M.; Villalta, M.; Estrada, R.; Cerdas, M.; Paiva, O.; Matainaho, T.; Jensen, S.D.; Winkel, K.D.; et al. Preclinical evaluation of caprylic acid-fractionated IgG antivenom for the treatment of Taipan (Oxyuranus scutellatus) envenoming in Papua New Guinea. PLoS Negl. Trop. Dis. 2011, 5, e1144. [Google Scholar] [CrossRef] [PubMed]
- Kondo, H.; Kondo, S.; Ikezawa, I.; Murata, R.; Ohsaka, A. Studies of the quantitative method for the determination of hemorrhagic activity of Habu snake venom. Japan J. Med. Sci. Biol. 1960, 13, 43–51. [Google Scholar] [CrossRef]
- Gutiérrez, J.M.; Gené, J.A.; Rojas, G.; Cerdas, L. Neutralization of proteolytic and hemorrhagic activities of Costa Rican snake venoms by a polyvalent antivenom. Toxicon 1985, 23, 887–893. [Google Scholar] [CrossRef] [PubMed]
- Escalante, T.; Núñez, J.; Moura da Silva, A.M.; Rucavado, A.; Theakston, R.D.G.; Gutiérrez, J.M. Pulmonary hemorrhage induced by jararhagin, a metalloproteinase from Bothrops jararaca venom. Toxicol. Appl. Pharmacol. 2003, 193, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, C.F.P.; Zamunér, S.R.; Zuliani, J.P.; Fernandes, C.M.; Cruz-Hofling, M.A.; Fernandes, I.; Chaves, F.; Gutiérrez, J.M. Neutrophils do not contribute to local tissue damage, but play a key role in skeletal muscle regeneration, in mice injected with Bothrops asper snake venom. Muscle Nerve 2003, 28, 449–459. [Google Scholar] [CrossRef] [PubMed]
- Rojas, E.; Quesada, L.; Arce, V.; Lomonte, B.; Rojas, G.; Gutiérrez, J.M. Neutralization of four Peruvian Bothrops sp snake venoms by polyvalent antivenoms produced in Perú and Costa Rica: Preclinical assessment. Acta Trop. 2005, 93, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Rojas, G.; Lomonte, B.; Gené, J.A.; Cerdas, L. Comparative study of the edema-forming activity of Costa Rican snake venoms and its neutralization by a polyvalent antivenom. Comp. Biochem. Physiol. 1986, 85, 171–175. [Google Scholar] [CrossRef]
- Barfaraz, A.; Harvey, A.L. The use of the chick biventer cervicis preparation to assess the protective activity of six international reference antivenoms on the neuromuscular effects of snake venoms in vitro. Toxicon 1994, 32, 267–272. [Google Scholar] [CrossRef] [PubMed]
- Rucavado, A.; Soto, M.; Escalante, T.; Loría, G.D.; Arni, R.D.; Gutiérrez, J.M. Thrombocytopenia and platelet hypoaggregation induced by Bothrops asper snake venom. Toxins involved and their contribution to metalloproteinase-induced pulmonary hemorrhage. Thromb. Haemost. 2005, 94, 123–131. [Google Scholar] [PubMed]
- Gutiérrez, J.; Sanz, L.; Escolano, J.; Fernández, J.; Lomonte, B.; Angulo, Y.; Rucavado, A.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics of the Lesser Antillean pit vipers Bothrops caribbaeus and Bothrops lanceolatus. Correlation with toxicological activities and immunoreactivity of a heterologous antivenom. J. Proteome Res. 2008, 7, 4396–4408. [Google Scholar] [CrossRef] [PubMed]
- Gené, J.A.; Gómez, M.; Gutiérrez, J.M.; Cerdas, L. Neutralization of hyaluronidase and indirect hemolytic activities of Costa Rican snake venoms by a polyvalent antivenom. Toxicon 1985, 23, 1015–1018. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Rojas, G.; Lomonte, B.; Gené, J.A.; Chaves, F.; Alvarado, J.; Rojas, E. Standardization of assays for testing the neutralizing ability of antivenoms. Toxicon 1990, 28, 1127–1129. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Juárez, P.; Sanz, L. Snake venomics. Strategy and applications. J. Mass Spectrom. 2007, 42, 1405–1414. [Google Scholar] [CrossRef] [PubMed]
- Fox, J.W.; Serrano, S.M.T. Exploring snake venom proteomes: multifaceted analyses for complex toxin mixtures. Proteomics 2008, 8, 909–920. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J. Next-generation snake venomics: protein-locus resolution through venom proteome decomplexation. Exp. Rev. Proteomics 2014, 11, 315–329. [Google Scholar] [CrossRef]
- Rokyta, D.R.; Lemmon, A.R.; Margres, M.J.; Aronow, K. The venom-gland transcriptome of the eastern diamondback rattlesnake (Crotalus adamanteus). BMC Genomics 2012, 13, 312. [Google Scholar] [CrossRef] [PubMed]
- Margres, M.J.; Aronow, K.; Loyacano, J.; Rokyta, D.R. The venom-gland transcriptome of the eastern coral snake (Micrurus fulvius) reveals high venom complexity in the intragenomic evolution of venoms. BMC Genomics 2013, 14, 531. [Google Scholar] [CrossRef] [PubMed]
- Margres, M.J.; McGivern, J.J.; Wray, K.P.; Seavy, M.; Calvin, K.; Rokyta, D.R. Linking the transcriptome and proteome to characterize the venom of the eastern diamondback rattlesnake (Crotalus adamanteus). J. Proteomics 2014, 96, 145–158. [Google Scholar] [CrossRef] [PubMed]
- Wagstaff, S.C.; Sanz, L.; Juárez, P.; Harrison, R.A.; Calvete, J.J. Combined snake venomics and venom gland transcriptomic analysis of the ocellated carpet viper, Echis ocellatus. J. Proteomics 2009, 71, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Paiva, O.; Pla, D.; Wright, C.E.; Beutler, M.; Sanz, L.; Gutiérrez, J.M.; Williams, D.J.; Calvete, J.J. Combined venom gland cDNA sequencing and venomics of the New Guinea small-eyed snake, Micropechis ikaheka. J. Proteomics 2014, 110, 209–229. [Google Scholar] [CrossRef] [PubMed]
- Alape-Girón, A.; Sanz, L.; Escolano, J.; Flores-Díaz, M.; Madrigal, M.; Sasa, M.; Calvete, J.J. Snake venomics of the lancehead pitviper Bothrops asper. Geographic, individual and ontogenetic variations. J. Proteome Res. 2008, 7, 3556–3571. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Sanz, L.; Pérez, A.; Borges, A.; Vargas, A.M.; Lomonte, B.; Angulo, Y.; Gutiérrez, J.M.; Chalkidis, H.M.; Mourão, R.H.; et al. Snake population venomics and antivenomics of Bothrops atrox: paedomorphism along its transamazonian dispersal and implications of geographic venom variability on snakebite management. J. Proteomics 2011, 74, 510–527. [Google Scholar] [CrossRef]
- Zelanis, A.; Tashima, A.K.; Pinto, A.F.; Leme, A.F.; Stuginski, D.R.; Furtado, M.F.; Sherman, N.E.; Ho, P.L.; Fox, J.W.; Serrano, S.M. Bothrops jararaca venom proteome rearrangement upon neonate to adult transition. Proteomics 2011, 11, 4218–4228. [Google Scholar] [CrossRef] [PubMed]
- Antunes, T.C.; Yamashita, K.M.; Barbaro, K.C.; Saiki, M.; Santoro, M.L. Comparative analysis of newborn and adult Bothrops jararaca snake venoms. Toxicon 2010, 56, 1443–1458. [Google Scholar] [CrossRef]
- Boldrini-França, J.; Corrêa-Netto, C.; Silva, M.M.S.; Rodrigues, R.S.; De La Torre, P.; Pérez, A.; Soares, A.M.; Zingali, R.B.; Nogueira, R.A.; Rodrigues, V.M.; et al. Snake venomics and antivenomics of Crotalus durissus subspecies from Brazil: Assessment of geographic variation and its implication on snakebite management. J. Proteomics 2010, 73, 1758–1776. [Google Scholar]
- Calvete, J.J.; Sanz, L.; Cid, P.; De La Torre, P.; Flores-Díaz, M.; Dos Santos, M.C.; Borges, A.; Bremo, A.; Angulo, Y.; Lomonte, B.; et al. Snake venomics of the Central American rattlesnake Crotalus simus and the South American Crotalus durissus complex points to neurotoxicity as an adaptive paedomorphic trend along Crotalus dispersal in South America. J. Proteome Res. 2010, 9, 528–544. [Google Scholar] [CrossRef] [PubMed]
- Sunagar, K.; Undheim, E.A.; Scheib, H.; Gren, E.C.; Cochran, C.; Person, C.E.; Koludarov, I.; Kelln, W.; Hayes, W.K.; King, G.F.; et al. Intraspecific venom variation in the medically significant Southern Pacific Rattlesnake (Crotalus oreganus helleri): Biodiscovery, clinical and evolutionary implications. J. Proteomics 2014, 99, 68–83. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, H.L.; Chiucchi, J.E. Deconstructing a complex molecular phenotype: Population-level variation in individual venom proteins in Eastern Massasauga rattlesnakes (Sistrurus c. catenatus). J. Mol. Evol. 2011, 72, 383–397. [Google Scholar] [CrossRef]
- Gibbs, H.L.; Sanz, L.; Chiucchi, J.E.; Farrell, T.M.; Calvete, J.J. Proteomic analysis of ontogenetic and diet-related changes in venom composition of juvenile and adult Dusky Pigmy rattlesnakes (Sistrurus miliarius barbouri). J. Proteomics 2011, 74, 2169–2179. [Google Scholar] [CrossRef] [PubMed]
- Madrigal, M.; Sanz, L.; Flores-Diaz, M.; Sasa, M.; Núñez, V.; Alape-Girón, A.; Calvete, J.J. Snake venomics across genus Lachesis. Ontogenetic changes in the venom composition of L. stenophrys and comparative proteomics of the venoms of adult L. melanocephala and L. acrochorda. J. Proteomics 2012, 77, 280–297. [Google Scholar] [CrossRef] [PubMed]
- Pla, D.; Sanz, L.; Molina-Sánchez, P.; Zorita, V.; Madrigal, M.; Flores-Díaz, M.; Alape-Girón, A.; Núñez, V.; Andrés, V.; Gutiérrez, J.M.; et al. Snake venomics of Lachesis muta rhombeata and genus-wide antivenomics assessment of the paraspecific immunoreactivity of two antivenoms evidence the high compositional and immunological conservation across Lachesis. J. Proteomics 2013, 89, 112–123. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.F.; Wang, J.; He, Y.; Qu, Y.F.; Lin, L.H.; Ma, X.M.; Ji, X. Proteomic and biochemical analyses of short-tailed pit viper (Gloydius brevicaudus) venom: Age-related variation and composition-activity correlation. J. Proteomics 2014, 105, 307–322. [Google Scholar] [CrossRef] [PubMed]
- Wooten, J.A.; Gibbs, H.L. Niche divergence and lineage diversification among closely related Sistrurus rattlesnakes. J. Evol. Biol. 2012, 25, 317–328. [Google Scholar] [CrossRef]
- Lomonte, B.; Escolano, J.; Fernández, J.; Sanz, L.; Angulo, Y.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics and antivenomics of the arboreal neotropical pitvipers Bothriechis lateralis and Bothriechis schlegelii. J. Proteome Res. 2008, 7, 2445–2457. [Google Scholar] [CrossRef] [PubMed]
- Pla, D.; Gutiérrez, J.M.; Calvete, J.J. Second generation snake antivenomics: Comparing immunoaffinity and immunodepletion protocols. Toxicon 2012, 60, 688–699. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Tsai, W.C.; Pla, D.; Solano, G.; Lomonte, B.; Sanz, L.; Angulo, Y.; Calvete, J.J. Preclinical assessment of a polyspecific antivenom against the venoms of Cerrophidion sasai, Porthidium nasutum and Porthidium ophryomegas: Insights from combined antivenomics and neutralization assays. Toxicon 2013, 64, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Lomonte, B.; Sanz, L.; Calvete, J.J.; Pla, D. Immunological profile of antivenoms: Preclinical analysis of the efficacy of a polyspecific antivenom through antivenomics and neutralization assays. J. Proteomics 2014, 105, 340–350. [Google Scholar] [CrossRef] [PubMed]
- Pla, D.; Paiva, O.K.; Sanz, L.; Beutler, M.; Wright, C.E.; Calvete, J.J.; Williams, D.J.; Gutiérrez, J.M. Preclinical efficacy of Australian antivenoms against the venom of the small-eyed snake, Micropechis ikaheka, from Papua New Guinea: An antivenomics and neutralization study. J. Proteomics 2014, 110, 198–208. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Rojas, E.; Quesada, L.; León, G.; Núñez, J.; Laing, G.D.; Sasa, M.; Renjifo, J.M.; Nasidi, A.; Warrell, D.A.; et al. Pan-African polyspecific antivenom produced by caprylic acid purification of horse IgG: An alternative to the antivenom crisis in Africa. Trans. R. Soc. Trop. Med. Hyg. 2005, 99, 468–475. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, E.; Eaker, D.L.; Porath, J. Purification of a neurotoxin from the venom of Naja nigricollis. Biochim. Biophys. Acta 1966, 127, 505–520. [Google Scholar]
- Branch, W.R. The venomous snakes of southern Africa. Part 2. Elapidae and Hydrophiidae. Snake 1979, 11, 199–225. [Google Scholar]
- Hughes, B. African snake faunas. Bonn Zool. Beitr. 1983, 34, 311–356. [Google Scholar]
- Wüster, W.; Broadley, D.G. A new species of spitting cobra from northeastern Africa (Serpentes: Elapidae: Naja). J. Zool. London 2003, 259, 345–359. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Calvete, J.J.; Sanz, L.; Pla, D.; Lomonte, B.; Gutiérrez, J.M. Omics Meets Biology: Application to the Design and Preclinical Assessment of Antivenoms. Toxins 2014, 6, 3388-3405. https://doi.org/10.3390/toxins6123388
Calvete JJ, Sanz L, Pla D, Lomonte B, Gutiérrez JM. Omics Meets Biology: Application to the Design and Preclinical Assessment of Antivenoms. Toxins. 2014; 6(12):3388-3405. https://doi.org/10.3390/toxins6123388
Chicago/Turabian StyleCalvete, Juan J., Libia Sanz, Davinia Pla, Bruno Lomonte, and José María Gutiérrez. 2014. "Omics Meets Biology: Application to the Design and Preclinical Assessment of Antivenoms" Toxins 6, no. 12: 3388-3405. https://doi.org/10.3390/toxins6123388
APA StyleCalvete, J. J., Sanz, L., Pla, D., Lomonte, B., & Gutiérrez, J. M. (2014). Omics Meets Biology: Application to the Design and Preclinical Assessment of Antivenoms. Toxins, 6(12), 3388-3405. https://doi.org/10.3390/toxins6123388






