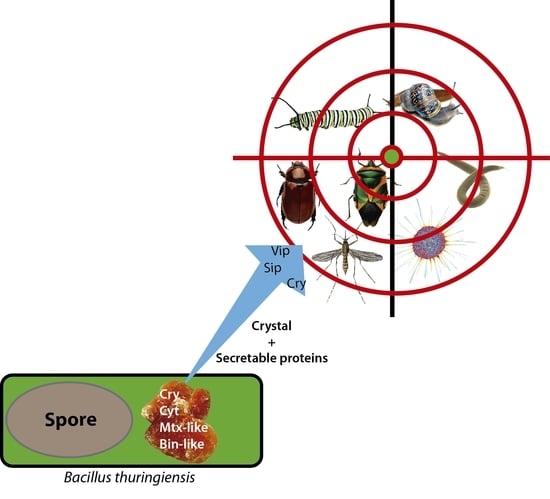
Bacillus thuringiensis Toxins: An Overview of Their Biocidal Activity
Abstract
:1. Introduction

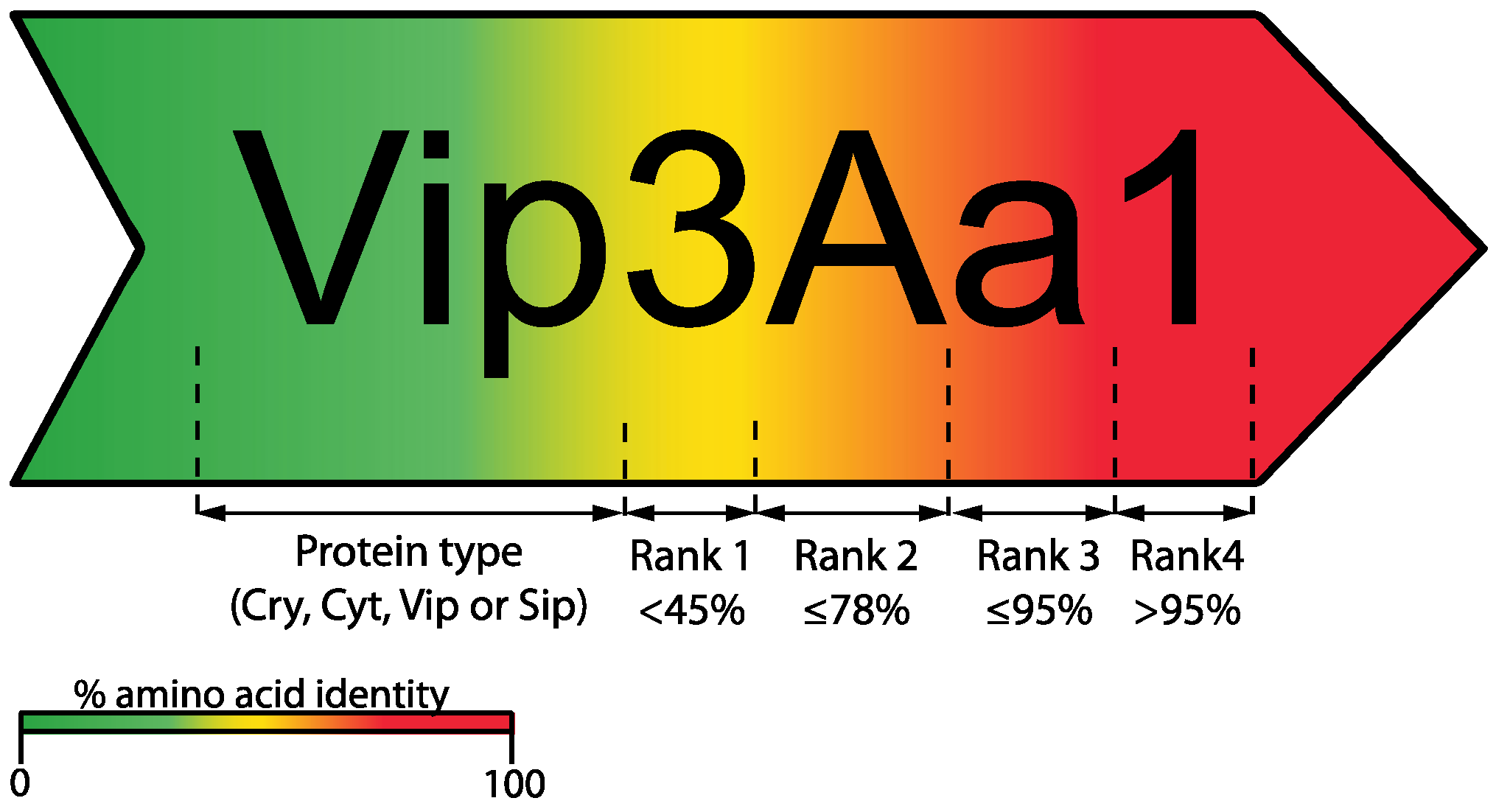
2. Bt Toxin Nomenclature
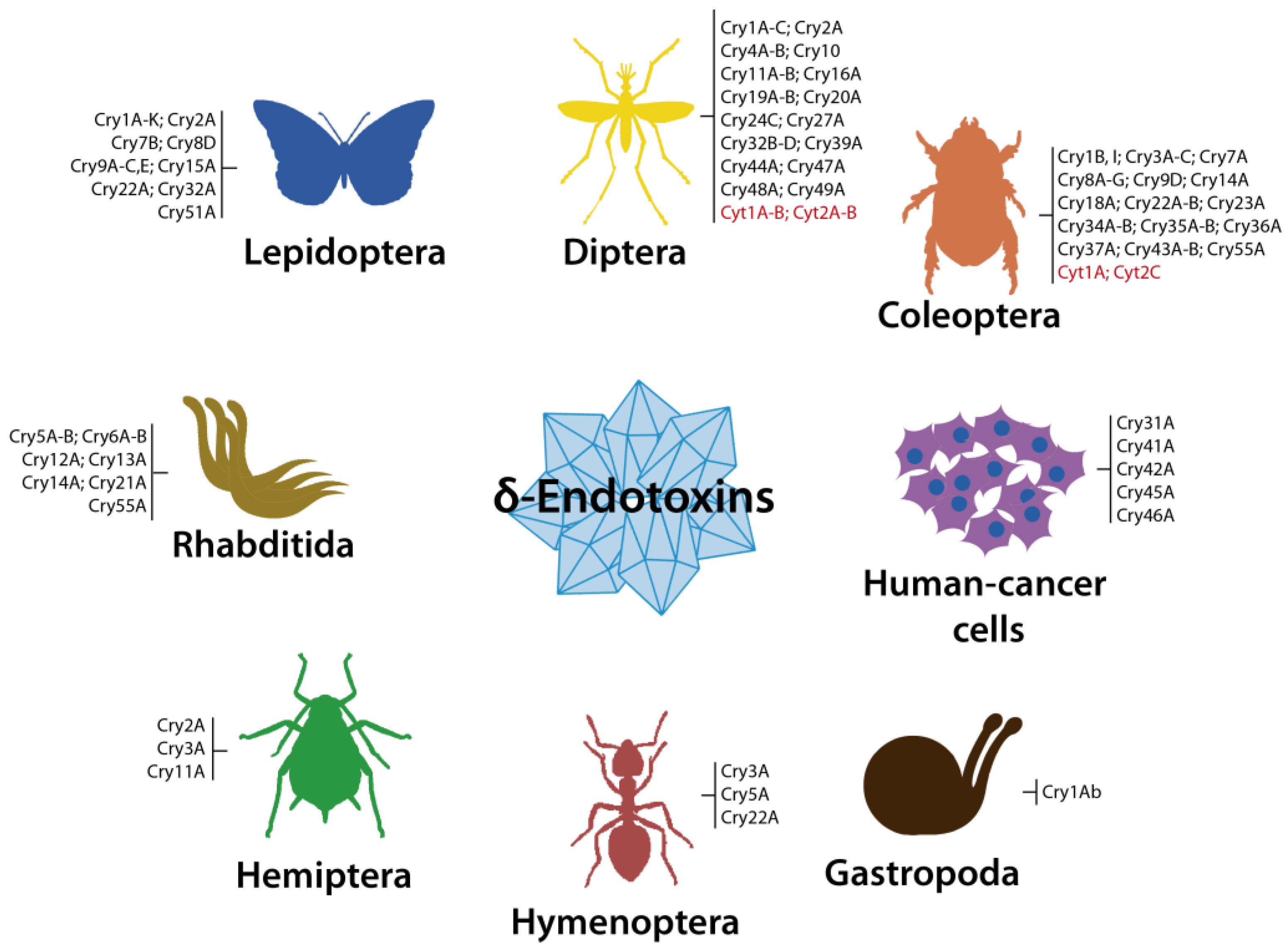
3. Crystal Toxins (δ-endotoxins)
3.1. Cry Toxins

3.1.1. Three-Domain Cry Toxins
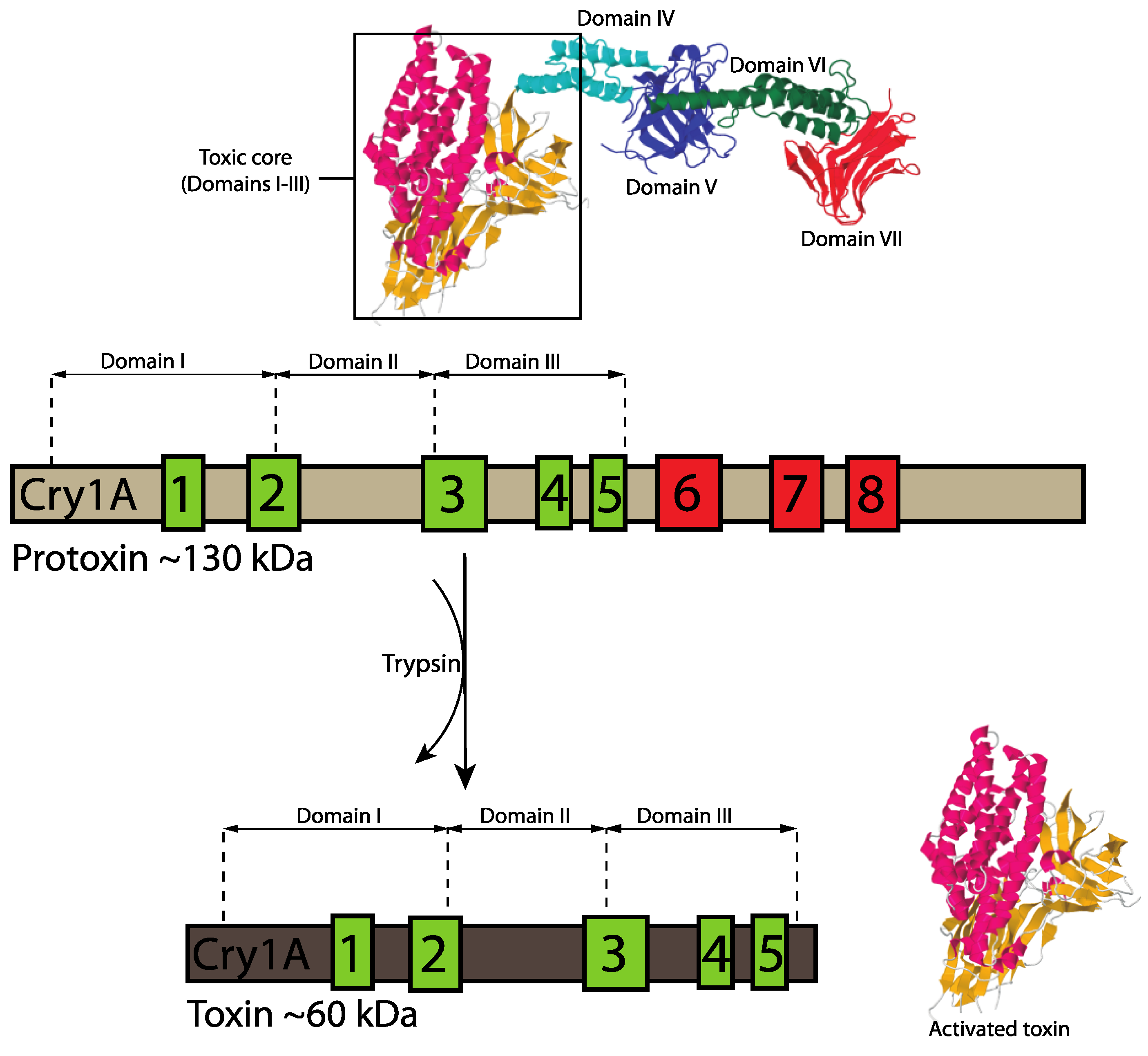
3.1.2. Non-Three-Domain Cry Toxins
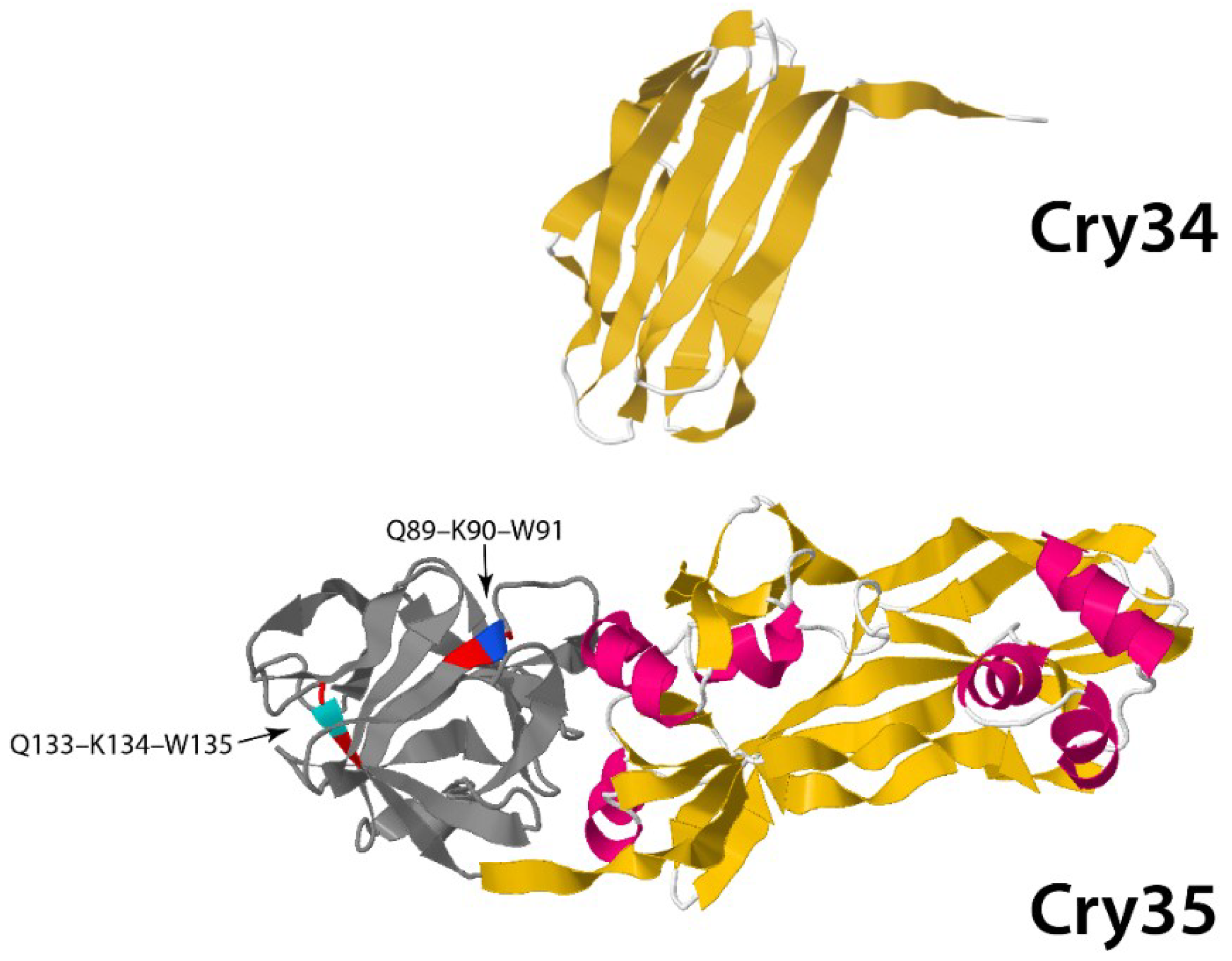
3.2. Cyt Toxins
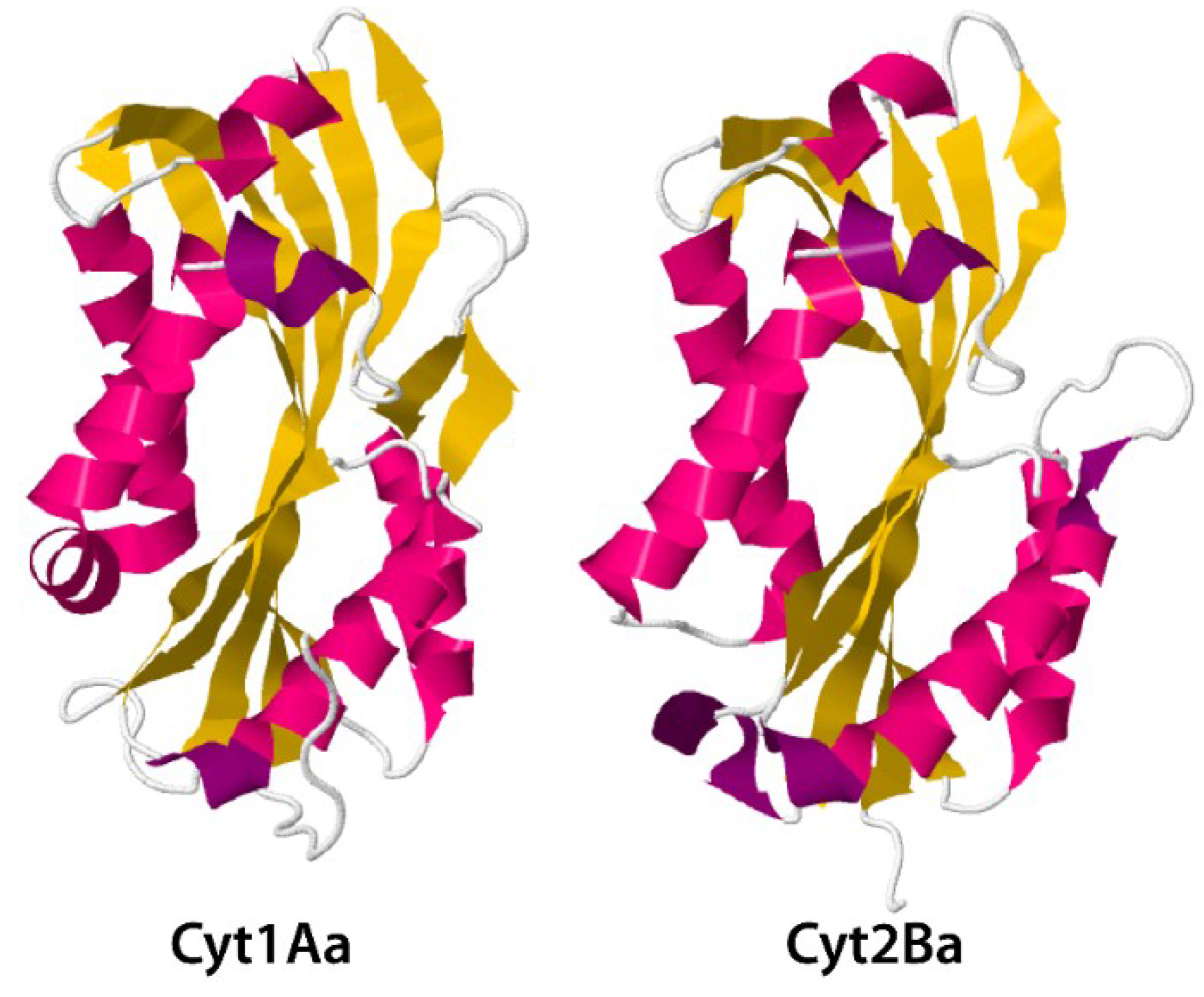
4. Secreted Toxins
4.1. Vip1/Vip2 (Binary) Toxins
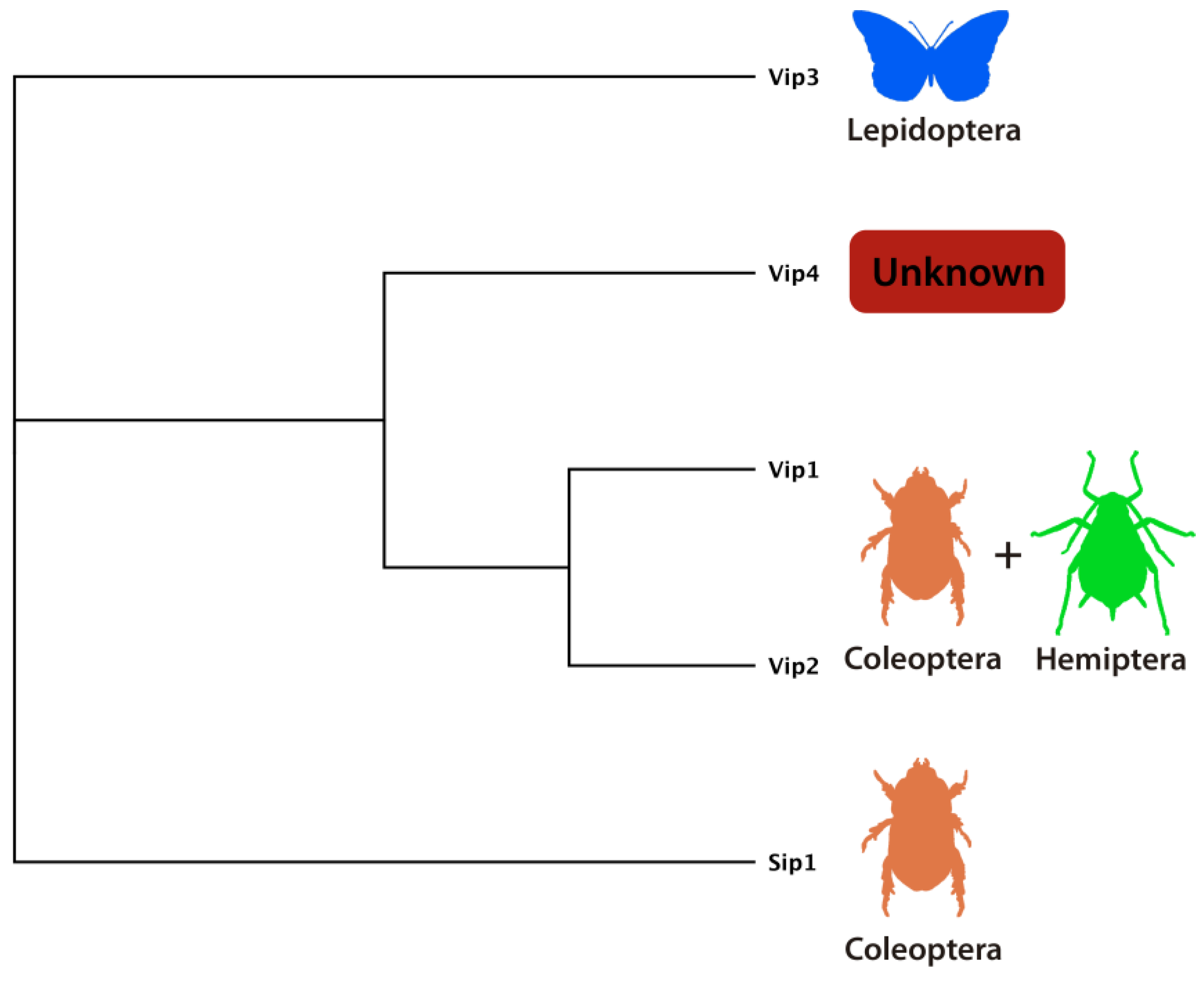
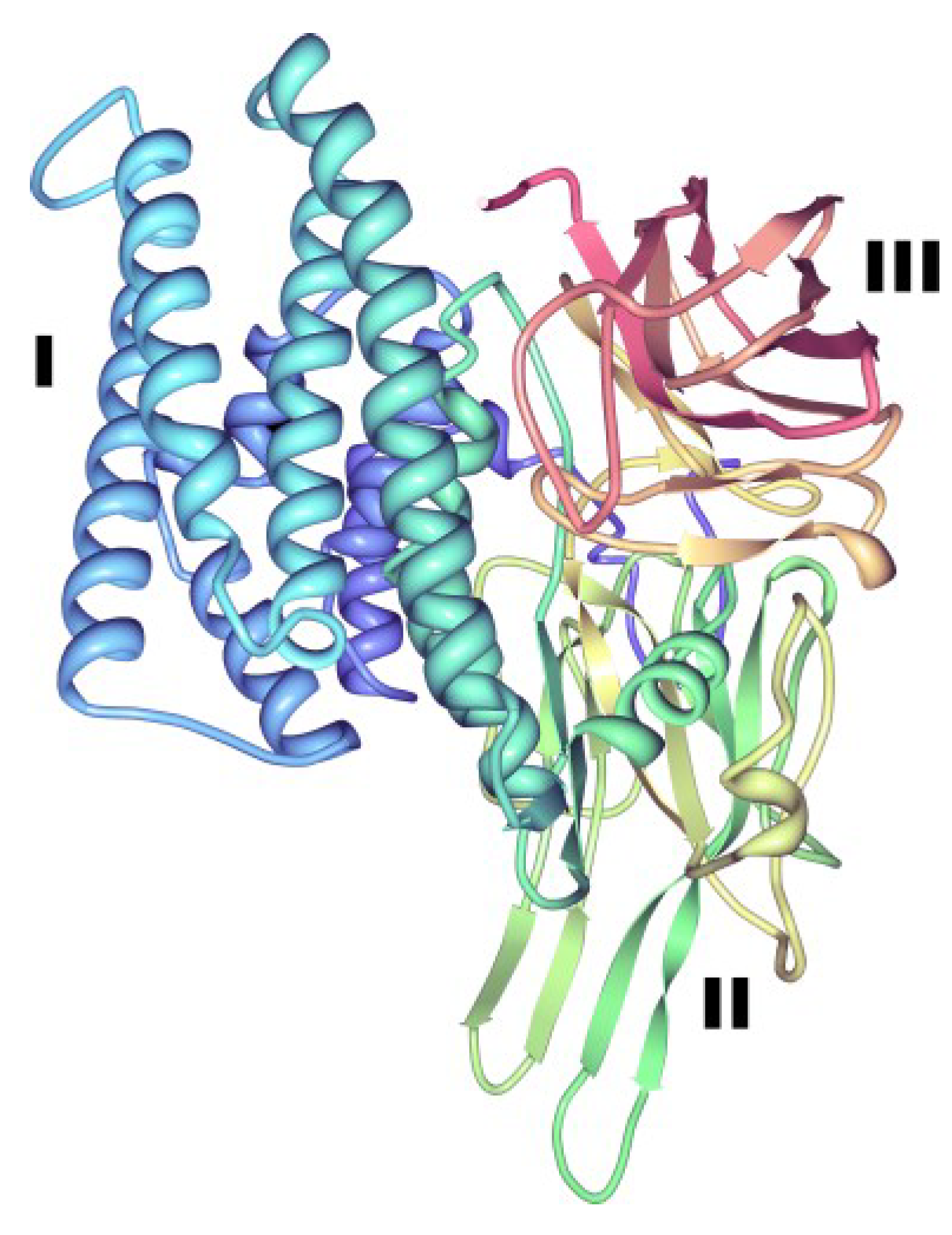
4.2. Vip3 Proteins
4.3. Vip4 Protein

4.4. Sip Toxin
5. Other Potential Insecticidal Toxins
5.1. A 41.9-kDa Protein
5.2. Sphaericolysins and Alveolysins
5.3. Beta Exotoxins
5.4. Enhancin-Like Proteins
5.5. P19 and P20 Helper Proteins
6. Mechanisms of Toxin Evolution
7. Novel Toxins and Next-Generation Sequencing (NGS) Technologies
8. Concluding Remarks
Acknowledgments
Authors Contribution
Abbreviations
| Bc | Bacillus cereus |
| Bt | Bacillus thuringiensis |
| Ls | Lysinibacillus sphaericus |
Conflicts of Interest
References
- Höfte, H.; Whiteley, H.R. Insecticidal Crystal Proteins of Bacillus thuringiensis. Microbiol. Rev. 1989, 53, 242–255. [Google Scholar]
- Knowles, B.H.; Dow, J.A.T. The crystal delta-endotoxins of Bacillus thuringiensis—Models for their mechanism of action on the insect gut. Bioessays 1993, 15, 469–476. [Google Scholar] [CrossRef]
- Raymond, B.; Johnston, P.R.; Nielsen-LeRoux, C.; Lereclus, D.; Crickmore, N. Bacillus thuringiensis: An impotent pathogen? Trends Microbiol. 2010, 18, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Roh, J.Y.; Choi, J.Y.; Li, M.S.; Jin, B.R.; Je, Y.H. Bacillus thuringiensis as a specific, safe, and effective tool for insect pest control. J. Mol. Biol. 2007, 17, 547–559. [Google Scholar]
- Schnepf, E.; Crickmore, N.; van Rie, J.; Lereclus, D.; Baum, J.; Feitelson, J.; Zeigler, D.R.; Dean, D.H. Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 1998, 62, 775–806. [Google Scholar] [PubMed]
- Sanchis, V. From microbial sprays to insect-resistant transgenic plants: History of the biospesticide Bacillus thuringiensis. A review. Agron. Sustain. Dev. 2011, 31, 217–231. [Google Scholar] [CrossRef]
- Bravo, A.; Gill, S.S.; Soberón, M. Mode of action of Bacillus thuringiensis Cry and Cyt toxins and their potential for insect control. Toxicon 2007, 49, 423–435. [Google Scholar] [CrossRef] [PubMed]
- Crickmore, N.; Zeigler, D.R.; Schnepf, E.; van Rie, J.; Lereclus, D.; Baum, J.; Bravo, A.; Dean, D.H. Bacillus thuringiensis Toxin Nomenclature. Available online: http://www.lifesci.sussex.ac.uk/Home/Neil_Crickmore/Bt/ (accessed on 7 November 2014).
- Van Frankenhuyzen, K. Insecticidal activity of Bacillus thuringiensis crystal proteins. J. Invertebr. Pathol. 2009, 101, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Sanchis, V.; Bourguet, D. Bacillus thuringiensis: Applications in agriculture and insect resistance management. A review. Agron. Sustain. Dev. 2008, 28, 11–20. [Google Scholar] [CrossRef]
- Ohba, M.; Mizuki, E.; Uemori, A. Parasporin, a new anticancer protein group from Bacillus thuringiensis. Anticancer Res. 2009, 29, 427–433. [Google Scholar] [PubMed]
- Okumura, S.; Ohba, M.; Mizuki, E.; Crickmore, N.; Coté, J.-C.; Nagamatsu, Y.; Kitada, S.; Sakai, H.; Harata, K.; Shin, T.; et al. Parasporin Nomenclature. Available online: http://parasporin.fitc.pref.fukuoka.jp/ (accessed on 7 November 2014).
- Estruch, J.J.; Warren, G.W.; Mullins, M.A.; Nye, G.J.; Craig, J.A.; Koziel, M.G. Vip3A, a novel Bacillus thuringiensis vegetative insecticidal protein with a wide spectrum of activities against lepidopteran insects. Proc. Natl. Acad. Sci. USA 1996, 93, 5389–5394. [Google Scholar] [CrossRef] [PubMed]
- Warren, G.W.; Koziel, M.G.; Mullins, M.A.; Nye, G.J.; Carr, B.; Desai, N.M.; Kostichka, K.; Duck, N.B.; Estruch, J.J. Auxiliary Proteins for Enhancing the Insecticidal Activity of Pesticidal Proteins. U.S. Patent 5,770,696, 23 June 1998. [Google Scholar]
- Donovan, W.P.; Engleman, J.T.; Donovan, J.C.; Baum, J.A.; Bunkers, G.J.; Chi, D.J.; Clinton, W.P.; English, L.; Heck, G.R.; Ilagan, O.M.; et al. Discovery and characterization of Sip1A: A novel secreted protein from Bacillus thuringiensis with activity against coleopteran larvae. Appl. Microbiol. Biotechnol. 2006, 72, 713–719. [Google Scholar] [CrossRef] [PubMed]
- Schnepf, H.E.; Whiteley, H.R. Cloning and expression of the Bacillus thuringiensis crystal protein gene in Escherichia coli. Proc. Natl. Acad. Sci. USA 1981, 78, 2893–2897. [Google Scholar] [CrossRef] [PubMed]
- Crickmore, N.; Zeigler, D.R.; Feitelson, J.; Schnepf, E.; van Rie, J.; Lereclus, D.; Baum, J.; Dean, D.H. Revision of the nomenclature for the Bacillus thuringiensis pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 1998, 62, 807–813. [Google Scholar] [PubMed]
- Ben-Dov, E. Bacillus thuringiensis subsp. israelensis and Its Dipteran-Specific Toxins. Toxins 2014, 6, 1222–1243. [Google Scholar]
- Butko, P. Cytolytic toxin Cyt1A and its mechanism of membrane damage: Data and hypotheses. Appl. Environ. Microbiol. 2003, 69, 2415–2422. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.; Albeck, S.; Ben-Dov, E.; Cahan, R.; Firer, M.; Zaritsky, A.; Dym, O. Cyt1Aa toxin: Crystal structure reveals implications for its membrane-perforating function. J. Mol. Biol. 2011, 413, 804–814. [Google Scholar] [CrossRef]
- De Maagd, R.A.; Bravo, A.; Berry, C.; Crickmore, N.; Schnepf, H.E. Structure, diversity, and evolution of protein toxins from spore-forming entomopathogenic bacteria. Annu. Rev. Genet. 2003, 37, 409–433. [Google Scholar]
- Soberon, M.; Lopez-Diaz, J.A.; Bravo, A. Cyt toxins produced by Bacillus thuringiensis: A protein fold conserved in several pathogenic microorganisms. Peptides 2013, 41, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Liu, T.; Sun, Z.; Guan, P.; Zhu, J.; Wang, S.; Li, S.; Deng, Q.; Wang, L.; Zheng, A.; et al. Co-expression and synergism analysis of Vip3Aa29 and Cyt2Aa3 insecticidal proteins from Bacillus thuringiensis. Curr. Microbiol. 2012, 64, 326–331. [Google Scholar] [CrossRef]
- Berry, C. The bacterium, Lysinibacillus sphaericus, as an insect pathogen. J. Invertebr. Pathol. 2012, 109, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Chougule, N.P.; Bonning, B.C. Toxins for transgenic resistance to hemipteran pests. Toxins 2012, 4, 405–429. [Google Scholar] [CrossRef]
- Van Frankenhuyzen, K. Cross-order and cross-phylum activity of Bacillus thuringiensis pesticidal proteins. J. Invertebr. Pathol. 2013, 114, 76–85. [Google Scholar]
- Ali, B.A.; Salem, H.H.; Wang, X.M.; Huang, T.H.; Xie, Q.D.; Zhang, X.Y. Effect of Bacillus thuringiensis var. israelensis endotoxin on the intermediate snail host of Schistosoma japonicum. Curr. Res. Bacteriol. 2010, 3, 37–41. [Google Scholar]
- Bravo, A.; Soberón, M. How to cope with insect resistance to Bt toxins? Trends Biotechnol. 2008, 26, 573–579. [Google Scholar] [CrossRef] [PubMed]
- Mizushiro, H.; Akao, T.; Yamashita, S.; Oba, M.; Kondo, S.; Maeda, M. Protien Having Antitrichomonal Activity and Derived from Bacillus thuringiensis and Method for Preparing the Same. Japanese Patent JP2002284800, 3 October 2002. [Google Scholar]
- Kondo, S.; Mizuki, E.; Akao, T.; Ohba, M. Antitrichomonal strains of Bacillus thuringiensis. Parasitol. Res. 2002, 88, 1090–1092. [Google Scholar] [CrossRef] [PubMed]
- Halima, H.S.; Bahy, A.A.; Tian, H.H.; Qing, D.X. Molecular characterization of novel Bacillus thuringiensis isolate with molluscicidal activity against the intermediate host of schistosomes. Biotechnology 2006, 5, 413–420. [Google Scholar]
- Ross, A.G.P.; Sleigh, A.C.; Li, Y.S.; Davis, G.M.; Williams, G.M.; Jiang, Z.; Feng, Z.; McManus, D.P. Schistosomiasis in the People’s Republic of China: Prospects and challenges for the 21st century. Clin. Microbiol. Rev. 2001, 14, 270–295. [Google Scholar] [CrossRef] [PubMed]
- Naimov, S.; Boncheva, R.; Karlova, R.; Dukiandjiev, S.; Minkov, I.; de Maagd, R.A. Solubilization, activation, and insecticidal activity of Bacillus thuringiensis serovar thompsoni HD542 crystal proteins. Appl. Environ. Microbiol. 2008, 74, 7145–7151. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.Z.; Hale, K.; Carta, L.; Platzer, E.; Wong, C.; Fang, S.C.; Aroian, R.V. Bacillus thuringiensis crystal proteins that target nematodes. Proc. Natl. Acad. Sci. USA 2003, 100, 2760–2765. [Google Scholar] [CrossRef] [PubMed]
- Yudina, T.G.; Brioukhanov, A.L.; Zalunin, I.A.; Revina, L.P.; Shestakov, A.I.; Voyushina, N.E.; Chestukhina, G.G.; Netrusov, A.I. Antimicrobial activity of different proteins and their fragments from Bacillus thuringiensis parasporal crystals against clostridia and archaea. Anaerobe 2007, 13, 6–13. [Google Scholar] [PubMed]
- Revina, L.P.; Kostina, L.I.; Dronina, M.A.; Zalunin, I.A.; Chestukhina, G.G.; Yudina, T.G.; Konukhova, A.V.; Izumrudova, A.V. Novel antibacterial proteins from entomocidal crystals of Bacillus thuringiensis ssp israelensis. Can. J. Microbiol. 2005, 51, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Yudina, T.G.; Konukhova, A.V.; Revina, L.P.; Kostina, L.I.; Zalunin, I.A.; Chestukhina, G.G. Antibacterial activity of Cry- and Cyt-proteins from Bacillus thuringiensis ssp israelensis. Can. J. Microbiol. 2003, 49, 37–44. [Google Scholar] [CrossRef]
- De Maagd, R.A.; Bravo, A.; Crickmore, N. How Bacillus thuringiensis has evolved specific toxins to colonize the insect world. Trends Genet. 2001, 17, 193–199. [Google Scholar]
- Pardo-López, L.; Soberón, M.; Bravo, A. Bacillus thuringiensis insecticidal three-domain Cry toxins: Mode of action, insect resistance and consequences for crop protection. FEMS Microbiol. Rev. 2013, 37, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Morse, R.J.; Yamamoto, T.; Stroud, R.M. Structure of Cry2Aa suggests an unexpected receptor binding epitope. Structure 2001, 9, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Wang, B.C.; Yu, Z.; Sun, M. Structural insights into Bacillus thuringiensis Cry, Cyt and parasporin toxins. Toxins 2014, 6, 2732–2770. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, J.L.; Dean, D.H. Exploring the mechanism of action of insecticidal proteins by genetic engineering methods. Genet. Eng. (N.Y.) 2000, 22, 33–54. [Google Scholar]
- Evdokimov, A.G.; Moshiri, F.; Sturman, E.J.; Rydel, T.J.; Zheng, M.; Seale, J.W.; Franklin, S. Structure of the full-length insecticidal protein Cry1Ac reveals intriguing details of toxin packaging into in vivo formed crystals. Protein Sci. 2014, 23, 1491–1497. [Google Scholar] [CrossRef] [PubMed]
- Du, C.; Martin, P.A.; Nickerson, K.W. Comparison of Disulfide Contents and Solubility at Alkaline pH of Insecticidal and Noninsecticidal Bacillus thuringiensis Protein Crystals. Appl. Environ. Microbiol. 1994, 60, 3847–3853. [Google Scholar] [PubMed]
- Creighton, T.E. Disulfide bonds as probes of protein folding pathways. Meth. Enzymol. 1986, 131, 83–106. [Google Scholar] [PubMed]
- Vachon, V.; Laprade, R.; Schwartz, J.L. Current models of the mode of action of Bacillus thuringiensis insecticidal crystal proteins: A critical review. J. Invertebr. Pathol. 2012, 111, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Kirouac, M.; Vachon, V.; Noel, J.F.; Girard, F.; Schwartz, J.L.; Laprade, R. Amino acid and divalent ion permeability of the pores formed by the Bacillus thuringiensis toxins Cry1Aa and Cry1Ac in insect midgut brush border membrane vesicles. Bba-Biomembranes 2002, 1561, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.B.; Candas, M.; Griko, N.B.; Taussig, R.; Bulla, L.A. A mechanism of cell death involving an adenylyl cyclase/PKA signaling pathway is induced by the Cry1Ab toxin of Bacillus thuringiensis. Proc. Natl. Acad. Sci. USA 2006, 103, 9897–9902. [Google Scholar] [CrossRef] [PubMed]
- Pigott, C.R.; Ellar, D.J. Role of receptors in Bacillus thuringiensis crystal toxin activity. Microbiol. Mol. Biol. Rev. 2007, 71, 255–281. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Luna, M.T.; Lanz-Mendoza, H.; Gill, S.S.; Bravo, A.; Soberon, M.; Miranda-Rios, J. An alpha-amylase is a novel receptor for Bacillus thuringiensis ssp. israelensis Cry4Ba and Cry11Aa toxins in the malaria vector mosquito Anopheles albimanus (Diptera: Culicidae). Environ. Microbiol. 2010, 12, 746–757. [Google Scholar] [PubMed]
- Zhang, Q.; Hua, G.; Bayyareddy, K.; Adang, M.J. Analyses of alpha-amylase and alpha-glucosidase in the malaria vector mosquito, Anopheles gambiae, as receptors of Cry11Ba toxin of Bacillus thuringiensis subsp. jegathesan. Insect Biochem. Mol. Biol. 2013, 43, 907–915. [Google Scholar] [CrossRef]
- Kuadkitkan, A.; Smith, D.R.; Berry, C. Investigation of the Cry4B-prohibitin interaction in Aedes aegypti cells. Curr. Microbiol. 2012, 65, 446–454. [Google Scholar] [CrossRef] [PubMed]
- Likitvivatanavong, S.; Chen, J.; Evans, A.M.; Bravo, A.; Soberon, M.; Gill, S.S. Multiple receptors as targets of Cry toxins in mosquitoes. J. Agric. Food Chem. 2011, 59, 2829–2838. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Hua, G.; Andacht, T.M.; Adang, M.J. A 106-kDa aminopeptidase is a putative receptor for Bacillus thuringiensis Cry11Ba toxin in the mosquito Anopheles gambiae. Biochemistry 2008, 47, 11263–11272. [Google Scholar] [CrossRef]
- Gonzalez, M.R.; Bischofberger, M.; Pernot, L.; van der Goot, F.G.; Freche, B. Bacterial pore-forming toxins: The (w)hole story? Cell. Mol. Life Sci. 2008, 65, 493–507. [Google Scholar] [CrossRef] [PubMed]
- Knapp, O.; Stiles, B.; Popoff, M.R. The aerolysin-like toxin family of cytolytic, pore-forming toxins. Open Toxinol. J. 2010, 3, 53–68. [Google Scholar] [CrossRef]
- Popoff, M.R. Epsilon toxin: A fascinating pore-forming toxin. FEBS J. 2011, 278, 4602–4615. [Google Scholar] [CrossRef] [PubMed]
- Kelker, M.S.; Berry, C.; Evans, S.L.; Pai, R.; McCaskill, D.G.; Wang, N.X.; Russell, J.C.; Baker, M.D.; Yang, C.; Pflugrath, J.W.; et al. Structural and biophysical characterization of Bacillus thuringiensis insecticidal proteins Cry34Ab1 and Cry35Ab1. PLoS One 2014, 9, e112555. [Google Scholar] [CrossRef] [PubMed]
- Srisucharitpanit, K.; Yao, M.; Promdonkoy, B.; Chimnaronk, S.; Tanaka, I.; Boonserm, P. Crystal structure of BinB: A receptor binding component of the binary toxin from Lysinibacillus sphaericus. Proteins 2014, 82, 2703–2712. [Google Scholar] [CrossRef]
- Akiba, T.; Abe, Y.; Kitada, S.; Kusaka, Y.; Ito, A.; Ichimatsu, T.; Katayama, H.; Akao, T.; Higuchi, K.; Mizuki, E.; et al. Crystal Structure of the Parasporin-2 Bacillus thuringiensis Toxin That Recognizes Cancer Cells. J. Mol. Biol. 2009, 386, 121–133. [Google Scholar] [CrossRef] [PubMed]
- Akiba, T.; Higuchi, K.; Mizuki, E.; Ekino, K.; Shin, T.; Ohba, M.; Kanai, R.; Harata, K. Nontoxic crystal protein from Bacillus thuringiensis demonstrates a remarkable structural similarity to beta-pore-forming toxins. Proteins 2006, 63, 243–248. [Google Scholar] [CrossRef]
- Jones, G.W.; Nielsen-Leroux, C.; Yang, Y.; Yuan, Z.; Dumas, V.F.; Monnerat, R.G.; Berry, C. A new Cry toxin with a unique two-component dependency from Bacillus sphaericus. FASEB J. 2007, 21, 4112–4120. [Google Scholar] [CrossRef] [PubMed]
- Hey, T.D.; Narva, K.; Woosley, A.T. Modified Bacillus thuringiensis Cry6 Proteins for Nematode Control. U.S. Patent 201,110,225,681, 15 September 2011. [Google Scholar]
- Yu, Z.; Luo, H.; Xiong, J.; Zhou, Q.; Xia, L.; Sun, M.; Li, L.; Yu, Z. Bacillus thuringiensis Cry6A exhibits nematicidal activity to Caenorhabditis elegans bre mutants and synergistic activity with Cry5B to C. elegans. Lett. Appl. Microbiol. 2014, 58, 511–519. [Google Scholar] [CrossRef] [PubMed]
- Guerchicoff, A.; Delecluse, A.; Rubinstein, C.P. The Bacillus thuringiensis cyt genes for hemolytic endotoxins constitute a gene family. Appl. Environ. Microbiol. 2001, 67, 1090–1096. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.; Dym, O.; Albeck, S.; Ben-Dov, E.; Cahan, R.; Firer, M.; Zaritsky, A. High-resolution crystal structure of activated Cyt2Ba monomer from Bacillus thuringiensis subsp. israelensis. J. Mol. Biol. 2008, 380, 820–827. [Google Scholar] [CrossRef]
- Berry, C.; O’Neil, S.; Ben-Dov, E.; Jones, A.F.; Murphy, L.; Quail, M.A.; Holden, M.T.; Harris, D.; Zaritsky, A.; Parkhill, J.; et al. Complete sequence and organization of pBtoxis, the toxin-coding plasmid of Bacillus thuringiensis subsp. israelensis. Appl. Environ. Microbiol. 2002, 68, 5082–5095. [Google Scholar] [CrossRef]
- Manasherob, R.; Itsko, M.; Sela-Baranes, N.; Ben-Dov, E.; Berry, C.; Cohen, S.; Zaritsky, A. Cyt1Ca from Bacillus thuringiensis subsp. israelensis: Production in Escherichia coli and comparison of its biological activities with those of other Cyt-like proteins. Microbiology 2006, 152, 2651–2659. [Google Scholar]
- Federici, B.A.; Bauer, L.S. Cyt1Aa protein of Bacillus thuringiensis is toxic to the cottonwood leaf beetle, Chrysomela scripta, and suppresses high levels of resistance to Cry3Aa. Appl. Environ. Microbiol. 1998, 64, 4368–4371. [Google Scholar] [PubMed]
- Wirth, M.C.; Delecluse, A.; Walton, W.E. Cyt1Ab1 and Cyt2Ba1 from Bacillus thuringiensis subsp. medellin and B. thuringiensis subsp. israelensis synergize Bacillus sphaericus against Aedes aegypti and resistant Culex quinquefasciatus (Diptera: Culicidae). Appl. Environ. Microbiol. 2001, 67, 3280–3284. [Google Scholar]
- Zhang, B.H.; Liu, M.; Yang, Y.K.; Yuan, Z.M. Cytolytic toxin Cyt1Aa of Bacillus thuringiensis synergizes the mosquitocidal toxin Mtx1 of Bacillus sphaericus. Biosci. Biotech. Bioch. 2006, 70, 2199–2204. [Google Scholar] [CrossRef]
- Donovan, W.P.; Donovan, J.C.; Engleman, J.T. Gene knockout demonstrates that vip3A contributes to the pathogenesis of Bacillus thuringiensis toward Agrotis ipsilon and Spodoptera exigua. J. Invertebr. Pathol. 2001, 78, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Milne, R.; Liu, Y.; Gauthier, D.; van Frankenhuyzen, K. Purification of Vip3Aa from Bacillus thuringiensis HD-1 and its contribution to toxicity of HD-1 to spruce budworm (Choristoneura fumiferana) and gypsy moth (Lymantria dispar) (Lepidoptera). J. Invertebr. Pathol. 2008, 99, 166–172. [Google Scholar] [CrossRef]
- Shi, Y.; Ma, W.; Yuan, M.; Sun, F.; Pang, Y. Cloning of vip1/vip2 genes and expression of Vip1Ca/Vip2Ac proteins in Bacillus thuringiensis. World J. Microbiol. Biotechnol. 2006, 23, 501–507. [Google Scholar] [CrossRef]
- Sattar, S.; Maiti, M.K. Molecular characterization of a novel vegetative insecticidal protein from Bacillus thuringiensis effective against sap-sucking insect pest. J. Microbiol. Biotechnol. 2011, 21, 937–946. [Google Scholar] [CrossRef]
- Gatehouse, J.A. Biotechnological prospects for engineering insect-resistant plants. Plant Physiol. 2008, 146, 881–887. [Google Scholar] [CrossRef] [PubMed]
- Barth, H.; Aktories, K.; Popoff, M.R.; Stiles, B.G. Binary bacterial toxins: Biochemistry, biology, and applications of common Clostridium and Bacillus proteins. Microbiol. Mol. Biol. Rev. 2004, 68, 373–402. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Craig, J.A.; Putnam, C.D.; Carozzi, N.B.; Tainer, J.A. Evolution and mechanism from structures of an ADP-ribosylating toxin and NAD complex. Nat. Struct. Biol. 1999, 6, 932–936. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Xu, W.; Yuan, M.; Tang, M.; Chen, J.; Pang, Y. Expression of vip1/vip2 genes in Escherichia coli and Bacillus thuringiensis and the analysis of their signal peptides. J. Appl. Microbiol. 2004, 97, 757–765. [Google Scholar] [CrossRef] [PubMed]
- MacIntosh, S.C.; Stone, T.B.; Sims, S.R.; Hunst, P.L.; Greenplate, J.T.; Marrone, P.G.; Perlak, F.J.; Fischhoff, D.A.; Fuchs, R.L. Specificity and efficacy of purified Bacillus thuringiensis proteins against agronomically important insects. J. Invertebr. Pathol. 1990, 56, 258–266. [Google Scholar] [CrossRef] [PubMed]
- Rang, C.; Gil, P.; Neisner, N.; van Rie, J.; Frutos, R. Novel Vip3-related protein from Bacillus thuringiensis. Appl. Environ. Microbiol. 2005, 71, 6276–6281. [Google Scholar]
- Palma, L.; Hernández-Rodríguez, C.S.; Maeztu, M.; Hernández-Martínez, P.; Ruiz de Escudero, I.; Escriche, B.; Muñoz, D.; van Rie, J.; Ferré, J.; Caballero, P.; et al. Vip3C, a novel class of vegetative insecticidal proteins from Bacillus thuringiensis. Appl. Environ. Microbiol. 2012, 78, 7163–7165. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Xu, N.; Huang, X.; Wang, W.; Cheng, J.; Wu, K.; Shen, Z. Bacillus thuringiensis Vip3 mutant proteins: Insecticidal activity and trypsin sensitivity. Biocontrol Sci. Technol. 2007, 17, 699–708. [Google Scholar] [CrossRef]
- Estruch, J.J.; Yu, C.G. Plant Pest Control. Patent WO 9,844,137, 17 December 1998. [Google Scholar]
- Bommireddy, P.L.; Leonard, B.R.; Emfinger, K. Heliothine larval behavior on transgenic cotton expressing a Bacillus thuringiensis insecticidal exotoxin, Vip3A. J. Cotton Sci. 2007, 11, 199–207. [Google Scholar]
- Palma, L.; Berry, C.; Caballero, P.; Universidad Pública de Navarra (Spain and Cardiff University) (United Kingdom), Pamplona, Spain. Unpublished Work. 2014.
- Lee, M.K.; Walters, F.S.; Hart, H.; Palekar, N.; Chen, J.S. The mode of action of the Bacillus thuringiensis vegetative insecticidal protein Vip3A differs from that of Cry1Ab delta-endotoxin. Appl. Environ. Microbiol. 2003, 69, 4648–4657. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.G.; Mullins, M.A.; Warren, G.W.; Koziel, M.G.; Estruch, J.J. The Bacillus thuringiensis vegetative insecticidal protein Vip3A lyses midgut epithelium cells of susceptible insects. Appl. Environ. Microbiol. 1997, 63, 532–536. [Google Scholar] [PubMed]
- Sena, J.A.D.; Hernández-Rodríguez, C.S.; Ferré, J. Interaction of Bacillus thuringiensis Cry1 and Vip3A proteins with Spodoptera frugiperda midgut binding sites. Appl. Environ. Microbiol. 2009, 75, 2236–2237. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.K.; Miles, P.; Chen, J.-S. Brush border membrane binding properties of Bacillus thuringiensis Vip3A toxin to Heliothis virescens and Helicoverpa zea midguts. Biochem. Biophys. Res. Commun. 2006, 339, 1043–1047. [Google Scholar] [CrossRef] [PubMed]
- Mike, C.; Ryan, J.; Maria, M.; Martin, T.; Dickerson, D.; Negrotto, D.; O’Reilly, D.; Chen, E.; Lee, M. Effective IRM with a novel insecticidal protein, Vip3A. In Proceedings of the Beltwide Cotton Conference, San Antonio, TX, USA, 3–6 January 2006; National Cotton Council: Memphis, TN, USA, 2006; pp. 1229–1235. [Google Scholar]
- Bravo, A.; Likitvivatanavong, S.; Gill, S.S.; Soberón, M. Bacillus thuringiensis: A story of a successful bioinsecticide. Insect Biochem. Mol. Biol. 2011, 41, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Mehlo, L.; Gahakwa, D.; Nghia, P.T.; Loc, N.T.; Capell, T.; Gatehouse, J.A.; Gatehouse, A.M.; Christou, P. An alternative strategy for sustainable pest resistance in genetically enhanced crops. Proc. Natl. Acad. Sci. USA 2005, 102, 7812–7816. [Google Scholar] [CrossRef] [PubMed]
- Palma, L.; Muñoz, D.; Berry, C.; Murillo, J.; Caballero, P. Draft genome sequences of two Bacillus thuringiensis strains and characterization of a putative 41.9-kDa insecticidal toxin. Toxins 2014, 6, 1490–1504. [Google Scholar] [CrossRef] [PubMed]
- Nishiwaki, H.; Nakashima, K.; Ishida, C.; Kawamura, T.; Matsuda, K. Cloning, functional characterization, and mode of action of a novel insecticidal pore-forming toxin, sphaericolysin, produced by Bacillus sphaericus. Appl. Environ. Microbiol. 2007, 73, 3404–3411. [Google Scholar] [CrossRef] [PubMed]
- Silva-Filha, M.H.N.L.; Berry, C.; Regis, L. Lysinibacillus sphaericus: Toxins and mode of action, applications for mosquito control and resistance management. In Advances in Insect Physiology: Insect Midgut and Insecticidal Proteins; Dhadialla, T.S., Gill, S.S., Eds.; Elsevier: Oxford, UK, 2014; Volume 47, pp. 89–176. [Google Scholar]
- Liu, X.Y.; Ruan, L.F.; Hu, Z.F.; Peng, D.H.; Cao, S.Y.; Yu, Z.N.; Liu, Y.; Zheng, J.S.; Sun, M. Genome-wide screening reveals the genetic determinants of an antibiotic insecticide in Bacillus thuringiensis. J. Biol. Chem. 2010, 285, 39191–39200. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Ruan, L.; Peng, D.; Li, L.; Sun, M.; Yu, Z. Thuringiensin: A Thermostable Secondary Metabolite from Bacillus thuringiensis with Insecticidal Activity against a Wide Range of Insects. Toxins 2014, 6, 2229–2238. [Google Scholar] [CrossRef] [PubMed]
- Levinson, B.L.; Kasyan, K.J.; Chiu, S.S.; Currier, T.C.; Gonzalez, J.M., Jr. Identification of beta-exotoxin production, plasmids encoding beta-exotoxin, and a new exotoxin in Bacillus thuringiensis by using high-performance liquid chromatography. J. Bacteriol. 1990, 172, 3172–3179. [Google Scholar] [PubMed]
- Gohar, M.; Perchat, S. Sample preparation for beta-exotoxin determination in Bacillus thuringiensis cultures by reversed-phase high-performance liquid chromatography. Anal. Biochem. 2001, 298, 112–117. [Google Scholar] [CrossRef] [PubMed]
- McClintock, J.; Stone, T.B.; Sjoblad, R.D. A comparative review of the mammalian toxicity of Bacillus thuringiensis-based pesticides. Pest Manag. Sci. 1995, 45, 95–105. [Google Scholar] [CrossRef]
- Hernandez, C.S.; Martinez, C.; Porcar, M.; Caballero, P.; Ferre, J. Correlation between serovars of Bacillus thuringiensis and type I beta-exotoxin production. J. Invertebr. Pathol. 2003, 82, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Glare, T.R.; O’Callaghan, M. Bacillus Thuringiensis: Biology, Ecology and Safety; Wiley: Chichester, UK, 2000. [Google Scholar]
- Fang, S.L.; Wang, L.; Guo, W.; Zhang, X.; Peng, D.H.; Luo, C.P.; Yu, Z.I.; Sun, M. Bacillus thuringiensis Bel protein enhances the toxicity of Cry1Ac protein to Helicoverpa armigera larvae by degrading insect intestinal mucin. Appl. Environ. Microbiol. 2009, 75, 5237–5243. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Guo, W.; Sun, W.; Xu, D.; Liu, D. Identification of a novel enhancin-like gene from Bacillus thuringiensis. Front. Agric. China 2011, 5, 423–429. [Google Scholar] [CrossRef]
- Agaisse, H.; Lereclus, D. How does Bacillus thuringiensis produce so much insecticidal crystal protein? J. Bacteriol. 1995, 177, 6027–6032. [Google Scholar] [PubMed]
- Ibrahim, M.A.; Griko, N.; Junker, M.; Bulla, L.A. Bacillus thuringiensis: A genomics and proteomics perspective. Bioeng. Bugs 2010, 1, 31–50. [Google Scholar] [CrossRef] [PubMed]
- Shao, Z.; Liu, Z.; Yu, Z. Effects of the 20-kilodalton helper protein on Cry1Ac production and spore formation in Bacillus thuringiensis. Appl. Environ. Microbiol. 2001, 67, 5362–5369. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Nagai, M.; Bagdasarian, M.; Smith, T.W.; Walker, E.D. Expression of the p20 gene from Bacillus thuringiensis H-14 increases Cry11A toxin production and enhances mosquito-larvicidal activity in recombinant gram-negative bacteria. Appl. Environ. Microbiol. 2001, 67, 3010–3015. [Google Scholar] [CrossRef]
- Wu, D.; Federici, B.A. A 20-kilodalton protein preserves cell viability and promotes CytA crystal formation during sporulation in Bacillus thuringiensis. J. Bacteriol. 1993, 175, 5276–5280. [Google Scholar] [PubMed]
- Nisnevitch, M.; Cohen, S.; Ben-Dov, E.; Zaritsky, A.; Sofer, Y.; Cahan, R. Cyt2Ba of Bacillus thuringiensis israelensis: Activation by putative endogenous protease. Biochem. Biophys. Res. Commun. 2006, 344, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Douek, J.; Einav, M.; Zaritsky, A. Sensitivity to plating of Escherichia coli cells expressing the cryA gene from Bacillus thuringiensis var. israelensis. Mol. Gen. Genet. 1992, 232, 162–165. [Google Scholar] [CrossRef]
- Peña, G.; Aguilar Jiménez, F.A.; Hallal-Calleros, C.; Morales-Montor, J.; Hernández-Velázquez, V.M.; Flores-Pérez, F.I. In vitro ovicidal and cestocidal efects of toxins from Bacillus thuringiensis on the canine and human parasite Dipylidium caninum. BioMed Res. Int. 2013, 2013, 174619. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Pattemore, J.; Ash, G.; Williams, A.; Hane, J. Draft genome sequence of Bacillus thuringiensis strain DAR 81934, which exhibits molluscicidal activity. J. Bacteriol. 2013, 1, e00175-12. [Google Scholar]
- Wright, M.K.; Brandt, S.L.; Coudron, T.A.; Wagner, R.M.; Habibi, J.; Backus, E.A.; Huesing, J.E. Characterization of digestive proteolytic activity in Lygus hesperus Knight (Hemiptera: Miridae). J. Insect Physiol. 2006, 52, 717–728. [Google Scholar] [CrossRef] [PubMed]
- Ferré, J.; van Rie, J. Biochemistry and genetics of insect resistance to Bacillus thuringiensis. Annu. Rev. Entomol. 2002, 47, 501–533. [Google Scholar] [CrossRef] [PubMed]
- Loeza-Lara, P.D.; Benintende, G.; Cozzi, J.; Ochoa-Zarzosa, A.; Baizabal-Aguirre, V.M.; Valdez-Alarcón, J.J.; López-Meza, J.E. The plasmid pBMBt1 from Bacillus thuringiensis subsp. darmstadiensis (INTA Mo14–4) replicates by the rolling-circle mechanism and encodes a novel insecticidal crystal protein-like gene. Plasmid 2005, 54, 229–240. [Google Scholar]
- Mesrati, L.A.; Tounsi, S.; Jaoua, S. Characterization of a novel vip3-type gene from Bacillus thuringiensis and evidence of its presence on a large plasmid. FEMS Microbiol. Lett. 2005, 244, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Jensen, G.B.; Andrup, L.; Wilcks, A.; Smidt, L.; Poulsen, O.M. The aggregation-mediated conjugation system of Bacillus thuringiensis subsp. israelensis: Host range and kinetics of transfer. Curr. Microbiol. 1996, 33, 228–236. [Google Scholar]
- Gammon, K.; Jones, G.W.; Hope, S.J.; Oliveira, C.M.F.; Regis, L.; Silva-Filha, M.H.; Dancer, B.M.; Berry, C. Conjugal transfer of a toxin-coding megaplasmid from Bacillus thuringiensis subsp. israelensis to mosquitocidal strains of Bacillus sphaericus. Appl. Environ. Microbiol. 2006, 73, 1766–1770. [Google Scholar] [CrossRef]
- Jarrett, P.; Stephenson, M. Plasmid Transfer between Strains of Bacillus thuringiensis Infecting Galleria mellonella and Spodoptera littoralis. Appl. Environ. Microbiol. 1990, 56, 1608–1614. [Google Scholar] [PubMed]
- Thomas, D.J.I.; Morgan, J.A.W.; Whipps, J.M.; Saunders, J.R. Plasmid transfer between the Bacillus thuringiensis subspecies kurstaki and tenebrionis in laboratory culture and soil and in lepidopteran and coleopteran larvae. Appl. Environ. Microbiol. 2000, 66, 118–124. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.J.I.; Morgan, J.A.W.; Whipps, J.M.; Saunders, J.R. Plasmid transfer between Bacillus thuringiensis subsp. israelensis strains in laboratory culture, river water, and dipteran larvae. Appl. Environ. Microbiol. 2001, 67, 330–338. [Google Scholar]
- González, J.M.; Brown, B.J.; Carlton, B.C. Transfer of Bacillus thuringiensis plasmids coding for delta-endotoxin among strains of B. thuringiensis and B. cereus. Proc. Natl. Acad. Sci. USA 1982, 79, 6951–6955. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Hodgman, T.C.; Krieger, L.; Schnetter, W.; Schairer, H.U. Cloning and analysis of the first cry gene from Bacillus popilliae. J. Bacteriol. 1997, 179, 4336–4341. [Google Scholar] [PubMed]
- Yokoyama, T.; Tanaka, M.; Hasegawa, M. Novel cry gene from Paenibacillus lentimorbus strain semadara inhibits ingestion and promotes insecticidal activity in Anomala cuprea larvae. J. Invertebr. Pathol. 2004, 85, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Barloy, F.; Delécluse, A.; Nicolas, L.; Lecadet, M.-M. Cloning and expression of the first anaerobic toxin gene from Clostridium bifermentans subsp. malaysia, encoding a new mosquitocidal protein with homologies to Bacillus thuringiensis delta-endotoxins. J. Bacteriol. 1996, 178, 3099–3105. [Google Scholar]
- Rigden, D.J. Does distant homology with Evf reveal a lipid binding site in Bacillus thuringiensis cytolytic toxins? FEBS Lett. 2009, 583, 1555–1560. [Google Scholar] [CrossRef] [PubMed]
- Costechareyre, D.; Dridi, B.; Rahbe, Y.; Condemine, G. Cyt toxin expression reveals an inverse regulation of insect and plant virulence factors of Dickeya dadantii. Environ. Microbiol. 2010, 12, 3290–3301. [Google Scholar] [CrossRef]
- Palma, L.; Muñoz, D.; Murillo, J.; Caballero, P. Draft Genome Sequence of Bacillus thuringiensis Serovar Tolworthi Strain Na205-3, an Isolate Toxic for Helicoverpa armigera. Genome Announc. 2014, 6, 1490–1504. [Google Scholar]
- Murawska, E.; Fiedoruk, K.; Swiecicka, I. Modular genetic architecture of the toxigenic plasmid pIS56–63 harboring cry1Ab21 in Bacillus thuringiensis subsp. thuringiensis strain IS5056. Pol. J. Microbiol. 2014, 63, 147–156. [Google Scholar]
- Wu, J.; Zhao, F.; Bai, J.; Deng, G.; Qin, S.; Bao, Q. Adaptive evolution of cry Genes in Bacillus thuringiensis: Implications for their specificity determination. Genomics Proteomics Bioinform. 2007, 5, 102–110. [Google Scholar] [CrossRef]
- Wu, J.; Zhao, F.; Bai, J.; Deng, G.; Qin, S.; Bao, Q. Evidence for positive Darwinian selection of vip gene in Bacillus thuringiensis. J. Genet. Genomics 2007, 34, 649–660. [Google Scholar]
- Chungjatupornchai, W. Expression of the Mosquitocidal-Protein Genes of Bacillus thuringiensis Subsp israelensis and the Herbicide-Resistance Gene Bar in Synechocystis Pcc6803. Curr. Microbiol. 1990, 21, 283–288. [Google Scholar] [CrossRef]
- Bravo, A. Phylogenetic relationships of Bacillus thuringiensis delta-endotoxin family proteins and their functional domains. J. Bacteriol. 1997, 179, 2793–2801. [Google Scholar] [PubMed]
- González, J.M., Jr.; Carlton, B.C. A large transmissible plasmid is required for crystal toxin production in Bacillus thuringiensis variety israelensis. Plasmid 1984, 11, 28–38. [Google Scholar] [CrossRef] [PubMed]
- Zghal, R.Z.; Jaoua, S. Evidence of DNA rearrangements in the 128-kilobase pBtoxis plasmid of Bacillus thuringiensis israelensis. Mol. Biotechnol. 2006, 33, 191–198. [Google Scholar] [PubMed]
- Doggett, N.A.; Stubben, C.J.; Chertkov, O.; Bruce, D.C.; Detter, J.C.; Johnson, S.L.; Han, C.S. Complete genome sequence of Bacillus thuringiensis serovar israelensis strain HD-789. Genome Announc. 2013, 1, e01023-13. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Fan, W.; Han, B.; Liu, H.; Zheng, D.; Li, Q.; Dong, W.; Yan, J.; Gao, M.; Berry, C.; et al. Complete genome sequences of the mosquitocidal bacterium Bacillus sphaericus C3-41 and comparisons with closely related Bacillus species. J. Bacteriol. 2008, 190, 2892–2902. [Google Scholar] [CrossRef] [PubMed]
- Poncet, S.; Bernard, C.; Dervyn, E.; Caylet, J.; Klier, A.; Rapoport, G. Improvement of Bacillus sphaericus toxicity against dipteran larvae by integration, via homologous recombination, of the Cry11A toxin gene from Bacillus thuringiensis subsp. israelensis. Appl. Environ. Microbiol. 1997, 63, 4413–4420. [Google Scholar]
- Ben-Dov, E.; Zaritsky, A.; Dahan, E.; Barak, D.; Sinai, R.; Manasherob, R.; Khamraev, A.; Triotskaya, E.; Dubitsky, A.; Berezina, N.; et al. Extended screening by PCR for seven cry-group genes from field-collected strains of Bacillus thuringiensis. Appl. Environ. Microbiol. 1997, 63, 4883–4890. [Google Scholar]
- Wang, J.; Boets, A.; van Rie, J.; Ren, G. Characterization of cry1, cry2, and cry9 genes in Bacillus thuringiensis isolates from China. J. Invertebr. Pathol. 2003, 82, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Granero, F.; Ballester, V.; Ferre, J. Bacillus thuringiensis crystal proteins Cry1Ab and Cry1Fa share a high affinity binding site in Plutella xylostella (L.). Biochem. Biophys. Res. Commum. 1996, 224, 779–783. [Google Scholar] [CrossRef]
- Li, H.; Gonzalez-Cabrera, J.; Oppert, B.; Ferre, J.; Higgins, R.A.; Buschman, L.L.; Radke, G.A.; Zhu, K.Y.; Huang, F. Binding analyses of Cry1Ab and Cry1Ac with membrane vesicles from Bacillus thuringiensis-resistant and -susceptible Ostrinia nubilalis. Biochem. Biophys. Res. Commun. 2004, 323, 52–57. [Google Scholar] [CrossRef] [PubMed]
- Hughes, P.A.; Stevens, M.M.; Park, H.-W.; Federici, B.A.; Dennis, E.S.; Akhurst, R. Response of larval Chironomus tepperi (Diptera: Chironomidae) to individual Bacillus thuringiensis var. israelensis toxins and toxin mixtures. J. Invertebr. Pathol. 2005, 88, 34–39. [Google Scholar]
- Lee, M.K.; Curtiss, A.; Alcantara, E.; Dean, D.H. Synergistic effect of the Bacillus thuringiensis toxins CryIAa and CryIAc on the gypsy moth, Lymantria dispar. Appl. Environ. Microbiol. 1996, 62, 583–586. [Google Scholar] [PubMed]
- Hernández-Rodríguez, C.S.; Boets, A.; van Rie, J.; Ferré, J. Screening and identification of vip genes in Bacillus thuringiensis strains. J. Appl. Microbiol. 2009, 107, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Kuo, W.S.; Chak, K.F. Identification of novel cry-type genes from Bacillus thuringiensis strains on the basis of restriction fragment length polymorphism of the PCR-amplified DNA. Appl. Environ. Microbiol. 1996, 62, 1369–1377. [Google Scholar] [PubMed]
- Liu, J.; Song, F.; Zhang, J.; Liu, R.; He, K.; Tan, J.; Huang, D. Identification of vip3A-type genes from Bacillus thuringiensis strains and characterization of a novel vip3A-type gene. Lett. Appl. Microbiol. 2007, 45, 432–438. [Google Scholar] [CrossRef] [PubMed]
- Palma, L.; Ruiz de Escudero, I.; Maeztu, M.; Caballero, P.; Muñoz, D. Screening of vip genes from a Spanish Bacillus thuringiensis collection and characterization of two novel Vip3 proteins highly-toxic to five lepidopteran crop pests. Biol. Control 2013, 66, 141–149. [Google Scholar] [CrossRef]
- Porcar, M.; Juárez-Pérez, V. PCR-based identification of Bacillus thuringiensis pesticidal crystal genes. FEMS Microbiol. Rev. 2003, 26, 419–432. [Google Scholar] [PubMed]
- Tan, F.R.; Zhu, J.; Tang, J.; Tang, X.M.; Wang, S.Q.; Zheng, A.P.; Li, P. Cloning and characterization of two novel crystal protein genes, cry54Aa1 and cry30Fa1, from Bacillus thuringiensis strain BtMC28. Curr. Microbiol. 2009, 58, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Oppert, B.; Kramer, K.J.; Beeman, R.W.; Johnson, D.; McGaughey, W.H. Proteinase-mediated insect resistance to Bacillus thuringiensis toxins. J. Biol. Chem. 1997, 272, 23473–23476. [Google Scholar] [CrossRef] [PubMed]
- Sayyed, A.H.; Haward, R.; Herrero, S.; Ferre, J.; Wright, D.J. Genetic and biochemical approach for characterization of resistance to Bacillus thuringiensis toxin Cry1Ac in a field population of the diamondback moth, Plutella xylostella. Appl. Environ. Microbiol. 2000, 66, 1509–1516. [Google Scholar] [CrossRef] [PubMed]
- Mardis, E.R. Next-generation DNA sequencing methods. Annu. Rev. Genomics Hum. Genet. 2008, 9, 387–402. [Google Scholar] [CrossRef] [PubMed]
- Metzker, M.L. Sequencing technologies—The next generation. Nat. Rev. Genet. 2010, 11, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Schuster, S.C. Next-generation sequencing transforms today’s biology. Nat. Methods 2008, 5, 16–18. [Google Scholar] [CrossRef] [PubMed]
- Shendure, J.; Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 2008, 26, 1135–1145. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Ren, L.; Meng, Q.; Li, Y.; Yu, Y.; Yu, J. The next-generation sequencing technology and application. Protein Cell 2010, 1, 520–536. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, D.; Francois, P.; Farinelli, L.; Osteras, M.; Schrenzel, J. De novo bacterial genome sequencing: Millions of very short reads assembled on a desktop computer. Genome Res. 2008, 18, 802–809. [Google Scholar] [CrossRef] [PubMed]
- Krampis, K.; Booth, T.; Chapman, B.; Tiwari, B.; Bicak, M.; Field, D.; Nelson, K.E. Cloud BioLinux: Pre-configured and on-demand bioinformatics computing for the genomics community. BMC Bioinform. 2012, 13, 42. [Google Scholar] [CrossRef]
- Zhang, W.Y.; Chen, J.J.; Yang, Y.; Tang, Y.F.; Shang, J.; Shen, B.R. A practical comparison of de novo genome assembly software tools for next-generation sequencing technologies. PLoS One 2011, 6, e17915. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Palma, L.; Muñoz, D.; Berry, C.; Murillo, J.; Ruiz de Escudero, I.; Caballero, P. Molecular and Insecticidal Characterization of a Novel Cry-Related Protein from Bacillus Thuringiensis Toxic against Myzus persicae. Toxins 2014, 6, 3144–3156. [Google Scholar] [CrossRef] [PubMed]
- Ye, W.; Zhu, L.; Liu, Y.; Crickmore, N.; Peng, D.; Ruan, L.; Sun, M. Mining new crystal protein genes from Bacillus thuringiensis based on mixed plasmid-enriched genome sequencing and a computational pipeline. Appl. Environ. Microbiol. 2012, 78, 4795–4801. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Palma, L.; Muñoz, D.; Berry, C.; Murillo, J.; Caballero, P. Bacillus thuringiensis Toxins: An Overview of Their Biocidal Activity. Toxins 2014, 6, 3296-3325. https://doi.org/10.3390/toxins6123296
Palma L, Muñoz D, Berry C, Murillo J, Caballero P. Bacillus thuringiensis Toxins: An Overview of Their Biocidal Activity. Toxins. 2014; 6(12):3296-3325. https://doi.org/10.3390/toxins6123296
Chicago/Turabian StylePalma, Leopoldo, Delia Muñoz, Colin Berry, Jesús Murillo, and Primitivo Caballero. 2014. "Bacillus thuringiensis Toxins: An Overview of Their Biocidal Activity" Toxins 6, no. 12: 3296-3325. https://doi.org/10.3390/toxins6123296
APA StylePalma, L., Muñoz, D., Berry, C., Murillo, J., & Caballero, P. (2014). Bacillus thuringiensis Toxins: An Overview of Their Biocidal Activity. Toxins, 6(12), 3296-3325. https://doi.org/10.3390/toxins6123296